Selection for both energy use efficiency and drought tolerance in canola leads to stable epilines with distinct transcriptome and chromatin modification profiles, supporting their superior performance.
Abstract
To increase both the yield potential and stability of crops, integrated breeding strategies are used that have mostly a direct genetic basis, but the utility of epigenetics to improve complex traits is unclear. A better understanding of the status of the epigenome and its contribution to agronomic performance would help in developing approaches to incorporate the epigenetic component of complex traits into breeding programs. Starting from isogenic canola (Brassica napus) lines, epilines were generated by selecting, repeatedly for three generations, for increased energy use efficiency and drought tolerance. These epilines had an enhanced energy use efficiency, drought tolerance, and nitrogen use efficiency. Transcriptome analysis of the epilines and a line selected for its energy use efficiency solely revealed common differentially expressed genes related to the onset of stress tolerance-regulating signaling events. Genes related to responses to salt, osmotic, abscisic acid, and drought treatments were specifically differentially expressed in the drought-tolerant epilines. The status of the epigenome, scored as differential trimethylation of lysine-4 of histone 3, further supported the phenotype by targeting drought-responsive genes and facilitating the transcription of the differentially expressed genes. From these results, we conclude that the canola epigenome can be shaped by selection to increase energy use efficiency and stress tolerance. Hence, these findings warrant the further development of strategies to incorporate epigenetics into breeding.
The need to improve crop yield in both quantity (yield potential) and stability (actual yield) to meet the increasing demand for food, feed, and plant-derived materials is a major challenge. Very different complementary technologies are used to optimize yield and to develop crops with increased resilience against adverse environmental growth conditions (Botella et al., 2008; Cattivelli et al., 2008; Jhaet al., 2014). Plant breeding programs are a basic component of this improvement process encompassing a wide range of technologies, such as exploration of the genetic potential by intraspecific and interspecific crosses, combination of genetic pools in hybrid breeding, mutational breeding, molecular breeding, and transgene technologies.
Rather recently, epigenetics has been investigated as a potential breeding platform (Springer, 2013). Epigenetic variations or heritable changes in gene expression that are not linked to changes in the DNA sequence, but associated with differences in DNA methylation or histone modifications, provide an alternative source of phenotypic variability. A vast and growing number of studies implicate DNA methylation and histone modification in the modulation of gene expression in general, control of developmental transitions, and plant responses to biotic and abiotic stresses (He et al., 2003; Raissig et al., 2011; Dowen et al., 2012; Perrella et al., 2013). However, our current understanding of the stability and heritability of epigenetic variation is limited. As such, how to implement epigenetics-related strategies into breeding remains an open question.
In plants, transgenerational inheritance has been shown for single-gene epialleles, such as SUPERMAN (Jacobsen and Meyerowitz, 1997) and CYCLOIDEA (Cubas et al., 1999), and for a transcription factor (TF) that controls fruit ripening in tomato (Solanum lycopersicum; Manning et al., 2006). For complex traits, such as flowering time, plant height, and primary root length, stable transgenerational epigenetic variations could be induced and linked to the identification of epigenetic quantitative trait loci (Johannes et al., 2009; Cortijo et al., 2014). Additional evidence supporting the impact of transgenerational epigenetic variation in breeding and the concept of shaping the epigenome to improve complex traits has been provided (Hauben et al., 2009). Starting from an isogenic canola (Brassica napus) population, individual plants were selected for reduced respiration, and subsequently, the self-fertilized progeny with the highest energy use efficiency were retained. This step was repeated twice with the progeny obtained from the previous selection cycle to fix the epigenome and to select epigenetically altered progeny with distinct physiological and agronomical properties that remained stable for at least eight generations.
Here, selection for energy use efficiency (Hauben et al., 2009) and drought tolerance were combined to isolate epilines. To unravel the biology of the enhanced energy use efficiency and drought stress tolerance, alterations in the transcriptome and epigenome in two of these epilines as well as one of the energy use-efficient epilines, the previously described line LR77 (Hauben et al., 2009), were analyzed in depth at the physiological, biochemical, and molecular levels. The LR77 line will be referred to hereafter as the EUE line.
RESULTS
Selection of Canola Lines with Improved Drought Stress Tolerance
Epilines were generated from the doubled haploid Athena spring canola cultivar by a repeated selection procedure similar to that described previously (Hauben et al., 2009; De Block and Van Lijsebettens, 2011). By screening explants of isogenic lines, variations in complex traits in the isolated progeny are expected to result from epigenetic differences (epialleles). Hypocotyl explants from 2-week-old seedlings were evaluated for respiration, and the shoot tips were kept for rooting. To induce a moderate drought stress, the hypocotyl explants were cultured on medium containing 5% (w/v) polyethylene glycol (PEG) 6000. Under these conditions, the respiration of the hypocotyl explants was higher than that of explants grown without PEG. Rooted shoot tips from 10 seedlings with the lowest hypocotyl explant respiration were transferred to the greenhouse for seed production by self-fertilization. Noteworthy, in this experimental setup, the plants were never exposed to PEG. Two of the self-fertilized progeny with the lowest respiration and the highest NAD(P)H content were retained. The selection procedure was repeated in three subsequent generations (Supplemental Fig. S1A). Respiration and NAD(P)H content were determined in the progeny of the last selection, and two epilines with low respiration and high NAD(P)H content, referred as the PEG1 and PEG2 lines, were kept for further analyses (Table I).
Table I. Respiration (TTC-H), energy content (NADH + NADPH), and energy use efficiency (EE) of hypocotyl explants.
Three repetitions per assay and 35 plants per repetition were used. Values shown are percentages versus control. EUE line data were as published by Hauben et al. (2009). Significance was determined by ANOVA with Dunnett’s posttest: *, P < 0.05; **, P < 0.01; ***, and P < 0.001; TTC-H, reduced 2,3,5-triphenyltetrazolium chloride; ND, not determined.
| Control Conditions |
+PEG |
|||||
|---|---|---|---|---|---|---|
| Lines | TTC-H | NAD(P)H | EE | TTC-H | NAD(P)H | EE |
| PEG1 | 91*** | 103* | 113*** | 95*** | 114*** | 120*** |
| PEG2 | 90*** | 99 | 110*** | 94*** | 108** | 115*** |
| EUE | 92.5*** | 103* | 111** | ND | ND | ND |
As described previously, lines with low respiration and high NAD(P)H content had reduced photorespiration (Hauben et al., 2009) under both control and PEG-induced stress conditions (Supplemental Table S1).
Under greenhouse conditions, the PEG lines could not be distinguished from the control plants regarding size, flowering time, and seed set. To assess their performance under drought stress, we subjected control and PEG plants to a drought stress regime starting on day 14 after sowing by withholding water supply until wilting symptoms (8 d) became apparent, followed by rewatering for 2 d. The PEG lines had increased rosette diameters (Fig. 1A). Under well-watered conditions, the shoot fresh weights were similar in PEG and control plants (Fig. 1B). The drought regime slowed down the growth of all lines, yet to a significantly smaller extent (25% versus 55% growth reduction) in the PEG lines, demonstrating an improved drought stress tolerance (Fig. 1B). The drought tolerance and energy use efficiency of the PEG lines were inherited stably for up to seven generations when seed upscaling was done by self-fertilization under nonselective conditions.
Figure 1.
Fresh weight of plants selected for high energy use efficiency (EUE lines) and drought tolerance (PEG lines). A, Control and PEG1 plants exposed to drought. The photograph was taken at the end of the experiment (24 d after sowing). For the experimental conditions, see “Materials and Methods.” B and C, Box-plot comparisons of the fresh weight of control, PEG1, and PEG2 plants (B) and of control and EUE plants (C) grown under control (blue) and drought (red) conditions 24 d after sowing. The whiskers are at the 10th and 90th percentiles, and the horizontal bar in each box shows the mean.
Nitrogen use efficiency (NUE) is an important agricultural trait that is possibly linked with drought tolerance (Sadras and Richards, 2014). The NUE of PEG and control lines was determined in a root length measurement assay on medium with low (35 µL L−1) or standard (210 µL L−1) nitrogen content. Plants with a high NUE form more lateral roots and have a higher total root length than plants with a low NUE. The seedlings of the PEG lines had a 40% higher total root length than control seedlings when grown on medium with low nitrogen content (Supplemental Fig. S1B).
In the EUE line, the seed yield was enhanced under optimal and moderate drought stress conditions, the photorespiration was reduced, the respiration had decreased, and the energy levels had increased. This line had been identified by selection for increased energy use efficiency and had been shown to have a decreased global DNA methylation level and altered histone modification patterns (Hauben et al., 2009). In this line, the growth was reduced by only 40% compared with 55% in its control upon an identical drought treatment as used for the PEG lines, which was significant but less pronounced than that of the drought tolerance measured in the PEG lines (Fig. 1C). Whereas the PEG lines had a pronounced NUE, the EUE line did not form significantly more roots than the control on medium with low nitrogen content and, thus, did not use nitrogen more efficiently.
Gene Expression Changes in PEG and EUE Lines
To assess the molecular footprint caused by the selection procedures for increased energy use efficiency in the presence or absence of PEG and to identify candidate genes responsible for the enhanced drought tolerance phenotypes of the selected lines, we performed genome-wide transcriptome analyses on control and selected plants. Both PEG lines (two biological replicates) and EUE lines (three biological replicates) with their respective controls were analyzed. Actively transcribed polysomal RNA pools from plants in different biological replicate experiments were isolated and subjected to RNA sequencing (RNA-seq) analysis.
