Bronchial asthma is a common inflammatory disease caused by the interaction of genetic and environmental factors1,2. Through a genome-wide association study (GWAS) and a replication study consisting of a total of 7,171 subjects with adult asthma and 27,912 controls in the Japanese population, we identified five candidate loci associated with susceptibility to adult asthma. In addition to the major histocompatibility complex (MHC) and TSLP/WDR36 loci reported previously, we identified three additional loci: a USP38/GAB1 locus on chromosome 4q31 (combined P = 1.87 × 10-12), a locus on chromosome 10p14 (P = 1.79 × 10-15) and a gene-rich region on chromosome 12q13 (P = 2.33 × 10-13). The most significant association with adult asthma was observed at rs404860 in the MHC region (P = 4.07 × 10-23), which is close to rs2070600 previously reported for association with FEV1/FVC by GWASs for lung function. Our findings may offer better understanding of the pathophysiology of asthma.
The precise mechanisms underlying the development and progression of bronchial asthma have not been fully elucidated. GWASs have contributed to identification of common genetic variants related to common diseases including asthma3. GWASs of asthma in Caucasian and African-ancestry populations4-8 have been conducted, and a recent large-scale, consortium-based GWAS among European populations reported ten loci strongly associated with asthma9. To provide further information for better understanding of the genetic basis of asthma, GWASs using different ethnic populations are also needed.
Since there have been few large-scale analyses of asthma in Asian populations, we conducted a GWAS with 1,532 cases and 3,304 controls in the Japanese population, using Illumina HumanHap 550v3/610-Quad BeadChips (Supplementary Table 1). After principal component analysis (PCA) and QC control, we excluded one outlier before the association analysis and subjected a total of 458,847 SNP loci to statistical analysis, and generated a quantile-quantile plot using the results of a Cochran-Armitage test (Supplementary Fig. 1a, b). The genomic inflation factor (λGC) was 1.07, and λ1000 adjusted by sample size was calculated to be 1.03, indicating a low possibility of false-positive associations resulting from population stratification. The strongest association (rs8192565) was observed in the MHC class III region on chromosome 6p21 (Fig. 1), and two variants in the NOTCH4 and C6orf10 loci reached the genome-wide significant level (P < 5 × 10-8; Supplementary Fig. 1c).
Figure 1.
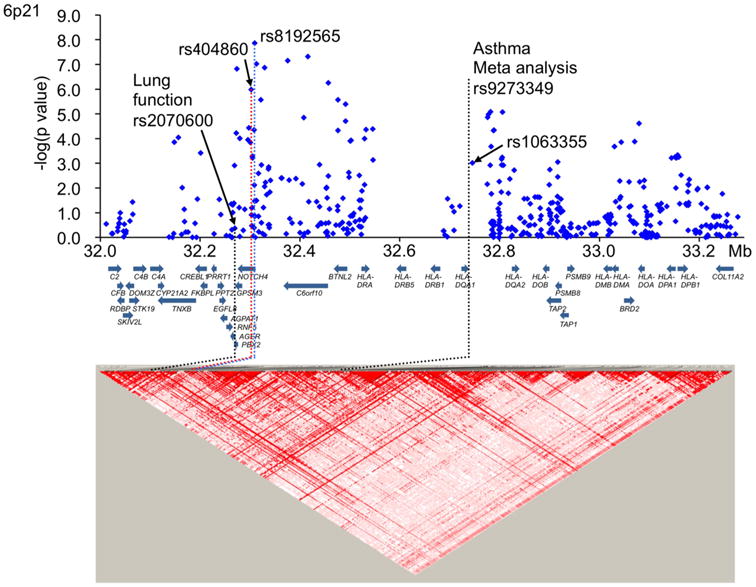
Case-control association results and LD map of the MHC region. The blue plots show the -log10 of Cochrane-Armitage trend P values for association results of the GWAS. The LD maps based on D′ were drawn using the genotype data of all cases and controls in the GWAS. Red dotted line, rs404860; blue dotted line, rs8192565; black dotted lines, locations of the SNPs giving significant signals for lung function (rs2070600), FEV1/FVC, in the GWAS and for bronchial asthma in a large-scale, consortium-based GWAS (rs9273349/rs1063355). rs1063355 is absolute LD with rs9273349 (r2 = 1) in the HapMap JPT and CEU populations. Blue arrows indicate the locations of genes. LD among the SNPs is shown in supplementary Table 4.
