Abstract
The proper functioning of extracytoplasmic proteins requires their export to, and productive folding in, the correct cellular compartment. All proteins in Escherichia coli are initially synthesized in the cytoplasm, then follow a pathway that depends upon their ultimate cellular destination. Many proteins destined for the periplasm are synthesized as precursors carrying an N-terminal signal sequence that directs them to the general secretion machinery at the inner membrane. After translocation and signal sequence cleavage, the newly exported mature proteins are folded and assembled in the periplasm. Maintaining quality control over these processes depends on chaperones, folding catalysts, and proteases. This article summarizes the general principles which control protein folding in the bacterial periplasm by focusing on the periplasmic maltose-binding protein.
Review
Although all the information necessary for a protein to reach a native structure is contained in its amino acid sequence, in vivo protein folding requires the participation of protein factors; molecular chaperones, folding catalysts and proteases, that monitor or regulate this process through many cellular functions. Under physiological conditions, these factors maintain quality control of protein biosynthesis, and errors or failures of the protein folding process are rare. However, incorrectly folded or misfolded proteins can appear as a result of (i) spontaneous or inducible mutations that affect protein folding pathways, (ii) exposure of the cell to environmental stress, such as elevated temperatures or hyperosmolarity, and (iii) overexpression of recombinant genes. In these cases, the polypeptide chain, instead of folding into the native biologically active state, misfolds, and eventually induces a stress response [1]. The resulting misfolded proteins may either be degraded by proteases, repaired by chaperones, or aggregated and sequestered as inclusion bodies (IBs), when escaping protein folding quality control [2]. In the cytoplasm of Escherichia coli, this control is performed mainly by a set of stress-inducible chaperones and proteases collectively known as heat shock proteins (Hsps). The presence of cellular membrane-bounded compartments in E. coli further complicates these issues of protein biosynthesis, and implies the existence of distinct folding and degradation machineries, quality control systems, for extracytoplasmic proteins [3]. The bacterial periplasm is separated from the extracellular milieu only by the porous outer membrane, and hence is more susceptible to changes in the environment than the cytoplasm. Thus, protein quality control in the periplasm is crucial for the survival of bacterial cells. Here we summarize the main features of protein folding in the periplasm of E. coli, and focus on studies using a model system, the maltose-binding protein.
Biosynthesis of periplasmic proteins
In E. coli, exported proteins with an ultimate destination of the periplasm and outer membrane are synthesized as precursors with a cleavable amino-terminal signal sequence. Depending on the nature of the precursors, different pathways exist for their transport across the inner membrane. Although the translocation of most exported proteins occurs by the general secretory (Sec) pathway [4], across a channel formed by the heterotrimeric membrane complex SecYEG, a subset of precursors, containing two consecutive arginine residues near the N-terminus of the signal sequences, utilizes the twin-arginine translocation (Tat) pathway. The Tat machinery, encoded by the tatABCE genes, exports precursors that must completely fold in the cytoplasm to bind cofactors [5].
Targeting of precursors to the SecYEG translocase
In marked contrast, the Sec pathway transports proteins which are almost unstructured, as it was confirmed by the recently solved crystal structure of the SecY complex from Methanococcus janansahii [6]. Since SecYEG forms a passive channel, it must be associated with other components to provide the driving force for translocating preprotein substrates. According to their degree of coupling to translation, two alternative modes of protein translocation have been identified. In cotranslational translocation, as the signal sequence emerges from a translating ribosome, it binds to the signal recognition particle (SRP), and, in concert with its membrane-bound receptor FtsY, the nascent polypeptide is targeted to SecYEG. The energy for translocation comes from GTP hydrolysis during translation. In posttranslational translocation, many newly synthesized precursors interact with the chaperone SecB (see below). Then, both SecB and preproteins provide binding sites for SecA, a peripheral membrane bound ATPase, that targets these initial complexes to the translocase. By mediating repeated cycles of ATP binding and hydrolysis, SecA pushes the polypeptide chain through the SecYEG channel [7]. The combined data available suggest that the SRP-dependent pathway is probably mainly involved in the cotranslational assembly of cytoplasmic membrane proteins [8,9], and that most periplasmic and outer membrane proteins are targeted posttranslationally to the SecA/SecB pathway [9]. However, recent studies indicate that some extracytoplasmic proteins exhibit a combined dependence on SRP/FtsY and SecA [10,11]. Even though protein translocation in E. coli has been well studied, the molecular mechanisms which allow the discrimination between both pathways converging at the SecYEG translocase remain unknown.
When a nascent chain emerges from the ribosome, a range of folding and targeting factors are waiting to bind the precursor. The first folding helper encountered is probably the trigger factor [12], a ribosome-bound peptidyl-prolyl isomerase [13] which displays chaperone activity in vitro. In addition to trigger factor, the emerging signal sequence is accessible to SRP and SecA [14]. These initial contacts are then modified upon further growth of the nascent chain and binding to SecYEG. In the Sec pathway, during elongation or after release from the ribosome, many, but not all, polypeptide chains bind to SecB, a tetrameric chaperone which stabilizes the precursor in a translocation-competent conformation [15]. The SecB-bound precursors, in cooperation with SecA, are then targeted to SecYEG. Although other cytoplasmic chaperones, like DnaK (Hsp70) and GroEL (Hsp60), may participate in protein export by preventing early folding of precursors [16,17], these general chaperones have a marginal role in targeting precursors to the Sec translocase. However, when translocation is impaired, by slowly exported proteins or defective signal peptides for example, precursors could accumulate in the cytoplasm. Structurally, a precursor in an export-competent conformation may be similar to an unfolded protein. Indeed, Wild et al. [17] showed that accumulation of precursors in strains lacking SecB induces a stress response mediated by the specific sigma factor, σ32. Under normal conditions, DnaK and DnaJ sequester σ32 through direct binding, inhibiting its association with RNA polymerase [18]. Accumulation of unfolded precursors displaces this complex and allows σ32 binding to RNA polymerase to direct transcription of heat shock genes. It has been suggested that the final distribution of the precursors among the different pathways is determined by a kinetic partitioning that is dependent on the rate of folding, relative to the rate of chaperone binding [19]. Finally, how these different factors compete will determine whether initiation of translocation occurs efficiently (Figure 1).
Figure 1.
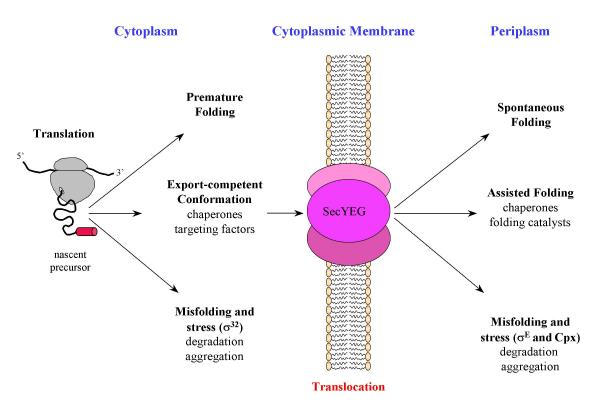
Alternative folding pathways for periplasmic protein in E. coli. This scheme illustrates the present discussion and tries to emphasize the alternative and competing process between folding and misfolding of periplasmic proteins before and after translocation across the cytoplasmic membrane.
In the next step, the precursor inserts as a loop into the SecYEG channel, with its signal sequence intercalated between two transmembrane helices of SecY, and with its mature region in the pore. The polypeptide chain is then transported through the pore, and the signal sequence is cleaved at some point during translocation by the signal peptidase, a membrane-bound endopeptidase with its catalytic domain facing the periplasm [20].
Protein folding in the periplasm
Once they reach the periplasm and after signal sequence cleavage, periplasmic proteins may encounter two types of protein folding catalysts: protein disulphide isomerases (Dsb proteins), which catalyse disulfide-bond formation, and peptidyl-prolyl isomerases (PPIase), which catalyse cis-trans isomerization of peptidyl bonds. These enzymes are known to catalyze rate-limiting steps in the in vitro protein folding process.
