Abstract
Recurrent Clostridium difficile infection (CDI) is of particular concern among health care-associated infections. The role of the microbiota in disease recovery is apparent given the success of fecal microbiota transplantation (FMT) for recurrent CDI. Here, we present a murine model of CDI relapse to further define the microbiota recovery following FMT. Cefoperazone-treated mice were infected with C. difficile 630 spores and treated with vancomycin after development of clinical disease. Vancomycin treatment suppressed both C. difficile colonization and cytotoxin titers. However, C. difficile counts increased within 7 days of completing treatment, accompanied by relapse of clinical signs. The administration of FMT immediately after vancomycin cleared C. difficile and decreased cytotoxicity within 1 week. The effects of FMT on the gut microbiota community were detectable in recipients 1-day posttransplant. Conversely, mice not treated with FMT remained persistently colonized with high levels of C. difficile, and the gut microbiota in these mice persisted at low diversity. These results suggest that full recovery of colonization resistance against C. difficile requires the restoration of a specific community structure.
INTRODUCTION
Recently, the Centers for Disease Control identified Clostridium difficile as an urgent nosocomial threat. In particular, recurrent C. difficile infection (CDI) is a major concern (1). Up to 30% of patients experience recurrent symptoms following cessation of antibiotic therapy, and subsequent recurrences are more likely after a primary recurrence incidence (2, 3). A healthy, diverse gastrointestinal microbial community, termed the gut microbiota, is important in resistance against the development of CDI. The normal gut microbiota provides colonization resistance (the ability to resist pathogen colonization) against C. difficile (4). In both human studies and animal models, perturbation of the indigenous gut microbiota by antibiotic administration has been correlated to a susceptible community structure, ultimately leading to CDI (5–8). Although it is clear that antibiotic use is correlated with the majority of CDI cases (9), the standard therapy for CDI is the administration of an antibiotic regimen with activity against C. difficile (typically vancomycin or metronidazole). Paradoxically, this may further perturb the intestinal microbial community (10, 11). Therefore, it is important to understand the impact of antibiotic treatment for CDI on recovery of the gut microbiota and colonization resistance.
In a murine model of persistent CDI, we have previously shown that the use of sequential antibiotics can delay recovery of bacterial diversity (11). Previous studies have suggested that patients who develop recurrence have a less diverse community than patients who have stable recovery (12). The idea that restoration of the gut microbiota is necessary to prevent or mitigate recurrence is supported by an ∼90% success rate of fecal microbiota transplantation (FMT) (13), and significant recovery of diversity and community membership has been observed following FMT treatment for recurrent CDI (14–16). Future treatment options have thus concentrated on approaches that aid in recovery of colonization resistance.
To study the effect of antibiotic treatment of primary CDI on the gut microbiota, we monitored the dynamics of the gut microbiota and C. difficile-induced colitis in mice following vancomycin treatment of primary CDI. Mice were rendered susceptible to CDI via antibiotic perturbation, infected with C. difficile, and then treated with vancomycin after the development of clinical disease. Without further intervention, mice exhibited CDI recurrence and remained stably colonized by C. difficile even after resolution of disease. Mice treated with FMT immediately after vancomycin treatment cleared C. difficile colonization within 10 days of treatment. Clearance of C. difficile was associated with immediate changes in the gut microbiota following FMT intervention, resulting in stable colonization resistance against C. difficile. These results imply that restoration of colonization resistance by bacterial intervention is necessary for ultimate clearance of C. difficile after longitudinal antibiotic insult to the gut community.
MATERIALS AND METHODS
Ethics statement.
The University Committee on Use and Care of Animals of the University of Michigan, Ann Arbor, approved all protocols of the present study (protocol PRO00006095). These guidelines adhere to the guidelines set by the Public Health Service policy on Humane Care and Use of Laboratory Animals.
Animals and housing.
We obtained 5- to 8-week-old male and female C57BL/6 mice from an established breeding colony at the University of Michigan, originally established with animals from Jackson Laboratories (Bar Harbor, ME). Animals were housed in specific-pathogen-free and biohazard AALAC-accredited facilities and assessed twice daily for physical health. Mice received autoclaved food, water, and bedding, with 12-h light/dark cycles. All cage changes, infection procedures, and sample collections were conducted under a laminar flow hood using appropriate, sterile personal protective equipment between cage contacts. The laminar flow hood work area was sterilized using Sanotracin RTU sporicidal disinfectant between experimental treatment groups to prevent cross-contamination of spores. Mice were housed in groups of three to five animals per cage, and multiple cages were included for each treatment in each of the experiments. Descriptions of mouse metadata for all sequenced samples, including cage numbers and treatment groups, can be found in Table S1 in the supplemental material. All mice were transferred to sterile cages following experimental manipulations throughout the study.
CDI relapse model.
A model of CDI recurrence was developed from our previously described mouse CDI model (11, 17, 18). In all three experiments (n = 82), mice received 0.5 mg of cefoperazone (MP Biochemicals, catalog no. 199695)/ml in sterile drinking water (Gibco Laboratories, catalog no. 15230) for 5 days to render them susceptible to C. difficile. Two days after antibiotic cessation (day 0), mice were challenged via oral gavage with 103 spores of C. difficile strain 630 (ATCC BAA-1382) in 20 μl of sterile phosphate-buffered saline (PBS; Gibco, catalog no. 10010). Control groups received a mock inoculum of PBS. Spores were prepared from a single colony of C. difficile strain 630 for inoculation as described previously (11). Spore pellets were resuspended in 1 ml of sterile water and stored at 4°C. The spore inoculum was heated at 65°C for 20 min to kill residual vegetative cells. The spore concentration was confirmed by enumerating the CFU/ml plated on taurocholate cycloserine cefoxitin fructose agar (TCCFA) plates before and after infection. A subset of mice (n = 62) was treated with 0.4 mg of vancomycin (Sigma, catalog no. V2002)/ml dissolved in sterile drinking water for 5 days (days 4 to 9 postinfection [p.i.]). After infection and vancomycin treatment, mice were monitored daily for clinical signs of CDI and weight loss. A subset of mice was euthanized at days 4, 9, and 16 p.i. for histopathological scoring and cecal toxin detection. Two mice fell below our experimental weight threshold (80% of preinfection weight) in the vancomycin-treated group (from different cages) on day 13 but were still considered an experimental endpoint of CDI relapse. Control mice (n = 3 for each experimental replicate) received both cefoperazone and vancomycin treatment concurrently with other treatment groups but were not challenged with C. difficile. We did not detect C. difficile in these mice throughout the experiments.