We validated the association of previously reported loci for bronchial asthma in GWAS. A recent large scale, consortium-based GWAS identified ten loci for asthma with genome-wide significance9. Our GWAS replicated three SNPs: rs1063355 on chromosome 6p21 (P = 0.0010), rs11071559 on chromosome 15q22 (P = 0.0036) and rs744910 on chromosome 15q22 (P = 0.0095) (Supplementary Table 2a, b). We also checked the associations of genes reported by previous GWASs for asthma5-7,9. The SNPs for which we observed replication of the association with P values < 0.05 in our GWAS are shown in Supplementary Table 2c.
To investigate additional susceptibility loci for adult asthma, we conducted a replication study using an independent set of 5,639 cases and 24,608 controls. We genotyped the 102 SNPs that showed Cochrane-Armitage trend P-values of < 1 × 10-4 in our GWAS after excluding 31 SNPs having r2 > 0.9 with some of the 102 SNPs. In the replication study, 24 of the 102 SNPs replicated their significant associations with Bonferroni corrected P values < 4.9 × 10-4 (0.05/102) (Supplementary Table 3). When we combined data of the GWAS and replication study by the Mantel-Haenszel method, all of the 24 SNPs in five genomic loci were demonstrated to be significantly associated with adult asthma at genome-wide significant levels (P < 5 × 10-8): chromosome 4q31 (rs7686660, combined P = 1.87 × 10-12, odds ratio (OR) = 1.16), 5q22 (rs1837253, P = 1.24 × 10-16, OR = 1.17), 6p21 (rs404860, P = 4.07 × 10-23, OR = 1.21), 10p14 (rs10508372, P = 1.79 × 10-15, OR = 1.16) and 12q13 (rs1701704, P = 2.33 × 10-13, OR = 1.19) (Table 1). A recent study by the candidate gene approach reported that the marker SNP rs1837253, which is present 5.7kb upstream of the transcription initiation site of the TSLP gene, was associated with asthma and airway hyperresponsiveness10. It was also reported that the HLA locus was associated with asthma and allergen sensitization9,11. Considering these previously reported results, our data identified three novel loci associated with adult asthma.
Table 1. Summary of the association results of the GWAS and the replication study.
| dbSNP Locus | Allele Risk | Case | Control | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Stage | Total | RAF | Total RAF | Pa,b | OR(95%CI)c | Phete | |||
| rs7686660 | T/G | GWAS | 1532 | 0.31 | 3301 | 0.27 | 1.33 × 10-4 | 1.21 (1.10-1.33) | |
| 4q31 | T | Replication | 5623 | 0.30 | 24606 | 0.27 | 3.33 × 10-9 | 1.15 (1.10-1.20) | |
| Combinedd | 7155 | 0.30 | 27907 | 0.27 | 1.87 × 10-12 | 1.16 (1.11-1.21) | 0.29 | ||
| rs1837253 | T/C | GWAS | 1532 | 0.40 | 3304 | 0.36 | 3.50 × 10-5 | 1.21 (1.11-1.32) | |
| 5q22 | C | Replication | 5629 | 0.39 | 24601 | 0.35 | 1.02 × 10-12 | 1.17 (1.12-1.22) | |
| Combinedd | 7161 | 0.39 | 27905 | 0.35 | 1.24 × 10-16 | 1.17 (1.13-1.22) | 0.43 | ||
| rs404860 | A/G | GWAS | 1532 | 0.53 | 3304 | 0.48 | 2.33 × 10-6 | 1.24 (1.13-1.35) | |
| 6p21 | A | Replication | 5583 | 0.55 | 24600 | 0.50 | 6.42 × 10-18 | 1.20 (1.15-1.25) | |
| Combinedd | 7115 | 0.55 | 27904 | 0.50 | 4.07 × 10-23 | 1.21 (1.16-1.25) | 0.54 | ||
| rs10508372 | T/C | GWAS | 1532 | 0.47 | 3304 | 0.43 | 3.47 × 10-5 | 1.21 (1.11-1.31) | |
| 10p14 | C | Replication | 5612 | 0.47 | 24608 | 0.43 | 1.31 × 10-11 | 1.15 (1.11-1.20) | |
| Combinedd | 7144 | 0.47 | 27912 | 0.43 | 1.79 × 10-15 | 1.16 (1.12-1.21) | 0.36 | ||
| rs1701704 | T/G | GWAS | 1532 | 0.23 | 3304 | 0.19 | 2.16 × 10-6 | 1.30 (1.17-1.45) | |
| 12q13 | G | Replication | 5590 | 0.21 | 24608 | 0.18 | 7.22 × 10-9 | 1.16 (1.11-1.22) | |
| Combinedd | 7122 | 0.21 | 27912 | 0.18 | 2.33 × 10-13 | 1.19 (1.14-1.25) | 0.06 | ||
RAF, risk allele frequency; OR, odds ratio; CI, confidence interval.