The formation of disulfide bonds is essential for the oxidative folding, activity, and stability of many proteins exported from the cytoplasm. Disulfide bonds formation results from elaborate electron transfer pathways between the Dsb oxydoreductases [21]. DsbA is the strongest thiol oxidant which catalyses oxidation of cysteines of folding proteins. This reaction occurs via formation of mixed disulfide complexes between DsbA and its substrates. DsbB, a membrane-bound enzyme, maintains DsbA oxidized by transferring electrons from reduced DsbA to quinones in the respiratory chain [22]. It has become clear that disulfide bond formation, far from being perfect, occurs between incorrect pairs of cysteines resulting in the presence of non-native disulfides in proteins. As periplasmic thiol isomerases, DsbC and DsbG promote the rearrangements of incorrect disulfide bonds. The maintenance of these thiol isomerases in the reduced state depends on the inner membrane protein DsbD, which utilizes the thioredoxin/thioredoxin reductase system in the cytoplasm as a source of reducing equivalents [23]. Recently, covalent intermediates between DsbA and exported substrate proteins were detected in vivo by using a DsbA variant carrying the Pro151 → Thr substitution in its active site [24]. This study opens the way to understand at what point during protein translocation and folding, disulfide bond formation takes place in the periplasm.
Representatives of the three unrelated families of PPIases have been identified in the periplasm: PpiA (also known as CypA or RotA) is a cyclophilin that is not inhibited by cyclosporin A [25]; FkpA is related to the FK506-binding protein family, or FKBP, [26]; and PpiD [27] and SurA [28], two parvulin homologs. Although all PPIases catalyze the rate-limiting step in the refolding of RNase T1 [29]in vitro, their cellular role remains enigmatic. With the exception of surA, the absence of a significant phenotype for null mutants indicates that these periplasmic PPIases are not essential for viability or have overlapping functions.
Aside from specific chaperones, such as PapD involved in pilus assembly [30], few periplasmic chaperones have been identified, and there are no classical Hsp chaperones, such as DnaK or GroEL, in this compartment. Indeed, as the periplasm lacks ATP, periplasmic chaperones must be mechanistically distinct from their cytoplasmic counterparts, most of which use ATP to drive their cycles of substrate binding and release. However, global searches using genetic selections based on σE activity (see below) have identified the surA and skp/ompH genes [28,31], which are involved in the folding and assembly of outer membrane proteins [32]. Furthermore, in vitro chaperone activity of several periplasmic proteins such as DsbC [33] and DsbG [34] or substrate-binding proteins [35,36] have already been reported, but their contribution has not yet been studied in vivo. It should be noted that studying the in vivo function of a putative chaperone is more difficult than assessing its activity in vitro, using a classical chaperone substrate. However, it is tempting to speculate from these observations that, because of their unique active sites, protein folding catalysts can perform chaperone functions in order to compensate for the apparent deficiency of general chaperones in the periplasm. Indeed, FkpA and SurA exhibit PPIase-independent chaperone activity in vivo [37,38]. Although these two PPIases are unrelated, their crystal structures share a similar organization: a chaperone activity residing in the N-terminal domains and the C-terminal catalytic domains tethered ~30Å apart, connected either by a long α helix [39], or an extended polypeptide [40]. In other foldases that exhibit both chaperone and protein folding catalytic activities, these two functions have also been assigned to distinct regions or domains of the molecule. For instance, the trigger factor comprises three domains, with the central FKBP-like domain displaying PPIase activity and the C-terminal domain being implicated in chaperone activity [41]. The structures of DsbC and FkpA present also further striking similarities. Both enzymes are homodimeric, with subunits divided into two domains where the N-domains form the dimeric interfaces. The C-domains, carrying the catalytic sites, are connected to the N-domains by a pliable helix, conferring a V-shaped form on the dimers, and orienting the two catalytic sites face inwards towards each other. The cleft formed in the DsbC dimer by association of the N-domains is hydrophobic in character, and it was proposed that this region forms the binding site of the unfolded polypeptide for the chaperone activity, sequestering the substrate from aggregation and, at the same time, giving it access to the catalytic sites to facilitate disulfide bond exchange [42]. In FkpA, the cleft formed by association of the N-domains has approximately the same dimensions to that formed in DsbC. In contrast to the hydrophobic nature of the DsbC cleft, that of FkpA has an intermediate negative potential, reflecting probably a difference in their substrate specificity. However, it is an attractive idea that the V-shaped form of these foldases cradles the unfolded polypeptide substrates while giving access to the isomerase catalytic sites. The innate flexibility of the C-domains would allow for adaptability between the chaperone and isomerase activities.
An alternative fate for misfolded periplasmic proteins is degradation by proteases (Figure 1). Although more than 10 periplasmic proteases have been identified in E. coli, DegP (also known as HtrA or Do) is the only protease identified as a heat shock protein involved in the degradation of misfolded proteins [43,44]. Indeed, degP, which is essential for the survival of bacteria above 40°C, is under the transcriptional regulation of the alternative heat shock sigma factor, σE [45]. The reason for this essential role is not clear, but presumably an increased number of misfolded proteins accumulate under these conditions. DegP is a widely conserved serine protease composed of an N-terminal domain believed to have regulatory functions, a trypsin-like protease domain, and two PDZ domains [46]. PDZ domains are protein modules that mediate specific protein-protein interactions. The crystal structure of DegP reveals a staggered association of two trimeric rings forming a hexamer [47]. The proteolytic sites are located in a central cavity that is accessible only laterally from the PDZ domains, which probably recognize and bind substrates. It has been shown that the activity of DegP could switch between chaperone and protease activities in a temperature-dependent manner [48]. At temperatures below 28°C, DegP acts as a chaperone, protecting misfolded proteins from irreversible aggregation, and above 28°C, its protease activity increases dramatically degrading misfolded proteins. However, in vivo DegP which does not act as a general chaperone in the periplasm [49], is the major proteolytic machine responsible for periplasmic protein quality control.
Extracytoplasmic stress response
Because optimal cellular growth requires that the cell be able to sense and respond to changes in subcellular compartments, it is not surprising that the stress response in Gram-negative bacteria is compartmentalized into cytoplasmic and extracytoplasmic responses. A common feature of inducers of the E. coli extracytoplasmic stress response is their potential to generate excessive amounts of misfolded proteins in the envelope, particularly outer membrane proteins, and many of the proteins involved in periplasmic protein quality control have been characterized from studies on the extracytoplasmic stress responses.
In contrast to cytoplasmic stress, where the sensing of misfolded proteins and the accompanying response take place in the same compartment, extracytoplasmic stress signals must cross the cytoplasmic membrane by a signal transduction system. E. coli senses and responds to extracytoplasmic stress via at least two overlapping, but distinct, transduction pathways, the Cpx two-component system and the heat shock σE pathway (Figure 2). Both regulatory systems control the expression of several genes whose products are envelope-localized protein folding catalysts, chaperones and proteases, as well as genes involved in lipid and lipopolysaccharide metabolism. Together, these proteins serve to ensure proper biogenesis of the bacterial envelope by preventing any perturbation in periplasmic protein folding [50].
Figure 2.
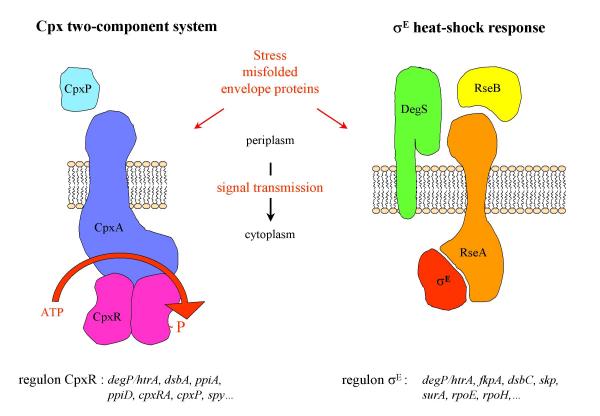
Two signaling pathways for extracytoplasmic stress responses in E. coli. Components of these pathways reside on each side of the cytoplasmic membrane (periplasm and cytoplasm). In the presence of misfolded periplasmic proteins, an unknown mechanism triggers a conformational change of the CpxA that leads to an activation of its kinase activity and promote an increase in CpxR phosphorylation. Under normal physiological conditions, σE is sequestred by RseA and DegS is inactive, with its PDZ domain inhibiting the protease activity. Recognition by the PDZ domain of misfolded periplasmic proteins relieves inhibition of the DegS protease. Activated DegS cleaves RseA, which releases σE. For both signal transduction systems, a partial list of the downstream targets is indicated.
