Abstract
Adenosine deaminases acting on RNA (ADARs) are involved in adenosine-to-inosine RNA editing and are implicated in development and diseases. Here we observed that ADAR1 deficiency in human embryonic stem cells (hESCs) significantly affected hESC differentiation and neural induction with widespread changes in mRNA and miRNA expression, including upregulation of self-renewal-related miRNAs, such as miR302s. Global editing analyses revealed that ADAR1 editing activity contributes little to the altered miRNA/mRNA expression in ADAR1-deficient hESCs upon neural induction. Genome-wide iCLIP studies identified that ADAR1 binds directly to pri-miRNAs to interfere with miRNA processing by acting as an RNA-binding protein. Importantly, aberrant expression of miRNAs and phenotypes observed in ADAR1-depleted hESCs upon neural differentiation could be reversed by an enzymatically inactive ADAR1 mutant, but not by the RNA-binding-null ADAR1 mutant. These findings reveal that ADAR1, but not its editing activity, is critical for hESC differentiation and neural induction by regulating miRNA biogenesis via direct RNA interaction.
Keywords: ADAR1, RNA editing, embryonic stem cell, neural induction, miRNA
Introduction
Adenosine deaminases acting on RNA (ADARs) catalyze the chemical conversion of adenosine (A) to inosine (I) in double-stranded RNA (dsRNA) substrates. As inosines preferentially base pair with cytidines (C), they change the primary sequence of RNA and are interpreted as guanosine (G) by the translational machinery1,2. Three ADAR gene family members (ADAR1-3) are found in mammals. ADAR1 and ADAR2 are widely expressed and catalytically active in mammals, while ADAR3 is considered to be inactive2.
ADARs interact with several types of dsRNA substrates. Short and imperfect RNA duplexes within pre-mRNAs can be site-specifically edited, leading to recoding of open reading frames or alterations in the pattern of splicing3,4,5,6,7. A-to-I editing events occur more frequently in noncoding sequences including repetitive elements and small RNA precursors. Alu and LINE elements, when positioned as sense and antisense pairs nearby one another in the genome, exhibit the potential to form long dsRNAs, and these dsRNAs can be hyper-edited at multiple adenosines8,9,10. Several studies suggest that the hyper-editing in repetitive elements is involved in the regulation of gene expression11,12,13. Primary miRNA transcripts fold into imperfect dsRNAs that are cleaved by DROSHA-DGCR8 into 60-70-nt long pre-miRNAs and subsequently exported to the cytoplasm and processed by Dicer to form mature miRNAs14. Editing of miRNA precursors can interfere with miRNA biogenesis15,16 or alter target specificity of edited mature miRNAs17,18.
Besides the catalytic activity of ADARs on miRNAs, it has been shown that ADARs can modulate the miRNA/siRNA pathways independently of the editing activity in fly19. ADARs are also important for maintaining many small RNAs in C. elegans through both editing and non-editing activities20. In addition, lack of ADARs in mouse embryos led to a general expansion of the miRNA repertoire, which seems unrelated to editing events18. However, the detailed mechanism(s) by which the non-catalytic activity of ADAR1 acts in regulating miRNA processing has remained elusive. Recently, two research groups showed that ADAR1 interacted with different components of the miRNA biogenesis pathway, and exerted different effects on miRNA production21,22. While Nemlich et al.21 showed that ADAR1 could interact with nuclear DGCR8, which would be mutually exclusive with the DROSHA-DGCR8 interaction and subsequently inhibit miRNA processing, Ota et al.22 showed that ADAR1 could interact with Dicer in the cytoplasm to promote miRNA processing in differentiated cells. Although these studies clearly suggest that the non-catalytic activity of ADAR1 and the miRNA biogenesis pathway are connected at multiple levels, the different results also strongly argue that the underlying mechanisms still require further investigation.
ADAR1 is an essential protein. ADAR1-deficient mice were embryonically lethal, and showed rapid liver disintegration and hematopoietic defects due to widespread apoptosis, which led to the notion that ADAR1 plays critical roles in the development of non-nervous tissues23,24. Moreover, hematopoietic lineage-specific ADAR1 knockout revealed that ADAR1 played an essential role in hematopoietic stem cell maintenance25; however, the detailed mechanism underlying embryonic lethality caused by ADAR1 deficiency has remained elusive. In addition, while ADAR1-deficient mice exhibited little aberrant neuronal phenotype prior to embryonic lethality23,24, mutations in human ADAR1 gene were shown to be associated with Aicardi-Goutières syndrome26, an early-onset encephalopathy that often results in severe and permanent neurological damage, indicating that ADAR1 may play an important role during neural development in humans.
Inner cell mass- or epiblast-derived embryonic stem cells (ESCs) can differentiate into cells/tissues of the three germ layers according to developmental principles27,28,29. Differentiation of human ESCs (hESCs) thus provides a way to investigate the regulatory mechanisms that facilitate cellular transitions in a human context30. We generated hESCs lacking ADAR1 and examined their ability to differentiate into specific types of neurons, followed by RNA-seq to systematically compare mRNA and miRNA changes between wild-type (WT) and ADAR1-deficient cells at several differentiation time points. While ADAR1 deficiency had little effects on hESC pluripotency, it affected embryoid body (EB) formation and neural induction. This strong phenotype was largely due to the altered expression of self-renewal and neural induction suppression-related miRNAs after disruption of ADAR1. Strikingly, our data revealed that ADAR1 could act as an RNA-binding protein to directly interfere with pri-miRNA processing and that the catalytically inert ADAR1 mutant, but not the RNA-binding-null ADAR1 mutant, rescued the observed differentiation defects.
Results
ADAR1 knockdown has little effect on hESC pluripotency
To determine the effect of ADAR1 on differentiation in a human cell culture model, we knocked down ADAR1 in H9 hESCs using lentiviral transduction. We used constructs that either constitutively expressed different small hairpin RNAs (shRNAs) directed against ADAR1 mRNA, or a mismatch shRNA as a scramble control (Supplementary information, Figure S1A). Both constructs also co-expressed GFP. GFP-positive hESC colonies were manually passaged to isolate individual subclones to form stable cell lines (Supplementary information, Figure S1B). High ADAR1 knockdown efficiency was achieved following different shRNA treatments (Supplementary information, Figure S1A) as revealed by subsequent RNA-seq analyses (Supplementary information, Table S1) and immunoblotting (Figure 1A). In addition, we found that several known ADAR1 substrates11 exhibited a significant reduction of A-to-I RNA editing level after ADAR1 knockdown (Supplementary information, Figure S1C), further confirming the successful generation of ADAR1-knockdown H9 cell lines. Results shown in the following analyses were mainly obtained from a stable ADAR1-knockdown cell line with sh4146 shRNA (Supplementary information, Figure S1A) that targets the 3′ UTR region of ADAR1 (ADAR1 KD hereafter).
Figure 1.
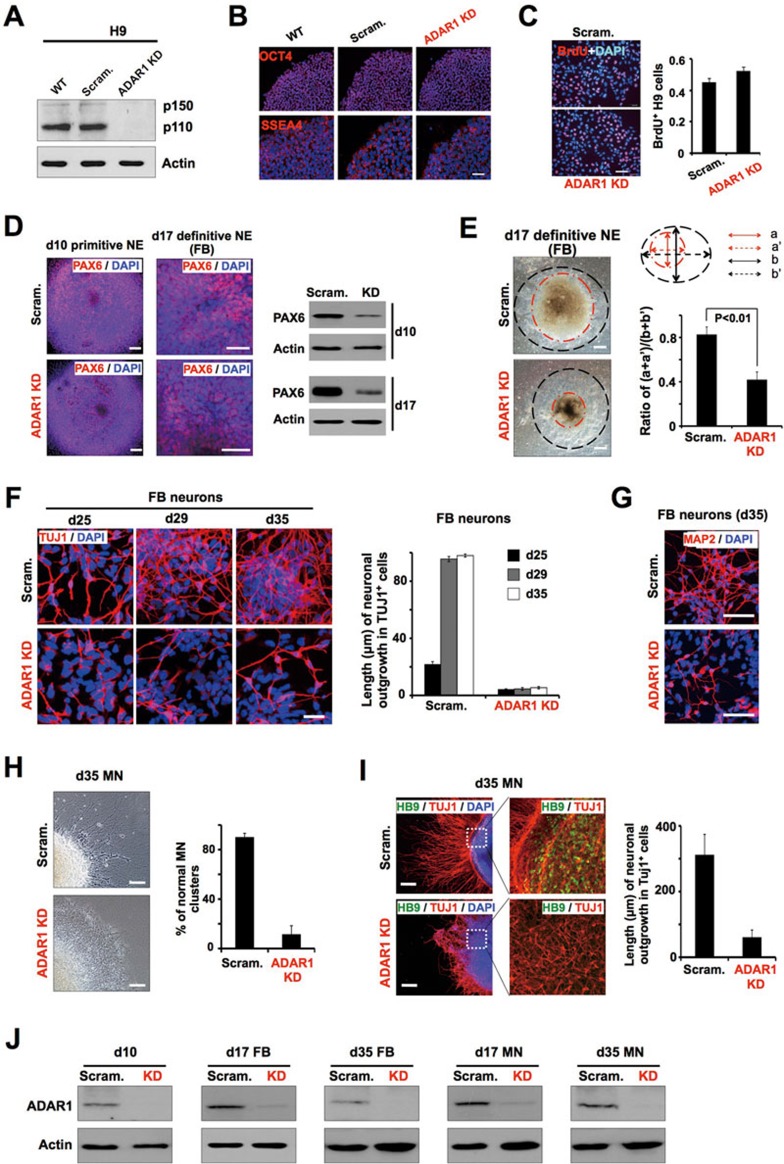
ADAR1 knockdown significantly represses neural induction and neuronal differentiation. (A) Generation of ADAR1 KD H9 stable cell lines. Western blotting (WB) confirms the stable knockdown of ADAR1 by shRNA targeting ADAR1 in H9 cells, but not in scramble shRNA-treated H9 cells (Scram). Actin was used as a loading control. (B) Loss of ADAR1 has little effect on pluripotency of hESCs. WT, scramble shRNA-treated (Scram) and ADAR1 KD H9 cells were stained with anti-Oct4 and anti-SSEA4. Nuclei were counterstained with DAPI. Scale bar, 50 μm. (C) Loss of ADAR1 has no significant effect on proliferation of hESCs. Left, representative images of proliferating cells with BrdU incorporation in scramble control and ADAR1 KD H9 cells. Right, statistical analyses for BrdU+ cells. Scale bar, 50 μm. (D) Columnar NE cells derived from ADAR1 KD H9 cells express a low level of PAX6. Immunostaining (left) and WB (right) show the expression of PAX6 in scramble control and ADAR1 KD H9 lines at neural induction d10 and d17. Scale bar, 50 μm. (E) The generation of definitive NE cells from ADAR1 KD H9 cells is less efficient than that from control cells. Left, representative images at d17 neural induction of scramble control and ADAR1 KD H9 cells. After EB attachment, columnar NE cells appear in the colony center at d10 and proliferate quickly and form multiple layers (red dotted line), whereas cells in the periphery gradually become flattened (between black and red dotted line). Right, a and a′ represent the diameter of the NE cluster, and b and b′ represent the diameter of the differentiating colony. Ratio of (a+a′)/(b+b′) indicated the approximate percentage of NE cells in the colony. P value from one-tail t-test is shown. (F, G) ADAR1 knockdown represses the formation of human FB neurons. Left, representative images of d25-d35 neural induction of scramble control and ADAR1 KD H9 cells. Cells were stained with anti-Tuj1. Right, the length of neurites in Tuj1+ cells was measured after the indicated differentiation days of control and ADAR1 KD H9 cells towards FB neuron differentiation (F). Neural induction of control and ADAR1 KD hESCs at d35 was shown by MAP2 staining (G). Scale bar, 50 μm. (H-I) ADAR1 knockdown retards the formation of human MNs. Left, representative images showing the repression of the formation of MN clusters derived from ADAR1 KD H9 cells. Right, the percentage of MN clusters derived from scramble control and ADAR1 KD H9 cells (H). Left, representative images showing the retardation of MN formation. Cells were stained with anti-Tuj1 and anti-HB9. Right, the length of neurites in Tuj1-positive cells was measured in MNs (d35) derived from control and ADAR1 KD H9 cells (I). (J) ADAR1 expression was maintained at very low level under ADAR1 knockdown condition upon neural induction and neuronal specification in H9 cells, as revealed by WB. Actin was used as a loading control. In C, E, F, H and I, error bars represent ± SD of triplicate experiments. Also see Supplementary information, Figures S1 and S2 and Table S1.
