Abstract
Somatic hypermutation (SHM) is critical for antibody affinity maturation and the generation of memory B cells. Somatic mutations consist mainly of single nucleotide changes with rare insertions and deletions. Such changes would be introduced during error-prone repair of lesions involving single-strand DNA breaks (SSBs) or, more likely, double-strand DNA breaks (DSBs), as DSBs occur exclusively in genes that have the potentials to undergo SHM. In the human, such genes include Ig V, BCL6, and c-MYC. In these germline genes, DSBs are blunt. In rearranged Ig V, BCL6, and translocated c-MYC genes, blunt DSBs are processed to yield resected DNA ends. This process is dependent on the expression of activation-induced cytidine deaminase (AID), which is selectively expressed upon CD40-signaling in hypermutating B cells. CD40-induced and AID-dependent free 5′- and 3′-staggered DNA ends critically channel the repair of DSBs through the homologous recombination (HR) repair pathway. During HR, the modulation of critical translesion DNA polymerases, as signaled by cross-linking of the B cell receptor (BCR) for antigen, leads to the insertions of mismatches, i.e., mutations. The nature of DSBs, the possible roles of AID in the modification of DSBs and that of the translesion DNA polymerases ζ and ι in the subsequent repair process that lead to the insertions of mutations are discussed here within the context of an integrated model of SHM.
Keywords: DSBs, AID, error-prone DNA repair, SHM
SHM AND ANTIBODY AFFINITY MATURATION
Antibody diversity is generated through processes occurring at sequential B cell developmental stages: (i) the rearrangement of different germline V (variable), D (diversity), and J (joining) genes to yield an enormous variety of VDJ gene segments in the heavy (H) chain locus; (ii) the rearrangement of V and J genes in the κ and λ chain loci, also giving rise to enormous assortment of rearranged VκJκ or VλJλ segments; and finally (iii) the random pairing of rearranged VHDJH and VκJκ or VλJλ gene segments. The staggering number of VHDJH/VκJκ or VHDJH/VλJλ pairs possible underlies the high degree of diversity of the primary antibody repertoire, which consists of IgM antibodies with, in general, a low affinity for antigens. The impact of antigen on this primary antibody repertoire, expressed as diversity of the B cell receptor (BCR) repertoire, results in the production of antibodies of higher affinities and different classes, i.e., IgG, IgA, IgE. The shift to high affinity and secondary isotypes is underpinned by two critical genetic processes: Ig somatic hypermutation (SHM) and class switch DNA recombination (CSR) (Fig. 1). Both SHM and CSR occur in the specialized microenvironment of the germinal center (GC), and both contribute to the maturation of the antibody response, although in different ways. By diversifying the binding strength of the surface receptor for antigen, SHM provides the structural basis for clonal selection by antigen of high affinity mutants and the affinity maturation of the antibody response. By changing the constant (C) region of the Ig H chain with a downstream CH region, CSR changes the antibody effector features to suit them to the new biological functions required by later stages of the antibody response.
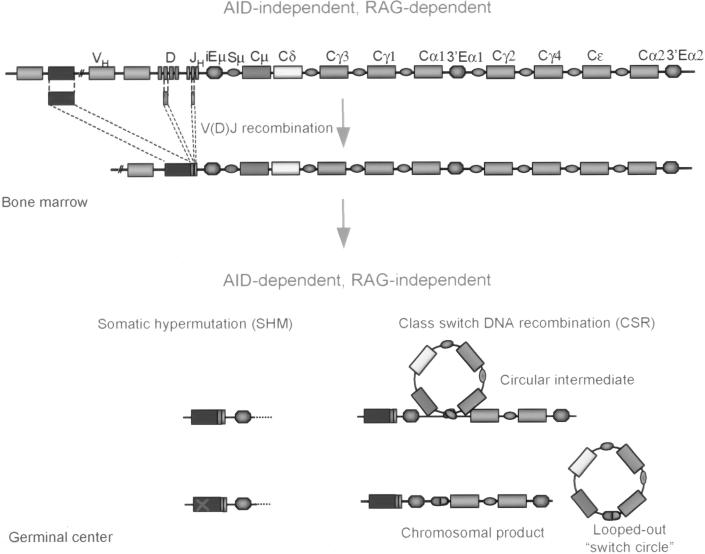
Fig. 1.
AID-independent and AID-dependent stages of B cell differentiation. Schematic diagram shows the Ig H chain locus. Rectangles and ovals represent V or C exons and switch regions. In the bone marrow, RAG-dependent V(D)J recombination selects one of each of the V, D, and J segments from respective gene pools and combines them into a single V(D)J exon (AID-independent and RAG-dependent stage of B cell differentiation). In peripheral lymphoid organs, the BCR is further diversified by SHM in an AID-dependent and RAG-independent fashion. SHM introduces mutations in the rearranged VDJ exon, yielding B cell submutants with diversified antigen specificities from which high affinity immunoglobulin producers are selected. CSR substitutes the upstream Cμ region with a downstream CH region by deletion of the DNA intervening between Sμ and the downstream S region 5′ of the target C region. Deleted DNA is released as a “switch circle.” Like SHM, CSR is AID-dependent and RAG-independent.
Somatic Ig V(D)J hypermutation is turned on at the centroblastic stage of GC, after mature B cells encounter antigen and in the presence of T cell help (1). At the centrocytic stage of GC, B clones expressing BCRs with higher antigen-binding affinity undergo antigen-driven positive selection and develop into memory B cells or antibody-secreting plasma cells, while autoreactive B cells or low affinity clones undergo negative selection through apoptosis (2), anergy (3), or receptor editing (4). Both in vivo (5) and in vitro (6–9) experiments have shown that SHM is induced in B cells that have received T cell help and sustained BCR cross-linking, consistent with the role of T cell and BCR engagement in GC formation. In human CL-01 cells (10), a monoclonal model of GC B lymphocyte differentiation that can be induced to switch to IgG, IgA, and IgE and effectively mutate the expressed VHDJH and VλJλ gene segments (11–15), it was shown that CD40 and CD80/CD86 coengagement by T cell CD40 ligand (CD40L) and CD28 is necessary in addition to BCR cross-linking for the induction of Ig SHM (8) (Fig. 2). The same stimuli also induce SHM in BCL6 (16–18), a proto-oncogene that is preferentially expressed in the GC. In contrast to SHM, induction of CSR requires only CD40 engagement by CD40L and exposure to cytokines (10, 19, 20). Although different in their induction requirements, both SHM and CSR are dependent on the activity of a recently characterized factor, a cytidine deaminase referred to activation-induced deaminase (AID) (21, 22). AID is not required for V(D)J rearrangement during B cell differentiation in the bone marrow, a process that is recombination-activating gene (RAG)-dependent, but plays a central role in the clonal expansion, selection and differentiation of B cells in the peripheral lymphoid organs. These processes are driven by antigen and are not dependent on RAG expression.
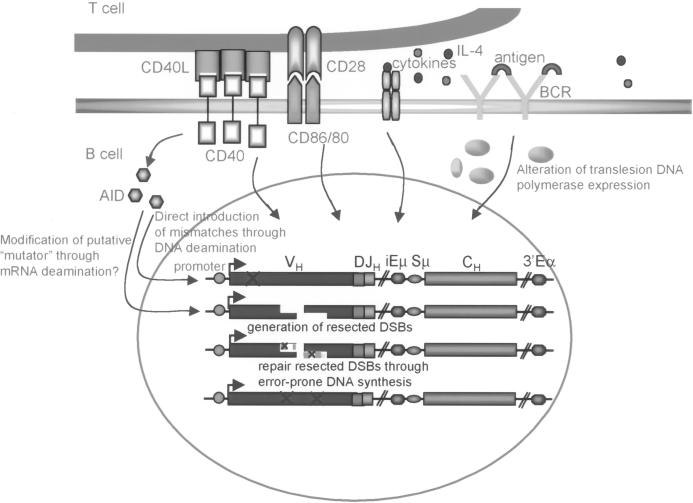
Fig. 2.