The Cpx pathway is a typical two-component signal transduction system, consisting of the hisitidine kinase CpxA and the response regulator CpxR [51]. The input signal is transduced across the cytoplasmic membrane by the sensor kinase CpxA, which is composed of a periplasmic domain and a conserved cytoplasmic signaling domain separated by two transmembrane α-helices [52]. Upon detection of envelope stress, CpxA autophosphorylates by using ATP at a conserved histidine and then transfers this phosphate to a conserved aspartate in the N-terminal domain of CpxR. Phosphorylated CpxR activates transcription of genes whose products are involved in envelope physiology. The Cpx pathway is induced by elevated pH [53], altered inner membrane composition [54], and overproduction of envelope proteins like the outer membrane lipoprotein NlpE [55] or pili subunits [56,57]. The Cpx regulon is positively autoregulated and is feedback inhibited by CpxP, a small periplasmic protein [58]. The response seems to be modulated by direct interaction of the periplasmic domain of CpxA, probably in concert with CpxP [59]. Some of the targets regulated by CpxR are the heat shock protease DegP [60], the peptidyl prolyl cis/trans isomerases PpiA [61] and PpiD [27], and the disulfide oxidoreductase DsbA [61]. Apart from the extracytoplasmic stress response the Cpx system is implicated in conjugation [62], invasion of host cells [63], and biofilm formation [64,65].
The σE envelope stress response is induced by heat shock [66,67] and by perturbations to outer membrane protein folding [31,68]. Under nonstress conditions, the activity of σE is negatively regulated by the inner membrane anti sigma factor RseA and by the periplasmic protein RseB [69-71]. In response to extracytoplasmic stress, RseA is rapidly degraded by the sequential action of DegS [72] and YaeL/EcfE proteases [73] that frees σE to associate with RNA polymerase and directs the transcription of its regulon. Recently, it was shown that peptides mimicking the C-terminal sequence of outer membrane proteins bind the DegS PDZ domain, activate DegS cleavage of RseA, and induce σE-dependent transcription [74]. These data suggest that DegS acts as a sensor of extracytoplasmic stress by binding unassembled outer membrane proteins, and that DegS activation involves relief of inhibitory interactions between its PDZ and protease domain. However, RseB, which interacts with misfolded periplasmic proteins [71], might influence the accessibility of RseA to cleavage by DegS. At least 20 genes were identified to be regulated by σE [75] which include rpoH, degP [45], yaeL/ecfE [76], and fkpA [77].
Focus on the maltose-binding protein
To study protein folding in the periplasm of E. coli, we use a model system, the maltose-binding protein (MalE or MBP). MalE is the soluble receptor for the high affinity transport of maltose and maltodextrins into the periplasm [78]. Because of its particular function in maltose transport, correct export and folding of MalE in this cellular compartment is essential for cells to utilize maltose as a carbon source [79]. This feature has facilitated the development of powerful genetic selections or of screens which led to the discovery of several genes composing the Sec pathway [80]. MalE has also been extensively used as a model of protein translocation [81]. Translocation of MalE requires that the nascent preMalE polypeptide chain reach a critical size corresponding to 80 % of the final length, and that the precursor exist in an export-competent conformation representing a partially unfolded state [82]. SecB binding mainly participates in the maintenance of this initial conformation. In addition to this chaperone activity, SecB is an important targeting factor for MalE. Indeed, when the signal sequence is deleted, translocation of MalE becomes entirely SecB dependent [83]. Then, MalE is translocated across the SecYEG channel, released into the periplasm after signal sequence cleavage, and the newly exported MalE folds into its native structure. The crystal structure of MalE consists of two discontinuous domains constructed from secondary structural βαβ units surrounding a cleft that forms the binding site for maltose [84]. These properties make MalE an ideal model with which a direct comparison of the mechanism of in vitro and in vivo folding is greatly facilitated.
MalE31, a model for periplasmic protein misfolding
For many years, and well before the resolution of its 3D-structure, the laboratory of M. Hofnung studied the MalE structure-function relationships. One genetic approach, using random insertion of a BamHI oligonucleotide linker, led to the identification of the so-called 'permissive sites' mutants of malE in which insertions or deletions were structurally tolerated without loss of biological properties [85]. In contrast, other random insertions in malE prevented both growth on maltose and periplasmic release of the corresponding proteins [86]. In many cases, a large deletion of several amino acid residues from the mature region of the protein could explain their defective phenotypes. However, in one case, malE31, the linker insertion did not modify the length of the protein and resulted in Gly32 → Asp and Ile33 → Pro substitutions. Structurally, these residues are located in a turn connecting the helix αI to strand βB of the first βαβ element within the N-domain (Figure 3A). The Ile33 → Pro substitution was identified as the major factor responsible for the Mal- phenotype [87]. Indeed, cells expressing a malE variant carrying the Gly32 → Asp substitution displayed biological properties similar to those of cells expressing wild-type malE. The subcellular location of these mutants revealed that MalE31 was entirely recovered in the membrane fraction. This result suggested that export of MalE31 was blocked somewhere in the membrane or, alternatively, that MalE31 formed inclusion bodies (IBs) either in the cytoplasm from the precursor, before translocation, or in the periplasm, from the mature protein, after release from the translocase. It is technically very difficult to rigorously differentiate between these cellular fates by cell-fractionation experiments only. Thus, to determine whether insoluble MalE31 which co-sedimented with membranes was consistent with aggregated proteins rather than membrane-associated proteins, a crude lysate was fractionated by flotation gradient centrifugation, and revealed that MalE31 remained near the bottom [88]. Although cell fractionation from spheroplasts is less sensitive to artifacts than the osmotic shock procedure, both methods cannot discriminate between cellular location (cytoplasmic or periplasmic) of IBs. However, processing of the precursors, a late step of export (see above), is a reliable indicator that the protein has, at least partially, crossed the inner membrane. Pulse-chase experiments showed that the MalE31 precursor was efficiently processed at the same rate as the wild-type protein and that no export interference or jamming was observed upon overexpression of malE31. Finally, transmission electron micrographs of cells overproducing MalE31 revealed the presence of small IBs between the inner and outer membranes (Figure 3B). Based on these data, it became clear that the mature MalE31 protein aggregates in the periplasm. As other aggregation-prone proteins, MalE31 could be purified from IBs, after a urea-solubilization step, and the renaturared protein in vitro exhibited complete maltose-binding activity. Thus, we concluded that the modified αβ turn, which is distant from the binding site, did not perturb the function but rather the periplasmic folding of MalE. Since these initial characterizations, several groups have used MalE31 as a protein model for cellular protein misfolding [71] or aggregation [89].
Figure 3.
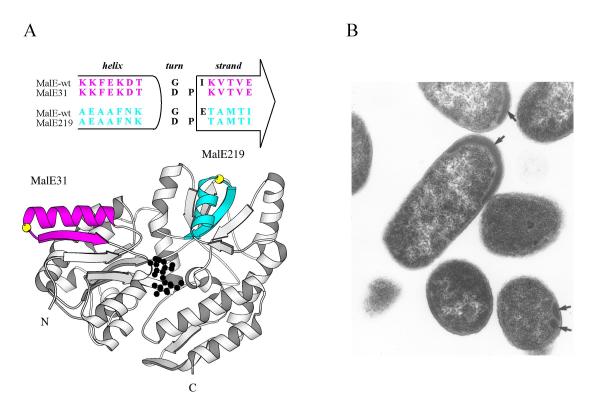
In vitro and in vivo fates for MalE31. A- The maltose-bound state of MalE31. The helix α1 and strand βB of the N-domain (MalE31), colored in magenta, and the helix αVII and strand βJ of the C-domain (MalE219), colored in blue, are located in the crystal structure of MalE. For both turns, the Gly residue is shown in yellow. The bound maltose substrate in black is located in a cleft between both domains. B- Periplasmic inclusion bodies of MalE31. Transmission electron micrograph of ultrathin sections of E. coli overproducing malE31 at 30°C. The black arrows indicate the position of periplasmic inclusion bodies.