To investigate the effect of FMT intervention on relapse prevention, a third experiment was conducted in which mice (n = 23) received one FMT via oral gavage on day 11 p.i. (n = 9) or 12 p.i. (n = 9) or two FMTs on both days (n = 5). Fecal samples for FMT were collected from 10 conventional, untreated, age-matched mice. A dilution of eight fecal pellets in 1.5 ml of prereduced PBS was homogenized by vortexing. A 100-μl portion of the mixture was administered to each animal by oral gavage.
All mice (with or without FMT) in this experiment received a single intraperitoneal (i.p.) injection of clindamycin (Sigma, catalog no. C5269) at day 35 p.i. to test susceptibility to C. difficile (18). A 1-mg/ml preparation of clindamycin dissolved in sterile 0.9% sodium chloride (Hospira, catalog no. RL-0496) was filter sterilized (0.22-μm pore size; Fisherbrand, catalog no. 097203), and animals received a 10-mg/kg i.p. injection. Animals were monitored until day 42 p.i., at which point all mice were euthanized. Animal weights were recorded daily for all three experiments, and signs of disease (diarrhea, hunched back, scruffiness) were recorded following primary infection and relapse. Humane CO2 euthanization was used if animals exhibited any moribund disease signs or lost more than 20% of their preinfection weight.
C. difficile isolation from stool samples.
Stool samples were collected directly from mice and immediately passed into an anaerobic chamber (Coy Laboratory Products). Samples were homogenized in a 1-to-10 ratio of prereduced, sterile PBS based on sample weight, and 20 μl of this mixture was further serially diluted up to a 10−6 dilution factor. Then, 100-μl portions of multiple dilutions of each sample were plated onto TCCFA and incubated anaerobically at 37°C overnight. The CFU/ml were enumerated the next day.
C. difficile cytotoxin assay.
Cytotoxicity assays were adapted as previously described (17). Briefly, Green African monkey kidney epithelial cells (Vero) were grown to confluence in Dulbecco modified Eagle medium (Gibco Laboratories, catalog no. 11965) with 10% fetal bovine serum (Gibco Laboratories, catalog no. 16140) and 1% penicillin-streptomycin solution (Gibco Laboratories, catalog no. 15070) and plated for a total cell density of 105 cells/well. Next, 10 μl of 0.22-μm-filtered murine fecal or cecal sample was serially diluted in sterile PBS to 10−6 and added to Vero cells for overnight incubation at 37°C. Filtered samples were tested in duplicate, each having a corresponding antitoxin well (goat antitoxin; TechLab, catalog no. T1000). A positive control of isolated C. difficile toxin B (List Biologicals, catalog no. 155D) was used. Each well of the cell culture plate was assessed for 80% cell rounding. The cytotoxic titer was determined for each sample by calculating the log10 reciprocal titer per gram of fecal or cecal content.
Histopathologic evaluation.
A board-certified veterinary pathologist, blinded to the experimental groups at the time of evaluation, performed light microscopic evaluation. Histological lesions were scored using a modification of previously defined criteria (17). In brief, slides were scored for edema (0 to 4), inflammation (0 to 4), and epithelial damage (0 to 4) and summed for a summary score (0 to 12). Previous inflammation scoring criteria were modified to reflect number of foci as follows: 0, no inflammation; 1, minimal, multifocal inflammation (few foci); 2, moderate, multifocal inflammation (numerous foci); 3, severe inflammation (larger and coalescing foci); and 4, marked (same as score 3 with abscesses or extensive mural involvement). Inflammation was also descriptively characterized with respect to predominant location (submucosal or mucosal) and predominant type: neutrophilic, mononuclear (macrophagic and lymphoplasmacytic), or mixed inflammation.
Fecal pellet DNA extraction and 16S rRNA sequencing.
For DNA extraction of fecal pellets, 200 to 300 μl of diluted material was extracted using a MoBio PowerMag soil isolation kit (MoBio Laboratories, catalog no. 271004EP) optimized for the epMotion 5075 TMX (Eppendorf). The DNA libraries were prepared by the University of Michigan Host Microbiome Core as described previously (19). Briefly, DNA was PCR amplified using a set of barcoded dual-index primers described by Kozich et al. (20) that are specific to the V4 regions of the 16S rRNA gene. Each 20-μl PCR was composed of 5 μl of a 4 μM equimolar primer set, 0.15 μl of Accuprime High-Fidelity Taq, 2 μl of 10× Accuprime PCR II buffer (Life Technologies, catalog no. 12346094), 11.85 μl of sterile PCR-grade water, and 1 μl of DNA template. The PCR cycle consisted of 2 min at 95°C, followed by 30 cycles of 95°C for 20 s, 55°C for 15 s, and 72°C for 5 min, followed in turn by 72°C for 10 min. A SequelPrep normalization plate kit (Life Technologies, catalog no. A10510-01) was used to normalize the DNA library to the lowest concentration of the pooled plates. The Kapa Biosystems Library Quantification kit for Illumina platforms (KapaBiosystems, catalog no. KK4854) was used to determine the pooled library concentration, and the Agilent Bioanalyzer high-sensitivity DNA analysis kit (catalog no. 5067-4626) was used to determine the amplicon size. Amplicons were sequenced on the Illumina MiSeq platform using a MiSeq Reagent 222 kit V2 (catalog no. MS-102-2003) for 500 cycles according to the manufacturer's instructions with modifications for the primer set. Libraries were prepared according to Illumina's protocol “Preparing Libraries for Sequencing on the MiSeq” (part 15039740, Rev. D) for 2 nM libraries. The final load concentration was 4 pM, spiked with 10% PhiX to add diversity. Sequencing reagents were prepared according to the “16S Sequencing with the Illumina MiSeq Personal Sequencer” protocol as previously described, with custom read 1/read 2 and index primers added to the reagent cartridge, generating FASTQ files with paired end reads (20).