The requirement for induction for SHM. BCR cross-linking and T cell contact through CD40:CD40L and CD86/CD80:CD28 coengagement are required for induction of SHM. AID, which is upregulated by CD40-signaling, could directly introduce mutations by DNA deamination or could edit mRNA for an exonuclease and/or endonuclease to modify the blunt DSB into the resected DSB. These are crucial to initiate DNA repair process that leads to SHM. BCR engagement upregulates translesion DNA polymerase ζ, which, together with other error-prone DNA polymerases, such as polymerase ι, is recruited to the repair pathway and inserts mutations.
The features of SHM support the postulation of a sequential process of SHM which includes targeting of hypermutation region, introduction of DNA lesions and repair of such lesion by error-prone DNA synthesis, eventually resulting in the introduction of mismatches, i.e. mutations (23–25). Critical questions are: what is the nature of the DNA lesions, what is the role of AID, and what is the DNA repair process resolving those DNA lesions? Here, we will review address the function of AID in light of our most recent findings on the nature of the DSBs and the translesion DNA polymerases involved in the DNA repair process, thereby providing an integrated model of SHM.
MODALITIES AND REGULATION OF SHM
SHM displays a prevalence of point-mutations with only occasional insertions and deletions and a rate of 10−3 to 10−4 change/base/cell division, which is about one million fold higher than spontaneous somatic DNA mutation rate of the genome at large (26). It targets the sequence immediately downstream of the Ig V promoter, stretching 1–2 kb and including the V(D)J region, while sparing the constant region (27–29). It is linked to transcription initiation (30–32) and favors transitions over transversions with an intrinsic target preference for the mutational RGY(W)/WRC(Y) hotspot (R = A or G, Y = C or T, W = A or T) (33–35).
The targeting preference of SHM has evolved to favor the accumulation of mutations in the complementarity-determining region (CDR), the area that comes in close contact with antigen and dictates the specificity of the antibody binding site. The use of mutability indexes to measure the tendency to mutate and generate maps of the most and least mutable di- and trinucleotide sequences in antibody V genes has shown that the CDRs comprise codons that are targeted by SHM at a high frequency. In addition, these CDR codons are more inherently susceptible to replacement mutation than it would be expected for a random sequence. Conversely, the framework (FR) sequences are targeted by SHM at a lower frequency and comprise codons inherently less susceptible to replacement mutation than expected for a random nucleotide sequence. Thus, the somatic hypervariability of the CDRs relies on a dual process: the “preferred” targeting of the mutational RGY(W)/WRC(Y) hotspot and the higher frequency of mutations resulting in amino acid replacement due to inherent difference of codon composition of the CDRs as opposed to the FRs (36). This suggests that the primary sequences of Ig V genes have evolved phylogenetically in response to a selection positive pressure for replacement mutations applied to the CDRs and a negative pressure for replacement mutations applied to the FRs (36–38). The CDRs must be diversified in order to maximize contact with antigen, while the FRs must be preserved in order to provide a sound scaffolding for the antigen-binding site.
The targeting preference of SHM is not due solely to the primary sequence of the target DNA, as RGY(W) motifs exist outside V(D)J, BCL6, or c-MYC sequence, indicating the existence of a mechanism that targets the SHM machinery specifically to these DNA regions. Experiments in transgenic mice have shown that Ig cis-regulatory elements, mainly the κ intronic enhancer (iEκ) and the 3′ enhancer (3′Eα), may be important for the targeting SHM to the transgenic substrates (39–41), while the Ig V promoter can be replaced by other promoters (39). However, our recent findings in human B cells suggest a more complex regulation of SHM. A construct consisting of a rearranged human Ig VHDJH gene segment, the complete H chain, intronic enhancer (iEμ)-switch (S)μ sequence, the second exon of Cγ1 and the 3′Eα hs1,2 and hs3/4 elements was used to address the contribution of the cis-regulatory elements to SHM. Different iterations of this construct containing different cis-elements showed that the mutations accumulating in the construct were mainly transitions, preferentially targeted the dA/dT residue, without mutational RGY(W)/WRC(Y) hotspot preference, irrespective of the presence of either or both Ig H enhancers (42). This contrasts with the preferential dC/dG targeting of endogenous mutations in Ramos and other lymphoblastoid cell lines and implies that cis-regulatory elements other than the iEμ, 3′Eα hs12, or hs3/4 are required for the expression of the canonical modalities of Ig SHM.
DNA LESIONS AND SHM
Somatic mutations could be inserted during error-prone repair of DNA lesions involving single-strand breaks (SSBs) or DSBs, as first hypothesized by Brenner and Milstein in 1966 (43). They could also be introduced as direct single nucleotide substitutions through a mechanism of base modification without interfering with the sugar-phosphate backbone of the DNA double helix. As suggested by experiments in E. coli, in the chicken lymphoma B cell line DT40 and in mice, AID could directly deaminate dC into dU residues and give rise to U-G mispairs, followed by removal of uracil by uracil–DNA–glycosylase (UDG) and the creation of an abasic site. When DNA synthesis occurs over the abasic site, a mismatched nucleotide i.e., mutation, can be introduced (44–46). In spite of the possible role of direct DNA deamination and abasic DNA lesion in SHM, DNA breaks and particularly DSBs are more likely the lesions that initiate the error-prone DNA repair and eventually lead to the insertion of mismatches, i.e., mutations. Deletions/insertions/duplications account for only 6% of the mutations introduced into rearranged VH genes in human GC B cells, but possibly represent important remnants of the DSBs that underlie the SHM process, as they are absent in surface IgM+IgD+ B cells and frequently include embedded point-mutations (47, 48). Since the generation of deletions/insertions must be associated with DNA cleavage, the occurrence of deletions/insertions hints that DNA cleavage is the intermediate or the by-product of the SHM process. Deletions/insertions can result from the removal/addition of nucleotides from/to DNA ends. Duplications can result from the fill-in followed by DNA ligation of broken ends that occur at different sites on opposed DNA strands. The existence of DNA breaks over hypermutating Ig V region was first inferred in terminal deoxynucleotidyl transferase (TdT) transfected IgM+ Burkitt's lymphoma cell line Ramos. When Ramos, which constitutively hypermutates the rearranged Ig V(D)J genes in culture, was transfected with TdT, short nontemplated nucleotide sequences were inserted over hypermutating domains. After the indication of the presence of TdT-accessible DNA ends embedded in the mutational region (49), Papavasiliou and Schatz and Bross and colleagues directly associated DSBs with SHM (50, 51). Free phosphorylated double-strand DNA ends were revealed by specific ligation-mediated PCR (LM-PCR) in the rearranged Ig V(D)J region but not in the C region of hypermutating GC B cells or spontaneously hypermutating Ramos B cells.
Like somatic mutations, DSBs preferentially target the RGY(W) hotspot (Fig. 3) and their generation is coupled to transcription and enhancer-dependent. Predominant occurrence of DSBs in the S/G2 phase of the cell cycle has suggested repair of these DNA lesions through HR (50, 51). Our recent experiments have clearly shown that dsDNA breakage is a highly specific function that is inherent only to the genes that can undergo SHM. In the human, these are three of such genes: Ig V, BCL6, and c-MYC Genes that cannot undergo SHM, including Cμ, PAX5, PIM1, and alpha-fetoprotein (AFP), do not “break.” Ig V, BCL6, and c-MYC DSBs occur equally well in T cells and even outside the lymphoid context, i.e., in fibroblasts, endothelial cells and cervix cells, further emphasizing that dsDNA breakage is an exquisite property of the genes that have potential to undergo SHM. However, DNA breakage alone is not sufficient to drive SHM, as Ig V, BCL6, or c-MYC break but do not accumulate mutations in non-B cells (52).