Compartmentalized cellular stress responses induced by malE31 mutants
To examine the alternative and competitive export and folding processes in the cytoplasm, we analyzed the cellular fates of MalE31 carrying either a wild-type or defective signal sequences [90]. Since we never observed MalE31 misfolding in the cytoplasm when it was expressed with its wild-type signal sequence, we concluded that the MalE31 precursor enters rapidly into the Sec pathway. The mutational deletion or substitution in signal sequences has been shown to strongly decrease the efficiency of MalE export [91]. The presence of these altered signal sequences in combination with the defective folding malE31 mutation resulted in the cytoplasmic accumulation of precursors. In the steady state, the cytoplasmic soluble fraction of MalE31, when expressed without a signal sequence (69%), is higher than the periplasmic soluble fraction of MalE31, when expressed with a wild-type signal sequence (5%). In the former case, the increased level of cytoplasmic chaperones (Dnak and GroEL), resulting from induction of the σ32 stress response, mainly suppresses the aggregation of MalE31 in the cytoplasm. Although overproduction of MalE31, carrying a wild-type signal sequence, induced an extracytoplasmic stress, the increased level of periplasmic Hsps failed to prevent the aggregation of MalE31. Thus, it appeared that there are insufficient or no general heat shock chaperones in the periplasm. Furthermore, we showed that aggregation of MalE31 in the periplasm did not induce a stress response via σ32 as it did when expressed with defective signal sequences, confirming that the stress responses induced by the presence of misfolded proteins in E. coli are compartmentalized and controlled by different pathways [90]. Technically, the determination of stress responses in E. coli is facilitated by using transcriptional gene fusions and allows the rapid discrimination of cellular location of the protein misfolding.
One physiological consequence for the cells overproducing MalE31 was to induce an extracytoplasmic stress response by increasing the expression of the heat shock protease DegP. Because the Cpx and σE regulatory systems both respond to the presence of misfolded exported proteins, we determined which pathway could be more specifically affected by MalE31. Therefore, by using two different transcriptional fusions which are independently regulated, either by the σE factor (P3rpoH-lacZ) or by CpxR (PcpxP-lacZ), we showed that overproduction of MalE31 induced the Cpx two-component system [49]. Because, the exact nature of inducers that activate this signaling pathway remained unclear, we examined whether the presence of periplasmic inclusion bodies is required. Interestingly, the Cpx system was also induced by the production of the destabilized MalE219 variant, which remains entirely soluble and functional in the periplasm (see below). This observation was further confirmed by producing truncated variants of MalE that did not accumulate at steady states [49]. It seems that the signal sensed by the Cpx pathway will be the presence of periplasmic unfolded or misfolded proteins rather than envelope alterations resulting from the formation of IBs.
Structure-dependence of the malE31 mutation
Despite a synergistic effect, the single Ile33 → Pro substitution was the major factor responsible for MalE31 aggregation. Thus, introduction of a proline residue in the first αβ turn of the N-domain could introduce either a conformational strain in the folded state or reduced conformational freedom of the polypeptide backbone in a critical folding intermediate state, because of the limited value of its dihedral angles. To probe genetically the structural role of this surface turn, we generated a library of random mutations at codons 32 and 33 of malE, and defective folding variants of MalE were screened [88]. Although many combinations of amino acid pairs replaced the native residues at both positions, underlining the passive role of this turn in determining the folded structure of MalE, the majority of critical folding substitutions, contained a proline residue at both positions. A second critical pair of residues involved the association of bulky hydrophobic residues (Phe, Leu, or Val) at position 32 and basic residues (Lys or Arg) at position 33. Close examination of the wild-type MalE crystal structure suggested that the combined negative effects of decreased conformational flexibility and exposed large hydrophobic side-chain residues at position 32, and a buried charged side-chain residue in the hydrophobic position 33, might create these defects. Since this study, the crystal structure of MalE31 was determined at 1.85Å resolution, and provided experimental evidence that MalE31 can attain a wild-type folded conformation in vitro and supports a model in which the structural effect of the malE31 mutation is exerted at the level of folding intermediates, rather than of the native conformation [92].
The secondary and tertiary context-dependence of the malE31 mutation was also assessed by probing the tolerance of an equivalent turn of the C-domain to the same double amino acid substitution. Among the four αβ turns of the MalE structure, we selected the turn connecting the helix αVII to strand βJ of the C-domain, and the corresponding Gly220 → Asp and Glu221 → Pro substitutions were introduced by site-directed mutagenesis [93], defining the MalE219 variant (Figure 3A). In contrast to MalE31, this modified-turn MalE219 variant was correctly folded and functionally active in the bacterial periplasm. Furthermore, in vitro unfolding/refolding experiments showed that both turn MalE variants were destabilized, but their intrinsic tendency to aggregate, correlated well with their periplasmic fates in E. coli, and arose from differences in refolding rates allowing long-term persisting folding intermediates of MalE31, probably kinetically trapped in an off-pathway folding reaction, to aggregate. These data supported the notion that the formation of this βαβ supersecondary structure of the N-domain is a rate-limiting step in the folding pathway of MalE. Indeed, previous intragenic suppressive mutations of export-defective signal peptides of MalE were identified in this structural element, and the analysis of refolding properties of the corresponding variants resulted in the same conclusion [94]. Together, these results argued that the information directing the polypeptide chain to the misfolding pathway is not only determined by a local amino acid sequence (the primary structure), but also by the tertiary structure surrounding the critical MalE31 substitution.
Cellular factors influencing inclusion body formation of MalE31
In a following series of experiments, we manipulated two cellular functions affecting protein folding pathways; the promoter activity of malE31 that modulates the rate of protein biosynthesis and the state of periplasmic DegP protease which determines cellular fates of misfolded MalE31 [95].
When expressed from its own promoter, the cellular level of MalE31 is controlled by the transcriptional activator MalT. Depending on the presence of either the wild type or constitutive MalT variant (MalTC), the production of MalE31 varied from a very low level to a high and constitutive level in the host cells. We took advantage of this regulation to modulate the biosynthetic rate of MalE31, and observed that at a low level of expression, MalE31 was rapidly degraded by the DegP protease, and that at a high level of expression, the misfolded protein formed IBs [95]. Surprisingly, the steaty-state level of soluble MalE31 was only dictated by the ratio of rate constants governing the misfolding and folding steps, and was independent on the promoter activity. The high production level of MalE31 in the malTC context had two important consequences. First, increased amounts of aggregating MalE31 species increases the formation of IBs, the compact nature of which renders them resistant to proteases. Second, although overproduction of MalE31 increased the rate of DegP synthesis via the Cpx system [49], the majority of that increased amount of protease was localized in the insoluble fraction. Because DegP is intimately involved in the degradation of misfolded proteins in the periplasm, it was conceivable that DegP co-aggregated with misfolded MalE31 during the formation of IBs. All these data were quantified and analyzed by numerical simulations [95] on the working model presented in Figure 4. Because the kinetics of signal sequence processing of precursors were unaffected by MalE31 overproduction, export to the periplasm does not constitute a rate-limiting step for the periplasmic folding pathway of MalE31. The extent of aggregation is determined by both the rate of protein biosynthesis, which is related for MalE31 to the activity of its gene promoter, and protein folding or misfolding rates. At a high level of expression, second order aggregation dominates over first order folding. The fate of misfolded proteins depends obviously on the existence of cellular proteases. At the misfolding stage a second kinetic competition between degradation and aggregation might occur, and a further increase in the rate of expression (as in the malTC background) will favor the aggregation reaction. At the time this model was proposed, we speculated that periplasmic chaperones and foldases might act directly on some kinetic partitioning steps.
Figure 4.
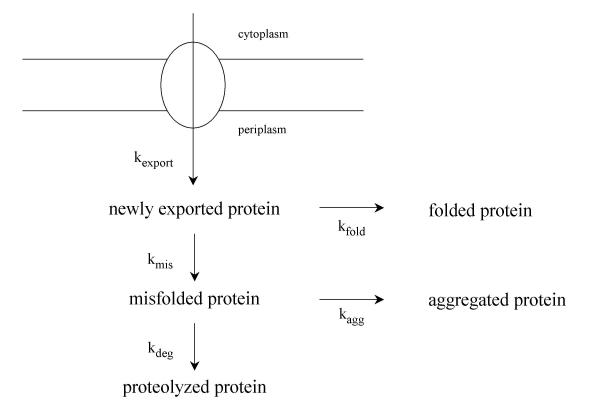
Kinetic competition between folding, degradation and aggregation in the periplasm. The model represents the export (zero order rate constant, kexport), folding (first order rate constant, kfold), degradation (first order rate constant, kdeg), and aggregation reactions (second order rate constant, kagg). In the postulated scheme, the newly exported protein is assumed to have two alternative pathways, either it correctly folds to give the folded protein or it proceeds via a side reaction leading to a misfolded protein. Once again, a kinetic competition is thought to occur between degradation and aggregation.
