Abstract
We present a general model of actin filament deformation and fragmentation in response to compressive forces. The elastic free energy density along filaments is determined by their shape and mechanical properties, which were modeled in terms of bending, twisting, and twist-bend coupling elasticities. The elastic energy stored in filament deformation (i.e., strain) tilts the fragmentation-annealing reaction free-energy profile to favor fragmentation. The energy gradient introduces a local shear force that accelerates filament intersubunit bond rupture. The severing protein, cofilin, renders filaments more compliant in bending and twisting. As a result, filaments that are partially decorated with cofilin are mechanically heterogeneous (i.e., nonuniform) and display asymmetric shape deformations and energy profiles distinct from mechanically homogenous (i.e., uniform), bare actin, or saturated cofilactin filaments. The local buckling strain depends on the relative size of the compliant segment as well as the bending and twisting rigidities of flanking regions. Filaments with a single bare/cofilin-decorated boundary localize energy and force adjacent to the boundary, within the compliant cofilactin segment. Filaments with small cofilin clusters were predicted to fragment within the compliant cofilactin rather than at boundaries. Neglecting contributions from twist-bend coupling elasticity underestimates the energy density and gradients along filaments, and thus the net effects of filament strain to fragmentation. Spatial confinement causes compliant cofilactin segments and filaments to adopt higher deformation modes and store more elastic energy, thereby promoting fragmentation. The theory and simulations presented here establish a quantitative relationship between actin filament fragmentation thermodynamics and elasticity, and reveal how local discontinuities in filament mechanical properties introduced by regulatory proteins can modulate both the severing efficiency and location along filaments. The emergent behavior of mechanically heterogeneous filaments, particularly under confinement, emphasizes that severing in cells is likely to be influenced by multiple physical and chemical factors.
Introduction
Assembly of the cytoskeletal protein actin into linear filaments and higher-order networks powers a variety of cell shape changes and movements, including leading-edge extension of migrating cells and cell division (1,2). Persistent motility requires continuous actin reorganization, a process coordinated through dynamic interactions with numerous regulatory proteins. Among these are contractile myosin motor proteins that generate force and work along filaments, and severing proteins that fragment filaments and increase the concentration of ends available for subunit addition and loss.
Members of the ADF/cofilin family of severing proteins (herein, referred to as cofilin) alter actin structure (3,4) and increase the filament bending and twisting compliance (5). Filaments partially decorated with cofilin display fragmentation more readily than bare or saturated ones (6–10), and severing occurs preferentially at or near boundaries (e.g., junctions) of bare and cofilin-decorated segments (5,8,11). These observations have favored a model in which local topological and mechanical discontinuities render boundaries along partially decorated filaments susceptible to fragmentation (4,12,13).
Active forces generated by myosin motor proteins also fragment filaments and accelerate actin network turnover, both in vivo and in reconstituted biomimetic systems (14–21). It has been postulated that severing and contractile protein activities are coordinated in vivo, such that motor protein-driven deformations in filament shape enhance fragmentation by cofilin severing proteins (12,16,17). Lacking, as of this writing, are predictive metric studies of how such active shape deformations promote filament fragmentation, as well as how local heterogeneities in filament mechanical properties introduced by cofilin occupancy influence the severing location and rate constant.
Here, we establish a theoretical framework of actin filament deformation and fragmentation in response to external, compressive forces generated by contractile motor proteins. We employ numerical simulations to quantitate the spatial elastic energy and local shear force of homogenous filaments (e.g., actin and cofilactin) and filaments with mechanical heterogeneity (e.g., partially decorated with cofilin) undergoing buckling deformation. We consider five filament classes: 1) native, bare actin filaments; 2) cofilactin filaments; 3) filaments with a single bare- and cofilin-decorated boundary; and filaments with two boundaries, defined by either 4) a single bare or 5) a cofilin-decorated segment, positioned at the filament center. Our approach and analysis provides a quantitative formalism between actin filament assembly thermodynamics and elasticity, and reveal how local alterations in mechanical properties introduced by cofilin occupancy can modulate both the severing efficiency and location along filaments.
Materials and Methods
Theory and simulation
In the forthcoming section we introduce the internal force density (f) and curvature strains (κ) representing bending and twisting deformations at a filament position, the moment vector (m), and Kirchhoff elastic free energy associated with filament bending and twisting strains. We develop a general model for the mechanical (force and moment) equilibrium of elastic filaments subjected to external forces and torques (General Equations for Mechanical Equilibrium of an Elastic Filament) and use Euler parameters (quaternions) to derive a set of equations suitable for simulations (Quaternions (Euler Parameters) and Componentwise Form of Equations). We present the derivation of mechanical equilibrium conditions accounting for coupling of actin filament bending and twisting modes (twist-bend coupling elasticity (22); see Fig. S1 in the Supporting Material). We provide the boundary conditions and external forces and torques applied to filaments (Boundary Conditions, and Cofilin Density and Parameter Dependence on Space). We then adapt the equations to filaments displaying heterogeneity and local discontinuities in mechanical properties (i.e., variable persistence length parameters) that result from cofilin binding (5,8,9,23–26) (Confinement in Compliant Cylindrical Chamber; see Fig. S2). This was followed by Verification of Numerical Simulations.
General equations for mechanical equilibrium of an elastic filament
We use the equations for elastic, inextensible rods (27) to simulate the conformations of individual actin filaments subjected to external buckling forces. At each point, the orientation of the filament cross-section is given by three unit, pairwise, orthogonal vectors (d1, d2, d3), called directors of the filament at position s. The internal force density, strain, and moment vectors (f, κ, m) are defined in the local coordinate system as f = f1d1 + f2d2 + f3d3 (similar relations for the other vectors). The moment scales linearly with the curvature strain such that m = B(κ−κ0), where κ0 is the resting filament strain. The first two components of κ0 are zero (i.e., no bending at rest). The third component, κ30, accounts for the intrinsic filament helicity. B is the bending rigidity matrix, which reads (expressed in the local frame of directors) as
| (1) |
where CB, CT, and CTB are the bending (flexural), twisting (torsional), and the twist-bend coupling rigidity moduli, respectively (note that rigidity moduli are just proportional to the persistence length times kT, with k as the Boltzmann constant and T as the temperature in Kelvin). Equation 1 assumes that bending rigidities are identical along d1 and d2 (i.e., bending stiffness is symmetrical and equal in all directions), and that twist-bend coupling is identical for both directors (i.e., the CTB term is identical for bending strains in the d1 and d2 directions). While this assumption may seem at variance with actin filament structure, which deviates locally from radial symmetry with a roughly elliptical cross-sectional area, the shape deformations evaluated here are on length-scales much greater than the filament helical repeat, so anisotropies arising from local, noncylindrical structure and associated elasticity will be averaged (11,24,28).