The undifferentiated H9 cells lacking ADAR1 appeared morphologically normal and stained positively for pluripotency markers of OCT4 and SSEA4 (Figure 1B). Expression of other known pluripotency genes was largely unchanged and the expression of marker genes of three germ layers remained undetectable (Supplementary information, Figure S1D and Table S2). In addition, the proliferation of hESCs remained largely unaltered after knockdown of ADAR1 (Figure 1C). Meanwhile, ADAR1 knockdown did not affect the ability of hESCs to differentiate into three germ layers in our teratoma assay (Supplementary information, Figure S1E), although the size of ADAR1 KD teratoma is reduced (data not shown). These observations suggested that ADAR1 knockdown had little effect on pluripotency and self-renewal. However, we have noticed that the clone formation efficiency was lower in ADAR1-knockdown H9 cells than that in control cells (Supplementary information, Figure S1F). RNA-seq and RT-qPCR results further revealed that some key pro-apoptotic genes, such as S100A6 and BAX31,32,33, were upregulated and the anti-apoptotic gene SFRP134,35 was downregulated in ADAR1-knockdown H9 cells (Supplementary information, Figure S1G and Table S2), suggesting that ADAR1 deficiency can promote apoptosis of hESCs under certain stresses, such as enzymatic detachment. This is consistent with a previous report that cells cultured from ADAR1-deficient embryos exhibited increased apoptosis when a stress response was induced24.
hESCs lacking ADAR1 exhibited significant defects in EB formation, neural induction and neuronal differentiation
As ADAR1 knockdown has little effect on pluripotency, we then proceeded to study the effect of ADAR1 deficiency on differentiation with a protocol summarized in Supplementary information, Figure S2A36,37,38. Briefly, hESCs were induced into the neural lineage by generating EBs followed by culturing in neural medium. We observed that about 30% more irregular EBs were formed in ADAR1-knockdown H9 lines than in controls during neural induction (Supplementary information, Figure S2B), suggesting that the general differentiation was affected by ADAR1 deficiency.
Morphologically normal EBs were manually collected and cultured in neural medium for subsequent differentiation. At 10 and 17 days of neural differentiation, hESC-derived primitive and definitive neuroepthelial (NE) cells with scramble shRNA treatment presented typical columnar NE morphology and were organized into rosettes36,37,38 (Figure 1D and 1E), similar to the structure of NE cells from WT hESC induction (as discussed below and data not shown). Noticeably, although differentiating hESCs with stable ADAR1 knockdown also formed rosette-like structures, these cells expressed low levels of PAX6, SOX1 and FOXG1 (Figure 1D and Supplementary information, Table S1), and formed much smaller round aggregates at d17 (Figure 1E). Consistent results were obtained with different lines (Supplementary information, Figure S2A) and different batches of differentiation (data not shown), indicating that the knockdown phenotype was not due to asynchronized differentiation or different viral integration. As PAX6 is a human neuroectoderm cell fate determinant and its expression reaches a peak on day 10 after differentiation38, our observation suggests that neural induction was retarded by ADAR1 knockdown.
To further investigate the fates of these NE cells produced from ADAR1 KD H9 cells, we differentiated NE cells into two types of neurons, forebrain (FB) neurons or spinal motor neurons (MNs). FB cells were generated using a well-established system without any caudalizing factors as described39,40. Strikingly, the formation of TUJ1- (Figure 1F) and MAP2-positive FB neurons (Figure 1G) was dramatically reduced from ADAR1 KD H9 cells. In addition, the TUJ1-positive FB cells derived from ADAR1 KD H9 cells exhibited much shorter growing neurites at all differentiation stages (Figure 1F). Meanwhile, although the differentiation efficiency of MNs was only about 30%-40% as reported previously37, we found that ADAR1 knockdown also dramatically repressed the formation of MN clusters (Figure 1H) and HB9-positive MNs (Figure 1I). The resulting MNs also exhibited a significant reduction of neurite outgrowth (Figure 1I). Furthermore, although differentiation of these NE cells into FB neurons and MNs was significantly retarded, they did not differentiate into midbrain or hindbrain neurons (Supplementary information, Figure S2C). Importantly, the ADAR1 expression in ADAR1 KD cells remained consistently low levels during all differentiation stages examined, as revealed by both protein (Figure 1J) and RNA (Supplementary information, Table S1) levels. Taken together, these results revealed that ADAR1 deficiency not only inhibited hESC differentiation into neural lineages, but also retarded the formation of different neuron types.
Repression of neuron-specific gene expression in ADAR1 KD H9 cells during differentiation
We have presented strong evidence that the deficiency of ADAR1 significantly repressed human neural induction (Figure 1). To understand the underlying mechanisms by which ADAR1 acts in this process, we performed a series of genome-wide transcriptome comparisons (Figure 2A and Supplementary information, Figure S3A) between the control (WT) and ADAR1 KD H9 cell lines at several landmark time points (d0, d10, d17 and d35) along FB and MN differentiation pathways (Supplementary information, Figure S3A). Such analyses not only allowed parallel and dynamic comparisons of gene expression profiles between ADAR1 KD and control cells, but also provided a rich source to analyze A-to-I RNA editing events catalyzed by ADAR1 upon hESC differentiation.
Figure 2.
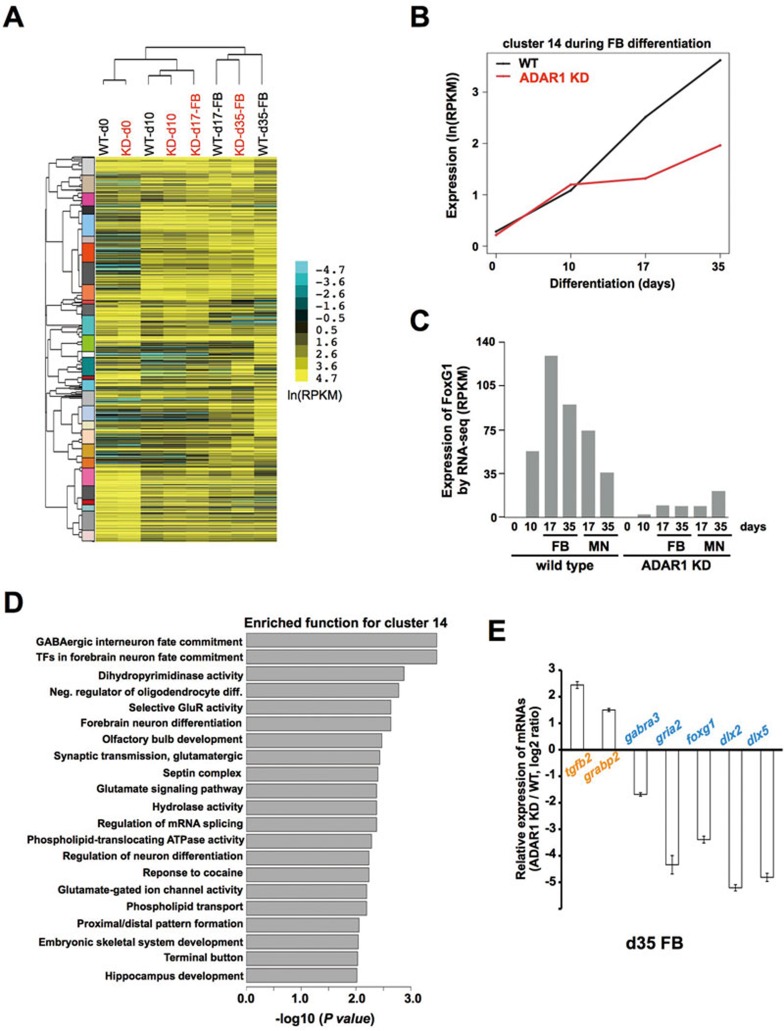
Effects of ADAR1 knockdown on gene expression in H9 cells upon FB neuron differentiation. (A) Repression of neuron-specific gene expression in ADAR1 KD H9 cells upon FB neuron differentiation. Heat map showing the hierarchical clustering of expression profiles of differentially expressed genes (DEGs) as measured by RNA-seq. Pairwise distances were measured by Pearson correlation coefficient (PCC). Different gene clusters are labeled beside the gene tree with different colors. The data shown in the maps are logarithm transformed expression values (ln(RPKM)). (B) ADAR1 KD hESCs exhibit clusters of repressed neuronal gene expression upon FB neuron differentiation. A representative cluster (#14) containing 102 genes was significantly repressed in ADAR1 KD H9 cells after the induction of NE (d10; P = 1.9e-7 for d17, or P < 2.2e-16 for d35, Student's t-test). (C) Histograph of RNA-seq results showing that FoxG1 was repressed during hESC differentiation into neurons with ADAR1 knockdown. (D) Cluster 14 was enriched for genes with neuronal functions. The enrichment P value cut-off is set to a significance value of 0.01 after Benjamini-Hochberg correction. (E) Verification of RNA-seq using RT-qPCR analysis. Error bars represent ± SD of triplicate experiments. Also see Supplementary information, Figure S3, Tables S1 and S2.
