Abstract
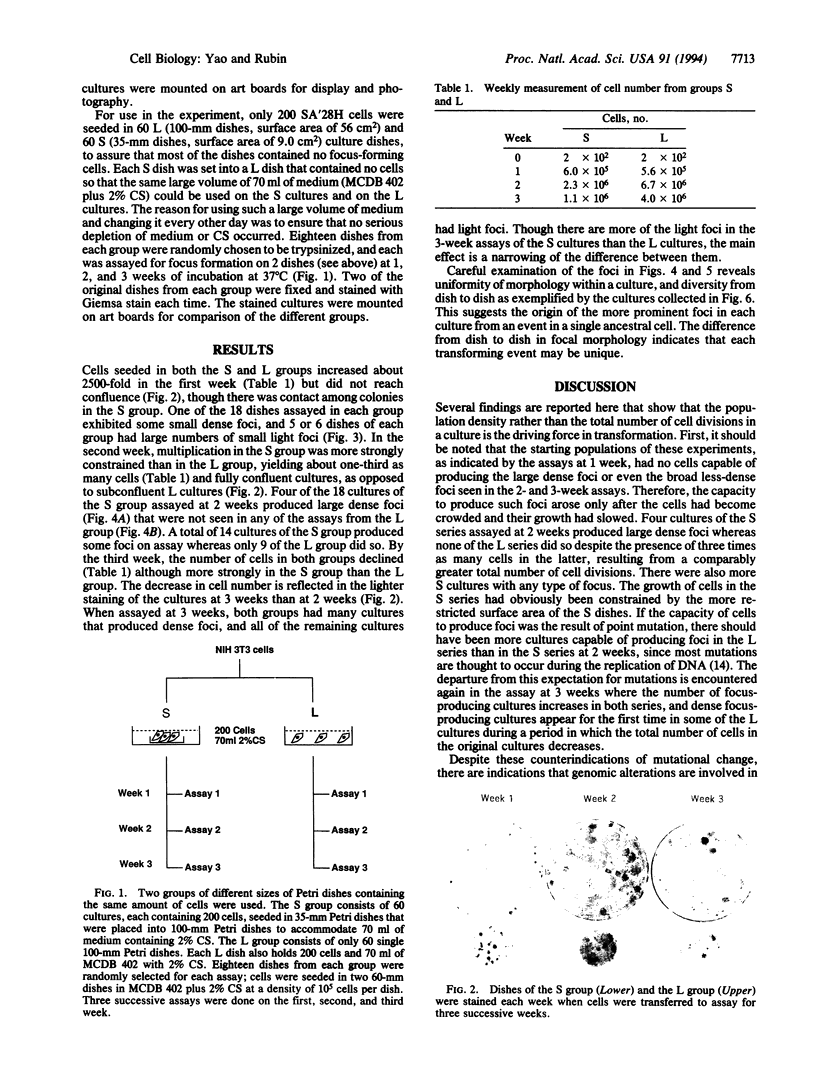
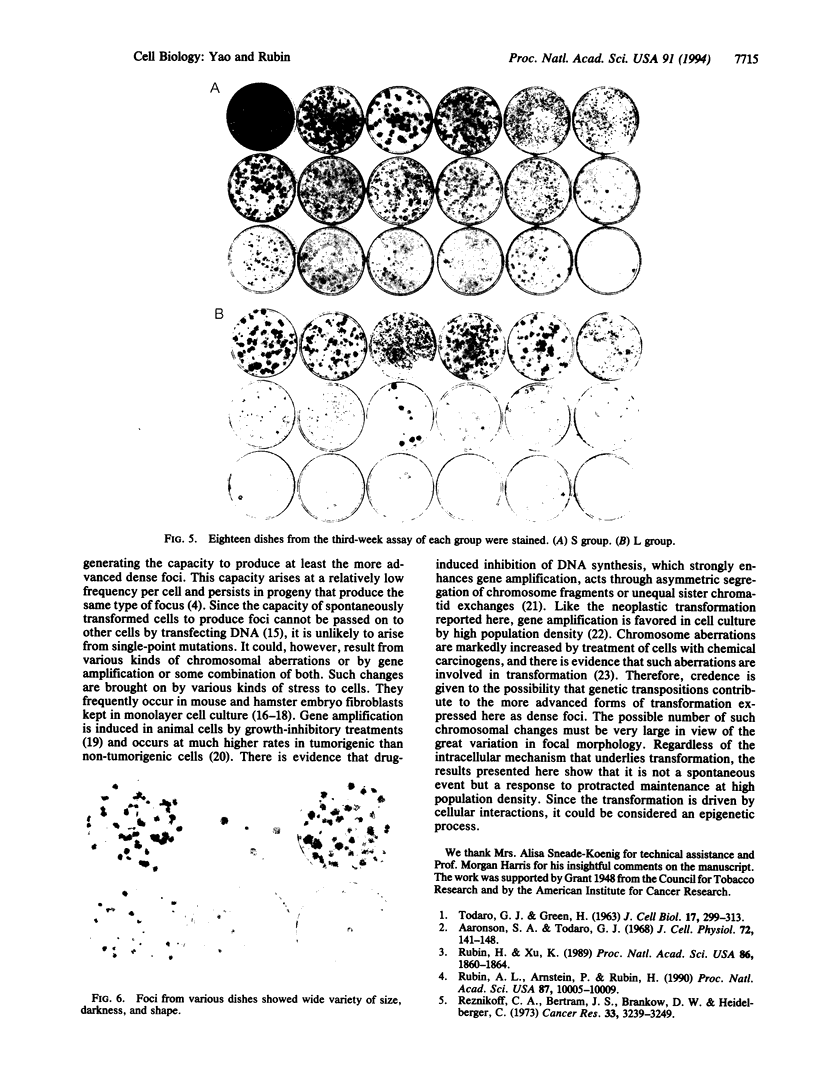
Cells of the NIH 3T3 line gain the capacity to produce neoplastically transformed foci when they are maintained at high density for more than 1 week and transferred in a standard assay for focus formation. This change in cell behavior has been variously attributed to an adaptive response to the constraint of the high population density or to a spontaneous genetic change that increases in probability for a culture with the increase in the total number of cell divisions. To distinguish between these alternatives, 200 cells of the 28H subline were seeded in many culture dishes of two size classes differing 6-fold in surface area and allowed to multiply for 1, 2, and 3 weeks. At each weekly interval, 18 dishes of each class were assayed for focus formation, and two of the original dishes were stained for focus formation. The cells in the small (S) and large (L) dishes multiplied to the same extent at 1 week and produced only a few small light foci in some of the assay dishes. At 2 weeks, cells in the S dishes had become confluent and had only one-third the number of cells as those in the nonconfluent L dishes. Upon assay, 14 of the 18 S cultures produced some foci whereas only 9 of the L cultures did so. In addition, 4 of the S cultures produced large dense foci while none of the L cultures did. By 3 weeks, the L cultures were confluent and had four times as many cells as the S cultures. When assayed at this time, both sets produced dense foci in many of the cultures and light foci in the remaining ones, indicating a narrowing of the differences between the S and L cultures between 2 and 3 weeks of incubation. There were differences in the morphology of the foci produced in parallel assays from different cultures. The results showed that transformation is a diverse graded response to the growth constraint of high population density and not a spontaneous event dependent on the number of cell divisions in a cell culture. Transformation thus is basically an epigenetic process since it represents a response to physiological restraint, but the final form of response may be modulated by genetic alterations.