The elastic free energy density (e) within segment s is given by
| (2) |
Positivity of free energy density is ensured by the following condition:
| (3) |
In simulations, we use the normalized coupling rigidity such that −1 ≤ θTB ≤ 1.
The balance of internal force (f) and moment (m) within segment s are given by
| (4) |
where fext and mext are, respectively, the external force and moment densities applied to the filament at position s. Using the definition of the moment, m(s) = B(s)(κ−κ0), the second equation in Eq. 4 can be expanded to explicitly emphasize the spatial derivative of rigidities (second term) and resting strain κ0 (third term):
| (5) |
We use fext = 0 and mext = 0 in simulations. The unit tangent vector, denoted d3, is given by
| (6) |
where r is a point along the filament. Because filaments move at low Reynolds number (∼10−5, see Table 1), we systematically eliminated from Eq. 4 all force and moment terms proportional to linear/angular acceleration.
Table 1.
Actin filament mechanical properties used in simulations
| Persistence Length | Actin | Cofilactin |
|---|---|---|
| Bend (LB) | 9.8 μma | 2.2 μmb |
| Twist (LT) | 0.5 μmc,d | 0.1 μmc |
| Twist-bend coupling (LTB) | 0.15 μmd | 0.03 μme |
Simulations were done at 25°C (kT = 4.1 × 10−21 J). The Reynolds number is typically 1.4 × 10−5 (for a filament of 10 μm moving at 1 μm.s−1 in a fluid of viscosity 10−3 Pa.s; the protein density is estimated at 1.4 kg.m−3 (56)).
McCullough et al. (24).
The filament torsional persistence length (LT) was calculated from the torsional rigidity (C) using LT = C/(kT) with kT = 4.1 × 10−21 J and C = 2.3 × 10−27 Nm2 rad−1 (23). Comparable values are predicted by molecular-dynamics simulations and normal mode analysis (C = 2.6 × 10−27 Nm2 rad−1 (26,57)).
De La Cruz et al. (22).
Calculated from |LTB| = (LTLB/2)0.5 (22)).
Quaternions (Euler parameters) and componentwise form of equations
Let q(s) = (q1(s), q2(s), q3(s), q4(s)) be a unit quaternion (i.e., ). The components of the three directors d1(s), d2(s), and d3(s) are given by
| (7) |
The curvature strain vector κ gives the rate of spatial change of the triad (d1(s), d2(s), and d3(s)) along the filament centerline: Using the expression of the director vectors (Eq. 7), and the spatial derivative of directors, yields the equation for quaternions:
| (8) |
Numerical simulations were performed using the balance of force/moment (Eq. 4), the inextensibility equations (Eq. 7, with d3 given by Eq. 6), and the direction vector quaternion (Eq. 8).
Boundary conditions
Solution of Eqs. 4–6 and 8 requires boundary conditions for the 13 variables of the system (three for position, three for internal force, four for the Euler parameter, and three for bending strains). Because these variables come in pairs (position/internal force and Euler parameter (orientation)/curvature strains)), either the position/orientation or the force/curvature strains should be constrained.
During buckling (Fig. S3), the end-to-end distance is progressively reduced by imposing the spatial position of both filament ends (six variables). The position of the barbed end is maintained at the origin while the other end is moved along the end-to-end axis in a stepwise manner. We define the position of filament ends as
| (9) |
where r0 (respectively, r1) is the position of the barbed end (respectively, of the pointed end) given by the initial configuration; L is the filament length; Δs is the buckling constraint (the actual end-to-end distance is L−Δs); and e is the unit vector along the end-to-end distance. The orientation at both ends is held constant during buckling,
| (10) |
where q0,L is a constant normalized quaternion. The normalization condition for Euler parameters,
| (11) |
holds at every position along the filament. Using Eq. 8, it is sufficient to impose this normalization at either one of the two filament ends. Hence, only three components of the quaternion are imposed at s = L (Eq. 10), yielding the 13 conditions required to find a solution to Eqs. 4–7.
Cofilin density and parameter dependence on space
The bound cofilin density is denoted by d(s), with 0 ≤ d(s) ≤ 1 (d = 0: no cofilin; d = 1: actin subunit with a bound cofilin). Transition from 0 (no cofilin bound) to 1 (cofilin occupancy) is given by (Fig. S2)
| (12) |
where s0 is the position of the transition; the term δ defines the transition steepness, given by the peak half-width Δ:
| (13) |
Because the bending rigidity coefficients (see Table 1 and Fig. S2) depend on cofilin occupancy (see Table 1, definition of bending rigidity matrix Eq. 1, and Fig. S2), the actual rigidity is interpolated according to
| (14) |
with CB1 (respectively, CB2), the bending rigidity value in absence of cofilin (respectively, in presence of cofilin). Similar relations hold for the torsional and twist-bend coupling rigidity coefficients as well as any parameter that scales with cofilin occupancy. All simulations presented here were carried out with a δ value of 10 nm.
Confinement in compliant cylindrical chamber
Model filaments were confined in a cylindrical chamber by constraining each point along the filament near the X axis during buckling with an applied force proportional to the lateral displacement of the filament in the (Y, Z) plane (Fig. S4), which enters the right-hand side of Eq. 4 as a force density term. All simulations are performed with a proportionality constant Kf of 50 pN.μm−2 (the actual force is KfD, where D is illustrated in Fig. S4), which corresponds to a moderately elastic surrounding medium (elastic modulus ∼300 Pa) (29).
Verification of numerical simulations
Numerical simulation outputs were verified by changing the step size along the filament using the MATLAB function bvp4c (The MathWorks, Natick, MA). This function solves boundary value problems and is well suited for the simulations carried out here. Our best choice for the step along the filament is 2.5 nm, which is much less than the transition width (∼2δ = 20 nm). To check the validity of our continuum approach, we simulated zero-width transition between pure actin and pure cofilactin segments using the MATLAB function bvp5c, which allows boundary points in the interval. At the internal boundary point we imposed the continuity of position (six conditions) and orientation (eight conditions, but only seven conditions apply because of quaternion normalization); no particular constraints are imposed upon the internal force and strain. No differences were found between the two approaches for all end-to-end distances and twist-bend coupling values considered in the article.
Effects of filament deformation on the severing-fragmentation free energy profile
We model filament severing as a two-state, reversible equilibrium reaction between an intact and fragmented filament, with a single transition state. We consider two mechanisms by which the elastic free energy stored in the filament-shape deformation promotes severing. In the first case, the elastic energy introduced by buckling destabilizes the intact filament (i.e., increases its energy), thereby lowering the transition state energy barrier height and facilitating filament breakage. In the second case, we treat the introduced elastic energy and its gradient along filaments as external work and force exerted on a filament subunit, respectively, analogous to single-molecule pulling-rupture experiments where applied loads modulate intersubunit bonds and intermolecular complex dissociation kinetics.