P value of the Cochran-Armitage trend test for each stage.
P values in the GWAS indicate λ-corrected probability PGC.
Odds ratios (OR) and confidence intervals (CI) were calculated using the non-susceptible allele as a reference.
Results of combined analyses were calculated by the Mantel-Haenszel method.
Results of Breslow-Day test.
After we combined the data of the GWAS and the replication study (Supplementary Table 3), 18 of the 26 SNPs in the MHC region showed association beyond the genome-wide significance threshold of P < 5 × 10-8. The strongest association was observed in the MHC class III region (Table 1, Fig. 1). A recent large scale European GWAS reported an association of the HLA-DQ locus (rs9273349) with asthma, and later-onset cases seemed to be influenced more significantly by this locus than childhood-onset cases9. To examine the relationship between rs9273349 and rs8192565, we conducted logistic regression analysis using GWAS data. Since rs9273349 was not included in our GWAS data, we used rs1063355 as a proxy SNP (r2 = 1 between rs1063355 and rs9273349 in the HapMap JPT and CEU populations) (Supplementary Table 2b). The strong association of SNPs in the MHC class III locus remained after adjustment for rs1063355 (Supplementary Fig. 2). This result indicated that the association observed in our study was independent of rs1063355- rs9273349 in the HLA-DQ locus.
Recent GWASs of natural variations in lung function have shown that a nonsynonymous coding SNP in AGER (rs2070600) on chromosome 6p21 is associated with forced expiratory volume in one second (FEV1)/forced vital capacity (FVC) 12,13. Reduction of FEV1/FVC is a characteristic of obstructive lung diseases such as asthma14. Rs2070600 showed relatively weak linkage disequilibrium (LD) with rs404860 in the Japanese population (r2 = 0.115 and D′ = 0.805; Supplementary Table 2b) and did not show association with adult asthma in our study. However, rs404860 is located close to rs2070600, 32.9 kb downstream of the SNP (Fig. 1). Asthma is a heterogenous disease with differences in severity and natural history, and lower pulmonary function is a feature of severe asthma. Thus, further genetic studies of asthma with detailed phenotypic characterization might help to clarify the relationship between genetic determinants of lung function and susceptibility to asthma.
To further investigate the gene(s) susceptible to adult asthma mapped to four loci, (excluding the MHC region), we performed detailed mapping analyses using 1,532 cases and 3,304 controls from the GWAS (Fig. 2a-d). We selected and genotyped tag SNPs (the minor allele frequency of ≥ 0.05 and r2 of < 0.8) using the data from the HapMap phase II+III (release 27) Japanese individuals. The fine mapping analysis of the region on chromosome 5q22 using 13 tag-SNPs revealed that rs1837253 represented an associated LD block spanning 88 kb that included two genes, TSLP and WDR36 (Fig. 2a and Supplementary Table 4a). Respiratory viruses, multiple protease allergens and inflammatory cytokines can induce TSLP expression in airway epithelial cells, and TSLP plays a critical role in sensing environmental agents and mediating T helper type 2 (TH2) cell responses following exposure15. Although further investigation is required to disclose the function of the related variants in asthma, current annotation suggests that TSLP is the most plausible susceptibility gene in this locus.
Figure 2.

Case-control association results and LD maps of the four candidate regions. (a-d) P value plot, genomic structures and LD maps of chromosomes 5q22 (a), 10p14 (b), 12q13 (c) and 4q31 (d). The blue plots show the -log10 of Cochrane-Armitage trend P values for association results of the GWAS. The LD maps based on D′ were drawn using the genotype data of all cases and controls in the GWAS. Black and red dotted lines indicate the ranges of the susceptible regions and positions of marker SNPs, respectively. Blue arrows indicate the locations of genes.