Hierarchical clustering of gene expression profiles (Supplementary information, Table S1) showed that WT and ADAR1 KD H9 cells were generally clustered together at the same differentiation time points, yet distinguished from those at other differentiation time points towards either FB neuron (Figure 2A) or MN (Supplementary information, Figure S3A). However, noticeable exceptions were observed for d17 and d35 FB neurons in ADAR1 KD H9 cells (Figure 2A). The sample from d17 “definitive NEs” derived from ADAR1 KD H9 cells was clustered with d10 primitive NEs derived from ADAR1 KD and WT cells. The sample from d35 “FB neurons” derived from ADAR1 KD H9 cells was clustered with d17 definitive NEs derived from WT cells (Figure 2A). Importantly, many of the altered genes have been reported to be important factors involved in neural development and neurogenesis (Figure 2B-2D, Supplementary information, Table S1 and Figure S3). Such altered gene expression was further confirmed by semi-quantitative RT-PCR (data not shown) and RT-qPCR (Figure 2E) in samples collected from independent differentiation experiments. Together, these altered gene expression signatures clearly suggest a delayed neural differentiation after human ADAR1 expression was repressed in hESCs as we have observed (Figure 1 and Supplementary information, Figure S2).
When ADAR1 KD H9 cells were differentiated to MNs, a delayed gene expression pattern towards this lineage differentiation was also observed (Supplementary information, Figure S3A). Importantly, consistent with the aberrant MN formation from ADAR1 KD H9 cells (Figure 1H and 1I), genes related with MNs and their functions were also clustered (Supplementary information, Figure S3B-S3D). Given that the differentiation efficiency of MNs is only about 30%-40%37,41, we focused on the differentiation to FB neurons for our subsequent analyses.
ADAR1 knockdown represses A-to-I RNA editing in general, but such altered editing sites do not correspond to the expression of genes related with neural induction and neurogenesis
We next asked what contributed to the retarded gene expression upon neural differentiation after knockdown of ADAR1 in H9 cells. ADAR1 is known for its A-to-I RNA editing activity. To our surprise, the most altered genes associated with neuron function (Figure 2B) were not shown to have significant A-to-I ratio changes between ADAR1 KD and WT cells (data not shown). We then took advantage of the RNA-seq datasets to genome-widely profile A-to-I RNA editing and their corresponding mRNA changes during neural differentiation under WT and ADAR1 KD conditions. Although the total number of high-confidence A-to-I RNA editing sites was significantly reduced in all stages of differentiations after depletion of ADAR1 (Supplementary information, Figure S4A and data not shown), the majority of selected high-confidence editing events were largely located in non-exon regions (43% in RefSeq introns, 30% in non-RefSeq regions, and 26% in RefSeq exons, Supplementary information, Figure S4B). Among those in RefSeq exons, only 68 A-to-I editing sites were located in coding exons (Supplementary information, Figure S4B and Table S3). However, expression of their host genes of these 68 A-to-I editing sites remained largely unchanged after ADAR1 knockdown (Supplementary information, Figure S4C, S4D and Table S1), suggesting that the editing activity of ADAR1 on mRNAs did not contribute to the observed retarded gene expression upon neural differentiation in ADAR1 KD hESCs.
ADAR1 knockdown dramatically alters miRNA repertoires
Our data revealed that RNA editing has little correlation with mRNA expression changes in ADAR1 KD H9 cells (Supplementary information, Figure S4 and data not shown), which is in consistency with recent reports in other organisms18,19,20. We hypothesized that ADAR1 deficiency may lead to an altered miRNA repertoire, which in turn regulates mRNA expression in H9 cells and during differentiation. We therefore performed small RNA-seq from the same aliquots used for polyadenylated RNA-seq described in Figure 2. This allowed us to perform not only an analysis of miRNA expression profiles, but also a comprehensive network analysis between miRNA and mRNA transcriptomes after ADAR1 knockdown (Figure 3A).
Figure 3.
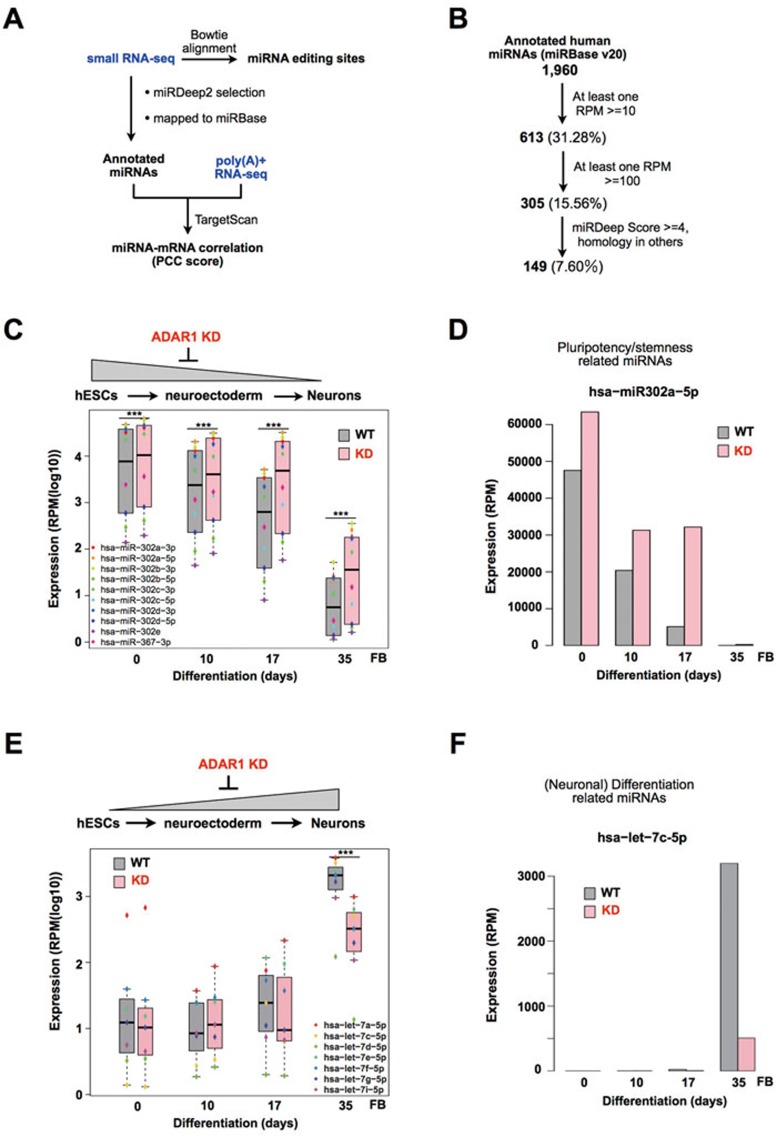
ADAR1 knockdown alteres miRNA expression. (A) A schematic depiction of miRNA analysis. Known miRNA A-to-I editing sites42 were first examined from our small RNA-seq data sets. miRNA expression was further evaluated with miRDeep2. Predicted regulatory targets of all highly expressed miRNAs were retrieved by TargetScan. The correlation of miRNA and mRNA expression was calculated by PCC using the paired RNA-seq and small RNA-seq data sets (Supplementary information, Table S4). (B) Selection of highly expressed miRNAs. About 1 960 known miRNAs were retrieved with default miRDeep2 pipeline. 305 highly expressed (≥ 100 RPM) known miRNAs (Supplementary information, Table S5) were subjected to further analysis. (C) Downregulation of the miR302 cluster is delayed in ADAR1 KD H9 cells upon differentiation into FB neurons. Each colored dot represents the expression level of a corresponding miR302 family miRNA (P = 4.5e-8 for d0, 2.4e-7 for d10, 1.4e-11 for d17 or 1.8e-4 for d35, pairwise t-test). (D) Expression of hsa-miR-302a-5p at differentiation time points examined in WT and ADAR1 KD H9 cells. (E) Upregulation of the let-7 cluster expression is repressed in ADAR1 KD H9 cells upon differentiation to FB neurons. Each colored dot represents the expression level of a corresponding let-7 family miRNA (P = 1.2e-5 for d35, pairwise t-test). (F) Expression of hsa-let-7c-5p at differentiation time points examined in WT and ADAR1 KD H9 cells. Also see Supplementary information, Figure S4.
In total, we obtained 5-12 million mapped small RNA-seq reads (Supplementary information, Table S4). By analyzing the reported A-to-I sites in mature miRNAs42, we found that the occurrence of RNA editing was either very low (6/19) or undetectable (data not shown), and the expression of host miRNAs remained unchanged after ADAR1 knockdown. This suggests that the frequency of A-to-I editing in mature miRNAs and the probability of re-targeting of mRNAs were limited in undifferentiated and differentiated hESCs.
To further evaluate the role of ADAR1 in miRNA production, we applied miRDeep2 to annotate both known miRNAs from miRBase (v20) and previously unreported miRNA candidates. Using the default miRDeep2 pipeline, 1 960 known miRBase (v20) miRNAs were identified (Figure 3B). With the expression cutoff at 100 RPM, about 16% (305 out of 1 960, Supplementary information, Table S5) of known miRNAs were selected for further analysis. Importantly, our data revealed that the expression of some key miRNAs was altered after ADAR1 knockdown and these could be categorized into two distinct groups according to their expression patterns.
