Abstract
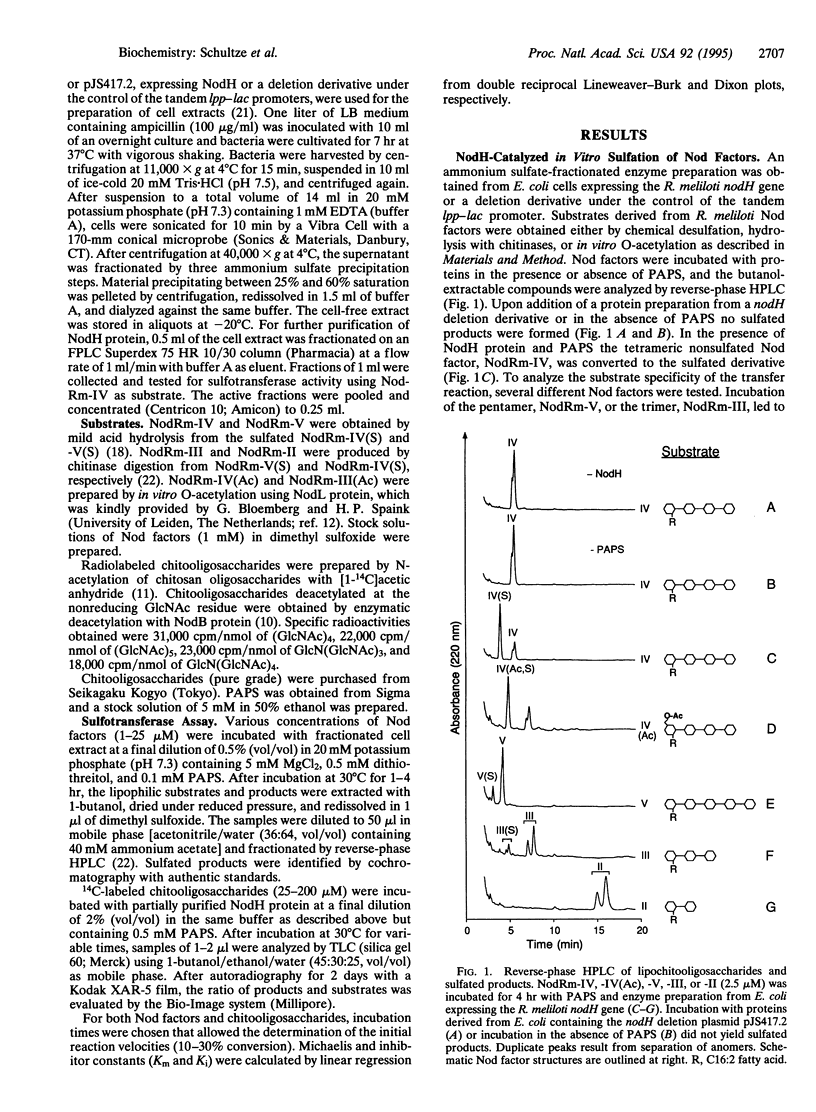
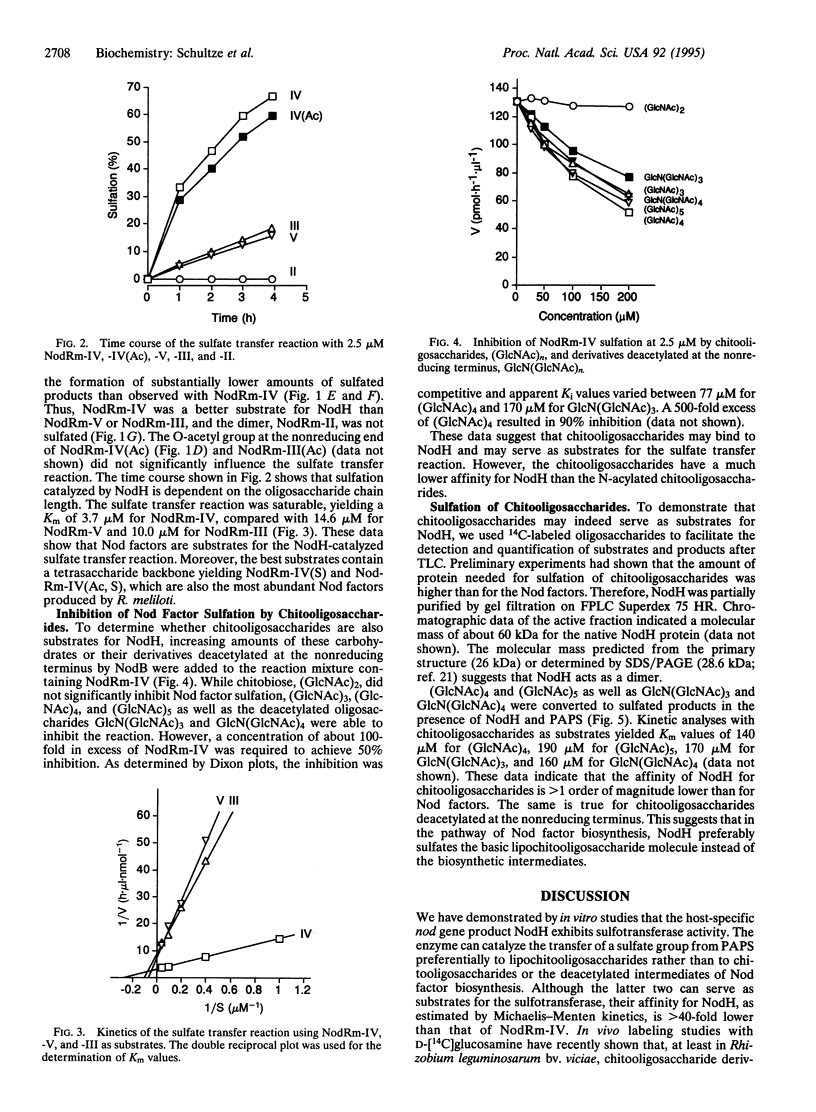
The Rhizobium common nod gene products NodABC are involved in the synthesis of the core lipochitooligosaccharide (Nod factor) structure, whereas the products of the host-specific nod genes are necessary for diverse structural modifications, which vary in different Rhizobium species. The sulfate group attached to the Rhizobium meliloti Nod signal is necessary for activity on the host plant alfalfa, while its absence renders the Nod factor active on the non-host plant vetch. This substituent is therefore a major determinant of host specificity. The exact biosynthetic pathway of Nod factors has not been fully elucidated. In particular, it is not known why some chemical modifications are introduced with high fidelity whereas others are inaccurate, giving rise to a family of different Nod factor structures produced by a single Rhizobium strain. Using protein extracts and partially purified recombinant NodH protein obtained from Escherichia coli expressing the R. meliloti nodH gene, we demonstrate here NodH-dependent in vitro sulfotransferase activity. Kinetic analyses with Nod factors, chitooligosaccharides, and their deacetylated derivatives revealed that Nod factors are the preferred substrate for the sulfate transfer. Moreover, the tetrameric Nod factor, NodRm-IV, was a better substrate than the trimer, NodRm-III, or the pentamer, NodRm-V. These data suggest that the core lipochitooligosaccharide structure must be synthesized prior to its host-specific modification with a sulfate group. Since in R. meliloti tetrameric Nod factors are the most abundant and the most active ones, high affinity of NodH for the appropriate tetrameric substrate guarantees its modification and thus contributes to the fidelity of host-specific behavior.
Full text
PDF

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Atkinson E. M., Palcic M. M., Hindsgaul O., Long S. R. Biosynthesis of Rhizobium meliloti lipooligosaccharide Nod factors: NodA is required for an N-acyltransferase activity. Proc Natl Acad Sci U S A. 1994 Aug 30;91(18):8418–8422. doi: 10.1073/pnas.91.18.8418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baev N., Endre G., Petrovics G., Banfalvi Z., Kondorosi A. Six nodulation genes of nod box locus 4 in Rhizobium meliloti are involved in nodulation signal production: nodM codes for D-glucosamine synthetase. Mol Gen Genet. 1991 Aug;228(1-2):113–124. doi: 10.1007/BF00282455. [DOI] [PubMed] [Google Scholar]