Fine mapping analysis of the locus on chromosome 10p14 with 22 tag-SNPs revealed that rs10508372 was located within a 112-kb LD block that contained no reported genes (Fig. 2b and Supplementary Table 4b). This region is a gene desert and located 1Mb downstream of the GATA3 gene, a master regulator of TH2-cell differentiation16. As enhancers might influence the expression of the gene with megabase-long genomic distance17, it should be verified whether the variants within the associated region affect the activity of long-range cis-regulatory elements controlling GATA3 gene expression.
The fourth locus was located in a gene-rich region on chromosome 12q13. Fine mapping analysis using 12 additional tag-SNPs indicated that rs1701704 was present within a 201-kb LD block including 13 genes (Fig. 2c and Supplementary Table 4c). Recent association studies have shown that rs1701704 at the locus is associated with type I diabetes18 and alopecia areata19. The SNP rs1701704 was located 2kb upstream from the IKZF4 gene (also known as Eos), which was shown to be involved in differentiation of regulatory T cells as a coregulator of FoxP3-directed gene silencing20. Regulatory T cells maintain pulmonary homeostasis and prevent harmful immune responses to innocuous inhaled antigens21.
The fifth locus on chromosome 4q31was finely mapped using 11 tag-SNPs. The 497-kb susceptibility LD block region including rs1397527 contained the USP38/GAB1 genes (Fig. 2d and Supplementary Table 4d). USP38 encodes ubiquitin specific peptidase 38, whose function is unclear. GAB1 is a scaffolding adopter protein that plays an important role in the signaling pathway activated by cytokine receptors for IL-3, IL-6, interferon α and γ, and B-cell and T-cell receptors22.
To examine the association of these variants with adult asthma in populations of different ancestries, we conducted an association study in 499 non-Hispanic white persons with mild to severe asthma from three adult asthma populations and 639 controls obtained from Illumina's iControl database. We found that the rs1837253 SNP on 5q22 showed a significant association with adult asthma (P = 0.023) (Supplementary Table 5). Hence, we conclude that the locus containing the TSLP gene may be a common susceptible region for asthma across the various ethnic groups.
In conclusion, we identified five loci susceptible to adult asthma in the Japanese population with genome-wide significant levels. The three novel loci identified in our GWAS were not observed in a large meta-analysis. In the previous meta-analysis, 65.4% of cases were subjects with childhood-onset asthma. In contrast, we recruited subjects with adult asthma, though 19.6% of the cases in the GWAS were subjects with childhood-onset asthma. Asthma has various clinical phenotypes that show age-related variation. The heterogeneity of phenotypes among studies might influence the results. The peak of association at 6p21 in this study was located close to rs2070600, which was associated with FEV1/FVC in GWASs of lung function in healthy individuals. We found for the first time an association of asthma with the TSLP/WDR36 locus at a genome-wide significant level. A SNP in the TSLP/WDR36 locus was associated with adult asthma in populations of both Japanese and non-Hispanic white decent. Further studies are necessary for better understanding of the clear role of genetic etiology and the pathophysiology of bronchial asthma.
URLs. The Leading Project for Personalized Medicine, http://biobankjp.org/; Haploview v4.2, http://www.broadinstitute.org/haploview/haploview; PLINK statistical software v1.06, http://pngu.mgh.harvard.edu/∼purcell/plink/.
Online Methods
Study subjects
Characteristics of each case-control group are shown in Supplementary Table 1.
RIKEN case samples
A total of 1,532 case subjects for the GWAS were recruited from the Miyatake Asthma Clinic, Miyagawa Clinic, Kyoto University Hospital, Tsukuba University Hospital and several other hospitals. All subjects with asthma were diagnosed by physicians according to the American Thoracic Society criteria as described23.
BioBank Japan cases samples
The BioBank Japan project has been running since 2003 aiming at collection of basic information of patients for personalized medicine24. The subjects were recruited from several medical institutes in Japan, including the Fukujuji Hospital, Iizuka Hospital, Juntendo University, Hospital Iwate Medical University School of Medicine, National Hospital Organization Osaka National Hospital, Nihon University, Nippon Medical School, Shiga University of Medical Science, the Cancer Institute Hospital of the Japanese Foundation for Cancer Research, Tokushukai Hospital and Tokyo Metropolitan Geriatric Hospital. We selected case samples from the subjects who participated in the BioBank Japan. A total of 5,639 cases for replication study were recruited. All subjects with bronchial asthma were diagnosed by physicians and recruited in hospitals.