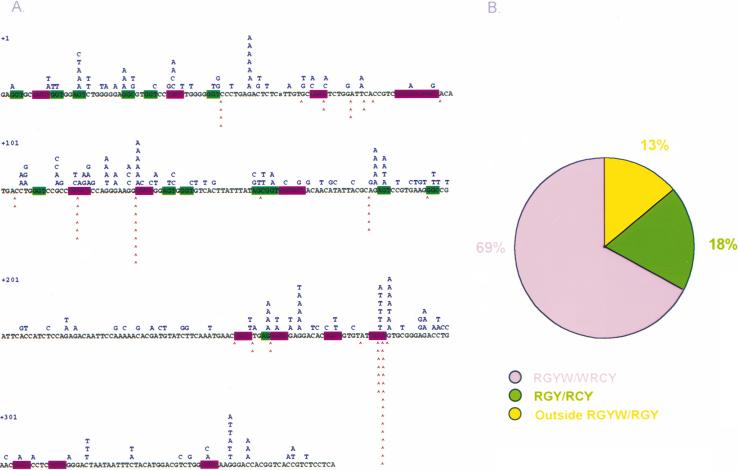
Fig. 3.
(A) Point-mutations and blunt DSBs target the same residues in the rearranged VHDJH gene segment expressed in human CL-01 B cells. Letters above the VHDJH sequence depict base changes; wedges below the VHDJH sequence indicate positions of DSBs. RGYW/WRCY are highlighted in purple and RGY/RCY in green. (B) Most DSBs in rearranged VHDJH genes target the mutational RGY(W)/WRC(Y) hotspot. DSBs within RGY(W)/WRC(Y) and those outside are 69, 18, and 13%, respectively of the total blunt DSBs.
The detection of DSBs by LM-PCR relies on the availability of 5′-phosphorylated blunt ends for DNA amplification, indicating that this is the nature of the DSBs characteristically found in genes that can undergo SHM. By treating genomic DNA with T4 DNA polymerase before LM-PCR linker ligation to trim back 3′-overhangs while filling in 3′-recessed ends, we have extended our analysis to the detection of resected DSBs, i.e., DSBs involving staggered 5′- and 3′-ends. The DSBs revealed by T4 polymerase pretreatment consist of both blunt and resected (total) DSBs and the level of resected DNA ends is deduced from the comparison of the amount of total DSBs with that of blunt DSBs. The application of this approach to the analysis of DNA from B cells at different stages of differentiation has shown that (i) resected DSBs are present only in rearranged Ig V genes of hypermutating GC B cells, but not in pre-GC or post-GC B cells; (ii) abundant blunt DSBs are present in nonhypermutating germline V genes but resected DSBs are present only in hypermutating rearranged V(D)J genes; (iii) blunt DSBs occur in c-MYC of both the germline and the translocated alleles in same B cell, but resected DSBs occur only in the translocated and hypermutating c-MYC allele (52, 53); and, finally, (iv) neither blunt nor resected DSBs occur in Cμ, PAX5, PIM1, AFP genes which cannot undergo SHM. Hence, resected DSBs are characteristic of hypermutating genes and likely constitute crucial intermediates in the SHM process (52).
The high relevance of DSBs to SHM is further emphasized by the demonstration that the DSBs detected by LM-PCR are not associated with replication stall or apoptotic DNA fragmentation, as enhanced B cell proliferation doesn't parallel increased DSBs in hypermutating genes and B cell apoptosis does not contribute to the overall level of detected Ig V DSBs (52). Therefore, the differential occurrence of DSBs at selected loci is distinct from the homogenous degradation of nuclear DNA, as occurring in apoptosis, a frequent event in the GC reaction (54). How the DNA breakage is introduced into Ig V, BCL6, and c-MYC loci remains to be determined, although it clear that RAG-1, which is needed to cleave DNA in VDJ recombination, is not involved in SHM and CSR (55), nor is Spo11, the generator of DSBs in meiotic recombination (56).
AID
AID is required for the two Ig DNA modification processes that are central to the maturation of the antibody response: SHM and CSR. AID was identified by subtraction of cDNAs from switch-induced and uninduced murine B lymphoma CH12F3-2 cells. It is a 198-amino acid protein that is homologous to the cytidine deaminase APOBEC-1, the catalytic subunit of the apolipoprotein B (apoB) mRNA-editing enzyme (57). AID-deficient mice have enlarged GC but defect in both CSR and SHM (21). In humans, AID mutations have been linked to autosomal recessive variant of hyper-IgM syndrome (HIGM2) (22). AID acts downstream of DSBs (58, 59). However, while AID is dispensable for the generation of blunt DSBs, it is critical for the generation of resected DSBs, which are intimately associated with SHM. As we have shown, AID expression is effectively induced in human B cells by CD40:CD40L engagement in a dose- and time-dependent fashion. In these B cells, overexpression of AID doesn't increase the level of blunt DSBs, but induces significantly high levels of resected-end DSBs (52). Further, expression of a dominant negative (DN) AID cDNA in Ramos B cells abrogates the generation of resected DSB and concomitant SHM in VHDJH, but doesn't affect the occurrence of blunt DSBs (52). How AID processes blunt DNA ends to yield resected ends is unclear. AID could generate 5′- and 3′-protruding ends by editing mRNA for an endonuclease that attacks blunt dsDNA termini, or could directly deaminate blunt DSBs. These two possibilities are not mutually exclusive, as recent data support both a mRNA editor and a direct DNA deaminator functions for AID (44–46, 60–62).
DSBS AND ERROR-PRONE DNA REPAIR
In mammalian cells, DSBs are repaired through two actively competing pathways. Opposing broken DNA ends can be rejoined with no or little regard for sequence homology by NHEJ or by a template-directed HR process. Papavasiliou and Schatz found that DSBs in hypermutating B cells are exclusively in S/G2 cell populations, and proposed a model in which mutations are introduced during DSB repair by HR through recruitment of error-prone DNA polymerases (50). Our recent findings have provided strong evidence for Ig VDJ DNA 3′-protruding end DSBs occurring in S and G2/M phase of hypermutating B cells. Since in HR the 3′-ended single-strand DNA tail is required for strand invasion of sister chromatids or homologues, the exclusive existence of Ig V genes or BCL6 3′-protruding ends in G2/M conforms with the role of HR in resolving such DNA ends. Further evidence that resected DNA ends are involved in directing repair process to HR has been provided by our recent experiment using a modified chromatin-immunoprecipitation (ChIP) assay to investigate binding of γ-H2AX, Nbs1, Mre11, Ku70/Ku86, Rad51, or Rad52 to nonhypermutating germline VH genes and hypermutating rearranged VHDJH genes. These experiments have shown that γ-H2AX, Nbs1, Mre11, and Ku70/Ku86 are bound to blunt ends of germline VH DNA in spontaneously hypermutating Ramos B cells, but Rad52 or Rad51 are bound only to resected DNA ends of rearranged VHDJH genes. Thus, blunt ends of nonhypermutating germline VH genes are generated in the absence of AID and recruit Ku70/Ku86 to then be religated through the NHEJ pathway (52), while resected DNA ends, which are generated through an AID-dependent process, recruit Rad52/Rad51, thereby directing the repair through HR repair pathway, in which mismatched nucleotides are introduced by recruitment of an error-prone DNA polymerase.
Because of the high frequency of base changes in SHM, the generation of a mismatched pair requires that a putative hypermutation DNA polymerase be capable of commonly inserting an incorrect base and also of extending from the newly generated mismatched terminus. Such putative SHM error-prone synthesis may occur outside of global replication and in the context of short patch error-prone repair (63). The unprecedented mutation frequency and the likely requirement mismatch extension is very difficult, if not impossible, to attain if the involved polymerases synthesize DNA with high fidelity and have proofreading capabilities. It is instead much more likely that the hypermutation polymerase is one (or several) of the growing family of DNA polymerases that have evolved to allow continued synthesis opposite DNA lesions that would otherwise stall a replication fork: the translesion synthesis polymerases. These DNA polymerases are in general error-prone and capable of extending, with various efficiencies, from a mismatched terminus (64–67).
