Abstract
The chloroplast genome (cpDNA) of plants has been a focus of research in plant molecular evolution and systematics. Several features of this genome have facilitated molecular evolutionary analyses. First, the genome is small and constitutes an abundant component of cellular DNA. Second, the chloroplast genome has been extensively characterized at the molecular level providing the basic information to support comparative evolutionary research. And third, rates of nucleotide substitution are relatively slow and therefore provide the appropriate window of resolution to study plant phylogeny at deep levels of evolution. Despite a conservative rate of evolution and a relatively stable gene content, comparative molecular analyses reveal complex patterns of mutational changes. Non-coding regions of cpDNA diverge through insertion/deletion changes that are sometimes site dependent. Coding genes exhibit different patterns of codon bias that appear to violate the equilibrium assumptions of some evolutionary models. Rates of molecular change often vary among plant families and orders in a manner that violates the assumption of a simple molecular clock. Finally, protein-coding genes exhibit patterns of amino acid change that appear to depend on protein structure, and these patterns may reveal subtle aspects of structure/function relationships. Only comparative studies of molecular sequences have the resolution to reveal this underlying complexity. A complete description of the complexity of molecular change is essential to a full understanding of the mechanisms of evolutionary change and in the formulation of realistic models of mutational processes.
Full text
PDF



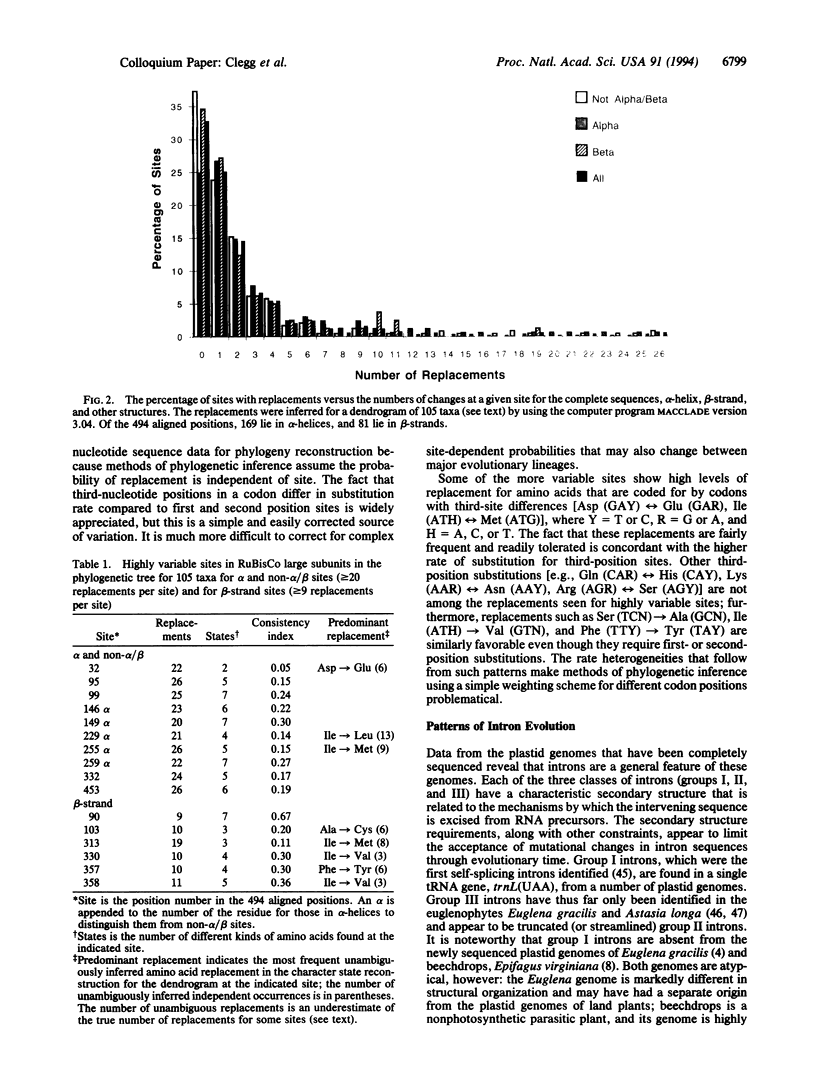


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bousquet J., Strauss S. H., Doerksen A. H., Price R. A. Extensive variation in evolutionary rate of rbcL gene sequences among seed plants. Proc Natl Acad Sci U S A. 1992 Aug 15;89(16):7844–7848. doi: 10.1073/pnas.89.16.7844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bousquet J., Strauss S. H., Li P. Complete congruence between morphological and rbcL-based molecular phylogenies in birches and related species (Betulaceae). Mol Biol Evol. 1992 Nov;9(6):1076–1088. doi: 10.1093/oxfordjournals.molbev.a040779. [DOI] [PubMed] [Google Scholar]
- Bowman C. M., Barker R. F., Dyer T. A. In wheat ctDNA, segments of ribosomal protein genes are dispersed repeats, probably conserved by nonreciprocal recombination. Curr Genet. 1988 Aug;14(2):127–136. doi: 10.1007/BF00569336. [DOI] [PubMed] [Google Scholar]
- Bulmer M., Wolfe K. H., Sharp P. M. Synonymous nucleotide substitution rates in mammalian genes: implications for the molecular clock and the relationship of mammalian orders. Proc Natl Acad Sci U S A. 1991 Jul 15;88(14):5974–5978. doi: 10.1073/pnas.88.14.5974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapman M. S., Suh S. W., Curmi P. M., Cascio D., Smith W. W., Eisenberg D. S. Tertiary structure of plant RuBisCO: domains and their contacts. Science. 1988 Jul 1;241(4861):71–74. doi: 10.1126/science.3133767. [DOI] [PubMed] [Google Scholar]