FkpA suppresses the formation of MalE31 inclusion bodies
The potential role of the three known periplasmic PPIases was studied both in vivo, by overproducing them, and in vitro by recording the refolding kinetics of MalE31 in the presence of purified PPIases. Among these folding catalysts, only FkpA prevented the periplasmic aggregation of MalE31 [37]. Using active site and deletion variants, we demonstrated that the chaperone activity of FkpA was independent of its prolyl isomerase catalytic activity, but requires an intact dimerization N-domain [39]. In vitro, the presence of FkpA at stoichiometric amounts in the refolding buffer did not change the rate of the slow refolding phase of MalE31, but increased its relative amplitude as a chaperone does in general [37]. These results suggested that the ability of FkpA to increase the yield of soluble MalE31 can be explained by the blocking of the initial misfolding step through its binding to newly translocated proteins rather than to misfolded proteins which leads to the second competition between aggregation and degradation, as shown in Figure 4. Indeed, in the absence or in the presence of FkpA, the steady state level of MalE31 remained unchanged, and any modification in the balance between the relative rate of aggregation and degradation would lead to different levels, as we observed with the protease DegP. Thus, FkpA would bind newly exported MalE31 proteins in the periplasm, consistent with the definition of 'holder' chaperones, which can prevent the aggregation of unfolded protein without mediating protein reactivation.
Temperature effect on MalE31 inclusion body formation
Generally, the extent of aggregation is greater at higher temperature due to the strong temperature-dependence of the hydrophobic interactions, which dominate protein aggregation. Although MalE31 aggregated at 30°C, the formation of periplasmic IBs did not affect bacterial growth in rich medium [87]. In contrast, at 37°C, accumulation of IBs became very toxic [49]. Surprisingly, at 42°C, IBs did not accumulate and bacterial growth was restored. To explain this toxicity we hypothesized that some periplasmic or outer membrane proteins, which are essential for envelope biogenesis or cell division, could be incorporated into IBs and therefore lost to the cells. Indeed, DegP [95] and RseB [71] have previously been found trapped with MalE31 aggregates at 30°C. Under heat shock conditions no IBs were detected, thus we examined whether a temperature shift up could suppress the growth defect by growing cells at 37°C and then shifting them to 42°C. Interestingly, after transfer to 42°C the bacterial growth resumed, while the turbidity of cells left at 37°C decreased. We showed that the suppression of MalE31 toxicity after the temperature upshift resulted in the degradation of pre-existing IBs accumulated at 37°C. The only clearly documented heat shock protease which degrades misfoded periplasmic proteins is DegP. The synthesis of DegP at 42°C, as for other σE regulon members, is activated and its proteolytic activity also significantly increases with rising temperature [96]. These cumulative effects could contribute to a further increase of proteolytic activity in the periplasm at 42°C, and the degradation of MalE31 would be favored over aggregation in the model (Figure 4). In addition, because the steady-state level of wild-type MalE was slightly lower at 42°C [49], a further decrease in the rate of MalE31 synthesis, at this temperature, will also facilitate the degradation reaction. However, it has been reported that highly aggregated proteins were not proteolyzed by DegP in vitro [97], and we do not know if DegP is able to degrade IBs. Alternatively, we have no evidence that misfolded folding intermediates of MalE31 can dissociate from IBs and then be degraded by DegP. Recent studies have revealed an unexpected dynamic process of IBs [98], from which aggregated proteins are steadily released or disaggregated to be further refolded or degraded by intracellular proteases [99]. Nevertheless, it is clear that under heat shock conditions, the progressive clearing of periplasmic IBs eliminated toxicity and restored bacterial growth.
Perspectives
Protein folding is one of the great unsolved problems of modern molecular biology. Not only is this question intrinsically fascinating, but it is closely tied to a wide range of applications and debilitating diseases, which are of critical importance in both biotechnology and molecular medicine. Most proteins that require chaperones to fold efficiently are poor folding models for biophysical experiments and, conversely, proteins suitable for in vitro folding studies do not require chaperone assistance. The complete characterization of a protein model, as MalE, and the availability of both in vivo and in vitro approaches, complementing each other, make it a particularly attractive system for mechanistic studies of this central issue. Our understanding of a number of cellular factors relevant to protein folding and assembly has increased enormously during the past five years, and quite remarkable progress has been made in the applications of this knowledge.
Although many alternative organisms and expression systems are now being developed, the most widely used host for high-level expression of recombinant proteins remains E. coli. Many important proteins for therapeutic or diagnostic applications including secreted enzymes contain disulfide bonds that are required for correct folding. Thus, studies on periplasmic protein folding have significant ramifications for biotechnology. However, many of these proteins are produced in biologically inactive and aggregated forms (IBs). Although solubilization and refolding procedures exist to obtain functionally soluble proteins, as well demonstrated with MalE31, they are lengthy processes, and not generalisable. For these proteins, modification of the amino acid sequences by mutagenesis, the folding code, may be one way to isolate a soluble form, and we can envision that genetic screens or selections for the rapid identification of correctly folded proteins expressed in the bacterial periplasm will be developed.
Other potential applications include the use of envelope folding factors as targets for the development of new antimicrobial drugs. Indeed, several proteins involved in periplasmic protein quality control are either essential or play an important role in the virulence of many pathogenic Gram-negative bacteria.
Acknowledgments
Acknowledgements
We are grateful to Emmett Johnson for critical reading of the manuscript. This work was supported by grants from the Institut Pasteur and the CNRS.
Contributor Information
Marika Miot, Email: mmiot@pasteur.fr.
Jean-Michel Betton, Email: jmbetton@pasteur.fr.
References
- Goff SA, Goldberg AL. Production of abnormal proteins in E. coli stimulates transcription of lon and other heat shock genes. Cell. 1985;41:587–595. doi: 10.1016/s0092-8674(85)80031-3. [DOI] [PubMed] [Google Scholar]
- Wickner S, Maurizi MR, Gottesman S. Posttranslational quality control: folding, refolding, and degrading proteins. Science. 1999;286:1888–1893. doi: 10.1126/science.286.5446.1888. [DOI] [PubMed] [Google Scholar]
- Raivio TL, Silhavy TJ. The sigmaE and Cpx regulatory pathways: overlapping but distinct envelope stress responses. Curr Opin Microbiol. 1999;2:159–65. doi: 10.1016/S1369-5274(99)80028-9. [DOI] [PubMed] [Google Scholar]