First mechanism: activation over a free energy barrier
Severing is modeled as a reversible, two-state transition between an intact and fragmented filament. We assume that the thermodynamic binding free-energy holding actin subunits together (ΔG°′) is modified by the elastic energy associated with filament deformation (ΔG°′Elastic), and therefore use the elastic energy terms (Eq. 2) to model contributions of filament strain to the severing-fragmentation reaction free-energy profile. We assume, as a first approximation, that all of the stored elastic energy contributes to the reduction of the fragmentation free-energy barrier. In doing so, we neglect potential dissipation of elastic energy to subunit deformation and/or unfolding. Therefore, the calculated changes in fragmentation rates correspond to upper-limit estimates for the effects of filament deformation on intersubunit bond rupture.
The total elastic free energy at position s, E(s), is given by the sum of local bending (EB), twisting (ET) and twist-bend coupling (ETB) elastic energies (Eq. 2):
| (15) |
For native configurations (relaxed filaments), the severing rate (kSev(Native)) is proportional to the intrinsic activation energy barrier height (ΔG‡′):
| (16) |
Filament buckling introduces elastic energy (ΔG°′Elastic) to the ground state so that the activation energy of the strained configuration(ΔG‡′Strained) becomes
| (17) |
Consequently, the stored elastic energy lowers the severing activation barrier height and accelerates severing. The strained configuration rate constant is given by
| (18) |
Combining Eqs. 16 and 17 yields the relative severing activity of strained and native filaments:
| (19) |
The predicted severing amplification factor follows the profile of ΔG°′Elastic along the filament.
Second mechanism: acceleration of filament segment dissociation by external force
To interpret and estimate contributions of the local elastic energy gradient and resultant external force to severing, we assume 1) ΔG°′Elastic introduced by compressive buckling loads is equivalent to the work done by applied external loads, as in single molecule pulling experiments, and 2) filament severing is analogous to intermolecular interface bond rupture (e.g., ligand dissociation or unfolding), for which analytical expressions have been derived (30). We adapted this theory of a single-well harmonic potential energy surface (Eq. 3 in Dudko et al. (30)) to our simulations, and express the ratio of the spontaneous (i.e., intrinsic) and loaded severing rates as
| (20) |
where kSev(Native) is the spontaneous severing rate of bare actin filaments. The activation energy ΔG‡′ is given by the general form of the Eyring-Polanyi formula,
| (21) |
using the experimentally determined values for spontaneous, bare actin filament severing (kSev(Native) = 1 × 10−6 s−1.subunit−1, ΔG‡′ = 43.27 kT (4,5). Altering the shape of the free-energy surface from a harmonic to a cubic potential (30,31) yielded identical results.
Results
Mechanically homogenous actin and cofilactin filaments
Bare and cofilin-decorated filaments (contour length = 1 μm) buckle symmetrically under ∼0.5–2 pN compressive loads (Fig. 1, A and B). These compressive forces are smaller than those generated by a single stroke of an individual myosin motor protein (∼5 pN (32)). However, the several 100-nm reductions in filament end-to-end distance require multiple myosin stepping events.
Figure 1.
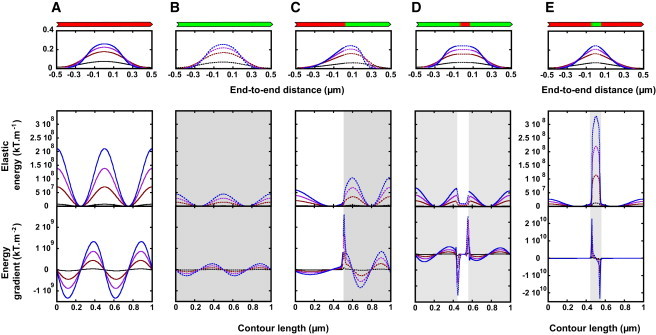
(A–E) The shape, spatial elastic energy, and local shear force of buckling (cofil)actin filaments. Filaments (length 1 μm), with different actin/cofilactin configurations (actin in red, cofilactin in green, illustrated above top panels) are buckled by reduction of the end-to-end distance (Fig. S3) by 1% (black curve), 10% (dark-red curve), 20% (purple curve), and 30% (blue curve). The shape of the filament changes with buckling (A–E, top row); note that the asymmetric configuration in (C) gives rise to asymmetric filament deformation. The total elastic energy stored in the filament configuration (middle row) and the elastic energy gradient (bottom row) for corresponding filaments (illustrated in the top row) is shown. To see this figure in color, go online.
The elastic energy stored in the filament configuration depends on the end-to-end filament length and peaks at regions of highest curvature (Fig. 1). The resulting shear force given by the energy gradient displays local maxima at the midpoints of stored elastic energy, such that the net force distributes equally at four distinct filament positions (Fig. 1). Cofilactin filaments, being more compliant (Table 1), buckle at lower loads and require less work to deform (Fig. S5), but store less energy and experience lower shear force than bare actin filaments at identical end-to-end lengths (compare Fig. 1, A and B).
Partially decorated cofilactin filaments with mechanical heterogeneity
Filaments that are partially decorated with cofilin are mechanically heterogeneous and display shape deformations and energy profiles distinct from mechanically homogenous bare (actin) or saturated (cofilactin) filaments (Fig. 1). Filaments with a single bare and cofilin-decorated segment of equal length (i.e., with a single boundary at center) buckle asymmetrically, with the compliant cofilactin segment deforming more than the stiff, bare filament segment (Fig. 1 C). This asymmetry introduces larger energy densities within the cofilactin segment and a maximal shear force adjacent to the boundary, within the compliant cofilactin segment (Fig. 1 C).
Filaments with two boundaries display two shear force peaks, with magnitudes that depend on the relative and spatial cofilin occupancy (Fig. 1, D and E). A bare filament with a small cluster of bound cofilin (cluster size in simulation is 0.05 μm, ∼16 subunits) stores an order-of-magnitude-more elastic energy and generates a 10-fold-higher shear force than a cofilactin filament with a bare segment of equivalent size (Fig. 1, D and E). Thus, all boundaries are not identical and the local buckling strain depends on the relative size as well as the bending and twisting rigidities of flanking regions.
These simulations assume filament composition is constant and that bound cofilin does not redistribute within the buckling time. Cofilin dissociation occurs on a timescale of tens of seconds (11,33). Nonmuscle myosins slide filaments at ∼170–340 nm.s−1 at near-physiological temperatures (30°C (34)), which corresponds to ∼1 s timescale to achieve the reductions in filament end-to-end lengths considered here. Therefore, the buckling times are considerably shorter than the intrinsic cofilin redistribution times, justifying the use of a constant cofilin segment length in the calculations.
Effects of buckling on filament severing
Filament deformation (i.e., strain) introduces elastic energy (Eq. 20) and tilts the fragmentation-annealing reaction free-energy profile (Fig. 2). The standard free-energy change between an intact and fragmented filament (ΔG°′) defines the severing-annealing equilibrium constant (35–37) and filament-length distribution. The severing and annealing rate constants (36–38) scale with the energy difference between ground and transition states.
