Abstract
Recombinant protein overexpression of large proteins in bacteria often results in insoluble and misfolded proteins directed to inclusion bodies. We report the application of shear stress in micrometer-wide, thin fluid films to refold boiled hen egg white lysozyme, recombinant hen egg white lysozyme, and recombinant caveolin-1. Furthermore, the approach allowed refolding of the much larger protein, cAMP-dependent protein kinase A (PKA). The reported methods require only minutes, which is >100-times faster than conventional, overnight dialysis. This rapid refolding technique could significantly shorten the times, lower costs, and reduce the waste streams associated with protein expression for a wide-range of industrial and research applications.
Keywords: proteins, protein folding, protein expression, green chemistry
Overexpressed recombinant proteins for industrial, pharmaceutical, environmental and agricultural applications annually represent a >$160 billion biotechnology world market.[1] Protein expression in yeast or Escherichia coli is highly preferred due to the organisms' rapid growth, low consumable costs, and high yields.[2, 3] However, large proteins overexpressed in bacteria often form aggregates and inclusion bodies.[4–7] Recovery of the correctly folded protein then requires laborious and expensive processing of inclusion bodies by conventional methods.[1, 8] The most common method for refolding such proteins, for example, involves multi-day dialysis with large volumes (typically 1-10 liters for mg quantities of protein).[9] Alternatively, high value proteins (e.g., therapeutic antibodies or GPCRs for structural biology) apply extensively optimized mammalian or insect cell lines, media and bioreactor conditions.[10–12] Recovery of correctly folded proteins from aggregates is inefficient and challenging for large-scale industrial processes. Mechanical methods to solve this problem have been reported. One approach applies very high hydrostatic pressures (400 bar) to refold recombinant proteins from inclusion bodies.[13–16] We report a novel method by applying finely controlled levels of shear stress to refold proteins trapped in the inclusion body. This method may be capable of broadening the utility of bacterial over-expression, and could transform industrial and research production of proteins.
We report using a vortex fluid device (VFD) to apply shear forces for rapid equilibration of protein folding and isolation of intermediates during protein folding. In this method, a glass cylinder (10 mm by 16 cm) is spun rapidly (5 krpm) at a 45° angle. At high rotational speeds, the solution within the sample tube forms micrometer-thick, thin fluid films, which flow with the same speed and direction as the wall of the glass tube. The rotating glass tube generates a velocity gradient within the thin fluid film, which introduces shear stress into the solution (Figure 1 a). We imagined applying the VFD with a similar range of input energies to refolding proteins; the device produces unusual shear within such films, and has been shown to be effective in disassembling molecular capsules,[15] and reducing the size of micelles used in the fabrication of SBA-15 mesoporous silica with control over the pore size of the material.[16] Importantly, the latter occurs at room temperature rather than requiring hydrothermal processing, and suggests that the constant ‘soft’ energy in the thin film has application in manipulating macromolecules, such as the refolding of proteins, without requiring heating. Other relevant applications and optimization of a similar vortex fluid device include exfoliating graphite and hexagonal boron-nitride to generate mono and multi-layer structures,[17, 18] controlling the formation of different calcium carbonate polymorphs,[19] and controlling chemical reactivity and selectivity.[20, 21]
Figure 1.
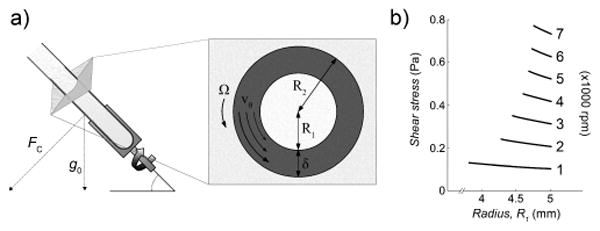
Protein refolding in vitro with the vortex fluid device (VFD), which generates shear flow inside thin fluid films (shaded). a) VFD schematic and parameters for modeling. The inset represents a two-dimensional slice of the tube through the axis of rotation. The inner diameter of the tube, R2, is 5 mm, and the thin fluid film thickness, δδ, has been previously measured (depicted not to scale).[17] The velocity of the solution, vθ, increases from the inner surface of the film to match the velocity of the tube, Ω. b). Shear stress calculated as a function of rotational speed (rpm) and radius, R1, from the center of the tube.