Full text
PDF

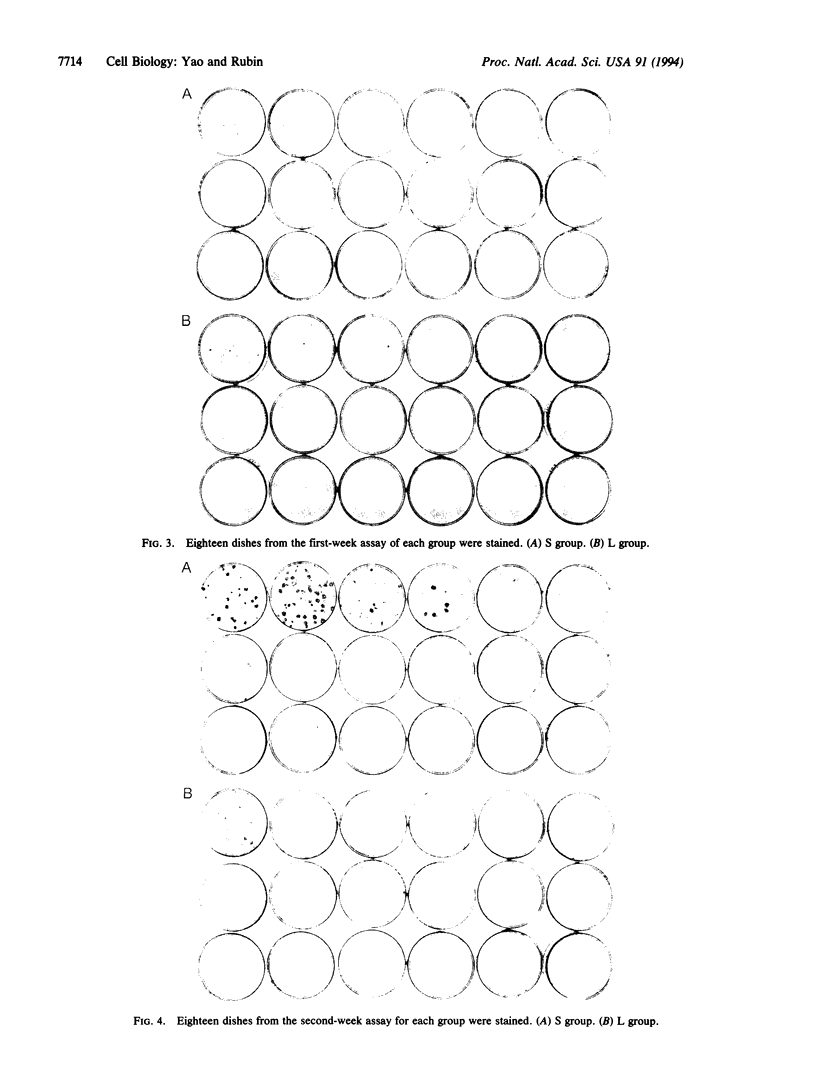


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aaronson S. A., Todaro G. J. Development of 3T3-like lines from Balb-c mouse embryo cultures: transformation susceptibility to SV40. J Cell Physiol. 1968 Oct;72(2):141–148. doi: 10.1002/jcp.1040720208. [DOI] [PubMed] [Google Scholar]
- Copeland N. G., Zelenetz A. D., Cooper G. M. Transformation of NIH/3T3 mouse cells by DNA of Rous sarcoma virus. Cell. 1979 Aug;17(4):993–1002. doi: 10.1016/0092-8674(79)90338-6. [DOI] [PubMed] [Google Scholar]
- Cram L. S., Bartholdi M. F., Ray F. A., Travis G. L., Kraemer P. M. Spontaneous neoplastic evolution of Chinese hamster cells in culture: multistep progression of karyotype. Cancer Res. 1983 Oct;43(10):4828–4837. [PubMed] [Google Scholar]
- Fernandez A., Mondal S., Heidelberger C. Probabilistic view of the transformation of cultured C3H/10T1/2 mouse embryo fibroblasts by 3-methylcholanthrene. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7272–7276. doi: 10.1073/pnas.77.12.7272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haber D. A., Fox D. A., Dynan W. S., Thilly W. G. Cell density dependence of focus formation in the C3H/10T1/2 transformation assay. Cancer Res. 1977 Jun;37(6):1644–1648. [PubMed] [Google Scholar]
- Hahn P., Kapp L. N., Morgan W. F., Painter R. B. Chromosomal changes without DNA overproduction in hydroxyurea-treated mammalian cells: implications for gene amplification. Cancer Res. 1986 Sep;46(9):4607–4612. [PubMed] [Google Scholar]
- Kennedy A. R., Cairns J., Little J. B. Timing of the steps in transformation of C3H 10T 1/2 cells by X-irradiation. Nature. 1984 Jan 5;307(5946):85–86. doi: 10.1038/307085a0. [DOI] [PubMed] [Google Scholar]
- Kennedy A. R., Fox M., Murphy G., Little J. B. Relationship between x-ray exposure and malignant transformation in C3H 10T1/2 cells. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7262–7266. doi: 10.1073/pnas.77.12.7262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kinsella A. R., Radman M. Inhibition of carcinogen-induced chromosomal aberrations by an anticarcinogenic protease inhibitor. Proc Natl Acad Sci U S A. 1980 Jun;77(6):3544–3547. doi: 10.1073/pnas.77.6.3544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LEVAN A., BIESELE J. J. Role of chromosomes in cancerogenesis, as studied in serial tissue culture of mammalian cells. Ann N Y Acad Sci. 1958 Sep 30;71(6):1022–1053. doi: 10.1111/j.1749-6632.1958.tb46820.x. [DOI] [PubMed] [Google Scholar]
- Reznikoff C. A., Bertram J. S., Brankow D. W., Heidelberger C. Quantitative and qualitative studies of chemical transformation of cloned C3H mouse embryo cells sensitive to postconfluence inhibition of cell division. Cancer Res. 1973 Dec;33(12):3239–3249. [PubMed] [Google Scholar]