As a complete and annotated genome sequence for canola (AACC genome) is not available yet, the genomes of the diploid species Brassica rapa (turnip; AA) and Brassica oleracea (cabbage; CC), of which the hybridization formed the amphidiploid canola species, were used as reference sequences. To optimize the mapping efficiency, RNA-seq reads were mapped to a set of 60,756 nonredundant B. rapa and B. oleracea transcripts (see “Materials and Methods”). Details of the mapping efficiency and coding sequence coverage are provided in Supplemental Table S2. Genes with low read coverage (see “Materials and Methods”) were excluded, finally retaining 35,892 genes (59% of all B. rapa and B. oleracea transcripts) to be examined. Differential transcript levels were identified using the edgeR package (false discovery rate < 0.05, fold change > 1.5; Robinson et al., 2010).
PEG1 plants displayed an altered expression in 1,158 genes, of which 706 induced and 452 repressed when compared with the control plants (Table II; Supplemental Table S3). In the PEG2 line, 478 genes were differentially expressed compared with the control line, of which 352 were up-regulated and 126 were down-regulated (Table II; Supplemental Table S3). The differential expression of arbitrarily selected genes was validated by quantitative reverse transcription-PCR (Fig. 2; Supplemental Table S4). The PEG1 and PEG2 lines displayed a significant overlap (355 genes; P < 1e-150) and a high expression correlation (r = 0.97) among the shared differentially expressed genes (DEGs; Fig. 2A), indicating that the selection altered the expression of a specific set of genes. In the EUE plants, screened solely for high energy use efficiency, a total of 1,099 DEGs were identified when compared with the corresponding control, among which 1,048 were up-regulated and 51 were down-regulated (Table II; Supplemental Table S2). Consistent with the partially similar selection strategy, a large number of DEGs (289 genes) were common in the EUE and PEG lines (Supplemental Fig. S2).
Table II. RNA-seq results.
|
Brassica spp. Linesa |
Arabidopsisb |
|||||
|---|---|---|---|---|---|---|
| Line | All | Up-Regulated | Down-Regulated | All | Up-Regulated | Down-Regulated |
| PEG1 | 1,158 | 706 | 452 | 844 | 503 | 341 |
| PEG2 | 478 | 352 | 126 | 360 | 257 | 103 |
| EUE | 1,099 | 1,048 | 51 | 768 | 719 | 49 |
Nonredundant B. rapa and B. oleracea genes. bUnique orthologous genes.
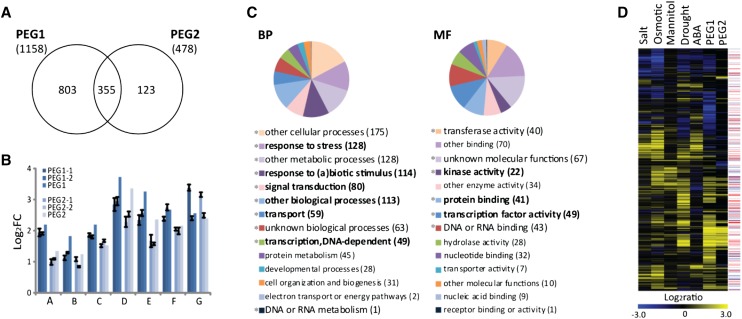
Figure 2.
Transcriptome changes related to energy use efficiency and drought tolerance. A, Venn diagram of DEGs in the PEG1 and PEG2 lines. B, Quantitative reverse transcription-PCR validation of the DEGs. Values are means ± sd (n = 3). For PEG1 and PEG2, log2 fold change (FC) values were calculated from RNA-seq data. C, Functional categorization of shared DEGs by annotation for the GO biological process (BP) and molecular function (MF) categories. The enriched (greater than 1.5-fold) functional categories are in boldface. *, P < 0.05. D, Hierarchical clustering analysis of the PEG Arabidopsis orthologous genes and a subset of drought stress-related transcriptome data sets (24 h of salt/osmotic/mannitol/drought and 3 h of ABA; Supplemental Table S7). TF (red) or kinase (blue) functional GO annotations for individual genes are indicated on the right.
To gain insight into the functional categories of the DEGs, we used their Arabidopsis (Arabidopsis thaliana) orthologs as a proxy (Table II; Supplemental Table S3). For 83% of all B. rapa and B. oleracea genes, an Arabidopsis orthologous gene was found. Comparison of the classification of the Arabidopsis orthologs of PEG1, PEG2, and EUE of the DEGs in the Gene Ontology (GO) terms related to the biological processes and molecular functions revealed the overrepresentation of the same terms, as expected given the high number of shared genes (Fig. 2C; Supplemental Table S5). Signal transduction, response to stress, transport, TF activity, protein binding, and kinase activity terms were significantly (more than 1.5-fold; P < 0.05) enriched, suggesting the onset of signaling events related to stress tolerance adaptation (Fig. 2C; Supplemental Table S5).
Accordingly, the DEGs of the PEG and EUE lines (Arabidopsis proxies) used as input for the signature tool in Genevestigator (see “Materials and Methods“; Hruz et al., 2008) demonstrated the relatedness with stress-linked perturbation profiles (Supplemental Table S6), including several drought stress experiments. The highest relative drought similarity scores were observed with the PEG1 line, followed by PEG2 and EUE (Supplemental Table S6). These transcriptomic signatures were largely in agreement with the higher drought stress tolerance of the PEG lines than that of the EUE lines (Fig. 1, B and C) and confirm the efficient selection procedure in the presence of PEG. A subsequent hierarchical clustering of the RNA-seq data with the Arabidopsis water stress-related transcriptome data sets (Supplemental Table S7) allowed a more detailed comparative analysis (Fig. 2D; Supplemental Fig. S3). There is a clear correlation with salt, osmotic, abscisic acid (ABA), mannitol, and drought treatments provoking expression changes (Fig. 2D; Supplemental Fig. S3). Indeed, of the responsive PEG1, PEG2, and EUE genes, 67.3%, 64.3%, and 62.7% were differentially expressed, respectively, in at least two of the compared subselected data sets (24-h salt, 24-h osmotic, 24-h mannitol, 3-h ABA, and 24-h drought treatments), confirming that PEG and energy use efficiency selection target, at least in part, drought tolerance-linked pathways.
Altered Histone 3 Lysine-4 Trimethylation Distribution in the Drought-Tolerant Canola Line
To investigate the epigenetic effect on the histone mark distribution of the energy use efficiency/PEG selection, we performed a chromatin immunoprecipitation sequencing (ChIP-seq) analysis. Dynamic changes in the occurrence of the histone 3 lysine-4 trimethylation (H3K4me3) mark are associated with the regulation of drought stress-responsive gene expression (Kim et al., 2008; Ding et al., 2012; Sani et al., 2013). Native ChIP with an anti-H3K4me3 antibody and without antibody (background control) was done on PEG1 and control plants. Details of the sequencing and mapping to the B. rapa and B. oleracea genomes separately are provided in Supplemental Table S8. The H3K4me3 distribution and differential binding in control and PEG1 lines were analyzed with the model-based analysis of chromatin immunoprecipitation sequencing (MACS; Zhang et al., 2008).
Using the MACS callpeak module, H3K4me3-enriched B. rapa and B. oleracea regions were identified in both the control and PEG1 lines and annotated to the nearest neighboring gene (Table III; Supplemental Tables S9 and S10). As expected for a histone protein modification, a uniform distribution of H3K4me3 was observed across all chromosomes of B. rapa (Fig. 3) and B. oleracea (Supplemental Fig. S4). More detailed analysis of the genomic H3K4me3 distribution in both B. oleracea and B. rapa in relation to the linked genes revealed that 26.2% of the enriched regions were located at the 5′ side (1 kb or intergenic), 66.2% in gene bodies, and 7.6% in the 3′ overlapping region (Supplemental Tables S9 and S10). A total of 73.4% of the H3K4me3 marks were located in the 5′ 1-kb (19.3%) and 5′ coding (54.1%) regions (Fig. 3B), consistent with the H3K4me3 distribution reported in Arabidopsis (Zhang et al., 2009; Roudier et al., 2011).
Table III. Number of total and differentially H3K4me3-bound regions (and genes) identified in PEG1 and control plants.
| Total Regions (Genes) |
Differential Regions (Genes) |
|||
|---|---|---|---|---|
| Species | Control | PEG1 | Control Enriched | PEG1 Enriched |
| B. rapa | 27,673 (21,844) | 31,464 (23,925) | 8 (8) | 3,131 (2,872) |
| B. oleracea | 36,371 (26,617) | 40,707 (28,492) | 38 (36) | 1,806 (1,748) |
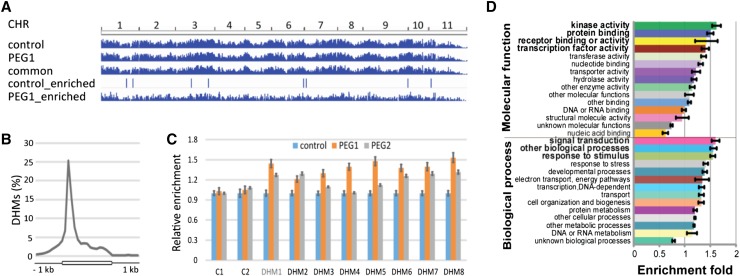
Figure 3.