Modeling the fluid behavior in the VFD allowed estimation of the shear forces experienced by proteins folding at various rotational speeds. Our analysis applies the solution for cylindrical Couette flow.[20, 21] The velocity of the solution, vθ, is a function of the radius, r (Figure 1 a). The boundary conditions for the liquid film interfaces are defined as follows. The inner air-liquid interface at r = R1 slips due to discontinuity in viscosity, and results in vanishingly low shear stress At the outer liquid-glass interface, the no-slip boundary dictates that the velocity of the liquid at r = R2 matches that of the inner wall of the glass tube (vθ = R2 · Ω), where Ω is the angular velocity of the tube. The resulting velocity profile is a nonlinear function of the form
where
and
From this velocity profile, shear stress can be calculated as
where μ is the viscosity of water at 20 °C. At a speed of 5 krpm, the calculated shear stress ranges from 0.5246 to 0.5574 Pa (Figure 1 b). While this model provides insight into the shear stress range, the fluid flow will be perturbed by the cross vector of gravity with centrifugal force for a tube orientated at 45°,[20] which may also facilitate protein folding; this angular setting also proved optimal for the aforementioned applications of the VFD. The calculated values of shear stress are similar to the levels previously reported to cause protein unfolding,[22] and we hypothesized that short periods of VFD processing could allow equilibration of protein folding.
Experiments with native hen egg white were conducted to determine if shear forces could refold denatured hen egg white lysozyme (HEWL) in complex environments. The separated whites were diluted in PBS, and heat-treated at 90 °C for 20 min. The resultant hard-boiled egg white was dissolved in 8 M urea, rapidly diluted and then VFD-processed at the indicated rotational speeds and times (Table S1 and Figure 2 a). The total protein concentration as determined by bicinchoninic acid assay was 44 μg/ml. The recovery of HEWL activity was then demonstrated by lysozyme activity assay (Figure S1). Refolding HEWL within the egg white at 5 krpm recovers glycosidase activity even after a short 2.5 min spin, but continued shear forces unfolds the protein. As mentioned above, shear stress-mediated protein unfolding has been observed for other proteins previously. Optimization of VFD times, speeds, and concentration of chaotropic additives is required to find conditions allowing re-equilibration of protein folding without causing loss of the protein structure. VFD processing for 5 minutes at 5 krpm in 1 M urea results in optimal HEWL refolding as monitored by lysozyme activity (Figure 2 a). This experiment establishes the basic parameters for protein refolding by VFD.
Figure 2.
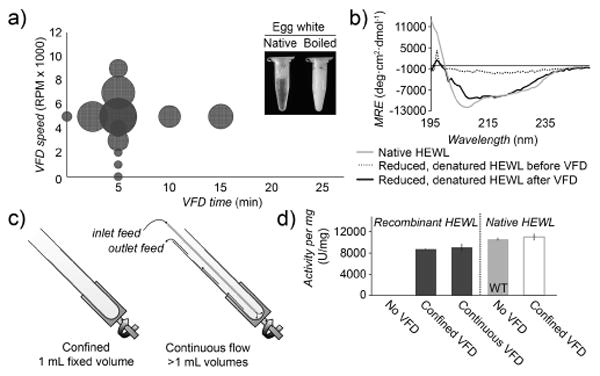
Determination of secondary structure and activity of hen egg white lysozyme (HEWL) processed by VFD. a) Lysozyme activity per mg protein following VFD processing of boiled egg white (90 °C, 20 min) at a fixed VFD speed of 5 krpm, or a fixed 5 min refolding time. The relative circle size represents the lysozyme activity with larger circles representing higher recovered activities, plotted as a function of VFD time and speed. b) CD spectra of recombinantly expressed, reduced, and denatured HEWL before (dotted) and after (dark gray) VFD refolding. c) In continuous flow mode, the protein solution is introduced through a thin, hollow metal tube to the bottom of the sample tube, and the folded protein can be collected at the top of the VFD. d) Lysozyme activity per mg protein following VFD refolding of recombinantly expressed HEWL and native HEWL. Under these conditions, VFD treatment of recombinant lysozyme recovered over 82% of activity in both confined and continuous flow modes, compared to wild-type protein (WT). VFD treatment of wild-type, active lysozyme isolated from eggs does not adversely affect its activity (white). Throughout this report, error bars indicate standard deviation (n = 3).
To demonstrate refolding of recombinantly expressed, reduced HEWL, the cell pellet was first reconstitued in lysis buffer containing 2-mercaptoethanol, denatured in 8 M urea and rapidly diluted into PBS (1:100). Buffers used for the expression and purifications of all recombinant proteins are described, in Table S2. Second, the diluted protein (1 ml, 44 μg/ml protein) was immediately transferred to the VFD sample tube and spun at 22 °C and 5 krpm for 5 min. Circular dichroism (CD) spectra of the VFD-refolded, recombinant HEWL demonstrates restoration of secondary structure from proteins isolated from inclusion bodies. After VFD processing, the CD spectra of identical HEWL samples demonstrates partial recovery of secondary structure compared to the native lysozyme (Figure 2 b). Yields of functional protein determined by lysozyme activity assay are shown in Figure 2 d.