- Pugsley AP. The complete general secretory pathway in Gram-Negative bacteria. Microbiol Rev. 1993;57:50–108. doi: 10.1128/mr.57.1.50-108.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeLisa MP, Tullman D, Georgiou G. Folding quality control in the export of proteins by the bacterial twin-arginine translocation pathway. Proc Natl Acad Sci USA. 2003;100:6115–6120. doi: 10.1073/pnas.0937838100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van den Berg B, Clemons WM, Jr, Collinson I, Modis Y, Hartmann E, Harrison SC, Rapoport TA. X-ray structure of a protein-conducting channel. Nature. 2004;427:36–44. doi: 10.1038/nature02218. [DOI] [PubMed] [Google Scholar]
- Economou A, Wickner W. SecA promotes preprotein translocation by undergoing ATP-driven cycles of membrane insertion and deinsertion. Cell. 1994;78:835–843. doi: 10.1016/s0092-8674(94)90582-7. [DOI] [PubMed] [Google Scholar]
- Luirink J, Dobberstein B. Mammalian and Escherichia coli signal recognition particles. Mol Microbiol. 1994;11:9–13. doi: 10.1111/j.1365-2958.1994.tb00284.x. [DOI] [PubMed] [Google Scholar]
- Danese PN, Silhavy TJ. Targeting and assembly of periplasmic and outer-membrane proteins in Escherichia coli. Annu Rev Genet. 1998;32:59–94. doi: 10.1146/annurev.genet.32.1.59. [DOI] [PubMed] [Google Scholar]
- Sijbrandi R, Urbanus ML, ten Hagen-Jongman CM, Bernstein HD, Oudega B, Otto BR, Luirink J. Signal recognition particule (SRP)-mediated targeting and Sec-dependent translocation of extracellular Escherichia coli protein. J Biol Chem. 2003;278:4654–4659. doi: 10.1074/jbc.M211630200. [DOI] [PubMed] [Google Scholar]
- Schierle CF, Berkmen M, Huber D, Kumamoto CA, Boyd D, Beckwith J. The DsbA signal sequence directs efficient, cotranslational export of passenger proteins to the Escherichia coli periplasm via the signal recognition particle pathway. J Bacteriol. 2003;185:5706–5713. doi: 10.1128/JB.185.19.5706-5713.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valent QA, Kendall DA, High S, Kusters R, Oudega B, Luirink J. Early events in preprotein recognition in E. coli: interaction of SRP and trigger factor with nascent polypeptides. EMBO J. 1995;14:5494–5505. doi: 10.1002/j.1460-2075.1995.tb00236.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hesterkamp T, Hauser S, Lutcke H, Bukau B. Escherichia coli trigger factor is a prolyl isomerase that associates with nascent polypeptide chains. Proc Natl Acad Sci USA. 1996;93:4437–4441. doi: 10.1073/pnas.93.9.4437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisner G, Koch H-G, Beck K, Brunner J, Müller M. Ligand crowding at a nascent signal sequence. J Cell Biol. 2003;163:35–44. doi: 10.1083/jcb.200306069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumamoto CA. SecB protein: a cytosolic export factor that associates with nascent exported proteins. J Bioenerg Biomembr. 1990;22:337–351. doi: 10.1007/BF00763171. [DOI] [PubMed] [Google Scholar]
- Kusukawa N, Yura T, Uegushi C, Akiayama Y, Ito Y. Effects of mutations in heat-shock genes groES and groEL on protein export in Escherichia coli. EMBO J. 1989;8:3517–3521. doi: 10.1002/j.1460-2075.1989.tb08517.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wild J, Altman E, Yura T, Gross CA. DnaK and DnaJ heat-shock proteins participate in protein export in Escherichia coli. Gene Dev. 1992;6:1165–1172. doi: 10.1101/gad.6.7.1165. [DOI] [PubMed] [Google Scholar]
- Bukau B. Regulation of the Escherichia coli heat-shock response. Mol Microbiol. 1993;9:671–680. doi: 10.1111/j.1365-2958.1993.tb01727.x. [DOI] [PubMed] [Google Scholar]
- Randall LL, Hardy SJ. High selectivity with low specificity: how SecB has solved the paradox of chaperone binding. Trends Biochem Sci. 1995;20:65–9. doi: 10.1016/S0968-0004(00)88959-8. [DOI] [PubMed] [Google Scholar]
- Paetzel M, Dalbey RE, Strynadka NCJ. Crystal structure of a bacterial signal peptidase in complex with a β-lactam inhibitor. Nature. 1998;396:186–190. doi: 10.1038/25403. [DOI] [PubMed] [Google Scholar]
- Kadokura H, Katzen F, Beckwith J. Protein disulfide bond formation in prokaryotes. Annu Rev Biochem. 2003;72:111–135. doi: 10.1146/annurev.biochem.72.121801.161459. [DOI] [PubMed] [Google Scholar]
- Bader M, Muse W, Ballou DP, Gassner C, Bardwell JCA. Oxidative protein folding is driven by the electron transport system. Cell. 1999;98:217–227. doi: 10.1016/S0092-8674(00)81016-8. [DOI] [PubMed] [Google Scholar]
- Rietsch A, Belin D, Martin N, Beckwith J. An in vivo pathway for disulfide bond isomerization in Escherichia coli. Proc Natl Acad Sci USA. 1996;93:13048–13053. doi: 10.1073/pnas.93.23.13048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadokura H, Tian H, Zander T, Bardwell JCA, Beckwith J. Snapshots of DsbA in action: detection of proteins in the process of oxidative folding. Science. 2004;303:534–537. doi: 10.1126/science.1091724. [DOI] [PubMed] [Google Scholar]
- Liu J, Walsh CT. Peptidyl-prolyl cis-trans isomerase from Escherichia coli: a periplasmic homolog of cyclophilin that is not inhibited by cyclosporin A. Proc Natl Acad Sci USA. 1990;87:4028–4032. doi: 10.1073/pnas.87.11.4028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horne SM, Young KD. Escherichia coli and other species of the Enterobacteriaceae encode a protein similar to the Mip-like FK506-binding protein. Arch Microbiol. 1995;163:357–365. doi: 10.1007/s002030050215. [DOI] [PubMed] [Google Scholar]
- Dartigalongue C, Raina S. A new heat-shock gene, ppiD, encodes a peptidyl-prolyl isomerase required for folding of outer membrane proteins in Escherichia coli. EMBO J. 1998;17:3968–3980. doi: 10.1093/emboj/17.14.3968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rouviere PE, Gross CA. SurA, a periplasmic protein with peptidyl-prolyl isomerase activity, participates in the assembly of outer membrane porins. Genes Dev. 1996;10:3170–3182. doi: 10.1101/gad.10.24.3170. [DOI] [PubMed] [Google Scholar]
- Scholz C, Scherer G, Mayr LM, Schindler T, Fischer G, Schmid FX. Prolyl isomerase do not catalyse isomerization of non-prolyl peptide bonds. Biol Chem. 1998;379:361–365. doi: 10.1515/bchm.1998.379.3.361. [DOI] [PubMed] [Google Scholar]
- Barnhart MM, Pinkner JS, Soto GE, Sauer FG, Langermann S, Waksman G, Frieden C, Hultgren SJ. PapD-like chaperones provide the missing information for folding of pilin proteins. Proc Natl Acad Sci USA. 2000;97:7709–7714. doi: 10.1073/pnas.130183897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Missiakas D, Betton J-M, Raina S. New components of protein folding in extracytoplasmic compartments of Escherichia coli SurA, FkpA and Skp/OmpH. Mol Microbiol. 1996;21:871–884. doi: 10.1046/j.1365-2958.1996.561412.x. [DOI] [PubMed] [Google Scholar]
- Schäfer U, Beck K, Müller M. Skp, a molecular chaperone of gram-negative bacteria is required for the formation of soluble periplasmic intermediates of outer membrane proteins. J Biol Chem. 1999;274:24567–24574. doi: 10.1074/jbc.274.35.24567. [DOI] [PubMed] [Google Scholar]
- Chen J, Song J-L, Zhang S, Wang Y, Cui D-F, Wang CC. Chaperone activity of DsbC. J Biol Chem. 1999;274:19601–19605. doi: 10.1074/jbc.274.28.19601. [DOI] [PubMed] [Google Scholar]
- Shao F, Bader MW, Jakob U, Bardwell JCA. DsbG, a protein disulfide isomerase with chaperone activity. J Biol Chem. 2000;275:13349–13352. doi: 10.1074/jbc.275.18.13349. [DOI] [PubMed] [Google Scholar]
- Richarme G, Caldas TD. Chaperone properties of the bacterial periplasmic substrate-binding proteins. J Biol Chem. 1997;272:15607–15612. doi: 10.1074/jbc.272.25.15607. [DOI] [PubMed] [Google Scholar]
- Matsuzaki M, Kiso Y, Yamamoto I, Satoh T. Isolation of a periplasmic molecular chaperone-like protein of Rhodobacter sphaeroides f. sp. denitrificans that is homologous to the dipeptide transport protein DppA of Escherichia coli. J Bacteriol. 1998;180:2718–2722. doi: 10.1128/jb.180.10.2718-2722.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arie J-P, Sassoon N, Betton J-M. Chaperone function of FkpA, a heat shock prolyl isomerase, in the periplasm of Escherichia coli. Mol Microbiol. 2001;39:199–210. doi: 10.1046/j.1365-2958.2001.02250.x. [DOI] [PubMed] [Google Scholar]