Analysis of microbiota community.
Raw sequence files have been deposited in the Sequence Read Archive database under the project PRJNA279891, samples SAMN03360503 to SAMN03360790. Sequences were analyzed using mothur v1.33.3 (21). Briefly, sequences were aligned to the V4 region using the SILVA rRNA database project, release 109 (22). Chimeric sequences were removed using UCHIME (23). All samples with <1,000 sequences were discarded. For phylotype identification, sequences were taxonomically classified at an 80% minimum bootstrap to the mothur-adapted RDP database (v9) using the Wang method (24). Operational taxonomic units (OTU) were identified using a 97% similarity rate and used for downstream community analyses. The inverse Simpson index was used to calculate alpha diversity, and the Yue and Clayton dissimilarity metric was used for beta diversity measures (25). Standard R packages were used for most statistical analyses and data visualization. The R package Vegan was used to calculate pairwise differences in heat map clustering (26) and visualized with the package gplots (27). A Wilcoxon rank sum test was used to detect significant differences in C. difficile colonization and cytotoxicity between two treatment groups. The pairwise Wilcoxon rank sum test in R was used to detect differences in phylotype relative abundance and diversity between three or more groups. The Friedman test was used to detect differences in relative abundance of phylotypes over time within mice that did or did not receive FMT. Analysis of molecular variance (AMOVA) was used to detect significant clustering of different treatment groups in nonmetric multidimensional scaling (NMDS) (28). Detailed steps in generating specific figures in the manuscript, as well as raw metadata used to create these graphs, can be found at https://github.com/aseekatz/mouseRM.
RESULTS
Vancomycin treatment of primary CDI leads to relapse.
In our previous studies, we have observed that infection of cefoperazone-treated mice with C. difficile strain 630 results in a self-limited colitis and chronic C. difficile colonization (11, 17). We extended this model to include treatment of established infection with vancomycin in order to determine how this would impact colonization and disease course. Mice were made susceptible to C. difficile colonization by administration of cefoperazone in drinking water for 5 days (Fig. 1A). Two days following cessation of the antibiotic, mice received 103 spores of C. difficile strain 630 via oral gavage. Starting 1 day p.i., C. difficile in feces was detected and enumerated by plating (Fig. 1B). As previously reported, infected animals developed clinical signs of disease, including lethargy and weight loss, within 4 days p.i. (see Fig. S1 in the supplemental material). Titers of C. difficile toxin in feces were high in infected animals by day 4 p.i. (Fig. 1C). At this time, a subset of mice (n = 27, representing two experimental replicates) was treated with vancomycin administered in drinking water for 5 days (days 4 to 9, Fig. 1A). Mice were observed for signs of CDI relapse after vancomycin cessation, and their recovery was compared to that of mice that did not receive vancomycin treatment (n = 20).
FIG 1.
Relapse of C. difficile infection (CDI) in mice treated with vancomycin. (A) Experimental timeline of murine model of CDI relapse. (B) Mean C. difficile colonization levels of untreated mice with primary CDI (no vancomycin; yellow) or vancomycin-treated mice that relapsed (blue). Error bars represent the standard deviations, and a horizontal line indicates the limit of detection. (C) Cytotoxin activity (expressed as the log10 reciprocal cytotoxin dilution per gram of cecal content) following C. difficile infection was determined by using a Vero cell cytotoxicity assay at experimental endpoints on days 4, 9, and 16. Two mice in the vancomycin-treated group fell below weight threshold on day 13. Points represent individual samples, and bars represent median and upper/lower quartiles. (D and E) Summary score of histopathological damage in the cecum (D) and colon (E) after CDI (days 13 to 16), with antibiotic-only treated mice (pink) as a control. Significance was determined by using a Wilcoxon rank sum test (*, P < 0.05; **, P < 0.005; ***, P < 0.0005). (F and G) Histopathology of the cecum (F) and colon (G) at days 13 to 16 p.i. for vancomycin-treated mice (top panels) and mice that did not receive vancomycin (bottom panels). All images were taken at ×200 magnification (scale bars, 100 μm).
During vancomycin treatment, both C. difficile colonization levels and cecal toxin activity decreased below levels of detection (Fig. 1B and C; Wilcoxon rank sum test, P < 0.0005). However, C. difficile rebounded by day 12 p.i. in vancomycin-treated mice, 3 days after the cessation of vancomycin. C. difficile toxin levels also returned to high levels by day 16 p.i., correlating with high C. difficile levels. Weight loss correlated with the observed increase in C. difficile colonization and toxin levels after the end of vancomycin treatment, a finding consistent with the development of disease relapse (see Fig. S1 in the supplemental material). Two vancomycin-treated mice fell below our weight threshold (80% of preinfection weight) on day 13 but were included in the final analysis of CDI relapse. In contrast, C. difficile colonization levels remained high in mice that did not receive vancomycin and persisted throughout the 16 days of infection, whereas the toxin levels decreased (Wilcoxon test, P < 0.0005) with the resolution of the initial weight loss (Fig. 1).