- Christopher D. A., Hallick R. B. Euglena gracilis chloroplast ribosomal protein operon: a new chloroplast gene for ribosomal protein L5 and description of a novel organelle intron category designated group III. Nucleic Acids Res. 1989 Oct 11;17(19):7591–7608. doi: 10.1093/nar/17.19.7591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clegg M. T. Chloroplast gene sequences and the study of plant evolution. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):363–367. doi: 10.1073/pnas.90.2.363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gantt J. S., Baldauf S. L., Calie P. J., Weeden N. F., Palmer J. D. Transfer of rpl22 to the nucleus greatly preceded its loss from the chloroplast and involved the gain of an intron. EMBO J. 1991 Oct;10(10):3073–3078. doi: 10.1002/j.1460-2075.1991.tb07859.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaut B. S., Muse S. V., Clark W. D., Clegg M. T. Relative rates of nucleotide substitution at the rbcL locus of monocotyledonous plants. J Mol Evol. 1992 Oct;35(4):292–303. doi: 10.1007/BF00161167. [DOI] [PubMed] [Google Scholar]
- Gillespie J. H. Natural selection and the molecular clock. Mol Biol Evol. 1986 Mar;3(2):138–155. doi: 10.1093/oxfordjournals.molbev.a040382. [DOI] [PubMed] [Google Scholar]
- Golenberg E. M., Clegg M. T., Durbin M. L., Doebley J., Ma D. P. Evolution of a noncoding region of the chloroplast genome. Mol Phylogenet Evol. 1993 Mar;2(1):52–64. doi: 10.1006/mpev.1993.1006. [DOI] [PubMed] [Google Scholar]
- Gray M. W. Origin and evolution of organelle genomes. Curr Opin Genet Dev. 1993 Dec;3(6):884–890. doi: 10.1016/0959-437x(93)90009-e. [DOI] [PubMed] [Google Scholar]
- Hallick R. B., Hong L., Drager R. G., Favreau M. R., Monfort A., Orsat B., Spielmann A., Stutz E. Complete sequence of Euglena gracilis chloroplast DNA. Nucleic Acids Res. 1993 Jul 25;21(15):3537–3544. doi: 10.1093/nar/21.15.3537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Higgins D. G., Bleasby A. J., Fuchs R. CLUSTAL V: improved software for multiple sequence alignment. Comput Appl Biosci. 1992 Apr;8(2):189–191. doi: 10.1093/bioinformatics/8.2.189. [DOI] [PubMed] [Google Scholar]
- Hiratsuka J., Shimada H., Whittier R., Ishibashi T., Sakamoto M., Mori M., Kondo C., Honji Y., Sun C. R., Meng B. Y. The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet. 1989 Jun;217(2-3):185–194. doi: 10.1007/BF02464880. [DOI] [PubMed] [Google Scholar]
- Jacquier A., Michel F. Multiple exon-binding sites in class II self-splicing introns. Cell. 1987 Jul 3;50(1):17–29. doi: 10.1016/0092-8674(87)90658-1. [DOI] [PubMed] [Google Scholar]
- Jarrell K. A., Dietrich R. C., Perlman P. S. Group II intron domain 5 facilitates a trans-splicing reaction. Mol Cell Biol. 1988 Jun;8(6):2361–2366. doi: 10.1128/mcb.8.6.2361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knight S., Andersson I., Brändén C. I. Reexamination of the Three-Dimensional Structure of the Small Subunit of RuBisCo from Higher Plants. Science. 1989 May 12;244(4905):702–705. doi: 10.1126/science.244.4905.702. [DOI] [PubMed] [Google Scholar]
