Abstract
Rhizobium bacteria synthesize N-acylated beta-1,4-N-acetylglucosamine lipooligosaccharides, called Nod factors, which act as morphogenic signal molecules to legume roots during development of nitrogen-fixing nodules. The biosynthesis of Nod factors is genetically dependent upon the nodulation (nod) genes, including the common nod genes nodABC. We used the Rhizobium meliloti NodH sulfotransferase to prepare 35S-labeled oligosaccharides which served as metabolic tracers for Nod enzyme activities. This approach provides a general method for following chitooligosaccharide modifications. We found nodAB-dependent conversion of N-acetylchitotetraose (chitotetraose) monosulfate into hydrophobic compounds which by chromatographic and chemical tests were equivalent to acylated Nod factors. Sequential incubation of labeled intermediates with Escherichia coli containing either NodA or NodB showed that NodB was required before NodA during Nod factor biosynthesis. The acylation activity was sensitive to oligosaccharide chain length, with chitotetraose serving as a better substrate than chitobiose or chitotriose. We constructed a putative Nod factor intermediate, GlcN-beta 1,4-(GlcNAc)3, by enzymatic synthesis and labeled it by NodH-mediated sulfation to create a specific metabolic probe. Acylation of this oligosaccharide required only NodA. These results confirm previous reports that NodB is an N-deacetylase and suggest that NodA is an N-acyltransferase.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Atkinson E. M., Long S. R. Homology of Rhizobium meliloti NodC to polysaccharide polymerizing enzymes. Mol Plant Microbe Interact. 1992 Sep-Oct;5(5):439–442. doi: 10.1094/mpmi-5-439. [DOI] [PubMed] [Google Scholar]
- Bibb M. J., Biró S., Motamedi H., Collins J. F., Hutchinson C. R. Analysis of the nucleotide sequence of the Streptomyces glaucescens tcmI genes provides key information about the enzymology of polyketide antibiotic biosynthesis. EMBO J. 1989 Sep;8(9):2727–2736. doi: 10.1002/j.1460-2075.1989.tb08414.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debellé F., Rosenberg C., Dénarié J. The Rhizobium, Bradyrhizobium, and Azorhizobium NodC proteins are homologous to yeast chitin synthases. Mol Plant Microbe Interact. 1992 Sep-Oct;5(5):443–446. doi: 10.1094/mpmi-5-443. [DOI] [PubMed] [Google Scholar]
- Demont N., Debellé F., Aurelle H., Dénarié J., Promé J. C. Role of the Rhizobium meliloti nodF and nodE genes in the biosynthesis of lipo-oligosaccharidic nodulation factors. J Biol Chem. 1993 Sep 25;268(27):20134–20142. [PubMed] [Google Scholar]
- Dénarié J., Cullimore J. Lipo-oligosaccharide nodulation factors: a minireview new class of signaling molecules mediating recognition and morphogenesis. Cell. 1993 Sep 24;74(6):951–954. doi: 10.1016/0092-8674(93)90717-5. [DOI] [PubMed] [Google Scholar]
- Dénarié J., Debellé F., Rosenberg C. Signaling and host range variation in nodulation. Annu Rev Microbiol. 1992;46:497–531. doi: 10.1146/annurev.mi.46.100192.002433. [DOI] [PubMed] [Google Scholar]
- Egelhoff T. T., Fisher R. F., Jacobs T. W., Mulligan J. T., Long S. R. Nucleotide sequence of Rhizobium meliloti 1021 nodulation genes: nodD is read divergently from nodABC. DNA. 1985 Jun;4(3):241–248. doi: 10.1089/dna.1985.4.241. [DOI] [PubMed] [Google Scholar]
- Egelhoff T. T., Long S. R. Rhizobium meliloti nodulation genes: identification of nodDABC gene products, purification of nodA protein, and expression of nodA in Rhizobium meliloti. J Bacteriol. 1985 Nov;164(2):591–599. doi: 10.1128/jb.164.2.591-599.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher R. F., Egelhoff T. T., Mulligan J. T., Long S. R. Specific binding of proteins from Rhizobium meliloti cell-free extracts containing NodD to DNA sequences upstream of inducible nodulation genes. Genes Dev. 1988 Mar;2(3):282–293. doi: 10.1101/gad.2.3.282. [DOI] [PubMed] [Google Scholar]
- Fisher R. F., Long S. R. Rhizobium--plant signal exchange. Nature. 1992 Jun 25;357(6380):655–660. doi: 10.1038/357655a0. [DOI] [PubMed] [Google Scholar]