Figure 2.
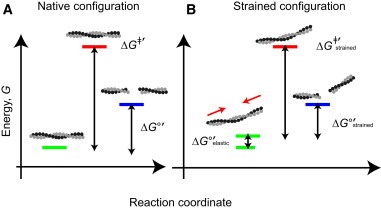
(A) Strain tilts the filament fragmentation-annealing reaction free-energy profile. Filament fragmentation is illustrated as a two-state system with intact (energy illustrated with green line) and severed (energy illustrated with blue line) filaments linked by a transition state (energy illustrated with red line). The spacings between energy states are drawn to scale and reflect relative heights estimated from the severing and annealing rate and equilibrium constants (4). (B) (Red arrows) Compressive forces applied at the two filament-ends during buckling. Note that considerable separation between the ground and transition states exists, even for strained filaments. Thus, the deformations are not irreversible (e.g., to rupture), but exist in a perturbative regime where fragmentation is likely to follow a nativelike pathway. To see this figure in color, go online.
Buckling strain destabilizes filaments (i.e., increases G), thereby lowering the net transition-state energy-barrier height for severing (ΔG‡′; green to red transition in Fig. 2 B) and accelerating filament fragmentation (39). For simplicity, we assume the transition state is not affected by strain. While this may not be an accurate assumption, it is a reasonable first approximation given that filament fragmentation is accelerated by strain (14,18,20,40). If filament strain increased the energy of the transition state equally or more than that of the ground state, loads would have no effect or inhibit severing, respectively. Filament annealing is diffusion-limited (36–38) and weakly affected by cofilin occupancy (4). If filament annealing remains diffusion-limited for bent filaments, which to the best of our knowledge is not known, the severing-annealing equilibrium constant is load-dependent and buckling strain reduces the energy difference between intact and fragmented filaments, which shortens the population average filament length.
Knowledge of the spatial filament elastic energy (Fig. 1) allows us to 1) estimate the extent to which filament-shape deformations (i.e., changes in end-to-end distance, radius of curvature or kink angle; relationship between these deformations is given in Fig. S6) accelerate fragmentation and 2) identify the most probable location of severing along strained filaments (Fig. 3). Assuming that filament severing results from rupture of one lateral and two longitudinal subunit contacts ((41); Fig. S7), and using values of −3.75 kT for lateral contacts and −12.15 kT for longitudinal contacts (Fig. S7; Table 2; (22,41)), we estimate filament severing of ADP-actin filaments to be associated with −28.05 kT (−16.61 kcal/mol) of energy.
Figure 3.
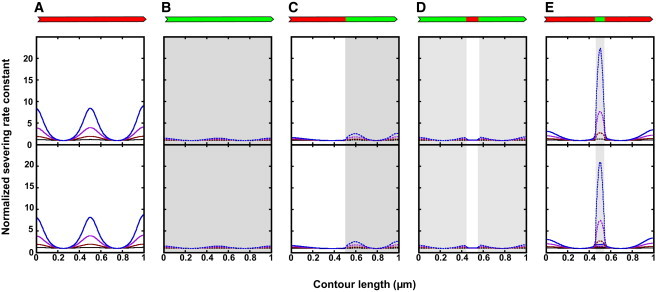
(A–E) Spatial distribution of buckled filament severing rate. The enhancement of filament severing was calculated by considering the contributions from the local elastic energy (Eq. 18; top row) and the energy gradient representing an effective force (Eq. 19; bottom row). The increase in the severing rate constant predicted from the spatial distribution of elastic energy, ksev(Strained)/ksev(Native), coincides with that of ΔG°′Elastic (top row). The distribution considering ΔG°′Elastic as the work done by a pulling force against a filament subunit yields comparable behavior. Note that both the activation energy barrier and force methods of analysis yield essentially identical results, with minor differences observed only in stiff actin filaments with a compliant cofilactin segment (right column). To see this figure in color, go online.
Table 2.
Interface and severing energy of actin filament
| Energy | Symbol | Value (kT) |
|---|---|---|
| Lateral contacta,b | E1 | −3.75 |
| Longitudinal contacta,b | E2 | −12.15 |
| Severing energyc | ΔG°′ = 2 E1 + E2 | −28.05 |
Lateral and longitudinal energies, respectively, E1 and E2, are obtained from values in Fig. S7, A and B, by E1 = ΔG°′/2 and E2 = ΔG°′2 − ΔG°′1/2.
Sept and McCammon (41).
Filament severing energy is calculated as the net rupture of two longitudinal and one lateral intersubunit contacts (Fig. S7C).
Bare actin filaments buckled to 70% of their contour lengths are predicted to fragment (approximately fivefold) more rapidly than relaxed filaments (Fig. 3), in agreement with nanomanipulation experiments in which the filament radius of curvature is continually controlled (40) or shortened with myosin motors (18). Flexible cofilactin filaments at comparable end-to-end lengths are less susceptible to fragmentation than stiff, bare filaments, as reported in McCullough et al. (5). Severing is enhanced at boundaries, although the contributions from filament strain are modest.
Filaments with small cofilin clusters are more susceptible to fragmentation under strain (Fig. 3 E) than relaxed filaments. The effect depends strongly on the size of the bound cofilin cluster. Filaments with small clusters accumulate more elastic energy, within the compliant cofilactin segment, than those with large clusters (Figs. 3 E and S6, top). Consequently, severing can be enhanced up to 1000-fold relative to the intrinsic fragmentation-rate constant of bare actin (Fig. 4). This amplification is highly nonlinear: an order-of-magnitude reduction in the cluster size yields a 100-fold increase in the maximal severing rate constant (Fig. 4). This local increase in the microscopic severing-rate constant exceeds the overall severing rate across the filament (i.e., many sites along a 1-μm filament that sever with a lower rate constant) when the bound cofilin cluster size is <0.04 μm. In other words, the local severing rate within the cluster exceeds the rate of severing elsewhere in the filament, accounting for potential severing sites, when the cluster is <0.04 μm.
Figure 4.

Spatial distribution of elastic energy density, showing severing rate along filaments with a central cofilin cluster. The total energy (top left) and the normalized severing rate constant (bottom left) values are displayed for three cofilin cluster sizes (inset shows color-code for corresponding cofilactin segment length, in micrometers). (Right) Maximal severing rate. To see this figure in color, go online.
We emphasize that these values represent lower estimates of acceleration, because boundaries between bare and cofilin-decorated segments fragment more readily than bare or cofilin-decorated (4,5), analogous to the adhesive/interfacial fracture of nonprotein materials (4,9,12).