- Kwakman J. H., Konings D., Pel H. J., Grivell L. A. Structure-function relationships in a self-splicing group II intron: a large part of domain II of the mitochondrial intron aI5 is not essential for self-splicing. Nucleic Acids Res. 1989 Jun 12;17(11):4205–4216. doi: 10.1093/nar/17.11.4205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Learn G. H., Jr, Shore J. S., Furnier G. R., Zurawski G., Clegg M. T. Constraints on the evolution of plastid introns: the group II intron in the gene encoding tRNA-Val(UAC). Mol Biol Evol. 1992 Sep;9(5):856–871. doi: 10.1093/oxfordjournals.molbev.a040765. [DOI] [PubMed] [Google Scholar]
- Li W. H. So, what about the molecular clock hypothesis? Curr Opin Genet Dev. 1993 Dec;3(6):896–901. doi: 10.1016/0959-437x(93)90011-d. [DOI] [PubMed] [Google Scholar]
- Michel F., Umesono K., Ozeki H. Comparative and functional anatomy of group II catalytic introns--a review. Gene. 1989 Oct 15;82(1):5–30. doi: 10.1016/0378-1119(89)90026-7. [DOI] [PubMed] [Google Scholar]
- Morton B. R. Chloroplast DNA codon use: evidence for selection at the psb A locus based on tRNA availability. J Mol Evol. 1993 Sep;37(3):273–280. doi: 10.1007/BF00175504. [DOI] [PubMed] [Google Scholar]
- Morton B. R., Clegg M. T. A chloroplast DNA mutational hotspot and gene conversion in a noncoding region near rbcL in the grass family (Poaceae). Curr Genet. 1993 Oct;24(4):357–365. doi: 10.1007/BF00336789. [DOI] [PubMed] [Google Scholar]
- Morton B. R. Codon use and the rate of divergence of land plant chloroplast genes. Mol Biol Evol. 1994 Mar;11(2):231–238. doi: 10.1093/oxfordjournals.molbev.a040105. [DOI] [PubMed] [Google Scholar]
- Mullet J. E., Klein R. R. Transcription and RNA stability are important determinants of higher plant chloroplast RNA levels. EMBO J. 1987 Jun;6(6):1571–1579. doi: 10.1002/j.1460-2075.1987.tb02402.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muse S. V., Weir B. S. Testing for equality of evolutionary rates. Genetics. 1992 Sep;132(1):269–276. doi: 10.1093/genetics/132.1.269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogihara Y., Terachi T., Sasakuma T. Intramolecular recombination of chloroplast genome mediated by short direct-repeat sequences in wheat species. Proc Natl Acad Sci U S A. 1988 Nov;85(22):8573–8577. doi: 10.1073/pnas.85.22.8573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogihara Y., Terachi T., Sasakuma T. Structural analysis of length mutations in a hot-spot region of wheat chloroplast DNAs. Curr Genet. 1992 Sep;22(3):251–258. doi: 10.1007/BF00351733. [DOI] [PubMed] [Google Scholar]
- Palmer J. D. Comparative organization of chloroplast genomes. Annu Rev Genet. 1985;19:325–354. doi: 10.1146/annurev.ge.19.120185.001545. [DOI] [PubMed] [Google Scholar]
- Rodermel S. R., Bogorad L. Molecular evolution and nucleotide sequences of the maize plastid genes for the alpha subunit of CF1 (atpA) and the proteolipid subunit of CF0 (atpH). Genetics. 1987 May;116(1):127–139. doi: 10.1093/genetics/116.1.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarich V. M., Wilson A. C. Generation time and genomic evolution in primates. Science. 1973 Mar 16;179(4078):1144–1147. doi: 10.1126/science.179.4078.1144. [DOI] [PubMed] [Google Scholar]