Genome-wide H3K4me3 distribution in the PEG lines. A, H3K4me3 distribution across the 11 B. rapa chromosomes (CHR). B, Gene body and flanking 1-kb region distribution of the H3K4me3 marks in B. oleracea and B. rapa. All genes were aligned from start to end and divided into 20 equal bins. Upstream and downstream 1-kb regions were divided into 10 equal bins. C, ChIP-quantitative PCR validation of enriched H3K4me3 binding (DHM genes) in PEG1 and PEG2 lines. The relative enrichment compared with the control was determined using two ChIP-seq-identified nonenriched regions (C1 and C2). Error bars indicate sd (n = 3). D, Functional categorization of PEG1-enriched H3K4me3 genes by annotation for the GO biological process and molecular function categories. The enriched (greater than 1.5-fold) functional categories are in boldface (P < 0.05).
For differential H3K4me3 distribution analysis, the MACS bdgdiff module was used (see “Materials and Methods”). Comparison of the MACS callpeak-generated H3K4me3 pileup tracks of the control and PEG1 lines revealed that the majority of the H3K4me3-bound regions were common and not significantly different between the two lines (Fig. 3A; Supplemental Fig. S4). A few regions displayed an enhanced H3K4me3 signal in the control line, but many appeared enriched in H3K4me3 signal in the PEG1 line (Table III; Fig. 3A; Supplemental Fig. S4; Supplemental Tables S11 and S12). The above analyses were done on one biological replicate, because the H3K4me3 association of different regions was validated by quantitative PCR after ChIP on independent biological replicates of plant material of the control, PEG1, and PEG2 lines. For all regions, enrichment was higher in the PEG1 line than in the control, confirming the differential H3K4me3 binding and illustrating that PEG selection alters the H3K4me3 distribution (Fig. 3C). Moreover, enriched binding was seen in the PEG2 line in the same regions as those found in the PEG1 line (Fig. 3C), suggesting that the episelection directs the H3K4me3 alteration of a specific set of genes. Remarkably, six out of the eight genes with an enriched binding in the PEG1 line were also enriched in the PEG2 line. That not all the tested genes give an enriched binding in the PEG2 line can be explained by the weaker drought tolerance of the PEG2 line than that of the PEG1 line (Fig. 1B), as also reflected in the RNA-seq analyses (see above).
Annotation of the PEG1-enriched H3K4me3 regions to the nearest gene, followed by functional enrichment analysis of the Arabidopsis orthologous genes, uncovered overrepresented GO terms similar to the DEGs (Fig. 3D; Supplemental Tables S5, S11, and S12), further demonstrating the selectivity of the energy use efficiency and PEG selection toward stress tolerance-related signaling processes. In agreement, exploration of the Arabidopsis drought stress-related transcriptomes (Supplemental Table S7) revealed substantial changes in the PEG1 H3K4me3-enriched or differentially H3K4me3-marked (DHM) orthologs (Supplemental Fig. S5). Approximately 45% of the DHMs were differentially expressed in the osmotic treatment (24-h) data set, representing a significantly higher percentage of all osmotic treatment-responsive genes than that of the PEG1 DEGs (20% versus 6%) and indicating that the DHM distribution targets additional drought stress-responsive genes (Supplemental Fig. S5). Together, our data demonstrate that the PEG lines display DHM distribution and that this epigenetic factor may constitute a causative factor of the increased PEG drought stress performance.
Interdependence of Differential Gene Expression and H3K4me3 Distribution
As the presence of specific histone marks might directly affect gene expression, the DHM data were searched for genes that were differentially expressed in the PEG1 line. In total, both data sets shared 219 genes (Fig. 4; Supplemental Table S13), revealing a small but significant overlap (P = 4.9e-10). Comparative analysis of this overlapping subset of genes with the Arabidopsis abiotic stress transcriptome data sets (Supplemental Table S7) strengthened, as confirmed by the GO classification, the link with the drought stress response and the overrepresentation of TFs and kinases (Fig. 4B; Supplemental Table S5). Among the differentially expressed TFs, the B. oleracea (ERC) ERC42165, B. rapa (ERF) ERF62736, and ERF5709 loci, encoding the Brassica WRKY33, ETHYLENE-RESPONSIVE FACTOR104 (ERF104), and NO APICAL MERISTEM, ARABIDOPSIS TRANSCRIPTION ACTIVATION FACTOR, AND CUP-SHAPED COTYLEDON090 orthologs, respectively, were found to be enriched with H3K4me3. Read coverage along the 5′ coding regions in the PEG1 sample was higher than that of the control (Fig. 4D). Likewise, enriched H3K4me3 occupancy was detected in the 5′ regions of ERC26284 and ERF77038 loci, the Brassica spp. MITOGEN-ACTIVATED PROTEIN KINASE KINASE9 and CREATINE PHOSPHOKINASE32 orthologs (Fig. 4D). In agreement with the described connection of H3K4me3 with gene activation (Roudier et al., 2011), most (76.7%) of the differentially marked genes were up-regulated (Fig. 4, B and C; Supplemental Table S13). Surprisingly, part of the H3K4me3-enriched genes appeared to be down-regulated in both the PEG1 line and the Arabidopsis drought stress-related expression profiling data (Fig. 4B). Interestingly, repressed genes, in contrast to induced ones, displayed an atypical H3K4me3 enrichment at the 3′ ends of transcripts (Supplemental Fig. S6). This unanticipated distribution and its effect on the expression of the generally considered activating H3K4me3 mark needs further investigation. Previously, a role for H3K4me3 in gene repression was reported (Weiner et al., 2012). Altogether, these data suggest that the differential H3K4me3 distribution sensitizes the PEG lines to enhanced drought stress tolerance, partly by directly affecting the expression of stress tolerance genes.
Figure 4.
Relationship between the differential H3K4me3 distribution and the transcript levels. A, Venn diagram of the annotated DHM genes and DEGs in the PEG1 line. B, Clustering of orthologous DEGs and DHM genes of Arabidopsis with drought stress-related transcriptome data sets (24 h of salt/osmotic/mannitol/drought and 3 h of ABA; Supplemental Table S7). TF (red) and kinase (blue) functions of GO annotations for individual genes are indicated on the right. C, Box-plot displaying the log2 fold change (FC) expression of the H3K4me3-enriched genes of the PEG1 line. D, GenomeView representation (Abeel et al., 2012) of selected DEGs and DHM genes showing the read coverage in PEG1 versus the control line. The reads are piled up with forward reads above and reverse reads below the axis. Total coverage is gray shaded. Scaling was done relative to the maximum number of reads. The read number is given in the middle of the enriched regions. The coding regions are indicated as gray boxes and the PEG1-enriched regions as pink boxes above the reads. Bars = 0.5 kb.
Identifying Drought-Related Signaling Networks
The enrichment of both kinases and TFs in the RNA-seq (DEG) and ChIP-seq (DHM) data sets and in their overlap (DHM+DEG; Figs. 2, C and D, 3D, and 4B; Supplemental Fig. S5; Supplemental Table S5) hints at a regulatory role in conferring the improved drought stress tolerance phenotypes. In the DEG and DHM+DEG data sets, there is a significant enrichment in kinases that are involved in osmotic stress signaling events, such as class I receptor-like, class IV calcium response (group 4.2), and mitogen-activated protein (groups 4.1 and 4.5) kinases (http://plantsp.genomics.purdue.edu; Supplemental Table S14; Marshall et al., 2012; Kissoudis et al., 2014). The TF classification shows an overrepresentation of primarily the APETALA2-ETHYLENE-RESPONSIVE ELEMENT-BINDING PROTEIN (AP2-EREBP) and WRKY families (Supplemental Table S15; Lindemose et al., 2013).
Screening of the 1,000-bp upstream promoter regions of all DEGs and DHM genes for the W-box, DROUGHT-RESPONSIVE (DRE) elements, and ABSCISIC ACID-RESPONSIVE (ABRE) motifs identified putative WRKY, AP2-EREBP DEHYDRATION-RESPONSIVE ELEMENT BINDING (DREB) family, and basic Leu zipper TFs downstream of drought-responsive genes (see “Materials and Methods”). The number and identity of the DEGs and DHM genes retrieved by both simple and conserved noncoding sequence (CNS)-based mapping (Van de Velde et al., 2014) of these cis-regulatory elements are provided in Supplemental Table S16. Regarding the DEGs, 55.1%, including 23.7% high-confidence CNS-displaying DEGs, appeared to be putative target genes, whereas for the DHM+DEG genes, higher mapping fractions were observed, namely 61.6% simple and 18.9% CNS-based targets. Moreover, DEGs and DHM genes containing W-box, DRE sites, and ABRE motifs retained TF, kinase, and drought stress-associated GO term enrichments. In addition, a significant overrepresentation for all three motifs was evident in the CNS-based DEG and DHM genes (Fig. 5; Supplemental Table S16), confirming roles for WRKY, AP2-EREBP DREB, and basic Leu zipper TFs as master regulators in the energy use efficiency/PEG selection-improved drought stress tolerance signaling.
Figure 5.