HEWL can also be refolded by continuous flow VFD. This approach delivers additional sample through an inlet at the cylinder base. The sample (50 ml), added at a flow-rate of 0.1 ml/min, demonstrates significant recovery of HEWL activity for scalable, high volume applications (Figure 2 c). The recombinant HEWL recovers >82% of its activity following VFD treatment. As expected, HEWL isolated from inclusion bodies without VFD processing fails to show any lysozyme activity (Figure 2 d). The continuous flow approach could be readily scaled up and parallelized for industrial applications requiring treatment of very large volumes.
After refolding denatured lysozyme in both complex (egg white) and simple (purified recombinant protein) environments, the next experiments focused on refolding the protein caveolin-1, as an example of a protein requiring an inordinate amount of processing time by conventional approaches (e.g., four days of dialysis). A caveolin variant without its transmembrane domain (caveolin-ΔTM) was recombinantly expressed, and the inclusion body was purified under denaturing conditions. Purified caveolin-ΔTM was diluted, and then given a short dialysis for 1 h to decrease the urea concentration to 1 M. The protein was then VFD-treated for 0, 10, or 30 min at 5 krpm at a concentration of 186 μg/ml. The CD spectra of the VFD processed caveolin-ΔTM shows a pronounced minima at 208 and 220 nm, which are indicative of α -helical secondary structure.[23] Solution turbidity also decreases sharply following VFD treatment, which illustrates VFD solubilization and refolding of partially aggregated proteins (Figure 3 b). ELISA experiments examined binding by the refolded caveolin-ΔTM to HIV glycoprotein 41 (gp41), a known caveolin binding partner.[24–26] VFD processing significantly restores protein function, as shown through binding to gp41 (Figure 3 c).
Figure 3.
Determination of secondary structure and activity of caveolin-ΔTM processed by VFD. a) Circular dichroism (CD) spectra of caveolin-ΔTM following VFD induced refolding or conventional dialysis. b) Turbidity of caveolin-ΔTM, measured by the fractional absorbance at 600 nm compared to the untreated sample also at 600 nm. The arrow indicates absorbance of caveolin-ΔTM following 4-days of dialysis. Both 30 min of VFD-based refolding and 96 h of dialysis significantly decrease the turbidity of the protein solution. c) Binding of caveolin-ΔTM to gp41 determined by ELISA using untreated, VFD-treated, and conventionally refolded by dialysis. All other conditions and buffers remained identical. Although some binding occurs without VFD treatment, caveolin-ΔTM binds with greater affinity after refolding by VFD.
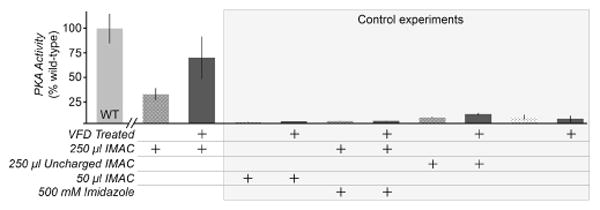
Larger-sized proteins initially failed to refold despite VFD treatment. For example, the catalytic domain of cAMP-dependent protein kinase A (PKA, 42 kDa) is significantly bulkier than HEWL (14 kDa) and caveolin-ΔTM (17 kDa), and did not refold from inclusion bodies after treatment with similar protocols to the above experiments. To refold full-length PKA in vitro, we adapted the VFD to provide a closer mimic of cellular folding. In cells, the nascent polypeptide can fold as the N-terminus extrudes from the ribosome, whereas in vitro refolding must address the entire protein at once.[27] Thus, we attempted to focus shear stress on the N-terminus of His-tagged PKA by binding the unfolded protein to Ni2+-charged immobilized metal affinity chromatography (IMAC) resin. The His6-PKA-IMAC complex (1 ml, 0.2 – 1 mg/ml) was then subjected to refolding by VFD. Following VFD treatment, His-PKA separated from the IMAC resin, and recovered 69% of its kinase activity (Figure 4). Interestingly, the remaining His-PKA eluted from the IMAC resin by elution with imidazole yielded far less active protein (Figure S3). We hypothesize that protein folding makes the His6 epitope less accessible for strong binding to the IMAC resin subjected to shear by the VFD. Under conventional conditions, IMAC resin binds to folded PKA, but we recovered protein remaining on the IMAC resin after VFD treatment to demonstrate its lack of folding and catalytic activity (Figure S3). Thus, as the VFD folds the protein, PKA elutes from the affinity resin. Negative control experiments with identical conditions, but with uncharged IMAC resin, decreased quantities of charged resin, or 500 mM imidazole to block the Ni2+-His6 tag interaction, showed only low levels of kinase activity (Figure 4).