A subset of the animals was euthanized, and histopathologic examination of intestinal tissue revealed characteristic inflammation and epithelial damage. Histological alterations, as previously described for this model, consisted of submucosal edema, inflammation, and epithelial damage (principally enterocyte vacuolation and erosion) in the cecum and colon (see Fig. S2 in the supplemental material) (17). At 4 days p.i., histopathological damage was prominent in all mice, particularly within the cecum (4 days p.i., Fig. 1; see also Fig. S2 in the supplemental material). Vancomycin treatment decreased summary histopathological scores for CDI (9 days p.i.), but these rebounded at 7 days following the cessation of vancomycin, reflecting CDI relapse (days 13 to 16 p.i., Fig. 1D and E). In contrast, mice that did not receive vancomycin had a slow decline in summary scores over the 13- to 16-day observation period (Fig. 1D and E; see also Fig. S2 in the supplemental material). In addition, we observed a difference in inflammatory cell type following vancomycin, indicative of recurrence of primary disease. Both groups had neutrophilic (acute) inflammation in the cecum during primary CDI (see Fig. S3 in the supplemental material). During relapse (days 13 to 16 p.i.), untreated mice transitioned to mononuclear (lymphocytic, plasmacytic, and macrophagic) or mixed (mononuclear and neutrophilic) inflammation, whereas vancomycin-treated mice re-experienced neutrophilic (acute) inflammation (see Fig. S3 in the supplemental material).
FMT results in C. difficile clearance.
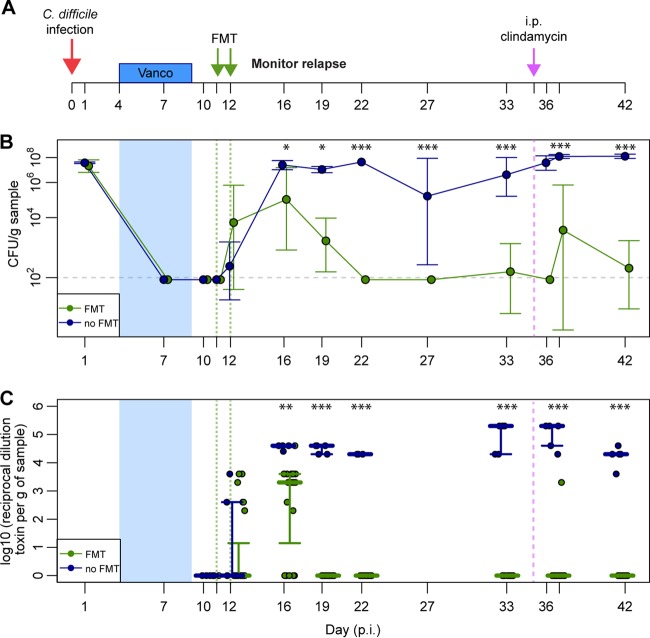
We next evaluated whether bacterial intervention altered the course of CDI relapse that we observed in vancomycin-treated animals. We administered a fecal microbiota transplant (FMT) to C. difficile-infected mice via oral gavage, prepared from the feces of conventional, untreated mice (n = 23; Fig. 2A). One FMT inoculum (on either day 11 or 12 p.i.) or two FMT inocula (on both day 11 and 12 p.i.) were administered to determine an optimal time point of treatment. Despite different inoculum days, fecal C. difficile levels were detectable by day 12 p.i. in all treatment groups (Fig. 2B). Both weight loss and increased cytotoxicity were observed in FMT-treated mice and did not differ significantly from results for vancomycin-treated mice that did not receive FMT (see Fig. S1 in the supplemental material and Fig. 2C; Wilcoxon test). Within 5 days of FMT (day 16 p.i.), C. difficile load and cytotoxicity were significantly decreased compared to results for mice that did not receive FMT (Fig. 2B; Wilcoxon test, P < 0.0005) and undetectable by day 22 following FMT.
FIG 2.
CDI in mice with or without FMT after vancomycin treatment of primary CDI. (A) Experimental design and timeline of mice with CDI relapse with or without FMT. (B) Mean C. difficile colonization levels (expressed as CFU/g of fecal pellets) of mice with relapse without FMT (blue) or with FMT (green). Error bars represent standard deviations, and the horizontal line indicates the limit of detection. (C) Cytotoxin activity (expressed as the log10 reciprocal cytotoxin dilution per gram of fecal content) was determined using a Vero cell cytotoxicity assay in mice treated with vancomycin only (blue) or vancomycin followed by FMT (green). Points represent samples collected from either mice that received only vancomycin (blue) or mice that also received FMT (green). Bars represent the median per group and lower/upper quartiles. Significance was determined by using a Wilcoxon rank sum test (*, P < 0.05; **, P < 0.005; ***, P < 0.0005).
To assess C. difficile resistance in mice that received FMT, an i.p. injection of clindamycin was administered 35 days after initial infection to both FMT and non-FMT mice. We previously demonstrated that a single i.p. administration of clindamycin in animals 6 weeks after a course of cefoperazone would result in susceptibility to C. difficile (18). However, in the absence of previous cefoperazone, a single dose of clindamycin would not be sufficient to overcome colonization resistance to C. difficile. Interestingly, clindamycin administration did not result in stable C. difficile colonization in mice that had received FMT, suggesting recovery of a community state resistant to C. difficile (Fig. 2B). Although irregular detection of C. difficile was observed randomly in some mice, colonization levels did not persist and cytotoxicity remained low or undetectable (Fig. 2C; Wilcoxon test, P < 0.0005). Conversely, mice that did not receive FMT intervention were persistently colonized by C. difficile throughout the study, and toxin levels remained at detectable levels. These mice remained stably colonized by C. difficile throughout clindamycin administration, which did not impact colonization or toxin levels.
FMT promotes recovery of microbial community and clearance of C. difficile.