Network analysis of DEGs enriched in H3K4me3. A, Cis-regulatory element enrichment analysis using 1-kb promoter sequences. Enrichment of W-box, DRE site, and ABRE motif occurrence in conserved noncoding sequences upstream of DEGs and/or DHM genes. Sequence logos used in mapping are shown below the graph. *, Not significant (P > 0.05). B, Gene regulatory and interaction subnetwork. The module represents pairwise first-neighbor interactions of the WRKY53 TFs with other PEG1 DEGs and/or DHM genes (see “Materials and Methods”; Supplemental Table S17). PPI, Protein-protein interaction.
To gain a hierarchical insight into the signaling and regulatory events, we used the DEG and DHM genes as query genes to retrieve pairwise regulatory (motif) interactions, protein interactions, and kinase-substrate interactions from the literature and several databases (see “Materials and Methods”; De Bodt et al., 2012; Zulawski et al., 2013; Jones et al., 2014; Lumba et al., 2014; Van de Velde et al., 2014). From the resulting 3,987 pairwise interactions, an integrative network was constructed, linking 1,841 genes/proteins (44.6% of all DEGs and DHMs). The generated network included 289 TFs that regulate 1,143 downstream DEGs and DHM genes and 190 kinases that interact with 125 proteins and phosphorylate 203 targets (Supplemental Table S17). Clustering (see “Materials and Methods”) allowed the isolation and closer inspection of discrete subnetworks. The most extensive subnetwork, comprising 875 genes/proteins, illustrated the high connectivity between the DEGs and DHM genes and retained GO term enrichments for TF and kinase activity (Supplemental Fig. S7). Interestingly, we observed a significant overrepresentation of genes displaying responses to water deprivation, to osmotic and salt stresses, and to ABA stimulus (Supplemental Fig. S7). When first-neighbor interactions of individual genes were extracted, PEG1 differentially expressed and/or H3K4me3-enriched kinases and TFs also appear as main regulators in several drought stress tolerance-related subnetworks (Fig. 5B; Supplemental Fig. S8; Supplemental Table S17). As an example, the network of WRKY53, a mitogen-activated protein kinase substrate, displays, besides regulatory interactions with other WRKY TFs, links with the dehydration stress-responsive TFs ERF5 and MYB15 and the REGULATORY COMPONENTS OF ABSCISIC ACID RECEPTOR1 (Miao et al., 2004; Ding et al., 2009; Park et al., 2009; Dubois et al., 2013; Fig. 5B). Moreover, WRKY53 is known to be regulated by histone methylation (Ay et al., 2009).
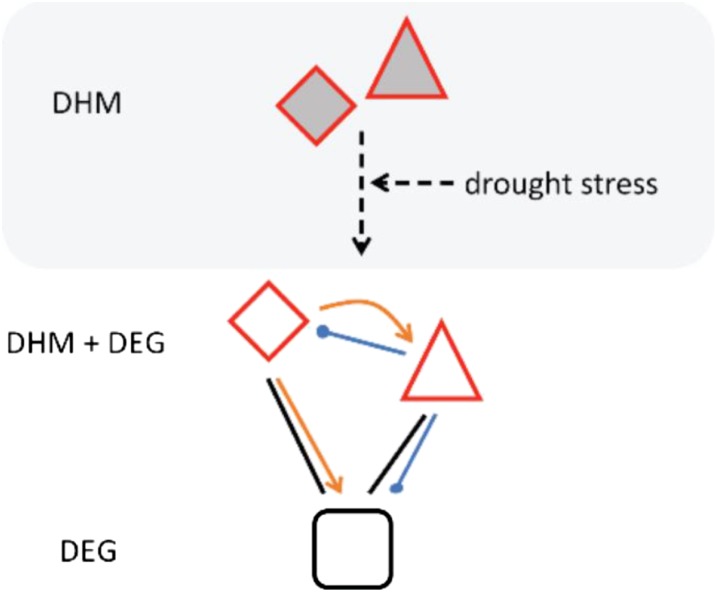
Together, these analyses support a model (Fig. 6) in which DEG and DHM key genes, primarily TFs and kinases, regulate downstream responses. Considering the role of the H3K4me3 dynamics in drought stress gene responsiveness (Kim et al., 2008; Ding et al., 2012), the strong regulatory interconnection observed among DEGs and DHM genes (Supplemental Fig. S7) suggests that the cascade conferring the improved drought stress tolerance in the PEG epilines becomes activated upon stress exposure (Fig. 6; Supplemental Fig. S8).
Figure 6.
Proposed network by CORNET (see “Materials and Methods”) conferring drought stress tolerance in the PEG epilines. Nodes depict DEGs (white) and/or DHM genes (red border), with triangles and diamonds representing TFs and kinases, respectively.
DISCUSSION
Epigenetic Selection for Energy Use Efficiency and Drought Tolerance
Molecular analysis of selected energy use-efficient (EUE lines) and drought-tolerant (PEG lines) plants highlights the directed epigenome and transcriptome changes and provides evidence for the episelection and an explanation for the enhanced drought stress tolerance of the epilines. RNA-seq analysis of control and PEG lines demonstrated the modified expression of a specific set of genes. In two independently selected PEG lines, a high overlap of shared DEGs was observed. Moreover, functional annotation and comparison with Arabidopsis transcriptome studies revealed a significant representation of genes linked to osmotic stress and ABA signaling. In favor of the efficiency of the combined selection strategy for both energy use efficiency and drought tolerance, the differential gene expression profiles of the PEG lines correlated better with water deprivation stress data than the EUE line selected solely for high energy use efficiency. Interestingly, the altered gene expression rendered the PEG lines more drought stress tolerant without negatively affecting growth or yield under optimal conditions (as observed in field trials), making them promising candidates for engineering enhanced stress tolerance.
The repeated selection for energy use efficiency solely or energy use efficiency and drought tolerance over several generations starting from isogenic populations of canola is an effective and fast process: from a population of approximately 200 plants, stable epilines with the selected phenotype were obtained after three generations. The altered gene expression in both the EUE and PEG lines obtained by screening explants of isogenic plants that are maintained and taken to the next generation when displaying the desired trait suggests an epigenetic feature. As already shown (Hauben et al., 2009), this screening resulted in the isolation of genetically identical, but epigenetically different, canola populations that were transgenerationally stable for over eight generations with respect to their phenotypes. Here, we demonstrate that this selection strategy enables the modulation of stress genes in the absence of a stimulus, indicating that the transgenerational trait inheritance occurs independently of any priming and memory (Molinier et al., 2006; Whittle et al., 2009). Similar to the EUE line, the drought tolerance of the PEG lines was inherited stably over at least seven generations. In isogenic populations, the epigenome is diverse and undergoes dynamic changes. During the screening process, some of these epigenetic states, epialleles, could be selected and fixed. Fixation of epialleles will reduce the epigenetic variation of the selected phenotype/trait in the population, as can be seen by the narrowing of the variation of the selected phenotype/trait in the population after each selection cycle (Hauben et al., 2009). It is very unlikely that mutations are responsible for the selected phenotypes, because the line to which the selections were applied was a double haploid that was derived from a single microspore and was maintained under controlled conditions to avoid unwanted cross-pollinations. After each seed upscaling, the progeny were tested for purity using specific phenotypic and molecular markers. Moreover, the high frequencies with which the epilines were selected (10−2) make selection by mutation very improbable. To assess potential genomic differences between a selected epiline and the original control line from which the epiline was derived, a variant-calling procedure was done on the available RNA-seq data of the PEG1 line and its control. Very few DNA polymorphisms were identified that probably do not cause mutations underlying the selected phenotypes (Supplemental Text S1).
Transcript Levels and H3K4me3 in the Epilines
The effectiveness of the episelection strategy was supported by the ChIP-seq analysis of control and PEG1 plants. Functional annotation of the H3K4me3-enriched genes in PEG1 plants and comparison with Arabidopsis transcriptome studies confirmed a significant representation of osmotic stress-linked genes. Moreover, in the PEG2 line, several regions with enriched H3K4me3 binding corresponded with those found in the PEG1 line, further favoring the specificity of the PEG selection to fix certain epialleles. In plants, changes in histone methylation mark distribution have been implicated in response to a range of abiotic stresses, including drought, osmotic, and salt stresses (Sokol et al., 2007; van Dijk et al., 2010; Kim et al., 2012; Sani et al., 2013; Zong et al., 2013). In all these studies, histone mark changes resulted from sensitization for stress responsiveness. In contrast, our selection strategy isolated epilines with a directed altered H3K4me3 distribution under optimal nonprimed conditions: only the hypocotyl explants of the seedlings were used for screening stress tolerance, whereas the shoot tips that were rooted and transferred to the greenhouse were never exposed to the stress. More research is needed to understand the mechanism of the epiallele selection, fixation, and transgenerational inheritance, for example by studying the epigenome of the populations generated in the three subsequent selections steps.