Figure 4.
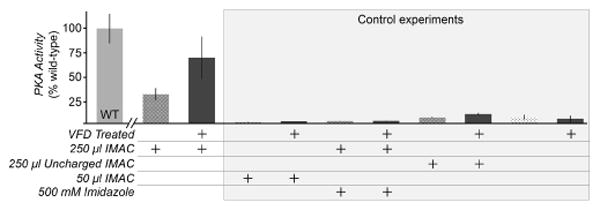
Facilitating VFD refolding of PKA by pre-binding the His6-taggged protein to IMAC resin. 1.7 mg PKA was pre-incubated with 250 or 50 μl IMAC resin in 6 M guanidine-HCl prior to dilution to 1 M guanidine-HCl and VFD treatment. Excess PKA was removed by a low imidazole (1 mM) wash buffer. The PKA activity per μg of protein was measured by a nicotinamide adenine dinucleotide (NADH) enzyme-linked assay, and shown here as a percentage of untreated, wild-type PKA activity (WT). Dark gray bars denote samples that are VFD-treated. In this assay, the consumption of ATP by PKA catalysis results in a lactate dehydrogenase-coupled decrease in the levels of NADH, monitored through measuring absorbance at 340 nm. The control experiments included low quantities of resin (50 μl IMAC), and two controls for non-specific binding to the resin in the absence of the Ni2+-His6 tag interaction (500 mM imidazole added and uncharged IMAC resin lacking Ni2+). Imidazole was diluted to 50 mM before kinase assay.
As reported here, protein refolding by VFD requires optimization for each protein. Buffers, chaotrope additive, protein concentration, and processing time were optimized for HEWL, caveolin-ΔTM, and PKA. The refolding of HEWL from the complex mixture of boiled egg whites appears less efficient than recovery of the folded protein isolated from inclusion bodies. In egg whites, the mechanical energy of the VFD could be misdirected to the other >96% of proteins present.
This report offers significant advantages over conventional approaches to protein refolding. First, VFD-mediated refolding requires much smaller solution volumes (approximately 1% of the volumes required for conventional dialysis). Second, this key step in protein production occurs >100-times faster than overnight dialysis with >1000-fold improvements for proteins such as a caveolin. Notably, introducing high shear in thin fluid films is a low energy, inexpensive process, and can be readily implemented for relatively low cost in conventional biochemistry laboratories.
The advantages of VFD refolding open new possibilities for increasing protein yields in simple cell lines. Using the constant soft energy of the thin microfluidic films, the VFD can untangle complex mixtures, aggregates and insoluble inclusion bodies. In industrial and academic applications, high concentrations of a chemical inducer like IPTG could drive overexpressed proteins into insoluble inclusion bodies before VFD-based refolding. Conversely, most conventional processes avoid inclusion bodies by optimization of growth conditions and special cell lines at the expense of higher yields and purer protein isolated directly from bacterial cells. As reported here, the continuous flow mode of the VFD or parallel processing with multiple VFD units could allow scale-up to accommodate much larger solution volumes. Thus, the approach could drastically lower the time and financial costs required to refold inactive proteins at an industrial scale. The VFD sample tube itself can also be modified to amplify or otherwise direct the intensity of shear forces applied; for example, modified surfaces with high contact angle and/or with textured features could enhance the turbulent flow, altering the applied shear stress. Harnessing shear forces to achieve rapid equilibration of protein folding could be expanded to a wide-range of applications for research and manufacturing.
Supplementary Material
Acknowledgments
We thank Professor Abraham Lee for discussion and advice on fluid modeling. This work was supported by the National Institute of General Medical Sciences of the NIH (R01 GM100700-01) and the Australian Research Council (ARC DP1092810 and DP130100066).
Footnotes
Supporting information for this article is given via a link at the end of the document.
Contributor Information
Colin L. Raston, Email: colin.raston@flinders.edu.au.
Gregory A. Weiss, Email: gweiss@uci.edu.