Several studies have shown that the translesion DNA polymerase β, μ, λ, κ, and δ are not involved in SHM (68–71). Rather, the translesion DNA polymerases ι and ζ likely play a central role in SHM. Whereas DNA polymerase ι is highly error-prone when copying undamaged DNA, the DNA polymerase ζ displays a relatively high fidelity and is very efficient at extending from a mismatched terminus. These two DNA polymerases can act in concert, where polymerase ι inserts an incorrect base, and the ubiquitous polymerase ζ extends from the mismatched pair (72, 73), thus stabilizing the mutation. Given the role of DSBs in SHM, the finding in yeast that increased incidences of base-substitutions near DSBs are entirely dependent on DNA polymerase ζ (74) further support the role of this polymerase is SHM (75). Polymerase ζ plays a critical role in the physiological repair of DNA damaged by physical agents, such as UV irradiation. In this process, that is referred to as “damaged-induced DNA mutagenesis,” it favors the insertion of mismatches by allowing for highly efficient DNA strand synthesis past mispairs, as introduced opposite to lesions by other translesion polymerases. The synthesis of a mutated DNA strand represents the price the cell pays to preserve its ability to duplicate DNA and, therefore, multiply after exposure to a DNA-damaging agent. Significant evidence exists on the critical role of polymerase ζ in both Ig V(D)J and BCL6 SHM. As shown by Zan et al. (76), BCR cross-linking, which is necessary to induce SHM, signals the upregulation of translesion DNA polymerase ζ (Fig. 4). Blocking of the DNA polymerase ζ REV3 catalytic subunit by specific antisense oligonucleotides impairs Ig and BCL6 hypermutation without affecting cell cycle or viability (76). The involvement of the DNA polymerase ζ REV3 in SHM has been further implied by Diaz et al. (77), who generated transgenic mice that expressed antisense RNA to a portion of mouse REV3 to reduce the level of REV3 transcript. In these transgenic animals, the accumulation of somatic mutations in the VH genes of memory B cells was decreased and the generation of high affinity antibodies was delayed. Gene inactivation of polymerase ι in Burkitt's lymphoma cell line BL2 abolished inducible Ig SHM, but didn't affect the background mutation frequency, suggesting that inducible hypermutation in BL2 is dependent on polymerase ι (78), and further supporting the postulated role of this polymerase as “mismatch inserter.” After incorporation of one or two mismatched nucleotides by DNA polymerase ι, polymerase would ζ extend from such nucleotides to preserve the mutation(s). The role of another translesion DNA polymerase, polymerase η, in SHM is controversial. In patients with the Xeroderma pigmentosum variant (XP-V) disease, who are deficient in polymerase η, somatic mutations were found to be normal in frequency but including fewer changes at A and T and increased changes at G and C, suggesting that polymerase η is a dA/dT mutator (79–81). However, this is not supported by other findings of the intact overall dA/dT targets in nonproductively rearranged Ig VH genes in a well studied XP-V patient (82) and by the demonstration that polymerase η is profoundly downregulated by BCR cross-linking in actively hypermutating B cells (76). In fact, the downregulation of polymerase η is likely critical to allow SHM to occur, as the low processivity of this translesion polymerase allows for extrinsic exonucleases to proofread the mismatched nucleotides it inserts. This, together with the inability of polymerase η to elongate a DNA strand after incorporating the incorrect nucleotide opposite a lesion, allows this polymerase to perform error-free translesion synthesis of damaged DNA.
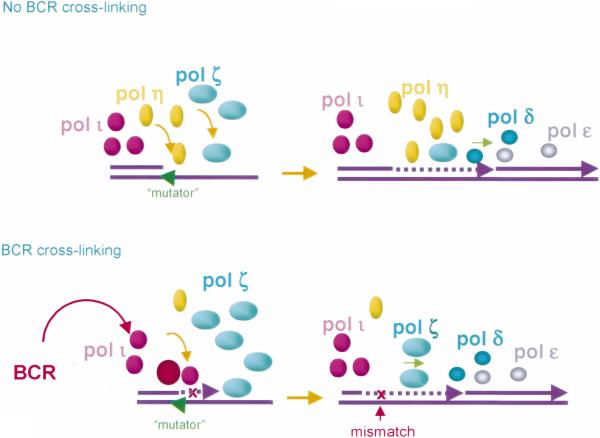
Fig. 4.
DNA polymerase ζ is regulated by BCR-signaling and plays a critical role in SHM, likely in concert with polymerase ι and concomitant with profound downregulation of polymerase η, as induced by BCR-signaling. Translesion DNA polymerase ι, polymerase η, and polymerase ζ are depicted in purple, yellow, and turquoise, respectively. The low processivity of polymerase η allows for extrinsic exonucleases to proofread the mismatched nucleotides it inserts. This together with the inability of polymerase η to elongate a DNA strand after incorporating the incorrect nucleotide opposite a lesion allows this polymerase to perform error-free translesion synthesis of damaged DNA. Polymerase ζ efficiently extends damaged DNA, in concert with the error-prone, low processivity polymerase ι, after the incorporation of one or two mismatched deoxynucleotides by this polymerase. Polymerase ι and polymerase ζ act sequentially: polymerase ι as a “mispair inserter” and polymerase ζ as “mispair extender,” and this lesion-bypass DNA synthesis is continued by polymerase δ and polymerase ε. BCR cross-linking also signals the upregulation of DNA polymerase ζ, thereby allowing for the mismatches inserted by polymerase ι to be efficiently extended, giving rise to a newly synthesized DNA strand containing point-mutations.
AN INTEGRATED MODEL FOR SHM
Analysis of the most recent findings on DNA lesions and error-prone repair allows us to integrate a model for SHM that satisfies the salient features of the process (Fig. 5). SHM would begin with the introduction of DSBs, mainly in the mutational RGWY hotspot. DSBs would then be subject to repair by NHEJ or HR. Blunt DNA ends would recruit the Ku70/Ku86 heterodimer and be repaired through NHEJ with no introduction of mismatches, i.e., mutations. As we show, Ku70/Ku86 as well as Mre11, Nbs1, and γ-H2AX bind to blunt DNA ends in both germline VH and rearranged VHDJH genes. In hypermutating B cells, resected DNA ends are characteristically generated in an AID-dependent fashion, as such or through modification of blunt DSBs. An error-prone DNA polymerase could introduce mismatches while filling in 3′-recessed termini possibly in G1, thereby allowing for ligation of the two free DNA ends by NHEJ. Accordingly, mutations have been suggested to be introduced at this stage of the cell cycle (83). Alternatively, 3′-single-strand DNA tails can recruit Rad52, which in turn mediates binding of Rad51, and direct the repair process toward HR. The activity of selected error-prone polymerases and the mispair extension ability of the translesion polymerase ζ, as modulated by BCR cross-linking, would then serve to introduce mutations at the sites of the DSBs. In the absence of BCR cross-linking, the resected DNA ends would be repaired in an error-free fashion through either HR or “fillin” of the DNA overhang and ligation through NHEJ. This implies that generation of 3′-protruding DNA ends does not necessarily lead to SHM unless error-prone DNA repair activity is also present. Our DSB-based SHM model explains the main features of SHM, including the gene targeting specificity, the intrinsic mutation hotspots, the requirement for AID, the transcription dependence and the occurrence of deletions/insertions (52). It implies three sequential steps, all of which necessary but none per se sufficient: (i) AID-independent occurrence of blunt DSBs in Ig V genes whether in germline or rearranged configuration; (ii) CD40-signaling induced and AID-dependent resection of blunt DSBs to generate 3′- and 5′-protruding ends in B cells undergoing cognate interaction with T cells; (iii) alteration of translesion DNA polymerase expression upon BCR cross-linking and subsequent error-prone repair of DSBs through HR (52).