The first group of miRNAs was reported to be highly expressed in undifferentiated hESCs but significantly downregulated upon differentiation, such as miR302s43,44,45. In our studies, these miRNAs were upregulated in undifferentiated H9 cells after ADAR1 knockdown. Upon neural induction, their expression exhibited a delayed downregulation in ADAR1 KD cells compared to WT cells (Figure 3C, 3D and Supplementary information, Figure S5A and Table S5). In addition, the repression on miR302s by ADAR1 was specific, as the expression of other miRNAs was not altered after ADAR1 depletion (Supplementary information, Figure S5B). It is well known that self-renewal-related miR-302s promote ESC proliferation and suppress differentiation45,46; thus the delayed downregulation of miR-302s upon ADAR1 depletion could lead to the suppression of hESC differentiation. Furthermore, some miRNAs, such as miR-96 and miR-200s, that repress neural induction36, were also upregulated in ADAR1 KD samples (Supplementary information, Table S5). This is in agreement with the notion that ADAR1 knockdown resulted in delayed neural induction (Figure 1).
The other group contains miRNAs such as let-7s, which are known to be highly expressed in differentiated cells47,48. In WT H9 cells, the expression of these miRNAs was low or undetectable in early stages of differentiation, but significantly upregulated in later stages of differentiation. Strikingly, their expression was remarkably repressed in ADAR1 KD cells upon neural induction (Figure 3E, 3F and Supplementary information, Table S5). However, as we observed reduced expression levels of transcription factors, such as PAX6, in day-10 NE cells (Figure 1D), whether this cluster of altered miRNAs was a cause or a consequence of neural differentiation inhibition by ADAR1 knockdown requires further investigation. Nevertheless, the downregulation of let-7 family expression after ADAR1 knockdown is consistent with a recent report that ADAR1 forms a complex with DICER to promote let-7 miRNA processing22.
ADAR1 knockdown promotes processing of pri-miR302s to mature miR302s
It is worth noting that knockdown of ADAR1 affected self-renewal-related miRNAs, such as miR302s, in undifferentiated H9 cells (Figure 3C and 3D). It is thus important to decipher how ADAR1 regulates miR302 maturation. We first excluded the role of transcriptional regulation of ADAR1 on miRNA processing. On one hand, loss of ADAR1 had no effect on the expression of oct4, sox2, nanog and rex1 (Supplementary information, Figure S1D and Table S1), which are known as upstream regulators of the pri-miR302 promoter45, excluding the possibility that the upregulation of miR302s is caused by upregulation of ESC-specific transcriptional factors in ADAR1 KD H9 cells. On the other hand, we found that knockdown of ADAR1 had no significant effect on the promoter activity of pri-miR302 (Supplementary information, Figure S5C), suggesting that ADAR1 is unlikely to be involved in the transcriptional regulation of pri-miR302. Furthermore, Sanger sequencing of precursors of these altered miRNAs (such as pri-miR302s, pri-miR-200, pri-miR-96, etc.) revealed that none of the examined transcripts was edited in undifferentiated H9 cells (data not shown), indicating that the editing activity of ADAR1 plays a limited role in the maturation of these miRNAs in hESCs.
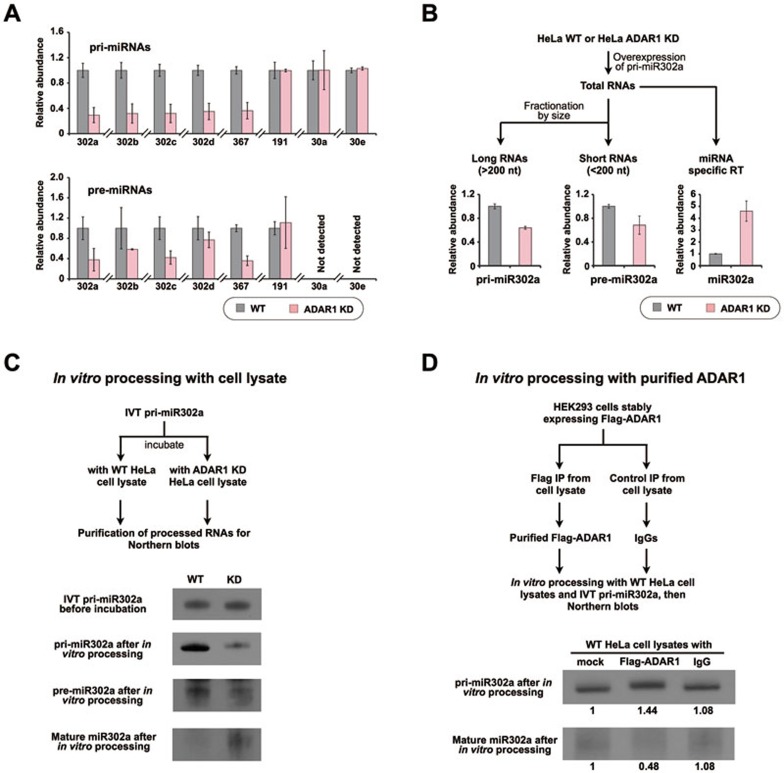
Then, at which step ADAR1 could interfere with miR302 processing? We found that both pri- and pre-miR302s were significantly reduced after ADAR1 knockdown in H9 cells (Figure 4A), suggesting that ADAR1 regulates the initial processing of pri-miR302s. Meanwhile, the abundance of other unaffected miRNAs, such as miR30a and miR30e, remained unaltered at their pri-/pre-miRNA levels after ADAR1 knockdown, indicating that the regulatory effect of ADAR1 on miR302s is specific. To further confirm the notion that ADAR1 can inhibit processing of pri-miR302s, we expressed pri-miR302a in either WT or ADAR1 KD HeLa cells, in which the expression of endogenous miR302s was undetectable. While the mature miR302a produced from exogenous pri-miR302 was significantly increased in ADAR1 KD HeLa cells, the amounts of pri- and pre-miR302a were reduced in ADAR1 KD HeLa cells, compared with those in WT HeLa cells (Figure 4B). Similar results were also obtained from the in vitro processing assays with in vitro transcribed (IVT) pri-miR302a. We observed that the lack of ADAR1 in the cell lysate promoted the processing of pri-miR302a to miR302a (Figure 4C). Consistently, addition of the partially purified Flag-ADAR1 inhibited pri-miR302a processing to miR302a (Figure 4D). Together, these assays confirmed that ADAR1 could inhibit pri-miR302 processing by its non-enzymatic activity in hESCs.
Figure 4.
ADAR1 knockdown promotes the processing from pri-miR302s to mature miR302s. (A) Knockdown of ADAR1 led to reduced expression of pri-miR302s (top, normalized to gapdh mRNA) and pre-miR302s (bottom, normalized to U6), but not miR191, miR30a and miR30e. Error bars represent ± SD of triplicate experiments. (B) Pri-miR302a processing in WT or ADAR1 KD HeLa cells. Long (> 200 nt) pri- and short (< 200 nt) pre-miRNAs were fractionated according to their sizes and were then used for RT-qPCR to quantify the relative abundance of pri- or pre-miR302a. Mature miRNA was measured from total RNAs with miScript II RT-qPCR. Error bars represent ± SD of triplicate experiments. (C) ADAR1 knockdown promotes miR302 maturation. IVT pri-miR302a was incubated with either WT or ADAR1 KD HeLa cell lysates. After incubation, RNAs were extracted for northern blot with denaturing PAGE gels. Pre-mixtures without incubation were loaded as controls. Note that matured miR302a was more abundant after the incubation with ADAR1 KD HeLa cell lysate than that with WT cell lysate. Triplicated experiments were performed and one representative is shown. (D) ADAR1 inhibits pri-miR302 processing. IVT pri-miR302a was incubated with ADAR1 KD HeLa cell lysates in the presence or absence of partially purified Flag-ADAR1 according to the protocol published previously57. After incubation, RNAs were extracted for northern blot with denaturing PAGE gels. Note that the processing of miR302a was repressed with the addition of ADAR1. Triplicated experiments were performed and one representative is shown. The relative abundance of pri-miR302a and mature miR302a was determined using Image J and labeled underneath. Also see Supplementary information, Figure S5.
ADAR1 acts as an RNA-binding protein that directly inhibits processing of pri-miR302s
To gain better insight into the mechanism, we further rescued undifferentiated ADAR1 KD H9 hESCs with WT or mutant ADAR1 proteins (Figure 5A), and performed a series of experiments in these lines to decipher how ADAR1 interferes with miRNA processing pathway independently of its catalytic activity.
Figure 5.
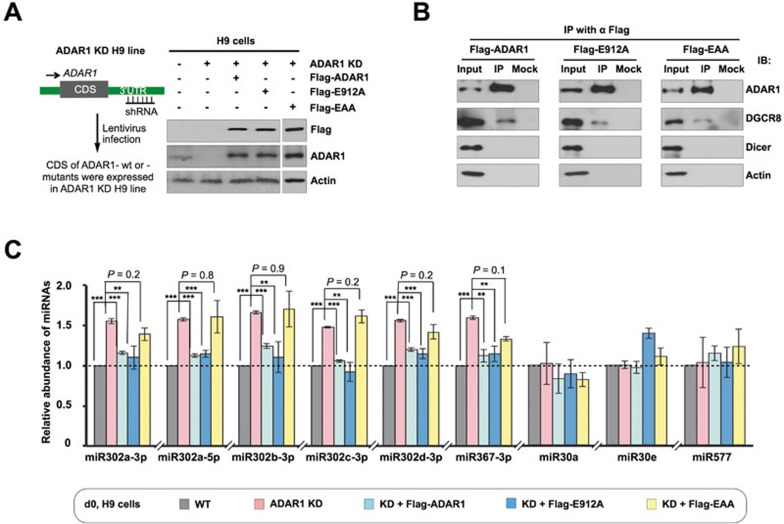
The RNA-binding property of ADAR1 affects miR302 processing from pri-miR302s. (A) Rescue of ADAR1 expression in H9 cells by Flag-ADAR1, Flag-E912A or Flag-EAA. Left, a drawing showing the generation of ADAR1-rescued ADAR1 KD H9 cells by lentiviral infection. Right, WB confirms their expression in ADAR1 KD H9 cells. (B) Immunoprecipitation of Flag-ADAR1, Flag-E912A (catalytically inactive mutant), or Flag-EAA (RNA-binding-null mutant) from HEK293 cells extracts, followed by immunoblotting with ADAR1, DGC8 and DICER antibodies. (C) Flag-E912A, but not Flag-EAA, reversed the upregulation of miR302s in ADAR1 KD H9 cells. RT-qPCR revealed the expression of miR302s, miR30a, miR30e and miR577 (normalized to U6) in control, ADAR1 KD, KD+Flag-ADAR1, KD+Flag-E912A and KD+Flag-EAA H9 lines. Error bars represent ± SD of triplicate experiments. P values from one-tail t-tests are shown. Also see Supplementary information, Figure S5.