Role of twist-bend coupling elasticity
Actin filament elasticity is dominated by three terms: bending (EB), twisting (ET), and coupling between twisting and bending (ETB (22)). Twist-bend coupling has a short persistence length (22) and is typically considered a local phenomenon (42) or ignored when modeling filament mechanical properties. However, twist-bend coupling causes bending strain in the (d1, d2) plane to generate rotation about the tangent direction along d3 (Figs. S1 and S2). As a result, pure bending deformations introduce twisting deformations and increase the shear stress about the tangent to centerline, as well as the total energy localization, thereby significantly diminishing the energy barrier for severing. Accordingly, filaments modeled without twist-bend coupling elasticity underestimate the filament elastic energy and gradients associated with deformation (Fig. 5).
Figure 5.
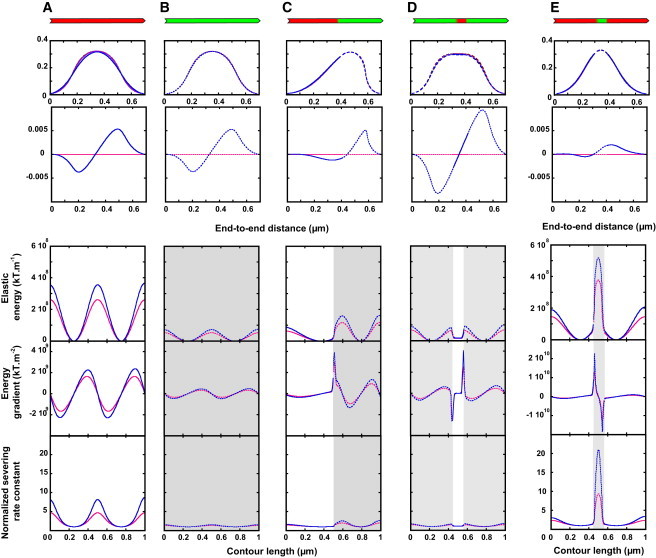
(A–E) Contributions of twist-bend coupling to filament energy profiles. Filaments (length, 1 μm), with different actin/cofilactin configurations (actin in red, cofilactin in green, illustrated above top panels), are buckled to an end-to-end distance of 0.7 μm. The two first rows represent the filament shape projected onto the X,Y and X,Z planes, respectively (Fig. S9). (Blue curves) Filaments with the mechanical properties of actin (Table 1); (red lines) filaments with neglected twist-bend coupling elasticity. To see this figure in color, go online.
Role of filament confinement
Filaments confined within a 0.2-μm radius cylinder defined by compliant walls, thereby mimicking an elastic matrix, adopt different shapes than unconstrained filaments (Fig. 6). Confinement forces compliant filaments and filament segments to adopt higher deformation modes and thus store more elastic energy. The energy within a cofilactin cluster of confined filaments is approximately twofold higher than for unconstrained filaments (Fig. 5 E). Consequently, the predicted severing rate is considerably higher during confinement than when allowed to bend freely.
Figure 6.

(A–E) Contributions of filament confinement. Filaments (length, 1 μm), with different actin/cofilactin configurations (actin in red, cofilactin in green, illustrated above top panels) are buckled to an end-to-end distance of 0.7 μm. (Blue curves) Filaments with the mechanical properties of actin (Table 1); (green lines) filament-shape deformations constrained within a rigid cylinder of radius 0.2 μm. To see this figure in color, go online.
Contributions from viscous drag
The filament end-to-end distance is reduced in a sequential stepwise manner during buckling. These stepping events correspond to myosin-driven contractile events in vivo, which are limited by ADP release kinetics of the motor (43). To evaluate potential contributions of viscous drag to filament energy and fragmentation, we consider buckling events occurring at a velocity of 300 nm.s−1, corresponding to buckling driven by double-headed nonmuscle myosin IIA or IIB (sliding velocities of ∼170–340 nm.s−1 at close to physiological temperatures; 30°C (34)). These velocities and forthcoming contributions from viscous drag represent upper estimates, because picoNewton loads can have slow ADP release from myosin motors, and thus the filament sliding velocity (44).
The velocity of any point M along a buckling filament is approximated by
| (22) |
where (t) and (t + τ) refer to two successive filament configurations separated by a single buckling step occurring within time τ (Fig. S10). At any point M on the filament, the drag force per unit length (force density) is proportional to the point velocity according to
| (23) |
where C1 (respectively, C2) is the drag coefficient per length (in units of Pa.s) associated with movement along the tangent at M (respectively, movement orthogonal to the tangent at M). Therefore, the new filament configuration r is given by the solution of Eq. 5 and a modified form of Eq. 4, which reads
| (24) |
where r0 is the known position of point M in the filament configuration computed at the buckling previous step. Note that the right-hand side of Eq. 23 is a force density term (in units of N.m−1).
The coupling between the equations of force (Eq. 23) and filament configuration (Eq. 6) is of importance in the numerical solution of equations. However, the viscous drag introduced by contractile protein-driven filament sliding velocities during buckling (Fig. S10, F–H; Table 3) is negligible. Consequently, the contributions of viscous drag to filament severing are also expected to be negligible.
Table 3.
Parameters using in simulations for viscous drag
| Persistence Length | Actin | Cofilactin |
|---|---|---|
| Filament radius, r | 3.5 nma | 4.2 nma |
| Fluid viscosity, η | 10−3 Pa.s | 10−3 Pa.s |
| Filament length, L | 1 μm | 1 μm |
| 0.0013 Pa.s | 0.0013 Pa.s | |
| 0.0025 Pa.s | 0.0025 Pa.s | |
| Time step during buckling, τ | 0.3 s | 0.3 s |
McCullough et al. (24).
Role of boundary conditions
Boundary conditions control the filament-end orientations, the associated bending and twisting strains, the elastic energy accumulation, and the severing probability. The simulations presented thus far constrain the orientation of both filament ends. To consider specific filament populations in cells that have free (e.g., by thermal fluctuations) or weakly constrained ends, such as those branching from dendritic networks, we compare the case where filament-end orientation directors are held constant (Fig. 1) with filaments that have only a single end constrained at s = 0, while bending vanishes at the other end (three conditions):
| (25) |
We consider two different actin/cofilactin distributions, corresponding either to a half-decorated filament (composition 1) or to a stretch of cofilin (size, 0.1 μm) surrounded by bare actin segments (composition 2). Because configuration 1 is asymmetric, we consider two cases, depending on whether the orientation is constrained at the bare (left; s = 0) or cofilin-decorated (right; s = L) end of the filament.
Relieving orientation constraints at one filament end lowers the accumulated elastic energy and the energy gradient (Fig. 7), because the elastic energy dissipates with reorientation of unconstrained filament ends. The extent of weakening in half-decorated filaments (Fig. 7, left) depends on which end is free to reorient, such that a free bare actin end dissipates less energy than a free cofilactin end (left column, compare blue and red curves). Therefore, the filament orientation and relative occupancy is likely a key factor in regulating fragmentation dynamics.
Figure 7.