The above results suggest that FMT restored long-term colonization resistance against C. difficile. Presumably, this resulted in changes to the gut microbiota that prevented subsequent C. difficile colonization. To assess the effect of antibiotic use and FMT on the gut microbiota, the fecal community from a subset of mice with or without FMT was characterized by 16S rRNA-encoding gene sequence analysis. 7,396,962 high-quality, filtered sequences (mean: 25,683 sequences/sample) were classified into OTU with 97% phylogenetic similarity and taxonomically classified to the genus level (see Tables S1 and S2 in the supplemental material).
To compare the beta diversity of the fecal community structure throughout the study, we calculated the dissimilarity between community structures using the Yue and Clayton dissimilarity metric (θYC), which accounts for the relative abundance of the OTU present (25). NMDS of these distances revealed distinct clustering of samples from mice that did or did not receive FMT (Fig. 3). Using AMOVA, no significant difference was detected between preantibiotic, donor, and post-FMT samples (28). However, these samples clustered separately from postantibiotic (vancomycin or cefoperazone), non-FMT samples (AMOVA; P < 0.001). Using a biplot of the NMDS axes, we identified which OTU were responsible for this community structure. Recovery of the community toward the baseline community structure was driven by five OTU that classified under the Porphyromonadaceae family, whereas postantibiotic and postinfection samples from non-FMT mice were mainly of Lactobacillus or C. difficile (Clostridium XI) itself. Postclindamycin samples from FMT mice clustered separately, driven by a single Enterobacteriaceae-classified OTU (AMOVA; P < 0.001).
FIG 3.
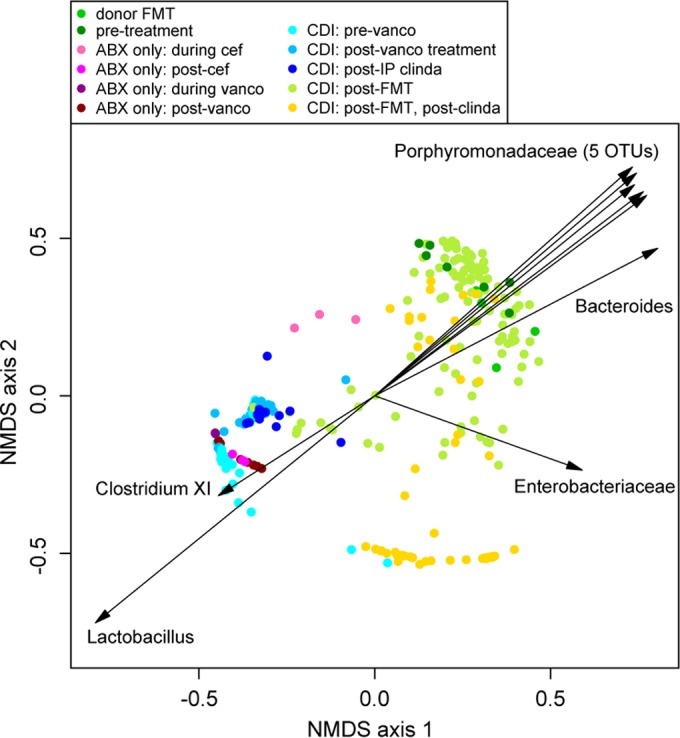
Effect of antibiotics and FMT on the microbial community of C. difficile-infected mice. NMDS ordination was calculated from the Yue and Clayton dissimilarity metric (θYC) on OTU at a 97% cutoff. Biplot values of the OTU driving the NMDS ordination were derived using a cutoff of P < 0.001 for at least one axis, a total sequence abundance of >100,000 and a length of >0.77.
To identify when community shifts occurred, we compared diversity and θYC distance over time with or without FMT intervention. Overall, diversity as measured by the inverse Simpson index was significantly decreased following cefoperazone (pairwise Wilcoxon test, P < 0.0005) (Fig. 4A). Without FMT, this low diversity persisted throughout the 42-day experiment. Conversely, the community diversity increased to levels similar to that of the donor mice at a similar rate for all FMT time points. In addition, with FMT intervention, the fecal microbiota shifted toward a community structure similar to its starting point within a day of FMT administration (Fig. 4A and B). Comparison of the θYC distance of FMT and non-FMT mice to the preantibiotic state over time remained significantly different between the two groups (Wilcoxon test; P < 0.0005) (Fig. 4B). Day-by-day comparison of the community structure revealed differences between the community structure by day 16 p.i. (Fig. 4C).
FIG 4.
Changes in community diversity and structure over time during relapse of CDI and FMT. (A) Community diversity of the microbiota in mice with CDI relapse (inverse Simpson index, y axis). Points represent individual samples, and lines represent the mean diversity per group. (B) Mean distance between the community structure on sampling day and conventional/untreated over time in mice without or with FMT, calculated using the Yue and Clayton dissimilarity metric (θYC). (C) Mean θYC distance between the community structure of mice with or without FMT over time. Error bars represent upper and lower quartiles. A pairwise Wilcoxon rank sum test was used to calculate significance between mice with or without FMT (*, P < 0.05; **, P < 0.005; ***, P < 0.0005).
Similar trends were observed in community membership. Prior to antibiotic exposure, the fecal microbiota consisted of a diverse population of Bacteroidetes, including members of the Porphyromonadaceae family (median relative abundance, 42%), Bacteroides (9%), and Barnesiella (3%), and Firmicutes, including unclassified Lachnospiraceae (9%) and Ruminococcaceae (2%) family members, Lactobacillus (5%), and Turicibacter (1%) (Fig. 5A, left panel; see Fig. S4 in the supplemental material). Significant changes to the fecal microbiota community were observed following both cefoperazone and vancomycin use (see Table S3 in the supplemental material; pairwise Wilcoxon test). After cefoperazone exposure, the community diversity was compressed to two OTU, Lactobacillus (66%) and Turicibacter (12%), a result consistent with previous studies using the same model (8, 19) (Fig. 5A, middle panel). In infected animals, C. difficile (classified as Clostridium XI, 14%) was detected prior to vancomycin treatment. This community structure was perpetuated following vancomycin treatment in mice that did not receive FMT, although an OTU classified as Akkermansia (47 to 69%) was detected in some cages (Fig. 5A, right panel; Fig. S4 in the supplemental material). Without FMT intervention, mice remained dominated by these three OTU throughout the study even after an i.p. dose of clindamycin, reflective of a prolonged alteration of the microbial community after the dual antibiotic exposure (Fig. 3B; Friedman rank sum test).
