Abstract
The protein encoded by the archaeal intron in the 23S rRNA gene of the hyperthermophile Desulfurococcus mobilis is a double-strand DNase that, like group I intron homing endonucleases, is capable of cleaving an intronless allele of the gene. This enzyme, I-Dmo I, is unusual among the intron endonucleases in that it is thermostable and is expressed only from linear and cyclized intron species and not from the precursor RNA. However, in analogy to its eukaryotic counterparts, but unlike the bacteriophage enzymes, I-Dmo I makes a staggered double-strand cut that generates 4-nt 3' extensions. Additionally, although the archaeal and group I introns have entirely different structural properties and splicing pathways, I-Dmo I shares sequence similarity, in the form of the LAGLI-DADG motif, with group I intron endonucleases of eukaryotes. These observations support the independent evolutionary origin of endonucleases and intron core elements and are consistent with the invasive potential of endonuclease genes.
Full text
PDF
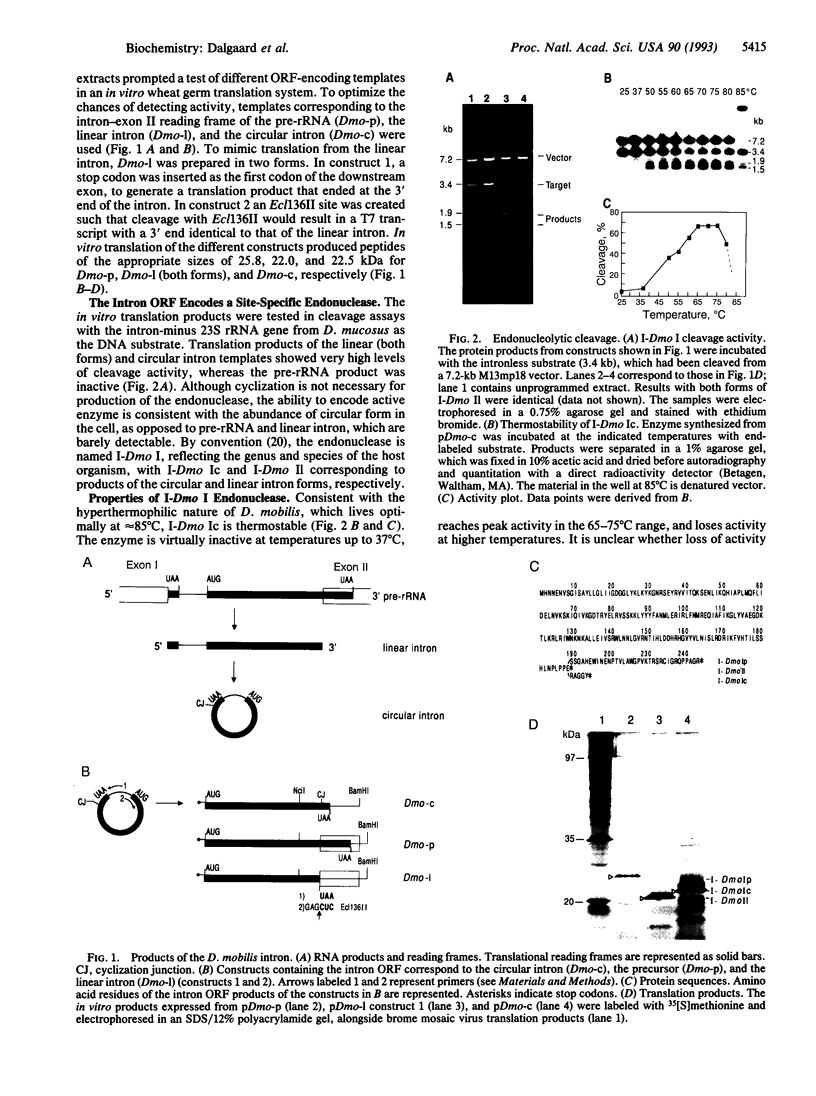


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Belfort M. Phage T4 introns: self-splicing and mobility. Annu Rev Genet. 1990;24:363–385. doi: 10.1146/annurev.ge.24.120190.002051. [DOI] [PubMed] [Google Scholar]
- Bell-Pedersen D., Quirk S., Clyman J., Belfort M. Intron mobility in phage T4 is dependent upon a distinctive class of endonucleases and independent of DNA sequences encoding the intron core: mechanistic and evolutionary implications. Nucleic Acids Res. 1990 Jul 11;18(13):3763–3770. doi: 10.1093/nar/18.13.3763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colleaux L., Michel-Wolwertz M. R., Matagne R. F., Dujon B. The apocytochrome b gene of Chlamydomonas smithii contains a mobile intron related to both Saccharomyces and Neurospora introns. Mol Gen Genet. 1990 Sep;223(2):288–296. doi: 10.1007/BF00265065. [DOI] [PubMed] [Google Scholar]
- Cummings D. J., Michel F., McNally K. L. DNA sequence analysis of the apocytochrome b gene of Podospora anserina: a new family of intronic open reading frame. Curr Genet. 1989 Dec;16(5-6):407–418. doi: 10.1007/BF00340720. [DOI] [PubMed] [Google Scholar]
- Dalgaard J. Z., Garrett R. A. Protein-coding introns from the 23S rRNA-encoding gene form stable circles in the hyperthermophilic archaeon Pyrobaculum organotrophum. Gene. 1992 Nov 2;121(1):103–110. doi: 10.1016/0378-1119(92)90167-n. [DOI] [PubMed] [Google Scholar]
- Daniels C. J., Gupta R., Doolittle W. F. Transcription and excision of a large intron in the tRNATrp gene of an archaebacterium, Halobacterium volcanii. J Biol Chem. 1985 Mar 10;260(5):3132–3134. [PubMed] [Google Scholar]
- Davis E. O., Jenner P. J., Brooks P. C., Colston M. J., Sedgwick S. G. Protein splicing in the maturation of M. tuberculosis recA protein: a mechanism for tolerating a novel class of intervening sequence. Cell. 1992 Oct 16;71(2):201–210. doi: 10.1016/0092-8674(92)90349-h. [DOI] [PubMed] [Google Scholar]
