Abstract
Bacteria lack subcellular compartments and harbor a single RNA polymerase that synthesizes both structural and protein-coding RNAs, which are cotranscriptionally processed by distinct pathways. Nascent rRNAs fold into elaborate secondary structures and associate with ribosomal proteins, whereas nascent mRNAs are translated by ribosomes. During elongation, nucleic acid signals and regulatory proteins modulate concurrent RNA-processing events, instruct RNA polymerase where to pause and terminate transcription, or act as roadblocks to the moving enzyme. Communications among complexes that carry out transcription, translation, repair, and other cellular processes ensure timely execution of the gene expression program and survival under conditions of stress. This network is maintained by auxiliary proteins that act as bridges between RNA polymerase, ribosome, and repair enzymes, blurring boundaries between separate information-processing steps and making assignments of unique regulatory functions meaningless. Understanding the regulation of transcript elongation thus requires genome-wide approaches, which confirm known and reveal new regulatory connections.
Keywords: RNA polymerase, pausing, antitermination, transcription-coupled DNA repair, translation, Nus factors
INTRODUCTION
RNA polymerase (RNAP) synthesizes an RNA copy of the template DNA, the first and often decisive step in gene expression. All RNAPs that transcribe cellular genomes are multisubunit enzymes that share homologous catalytic cores. Bacterial core RNAP, a five-subunit complex (α2ββ′ω), is the simplest model system for studies of fundamental properties of all multisubunit RNAPs. Following initiation, which requires a σ factor, the core enzyme can elongate the RNA chain and terminate transcription guided only by nucleic acid signals in vitro, but even a single accessory protein can significantly alter the pattern of RNA synthesis. By contrast, in vivo transcribing RNAP shares its template and product with other information-processing complexes and associates with regulators that fine-tune the balance between processive elongation, pausing, and termination; contribute to proofreading; and help it recover from arrested states. This regulatory complexity prompted development of whole-cell approaches to monitor transcription across the entire genome. These studies revealed that RNAP and its associated proteins cross talk with many parallel regulatory pathways. Consequently, transcriptional regulators with well-established functions were discovered to have essential roles in translation and repair, among other functions. A holistic view, in which multiple players act together to enable cells to function optimally under drastically different conditions, is rapidly emerging. In this review, we briefly cover recent advances in our understanding of the molecular mechanism and regulation of transcript elongation by bacterial RNAP and the coupled processes that enable RNAP to carry out its function in the crowded cellular environment. We apologize to many colleagues whose work we cannot cite because of space limitations.
OVERALL STRUCTURE AND CHEMICAL CATALYSIS
In contrast to replicative polymerases, transcribing RNAPs do not displace the nontemplate strand; rather, they maintain an 8- to 9-base pair (bp) RNA-DNA hybrid within the 11- to 12-bp melted DNA bubble (Figure 1). The overall shape of the RNAP molecule resembles a crab claw (153), with the two largest β′ and β subunits forming a cleft that acts as a binding site for the RNA-DNA hybrid and the downstream DNA duplex (134). The upstream DNA duplex is located outside the cleft and is only loosely associated with RNAP (2). The two α subunits are located at the periphery, serving largely structural roles, except for their C-terminal domains (CTDs), which extend 40 Å from the RNAP body to establish regulatory interactions with the upstream DNA and transcription factors (76, 107). A small ω subunit binds near the RNA exit channel and facilitates RNAP assembly and regulation (137, 155).
Figure 1.
Structural overview of the bacterial TEC. The βsubunit is rendered semitransparent, and the β′ subunit surface is selectively carved to reveal the path of nucleic acids through RNAP and the active site elements. The composite TEC model was generated using elements from PDB ID 2O5J and 4IGC; upstream DNA and single-stranded nontemplate DNA were modeled using data from Reference 2. The schematic and molecular surfaces were drawn using the PyMOL Molecular Graphics System, Version 1.6.4.0 Schrödinger, LLC, and further simplified using the computer-aided design software MeshLab, MeshMixer, Tinkercad, and Rhinoceros 3D. Abbreviations: BH, bridge helix; CTD, C-terminal domain; NTP, nucleoside triphosphate; RNAP, RNA polymerase; TEC, transcription elongation complex; TL, trigger loop.