Controls
We used genome-wide screening data of the BioBank Japan project as controls. Individuals with bronchial asthma, allergic rhinitis, atopic dermatitis, lung fibrosis or chronic obstructive pulmonary disease were excluded from controls. Controls for the GWAS consisted of 2,403 cases in BioBank Japan and 901 healthy volunteers from members of the Rotary Club of Osaka-Midosuji District 2660 Rotary International in Japan25. Controls for the replication study consisted of 24,608 cases registered in BioBank Japan.
All individuals were Japanese and gave written informed consent to participate in the study. This research project was approved by the ethical committees at the Institute of Medical Science, the University of Tokyo and RIKEN Yokohama Institute.
Sample descriptions of Sepracor/LOCCS/LODO/Illumina
This cohort consisted of 499 non-Hispanic white cases with mild to severe asthma from three adult asthma populations: (1) a medication trial conducted by Sepracor, Inc., US26, 27; (2) the Leukotriene Modifier or Corticosteroid Salmeterol (LOCCS) study28; and (3) the Effectiveness of Low Dose Theophylline as an Add-on Treatment in Asthma (LODO) trial28. Cases were matched with 639 population controls obtained from Illumina's IconDB resource (http://www.illumina.com/science/icontroldb.ilmn) using the genetic matching (GEM) algorithm29. Genome-wide genotyping of cases was performed on the Illumina 610 quad platform. Genome-wide genotyping of controls was performed on the Illumina HumanHap 550K v3 platform. Allelic model association statistics were calculated with PLINK30. The corresponding genetic inflation factor was 1.03, demonstrating minimal population stratification. Power calculations were performed using the Genetic Power Calculator by Purcell S, et al31 with effect size as estimated in primary combined GWAS, disease prevalence of 0.05, D-prime of 1, marker allele frequency of cases, assuming use of unselected controls, and with default error rates (alpha=0.05, power=0.80).
Genotyping and quality control
For the GWAS, we genotyped 1,532 cases using the Illumina HumanHap610-Quad BeadChip and 3,304 controls using the Illumina HumanHap550 BeadChip. We excluded related samples by allele sharing analysis and performed principal component analysis for the genotype data of the samples along with European (CEU), African (YRI) and East Asian (Japanese [JPT], and Han Chinese [CHB]) individuals obtained from the Phase II HapMap database by using smartpca32. We excluded outliers, who were away from the East Asian cluster (Supplementary Fig. 1a). We also excluded SNPs with minor allele frequencies of less than 0.01 in both cases and controls. SNPs having call rates ≥99% in both cases and controls were used for the association study. We conducted exact Hardy-Weinberg equilibrium (HWE) analysis, and SNPs with P values less than the cut-off values of the HWE test (P < 10-6 in controls) were excluded from the analysis.
In the replication study, we genotyped patients with asthma using the TaqMan assay (Applied Biosystems) or multiplex-PCR based Invader assay (Third Wave Technologies). We calculated the concordance rates of the five SNPs in Table 1 between genotypes determined by the Illumina HumanHap610-Quad BeadChip and those determined by the TaqMan assay (rs1837253, rs10508372 and rs1701704) or multiplex-PCR based Invader assay (rs7686660 and rs404860). We genotyped a total of 1532 cases in the GWAS, and the concordance rates were 1.000, 0.999, 1.000, 1.000 and 1.000, respectively. The concordance rate between the genotypes determined by the Illumina Human610-Quad BeadChip and Illumina HumanHap550v3 BeadChip among 182 duplicated samples was 0.9999825. Cluster plots of the GWAS for the five SNPs are shown in Supplementary Fig. 3.
Statistical analysis
In the GWAS and replication study, the statistical significance of the association with each SNP was assessed using a one-degree-of-freedom Cochran-Armitage trend test. OR and CI were calculated from a two-by-two allele frequency table. Combined analysis was conducted using the Mantel-Haenszel method and heterogeneity among studies was examined with the Breslow-Day test. We used Haploview 4.2 software to analyze the LD values between SNPs.