- Geiger O., Spaink H. P., Kennedy E. P. Isolation of the Rhizobium leguminosarum NodF nodulation protein: NodF carries a 4'-phosphopantetheine prosthetic group. J Bacteriol. 1991 May;173(9):2872–2878. doi: 10.1128/jb.173.9.2872-2878.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobs T. W., Egelhoff T. T., Long S. R. Physical and genetic map of a Rhizobium meliloti nodulation gene region and nucleotide sequence of nodC. J Bacteriol. 1985 May;162(2):469–476. doi: 10.1128/jb.162.2.469-476.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- John M., Röhrig H., Schmidt J., Wieneke U., Schell J. Rhizobium NodB protein involved in nodulation signal synthesis is a chitooligosaccharide deacetylase. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):625–629. doi: 10.1073/pnas.90.2.625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- John M., Schmidt J., Wieneke U., Krüssmann H. D., Schell J. Transmembrane orientation and receptor-like structure of the Rhizobium meliloti common nodulation protein NodC. EMBO J. 1988 Mar;7(3):583–588. doi: 10.1002/j.1460-2075.1988.tb02850.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson D., Roth L. E., Stacey G. Immunogold localization of the NodC and NodA proteins of Rhizobium meliloti. J Bacteriol. 1989 Sep;171(9):4583–4588. doi: 10.1128/jb.171.9.4583-4588.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kafetzopoulos D., Thireos G., Vournakis J. N., Bouriotis V. The primary structure of a fungal chitin deacetylase reveals the function for two bacterial gene products. Proc Natl Acad Sci U S A. 1993 Sep 1;90(17):8005–8008. doi: 10.1073/pnas.90.17.8005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lerouge P., Roche P., Faucher C., Maillet F., Truchet G., Promé J. C., Dénarié J. Symbiotic host-specificity of Rhizobium meliloti is determined by a sulphated and acylated glucosamine oligosaccharide signal. Nature. 1990 Apr 19;344(6268):781–784. doi: 10.1038/344781a0. [DOI] [PubMed] [Google Scholar]
- Palcic M. M., Hindsgaul O. Flexibility in the donor substrate specificity of beta 1,4-galactosyltransferase: application in the synthesis of complex carbohydrates. Glycobiology. 1991 Mar;1(2):205–209. doi: 10.1093/glycob/1.2.205. [DOI] [PubMed] [Google Scholar]
- Price N. P., Relić B., Talmont F., Lewin A., Promé D., Pueppke S. G., Maillet F., Dénarié J., Promé J. C., Broughton W. J. Broad-host-range Rhizobium species strain NGR234 secretes a family of carbamoylated, and fucosylated, nodulation signals that are O-acetylated or sulphated. Mol Microbiol. 1992 Dec;6(23):3575–3584. doi: 10.1111/j.1365-2958.1992.tb01793.x. [DOI] [PubMed] [Google Scholar]
- Reuber T. L., Walker G. C. Biosynthesis of succinoglycan, a symbiotically important exopolysaccharide of Rhizobium meliloti. Cell. 1993 Jul 30;74(2):269–280. doi: 10.1016/0092-8674(93)90418-p. [DOI] [PubMed] [Google Scholar]
- Roche P., Debellé F., Maillet F., Lerouge P., Faucher C., Truchet G., Dénarié J., Promé J. C. Molecular basis of symbiotic host specificity in Rhizobium meliloti: nodH and nodPQ genes encode the sulfation of lipo-oligosaccharide signals. Cell. 1991 Dec 20;67(6):1131–1143. doi: 10.1016/0092-8674(91)90290-f. [DOI] [PubMed] [Google Scholar]
- Roche P., Lerouge P., Ponthus C., Promé J. C. Structural determination of bacterial nodulation factors involved in the Rhizobium meliloti-alfalfa symbiosis. J Biol Chem. 1991 Jun 15;266(17):10933–10940. [PubMed] [Google Scholar]
- Sanjuan J., Carlson R. W., Spaink H. P., Bhat U. R., Barbour W. M., Glushka J., Stacey G. A 2-O-methylfucose moiety is present in the lipo-oligosaccharide nodulation signal of Bradyrhizobium japonicum. Proc Natl Acad Sci U S A. 1992 Sep 15;89(18):8789–8793. doi: 10.1073/pnas.89.18.8789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultze M., Quiclet-Sire B., Kondorosi E., Virelizer H., Glushka J. N., Endre G., Géro S. D., Kondorosi A. Rhizobium meliloti produces a family of sulfated lipooligosaccharides exhibiting different degrees of plant host specificity. Proc Natl Acad Sci U S A. 1992 Jan 1;89(1):192–196. doi: 10.1073/pnas.89.1.192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwedock J. S., Long S. R. Rhizobium meliloti genes involved in sulfate activation: the two copies of nodPQ and a new locus, saa. Genetics. 1992 Dec;132(4):899–909. doi: 10.1093/genetics/132.4.899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwedock J., Long S. R. ATP sulphurylase activity of the nodP and nodQ gene products of Rhizobium meliloti. Nature. 1990 Dec 13;348(6302):644–647. doi: 10.1038/348644a0. [DOI] [PubMed] [Google Scholar]
- Shearman C. A., Rossen L., Johnston A. W., Downie J. A. The Rhizobium leguminosarum nodulation gene nodF encodes a polypeptide similar to acyl-carrier protein and is regulated by nodD plus a factor in pea root exudate. EMBO J. 1986 Apr;5(4):647–652. doi: 10.1002/j.1460-2075.1986.tb04262.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
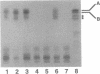
- Spaink H. P., Sheeley D. M., van Brussel A. A., Glushka J., York W. S., Tak T., Geiger O., Kennedy E. P., Reinhold V. N., Lugtenberg B. J. A novel highly unsaturated fatty acid moiety of lipo-oligosaccharide signals determines host specificity of Rhizobium. Nature. 1991 Nov 14;354(6349):125–130. doi: 10.1038/354125a0. [DOI] [PubMed] [Google Scholar]