- Behrens S, Maier R, de Cock H, Schmid FX, Gross CA. The SurA periplasmic PPIase lacking its parvulin domains functions in vivo and has chaperone activity. EMBO J. 2001;20:285–294. doi: 10.1093/emboj/20.1.285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saul FA, Arié J-P, Vulliez-le Normand B, Kahn R, Betton J-M, Bentley GA. Structural and functional studies of FkpA from Escherichia coli, a cis/trans peptidyl-prolyl isomerase with chaperone activity. J Mol Biol. 2004;335:595–608. doi: 10.1016/j.jmb.2003.10.056. [DOI] [PubMed] [Google Scholar]
- Bitto E, McKay DB. Crystallographic structure of SurA, a molecular chaperone that facilitates folding of outer membrane porins. Structure. 2002;10:1489–1498. doi: 10.1016/S0969-2126(02)00877-8. [DOI] [PubMed] [Google Scholar]
- Zarnt T, Tradler T, Stoller G, Scholz C, Schmid FX, Fischer G. Modular structure of the trigger factor required for high activity in ptotein folding. J Mol Biol. 1997;271:827–837. doi: 10.1006/jmbi.1997.1206. [DOI] [PubMed] [Google Scholar]
- McCarthy AA, Haebel PW, Törrönen A, Rybin V, Baker EN, Metcalf P. Crystal structure of the protein disulfide bond isomerase, DsbC, from Escherichia coli. Nature Struct Biol. 2000;7:196–199. doi: 10.1038/73295. [DOI] [PubMed] [Google Scholar]
- Lipinska B, Zylicz M, Georgopoulos C. The HtrA (DegP) protein, essential for Escherichia coli survival at high temperatures, is an endopeptidase. J Bacteriol. 1990;172:1791–1797. doi: 10.1128/jb.172.4.1791-1797.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strauch KL, Johnson K, Beckwith J. Characterization of degP, a gene required for proteolysis in the cell envelope and essential for growth of Escherichia coli at high temperature. J Bacteriol. 1989;171:2689–2696. doi: 10.1128/jb.171.5.2689-2696.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erickson JW, Gross CA. Identification of the σE subunit of Escherichia coli RNA polymerase: a second alternate σ factor involved in high-temperature gene expression. Genes Dev. 1989;3:1462–1471. doi: 10.1101/gad.3.9.1462. [DOI] [PubMed] [Google Scholar]
- Clausen T, Southan C, Ehrmann M. The HtrA family of proteases: implications for protein composition and cell fate. Mol Cell. 2002;10:443–455. doi: 10.1016/S1097-2765(02)00658-5. [DOI] [PubMed] [Google Scholar]
- Krojer T, Garrido-Franco M, Huber H, Ehrmann M, Clausen T. Crystal structure of DegP (HtrA) reveals a new protease-chaperone machine. Nature. 2002;416:455–459. doi: 10.1038/416455a. [DOI] [PubMed] [Google Scholar]
- Spiess C, Beil A, Ehrmann M. A temperature-dependent switch from chaperone to protease in a widely conserved heat shock protein. Cell. 1999;97:339–347. doi: 10.1016/s0092-8674(00)80743-6. [DOI] [PubMed] [Google Scholar]
- Hunke S, Betton J-M. Temperature effect on inclusion body formation and stress response in the periplasm of Escherichia coli. Mol Microbiol. 2003;50:1579–1589. doi: 10.1046/j.1365-2958.2003.03785.x. [DOI] [PubMed] [Google Scholar]
- Raivio TL, Silhavy TJ. Periplasmic stress and ECF sigma factors. Annu Rev Microbiol. 2001;55:591–624. doi: 10.1146/annurev.micro.55.1.591. [DOI] [PubMed] [Google Scholar]
- Dong J, Iuchi S, Kwan H-S, Lu Z, Lin ECC. The deduced amino-acid sequence of the cloned cpxR gene suggests the protein is the cognate regulator for the membrane sensor, CpxA, in a two-component signal transduction system of Escherichia coli. Gene. 1993;136:227–230. doi: 10.1016/0378-1119(93)90469-J. [DOI] [PubMed] [Google Scholar]
- Weber RF, Silverman PM. The Cpx proteins of Escherichia coli K 12: structure of the CpxA polypeptide as an inner membrane component. J Mol Biol. 1988;203:467–478. doi: 10.1016/0022-2836(88)90013-7. [DOI] [PubMed] [Google Scholar]
- Nakayama S-I, Watanabe H. Involvement of cpxA, a sensor of a two-component regulatory system, in the pH-dependent regulation of expression of Shigella sonnei virF gene. J Bacteriol. 1995;177:5062–5069. doi: 10.1128/jb.177.17.5062-5069.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mileykosvskaya E, Dowhan W. The Cpx two-component signal transduction pathway is activated in Escherichia coli mutant strains lacking phosphatidylethanolamine. J Bacteriol. 1997;179:1029–1034. doi: 10.1128/jb.179.4.1029-1034.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Snyder WB, Davis LJB, Danese PN, Cosma CL, Silhavy TJ. Overproduction of NlpE, a new membrane lipoprotein, suppresses the toxicity of periplasmic LacZ by activation of the Cpx signal transduction pathway. J Bacteriol. 1995;177:4216–4223. doi: 10.1128/jb.177.15.4216-4223.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones CH, Danese PN, Pinkner JS, Silhavy TJ, Hultgren SJ. The chaperone-assisted membrane release and folding pathway is sensed by two signal transduction systems. EMBO J. 1997;16:6394–6406. doi: 10.1093/emboj/16.21.6394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hung DL, Raivio TL, Jones CH, Silhavy TJ, Hultgren SJ. Cpx signaling pathway monitors biogenesis and affects assembly and expression of P pili. EMBO J. 2001;20:1508–1518. doi: 10.1093/emboj/20.7.1508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raivio TL, Popkin DL, Silhavy TJ. The Cpx envelope stress response is controlled by amplification and feedback inhibition. J Bacteriol. 1999;181:5263–72. doi: 10.1128/jb.181.17.5263-5272.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raivio TL, Laird MW, Joly JC, Silhavy TJ. Tethering of CpxP to the inner membrane prevents spheroplast induction of the Cpx envelope stress response. Mol Microbiol. 2000;37:1186–1197. doi: 10.1046/j.1365-2958.2000.02074.x. [DOI] [PubMed] [Google Scholar]
- Danese PN, Snyder WB, Cosma CL, Davis LJB, Silhavy TJ. The Cpx two-component signal transduction pathway of Escherichia coli regulates transcription of the gene specifying the stress-inducible periplasmic protease, DegP. Genes Dev. 1995;9:387–398. doi: 10.1101/gad.9.4.387. [DOI] [PubMed] [Google Scholar]
- Pogliano J, Lynch AS, Belin D, Lin ECC, Beckwith J. Regulation of Escherichia coli cell envelope proteins involved in protein folding and degradation by the Cpx two-component system. Genes Dev. 1997;11:1169–1182. doi: 10.1101/gad.11.9.1169. [DOI] [PubMed] [Google Scholar]
- Silverman PM. Host cell-plasmid interactions in the expression of DNA donor activity by F+ strains of Escherichia coli K-12. Bioessays. 1985;2:254–259. [Google Scholar]
- Leclerc GJ, Tartera C, Metcalf ES. Environmental regulation of Salmonella typhi invasion-defective mutants. Infect Immun. 1998;66:682–691. doi: 10.1128/iai.66.2.682-691.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorel C, Vidal O, Prigent-Combaret C, Vallet I, Lejeune P. Involvement of the Cpx signal transduction pathway of E. coli in biofilm formation. FEMS Microbiol Lett. 1999;178:169–175. doi: 10.1016/S0378-1097(99)00347-X. [DOI] [PubMed] [Google Scholar]
- Otto K, Silhavy TJ. Surface sensing and adhesion of Escherichia coli controlled by the Cpx-signaling pathway. Proc Natl Acad Sci. 2002;99:2287–2292. doi: 10.1073/pnas.042521699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raina S, Missiakas D, Georgopoulos C. The rpoE gene encoding the σE (σ24) heat shock sigma factor of Escherichia coli. EMBO J. 1995;14:1043–1055. doi: 10.1002/j.1460-2075.1995.tb07085.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rouvière PE, De Las Penas A, Mecsas J, Lu CZ, Rudd KE, Gross CA. rpoE, the gene encoding the secon heat-shock sigma factor, σE, in Escherichia coli. EMBO J. 1995;14:1032–1042. doi: 10.1002/j.1460-2075.1995.tb07084.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mecsas J, Rouvière PE, Erickson JW, Donohue TJ, Gross CA. The activity of σE, an Escherichia coli heat-inducible σ-factor, is modulated by expression of outer membrane proteins. Genes Dev. 1993;7:2618–2628. doi: 10.1101/gad.7.12b.2618. [DOI] [PubMed] [Google Scholar]