Supplementary Material
Acknowledgments
We thank all the patients for participating in the study and also thank the collaborating physicians for helping with sample collection. We are grateful to the members of BioBank Japan and the Rotary Club of Osaka-Midosuji District 2660 Rotary International in Japan for supporting our study. We thank M. T. Shimizu, H. Sekiguchi, A. I. Jodo, N. Kawaraichi and the technical staff of the Center for Genomic Medicine for providing technical assistance. This work was conducted as a part of the BioBank Japan Project supported by of Ministry of Education, Culture, Sports, Science and Technology, Japan. This work was also partly supported by grants from the Ministry of Health, Labour and Welfare, Japan. We acknowledge the American Lung Association (ALA) and the ALA's Asthma Clinical Research Centers investigators and research teams for use of LOCCS and LoDo data, with additional funding from HL071394 and HL074755 from the NHLBI, and Nemours Childrens’ Clinic. GlaxoSmithKline supported the conduct of the LOCCS Trial by an unrestricted grant to the ALA. We acknowledge Sepracor, Inc. for use of the Asthma Trial data. C.G.I. was supported by grants from the NIH NCRR RR022675 and RR015557 as well as the ALA, Asthma Clinical Research Center award.
Footnotes
Author Contributions: T.H. and M.T. designed the study and drafted the manuscript. A.T. and T.T. analyzed the GWAS data. T.H., K.T. and M.K. performed the genotyping for the GWAS. S.D., K.F., A.M., T.E., T.M., M.A., H.T., A.N., H.M., I.I., H.M., T.S., N. Hizawa, and M. Taniguchi collected subjects and participated in the diagnostic evaluations. B.E.H., A.A.L., K.G.T., J.J.L., C.G.I., S.P.P., and S.T.W. conducted an association study in a non-Hispanic white population. M.T. and S.T.W. wrote the manuscript. M.K., N.K. and Y.N. contributed to the overall GWAS study design.
Competing Financial Interests: The authors declare no competing financial interests.
Note: supplementary information is available on the Nature Genetics website.
References
- 1.Barnes PJ. Immunology of asthma and chronic obstructive pulmonary disease. Nat Rev Immunol. 2008;8:183–192. doi: 10.1038/nri2254. [DOI] [PubMed] [Google Scholar]
- 2.Vercelli D. Discovering susceptibility genes for asthma and allergy. Nat Rev Immunol. 2008;8:169–182. doi: 10.1038/nri2257. [DOI] [PubMed] [Google Scholar]
- 3.Kabesch M. Novel asthma-associated genes from genome-wide association studies: what is their significance? Chest. 2010;137:909–915. doi: 10.1378/chest.09-1554. [DOI] [PubMed] [Google Scholar]
- 4.Moffatt MF, et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature. 2007;448:470–473. doi: 10.1038/nature06014. [DOI] [PubMed] [Google Scholar]
- 5.Himes BE, et al. Genome-wide association analysis identifies PDE4D as an asthma-susceptibility gene. Am J Hum Genet. 2009;84:581–593. doi: 10.1016/j.ajhg.2009.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sleiman PM, et al. Variants of DENND1B associated with asthma in children. N Engl J Med. 2010;362:36–44. doi: 10.1056/NEJMoa0901867. [DOI] [PubMed] [Google Scholar]
- 7.Li X, et al. Genome-wide association study of asthma identifies RAD50-IL13 and HLA-DR/DQ regions. J Allergy Clin Immunol. 2010;125:328–335. doi: 10.1016/j.jaci.2009.11.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mathias RA, et al. A genome-wide association study on African-ancestry populations for asthma. J Allergy Clin Immunol. 2010;125:336–346. doi: 10.1016/j.jaci.2009.08.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Moffatt MF, et al. A large-scale, consortium-based genomewide association study of asthma. N Engl J Med. 2010;363:1211–1221. doi: 10.1056/NEJMoa0906312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.He JQ, et al. A thymic stromal lymphopoietin gene variant is associated with asthma and airway hyperresponsiveness. J Allergy Clin Immunol. 2009;124:222–229. doi: 10.1016/j.jaci.2009.04.018. [DOI] [PubMed] [Google Scholar]
- 11.Moffatt MF, et al. Association between quantitative traits underlying asthma and the HLA-DRB1 locus in a family-based population sample. Eur J Hum Genet. 2001;9:341–346. doi: 10.1038/sj.ejhg.5200636. [DOI] [PubMed] [Google Scholar]