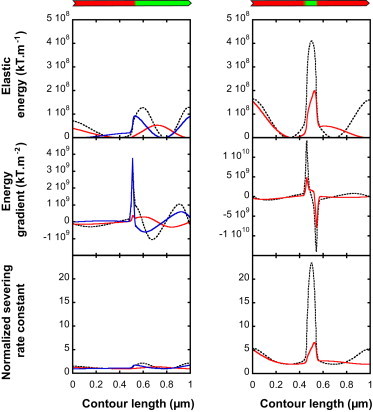
(Left) Zero-strain conditions weaken elastic energy and severing. (Color-code: dashed black curve, boundary conditions with both ends constrained; red solid curves, filaments with unconstrained, i.e., green segment, cofilactin ends (s = L); blue, filaments with unconstrained, i.e., red segment, actin ends (s = 0).) For the symmetric actin/cofilin configuration (right), only the condition corresponding to zero strain at s = 0 is shown. Note that the energy and its gradient vanish for the zero strain condition, coherent with the definition of energy (Eq. 1). To see this figure in color, go online.
Discussion
Accumulating experimental evidence, both in cells and in vitro, supports a role for myosin contractile protein-driven filament deformations in accelerating fragmentation and network reorganization (14,16–18,45). Predictions made by the theory and simulations developed here compare quantitatively with experimental measurements of contractile protein-driven filament severing events, and make additional predictions that can be tested through future experiment. The proposed mechanism of contractile protein-driven filament strain and severing would apply to the lamellipodium-lamellum junction, where both cofilin and myosin drive actin-network reorganization (16,45).
Shape deformations promote filament fragmentation
Actin and cofilactin filaments deform symmetrically under low picoNewton compressive buckling forces (Fig. 1), magnitudes comparable to the forces generated by individual myosin motors (32) and growing filaments (46). Filament buckling localizes elastic energy at regions of largest deformation and introduces a shear force (i.e., energy gradient) at several foci along filaments (Fig. 1).
Our analyses and results predict that buckling promotes bare actin filament fragmentation at curvature radii <300 nm (Fig. S8) as observed in reconstituted contractile actomyosin systems (18). Similarly, the theory and simulations predict that filament fragmentation is enhanced at kink angles >50°, as reported in McCullough et al. (5). Cofilactin filament fragmentation requires larger deformations than bare actin filaments, consistent with direct visualization of actin and cofilactin filament severing events (5).
Mechanical heterogeneity introduced with cofilin occupancy can promote severing
Filament shape deformations are predicted to enhance cofilin severing activity, lending credence to proposed in vivo severing mechanisms in which contractile and severing proteins coordinate to accelerate filament turnover (15,16,47). Filaments partially decorated with cofilin deform asymmetrically, with strain-energy and shear-force peaking adjacent to regions of mechanical discontinuities defined by boundaries of bare and cofilin-decorated segments. As a result, buckling promotes severing at boundaries within the compliant cofilactin segment, analogous to the manual snapping of uncooked asparagus.
The enhanced compliance of cofilactin allows partially decorated filaments with large cofilin clusters (e.g., a single boundary or a small bare cluster; Fig. 3) to tolerate larger deformations before fragmenting, as observed in wet-lab experiments (5). In contrast, small bound cofilin clusters within a bare filament localize the majority of buckled filament strain energy (Fig. 3) and are predicted to fragment within the cofilin cluster. Thus, filament severing efficiency and location depend on the composition and mechanical properties of regions flanking the boundaries. This behavior implicates net cofilin occupancy, rather than just the boundary density, in regulating severing under compressive loads. Accordingly, strain-dependent cofilin occupancy (48,49) or dissociation through competition with regulatory proteins (9) can potentially introduce distinct boundary species with variable flanking regions and thus modulate the overall severing efficiency. For example, preferential cofilin binding to deformed filament segments would target strained filaments for severing (e.g., Fig. 5 E). Such a mechanism would introduce an efficient feedback response for locally and rapidly reorganizing strained filament networks.
Factors contributing to filament severing by cofilin
Multiple factors—structure, elasticity, deformation, and confinement—contribute to the actin-filament-severing activity of cofilin. Structural discontinuities render boundaries between bare and cofilin-decorated segments more susceptible to fragmentation than continuous regions within bare or cofilin-decorated segments, analogous to adhesive/interface of nonprotein materials (i.e., boundaries are weak joints (4–7,12,13). The theory and simulations presented here support a potential regulatory role for contractile protein-driven filament shape deformations and cofilin-dependent alterations in filament mechanical properties. Filament strain introduced by contractile motor proteins localizes elastic energy and locally enhances fragmentation at regions of greatest deformation. Cofilin-linked alterations in filament mechanics introduce an asymmetry in filaments that modulates the energy density and gradient along strained filaments and the spatial probability of severing. Thus, discontinuities in both mechanics and structure can contribute to cofilin-dependent filament severing, with the latter presumably dominating thermally driven fragmentation events in the absence of active deformation and strain.
The calculations carried out here consider the coupling between filament bending and twisting elasticities. Although this twist-bend coupling elasticity has a rather short persistence length (<0.2 μm; Table 1) and dissipates over longer length scales, it makes significant contributions to the total and local filament elastic energy density, energy gradient, and filament severing probability. Accordingly, elasticity models of actin filaments that ignore twist-bend coupling underestimate the elastic energy profile and, presumably, its consequences.
Filaments with their end orientations constrained, as would occur with attachment to other filaments and cellular structures via cross-linking, branching, adaptor, and/or contractile proteins, store more elastic energy and are predicted to sever more readily than filaments with one or both ends free (Fig. 7). Similarly, restricting filament-bending amplitudes through confinement within a chamber (Fig. 6) or a cytoskeleton network (50,51) forces filaments to populate higher bending modes, as observed for microtubules (29) and intermediate filaments (52), store more elastic energy, and sever with distinct probabilities and locations along filaments. Thus, filaments constrained and confined within networks are predicted to be more susceptible to strain-induced fragmentation than individual filaments with one or both ends free. Consistent with this behavior, cross-linking proteins inhibit filament severing by cofilin (53) but the (cofilin)-dependent severing rate, after considering contributions from reduced filament site accessibility, appears to be accelerated in some cases (54). Both spatial confinement and local filament stiffening by cross-linking proteins could potentially account for the acceleration (13,55).
Author Contributions
E.M.D.L.C. and J.-L.M. designed and performed research; E.M.D.L.C., J.-L.M., and L.B. wrote the article.
Acknowledgments
The authors thank Ziad Ganim (Yale University) for comments on the article and discussions regarding mechanical bond rupture, Steve Girvin for his culinary insight, and the three anonymous reviewers for careful evaluation of the work.
This work was supported by grants from the National Institutes of Health (No. GM097348 awarded to E.M.D.L.C.) and the program of Complex Systems, IXXI (University of Grenoble awarded to J.-L.M.). E.M.D.L.C. and L.B. are supported by the International Projects for Scientific Cooperation from Centre National de la Recherche Scientifique.