In plants, a positive correlation has been found between H3K4me3 and gene transcription (Roudier et al., 2011). Accordingly, most DEGs with enriched H3K4me3 mark distribution are up-regulated in the PEG lines. However, until now, it has been unclear whether the H3K4me3 pattern is the cause or the consequence of the increased gene expression (Sani et al., 2013). Besides the overall positive correlation between H3K4me3 and gene expression, a small set of down-regulated PEG genes also display H3K4me3 enrichment. The lack of correlation between H3K4me3 and enhanced gene expression was also described in other studies (Weiner et al., 2012; Sani et al., 2013). Moreover, the positive correlation between H3K4me3 and gene expression is much weaker for genes with a high expression than for those with a low expression (Sani et al., 2013). In budding yeast (Saccharomyces cerevisiae), a repressive role for H3K4me3 has been reported (Weiner et al., 2012). A limited, but substantial, set of diamide stress-repressed genes were found to have increased H3K4me3 levels. Interestingly, this unexpected chromatin mark enrichment appeared to target a specific gene class, the ribosomal protein genes (Weiner et al., 2012). Among the small number of H3K4me3-enriched transcriptionally repressed PEG genes (47 genes), no specific gene group seems overrepresented. Notably, whereas H3K4me3 occurs at the 5′ ends of transcribed and PEG-induced genes, the repressed PEG genes displayed an atypical H3K4me3 enrichment distribution at the 3′ ends. This unanticipated distribution and its possible effect on expression need further investigation. Additionally, studies might connect the H3K4me3 enrichment of PEG lines with osmotic stress-responsive up-regulated and, possibly, down-regulated genes.
Chromatin modification is not necessarily correlated directly with transcriptional regulation, but it can generate a context for the interplay with other factors at a developmental stage or in an environmental condition. Elevated H3K4me3 levels may not affect transcription immediately, but could act as preparatory/sensitizing marks for a fast and genome-wide gene transcription responsiveness upon stress perception (Jaskiewicz et al., 2011). Considering the high number of H3K4me3-enriched osmotic stress-responsive genes without altered expression in the PEG1 line, examination and comparison of their expression profiles upon stress exposure might be interesting. In summary, our study demonstrates that epilines with distinct characteristics can be selected from a pool of isogenic seeds, implying that the implementation of epigenetics in breeding is promising.
MATERIALS AND METHODS
Plant Material, Generation of Lines, and Physiological and Biochemical Analyses
Athena is an elite canola (Brassica napus) breeding line of Bayer CropScience. Canola lines with improved energy use efficiency and drought stress tolerance were selected as described previously with some minor modifications (Hauben et al., 2009). Seedlings from an isogenic doubled haploid line were grown for 2 weeks on agar in one-half-strength Murashige and Skoog medium supplemented with 2% (w/v) Suc. Seedling shoot tips were placed on the medium for rooting, whereas five hypocotyl explants per seedling were cultured for 5 d on callus-inducing medium containing 5% (w/v) PEG-6000 (Fluka BioUltra; Sigma-Aldrich) before cellular respiration was measured (Hauben et al., 2009). Rooted shoot tips of seedlings with the lowest respiration were transferred to the greenhouse for seed production by self-fertilization and subjected to two additional identical selection rounds. Respiration and photorespiration of the obtained lines were measured as described (Hauben et al., 2009).
Intracellular NADH and NADPH were quantified as described (Nakamura et al., 2003) with the following modifications. For the reaction solution, 5% (v/v) of the Cell Counting Kit-8 (Dojindo Laboratories) was diluted in 25 mm potassium phosphate buffer (pH 7.4) containing 0.1 mm 1-methoxyphenazine methosulfate (Sigma-Aldrich) and 0.1% (v/v) PEG sorbitan monolaurate (Tween 20). The explants were placed in the reaction solution and incubated at 26°C for 30 min to 1 h. The absorbance of the reaction solution was measured at 450 nm [optical density at 450 nm = 0.281 for 10 µm NAD(P)H].
NUE was scored by quantifying the root formation on medium with a low nitrogen content (35 µL L−1) versus medium with a standard nitrogen content (210 µL L−1). Hoagland medium (Hoagland and Arnon, 1938) with 1.2% (w/v) plant agar (Duchefa) was poured into square Nunc bioassay dishes (Thermo Scientific). After solidification of the agar, the top half of the medium was removed and seeds were sown on top of the agar layer. The plates were incubated vertically for 8 d (at 23°C, 80 µmol m−2 s−1 light, and a 16-h-light/8-h-dark regime). The roots that grew on the surface of the agar were quantified by scanning the plates and analyzed with ImageJ (http://imagej.nih.gov/ij/).
Drought tolerance was analyzed by measuring the fresh weight of shoots of 24-d-old normally watered and drought regime-treated plants. After 13 d, plants under the drought regime were no longer watered for 8 d, until wilting symptoms became apparent, followed by watering for 2 d, after which the fresh weight was determined.
Polysomal RNA Isolation
Polysomal RNA was isolated according to an adapted protocol (de Jong et al., 2006). Ground and liquid nitrogen frozen material of leaf 4 from 26-d-old control and selected lines was solubilized in an equal volume of extraction buffer (pH 8.5) containing 0.2 m Tris, 0.1 m KCl, 70 mm magnesium acetate, 50 mm EGTA, 0.25 m Suc, 10 mm dithiothreitol, 100 µg mL−1 cycloheximide (Sigma-Aldrich), and 100 units mL−1 SUPERase-In (Ambion). The slurry was allowed to thaw on ice and gently mixed to enhance homogenization. The suspension was strained through two layers of Miracloth (Merck Millipore) via successive centrifugation for 1 min at 100g (4°C) and 5 min at 1,000g (4°C) in 50-mL sterile tubes. The cleared supernatant was centrifuged for 15 min at 8,400g (4°C) to pellet additional debris, made to 1% (v/v) Triton X-100, and incubated on ice for 10 min. After centrifugation at 20,800g (4°C) in an Eppendorf tabletop centrifuge, the supernatant was loaded in SW41 Ultra-Clear tubes (Beckman Coulter) on discontinuous Suc gradients that consisted of 0.6 mL of 1.65 m Suc and 1 mL of 1 m Suc, both buffered with 10× concentrated Suc salt buffer (pH 8.5, autoclaved) containing 0.4 m Tris, 1 m KCl, 0.3 m magnesium acetate, and 50 mm EGTA. To maintain RNA integrity, 10 mm dithiothreitol, 100 µg mL−1 cycloheximide, and 100 units mL−1 SUPERase-In (Ambion) were added to these Suc solutions. The gradients were centrifuged for 4 h at 150,000g (4°C) in an ultracentrifuge (type L8-50 M/E; Beckman Coulter) to pellet the polysomes. The polysomal pellets were resuspended in 100 µL of extraction buffer, and polysomal RNA was isolated with an RNeasy kit (Qiagen). The quantity of polysomal RNA was measured with a Nanodrop spectrophotometer (Thermo Scientific), and the quality was examined with a Bioanalyzer 2100 (Agilent Technologies) and the RNA 6000 Nano Assay kit (Agilent Technologies).
RNA-seq Analysis
The RNA-seq analysis was done at the Beijing Genome Institute. Complementary DNA (cDNA) sequencing libraries were prepared from polysomal RNA by means of the mRNA-Seq Sample Preparation Kit (Illumina) and run on a genome analyzer (GAIIx; Illumina), yielding 90-bp paired-end reads that were quality controlled with FastQC (version 0.10.0; http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc/), which showed that the quality decreased in the last 10 bp of the reads. With the FASTX trimmer of the FASTX toolkit (version 0.0.13; http://hannonlab.cshl.edu/fastx_toolkit/), 10 bases were trimmed at the 3′ side, yielding high-quality 80-bp paired-end reads.
As a reference for the read mapping, a nonredundant gene set was built as follows. Orthologous pairs were formed by an all-against-all BLASTN (Altschul et al., 1990) comparison between the nucleotide sequences of the in silico predicted gene sequences of Brassica rapa and Brassica oleracea (in-house data) and by retrieval of the reciprocal best hits, resulting in 21,260 orthologous pairs and 39,496 singletons (nonorthologs). Because of the higher annotation quality of B. rapa, the B. rapa ortholog was taken as the orthologous pair representative in every orthologous pair. These 21,260 orthologous pair representatives together with the 39,496 singletons form the nonredundant reference gene set that contains 60,756 genes.
In the next step, the high-quality 80-bp paired-end reads were mapped to the nonredundant gene set with GSNAP [unique mapping with maximum number of paths to print set to 1 (-n 1 -Q) and maximum of five mismatches (-m 5) allowed]. These options enforce, when used together, that reads that cannot be mapped in a unique manner, are not written to the output file (Wu and Nacu, 2010). All reads were retained that mapped uniquely and concordantly, and the number of reads mapping to each transcript was calculated with samtools idxstats (Li et al., 2009). The corresponding count table was filtered on a minimum average read count of five over the lines analyzed and then was given as an input to edgeR (Robinson et al., 2010) for differential expression testing.
Quantitative Reverse Transcription-PCR
Total RNA was extracted with the RNeasy Plant Mini Kit (Qiagen) and treated with DNase I prior to cDNA synthesis. cDNA was prepared from DNase I-treated total RNA with the iScript cDNA Synthesis Kit (Bio-Rad) according to the manufacturer’s instructions. The relative transcript abundance of selected genes (for genes and primers used, see Supplemental Table S4) was determined with the LightCycler 480 system and the LC480 SYBR Green I Master kit (Roche Diagnostics). Two biological and three technical repeats were measured. To obtain relative expression values, the amplification data were analyzed by means of qBase+ (Hellemans et al., 2007) with the second derivative maximum method and three reference genes for normalization.