FIG 5.
Taxonomic profile of the microbiota in mice treated with FMT after vancomycin treatment of CDI. (A) Median relative abundance (bars) of select dominant phylotypes in microbiota community of untreated mice (left panel, n = 11), during CDI (middle, n = 24, days 1 and 4), and following vancomycin (right, n = 20) before relapse. Error bars represent interquartile range. (B and C) Median relative abundance of dominant phylotypes in mice without FMT (B) and mice with FMT (C) after CDI relapse (left panel, n = 3 without FMT and n = 19 with FMT), 1 day after clindamycin (middle panel, n = 5 and n = 19), and 1 week after clindamycin (right panel, n = 5 and n = 18). A Friedman rank sum test (paired) was used to calculate significance over time in panels B and C (***, P < 0.0005).
In comparison, we observed an immediate shift of the microbiota community to the baseline (preantibiotic state) in mice treated with FMT after vancomycin treatment. Several Bacteroidetes members that were present prior to antibiotic use were detectable within 1-day post-FMT (see Fig. S4 in the supplemental material). By day 27 p.i., the most dominant members within the community resembled those present in the donor (Fig. 5A and C). In particular, Bacteroidetes members, such as Porphyromonadaceae family members (median, 53%), Bacteroides (7%), or Barnesiella (4%), were observed (Fig. 5C; see also Fig. S4 in the supplemental material). Unsupervised clustering of OTU classified at a 97% cutoff using the Morisita similarity index revealed that preantibiotic samples and the majority of post-FMT samples clustered together (see Fig. S5 in the supplemental material). These were distinct from postantibiotic, non-FMT samples, which clustered separately.
Lawley and others have used clindamycin to unmask C. difficile colonization in a model of chronic colonization (29, 30), and other investigators have observed prolonged susceptibility to C. difficile after clindamycin exposure alone (31). Although clindamycin disrupted the community of FMT-treated mice, the community was capable of recovering within 7 days. At 1-day postclindamycin (day 36 p.i.), nearly all mice were dominated by a single OTU classified under the Enterobacteriaceae family (91%) (Fig. 5C). However, this disruption did not lead to persistent C. difficile colonization (day 42 p.i., Fig. 2B), and community recovery was observed rapidly (Fig. 5C). This is in stark contrast to the distinct community shifts following cefoperazone or vancomycin, which were incapable of recovery or clearing C. difficile without FMT intervention (Fig. 5 and see Fig. S5 in the supplemental material; Friedman rank sum test). Mice that did not receive FMT remained unaffected by clindamycin, and comparison of the fecal microbiota community between FMT and non-FMT mice after clindamycin suggests that the microbiota remains distinct between the two groups (Fig. 4C). Despite pronounced changes in diversity and community structure after clindamycin in FMT-treated mice, we observed only transient C. difficile colonization, suggesting recovery of colonization resistance.
DISCUSSION
In patients who develop C. difficile infection, primary therapy currently consists of the administration of antibiotics with intrinsic activity against the pathogen (32). The goal of this therapy is to decrease the pathogen burden, therefore decreasing toxin levels and limiting damage to the intestinal epithelium. Once ongoing damage to the epithelium is limited, restoration of normal intestinal function can occur. In order to prevent recurrence of disease once the therapy targeted against C. difficile is discontinued, restoration of the indigenous microbiome and functional colonization resistance is also required. Current antibiotic therapy does not facilitate recovery of the microbiome in patients with CDI. In fact, antibiotic treatment of CDI may paradoxically contribute to further disruption of the indigenous microbiota, leading to recurrences of disease. We and others have shown that disruption of the indigenous intestinal microbiota with antibiotics that have intrinsic activity against C. difficile can actually eliminate colonization resistance permitting development of CDI once these antibiotics are discontinued (8, 11, 33).
Recovery of the microbiota and restoration of colonization resistance in patients with CDI can occur via two nonexclusive routes. During antibiotic treatment, the residual microbiota can begin to recover during the time C. difficile levels are being suppressed. Once the anti-C. difficile treatment was discontinued, the resultant intestinal community would be resistant to outgrowth of residual C. difficile within the gut or to re-exposure from environmental spores. In addition, exogenous exposure to microbes from the environment and food could contribute to the restoration of the microbiota and colonization resistance. Fecal microbiota transplantation represents the ultimate restoration of colonization assistance via exposure to exogenous microbes, as exemplified by the observed changes in community structure following FMT (10, 15, 34). Here, we describe a novel model of relapsing CDI in mice in which restoration of full colonization resistance against C. difficile requires exogenous reconstitution of the gut microbiota after antimicrobial treatment. In this model, pathogen elimination and microbiota restoration did not spontaneously occur following treatment with vancomycin, an antibiotic with activity against C. difficile. Once vancomycin treatment was discontinued, C. difficile infection rebounded, accompanied by increasing levels of toxin and the reappearance of acute inflammation. This suggests that the gut microbial community present at the end of vancomycin treatment is not sufficient to restore colonization resistance.