Contributor Information
Enrique M. De La Cruz, Email: enrique.delacruz@yale.edu.
Jean-Louis Martiel, Email: jean-louis.martiel@cea.fr.
Supporting Material
References
- 1.Pollard T.D., Borisy G.G. Cellular motility driven by assembly and disassembly of actin filaments. Cell. 2003;112:453–465. doi: 10.1016/s0092-8674(03)00120-x. [DOI] [PubMed] [Google Scholar]
- 2.Blanchoin L., Boujemaa-Paterski R., Plastino J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 2014;94:235–263. doi: 10.1152/physrev.00018.2013. [DOI] [PubMed] [Google Scholar]
- 3.Galkin V.E., Orlova A., Egelman E.H. Remodeling of actin filaments by ADF/cofilin proteins. Proc. Natl. Acad. Sci. USA. 2011;108:20568–20572. doi: 10.1073/pnas.1110109108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kang H., Bradley M.J., De La Cruz E.M. Site-specific cation release drives actin filament severing by vertebrate cofilin. Proc. Natl. Acad. Sci. USA. 2014;111:17821–17826. doi: 10.1073/pnas.1413397111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.McCullough B.R., Grintsevich E.E., De La Cruz E.M. Cofilin-linked changes in actin filament flexibility promote severing. Biophys. J. 2011;101:151–159. doi: 10.1016/j.bpj.2011.05.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Andrianantoandro E., Pollard T.D. Mechanism of actin filament turnover by severing and nucleation at different concentrations of ADF/cofilin. Mol. Cell. 2006;24:13–23. doi: 10.1016/j.molcel.2006.08.006. [DOI] [PubMed] [Google Scholar]
- 7.De La Cruz E.M. Cofilin binding to muscle and non-muscle actin filaments: isoform-dependent cooperative interactions. J. Mol. Biol. 2005;346:557–564. doi: 10.1016/j.jmb.2004.11.065. [DOI] [PubMed] [Google Scholar]
- 8.Suarez C., Roland J., Blanchoin L. Cofilin tunes the nucleotide state of actin filaments and severs at bare and decorated segment boundaries. Curr. Biol. 2011;21:862–868. doi: 10.1016/j.cub.2011.03.064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Elam W.A., Kang H., De La Cruz E.M. Competitive displacement of cofilin can promote actin filament severing. Biochem. Biophys. Res. Commun. 2013;438:728–731. doi: 10.1016/j.bbrc.2013.07.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nadkarni A.V., Brieher W.M. Aip1 destabilizes cofilin-saturated actin filaments by severing and accelerating monomer dissociation from ends. Curr. Biol. 2014;24:2749–2757. doi: 10.1016/j.cub.2014.09.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.De La Cruz E.M., Sept D. The kinetics of cooperative cofilin binding reveals two states of the cofilin-actin filament. Biophys. J. 2010;98:1893–1901. doi: 10.1016/j.bpj.2010.01.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.De La Cruz E.M. How cofilin severs an actin filament. Biophys. Rev. 2009;1:51–59. doi: 10.1007/s12551-009-0008-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Elam W.A., Kang H., De la Cruz E.M. Biophysics of actin filament severing by cofilin. FEBS Lett. 2013;587:1215–1219. doi: 10.1016/j.febslet.2013.01.062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Reymann A.C., Boujemaa-Paterski R., Blanchoin L. Actin network architecture can determine myosin motor activity. Science. 2012;336:1310–1314. doi: 10.1126/science.1221708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wilson C.A., Tsuchida M.A., Theriot J.A. Myosin II contributes to cell-scale actin network treadmilling through network disassembly. Nature. 2010;465:373–377. doi: 10.1038/nature08994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.van Goor D., Hyland C., Forscher P. The role of actin turnover in retrograde actin network flow in neuronal growth cones. PLoS ONE. 2012;7:e30959. doi: 10.1371/journal.pone.0030959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Medeiros N.A., Burnette D.T., Forscher P. Myosin II functions in actin-bundle turnover in neuronal growth cones. Nat. Cell Biol. 2006;8:215–226. doi: 10.1038/ncb1367. [DOI] [PubMed] [Google Scholar]
- 18.Murrell M.P., Gardel M.L. F-actin buckling coordinates contractility and severing in a biomimetic actomyosin cortex. Proc. Natl. Acad. Sci. USA. 2012;109:20820–20825. doi: 10.1073/pnas.1214753109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Abu Shah E., Keren K. Symmetry breaking in reconstituted actin cortices. eLife. 2014;3:e01433. doi: 10.7554/eLife.01433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vogel S.K., Petrasek Z., Schwille P. Myosin motors fragment and compact membrane-bound actin filaments. eLife. 2013;2:e00116. doi: 10.7554/eLife.00116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Haviv L., Gillo D., Bernheim-Groswasser A. A cytoskeletal demolition worker: myosin II acts as an actin depolymerization agent. J. Mol. Biol. 2008;375:325–330. doi: 10.1016/j.jmb.2007.09.066. [DOI] [PubMed] [Google Scholar]
- 22.De La Cruz E.M., Roland J., Martiel J.L. Origin of twist-bend coupling in actin filaments. Biophys. J. 2010;99:1852–1860. doi: 10.1016/j.bpj.2010.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Prochniewicz E., Janson N., De la Cruz E.M. Cofilin increases the torsional flexibility and dynamics of actin filaments. J. Mol. Biol. 2005;353:990–1000. doi: 10.1016/j.jmb.2005.09.021. [DOI] [PubMed] [Google Scholar]
- 24.McCullough B.R., Blanchoin L., De la Cruz E.M. Cofilin increases the bending flexibility of actin filaments: implications for severing and cell mechanics. J. Mol. Biol. 2008;381:550–558. doi: 10.1016/j.jmb.2008.05.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pfaendtner J., De La Cruz E.M., Voth G.A. Actin filament remodeling by actin depolymerization factor/cofilin. Proc. Natl. Acad. Sci. USA. 2010;107:7299–7304. doi: 10.1073/pnas.0911675107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fan J., Saunders M.G., Voth G.A. Molecular origins of cofilin-linked changes in actin filament mechanics. J. Mol. Biol. 2013;425:1225–1240. doi: 10.1016/j.jmb.2013.01.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Berro J., Michelot A., Martiel J.L. Attachment conditions control actin filament buckling and the production of forces. Biophys. J. 2007;92:2546–2558. doi: 10.1529/biophysj.106.094672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Howard J. Sinauer; Sunderland, MA: 2001. Mechanics of Motor Protein and the Cytoskeleton. [Google Scholar]