Native ChIP
The native ChIP protocol was adapted (Bernatavichute et al., 2008) as follows. Material from leaf 4 of 26-d-old wild-type and PEG1 plants was harvested, frozen in liquid nitrogen, and ground to powder (1.5 g). Plant tissue was resuspended in 15 mL of HBM buffer (25 mm Tris-HCl, pH 7.5, 440 mm Suc, 10 mm MgCl2, 0.1% [v/v] Triton-X, 10 mm β-mercaptoethanol, 2 mm spermine, 1 mm phenylmethanesulfonyl fluoride [PMSF], 1 µg mL−1 pepstatin, 1 µg mL−1 aprotinin, and 1 µg mL−1 leupeptin), homogenized, and filtered twice through Miracloth (Merck Millipore). After centrifugation at 1,000g for 5 min (4°C; SS-34 Sorvall; Thermo Scientific), the pellet was resuspended in 5 mL of NIB buffer (20 mm Tris-HCl, pH 7.5, 250 mm Suc, 5 mm MgCl2, 5 mm KCl, 0.1% [v/v] Triton-X, 10 mm β-mercaptoethanol, 1 mm PMSF, 1 µg mL−1 pepstatin, 1 µg mL−1 aprotinin, and 1 µg mL−1 leupeptin), applied to a 15%/50% Percoll (GE-Healthcare) gradient in NIB buffer, and centrifuged at 500g for 20 min (4°C; SS-34 Sorvall). Isolated nuclei were washed three times in NIB buffer and flash frozen in liquid nitrogen in HBC buffer (25 mm Tris-HCl, pH 7.5, 440 mm Suc, 10 mm MgCl2, 0.1% [v/v] Triton-X, 10 mm β-mercaptoethanol, and 20% [v/v] glycerol). Nuclei from each preparation were diluted 2-fold in digestion buffer (20 mm Tris-HCl, pH 7.5, 0.22 m Suc, 50 mm NaCl, 5 mm MgCl2, 1 mm CaCl2, and 5 mm sodium butyrate), centrifuged at 600g for 5 min (4°C; model 5415R; Eppendorf), and resuspended in 0.25 mL of digestion buffer for micrococcal nuclease (30 units; Affymetrix) digestion for 10 min at 37°C stopped with 10 mm EDTA. Mononucleosomes were released by treating nuclei with 0.1% (v/v) Triton-X for 2 h in the cold and then pelleting the debris by centrifugation at 9,300g for 5 min (4°C; 5415R; Eppendorf). The supernatants were diluted to 0.5 mL in incubation buffer (20 mm Tris-HCl, pH 7.5, 50 mm NaCl, 5 mm EDTA, 0.1% [v/v] Triton-X, 20 mm sodium butyrate, 0.1 mm PMSF, 1 µg mL−1 pepstatin, 1 µg mL−1 aprotinin, and 1 µg mL−1 leupeptin) and incubated overnight in the cold with and without 2.5 µg of the anti-H3K4m3 antibody (#07-473; Merck Millipore). Then, 50 µL of Dynabeads Protein G (Invitrogen) was added, incubated for 3 h in the cold, and washed three times with 500 µL of wash buffer (50 mm Tris-HCl, pH 7.5, 10 mm EDTA, 5 mm sodium butyrate, 0.1 mm PMSF, 1 µg mL−1 pepstatin, 1 µg mL−1 aprotinin, and 1 µg mL−1 leupeptin) with increasing concentrations of NaCl (50, 100, and 150 mm) subsequently. The final wash was done in TE buffer (10 mm Tris-Cl, pH 8, 500 mm NaCl, and 1 mm EDTA), and immunocomplexes were eluted and reverse cross-linked by incubation overnight at 65°C in 0.25 mL of 10 mm Tris-HCl (pH 8), 1 mm EDTA, 0.5 m NaCl, 1% (w/v) SDS, and 1 µL of RNase A (100 mg mL−1). Eluates were incubated with 100 µg of Proteinase K for 2 h at 42°C, and the DNA was extracted by phenol/chloroform/indole-3-acetic acid, followed by purification with the Qiaquick PCR purification kit and DNA quantification with the Quant-iT dsDNA High Sensitivity kit (Invitrogen).
ChIP-seq Analysis
ChIP DNA libraries were prepared according to the manufacturer’s protocol and sequenced on a HiSeq2500 (100 paired-end reads; Illumina). The sequencing data were quality controlled with FastQC. Overrepresented sequences were removed using FASTX-clipper from the FASTX toolkit. The reads were mapped to the reference genomes of B. rapa and B. oleracea using GSNAP with default settings (Wu and Nacu, 2010). Reads that could not be assigned to a unique position in the genome were removed.
Peak calling was done with MACS version 2.0 (Zhang et al., 2008). The genome size (-g) was set at 2.0e8 and 3.7e8 for B. rapa and B. oleracea, respectively. Other parameters were set at their default values using the BAMPE format option. For differential histone mark-binding analysis, the MACS version 2.0 bdgdiff module with default settings was used. Peaks were annotated by overlapping the peak summits reported by MACS with the B. rapa or B. oleracea annotation. A peak was assigned to the closest gene, taking into account both upstream and downstream regions of the genes. When a peak was located within the boundaries of a gene, it was assigned to this gene.
Quantitative ChIP-PCR
Quantitative PCR experiments were carried out on H3K4me3 ChIP-isolated genomic DNA from control and selected epilines and done as technical triplicates on equal DNA concentrations with the LightCycler 480 system and the LC480 SYBR Green I Master kit (Roche Diagnostics). Primers used for PCR amplification were designed surrounding the mid region of nonenriched (C1 and C2) and enriched regions identified by the sequencing (Supplemental Table S4). ChIP values of H3K4me3 binding were normalized by means of nonenriched regions.
Computational Analyses
The overlap significance of gene sets was calculated with a hypergeometric distribution test. Orthologous Arabidopsis genes (The Arabidopsis Information Resource 10) of the B. rapa and B. oleracea genes were identified by an all-against-all BLASTN (E-value cutoff of 1e-5; Altschul et al., 1990) and by retrieval of the reciprocal best hits. Functional GO classification and enrichment analysis of the Arabidopsis orthologs were done with BAR (Toufighi et al., 2005) and PLAZA 2.5 (Van Bel et al., 2012).
For comparative transcriptome analysis, the Genevestigator signature tool was utilized and hierarchical clustering analyses were performed. The 300 strongest DEGs (highest absolute log2 fold changes) were extracted separately for the PEG1, PEG2, and EUE lines and used as input for the signature tool in Genevestigator (Hruz et al., 2008), thereby selecting all perturbation experiments on the ATH1 microarray platform (relative log2 fold scaled data) to retrieve similar experiments. Raw Affymetrix Cell Intensity files were obtained from the Gene Expression Omnibus database (ncbi.nlm.nih.gov/geo/; GSE5628, GSE5621, GSE5622, GSE5623, GSE39384, GSE16474, GSE36789, and GSE22107) and ArrayExpress (ebi.ac.uk/arrayexpress/; E-MEXP-2435 and E-MEXP-2377). Data were normalized with Robust Multiarray Average (Bolstad et al., 2003; Irizarry et al., 2003a, 2003b) with the affy package of R/Bioconductor (Gautier et al., 2004). Probe sets were annotated according to the latest Brainarray (http://brainarray.mbni.med.umich.edu/Brainarray/; Dai et al., 2005) custom chip definition file provided for Arabidopsis (The Arabidopsis Information Resource G version 18.0.0). Differential gene expression was analyzed by the limma package (Smyth, 2004). Genes were hierarchically clustered (single linkage) by Pearson correlation in the open-source analysis software MultiExperiment Viewer version 4.9.0 (Saeed et al., 2003).
Genes were annotated as TFs when found at least twice in the databases DATF (Guo et al., 2005), PlnTFDB (Pérez-Rodríguez et al., 2010), AtTFDB (Yilmaz et al., 2011), and PlantTFDB (Jin et al., 2014). For the annotation of kinases, the PlantsP database (Tchieu et al., 2003) was used.
For promoter analysis, sequences were restricted to the first 1,000 bp upstream from the translation start site or to a shorter region when the adjacent upstream gene was located at a distance smaller than 1,000 bp. Known motifs from the integrated AGRIS, PLACE, and AthaMap (Van de Velde et al., 2014) databases were mapped with the dna-pattern program without allowing mismatches (Thomas-Chollier et al., 2008).
For network analysis, Arabidopsis orthologous DHM genes and DEGs were used as query genes to retrieve pairwise interaction information from different sources. Protein-protein interactions were extracted from CORNET2.0 (De Bodt et al., 2012; cornet.psb.ugent.be/) using experimental and predicted interactions from all available databases. Predicted interactions were filtered to a minimum of two independent references. Additional protein interaction sources were MIND (Jones et al., 2014; associomics.org/), eTRAIN (Lumba et al., 2014), and kinase-substrate interactions from PhosPhAt4.0 (Zulawski et al., 2013; phosphat.uni-hohenheim.de/). Regulatory interactions were downloaded from AGRIS (Yilmaz et al., 2011; Arabidopsis.med.ohio-state.edu/) and supplemented with predicted conserved TF motif interactions (Van de Velde et al., 2014). Interactions were filtered on a minimum of two independent references (confirmed; AGRIS) and motif conservation among four species. Additional regulatory interactions were retrieved from the EVEX database (Van Landeghem et al., 2012; evexdb.org/). All interactions were loaded into Cytoscape, and the FAG-EC algorithm (Li et al., 2008) was run with default settings for network clustering.
The data discussed in this article have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (Edgar et al., 2002) and are accessible through the Gene Expression Omnibus accession number GSE65578.