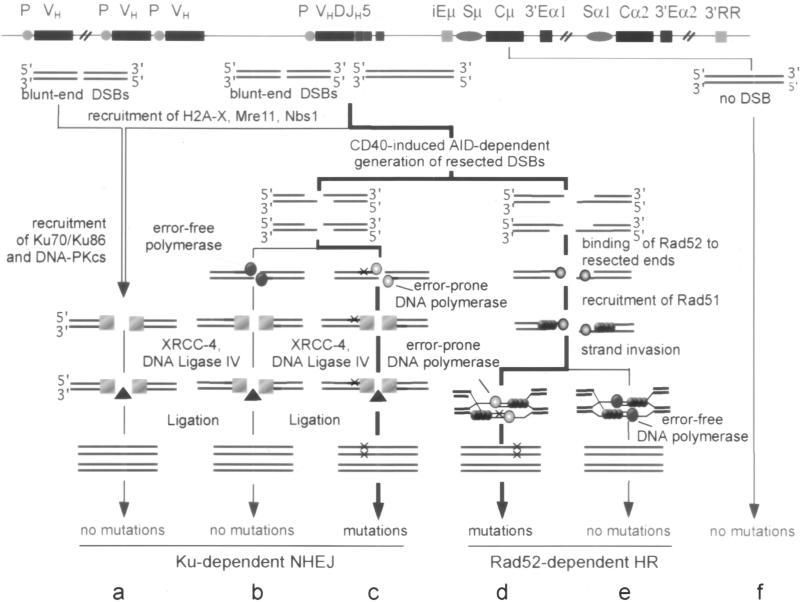
Fig. 5.
An integrated model for SHM. DSBs target V(D)J, BCL6, and c-MYC genes and initiate the process that leads to the introduction of somatic mutations through error-prone repair. In germline V and c-MYC genes of hypermutating or nonhyper-mutating B cells and in BCL6 of nonhypermutating B cells, DSBs are blunt and are repaired through NHEJ in a faultless fashion (a). In hypermutating rearranged V(D)J genes or germline BCL6 or c-MYC translocated into the Ig locus, blunt DSBs are either similarly resolved (a) or are modified in a CD40-induced and AID-dependent fashion to yield 5′- and 3′-protruding DNA ends. Other resected DSBs may also be generated as such. Generation of resected DSBs as such or through modification of blunt DSBs would occur through intervention of an AID-edited endonuclease and/or exonuclease or direct AID-dependent deamination. 3′-Recessed termini may be filled in by an error-free DNA polymerase (in absence of BCR cross-linking) (b) or by an error-prone DNA polymerase (upon BCR cross-linking) with introduction of mismatches (c), and then resolved by NHEJ. 3′-Protruding ends can invade sister chromatid strands and initiate a HR process which, upon BCR cross-linking, involves the activity of the translesion DNA polymerase ζ. Polymerase ζ, possibly in concern with polymerase ι, extends DNA strands past mispair, eventually leading to SHM (d). In the absence of BCR cross-linking, HR proceeds in an error-free fashion (e). DSBs do not target C regions and no mutations can be introduced in this DNA (f)
CONCLUSIONS AND PERSPECTIVES
SHM has long been a black box for immunologists and molecular biologists for over 30 years. Recent findings have defined the roles of AID, DSBs, and error-prone DNA polymerases in SHM. In particular, it has become clear that the DNA lesion underlying SHM contains many of DSBs with staggered ends and such staggered free DNA ends emerge through processing of the blunt DSBs through direct or indirect activity of AID (52). It has also become clear that the B cell utilizes the very physiological repair pathway which is used to repair damaged DNA for the introduction of somatic mutations, as reflected in the central role of DNA polymerase ζ in both damage-induced mutagenesis and SHM.
The issues that still need to be addressed in SHM are (i) the precise role of different cis-elements and trans-factors in targeting SHM to Ig or BCL6 loci; (ii) the nature of the nuclease(s) that effect blunt DSBs in these genes and, conversely, the inherent gene features that determine the recognition by such nuclease(s); (iii) the identification of the specific target of AID, i.e., DNA or pre-mRNA or both; (iv) the nature of the AID targeting process that transforms blunt DNA ends into resected-ends in rearranged Ig V, BCL6, and translocated c-MYC genes; and, finally, (v) the precise mechanisms by which the HR repair pathway, which is usually accurate and involves high-fidelity DNA polymerases, is geared in SHM to insert mismatches by recruiting error-prone DNA polymerases. The increasingly detailed knowledge of the human Ig locus, inherent cis-elements and relevant transcription factors together with the recent availability of useful in vitro models of B cell differentiation should help make significant progress in the elucidation of the above and still unaddressed issues in SHM.
ACKNOWLEDGMENTS
This work was supported by N. I. H. grants AR 40908, AI 45011, AG 13910, and AI 0762 to P.C.
Abbreviations
- AID
activation induced deaminase
- BCR
B cell receptor
- CSR
class switch DNA recombination
- C
constant region
- CDR
complementarity-determining region
- ChIP
chromatin-immunoprecipitation
- DN
dominant negative
- DSBs
double-strand DNA breaks
- AFP
alpha-fetoprotein
- FR
framework
- GC
germinal center
- HR
homologous recombination
- iEμ
intronic enhancer
- Ig
immunoglobulin
- LM-PCR
ligation-mediated PCR
- NHEJ
nonhomologous end joining
- RAG-1
recombination-activating gene-1
- SHM
somatic hypermutation
- SSB
single-strand DNA breaks
- TdT
terminal deoxynucleotidyl transferase
- UDG
uracil DNA glycosylase
- V(D)J
variable (diversity) joining
- XP-V
Xeroderma pigmentosum variant
REFERENCES
- 1.Pascual V, Liu YL, Magalski A, de Bouteiller O, Banchereau J, Capra JD. Analysis of somatic mutation in five B cell subsets of human tonsil. J Exp Med. 1994;180:329–339. doi: 10.1084/jem.180.1.329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Pulendran B, Kannourakis G, Nouri S, Smith KG, Nossal GJ. Soluble antigen can cause enhanced apoptosis of germinal-centre B cells. Nature. 1995;375:331–334. doi: 10.1038/375331a0. [DOI] [PubMed] [Google Scholar]
- 3.Cornall RJ, Goodnow CC, Cyster JG. The regulation of self-reactive B cells. Curr Opin Immunol. 1995;7:804–811. doi: 10.1016/0952-7915(95)80052-2. [DOI] [PubMed] [Google Scholar]
- 4.Hertz M, Nemazee D. Receptor editing and commitment in B lymphocytes. Curr Opin Immunol. 1998;10:208–213. doi: 10.1016/s0952-7915(98)80250-1. [DOI] [PubMed] [Google Scholar]
- 5.Miller C, Stedra J, Kelsoe G, Cerny J. Facultative role of germinal centers and T cells in the somatic diversification of IgVH genes. J Exp Med. 1995;181:1319–1331. doi: 10.1084/jem.181.4.1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Razanajaona D, Denepoux S, Blanchard D, de Bouteiller O, Liu YJ, Banchereau J, Lebecque S. In vitro triggering of somatic mutation in human naive B cells. J Immunol. 1997;159:3347–3353. [PubMed] [Google Scholar]
- 7.Denepoux S, Razanajaona D, Blanchard D, Meffre G, Capra JD, Banchereau J, Lebecque S. Induction of somatic mutation in a human B cell line in vitro. Immunity. 1997;6:35–46. doi: 10.1016/s1074-7613(00)80240-x. [DOI] [PubMed] [Google Scholar]
- 8.Zan H, Cerutti A, Dramitinos P, Schaffer A, Li Z, Casali P. Induction of Ig somatic hypermutation and class switching in a human monoclonal IgM+ IgD+ B cell line in vitro: Definition of the requirements and modalities of hypermutation. J Immunol. 1999;162:3437–3447. [PMC free article] [PubMed] [Google Scholar]
- 9.Decker DJ, Linton PJ, Zaharevitz S, Biery M, Gingeras TR, Klinman NR. Defining subsets of naive and memory B cells based on the ability of their progeny to somatically mutate in vitro. Immunity. 1995;2:195–203. doi: 10.1016/s1074-7613(95)80092-1. [DOI] [PubMed] [Google Scholar]