It was recently reported that ADAR1 could either bind with DICER22 or DGCR821 to regulate miRNA processing. Interestingly, we found that ADAR1 in hESCs was predominately localized to the nucleus (data not shown) as previously reported49. Consistent with its nuclear localization, ADAR1 interacted with endogenous DGCR8 (Figure 5B), but its interaction with DICER was barely detectable under our experimental conditions. In addition, both the catalytically inactive ADAR1 mutant (E912A)22,50 and the RNA-binding-null ADAR1 mutant (EAA)51 acted in a similar manner in the association with DGCR8 (Figure 5B and data not shown), indicating that these ADAR1 mutations do not affect its association with DGCR8.
While the expression of all key components of the Microprocessor remained unchanged in ADAR1 KD H9 cells (Supplementary information, Table S1), introducing WT ADAR1 or catalytically inactive ADAR1-E912A mutant, but not RNA-binding-null ADAR1-EAA mutant, into ADAR1 KD H9 cells restored the expression of miR302s, but not those unaltered miRNAs, to their normal levels (Figure 5C). These results clearly suggest that catalytically inert ADAR1 plays a key role in the regulation of some self-renewal-related miRNAs in H9 cells and that the RNA-binding activity of ADAR1, rather than its interaction with components of the Microprocessor, is more important for miR302 processing.
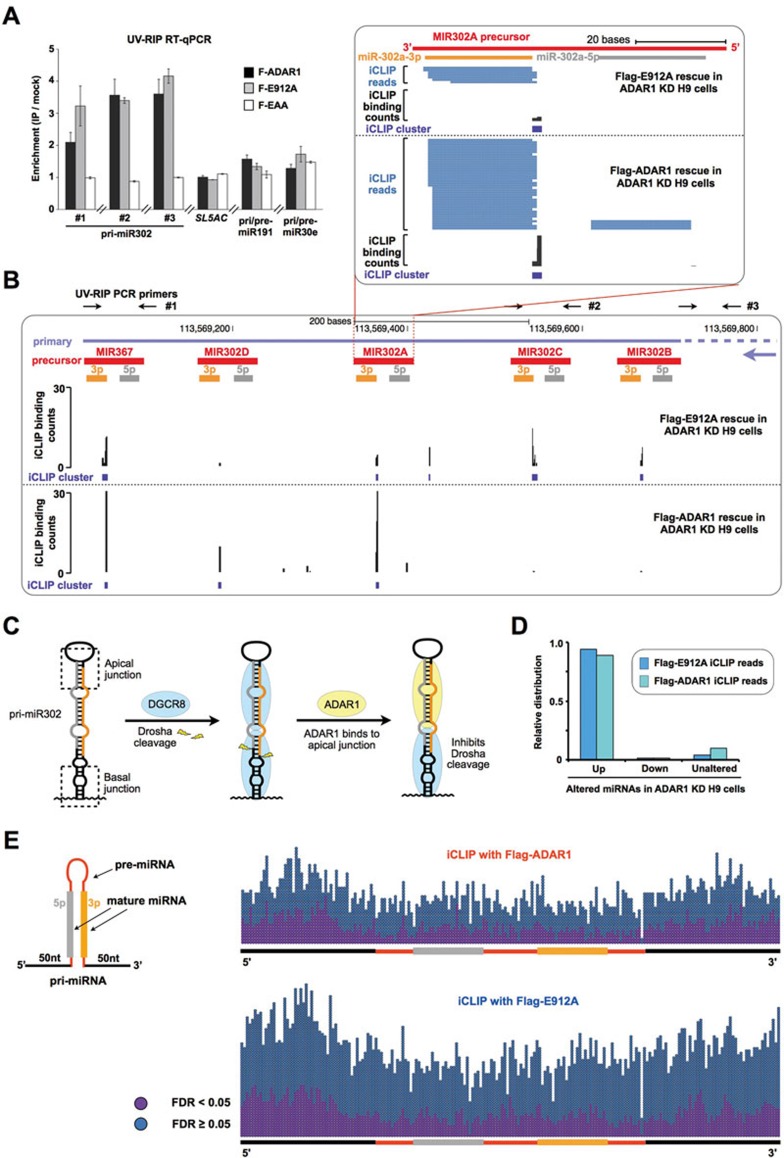
We further identified that both ADAR1-WT and ADAR1-E912A associated with the polycistronic primary transcript of miR302s, while ADAR1-EAA did not (Figure 6A). UV crosslinking and immunoprecipitation coupled with high-throughput sequencing (iCLIP-seq) in H9 cells were then performed to identify ADAR1-binding sites globally (Supplementary information, Figure S5D and S5E). Our iCLIP datasets revealed that both Flag-ADAR1 and Flag-ADAR1-E912A bound directly to multiple apical regions of the hairpin of pri-miR302s (Figure 6B), consistent with the notion that ADAR1 is a dsRNA-binding protein. As the direct binding of DGCR8 to apical junctions of pri-miRNAs is essential for processing and that two DGCR8 dimers clamp a pri-miRNA hairpin using their RNA-binding heme domain to activate DROSHA cleavage52,53 (Figure 6C), the binding of ADAR1 to the same stem region of pri-miR302s suggests that ADAR1 functions as an RNA-binding protein to compete with DGCR8, resulting in an inhibitory effect on miRNA processing in hESCs (Figure 6C). In addition, iCLIP reads were generally enriched in highly expressed primary miRNAs (Supplementary information, Figure S6A, S6B and Table S6), and the majority of reads were shown in the primary transcripts of upregulated miRNAs caused by ADAR1 knockdown (Figure 6D and Supplementary information, Figure S6 and Table S6).
Figure 6.
ADAR1 binds to pri-miR302s and interferes with processing. (A) UV crosslinking RIP confirms that both Flag-ADAR1 and Flag-E912A, but not Flag-EAA, are associated with the polycistronic primary transcript of miR302s. Note that non-altered miRNAs, such as miR191 and miR30e, were not precipitated by ADAR1 proteins. qPCR primers for pri-miR302 detection are labeled as black arrows in B. Error bars represent ± SD of triplicate experiments. (B) iCLIP-seq defines that Flag-ADAR1- and Flag-E912A-binding sites are located within pri-miR302 hairpins. Individual Flag-ADAR1 and Flag-E912A iCLIP-seq reads (in blue) or iCLIP binding counts (in black) were aligned to pri-miR302s, with the mature miRNA boundaries depicted below (orange and gray bars). The iCLIP clusters are depicted as purple rectangles. (C) Two DGCR8 dimers clamp a pri-miRNA hairpin and activate Drosha cleavage (middle)52. The binding of ADAR1 to hairpins of pri-miR302s competes with DGCR8 binding, resulting in an inhibitory effect on pri-miR302 processing (right). (D) The majority of Flag-ADAR1 and Flag-E912A iCLIP reads are located in pri-miRNA regions of the upregulated miRNA targets after ADAR1 knockdown in H9 cells. (E) Flag-ADAR1 and Flag-E912A bind to pri-miRNAs with no sequence specificity. iCLIP clusters, significant (purple points) or non-significant (blue points), were scattered evenly on pri-miRNAs for both Flag-ADAR1 (right panel, top) and Flag-E912A (right panel, bottom). Also see Supplementary information, Figures S5, S6 and Table S6.
Interestingly, the genome-wide analysis of human ADAR1 iCLIP signals also revealed that ADAR1 could bind both miRNA and other annotated RefGene regions (Supplementary information, Figure S6C and S6D). Comprehensive evaluation of ADAR1 binding on pri-miRNAs suggests that there was no sequence specificity for the binding (Figure 6E), and ADAR1 might bind via its RNA-binding domain to structured pri-miRNAs. Such binding could be readily examined in highly expressed pri-miRNAs, such as pri-miR302s (Figure 6B) and pri-miR1260s (Supplementary information, Figure S6A, S6B and Table S6).
Catalytically inactive ADAR1-E912A rescues retarded neural induction and aberrant miRNA expression upon differentiation of ADAR1 KD H9 cells
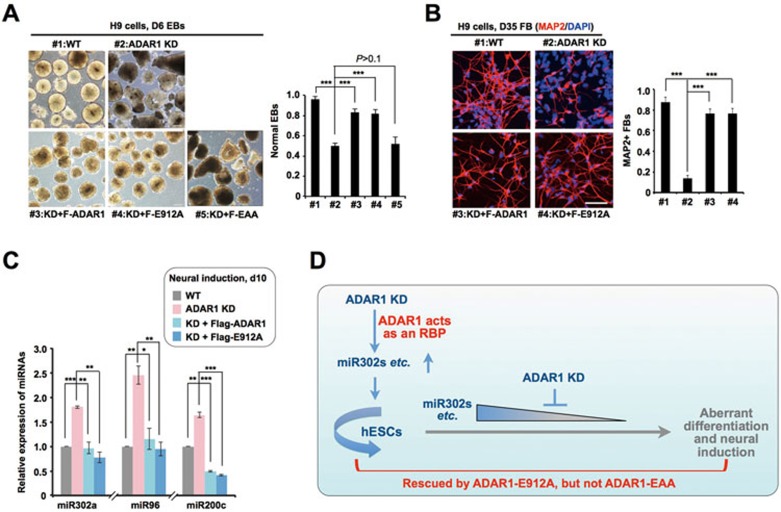
Consistent with the important role of the non-enzymatic ADAR1 acting as an RNA-binding protein in regulating self-renewal-related miRNAs in hESCs (Figures 5 and 6), the catalytically inactive ADAR1-E912A, but not the RNA-binding-deficient ADAR1-EAA, could rescue aberrant EB formation in ADAR1 KD H9 cells (Figure 7A). Importantly, ADAR1-E912A could rescue the phenotype of retarded neural differentiation caused by the loss of ADAR1 (Figure 7B). These observations strongly suggest that the RNA-binding property of ADAR1 is essential for hESC differentiation and neural induction.