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. PEG line selection scheme and NUE analysis.
Supplemental Figure S2. Comparative transcriptome analysis of EUE and PEG plants.
Supplemental Figure S3. Hierarchical clustering analysis of RNA-seq and Arabidopsis drought stress-related transcriptome studies.
Supplemental Figure S4. H3K4me3 distribution across the B. oleracea reference.
Supplemental Figure S5. Heat map of differential expression levels of the Arabidopsis orthologous DHM genes in drought stress-related transcriptome data sets and the PEG1 line.
Supplemental Figure S6. Enriched H3K4me3 mark distribution in DEGs of PEG1.
Supplemental Figure S7. Cytoscape representation of the largest module obtained by clustering of the compiled interactome, kinase substrate, and regulatory data of the PEG1 DEGs and/or enriched DHM genes.
Supplemental Figure S8. Gene regulatory and interaction subnetwork of the CREATINE PHOSPHOKINASE32 kinase.
Supplemental Table S1. Photorespiration of selected epilines.
Supplemental Table S2. RNA-seq mapping details.
Supplemental Table S3. DEGs in the PEG and EUE lines.
Supplemental Table S4. List of primers.
Supplemental Table S5. Functional categorization of DEGs and DHM genes.
Supplemental Table S6. Genevestigator similarity analysis report of the DEGs of the PEG and EUE lines.
Supplemental Table S7. Arabidopsis transcriptome data sets (used for generating Supplemental Table S6).
Supplemental Table S8. ChIP-seq count and mapping results.
Supplemental Table S9. H3K4me3 distribution in the PEG control line.
Supplemental Table S10. H3K4me3 distribution in the PEG1 line.
Supplemental Table S11. H3K4me3 enrichment in the PEG control line.
Supplemental Table S12. H3K4me3 enrichment in the PEG1 line.
Supplemental Table S13. List of DEGs and DHM genes in the PEG1 line.
Supplemental Table S14. Kinase classification and enrichments.
Supplemental Table S15. Transcription factor classification and enrichments.
Supplemental Table S16. Simple and conserved cis-regulatory motif mapping.
Supplemental Table S17. List of pairwise protein and regulatory interactions between DEGs and/or DHM genes.
Supplemental Text S1. Variant calling between the PEG1 line and its control.
Supplementary Material
Acknowledgments
We thank Frederik Coppens, Ken Heyndrickx, Daisuke Miki, and Michiel Van Bel for assistance and useful suggestions and Martine De Cock for help in preparing the article.
Glossary
- PEG
polyethylene glycol
- NUE
nitrogen use efficiency
- RNA-seq
RNA sequencing
- DEG
differentially expressed gene
- GO
Gene Ontology
- TF
transcription factor
- ABA
abscisic acid
- ChIP-seq
chromatin immunoprecipitation sequencing
- ChIP
chromatin immunoprecipitation
- H3K4me3
histone 3 lysine-4 trimethylation
- MACS
model-based analysis of chromatin immunoprecipitation sequencing
- DHM
differentially histone 3 lysine-4 trimethylation marked
- CNS
conserved noncoding sequence
- cDNA
complementary DNA
- PMSF
phenylmethanesulfonyl fluoride
Footnotes
This work was supported by the Agency for Innovation by Science and Technology (grant no. IWT100268), the Ghent University Special Research Fund (grant no. 01J11311), the European Training and Mobility Network (grant no. FP7–PEOPLE–2013–ITN 607880), the Ghent University Multidisciplinary Research Partnership (grant nos. 01MR0410W and 01MRB510W), and the Interuniversity Attraction Poles Program (grant no. VII/29), initiated by the Belgian State, Science Policy Office.
Articles can be viewed without a subscription.
References
- Abeel T, Van Parys T, Saeys Y, Galagan J, Van de Peer Y (2012) GenomeView: a next-generation genome browser. Nucleic Acids Res 40: e12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215: 403–410 [DOI] [PubMed] [Google Scholar]
- Ay N, Irmler K, Fischer A, Uhlemann R, Reuter G, Humbeck K (2009) Epigenetic programming via histone methylation at WRKY53 controls leaf senescence in Arabidopsis thaliana. Plant J 58: 333–346 [DOI] [PubMed] [Google Scholar]
- Bernatavichute YV, Zhang X, Cokus S, Pellegrini M, Jacobsen SE (2008) Genome-wide association of histone H3 lysine nine methylation with CHG DNA methylation in Arabidopsis thaliana. PLoS ONE 3: e3156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolstad BM, Irizarry RA, Åstrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19: 185–193 [DOI] [PubMed] [Google Scholar]
- Botella MA, Rosada A, Bressan RA, Hasegawa PM (2008) Plant adaptive responses to salinity stress. In Jenks MA, Hasegawa PM, eds, Plant Abiotic Stress. Blackwell, Oxford, pp 37–70 [Google Scholar]
- Cattivelli L, Rizza F, Badeck FW, Mazzucotelli E, Mastrangelo AM, Francia E, Marè C, Tondelli A, Stanca AM (2008) Drought tolerance improvement in crop plants: an integrated view from breeding to genomics. Field Crops Res 105: 1–14 [Google Scholar]
- Cortijo S, Wardenaar R, Colomé-Tatché M, Gilly A, Etcheverry M, Labadie K, Caillieux E, Hospital F, Aury JM, Wincker P, et al. (2014) Mapping the epigenetic basis of complex traits. Science 343: 1145–1148 [DOI] [PubMed] [Google Scholar]
- Cubas P, Vincent C, Coen E (1999) An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401: 157–161 [DOI] [PubMed] [Google Scholar]
- Dai M, Wang P, Boyd AD, Kostov G, Athey B, Jones EG, Bunney WE, Myers RM, Speed TP, Akil H, et al. (2005) Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res 33: e175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Block M, Van Lijsebettens M (2011) Energy efficiency and energy homeostasis as genetic and epigenetic components of plant performance and crop productivity. Curr Opin Plant Biol 14: 275–282 [DOI] [PubMed] [Google Scholar]
- De Bodt S, Hollunder J, Nelissen H, Meulemeester N, Inzé D (2012) CORNET 2.0: integrating plant coexpression, protein–protein interactions, regulatory interactions, gene associations and functional annotations. New Phytol 195: 707–720 [DOI] [PubMed] [Google Scholar]
- de Jong M, van Breukelen B, Wittink FR, Menke FLH, Weisbeek PJ, Van den Ackerveken G (2006) Membrane-associated transcripts in Arabidopsis: their isolation and characterization by DNA microarray analysis and bioinformatics. Plant J 46: 708–721 [DOI] [PubMed] [Google Scholar]
- Ding Y, Fromm M, Avramova Z (2012) Multiple exposures to drought ‘train’ transcriptional responses in Arabidopsis. Nat Commun 3: 740. [DOI] [PubMed] [Google Scholar]
- Ding Z, Li S, An X, Liu X, Qin H, Wang D (2009) Transgenic expression of MYB15 confers enhanced sensitivity to abscisic acid and improved drought tolerance in Arabidopsis thaliana. J Genet Genomics 36: 17–29 [DOI] [PubMed] [Google Scholar]
- Dowen RH, Pelizzola M, Schmitz RJ, Lister R, Dowen JM, Nery JR, Dixon JE, Ecker JR (2012) Widespread dynamic DNA methylation in response to biotic stress. Proc Natl Acad Sci USA 109: E2183–E2191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubois M, Skirycz A, Claeys H, Maleux K, Dhondt S, De Bodt S, Vanden Bossche R, De Milde L, Yoshizumi T, Matsui M, et al. (2013) ETHYLENE RESPONSE FACTOR6 acts as a central regulator of leaf growth under water-limiting conditions in Arabidopsis. Plant Physiol 162: 319–332 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30: 207–210 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) affy: analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20: 307–315 [DOI] [PubMed] [Google Scholar]
- Guo A, He K, Liu D, Bai S, Gu X, Wei L, Luo J (2005) DATF: a database of Arabidopsis transcription factors. Bioinformatics 21: 2568–2569 [DOI] [PubMed] [Google Scholar]
- Hauben M, Haesendonckx B, Standaert E, Van Der Kelen K, Azmi A, Akpo H, Van Breusegem F, Guisez Y, Bots M, Lambert B, et al. (2009) Energy use efficiency is characterized by an epigenetic component that can be directed through artificial selection to increase yield. Proc Natl Acad Sci USA 106: 20109–20114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Y, Michaels SD, Amasino RM (2003) Regulation of flowering time by histone acetylation in Arabidopsis. Science 302: 1751–1754 [DOI] [PubMed] [Google Scholar]
- Hellemans J, Mortier G, De Paepe A, Speleman F, Vandesompele J (2007) qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 8: R19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoagland DR, Arnon DI (1938) The water-culture method for growing plants without soil. Calif Agric Exp Stn Circ 347: 1–39 [Google Scholar]
- Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinformatics 2008: 420747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP (2003a) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31: e15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003b) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4: 249–264 [DOI] [PubMed] [Google Scholar]
- Jacobsen SE, Meyerowitz EM (1997) Hypermethylated SUPERMAN epigenetic alleles in Arabidopsis. Science 277: 1100–1103 [DOI] [PubMed] [Google Scholar]
- Jaskiewicz M, Conrath U, Peterhänsel C (2011) Chromatin modification acts as a memory for systemic acquired resistance in the plant stress response. EMBO Rep 12: 50–55 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jha UC, Bohra A, Singh NP (2014) Heat stress in crop plants: its nature, impacts and integrated breeding strategies to improve heat tolerance. Plant Breed 133: 679–701 [Google Scholar]