- 10.Cerutti A, Zan H, Schaffer A, Bergsagel L, Harindranath N, Max EE, Casali P. CD40 ligand and appropriate cytokines induce switching to IgG, IgA, and IgE and coordinated germinal center and plasmacytoid phenotypic differentiation in a human monoclonal IgM+IgD+B cell line. J Immunol. 1998;160:2145–2157. [PMC free article] [PubMed] [Google Scholar]
- 11.Zan H, Cerutti A, Dramitinos P, Schaffer A, Casali P. CD40 engagement triggers switching to IgA1 and IgA2 in human B cells through induction of endogenous TGF-β: Evidence for TGF-β but not IL-10-dependent direct S μ->S γ and sequential S μ->S γ , S γ -> S α DNA recombination. J Immunol. 1998;161:5217–5225. [PMC free article] [PubMed] [Google Scholar]
- 12.Cerutti A, Schaffer A, Shah S, Zan H, Liou HC, Goodwin RG, Casali P. CD30 is a CD40-inducible molecule that negatively regulates CD40-mediated immunoglobulin class switching in non-antigen-selected human B cells. Immunity. 1998;9:247–256. doi: 10.1016/s1074-7613(00)80607-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Schaffer A, Cerutti A, Shah S, Zan H, Casali P. The evolutionarily conserved sequence upstream of the human Ig heavy chain S γ 3 region is an inducible promoter: Synergistic activation by CD40 ligand and IL-4 via cooperative NF-κ B and STAT-6 binding sites. J Immunol. 1999;162:5327–5336. [PubMed] [Google Scholar]
- 14.Schaffer A, Kim EC, Wu X, Zan H, Testoni L, Salamon S, Cerutti A, Casali P. Selective inhibition of class switching to IgG and IgE by recruitment of the HoxC4 and Oct-1 homeodomain proteins and Ku70/Ku86 to newly identified ATTT cis-elements. J Biol Chem. doi: 10.1074/jbc.M212952200. in press. [DOI] [PubMed] [Google Scholar]
- 15.Cerutti A, Schaffer A, Goodwin RG, Shah S, Zan H, Ely S, Casali P. Engagement of CD153 (CD30 ligand) by CD30+ T cells inhibits class switch DNA recombination and antibody production in human IgD+ IgM+B cells. J Immunol. 2000;165:786–794. doi: 10.4049/jimmunol.165.2.786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zan H, Li Z, Yamaji K, Dramitinos P, Cerutti A, Casali P. B cell receptor engagement and T cell contact induce Bcl-6 somatic hy-permutation in human B cells: Identity with Ig hypermutation. J Immunol. 2000;165:830–839. doi: 10.4049/jimmunol.165.2.830. [DOI] [PubMed] [Google Scholar]
- 17.Shen HM, Peters A, Baron B, Zhu X, Storb U. Mutation of BCL-6 gene in normal B cells by the process of somatic hypermutation of Ig genes. Science. 1998;280:1750–1752. doi: 10.1126/science.280.5370.1750. [DOI] [PubMed] [Google Scholar]
- 18.Migliazza A, Martinotti S, Chen W, Fusco C, Ye BH, Knowles DM, Offit K, Chaganti RS, Dalla-Favera R. Frequent somatic hypermutation of the 5′ noncoding region of the BCL-6 gene in B-cell lymphoma. Proc Natl Acad Sci USA. 1995;92:12520–12524. doi: 10.1073/pnas.92.26.12520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fujieda S, Zhang K, Saxon A. IL-4 plus CD40 monoclonal antibody induces human B cells γ subclass-specific isotype switch: Switching to γ 1, γ 3, and γ 4, but not gamma 2. J Immunol. 1995;155:2318–2328. [PubMed] [Google Scholar]
- 20.Malisan F, Briere F, Bridon JM, Harindranath N, Mills FC, Max EE, Banchereau J, Martinez-Valdez H. Interleukin-10 induces immunoglobulin G isotype switch recombination in human CD40-activated naive B lymphocytes. J. Exp Med. 1996;183:937–947. doi: 10.1084/jem.183.3.937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Muramatsu M, Kinoshita K, Fagarasan S, Yamada S, Shinkai Y, Honjo T. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell. 2000;102:553–563. doi: 10.1016/s0092-8674(00)00078-7. [DOI] [PubMed] [Google Scholar]
- 22.Revy P, Muto T, Levy Y, Geissmann F, Plebani A, Sanal O, Catalan N, Forveille M, Dufourcq-Labelouse R, Gennery A, Tezcan I, Ersoy F, Kayserili H, Ugazio AG, Brousse N, Muramatsu M, Notarangelo LD, Kinoshita K, Honjo T, Fischer A, Durandy A. Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the Hyper IgM syndrome (HIGM2). Cell. 2000;102:565–575. doi: 10.1016/s0092-8674(00)00079-9. [DOI] [PubMed] [Google Scholar]
- 23.Kong Q, Harris RS, Maizels N. Recombination-based mechanisms for somatic hypermutation. Immunol Rev. 1998;162:67–76. doi: 10.1111/j.1600-065x.1998.tb01430.x. [DOI] [PubMed] [Google Scholar]
- 24.Papavasiliou FN, Schatz DG. Somatic hypermutation of immunoglobulin genes: Merging mechanisms for genetic diversity. Cell. 2002;109(Suppl):S35–S44. doi: 10.1016/s0092-8674(02)00706-7. [DOI] [PubMed] [Google Scholar]
- 25.Diaz M, Casali P. Somatic immunoglobulin hypermutation. Curr Opin Immunol. 2002;14:235–240. doi: 10.1016/s0952-7915(02)00327-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Levy NS, Malipiero UV, Lebecque SG, Gearhart PJ. Early onset of somatic mutation in immunoglobulin VH genes during the primary immune response. J Exp Med. 1989;169:2007–2019. doi: 10.1084/jem.169.6.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lebecque SG, Gearhart PJ. Boundaries of somatic mutation in rearranged immunoglobulin genes: 5′ boundary is near the promoter, and 3′ boundary is approximately 1 kb from V(D)J gene. J Exp Med. 1990;172:1717–1727. doi: 10.1084/jem.172.6.1717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Rogerson BJ. Mapping the upstream boundary of somatic mutations in rearranged immunoglobulin transgenes and endogenous genes. Mol Immunol. 1994;31:83–98. doi: 10.1016/0161-5890(94)90081-7. [DOI] [PubMed] [Google Scholar]
- 29.Rada C, Gonzalez-Fernandez A, Jarvis JM, Milstein C. The 5′ boundary of somatic hypermutation in a V κ gene is in the leader intron. Eur J Immunol. 1994;24:1453–1457. doi: 10.1002/eji.1830240632. [DOI] [PubMed] [Google Scholar]
- 30.Peters A, Storb U. Somatic hypermutation of immunoglobulin genes is linked to transcription initiation. Immunity. 1996;4:57–65. doi: 10.1016/s1074-7613(00)80298-8. [DOI] [PubMed] [Google Scholar]
- 31.Fukita Y, Jacobs H, Rajewsky K. Somatic hypermutation in the heavy chain locus correlates with transcription. Immunity. 1998;9:105–114. doi: 10.1016/s1074-7613(00)80592-0. [DOI] [PubMed] [Google Scholar]
- 32.Reynaud CA, Frey S, Aoufouchi S, Faili A, Bertocci B, Dahan A, Flatter E, Delbos F, Storck S, Zober C, Weill JC. Transcription, β-like DNA polymerases and hypermutation. Philos Trans R Soc Lond B Biol Sci. 2001;356:91–97. doi: 10.1098/rstb.2000.0753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Rogozin IB, Kolchanov NA. Somatic hypermutagenesis in immunoglobulin genes. II. Influence of neighbouring base sequences on mutagenesis. Biochim Biophys Acta. 1992;1171:11–18. doi: 10.1016/0167-4781(92)90134-l. [DOI] [PubMed] [Google Scholar]