- 29.Brangwynne C.P., MacKintosh F.C., Weitz D.A. Microtubules can bear enhanced compressive loads in living cells because of lateral reinforcement. J. Cell Biol. 2006;173:733–741. doi: 10.1083/jcb.200601060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dudko O.K., Hummer G., Szabo A. Intrinsic rates and activation free energies from single-molecule pulling experiments. Phys. Rev. Lett. 2006;96:108101. doi: 10.1103/PhysRevLett.96.108101. [DOI] [PubMed] [Google Scholar]
- 31.Pierse C.A., Dudko O.K. Kinetics and energetics of biomolecular folding and binding. Biophys. J. 2013;105:L19–L22. doi: 10.1016/j.bpj.2013.09.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.De La Cruz E.M., Ostap E.M. Relating biochemistry and function in the myosin superfamily. Curr. Opin. Cell Biol. 2004;16:61–67. doi: 10.1016/j.ceb.2003.11.011. [DOI] [PubMed] [Google Scholar]
- 33.Cao W., Goodarzi J.P., De La Cruz E.M. Energetics and kinetics of cooperative cofilin-actin filament interactions. J. Mol. Biol. 2006;361:257–267. doi: 10.1016/j.jmb.2006.06.019. [DOI] [PubMed] [Google Scholar]
- 34.Kim K.Y., Kawamoto S., Adelstein R.S. The B2 alternatively spliced isoform of nonmuscle myosin II-B lacks actin-activated MgATPase activity and in vitro motility. Biochem. Biophys. Res. Commun. 2008;369:124–134. doi: 10.1016/j.bbrc.2007.11.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kinosian H.J., Selden L.A., Gershman L.C. Actin filament annealing in the presence of ATP and phalloidin. Biochemistry. 1993;32:12353–12357. doi: 10.1021/bi00097a011. [DOI] [PubMed] [Google Scholar]
- 36.Hill T.L. Length dependence of rate constants for end-to-end association and dissociation of equilibrium linear aggregates. Biophys. J. 1983;44:285–288. doi: 10.1016/S0006-3495(83)84301-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Sept D., Xu J., McCammon J.A. Annealing accounts for the length of actin filaments formed by spontaneous polymerization. Biophys. J. 1999;77:2911–2919. doi: 10.1016/s0006-3495(99)77124-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Andrianantoandro E., Blanchoin L., Pollard T.D. Kinetic mechanism of end-to-end annealing of actin filaments. J. Mol. Biol. 2001;312:721–730. doi: 10.1006/jmbi.2001.5005. [DOI] [PubMed] [Google Scholar]
- 39.Davis L.J., Odde D.J., Gross S.P. The importance of lattice defects in katanin-mediated microtubule severing in vitro. Biophys. J. 2002;82:2916–2927. doi: 10.1016/S0006-3495(02)75632-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Arai Y., Yasuda R., Itoh H. Tying a molecular knot with optical tweezers. Nature. 1999;399:446–448. doi: 10.1038/20894. [DOI] [PubMed] [Google Scholar]
- 41.Sept D., McCammon J.A. Thermodynamics and kinetics of actin filament nucleation. Biophys. J. 2001;81:667–674. doi: 10.1016/S0006-3495(01)75731-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Yogurtcu O.N., Kim J.S., Sun S.X. A mechanochemical model of actin filaments. Biophys. J. 2012;103:719–727. doi: 10.1016/j.bpj.2012.07.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Siemankowski R.F., Wiseman M.O., White H.D. ADP dissociation from actomyosin subfragment 1 is sufficiently slow to limit the unloaded shortening velocity in vertebrate muscle. Proc. Natl. Acad. Sci. USA. 1985;82:658–662. doi: 10.1073/pnas.82.3.658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Greenberg M.J., Shuman H., Ostap E.M. Inherent force-dependent properties of β-cardiac myosin contribute to the force-velocity relationship of cardiac muscle. Biophys. J. 2014;107:L41–L44. doi: 10.1016/j.bpj.2014.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Burnette D.T., Ji L., Forscher P. Myosin II activity facilitates microtubule bundling in the neuronal growth cone neck. Dev. Cell. 2008;15:163–169. doi: 10.1016/j.devcel.2008.05.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kovar D.R., Pollard T.D. Insertional assembly of actin filament barbed ends in association with formins produces picoNewton forces. Proc. Natl. Acad. Sci. USA. 2004;101:14725–14730. doi: 10.1073/pnas.0405902101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wiggan O., Shaw A.E., Bamburg J.R. ADF/cofilin regulates actomyosin assembly through competitive inhibition of myosin II binding to F-actin. Dev. Cell. 2012;22:530–543. doi: 10.1016/j.devcel.2011.12.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hayakawa K., Tatsumi H., Sokabe M. Actin filaments function as a tension sensor by tension-dependent binding of cofilin to the filament. J. Cell Biol. 2011;195:721–727. doi: 10.1083/jcb.201102039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hayakawa K., Tatsumi H., Sokabe M. Mechano-sensing by actin filaments and focal adhesion proteins. Commun. Integr. Biol. 2012;5:572–577. doi: 10.4161/cib.21891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Vonna L., Limozin L., Sackmann E. Single-filament dynamics and long-range ordering of semiflexible biopolymers under flow and confinement. Langmuir. 2005;21:9635–9643. doi: 10.1021/la050986w. [DOI] [PubMed] [Google Scholar]
- 51.Arsenault M.E., Zhao H., Bau H.H. Confinement and manipulation of actin filaments by electric fields. Biophys. J. 2007;93:L42–L44. doi: 10.1529/biophysj.107.114538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Nöding B., Köster S. Intermediate filaments in small configuration spaces. Phys. Rev. Lett. 2012;108:088101. doi: 10.1103/PhysRevLett.108.088101. [DOI] [PubMed] [Google Scholar]
- 53.Schmoller K.M., Semmrich C., Bausch A.R. Slow-down of actin depolymerization by cross-linking molecules. J. Struct. Biol. 2011;173:350–357. doi: 10.1016/j.jsb.2010.09.003. [DOI] [PubMed] [Google Scholar]
- 54.Breitsprecher D., Koestler S.A., Faix J. Cofilin cooperates with fascin to disassemble filopodial actin filaments. J. Cell Sci. 2011;124:3305–3318. doi: 10.1242/jcs.086934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Miklavc P., Ehinger K., Frick M. Actin depolymerization and crosslinking join forces with myosin II to contract actin coats on fused secretory vesicles. J. Cell Sci. 2015;128:1193–1203. doi: 10.1242/jcs.165571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Demé B., Hess D., Sackmann E. Binding of actin filaments to changed lipid monolayers: film balance experiments combined with neutron reflectivity. Eur. Phys. J. E. 2000;2:125–136. [Google Scholar]
- 57.Deriu M.A., Bidone T.C., Morbiducci U. Biomechanics of actin filaments: a computational multi-level study. J. Biomech. 2011;44:630–636. doi: 10.1016/j.jbiomech.2010.11.014. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