Figure 7.
Catalytically inactive Flag-E912A rescues the retarded neural induction and aberrant miRNA expression upon differentiation in ADAR1 KD H9 cells. (A) Flag-E912A, but not Flag-EAA, could reverse the aberrant EB formation derived from ADAR1 KD H9 cells. Left, representative images of EBs derived from different H9 cell lines. Right, the percentage of morphologically normal EBs derived from different H9 cell lines (n > 200). (B) Flag-E912A restored the repressed neural induction in ADAR1 KD H9 cells. Left, representative images of MAP2+ FB neurons derived from different H9 cell lines. Right, the percentage of MAP2+ cells derived from different H9 cell lines (n > 200). (C) Flag-E912A reversed the upregulation of neural induction repression-related miRNAs in ADAR1 KD H9 cells at d10 upon neural induction, as revealed by RT-qPCR (normalized to U6). (D) A model for ADAR1 regulation in hESC self-renewal and differentiation. ADAR1 inhibits the processing of pri-mi302s and other miRNAs in hESCs upon differentiation by acting as a dsRNA-binding protein. In A, B and C, error bars represent ± SD of triplicate experiments. P values from one-tail t-tests are shown.
In addition to miR302s, other miRNAs, such as miR-96 and miR-200 that suppress neural induction, were also upregulated in ADAR1 KD samples (Supplementary information, Table S5). miR-96 and miR-200 families are known to suppress neural differentiation by targeting key neural transcription factors PAX6 and ZEB136. Consistent with their upregulation (Supplementary information, Table S5), we observed that pax6 and zeb1 mRNAs were significantly downregulated in ADAR1 KD H9 cells upon differentiation (data not shown). Thus, the upregulated miR-96 and miR-200s in differentiated ADAR1 KD H9 cells may lead to the suppression of neural induction. Importantly, the observed aberrant expression of miR-96 and miR-200 (Figure 7C) and their targeted mRNAs (data not shown) could be rescued by introducing catalytically inactive ADAR1. We thus suggest that non-enzymatic ADAR1 participates in the regulation of these miRNA-mRNA networks during hESC differentiation and neural induction. However, we were unable to obtain iCLIP signals of ADAR1 binding to primary transcripts of miR-96 and miR-200, most likely due to their low expression (about 50-1 000-fold lower than miR302s) in undifferentiated H9 cells.
Discussion
ADAR1 deficiency is embryonically lethal in mice, showing rapid liver disintegration and hematopoietic defects23,24,25 by an unknown mechanism. This has led to the notion that ADAR1 is critical for development of some non-nervous tissues. Here we identified an unexpected non-enzymatic role of ADAR1 in ESC differentiation and neural induction in a human context. We found that ADAR1 deficiency in hESCs significantly affected EB formation and neural induction (Figures 1 and 7) by regulating miRNA maturation (Figures 3, 4, 5, 6, 7). This regulation was dependent on ADAR1 RNA-binding ability, but independent of ADAR1 catalytic activity, as the neuron induction retardation and aberrant miRNA biogenesis could be largely rescued by the catalytically inactive ADAR1-E912A but not the RNA-binding-null ADAR1-EAA (Figures 5 and 7). Mechanistically, we found that ADAR1 directly binds to stem regions of some pri-miRNAs, such as pri-miR302s, and interferes with Microprocessor processing (Figure 6 and Supplementary information, Figure S6). Together, our studies have uncovered an important role of human ADAR1 in hESC differentiation and neural induction by regulating the maturation of miRNAs via direct RNA interaction independent of its editing activity (Figure 7D).
It is well known that site-specific A-to-I RNA editing can interfere with miRNA pathways15,16,18,42,54. Recent studies reported that non-enzymatic ADAR1 participates in small RNA biogenesis in different organisms19,20. ADAR1 was suggested to interact with DICER to promote miRNA processing22 or to form a complex with DGCR8 to inhibit pri-miRNA processing21. We found that ADAR1 binds directly to pri-miR302s, competes with DGCR8 and thus inhibits miRNA processing (Figure 6). While it remains unknown how ADAR1 selects its pri-miRNAs targets, the binding sites of ADAR1 on pri-miRNAs appear to be degenerate. For example, ADAR1 largely binds to apical junction regions of pri-miR302s (Figure 6), but it binds to the predicted DROSHA cleavage sites of pri-miR1260s (Supplementary information, Figure S6). In addition, genome-wide analysis of human ADAR1 iCLIP signals revealed that ADAR1 could bind pri-miRNAs without sequence specificity (Figure 6E). Although so far little is known on how the selectivity of ADAR1 binding to different pri-miRNAs is achieved, these observations are consistent with the notion that ADAR1 prefers to bind dsRNAs, but with little sequence specificity. Importantly, it was recently reported that two DGCR8 dimers clamp a pri-miRNA hairpin using their RNA-binding heme domain to activate DROSHA cleavage52 (Figure 6C), thus the ADAR1 binding to pri-miRNA hairpin can compete with DGCR8 to inhibit pri-miRNA processing. Finally, as no A-to-I editing was identified in primary miR302s (data not shown), it further suggests that ADAR1 binding may not always lead to adenosine deamination; for instance, ADAR1 may bind to dsRNAs formed across introns and promote exon circularization55.
Depletion of ADAR1 led to the upregulation of self-renewal-related miRNAs, such as miR302s, in undifferentiated hESCs (Figures 4 and 5). In addition, such inhibition of miR302 maturation by ADAR1 can be even stronger upon differentiation (Figure 3C and 3D), when less miR302 primary transcripts are produced, resulting in an additional layer of processing regulation to facilitate a rapid downregulation of miR302s upon differentiation. Importantly, loss of ADAR1 also resulted in an increased expression of neural induction suppression-related miRNAs upon neural induction (Figure 7). Interestingly, this general upregulation of miRNAs after ADAR1 depletion is consistent with a recent study that the lack of ADARs in mouse embryos led to a general increase in miRNA numbers and changes in miRNA abundance seemed unrelated to editing events18.
We have also observed that the expression of some miRNAs was remarkably repressed during neural differentiation in ADAR1-deficient hESCs. For instance, let-7 family miRNAs were significantly downregulated in ADAR1 KD samples in late stage of neuronal differentiation (Figure 3E and 3F). This is consistent with a reported role of ADAR1 in promoting let-7 miRNA processing observed in HEK293 cells and growing mouse embryos22. However, as loss of ADAR1 significantly repressed the neural induction of hESCs (Figure 1), we cannot exclude the possibility that the downregulation of let-7 was indirect and caused by neural differentiation repression by ADAR1 depletion.
Importantly, the role of catalytically inactive ADAR1 in miRNA biogenesis seems to be evolutionarily conserved. It has been shown that at least 40% of miRNAs have altered levels in ADAR mutant C. elegans strains, although editing in small RNAs is rare20. Interestingly, a recent report found that despite the presence of a few sites in which editing is clearly beneficial, human RNA editing sites in coding regions are generally nonadaptive and evolutionarily poorly conserved, arguing against the physiological significance of these editing events in human56. Strikingly, the observations that ADAR1-deficient mice exhibited embryonic lethality23,24,25 and that ADAR1 KD hESCs exhibited significant differentiation defects (Figure 1) clearly demonstrate that ADAR1 is highly likely to be crucial for mammals. As we have observed that ADAR1 acts as an RNA-binding protein that interferes with miRNA processing in hESCs (Figures 3, 4, 5, 6) and that the retarded neural differentiation in ADAR1-deficient H9 cells could be rescued by introducing the catalytically inactive ADAR1-E912A but not the RNA-binding-deficient ADAR1-EAA (Figure 7), we suggest that the RNA-binding activity of ADAR1 is essential for development apart from its editing activity.
Materials and Methods
Plasmid construction
Plasmids used in a number of experiments were constructed using different vectors by standard molecular cloning protocols. Mutants were generated using Quikchange site-directed mutagenesis kit (Stratagene) with mutated gene-specific primers. All primers used were listed in Supplementary information, Table S7.
hESC maintenance, differentiation and lentivirus infection
H9 hESCs were maintained on irradiated-MEF feeder cells and passaged weekly as described previously58. H9 cells were induced to differentiate into the region-specific neural cells according to the published protocols37,38,39,40,41. To infect hESCs with lentivirus, H9 cells grown on Matrigel (BD Biosciences) were digested into single cells with Accutase (Millipore). Cells were incubated with MEF-condition medium (MEF-CM) containing 10 μl concentrated lentivirus, 5 μg/ml polybrene (Sigma) and 10 μM Y-27632 (Tocris) at 37 °C for 1 h in a 15 ml-conical tube and were then seeded onto Matrigel-coated culture dishes. 24 h later, cells were fed with pre-warmed MEF-CM containing 5 μg/ml polybrene for 15 min and then infected the second time on the plates. Four days after the first infection, cells were split and cultured for 5 additional days. GFP-positive colonies were mechanically detached and cultured individually.
Immunocytochemistry and western blotting
H9 cells or H9 cells differentiated into specific lineages were fixed in 4% PFA for 10 min and permeabilizd in 0.5% Triton X-100 for 10-15 min. After blocking with 1% BSA for 30 min, cells were sequentially incubated with specific primary antibodies overnight at 4 °C, and the second antibody for 1 h at 37 °C. Nuclei were counterstained with DAPI. The images were collected on a confocal microscope (Nikon). Primary antibodies used were anti-OCT-4 (1:200, Santa Cruz Biotechnology), anti-SSEA4 (1:100, Millipore), anti-PAX6 (1:300, Covance), anti-Tuj1 (1:1 000, Sigma), anti-MAP2 (1:500, Millipore) and anti-HB9 (1:50, Developmental Hybridoma Bank). For western blotting analyses, equal amount of proteins were subjected to SDS-PAGE analysis. Primary antibodies used were anti-ADAR1 (L-15; 1:200, Santa Cruz Biotechnology), anti-PAX6 (1:1 000, Covance) and anti-β-actin (1:50 000, Sigma).