- Delahodde A., Goguel V., Becam A. M., Creusot F., Perea J., Banroques J., Jacq C. Site-specific DNA endonuclease and RNA maturase activities of two homologous intron-encoded proteins from yeast mitochondria. Cell. 1989 Feb 10;56(3):431–441. doi: 10.1016/0092-8674(89)90246-8. [DOI] [PubMed] [Google Scholar]
- Dujon B., Belfort M., Butow R. A., Jacq C., Lemieux C., Perlman P. S., Vogt V. M. Mobile introns: definition of terms and recommended nomenclature. Gene. 1989 Oct 15;82(1):115–118. doi: 10.1016/0378-1119(89)90035-8. [DOI] [PubMed] [Google Scholar]
- Dujon B. Group I introns as mobile genetic elements: facts and mechanistic speculations--a review. Gene. 1989 Oct 15;82(1):91–114. doi: 10.1016/0378-1119(89)90034-6. [DOI] [PubMed] [Google Scholar]
- Dürrenberger F., Rochaix J. D. Chloroplast ribosomal intron of Chlamydomonas reinhardtii: in vitro self-splicing, DNA endonuclease activity and in vivo mobility. EMBO J. 1991 Nov;10(11):3495–3501. doi: 10.1002/j.1460-2075.1991.tb04913.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garrett R. A., Dalgaard J., Larsen N., Kjems J., Mankin A. S. Archaeal rRNA operons. Trends Biochem Sci. 1991 Jan;16(1):22–26. doi: 10.1016/0968-0004(91)90011-j. [DOI] [PubMed] [Google Scholar]
- Gauthier A., Turmel M., Lemieux C. A group I intron in the chloroplast large subunit rRNA gene of Chlamydomonas eugametos encodes a double-strand endonuclease that cleaves the homing site of this intron. Curr Genet. 1991 Jan;19(1):43–47. doi: 10.1007/BF00362086. [DOI] [PubMed] [Google Scholar]
- Gimble F. S., Thorner J. Homing of a DNA endonuclease gene by meiotic gene conversion in Saccharomyces cerevisiae. Nature. 1992 May 28;357(6376):301–306. doi: 10.1038/357301a0. [DOI] [PubMed] [Google Scholar]
- Hensgens L. A., Bonen L., de Haan M., van der Horst G., Grivell L. A. Two intron sequences in yeast mitochondrial COX1 gene: homology among URF-containing introns and strain-dependent variation in flanking exons. Cell. 1983 Feb;32(2):379–389. doi: 10.1016/0092-8674(83)90457-9. [DOI] [PubMed] [Google Scholar]
- Hirata R., Ohsumk Y., Nakano A., Kawasaki H., Suzuki K., Anraku Y. Molecular structure of a gene, VMA1, encoding the catalytic subunit of H(+)-translocating adenosine triphosphatase from vacuolar membranes of Saccharomyces cerevisiae. J Biol Chem. 1990 Apr 25;265(12):6726–6733. [PubMed] [Google Scholar]
- Hodges R. A., Perler F. B., Noren C. J., Jack W. E. Protein splicing removes intervening sequences in an archaea DNA polymerase. Nucleic Acids Res. 1992 Dec 11;20(23):6153–6157. doi: 10.1093/nar/20.23.6153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaine B. P., Gupta R., Woese C. R. Putative introns in tRNA genes of prokaryotes. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3309–3312. doi: 10.1073/pnas.80.11.3309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kane P. M., Yamashiro C. T., Wolczyk D. F., Neff N., Goebl M., Stevens T. H. Protein splicing converts the yeast TFP1 gene product to the 69-kD subunit of the vacuolar H(+)-adenosine triphosphatase. Science. 1990 Nov 2;250(4981):651–657. doi: 10.1126/science.2146742. [DOI] [PubMed] [Google Scholar]