- 34.Foster SJ, Dorner T, Lipsky PE. Somatic hypermutation of VκJκ rearrangements: Targeting of RGYW motifs on both DNA strands and preferential selection of mutated codons within RGYW motifs. Eur J Immunol. 1999;29:4011–4021. doi: 10.1002/(SICI)1521-4141(199912)29:12<4011::AID-IMMU4011>3.0.CO;2-W. [DOI] [PubMed] [Google Scholar]
- 35.Wagner SD, Milstein C, Neuberger MS. Codon bias targets mutation. Nature. 1995;376:732. doi: 10.1038/376732a0. [DOI] [PubMed] [Google Scholar]
- 36.Chang B, Casali P. The CDR1 sequences of a major proportion of human germline Ig VH genes are inherently susceptible to amino acid replacement. Immunol Today. 1994;15:367–373. doi: 10.1016/0167-5699(94)90175-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Dorner T, Brezinschek HP, Brezinschek RI, Foster SJ, Domiati-Saad R, Lipsky PE. Analysis of the frequency and pattern of somatic mutations within nonproductively rearranged human variable heavy chain genes. J Immunol. 1997;158:2779–2789. [PubMed] [Google Scholar]
- 38.Shapiro GS, Aviszus K, Murphy J, Wysocki LJ. Evolution of Ig DNA sequence to target specific base positions within codons for somatic hypermutation. J Immunol. 2002;168:2302–2306. doi: 10.4049/jimmunol.168.5.2302. [DOI] [PubMed] [Google Scholar]
- 39.Betz AG, Milstein C, Gonzalez-Fernandez A, Pannell R, Larson T, Neuberger MS. Elements regulating somatic hypermutation of an immunoglobulin κ gene: Critical role for the intron enhancer/matrix attachment region. Cell. 1994;77:239–248. doi: 10.1016/0092-8674(94)90316-6. [DOI] [PubMed] [Google Scholar]
- 40.Goyenechea B, Klix N, Yelamos J, Williams GT, Riddell A, Neuberger MS, Milstein C. Cells strongly expressing Ig(κ) trans-genes show clonal recruitment of hypermutation: A role for both MAR and the enhancers. EMBO J. 1997;16:3987–3994. doi: 10.1093/emboj/16.13.3987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Klix N, Jolly CJ, Davies SL, Bruggemann M, Williams GT, Neuberger MS. Multiple sequences from downstream of the J κ cluster can combine to recruit somatic hypermutation to a heterologous, upstream mutation domain. Eur J Immunol. 1998;28:317–326. doi: 10.1002/(SICI)1521-4141(199801)28:01<317::AID-IMMU317>3.0.CO;2-S. [DOI] [PubMed] [Google Scholar]
- 42.Komori A, Wu X, Kim E, Zan H, Casali P. Dispensability of the Ig heavy chain iEm and 3′Ea enhancers and preferential A/T targeting in the hypermutation of a human VHDJH-Cg1 construct. submitted [Google Scholar]
- 43.Brenner S, Milstein C. Origin of antibody variation. Nature. 1966;211:242–243. [PubMed] [Google Scholar]
- 44.Petersen-Mahrt SK, Harris RS, Neuberger MS. AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature. 2002;418:99–103. doi: 10.1038/nature00862. [DOI] [PubMed] [Google Scholar]
- 45.Di Noia J, Neuberger MS. Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature. 2002;419:43–48. doi: 10.1038/nature00981. [DOI] [PubMed] [Google Scholar]
- 46.Rada C, Williams GT, Nilsen H, Barnes DE, Lindhal T, Neuberger MS. Immunoglobulin isotype switching is inhibited and somatic hy-permutation perturbed in UNG-deficient mice. Curr Biol. 2002;12:1748–1755. doi: 10.1016/s0960-9822(02)01215-0. [DOI] [PubMed] [Google Scholar]
- 47.Goossens T, Klein U, Kuppers R. Frequent occurrence of deletions and duplications during somatic hypermutation: Implications for oncogene translocations and heavy chain disease. Proc Natl Acad Sci USA. 1998;95:2463–2468. doi: 10.1073/pnas.95.5.2463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wilson PC, de Bouteiller O, Liu YJ, Potter K, Banchereau J, Capra JD, Pascual V. Somatic hypermutation introduces insertions and deletions into immunoglobulin V genes. J Exp Med. 1998;187:59–70. doi: 10.1084/jem.187.1.59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Sale JE, Neuberger MS. TdT-accessible breaks are scattered over the immunoglobulin V domain in a constitutively hypermutating B cell line. Immunity. 1998;9:859–869. doi: 10.1016/s1074-7613(00)80651-2. [DOI] [PubMed] [Google Scholar]
- 50.Papavasiliou FN, Schatz DG. Cell-cycle-regulated DNA double-stranded breaks in somatic hypermutation of immunoglobulin genes. Nature. 2000;408:216–221. doi: 10.1038/35041599. [DOI] [PubMed] [Google Scholar]
- 51.Bross L, Fukita Y, McBlane F, Demolliere C, Rajewsky K, Jacobs H. DNA double-strand breaks in immunoglobulin genes undergoing somatic hypermutation. Immunity. 2000;13:589–597. doi: 10.1016/s1074-7613(00)00059-5. [DOI] [PubMed] [Google Scholar]
- 52.Zan H, Wu X, Komori A, Holloman WK, Casali P. AID-dependent generation of resected double-strand DNA breaks and recruitment of Rad52/Rad51 in somatic hypermutation. Immunity. doi: 10.1016/s1074-7613(03)00151-1. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Bemark M, Neuberger MS. The c-MYC allele that is translocated into the Ig H locus undergoes constitutive hypermutation in a Burkitt's lymphoma line. Oncogene. 2000;19:3404–3410. doi: 10.1038/sj.onc.1203686. [DOI] [PubMed] [Google Scholar]
- 54.MacLennan IC. Germinal centers. Annu Rev Immunol. 1994;12:117–139. doi: 10.1146/annurev.iy.12.040194.001001. [DOI] [PubMed] [Google Scholar]
- 55.Bemark M, Sale J, Kim HJ, Berek C, Cosgrove RA, Neuberger MS. Somatic hypermutation in the absence of DNA-dependent protein kinase catalytic subunit (DNA-PK(cs)) or recombination-activating gene (RAG)1 activity. J Exp Med. 2000;192:1509–1514. doi: 10.1084/jem.192.10.1509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Klein U, Esposito G, Baudat F, Keeney S, Jasin M. Mice deficient for the type II topoisomerase-like DNA transesterase Spo11 show normal immunoglobulin somatic hypermutation and class switching. Eur J Immunol. 2002;32:316–321. doi: 10.1002/1521-4141(200202)32:2<316::AID-IMMU316>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- 57.Muramatsu M, Sankaranand VS, Anant S, Sugai M, Kinoshita K, Davidson NO, Honjo T. Specific expression of activation-induced cytidine deaminase (AID), a novel member of the RNA-editing deaminase family in germinal center B cells. J Biol Chem. 1999;274:18470–18476. doi: 10.1074/jbc.274.26.18470. [DOI] [PubMed] [Google Scholar]