- Schmelzer C., Müller M. W. Self-splicing of group II introns in vitro: lariat formation and 3' splice site selection in mutant RNAs. Cell. 1987 Dec 4;51(5):753–762. doi: 10.1016/0092-8674(87)90098-5. [DOI] [PubMed] [Google Scholar]
- Schneider G., Lindqvist Y., Lundqvist T. Crystallographic refinement and structure of ribulose-1,5-bisphosphate carboxylase from Rhodospirillum rubrum at 1.7 A resolution. J Mol Biol. 1990 Feb 20;211(4):989–1008. doi: 10.1016/0022-2836(90)90088-4. [DOI] [PubMed] [Google Scholar]
- Shinozaki K., Ohme M., Tanaka M., Wakasugi T., Hayashida N., Matsubayashi T., Zaita N., Chunwongse J., Obokata J., Yamaguchi-Shinozaki K. The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J. 1986 Sep;5(9):2043–2049. doi: 10.1002/j.1460-2075.1986.tb04464.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siemeister G., Buchholz C., Hachtel W. Genes for the plastid elongation factor Tu and ribosomal protein S7 and six tRNA genes on the 73 kb DNA from Astasia longa that resembles the chloroplast DNA of Euglena. Mol Gen Genet. 1990 Feb;220(3):425–432. doi: 10.1007/BF00391749. [DOI] [PubMed] [Google Scholar]
- Soltis D. E., Soltis P. S., Clegg M. T., Durbin M. rbcL sequence divergence and phylogenetic relationships in Saxifragaceae sensu lato. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4640–4644. doi: 10.1073/pnas.87.12.4640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tajima F. Simple methods for testing the molecular evolutionary clock hypothesis. Genetics. 1993 Oct;135(2):599–607. doi: 10.1093/genetics/135.2.599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Umesono K., Inokuchi H., Shiki Y., Takeuchi M., Chang Z., Fukuzawa H., Kohchi T., Shirai H., Ohyama K., Ozeki H. Structure and organization of Marchantia polymorpha chloroplast genome. II. Gene organization of the large single copy region from rps'12 to atpB. J Mol Biol. 1988 Sep 20;203(2):299–331. doi: 10.1016/0022-2836(88)90002-2. [DOI] [PubMed] [Google Scholar]
- Wilson M. A., Gaut B., Clegg M. T. Chloroplast DNA evolves slowly in the palm family (Arecaceae). Mol Biol Evol. 1990 Jul;7(4):303–314. doi: 10.1093/oxfordjournals.molbev.a040605. [DOI] [PubMed] [Google Scholar]
- Wolfe K. H., Morden C. W., Palmer J. D. Function and evolution of a minimal plastid genome from a nonphotosynthetic parasitic plant. Proc Natl Acad Sci U S A. 1992 Nov 15;89(22):10648–10652. doi: 10.1073/pnas.89.22.10648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C. I., Li W. H. Evidence for higher rates of nucleotide substitution in rodents than in man. Proc Natl Acad Sci U S A. 1985 Mar;82(6):1741–1745. doi: 10.1073/pnas.82.6.1741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaug A. J., Cech T. R. In vitro splicing of the ribosomal RNA precursor in nuclei of Tetrahymena. Cell. 1980 Feb;19(2):331–338. doi: 10.1016/0092-8674(80)90507-3. [DOI] [PubMed] [Google Scholar]