- Kjems J., Garrett R. A. Novel splicing mechanism for the ribosomal RNA intron in the archaebacterium Desulfurococcus mobilis. Cell. 1988 Aug 26;54(5):693–703. doi: 10.1016/s0092-8674(88)80014-x. [DOI] [PubMed] [Google Scholar]
- Kjems J., Garrett R. A. Ribosomal RNA introns in archaea and evidence for RNA conformational changes associated with splicing. Proc Natl Acad Sci U S A. 1991 Jan 15;88(2):439–443. doi: 10.1073/pnas.88.2.439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kjems J., Jensen J., Olesen T., Garrett R. A. Comparison of transfer RNA and ribosomal RNA intron splicing in the extreme thermophile and archaebacterium Desulfurococcus mobilis. Can J Microbiol. 1989 Jan;35(1):210–214. doi: 10.1139/m89-033. [DOI] [PubMed] [Google Scholar]
- Lambowitz A. M., Belfort M. Introns as mobile genetic elements. Annu Rev Biochem. 1993;62:587–622. doi: 10.1146/annurev.bi.62.070193.003103. [DOI] [PubMed] [Google Scholar]
- Lambowitz A. M. Infectious introns. Cell. 1989 Feb 10;56(3):323–326. doi: 10.1016/0092-8674(89)90232-8. [DOI] [PubMed] [Google Scholar]
- Michel F., Dujon B. Genetic exchanges between bacteriophage T4 and filamentous fungi? Cell. 1986 Aug 1;46(3):323–323. doi: 10.1016/0092-8674(86)90651-3. [DOI] [PubMed] [Google Scholar]
- Michel F., Jacquier A., Dujon B. Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie. 1982 Oct;64(10):867–881. doi: 10.1016/s0300-9084(82)80349-0. [DOI] [PubMed] [Google Scholar]
- Moran J. V., Wernette C. M., Mecklenburg K. L., Butow R. A., Perlman P. S. Intron 5 alpha of the COXI gene of yeast mitochondrial DNA is a mobile group I intron. Nucleic Acids Res. 1992 Aug 11;20(15):4069–4076. doi: 10.1093/nar/20.15.4069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mota E. M., Collins R. A. Independent evolution of structural and coding regions in a Neurospora mitochondrial intron. Nature. 1988 Apr 14;332(6165):654–656. doi: 10.1038/332654a0. [DOI] [PubMed] [Google Scholar]
- Muscarella D. E., Vogt V. M. A mobile group I intron in the nuclear rDNA of Physarum polycephalum. Cell. 1989 Feb 10;56(3):443–454. doi: 10.1016/0092-8674(89)90247-x. [DOI] [PubMed] [Google Scholar]
- Nakagawa K., Morishima N., Shibata T. A maturase-like subunit of the sequence-specific endonuclease endo.SceI from yeast mitochondria. J Biol Chem. 1991 Jan 25;266(3):1977–1984. [PubMed] [Google Scholar]
- Nakagawa K., Morishima N., Shibata T. An endonuclease with multiple cutting sites, Endo.SceI, initiates genetic recombination at its cutting site in yeast mitochondria. EMBO J. 1992 Jul;11(7):2707–2715. doi: 10.1002/j.1460-2075.1992.tb05336.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perler F. B., Comb D. G., Jack W. E., Moran L. S., Qiang B., Kucera R. B., Benner J., Slatko B. E., Nwankwo D. O., Hempstead S. K. Intervening sequences in an Archaea DNA polymerase gene. Proc Natl Acad Sci U S A. 1992 Jun 15;89(12):5577–5581. doi: 10.1073/pnas.89.12.5577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlman P. S., Butow R. A. Mobile introns and intron-encoded proteins. Science. 1989 Dec 1;246(4934):1106–1109. doi: 10.1126/science.2479980. [DOI] [PubMed] [Google Scholar]