The active site is located within the cleft, 30–40 Å beneath the protein surface at the bottom of a funnel-shaped secondary channel that serves as an entry route for nucleoside triphosphate (NTP) substrates (134, 140). The active site can be schematically subdivided into three sections: an i site, an i+1 site, and a mobile β′ domain called trigger loop (TL, a.k.a. G-loop). The i site features a β′ loop with three universally conserved aspartate residues that tightly chelate a Mg2+ ion, which binds the 3′ OH of the nascent RNA and activates it for SN2 nucleophilic attack on the α-phosphate of the NTP substrate (121). The incoming NTP in complex with a second Mg2+ ion binds to the i+1 site, which contains a vacant template DNA base that pairs with an NTP base, several basic β and β′ residues that interact with NTP triphosphate moiety, and conserved β′ asparagine and arginine residues that interact with NTP ribose (122, 135). Substrate loading is thought to be a multistep process in which initial binding in a catalytically incompetent conformation is followed by rate-limiting isomerization into a catalytically proficient conformation that involves the closure of the active site (135).
The TL is typically unresolved in structures of transcription elongation complexes (TECs), indicating its mobility (134). In the presence of the substrate, the TL adopts a helical hairpin conformation and closes the active site by obstructing the secondary channel with a highly conserved N-terminal helix, which forms multiple stabilizing interactions with NTP (135, 140). TL folding also involves the formation of a triple-helix bundle with the β′ bridge helix (BH), a long metastable α-helix that spans the active site cleft and molds into a groove in the β subunit (52). Finally, the stability of the helical TL is also modulated by interactions of the TL tip with the β′ F-loop, an N-terminal extension of the BH (82). TL closure aligns NTP for efficient catalysis (135) and introduces into the i+1 site a conserved histidine residue capable of protonating the leaving PPi group (140). Stabilization of the closed conformation requires Watson-Crick base-pairing between NTP and acceptor template base and ribose as the NTP sugar. Accordingly, the TL contributes 100- to 1,000-fold to the rate of phosphodiester bond formation and fidelity of transcription (61, 65,72, 140, 151).
POSTCATALYTIC RELAXATION OF THE TRANSCRIPTION ELONGATION COMPLEX: PYROPHOSPHATE RELEASE AND TRANSLOCATION
Nucleotide addition extends the RNA-DNA hybrid by one base pair, resulting in a pretranslocated state in which the i+1 site is occupied by newly incorporated nucleoside monophosphate and pyrophosphate (Figure 2). Postcatalytic relaxation into a catalytically proficient posttranslocated state involves the release of PPi, opening of the active site, and RNAP translocation. During translocation, the hybrid and the bubble move forward by one nucleotide, the newly generated RNA 3′ end moves into the i site, and a new template DNA base enters the i+1 site (62). Although it had been initially assumed that RNAP equilibrates rapidly between the pre- and posttranslocated states after each round of nucleotide addition, recent reports suggest that translocation rates are comparable to catalytic rates (24, 78). Thus, the overall kinetics of elongation likely represent a complex balance of nucleotide addition and forward and backward translocation rates at each sequence position. Similarly, whereas early models calculated the translocation bias on the basis of base-pairing energies of nucleic acids within the TEC, recent studies argue that interactions of the RNA 3′ end with the active site may be pivotal (36, 53, 72, 78): The open active site favors the posttranslocated state because of the intrinsically high affinity of the 3′ end to the i site, whereas the closed active site favors the pretranslocated state, in which the 3′ end interacts with the i+1 site.
Figure 2.
Schematic overview of the nucleotide addition cycle. NTP binding is abbreviated into a single transition. TL folding into a triple-helix bundle with the BH is depicted as a transition from an isolated shape into a dumbbell. Abbreviations: BH, bridge helix; NMP, nucleoside monophosphate; NTP, nucleoside triphosphate; TL, trigger loop.