RNA isolation and sequencing
Total RNAs from cultured cells were extracted with Trizol (Invitrogen) according to the manufacturer's instruction, followed by DNase I treatment at 37 °C for 30 min (Ambion, DNA-freeTM Kit). Polyadenylated RNA-seq libraries were prepared from 5 μg of total RNAs with Illumina TruSeq Stranded Total RNA HT Sample Prep Kit, and the same aliquots of total RNAs were also used to generate small RNA libraries with Illumina TruSeq Small Sample Prep Kit. All mRNA and small RNA libraries were subjected to deep sequencing with Illumina HiSeq 2000 at CAS-MPG Partner Institute for Computational Biology Omics Core, Shanghai, China. Total and mapped read numbers were listed in Supplementary information, Table S4.
In vitro processing assays
In vitro processing assay was carried out as described previously57. Briefly, pri-miR302 was IVT with the DIG RNA Labeling Mix (Roche) and AmpliScripe T7/SP6-flash Transcription Kit (Epicentre). 2 ng of Dig-labeled RNA was used for each in vitro processing assay. The same number of WT and ADAR1 KD HeLa cells was used to prepare the cell lysates. Pre-mixed reaction was first aliquoted, followed by adding either WT or ADAR1 KD HeLa cell lysates. In vitro processing assays were performed at 37 °C for 1 h, followed by RNA purification. In addition, in vitro processing assays were also performed in ADAR1 KD HeLa cell lysate in the presence or absence of the purified Flag-ADAR1. Pre-mixed reaction with ADAR1 KD HeLa cell lysate was first aliquoted to three individual tubes, followed by adding mock (lysis buffer only), control (IgG) or purified Flag-ADAR1. The reactions were performed at 37 °C for 30 min followed by RNA purification. All purified products were detected by northern blot on PAGE gels as described59.
RT-qPCRs with fractionated pri-/pre-miRNAs
Pri-miRNAs and pre-miRNAs were separated as long (> 200 nt) or short (< 200 nt) RNA fractions according to the standard protocol (Ambion)60. Briefly, 50 μg of total RNAs were mixed with lysis/binding buffer, miRNA homogenate additive and ethanol. After being passed through filter cartridge, filtrate was collected for short RNA purification, and long RNA fraction was obtained from the filter membrane followed by a series of purification steps. The fractionated RNAs were used for RT-qPCR. RT-qPCR of matured miRNA was performed with miScript II RT Kit (QIAGEN) according to the manufacturer's instructions.
Individual-nucleotide resolution UV crosslinking and immunoprecipitation (iCLIP) and UV-crosslinking RNA immunoprecipitation (RIP)
iCLIP was carried out according to the standard protocol61 with ADAR1 KD H9 cells infected with Flag-ADAR1 or Flag-E912A. See details in Supplementary information, Data S1. UV-crosslinking RIP was carried out as described previously59 with primers listed in Supplementary information, Table S7.
Gene expression and miRNA expression analyses
RNA-seq reads of each library were mapped to hg19 reference genome using TopHat262. Expression for each known gene from RefSeq was determined by covered reads and normalized with RPKM. miRNA expression was quantified by miRDeep263 pipeline. Briefly, small RNA-seq reads were trimmed to remove 3′ adapters, mapped to the genome (hg19) and piped up for miRNA expression.
Differential gene expression (DEG) and gene cluster analyses
DEG was defined with several parameters (P < 0.01 (wald test), fold change ≥ 2 or ≤ 0.5, and expression level ≥ 5 RPKM in at least one sample). DEGs were identified by gene expression comparison between WT and ADAR1 KD H9 cells at the same induction time points (d0, d10, d17 and d35). A customized pipeline was applied to obtain different gene clusters. In brief, an unsupervised hierarchical clustering for DEG genes was performed in cluster 3.064,65, and the cluster tree was manually divided to obtain about 30 gene clusters. The cluster results were shown by Java Treeview 1.0.1265,66.
Genome-wide A-to-I RNA editing analysis
To determine A-to-I editing level, RNA-seq reads were uniquely mapped to human genome hg19 and its exon junctions by bwa67 and mismatches at all reported A-to-I sites were piped up with samtools68. High-confidence A-to-I sites were obtained by stringent cutoff criteria as described previously69. For A-to-I editing in matured miRNAs, 19 known sites were examined from our small RNA-seq data sets with similar strategy as described previously42.
Accession numbers
Raw sequencing datasets for mRNAs, miRNAs and iCLIP can be accessed from the NCBI Sequence Read Archive by Gene Expression Omnibus accession numbers GSE56152 and GSE63709.
Acknowledgments
We are grateful to G Carmichael for critical reading of the manuscript, and all the lab members for helpful discussion. H9 cells were obtained from the WiCell Research Institute. This work was supported by the Ministry of Science and Technology of China (2014CB964802 and 2011CBA01105), Chinese Academy of Sciences (XDA01010206), the National Natural Science Foundation of China (91440202, 31271390, 31322018 and 31271376).
Footnotes
(Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary Information
Generation and characterization of ADAR1 stable knockdown H9 lines.
Analyses of ADAR1 stable knockdown H9 lines upon differentiation.
Effects of ADAR1 knockdown in H9 cells upon motor neuron differentiation.
Genome wide analyses of A-to-I editing sites after ADAR1 KD.
Characterization of ADAR1 on miRNA processing.
Genome-wide analysis of ADAR1 binding sites from iCLIP-seq datasets in H9 cells.
List of RefSeq genes and their expression.
List of important pathway related genes and their expression.
68 A-to-I RNA editing sites in coding regions.
Summary of RNA-seq and small RNA-seq datasets.
List of highly expressed miRNAs.
Aligned iCLIP reads to primary transcripts related to miRNAs.
Primer Sequences.
Experimental Procedures
References
- Bass BL. RNA editing by adenosine deaminases that act on RNA. Ann Rev Biochem. 2002;71:817–846. doi: 10.1146/annurev.biochem.71.110601.135501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishikura K. Functions and regulation of RNA editing by ADAR deaminases. Ann Rev Biochem. 2010;79:321–349. doi: 10.1146/annurev-biochem-060208-105251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burns CM, Chu H, Rueter SM, et al. Regulation of serotonin-2C receptor G-protein coupling by RNA editing. Nature. 1997;387:303–308. doi: 10.1038/387303a0. [DOI] [PubMed] [Google Scholar]
- Higuchi M, Maas S, Single FN, et al. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature. 2000;406:78–81. doi: 10.1038/35017558. [DOI] [PubMed] [Google Scholar]
- Hoopengardner B, Bhalla T, Staber C, Reenan R. Nervous system targets of RNA editing identified by comparative genomics. Science. 2003;301:832–836. doi: 10.1126/science.1086763. [DOI] [PubMed] [Google Scholar]
- Ohlson J, Pedersen JS, Haussler D, Ohman M. Editing modifies the GABA(A) receptor subunit alpha3. Rna. 2007;13:698–703. doi: 10.1261/rna.349107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sommer B, Kohler M, Sprengel R, Seeburg PH. RNA editing in brain controls a determinant of ion flow in glutamate-gated channels. Cell. 1991;67:11–19. doi: 10.1016/0092-8674(91)90568-j. [DOI] [PubMed] [Google Scholar]
- Athanasiadis A, Rich A, Maas S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2004;2:e391. doi: 10.1371/journal.pbio.0020391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim DD, Kim TT, Walsh T, et al. Widespread RNA editing of embedded alu elements in the human transcriptome. Genome Res. 2004;14:1719–1725. doi: 10.1101/gr.2855504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramaswami G, Lin W, Piskol R, Tan MH, Davis C, Li JB. Accurate identification of human Alu and non-Alu RNA editing sites. Nat Methods. 2012;9:579–581. doi: 10.1038/nmeth.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen LL, Carmichael GG. Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear noncoding RNA. Mol Cell. 2009;35:467–478. doi: 10.1016/j.molcel.2009.06.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen LL, DeCerbo JN, Carmichael GG. Alu element-mediated gene silencing. EMBO J. 2008;27:1694–1705. doi: 10.1038/emboj.2008.94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prasanth KV, Prasanth SG, Xuan Z, et al. Regulating gene expression through RNA nuclear retention. Cell. 2005;123:249–263. doi: 10.1016/j.cell.2005.08.033. [DOI] [PubMed] [Google Scholar]
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. doi: 10.1016/s0092-8674(04)00045-5. [DOI] [PubMed] [Google Scholar]
- Kawahara Y, Megraw M, Kreider E, et al. Frequency and fate of microRNA editing in human brain. Nucleic Acids Res. 2008;36:5270–5280. doi: 10.1093/nar/gkn479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawahara Y, Zinshteyn B, Chendrimada TP, Shiekhattar R, Nishikura K. RNA editing of the microRNA-151 precursor blocks cleavage by the Dicer-TRBP complex. EMBO Rep. 2007;8:763–769. doi: 10.1038/sj.embor.7401011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawahara Y, Zinshteyn B, Sethupathy P, Iizasa H, Hatzigeorgiou AG, Nishikura K. Redirection of silencing targets by adenosine-to-inosine editing of miRNAs. Science. 2007;315:1137–1140. doi: 10.1126/science.1138050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vesely C, Tauber S, Sedlazeck FJ, von Haeseler A, Jantsch MF. Adenosine deaminases that act on RNA induce reproducible changes in abundance and sequence of embryonic miRNAs. Genome Res. 2012;22:1468–1476. doi: 10.1101/gr.133025.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heale BS, Keegan LP, McGurk L, et al. Editing independent effects of ADARs on the miRNA/siRNA pathways. EMBO J. 2009;28:3145–3156. doi: 10.1038/emboj.2009.244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warf MB, Shepherd BA, Johnson WE, Bass BL. Effects of ADARs on small RNA processing pathways in C. elegans. Genome Res. 2012;22:1488–1498. doi: 10.1101/gr.134841.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nemlich Y, Greenberg E, Ortenberg R, et al. MicroRNA-mediated loss of ADAR1 in metastatic melanoma promotes tumor growth. J Clin Invest. 2013;123:2703–2718. doi: 10.1172/JCI62980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ota H, Sakurai M, Gupta R, et al. ADAR1 forms a complex with Dicer to promote microRNA processing and RNA-induced gene silencing. Cell. 2013;153:575–589. doi: 10.1016/j.cell.2013.03.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartner JC, Schmittwolf C, Kispert A, Muller AM, Higuchi M, Seeburg PH. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J Biol Chem. 2004;279:4894–4902. doi: 10.1074/jbc.M311347200. [DOI] [PubMed] [Google Scholar]