Although we cannot determine the source of spores that resulted in recolonization and disease relapse, the administration of FMT was robust enough to restore colonization resistance and decrease C. difficile to undetectable levels, be it within the gastrointestinal tract or elsewhere on the animal. Further, we used clindamycin dosing to “unmask” low levels of colonization in animals that received FMT analogous to what has been observed by Lawley et al. (29, 35). Although we did observe transient shedding of C. difficile in a minority of animals that received clindamycin, the C. difficile levels did not persist, suggesting restored colonization resistance. This is in line with our previous results that demonstrated that clindamycin exposure up to 6 weeks after cefoperazone exposure has resulted in shedding of low numbers of C. difficile without stable colonization or the development of disease (18). Despite other studies that have observed prolonged susceptibility to C. difficile after only a single dose of clindamycin (31), we did not observe prolonged effects on the microbiota in our mice. This is likely a reflection of differences in the microbiota composition across facilities.
Taken together, our results indicate that while antibiotic treatment can suppress C. difficile to undetectable levels, full restoration of colonization resistance and thus the prevention of recurrence require recovery of the intestinal microbiome. Our results in a murine model of CDI relapse corroborate those observed by others in both animal models (33, 35) and humans (14–16), suggesting that additional assistance in restoration of the microbiota may be required to allow complete treatment of CDI without the development of recurrence in patients. In the future, it may be possible to identify the subset of patients susceptible to recurrent CDI based on the extent of microbial community recovery in particular individuals. In these patients, the administration of additional microbes potentially in the form of rationally selected probiotics, along with antibiotic therapy targeted against C. difficile, would be able to limit the development of recurrent disease. This would represent a significant advance in our ability to address this important nosocomial threat.
Supplementary Material
ACKNOWLEDGMENTS
This study was supported by the National Institutes of Health (NIH) grants U19A090871 and P30DK034922 from the Michigan Gastrointestinal Peptide Research Center. A.M.S. is supported by grant 2L1TR00043 from the National Center for Advancing Translational Sciences. C.M.T. is supported by the NIH Metabolomics Common Fund and National Institute of General Medical Science under award K01GM109236. This research was also aided by the Host Microbiome Initiative and computational resources and services provided by Advanced Research Computing at the University of Michigan, Ann Arbor.
Footnotes
Supplemental material for this article may be found at http://dx.doi.org/10.1128/IAI.00459-15.
REFERENCES
- 1.Lessa FC, Mu Y, Bamberg WM, Beldavs ZG, Dumyati GK, Dunn JR, Farley MM, Holzbauer SM, Meek JI, Phipps EC, Wilson LE, Winston LG, Cohen JA, Limbago BM, Fridkin SK, Gerding DN, McDonald LC. 2015. Burden of Clostridium difficile infection in the United States. N Engl J Med 372:825–834. doi: 10.1056/NEJMoa1408913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Musher DM, Aslam S, Logan N, Nallcheru S, Bhaila I, Borchert F, Hamill RJ. 2005. Relatively poor outcome after treatment of Clostridium difficile colitis with metronidazole. Clin Infect Dis 40:1586–1590. doi: 10.1086/430311. [DOI] [PubMed] [Google Scholar]
- 3.Ananthakrishnan AN. 2011. Clostridium difficile infection: epidemiology, risk factors and management. Nat Rev Gastroenterol Hepatol 8:17–26. doi: 10.1038/nrgastro.2010.190. [DOI] [PubMed] [Google Scholar]
- 4.van der Waaji D. 1989. The ecology of the human intestine and its consequences for overgrowth by pathogens such as Clostridium difficile. Annu Rev Microbiol 43:69–87. doi: 10.1146/annurev.mi.43.100189.000441. [DOI] [PubMed] [Google Scholar]
- 5.Theriot CM, Koenigsknecht MJ, Carlson PE Jr, Hatton GE, Nelson AM, Li B, Huffnagle GBZLJ, Young VB. 2014. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat Commun 5:3114. doi: 10.1038/ncomms4114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schubert AM, Rogers MA, Ring C, Mogle J, Petrosino JP, Young VB, Aronoff DM, Schloss PD. 2014. Microbiome data distinguish patients with Clostridium difficile infection and non-C. difficile-associated diarrhea from healthy controls. mBio 5:e01021-14. doi: 10.1128/mBio.01021-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Antharam VC, Li EC, Ishmael A, Sharma A, Mai V, Rand KH, Wang GP. 2013. Intestinal dysbiosis and depletion of butyrogenic bacteria in Clostridium difficile infection and nosocomial diarrhea. J Clin Microbiol 51:2884–2892. doi: 10.1128/JCM.00845-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bassis CM, Theriot CM, Young VB. 2014. Alteration of the murine gastrointestinal microbiota by tigecycline leads to increased susceptibility to Clostridium difficile infection. Antimicrob Agents Chemother 58:2767–2774. doi: 10.1128/AAC.02262-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Owens RC Jr, Donskey CJ, Gaynes RP, Loo VG, Muto CA. 2008. Antimicrobial-associated risk factors for Clostridium difficile infection. Clin Infect Dis 46(Suppl 1):S19–S31. doi: 10.1086/521859. [DOI] [PubMed] [Google Scholar]
- 10.Seekatz AM, Young VB. 2014. Clostridium difficile and the microbiota. J Clin Invest 124:4182–4189. doi: 10.1172/JCI72336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Theriot CM, Schumacher CA, Bassis CM, Seekatz AM, Young VB. 2014. The effects of tigecycline and vancomycin administration on established Clostridium difficile infection. Antimicrob Agents Chemother 59:1596–1604. doi: 10.1128/AAC.04296-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chang JY, Antonopoulos DA, Kalra A, Tonelli A, Khalife WT, Schmidt TM, Young VB. 2008. Decreased diversity of the fecal microbiome in recurrent Clostridium difficile-associated diarrhea. J Infect Dis 197:435–438. doi: 10.1086/525047. [DOI] [PubMed] [Google Scholar]