- Reznikoff C. A., Brankow D. W., Heidelberger C. Establishment and characterization of a cloned line of C3H mouse embryo cells sensitive to postconfluence inhibition of division. Cancer Res. 1973 Dec;33(12):3231–3238. [PubMed] [Google Scholar]
- Rubin A. L., Arnstein P., Rubin H. Physiological induction and reversal of focus formation and tumorigenicity in NIH 3T3 cells. Proc Natl Acad Sci U S A. 1990 Dec;87(24):10005–10009. doi: 10.1073/pnas.87.24.10005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin A. L., Yao A., Rubin H. Relation of spontaneous transformation in cell culture to adaptive growth and clonal heterogeneity. Proc Natl Acad Sci U S A. 1990 Jan;87(1):482–486. doi: 10.1073/pnas.87.1.482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin H. 'Spontaneous' transformation as aberrant epigenesis. Differentiation. 1993 Jun;53(2):123–137. doi: 10.1111/j.1432-0436.1993.tb00652.x. [DOI] [PubMed] [Google Scholar]
- Rubin H. Cellular epigenetics: effects of passage history on competence of cells for "spontaneous" transformation. Proc Natl Acad Sci U S A. 1993 Nov 15;90(22):10715–10719. doi: 10.1073/pnas.90.22.10715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin H., Xu K. Evidence for the progressive and adaptive nature of spontaneous transformation in the NIH 3T3 cell line. Proc Natl Acad Sci U S A. 1989 Mar;86(6):1860–1864. doi: 10.1073/pnas.86.6.1860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schimke R. T. Gene amplification in cultured animal cells. Cell. 1984 Jul;37(3):705–713. doi: 10.1016/0092-8674(84)90406-9. [DOI] [PubMed] [Google Scholar]
- TODARO G. J., GREEN H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J Cell Biol. 1963 May;17:299–313. doi: 10.1083/jcb.17.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terzi M., Hawkins T. S. Chromosomal variation and the establishment of somatic cell lines in vitro. Nature. 1975 Jan 31;253(5490):361–362. doi: 10.1038/253361a0. [DOI] [PubMed] [Google Scholar]
- Tlsty T. D., Margolin B. H., Lum K. Differences in the rates of gene amplification in nontumorigenic and tumorigenic cell lines as measured by Luria-Delbrück fluctuation analysis. Proc Natl Acad Sci U S A. 1989 Dec;86(23):9441–9445. doi: 10.1073/pnas.86.23.9441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varshavsky A. Phorbol ester dramatically increases incidence of methotrexate-resistant mouse cells: possible mechanisms and relevance to tumor promotion. Cell. 1981 Aug;25(2):561–572. doi: 10.1016/0092-8674(81)90074-x. [DOI] [PubMed] [Google Scholar]