- Wang Q, Miyakoda M, Yang W, et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J Biol Chem. 2004;279:4952–4961. doi: 10.1074/jbc.M310162200. [DOI] [PubMed] [Google Scholar]
- Hartner JC, Walkley CR, Lu J, Orkin SH. ADAR1 is essential for the maintenance of hematopoiesis and suppression of interferon signaling. Nat Immunol. 2009;10:109–115. doi: 10.1038/ni.1680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rice GI, Kasher PR, Forte GM, et al. Mutations in ADAR1 cause Aicardi-Goutieres syndrome associated with a type I interferon signature. Nat Genet. 2012;44:1243–1248. doi: 10.1038/ng.2414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murry CE, Keller G. Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell. 2008;132:661–680. doi: 10.1016/j.cell.2008.02.008. [DOI] [PubMed] [Google Scholar]
- Stern CD. Neural induction: old problem, new findings, yet more questions. Development. 2005;132:2007–2021. doi: 10.1242/dev.01794. [DOI] [PubMed] [Google Scholar]
- Zhang SC. Neural subtype specification from embryonic stem cells. Brain Pathol. 2006;16:132–142. doi: 10.1111/j.1750-3639.2006.00008.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gifford CA, Ziller MJ, Gu H, et al. Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell. 2013;153:1149–1163. doi: 10.1016/j.cell.2013.04.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joo JH, Yoon SY, Kim JH, et al. S100A6 (calcyclin) enhances the sensitivity to apoptosis via the upregulation of caspase-3 activity in Hep3B cells. J Cell Biochem. 2008;103:1183–1197. doi: 10.1002/jcb.21496. [DOI] [PubMed] [Google Scholar]
- Pawlowski J, Kraft AS. Bax-induced apoptotic cell death. Proc Natl Acad Sci USA. 2000;97:529–531. doi: 10.1073/pnas.97.2.529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slomnicki LP, Nawrot B, Lesniak W. S100A6 binds p53 and affects its activity. Int J Biochem Cell Biol. 2009;41:784–790. doi: 10.1016/j.biocel.2008.08.007. [DOI] [PubMed] [Google Scholar]
- Han X, Amar S. Secreted frizzled-related protein 1 (SFRP1) protects fibroblasts from ceramide-induced apoptosis. J Biol Chem. 2004;279:2832–2840. doi: 10.1074/jbc.M308102200. [DOI] [PubMed] [Google Scholar]
- Joesting MS, Perrin S, Elenbaas B, et al. Identification of SFRP1 as a candidate mediator of stromal-to-epithelial signaling in prostate cancer. Cancer Res. 2005;65:10423–10430. doi: 10.1158/0008-5472.CAN-05-0824. [DOI] [PubMed] [Google Scholar]
- Du ZW, Ma LX, Phillips C, Zhang SC. miR-200 and miR-96 families repress neural induction from human embryonic stem cells. Development. 2013;140:2611–2618. doi: 10.1242/dev.092809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang ZB, Zhang X, Li XJ. Recapitulation of spinal motor neuron-specific disease phenotypes in a human cell model of spinal muscular atrophy. Cell Res. 2013;23:378–393. doi: 10.1038/cr.2012.166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Huang CT, Chen J, et al. Pax6 is a human neuroectoderm cell fate determinant. Cell Stem Cell. 2010;7:90–100. doi: 10.1016/j.stem.2010.04.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boisvert EM, Denton K, Lei L, Li XJ.The specification of telencephalic glutamatergic neurons from human pluripotent stem cells J Vis Exp 2013. Apr 14; doi: 10.3791/50321 [DOI] [PMC free article] [PubMed]
- Li XJ, Zhang X, Johnson MA, Wang ZB, Lavaute T, Zhang SC. Coordination of sonic hedgehog and Wnt signaling determines ventral and dorsal telencephalic neuron types from human embryonic stem cells. Development. 2009;136:4055–4063. doi: 10.1242/dev.036624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li XJ, Du ZW, Zarnowska ED, et al. Specification of motoneurons from human embryonic stem cells. Nat Biotechnol. 2005;23:215–221. doi: 10.1038/nbt1063. [DOI] [PubMed] [Google Scholar]
- Alon S, Mor E, Vigneault F, et al. Systematic identification of edited microRNAs in the human brain. Genome Res. 2012;22:1533–1540. doi: 10.1101/gr.131573.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barroso-del Jesus A, Lucena-Aguilar G, Menendez P. The miR-302-367 cluster as a potential stemness regulator in ESCs. Cell Cycle. 2009;8:394–398. doi: 10.4161/cc.8.3.7554. [DOI] [PubMed] [Google Scholar]
- Houbaviy HB, Murray MF, Sharp PA. Embryonic stem cell-specific microRNAs. Dev Cell. 2003;5:351–358. doi: 10.1016/s1534-5807(03)00227-2. [DOI] [PubMed] [Google Scholar]
- Wang Y, Melton C, Li YP, et al. miR-294/miR-302 promotes proliferation, suppresses G1-S restriction point, and inhibits ESC differentiation through separable mechanisms. Cell Rep. 2013;4:99–109. doi: 10.1016/j.celrep.2013.05.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Card DA, Hebbar PB, Li L, et al. Oct4/Sox2-regulated miR-302 targets cyclin D1 in human embryonic stem cells. Mol Cell Biol. 2008;28:6426–6438. doi: 10.1128/MCB.00359-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asuelime GE, Shi Y. The little molecules that could: a story about microRNAs in neural stem cells and neurogenesis. Front Neurosci. 2012;6:176. doi: 10.3389/fnins.2012.00176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawahara H, Imai T, Okano H. MicroRNAs in neural stem cells and neurogenesis. Front Neurosci. 2012;6:30. doi: 10.3389/fnins.2012.00030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- George CX, John L, Samuel CE. An RNA editor, adenosine deaminase acting on double-stranded RNA (ADAR1) J Interferon Cytokine Res. 2014;34:437–446. doi: 10.1089/jir.2014.0001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai F, Drakas R, Nishikura K. Mutagenic analysis of double-stranded RNA adenosine deaminase, a candidate enzyme for RNA editing of glutamate-gated ion channel transcripts. J Biol Chem. 1995;270:17098–17105. doi: 10.1074/jbc.270.29.17098. [DOI] [PubMed] [Google Scholar]
- Valente L, Nishikura K. RNA binding-independent dimerization of adenosine deaminases acting on RNA and dominant negative effects of nonfunctional subunits on dimer functions. J Biol Chem. 2007;282:16054–16061. doi: 10.1074/jbc.M611392200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quick-Cleveland J, Jacob JP, Weitz SH, Shoffner G, Senturia R, Guo F. The DGCR8 RNA-binding heme domain recognizes primary microRNAs by clamping the hairpin. Cell Rep. 2014;7:1994–2005. doi: 10.1016/j.celrep.2014.05.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng Y, Yi R, Cullen BR. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 2005;24:138–148. doi: 10.1038/sj.emboj.7600491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang W, Chendrimada TP, Wang Q, et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat Struct Mol Biol. 2006;13:13–21. doi: 10.1038/nsmb1041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L. Complementary sequence-mediated exon circularization. Cell. 2014;159:134–147. doi: 10.1016/j.cell.2014.09.001. [DOI] [PubMed] [Google Scholar]
- Xu G, Zhang J. Human coding RNA editing is generally nonadaptive. Proc Natl Acad Sci USA. 2014;111:3769–3774. doi: 10.1073/pnas.1321745111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Y, Kim VN. In vitro and in vivo assays for the activity of Drosha complex. Methods Enzymol. 2007;427:89–106. doi: 10.1016/S0076-6879(07)27005-3. [DOI] [PubMed] [Google Scholar]
- Yin QF, Yang L, Zhang Y, et al. Long noncoding RNAs with snoRNA ends. Mol Cell. 2012;48:219–230. doi: 10.1016/j.molcel.2012.07.033. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Zhang XO, Chen T, et al. Circular intronic long noncoding RNAs. Mol Cell. 2013;51:792–806. doi: 10.1016/j.molcel.2013.08.017. [DOI] [PubMed] [Google Scholar]
- Cheng TL, Wang Z, Liao Q, et al. MeCP2 suppresses nuclear microRNA processing and dendritic growth by regulating the DGCR8/Drosha complex. Dev Cell. 2014;28:547–560. doi: 10.1016/j.devcel.2014.01.032. [DOI] [PubMed] [Google Scholar]
- Konig J, Zarnack K, Rot G, et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat Struct Mol Biol. 2010;17:909–915. doi: 10.1038/nsmb.1838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14:R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012;40:37–52. doi: 10.1093/nar/gkr688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Hoon MJ, Imoto S, Nolan J, Miyano S. Open source clustering software. Bioinformatics. 2004;20:1453–1454. doi: 10.1093/bioinformatics/bth078. [DOI] [PubMed] [Google Scholar]
- Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998;95:14863–14868. doi: 10.1073/pnas.95.25.14863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saldanha AJ. Java Treeview ― extensible visualization of microarray data. Bioinformatics. 2004;20:3246–3248. doi: 10.1093/bioinformatics/bth349. [DOI] [PubMed] [Google Scholar]
- Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu S, Xiang JF, Chen T, Chen LL, Yang L. Prediction of constitutive A-to-I editing sites from human transcriptomes in the absence of genomic sequences. BMC genomics. 2013;14:206. doi: 10.1186/1471-2164-14-206. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Generation and characterization of ADAR1 stable knockdown H9 lines.
Analyses of ADAR1 stable knockdown H9 lines upon differentiation.
Effects of ADAR1 knockdown in H9 cells upon motor neuron differentiation.
Genome wide analyses of A-to-I editing sites after ADAR1 KD.
Characterization of ADAR1 on miRNA processing.
Genome-wide analysis of ADAR1 binding sites from iCLIP-seq datasets in H9 cells.
List of RefSeq genes and their expression.
List of important pathway related genes and their expression.
68 A-to-I RNA editing sites in coding regions.
Summary of RNA-seq and small RNA-seq datasets.
List of highly expressed miRNAs.
Aligned iCLIP reads to primary transcripts related to miRNAs.
Primer Sequences.
Experimental Procedures