- Sargueil B., Delahodde A., Hatat D., Tian G. L., Lazowska J., Jacq C. A new specific DNA endonuclease activity in yeast mitochondria. Mol Gen Genet. 1991 Feb;225(2):340–341. doi: 10.1007/BF00269867. [DOI] [PubMed] [Google Scholar]
- Sharma M., Ellis R. L., Hinton D. M. Identification of a family of bacteriophage T4 genes encoding proteins similar to those present in group I introns of fungi and phage. Proc Natl Acad Sci U S A. 1992 Jul 15;89(14):6658–6662. doi: 10.1073/pnas.89.14.6658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shub D. A., Goodrich-Blair H. Protein introns: a new home for endonucleases. Cell. 1992 Oct 16;71(2):183–186. doi: 10.1016/0092-8674(92)90345-d. [DOI] [PubMed] [Google Scholar]
- Séraphin B., Faye G., Hatat D., Jacq C. The yeast mitochondrial intron aI5 alpha: associated endonuclease activity and in vivo mobility. Gene. 1992 Apr 1;113(1):1–8. doi: 10.1016/0378-1119(92)90663-a. [DOI] [PubMed] [Google Scholar]
- Thompson A. J., Yuan X., Kudlicki W., Herrin D. L. Cleavage and recognition pattern of a double-strand-specific endonuclease (I-creI) encoded by the chloroplast 23S rRNA intron of Chlamydomonas reinhardtii. Gene. 1992 Oct 1;119(2):247–251. doi: 10.1016/0378-1119(92)90278-w. [DOI] [PubMed] [Google Scholar]
- Thompson L. D., Brandon L. D., Nieuwlandt D. T., Daniels C. J. Transfer RNA intron processing in the halophilic archaebacteria. Can J Microbiol. 1989 Jan;35(1):36–42. doi: 10.1139/m89-006. [DOI] [PubMed] [Google Scholar]
- Thompson L. D., Daniels C. J. A tRNA(Trp) intron endonuclease from Halobacterium volcanii. Unique substrate recognition properties. J Biol Chem. 1988 Dec 5;263(34):17951–17959. [PubMed] [Google Scholar]
- Thompson L. D., Daniels C. J. Recognition of exon-intron boundaries by the Halobacterium volcanii tRNA intron endonuclease. J Biol Chem. 1990 Oct 25;265(30):18104–18111. [PubMed] [Google Scholar]
- Waring R. B., Davies R. W., Scazzocchio C., Brown T. A. Internal structure of a mitochondrial intron of Aspergillus nidulans. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6332–6336. doi: 10.1073/pnas.79.20.6332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wenzlau J. M., Saldanha R. J., Butow R. A., Perlman P. S. A latent intron-encoded maturase is also an endonuclease needed for intron mobility. Cell. 1989 Feb 10;56(3):421–430. doi: 10.1016/0092-8674(89)90245-6. [DOI] [PubMed] [Google Scholar]
- Woese C. R., Kandler O., Wheelis M. L. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4576–4579. doi: 10.1073/pnas.87.12.4576. [DOI] [PMC free article] [PubMed] [Google Scholar]