- 13.Gough E, Shaikh H, Manges AR. 2011. Systematic review of intestinal microbiota transplantation (fecal bacteriotherapy) for recurrent Clostridium difficile infection. Clin Infect Dis 53:994–1002. doi: 10.1093/cid/cir632. [DOI] [PubMed] [Google Scholar]
- 14.Seekatz AM, Aas J, Gessert CE, Rubin TA, Saman DM, Bakken JS, Young VB. 2014. Recovery of the gut microbiome following fecal microbiota transplantation. mBio 5:e00893-14. doi: 10.1128/mBio.00893-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hamilton MJ, Weingarden AR, Unno T, Khoruts A, Sadowsky MJ. 2013. High-throughput DNA sequence analysis reveals stable engraftment of gut microbiota following transplantation of previously frozen fecal bacteria. Gut Microbes 4:125–135. doi: 10.4161/gmic.23571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Song Y, Garg S, Girotra M, Maddox C, von Rosenvinge EC, Dutta A, Dutta S, Fricke FW. 2013. Microbiota dynamics in patients treated with fecal microbiota transplantation for recurrent Clostridium difficile infection. PLoS One 8:e81330. doi: 10.1371/journal.pone.0081330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Theriot CM, Koumpouras CC, Carlson PE, Bergin II, Aronoff DM, Young VB. 2011. Cefoperazone-treated mice as an experimental platform to assess differential virulence of Clostridium difficile strains. Gut Microbes 2:326–334. doi: 10.4161/gmic.19142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Reeves AE, Theriot CM, Bergin IL, Huffnagle GB, Schloss PD, Young VB. 2011. The interplay between microbiome dynamics and pathogen dynamics in a murine model of Clostridium difficile infection. Gut Microbes 2:145–158. doi: 10.4161/gmic.2.3.16333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Koenigsknecht MJ, Theriot CM, Bergin IL, Schumacher CA, Schloss PD, Young VB. 2015. Dynamics and establishment of Clostridium difficile infection in the murine gastrointestinal tract. Infect Immun 83:934–941. 10.1128/IAI.02768-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD. 2013. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. doi: 10.1128/AEM.01043-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF. 2009. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. doi: 10.1128/AEM.01541-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glockner FO. 2007. SILVA: a comprehensive online resource for quality checked and aligned rRNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196. doi: 10.1093/nar/gkm864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. 2011. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi: 10.1093/bioinformatics/btr381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wang Q, Garrity GM, Tiedje JM, Cole JR. 2007. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. doi: 10.1128/AEM.00062-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yue JC, Clayton MK. 2005. A similarity measure based on species proportions. Commun Stat Theory Methods 34:2123–2131. doi: 10.1080/STA-200066418. [DOI] [Google Scholar]
- 26.Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MH, Wagner H. 2013. Community Ecology Package, v2.0-10. https://cran.r-project.org/. [Google Scholar]
- 27.Warnes GR, Bolker B, Bonebakker L, Gentleman R, Liaw WH, Lumley T, Maechler M, Magnusson A, Moeller S, Schwartz M, Venables B. 2013. gplots: various R programming tools for plotting data (v2.12.1). http://cran.r-project.org/web/packages/gplots/gplots.pdf. [Google Scholar]
- 28.Excoffier L, Smouse PE, Quattro JM. 1992. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lawley TD, Clare S, Walker AW, Goulding D, Stabler RA, Croucher N, Mastroeni P, Scott P, Raisen C, Mottram L, Fairweather NF, Wren BW, Parkhill J, Dougan G. 2009. Antibiotic treatment of Clostridium difficile carrier mice triggers a supershedder state, spore-mediated transmission, and severe disease in immunocompromised hosts. Infect Immun 77:3661–3669. doi: 10.1128/IAI.00558-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chen X, Katchar K, Goldsmith JD, Nanthakumar N, Cheknis A, Gerding DN, Kelly CP. 2008. A mouse model of Clostridium difficile-associated disease. Gastroenterology 135:1984–1992. doi: 10.1053/j.gastro.2008.09.002. [DOI] [PubMed] [Google Scholar]
- 31.Buffie CG, Jarchum I, Equinda M, Lipuma L, Gobourne A, Viale A, Ubeda C, Xavier J, Pamer EG. 2012. Profound alterations of intestinal microbiota following a single dose of clindamycin results in sustained susceptibility to Clostridium difficile-induced colitis. Infect Immun 80:62–73. doi: 10.1128/IAI.05496-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cohen SH, Gerding DN, Johnson S, Kelly CP, Loo VG, McDonald LC, Pepin J, Wilcox MH. 2010. Clinical practice guidelines for Clostridium difficile infection in adults: 2010 update by the Society for Healthcare Epidemiology of America (SHEA) and the Infectious Diseases Society of America (IDSA). Infect Control Hosp Epidemiol 31:431–455. doi: 10.1086/651706. [DOI] [PubMed] [Google Scholar]
- 33.Buffie CG, Bucci V, Stein RR, McKenney PT, Ling L, Gobourne A, No D, Liu H, Kinnebrew M, Viale A, Littmann E, van den Brink MR, Jenq RR, Taur Y, Sander C, Cross JR, Toussaint NC, Xavier JB, Pamer EG. 2015. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517:205–208. doi: 10.1038/nature13828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Rao K, Young VB. 2015. Fecal microbiota transplantation for the management of Clostridium difficile infection. Infect Dis Clin North Am 29:109–122. doi: 10.1016/j.idc.2014.11.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lawley TD, Clare S, Walker AW, Stares MD, Connor TR, Raisen C, Goulding D, Rad R, Schreiber F, Brandt C, Deakin LJ, Pickard DJ, Duncan SH, Flint HJ, Clark TG, Parkhill J, Dougan G. 2012. Targeted restoration of the intestinal microbiota with a simple, defined bacteriotherapy resolves relapsing Clostridium difficile disease in mice. PLoS Pathog 8:e1002995. doi: 10.1371/journal.ppat.1002995. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.