- 12.Repapi E, et al. Genome-wide association study identifies five loci associated with lung function. Nat Genet. 2010;42:36–44. doi: 10.1038/ng.501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hancock DB, et al. Meta-analyses of genome-wide association studies identify multiple loci associated with pulmonary function. Nat Genet. 2010;42:45–52. doi: 10.1038/ng.500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Weiss ST. Lung function and airway diseases. Nat Genet. 2010;42:14–16. doi: 10.1038/ng0110-14. [DOI] [PubMed] [Google Scholar]
- 15.Ziegler SF, Artis D. Sensing the outside world: TSLP regulates barrier immunity. Nat Immunol. 2010;11:289–293. doi: 10.1038/ni.1852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ho IC, et al. GATA3 and the T-cell lineage: essential functions before and after T-helper-2-cell differentiation. Nat Rev Immunol. 2009;9:125–135. doi: 10.1038/nri2476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wasserman NF, et al. An 8q24 gene desert variant associated with prostate cancer risk confers differential in vivo activity to a MYC enhancer. Genome Res. 2010;20:1191–1197. doi: 10.1101/gr.105361.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hakonarson H, et al. A novel susceptibility locus for type 1 diabetes on Chr12q13 identified by a genome-wide association study. Diabetes. 2008;57:1143–1146. doi: 10.2337/db07-1305. [DOI] [PubMed] [Google Scholar]
- 19.Petukhova L, et al. Genome-wide association study in alopecia areata implicates both innate and adaptive immunity. Nature. 2010;466:113–117. doi: 10.1038/nature09114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pan F, et al. Eos mediates Foxp3-dependent gene silencing in CD4+ regulatory T cells. Science. 2009;325:1142–1146. doi: 10.1126/science.1176077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lambrecht BN, Hammad H. The role of dendritic and epithelial cells as master regulators of allergic airway inflammation. Lancet. 2010;376:835–843. doi: 10.1016/S0140-6736(10)61226-3. [DOI] [PubMed] [Google Scholar]
- 22.Sármay G, et al. The multiple function of Grb2 associated binder (Gab) adaptor/scaffolding protein in immune cell signaling. Immunol Lett. 2006;104:76–82. doi: 10.1016/j.imlet.2005.11.017. [DOI] [PubMed] [Google Scholar]
- 23.Harada M, et al. A functional polymorphism in IL-18 is associated with severity of bronchial asthma. Am J Respir Crit Care Med. 2009;180:1048–1055. doi: 10.1164/rccm.200905-0652OC. [DOI] [PubMed] [Google Scholar]
- 24.Nakamura Y. The BioBank Japan Project. Clin Adv Hematol Oncol. 2007;5:696–697. [PubMed] [Google Scholar]
- 25.Takata R, et al. Genome-wide association study identifies five new susceptibility loci for prostate cancer in the Japanese population. Nat Genet. 2010;42:751–754. doi: 10.1038/ng.635. [DOI] [PubMed] [Google Scholar]
- 26.Silverman ES, et al. Transforming growth factor-beta1 promoter polymorphism C-509T is associated with asthma. Am J Respir Crit Care Med. 2004;169:214–219. doi: 10.1164/rccm.200307-973OC. [DOI] [PubMed] [Google Scholar]
- 27.Silverman ES, et al. Constitutive and cytokine-induced expression of the ETS transcription factor ESE-3 in the lung. Am J Respir Cell Mol Biol. 2002;27:697–704. doi: 10.1165/rcmb.2002-0011OC. [DOI] [PubMed] [Google Scholar]
- 28.Peters SP, et al. Randomized comparison of strategies for reducing treatment in mild persistent asthma. N Engl J Med. 2007;356:2027–2039. doi: 10.1056/NEJMoa070013. [DOI] [PubMed] [Google Scholar]
- 29.Luca D, et al. On the use of general control samples for genome-wide association studies: genetic matching highlights causal variants. Am J Hum Genet. 2008;82:453–463. doi: 10.1016/j.ajhg.2007.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Purcell S, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Purcell S, et al. Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics. 2003;19:149–150. doi: 10.1093/bioinformatics/19.1.149. [DOI] [PubMed] [Google Scholar]
- 32.Price AL, et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 2006;38:904–909. doi: 10.1038/ng1847. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