- 58.Papavasiliou FN, Schatz DG. The activation-induced deaminase functions in a postcleavage step of the somatic hypermutation process. J Exp Med. 2002;195:1193–1198. doi: 10.1084/jem.20011858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Bross L, Muramatsu M, Kinoshita K, Honjo T, Jacobs H. DNA double-strand breaks: Prior to but not sufficient in targeting hyper-mutation. J Exp Med. 2002;195:1187–1192. doi: 10.1084/jem.20011749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Jacobs H, Bross L. Towards an understanding of somatic hypermutation. Curr Opin Immunal. 2001;13:208–218. doi: 10.1016/s0952-7915(00)00206-5. [DOI] [PubMed] [Google Scholar]
- 61.Doi T, Kinoshita K, Ikegawa M, Muramatsu M, Honjo T. Inaugural Article: De novo protein synthesis is required for the activation-induced cytidine deaminase function in class-switch recombination. Proc Natl Acad Sci USA. 2003;100:2634–2638. doi: 10.1073/pnas.0437710100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bransteitter R, Pham P, Scharff MD, Goodman MF. Activation-induced cytidine deaminase deaminates deoxycytidine on single-stranded DNA but requires the action of RNase. Proc Natl Acad Sci USA. 2003;100:4102–4107. doi: 10.1073/pnas.0730835100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bertocci B, Quint L, Delbos F, Garcia C, Reynaud CA, Weill JC. Probing immunoglobulin gene hypermutation with microsatellites suggests a nonreplicative short patch DNA synthesis process. Immunity. 1998;9:257–265. doi: 10.1016/s1074-7613(00)80608-1. [DOI] [PubMed] [Google Scholar]
- 64.Lawrence CW, Maher VM. Mutagenesis in eukaryotes dependent on DNA polymerase zetaand Rev1p. Philos Trans R Soc London B Biol Sci. 2001;356:41–46. doi: 10.1098/rstb.2000.0746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Matsuda T, Bebenek K, Masutani C, Rogozin IB, Hanaoka F, Kunkel TA. Error rate and specificity of human and murine DNA polymerase eta. J Mol Biol. 2001;312:335–346. doi: 10.1006/jmbi.2001.4937. [DOI] [PubMed] [Google Scholar]
- 66.McDonald JP, Tissier A, Frank EG, Iwai S, Hanaoka F, Woodgate R. DNA polymerase iota and related rad30-like enzymes. Philos Trans R Soc Lond B Biol Sci. 2001;356:53–60. doi: 10.1098/rstb.2000.0748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Matsuda T, Bebenek K, Masutani C, Hanaoka F, Kunkel TA. Low fidelity DNA synthesis by human DNA polymerase-η. Nature. 2000;404:1011–1013. doi: 10.1038/35010014. [DOI] [PubMed] [Google Scholar]
- 68.Bertocci B, De Smet A, Flatter E, Dahan A, Bories JC, Landreau C, Weill JC, Reynaud CA. Cutting edge: DNA polymerases μ and λ are dispensable for Ig gene hypermutation. J Immunol. 2002;168:3702–3706. doi: 10.4049/jimmunol.168.8.3702. [DOI] [PubMed] [Google Scholar]
- 69.Esposito G, Texido G, Betz UA, Gu H, Muller W, Klein U, Rajewsky K. Mice reconstituted with DNA polymerase β-deficient fetal liver cells are able to mount a T cell-dependent immune response and mutate their Ig genes normally. Proc Natl Acad Sci USA. 2000;97:1166–1171. doi: 10.1073/pnas.97.3.1166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Schenten D, Gerlach VL, Guo C, Velasco-Miguel S, Hladik CL, White CL, Friedberg EC, Rajewsky K, Esposito G. DNA polymerase κ deficiency does not affect somatic hypermutation in mice. Eur J Immunol. 2002;32:3152–3160. doi: 10.1002/1521-4141(200211)32:11<3152::AID-IMMU3152>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 71.Longacre A, Sun T, Goldsby RE, Preston BD, Storb U. Ig gene somatic hypermutation in mice defective for DNA polymerase ± proofreading. Int Immunol. 2003;15:477–481. doi: 10.1093/intimm/dxg047. [DOI] [PubMed] [Google Scholar]
- 72.Yuan F, Zhang Y, Rajpal DK, Wu X, Guo D, Wang W, Taylor JS, Wang Z. Specificity of DNA lesion bypass by the yeast DNA polymerase η. J Biol Chem. 2001;275:8233–8293. doi: 10.1074/jbc.275.11.8233. [DOI] [PubMed] [Google Scholar]
- 73.Johnson RE, Washington MT, Haracska L, Prakash S, Prakash L. Eukaryotic polymerases ι and ζ act sequentially to bypass DNA lesions. Nature. 2000;406:1015–1019. doi: 10.1038/35023030. [DOI] [PubMed] [Google Scholar]
- 74.Holbeck SL, Strathern JN. A role for REV3 in mutagenesis during double-strand break repair in Saccharomyces cerevisiae. Genetics. 1997;147:1017–1024. doi: 10.1093/genetics/147.3.1017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Diaz M, Flajnik MF. Evolution of somatic hypermutation and gene conversion in adaptive immunity. Immunol Rev. 1998;162:13–24. doi: 10.1111/j.1600-065x.1998.tb01425.x. [DOI] [PubMed] [Google Scholar]
- 76.Zan H, Komori A, Li Z, Cerutti A, Schaffer A, Flajnik MF, Diaz M, Casali P. The translesion DNA polymerase ζ plays a major role in Ig and bcl-6 somatic hypermutation. Immunity. 2001;14:634–653. doi: 10.1016/s1074-7613(01)00142-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Diaz M, Verkoczy LK, Flajnik MF, Kliman NR. Decreased frequency of somatic hypermutation and impaired affinity maturation but intact germinal center formation in nice expressing antisense RNA to DNA polymerase ζ. J Immunol. 2001;167:327–335. doi: 10.4049/jimmunol.167.1.327. [DOI] [PubMed] [Google Scholar]
- 78.Faili A, Aoufouchi S, Flatter E, Gueranger Q, Reynaud CA, Weill JC. Induction of sonatic hypermutation in immunoglobulin genes is dependent on DNA polymerase ι. Nature. 2002;419:944–947. doi: 10.1038/nature01117. [DOI] [PubMed] [Google Scholar]
- 79.Zeng X, Winter DB, Kasmer C, Kraemer KH, Lehmann AR, Gearhart PJ. DNA polymerase η is an A-T mutator in somatic hyper-mutation of immunoglobulin variable genes. Nat Immunol. 2001;2:537–541. doi: 10.1038/88740. [DOI] [PubMed] [Google Scholar]
- 80.Rogozin IB, Pavlov YI, Bebenek K, Matsuda T, Kunkel TA. Somatic mutation hotspots correlate with DNA polymerase η error spectrum. Nat Immunol. 2001;2:530–536. doi: 10.1038/88732. [DOI] [PubMed] [Google Scholar]
- 81.Pavlov YI, Rogozin IB, Galkin AP, Aksenova AY, Hanaoka F, Rada C, Kunkel TA. Correlation of somatic hypermutation specificity and A-T base pair substitution errors by DNA polymerase η during copying of a mouse immunoglobulin κ light chain transgene. Proc Natl Acad Sci USA. 2002;99:9954–9959. doi: 10.1073/pnas.152126799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Yavuz S, Yavuz AS, Kraemer KH, Lipsky PE. The role of polymerase η in somatic hypermutation determined by analysis of mutations in a patient with xeroderma pigmentosum variant. J Immunol. 2002;169:3825–3830. doi: 10.4049/jimmunol.169.7.3825. [DOI] [PubMed] [Google Scholar]
- 83.Faili A, Aoufouchi S, Gueranger Q, Zober C, Leon A, Bertocci B, Weill JC, Reynaud CA. AID-dependent somatic hypermutation occurs as a DNA single-strand event in the BL2 cell line. Nat Immunol. 2002;3:815–821. doi: 10.1038/ni826. [DOI] [PubMed] [Google Scholar]