- De Las Penas A, Connolly L, Gross CA. The σE-mediated response to extracytoplasmic stress in Escherichia coli is transduced by RseA and RseB, two negative regulators of σE. Mol Microbiol. 1997;24:373–385. doi: 10.1046/j.1365-2958.1997.3611718.x. [DOI] [PubMed] [Google Scholar]
- Missiakas D, Mayer MP, Lemaire M, Georgopoulos C, Raina S. Modulation of the Escherichia coli σE(RpoE) heat-shock transcription-factor activity by the RseA, RseB and RseC proteins. Mol Microbiol. 1997;24:355–371. doi: 10.1046/j.1365-2958.1997.3601713.x. [DOI] [PubMed] [Google Scholar]
- Collinet B, Yuzawa H, Chen T, Herrera C, Missiakas D. RseB binding to the periplasmic domain of RseA modulates the RseA: sigmaE interaction in the cytoplasm and the availability of sigmaE.RNA polymerase. J Biol Chem. 2000;275:33898–33904. doi: 10.1074/jbc.M006214200. [DOI] [PubMed] [Google Scholar]
- Ades SE, Connolly LE, Alba BM, Gross CA. The Escherichia coli σE-dependent extracytoplasmic stress response is controlled by the regulated proteolysis of an anti-sigma factor. Gene Dev. 1999;13:2449–2461. doi: 10.1101/gad.13.18.2449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alba BM, Leeds JA, Onufryk C, Lu CZ, Gross CA. DegS and Yael participate sequentially in the cleavage of RseA to activate the σE-dependent extracytoplasmic stress response. Gene Dev. 2002;16:2156–2168. doi: 10.1101/gad.1008902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walsh NP, Alba BM, Bose B, Gross CA, Sauer RT. OMP peptide signals initiate the envelope-stress response by activating DegS protease via relief of inhibition mediated by its PDZ domain. Cell. 2003;113:61–71. doi: 10.1016/S0092-8674(03)00203-4. [DOI] [PubMed] [Google Scholar]
- Dartigalongue C, Missiakas D, Raina S. Characterization of the Escherichia coli sigma E regulon. J Biol Chem. 2001;276:20866–20875. doi: 10.1074/jbc.M100464200. [DOI] [PubMed] [Google Scholar]
- Dartigalongue C, Loferer H, Raina S. EcfE, a new essential inner membrane protease: its role in the regulation of heat shock response in Escherichia coli. EMBO J. 2001;20:5908–5918. doi: 10.1093/emboj/20.21.5908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Danese PN, Silhavy TJ. The σE and the Cpx signal transduction systems control the synthesis of periplasmic protein-folding enzymes in Escherichia coli. Genes Dev. 1997;11:1183–1193. doi: 10.1101/gad.11.9.1183. [DOI] [PubMed] [Google Scholar]
- Boos W, Shuman H. Maltose/maltodextrin system of Escherichia coli: transport, metabolism, and regulation. Microbiol Mol Biol Rev. 1998;62:204–229. doi: 10.1128/mmbr.62.1.204-229.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shuman HA. Active transport of maltose in Escherichia coli K12. J Biol Chem. 1982;257:5455–5461. [PubMed] [Google Scholar]
- Schatz PJ, Beckwith J. Genetic analysis of protein export in Escherichia coli. Ann Rev Genet. 1990;24:215–248. doi: 10.1146/annurev.ge.24.120190.001243. [DOI] [PubMed] [Google Scholar]
- Bassford PJ. Export of the periplasmic maltose-binding protein of Escherichia coli. J Bioenerg Biomembr. 1990;22:401–439. doi: 10.1007/BF00763175. [DOI] [PubMed] [Google Scholar]
- Randall LL, Hardy SJ. Correlation of competence for export with lack of tertiary structure of the mature species: a study in vivo of maltose-binding protein in E. coli. Cell. 1986;46:921–8. doi: 10.1016/0092-8674(86)90074-7. [DOI] [PubMed] [Google Scholar]
- Prinz WA, Spiess C, Ehrmann M, Schierle CF, Beckwith J. Targeting of signal sequenceless proteins for export in Escherichia coli with altered protein translocase. EMBO J. 1996;15:5209–5217. [PMC free article] [PubMed] [Google Scholar]
- Spurlino JC, Lu GY, Quiocho FA. The 2.3-Å resolution structure of the maltose-or maltodextrin-binding protein, a primary receptor of bacterial active transport and chemotaxis. J Biol Chem. 1991;266:5202–5219. doi: 10.2210/pdb1mbp/pdb. [DOI] [PubMed] [Google Scholar]
- Duplay P, Szmelcman S, Bedouelle H, Hofnung M. Silent and functional changes in the periplasmic maltose-binding protein of Escherichia coli K12. J Mol Biol. 1987;194:663–673. doi: 10.1016/0022-2836(87)90243-9. [DOI] [PubMed] [Google Scholar]
- Duplay P, Hofnung M. Two regions of mature periplasmic maltose-binding protein of Escherichia coli involved in secretion. J Bacteriol. 1988;170:4445–4450. doi: 10.1128/jb.170.10.4445-4450.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Betton J-M, Hofnung M. Folding of a mutant maltose binding protein of E. coli which forms inclusion bodies. J Biol Chem. 1996;271:8046–8052. doi: 10.1074/jbc.271.14.8046. [DOI] [PubMed] [Google Scholar]
- Betton J-M, Boscus D, Missiakas D, Raina S, Hofnung M. Probing the structural role of an αβ loop of maltose-binding protein by mutagenesis: heat-shock induction by loop variants of the maltose-binding protein that form periplasmic inclusion bodies. J Mol Biol. 1996;262:140–150. doi: 10.1006/jmbi.1996.0504. [DOI] [PubMed] [Google Scholar]
- Wigley WC, Stidham RD, Smith NM, Hunt JF, Thomas PJ. Protein solubility and folding monitored in vivo by structural complementation of a genetic marker protein. Nat Biotechnol. 2001;19:131–136. doi: 10.1038/84389. [DOI] [PubMed] [Google Scholar]
- Betton J-M, Phichith D, Hunke S. Folding and aggregation of export-defective mutants of the maltose-binding protein. Res Microbiol. 2002;153:399–404. doi: 10.1016/S0923-2508(02)01338-4. [DOI] [PubMed] [Google Scholar]
- Bedouelle H, Bassford PJ, Fowler AV, Zabin I, Beckwith J, Hofnung M. Mutations which alter the function of the signal sequence of the maltose binding protein of Escherichia coli. Nature. 1980;285:78–81. doi: 10.1038/285078a0. [DOI] [PubMed] [Google Scholar]
- Saul FA, Mourez M, Vulliez-Le Normand B, Sassoon N, Bentley GA, Betton J-M. Crystal structure of a defective folding protein. Protein Sci. 2003;12:577–85. doi: 10.1110/ps.0235103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raffy S, Sassoon N, Hofnung H, Betton J-M. Tertiary structure-dependence of misfolding substitutions in loops of the maltose-binding protein. Protein Sci. 1998;7:2136–2142. doi: 10.1002/pro.5560071010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chun S-Y, Strobel S, Bassford PJ, Randall LL. Folding of maltose-binding protein. J Biol Chem. 1993;268:20855–20862. [PubMed] [Google Scholar]
- Betton J-M, Sassoon N, Hofnung M, Laurent M. Degradation versus aggregation of misfolded maltose-binding protein in the periplasm of Escherichia coli. J Biol Chem. 1998;273:8897–8902. doi: 10.1074/jbc.273.15.8897. [DOI] [PubMed] [Google Scholar]
- Skorko-Glonek J, Krzewski K, Lipinska B, Bertoli E, Tanfani F. Comparaison of the structure of wild-type HtrA heat shock protease and mutant HtrA proteins. J Biol Chem. 1995;270:11140–11146. doi: 10.1074/jbc.270.19.11140. [DOI] [PubMed] [Google Scholar]
- Kim KI, Park S-C, Kang SH, Cheong G-W, Chung CH. Selective degradation of unfolded proteins by the self-compartmentalizing HtrA protease, a periplasmic heat shock protein in Escherichia coli. J Mol Biol. 1999;294:1363–1374. doi: 10.1006/jmbi.1999.3320. [DOI] [PubMed] [Google Scholar]
- Carrio MM, Villaverde A. Construction and deconstruction of bacterial inclusion bodies. J Biotechnol. 2002;96:3–12. doi: 10.1016/S0168-1656(02)00032-9. [DOI] [PubMed] [Google Scholar]
- Carrio MM, Corchero JL, Villaverde A. Proteolytic digestion of bacterial inclusion body proteins during dynamic transition between soluble and insoluble forms. Biochim Biophys Acta. 1999;1434:170–6. doi: 10.1016/s0167-4838(99)00177-6. [DOI] [PubMed] [Google Scholar]