- Baev N., Schultze M., Barlier I., Ha D. C., Virelizier H., Kondorosi E., Kondorosi A. Rhizobium nodM and nodN genes are common nod genes: nodM encodes functions for efficiency of nod signal production and bacteroid maturation. J Bacteriol. 1992 Dec;174(23):7555–7565. doi: 10.1128/jb.174.23.7555-7565.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bloemberg G. V., Thomas-Oates J. E., Lugtenberg B. J., Spaink H. P. Nodulation protein NodL of Rhizobium leguminosarum O-acetylates lipo-oligosaccharides, chitin fragments and N-acetylglucosamine in vitro. Mol Microbiol. 1994 Feb;11(4):793–804. doi: 10.1111/j.1365-2958.1994.tb00357.x. [DOI] [PubMed] [Google Scholar]
- Demont N., Debellé F., Aurelle H., Dénarié J., Promé J. C. Role of the Rhizobium meliloti nodF and nodE genes in the biosynthesis of lipo-oligosaccharidic nodulation factors. J Biol Chem. 1993 Sep 25;268(27):20134–20142. [PubMed] [Google Scholar]
- Geiger O., Thomas-Oates J. E., Glushka J., Spaink H. P., Lugtenberg B. J. Phospholipids of Rhizobium contain nodE-determined highly unsaturated fatty acid moieties. J Biol Chem. 1994 Apr 15;269(15):11090–11097. [PubMed] [Google Scholar]
- Geremia R. A., Mergaert P., Geelen D., Van Montagu M., Holsters M. The NodC protein of Azorhizobium caulinodans is an N-acetylglucosaminyltransferase. Proc Natl Acad Sci U S A. 1994 Mar 29;91(7):2669–2673. doi: 10.1073/pnas.91.7.2669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horvath B., Kondorosi E., John M., Schmidt J., Török I., Györgypal Z., Barabas I., Wieneke U., Schell J., Kondorosi A. Organization, structure and symbiotic function of Rhizobium meliloti nodulation genes determining host specificity for alfalfa. Cell. 1986 Aug 1;46(3):335–343. doi: 10.1016/0092-8674(86)90654-9. [DOI] [PubMed] [Google Scholar]
- John M., Röhrig H., Schmidt J., Wieneke U., Schell J. Rhizobium NodB protein involved in nodulation signal synthesis is a chitooligosaccharide deacetylase. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):625–629. doi: 10.1073/pnas.90.2.625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lerouge P., Roche P., Faucher C., Maillet F., Truchet G., Promé J. C., Dénarié J. Symbiotic host-specificity of Rhizobium meliloti is determined by a sulphated and acylated glucosamine oligosaccharide signal. Nature. 1990 Apr 19;344(6268):781–784. doi: 10.1038/344781a0. [DOI] [PubMed] [Google Scholar]