- Jin J, Zhang H, Kong L, Gao G, Luo J (2014) PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res 42: D1182–D1187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johannes F, Porcher E, Teixeira FK, Saliba-Colombani V, Simon M, Agier N, Bulski A, Albuisson J, Heredia F, Audigier P, et al. (2009) Assessing the impact of transgenerational epigenetic variation on complex traits. PLoS Genet 5: e1000530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones AM, Xuan Y, Xu M, Wang RS, Ho CH, Lalonde S, You CH, Sardi MI, Parsa SA, Smith-Valle E, et al. (2014) Border control: a membrane-linked interactome of Arabidopsis. Science 344: 711–716 [DOI] [PubMed] [Google Scholar]
- Kim JM, To TK, Ishida J, Matsui A, Kimura H, Seki M (2012) Transition of chromatin status during the process of recovery from drought stress in Arabidopsis thaliana. Plant Cell Physiol 53: 847–856 [DOI] [PubMed] [Google Scholar]
- Kim JM, To TK, Ishida J, Morosawa T, Kawashima M, Matsui A, Toyoda T, Kimura H, Shinozaki K, Seki M (2008) Alterations of lysine modifications on the histone H3 N-tail under drought stress conditions in Arabidopsis thaliana. Plant Cell Physiol 49: 1580–1588 [DOI] [PubMed] [Google Scholar]
- Kissoudis C, van de Wiel C, Visser RGF, van der Linden G (2014) Enhancing crop resilience to combined abiotic and biotic stress through the dissection of physiological and molecular crosstalk. Front Plant Sci 5: 207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25: 2078–2079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M, Wang J, Chen J (2008) A fast agglomerate algorithm for mining functional modules in protein interaction networks. In Peng YF, Zhang YF, eds, BioMedical Engineering and Informatics: New Development and the Future, Vol 1 Institute of Electrical and Electronics Engineers, Los Alamitos, CA, pp 3–7 [Google Scholar]
- Lindemose S, O’Shea C, Jensen MK, Skriver K (2013) Structure, function and networks of transcription factors involved in abiotic stress responses. Int J Mol Sci 14: 5842–5878 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lumba S, Toh S, Handfield LF, Swan M, Liu R, Youn JY, Cutler SR, Subramaniam R, Provart N, Moses A, et al. (2014) A mesoscale abscisic acid hormone interactome reveals a dynamic signaling landscape in Arabidopsis. Dev Cell 29: 360–372 [DOI] [PubMed] [Google Scholar]
- Manning K, Tör M, Poole M, Hong Y, Thompson AJ, King GJ, Giovannoni JJ, Seymour GB (2006) A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat Genet 38: 948–952 [DOI] [PubMed] [Google Scholar]
- Marshall A, Aalen RB, Audenaert D, Beeckman T, Broadley MR, Butenko MA, Caño-Delgado AI, de Vries S, Dresselhaus T, Felix G, et al. (2012) Tackling drought stress: RECEPTOR-LIKE KINASES present new approaches. Plant Cell 24: 2262–2278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miao Y, Laun T, Zimmermann P, Zentgraf U (2004) Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol Biol 55: 853–867 [DOI] [PubMed] [Google Scholar]
- Molinier J, Ries G, Zipfel C, Hohn B (2006) Transgeneration memory of stress in plants. Nature 442: 1046–1049 [DOI] [PubMed] [Google Scholar]
- Nakamura J, Asakura S, Hester SD, de Murcia G, Caldecott KW, Swenberg JA (2003) Quantitation of intracellular NAD(P)H can monitor an imbalance of DNA single strand break repair in base excision repair deficient cells in real time. Nucleic Acids Res 31: e104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park SY, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Chow TF, et al. (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324: 1068–1071 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pérez-Rodríguez P, Riaño-Pachón DM, Corrêa LG, Rensing SA, Kersten B, Mueller-Roeber B (2010) PlnTFDB: updated content and new features of the plant transcription factor database. Nucleic Acids Res 38: D822–D827 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perrella G, Lopez-Vernaza MA, Carr C, Sani E, Gosselé V, Verduyn C, Kellermeier F, Hannah MA, Amtmann A (2013) Histone Deacetylase Complex1 expression level titrates plant growth and abscisic acid sensitivity in Arabidopsis. Plant Cell 25: 3491–3505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raissig MT, Baroux C, Grossniklaus U (2011) Regulation and flexibility of genomic imprinting during seed development. Plant Cell 23: 16–26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26: 139–140 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roudier F, Ahmed I, Bérard C, Sarazin A, Mary-Huard T, Cortijo S, Bouyer D, Caillieux E, Duvernois-Berthet E, Al-Shikhley L, et al. (2011) Integrative epigenomic mapping defines four main chromatin states in Arabidopsis. EMBO J 30: 1928–1938 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadras VO, Richards RA (2014) Improvement of crop yield in dry environments: benchmarks, levels of organisation and the role of nitrogen. J Exp Bot 65: 1981–1995 [DOI] [PubMed] [Google Scholar]
- Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, et al. (2003) TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34: 374–378 [DOI] [PubMed] [Google Scholar]
- Sani E, Herzyk P, Perrella G, Colot V, Amtmann A (2013) Hyperosmotic priming of Arabidopsis seedlings establishes a long-term somatic memory accompanied by specific changes of the epigenome. Genome Biol 14: R59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smyth GK. (2004) Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3: Article 3. [DOI] [PubMed] [Google Scholar]
- Sokol A, Kwiatkowska A, Jerzmanowski A, Prymakowska-Bosak M (2007) Up-regulation of stress-inducible genes in tobacco and Arabidopsis cells in response to abiotic stresses and ABA treatment correlates with dynamic changes in histone H3 and H4 modifications. Planta 227: 245–254 [DOI] [PubMed] [Google Scholar]
- Springer NM. (2013) Epigenetics and crop improvement. Trends Genet 29: 241–247 [DOI] [PubMed] [Google Scholar]
- Tchieu JH, Fana F, Fink JL, Harper J, Nair TM, Niedner RH, Smith DW, Steube K, Tam TM, Veretnik S, et al. (2003) The PlantsP and PlantsT functional genomics databases. Nucleic Acids Res 31: 342–344 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas-Chollier M, Sand O, Turatsinze JV, Janky R, Defrance M, Vervisch E, Brohée S, van Helden J (2008) RSAT: regulatory sequence analysis tools. Nucleic Acids Res 36: W119–W127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toufighi K, Brady SM, Austin R, Ly E, Provart NJ (2005) The Botany Array Resource: e-Northerns, Expression Angling, and promoter analyses. Plant J 43: 153–163 [DOI] [PubMed] [Google Scholar]
- Van Bel M, Proost S, Wischnitzki E, Movahedi S, Scheerlinck C, Van de Peer Y, Vandepoele K (2012) Dissecting plant genomes with the PLAZA comparative genomics platform. Plant Physiol 158: 590–600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van de Velde J, Heyndrickx KS, Vandepoele K (2014) Inference of transcriptional networks in Arabidopsis through conserved noncoding sequence analysis. Plant Cell 26: 2729–2745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Dijk K, Ding Y, Malkaram S, Riethoven JJM, Liu R, Yang J, Laczko P, Chen H, Xia Y, Ladunga I, et al. (2010) Dynamic changes in genome-wide histone H3 lysine 4 methylation patterns in response to dehydration stress in Arabidopsis thaliana. BMC Plant Biol 10: 238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Landeghem S, Hakala K, Rönnqvist S, Salakoski T, Van de Peer Y, Ginter F (2012) Exploring biomolecular literature with EVEX: connecting genes through events, homology, and indirect associations. Adv Bioinformatics 2012: 582765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiner A, Chen HV, Liu CL, Rahat A, Klien A, Soares L, Gudipati M, Pfeffner J, Regev A, Buratowski S, et al. (2012) Systematic dissection of roles for chromatin regulators in a yeast stress response. PLoS Biol 10: e1001369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whittle CA, Otto SP, Johnston MO, Krochko JE (2009) Adaptive epigenetic memory of ancestral temperature regime in Arabidopsis thaliana. Botany 87: 650–657 [Google Scholar]
- Wu TD, Nacu S (2010) Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26: 873–881 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yilmaz A, Mejia-Guerra MK, Kurz K, Liang X, Welch L, Grotewold E (2011) AGRIS: the Arabidopsis Gene Regulatory Information Server, an update. Nucleic Acids Res 39: D1118–D1122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Bernatavichute YV, Cokus S, Pellegrini M, Jacobsen SE (2009) Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol 10: R62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, et al. (2008) Model-based analysis of ChIP-Seq (MACS). Genome Biol 9: R137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zong W, Zhong X, You J, Xiong L (2013) Genome-wide profiling of histone H3K4-tri-methylation and gene expression in rice under drought stress. Plant Mol Biol 81: 175–188 [DOI] [PubMed] [Google Scholar]
- Zulawski M, Braginets R, Schulze WX (2013) PhosPhAt goes kinases: searchable protein kinase target information in the plant phosphorylation site database PhosPhAt. Nucleic Acids Res 41: D1176–D1184 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.