References
- 1.Meyer H, Schmidhalter D. Innov Biotechnol. 2012:211–250. [Google Scholar]
- 2.Freire E, Schön A, Velazquez-Campoy A. Methods Enzymol. 2009;455:127–155. doi: 10.1016/S0076-6879(08)04205-5. [DOI] [PubMed] [Google Scholar]
- 3.Futaki S. Adv Drug Deliv Rev. 2005;57:547–558. doi: 10.1016/j.addr.2004.10.009. [DOI] [PubMed] [Google Scholar]
- 4.Radford SE. Trends Biochem Sci. 2000;25:611–618. doi: 10.1016/s0968-0004(00)01707-2. [DOI] [PubMed] [Google Scholar]
- 5.Jemth P, Gianni S, Day R, Li B, Johnson CM, Daggett V, Fersht AR. Proc Natl Acad Sci U S A. 2004;101:6450–6455. doi: 10.1073/pnas.0401732101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wang L, Maji SK, Sawaya MR, Eisenberg D, Riek R. PLoS Biol. 2008;6:e195. doi: 10.1371/journal.pbio.0060195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wang L, Schubert D, Sawaya MR, Eisenberg D, Riek R. Angew Chem Int Ed Engl. 2010;49:3904–3908. doi: 10.1002/anie.201000068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Vallejo LF, Rinas U. Microb Cell Fact. 2004;3:11. doi: 10.1186/1475-2859-3-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tsumoto K, Ejima D, Kumagai I, Arakawa T. Protein Expr Purif. 2003;28:1–8. doi: 10.1016/s1046-5928(02)00641-1. [DOI] [PubMed] [Google Scholar]
- 10.Li F, Vijayasankaran N, Shen A, Kiss R, Amanullah A. MAbs. 2010;2:466–479. doi: 10.4161/mabs.2.5.12720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hannig G, Makrides SC. Trends Biotechnol. 1998;16:54–60. doi: 10.1016/s0167-7799(97)01155-4. [DOI] [PubMed] [Google Scholar]
- 12.Andersen DC, Krummen L. Curr Opin Biotechnol. 2002;13:117–123. doi: 10.1016/s0958-1669(02)00300-2. [DOI] [PubMed] [Google Scholar]
- 13.St John RJ, Carpenter JF, Randolph TW. Proc Natl Acad Sci U S A. 1999;96:13029–13033. doi: 10.1073/pnas.96.23.13029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Qoronfleh MW, Hesterberg LK, Seefeldt MB. Protein Expr Purif. 2007;55:209–224. doi: 10.1016/j.pep.2007.05.014. [DOI] [PubMed] [Google Scholar]
- 15.Lefebvre BG, Gage MJ, Robinson AS. Biotechnol Prog. 2004;20:623–629. doi: 10.1021/bp034221v. [DOI] [PubMed] [Google Scholar]
- 16.Lefebvre BG, Robinson AS. Biotechnol Bioeng. 2003;82:595–604. doi: 10.1002/bit.10607. [DOI] [PubMed] [Google Scholar]
- 17.Chen X, Dobson JF, Raston CL. Chem Commun. 2012;48:3703–3705. doi: 10.1039/c2cc17611d. [DOI] [PubMed] [Google Scholar]
- 18.Wahid MH, Eroglu E, Chen X, Smith SSM, Raston CCL. Green Chem. 2013;15:650–655. [Google Scholar]
- 19.Boulos RA, Zhang F, Tjandra ES, Martin AD, Spagnoli D, Raston CL. Sci Rep. 2014;4:3616. doi: 10.1038/srep03616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mallock A. Proc R Soc A. 1888;45:126–132. [Google Scholar]
- 21.Rayleigh L. Proc R Soc A. 1917;93:148–154. [Google Scholar]
- 22.Bekard IB, Asimakis P, Bertolini J, Dunstan DE. Biopolymers. 2011;95:733–745. doi: 10.1002/bip.21646. [DOI] [PubMed] [Google Scholar]
- 23.Saxena VP, Wetlaufer DB. Proc Natl Acad Sci. 1971;68:969–972. doi: 10.1073/pnas.68.5.969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hovanessian AG, Briand JP, a Said E, Svab J, Ferris S, Dali H, Muller S, Desgranges C, Krust B. Immunity. 2004;21:617–627. doi: 10.1016/j.immuni.2004.08.015. [DOI] [PubMed] [Google Scholar]
- 25.Wang XM, Nadeau PE, Lo YT, Mergia A. J Virol. 2010;84:6515–6526. doi: 10.1128/JVI.02722-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Majumdar S, Hajduczki A, Vithayathil R, Olsen TJ, Spitler RM, Mendez AS, Thompson TD, a Weiss G. J Am Chem Soc. 2011;133:9855–9862. doi: 10.1021/ja201792q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Evans MS, Sander IM, Clark PL. J Mol Biol. 2008;383:683–692. doi: 10.1016/j.jmb.2008.07.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