- Marie C., Barny M. A., Downie J. A. Rhizobium leguminosarum has two glucosamine synthases, GlmS and NodM, required for nodulation and development of nitrogen-fixing nodules. Mol Microbiol. 1992 Apr;6(7):843–851. doi: 10.1111/j.1365-2958.1992.tb01535.x. [DOI] [PubMed] [Google Scholar]
- Roche P., Debellé F., Maillet F., Lerouge P., Faucher C., Truchet G., Dénarié J., Promé J. C. Molecular basis of symbiotic host specificity in Rhizobium meliloti: nodH and nodPQ genes encode the sulfation of lipo-oligosaccharide signals. Cell. 1991 Dec 20;67(6):1131–1143. doi: 10.1016/0092-8674(91)90290-f. [DOI] [PubMed] [Google Scholar]
- Röhrig H., Schmidt J., Wieneke U., Kondorosi E., Barlier I., Schell J., John M. Biosynthesis of lipooligosaccharide nodulation factors: Rhizobium NodA protein is involved in N-acylation of the chitooligosaccharide backbone. Proc Natl Acad Sci U S A. 1994 Apr 12;91(8):3122–3126. doi: 10.1073/pnas.91.8.3122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultze M., Quiclet-Sire B., Kondorosi E., Virelizer H., Glushka J. N., Endre G., Géro S. D., Kondorosi A. Rhizobium meliloti produces a family of sulfated lipooligosaccharides exhibiting different degrees of plant host specificity. Proc Natl Acad Sci U S A. 1992 Jan 1;89(1):192–196. doi: 10.1073/pnas.89.1.192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwedock J. S., Long S. R. Rhizobium meliloti genes involved in sulfate activation: the two copies of nodPQ and a new locus, saa. Genetics. 1992 Dec;132(4):899–909. doi: 10.1093/genetics/132.4.899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwedock J., Long S. R. ATP sulphurylase activity of the nodP and nodQ gene products of Rhizobium meliloti. Nature. 1990 Dec 13;348(6302):644–647. doi: 10.1038/348644a0. [DOI] [PubMed] [Google Scholar]
- Spaink H. P., Sheeley D. M., van Brussel A. A., Glushka J., York W. S., Tak T., Geiger O., Kennedy E. P., Reinhold V. N., Lugtenberg B. J. A novel highly unsaturated fatty acid moiety of lipo-oligosaccharide signals determines host specificity of Rhizobium. Nature. 1991 Nov 14;354(6349):125–130. doi: 10.1038/354125a0. [DOI] [PubMed] [Google Scholar]
- Spaink H. P., Wijfjes A. H., van der Drift K. M., Haverkamp J., Thomas-Oates J. E., Lugtenberg B. J. Structural identification of metabolites produced by the NodB and NodC proteins of Rhizobium leguminosarum. Mol Microbiol. 1994 Sep;13(5):821–831. doi: 10.1111/j.1365-2958.1994.tb00474.x. [DOI] [PubMed] [Google Scholar]