The loss of translocation register can generate other translocation states. A hypertranslocated state, in which the RNA 3′ end moves upstream into the RNAP main channel, has been suggested as an intermediate during termination (93) and may also occur during processive elongation (86). A backtracked state (88), in which the RNAP has retreated along the template, has been characterized biochemically (66) and structurally (139). RNAP backtracked by one or a few residues can slowly reestablish the active conformation via successive rounds of forward translocation and/or RNA cleavage by weak intrinsic RNase activity of RNAP (151).
Backtracking a greater distance extrudes the 3′ end of the RNA segment into the secondary channel and leads to arrest. Bacteria encode several regulatory proteins, such as GreA/B and DksA, that bind within the secondary channel (90, 96). These proteins interact with RNAP via their globular domains and extend their coiled-coil domains through the secondary channel all the way toward the active site to prevent arrest (96) or to rescue arrested TECs via cleavage of the nascent RNA (70).
PAUSING
RNAP movement along the DNA can be hindered by a DNA-bound protein, by a lesion in the template strand, or upon misincorporation. However, transcription is interrupted more often by pauses, which slow nucleotide addition 10- to 1,000-fold even on intact templates in vitro. Pausing likely determines the overall rate of RNA chain synthesis, is an obligatory step in termination, and may be necessary for timely recruitment of regulatory factors, attenuation control, cotranscriptional folding of the nascent RNA, and efficient coupling of transcription and translation (see 68 for review).
Pauses are events that branch off the main elongation pathway: Although RNAP has a low probability of isomerizing into a short-lived paused state at every position (87), a fraction of the RNAP bypasses even strong pauses without delay (18). Pausing is thus an intrinsic property of RNAP augmented by sequence context; certain sequences dramatically increase the probability (efficiency in ensemble experiments) of RNAP isomerizing into an elemental paused state (a non-backtracked inactivated state accessible for all TECs, irrespective of the sequence context) and often cause additional structural rearrangements that lead to long-lived stabilized pauses. The initial isomerization is thought to proceed similarly during the majority of pause events, whereas subsequent stabilization follows distinct, relatively well-understood pathways, which include RNAP backtracking (66) or stabilization of the RNA 3′ end in either posttranslocated (64) or pretranslocated (130) register. In the latter, the hairpin-stabilized-pathway TL plays a critical role (130). Interactions of the unpaired nontemplate DNA strand with the β subunit (63, 138) or regulatory proteins, such as NusG, RfaH, and σ (5, 105, 113, 147), may also modulate pause efficiency and duration.
In contrast, isomerization into elemental pause remains obscure because of the transient nature of this state, but it is thought to involve concerted bending of the BH, which is modulated by conformation cycling of the interacting TL, and opening of the β′ clamp domain (51). Both transitions favor distortion of the template DNA in the active site. It is commonly anticipated that isomerization originates from the pretranslocated state when a block to translocation is encountered; the failure of the RNA-DNA hybrid to translocate has been proposed as one mechanism (11). It has been long anticipated that fraying of the RNA 3′ end before or during translocation results in short-term inactivation (4). However, recent structural studies suggest that incomplete translocation of the downstream DNA leads to a state in which the RNA 3′ end is properly positioned but the acceptor template base is unavailable (143). Genome-wide analyses of pause sites (71, 138) established that a pyrimidine/purine active site combination is the most conserved feature, consistent with template-strand isomerization being the principal cause of elemental pauses. However, efficient, physiologically relevant pauses encompass multiple interactions along the entire RNAP footprint on the DNA and beyond (18, 71, 138).
Subsequent rearrangements of a paused complex may also lead to termination, an irreversible process triggered by the formation of a nascent RNA hairpin (47) or the ATP-dependent RNA translocase Rho (104) (see 99 for a comprehensive review of the mechanisms and regulation of termination).
Nus FACTORS AND rRNA ANTITERMINATION COMPLEXES
Translating ribosomes ensure uninterrupted RNA chain elongation by inhibiting backtracking (102) and premature termination by Rho (104). Rho, aided by NusG, suppresses expression of poorly translated foreign DNA (16) and antisense transcription (97). However, Escherichia coli RNAP transcribes the noncoding rRNA genes twice as fast as it transcribes mRNA genes and it is resistant to Rho (127, 152). These properties are conferred by the assembly of a ribonucleoprotein antitermination complex composed of NusA, NusB, NusG, and NusE (ribosomal protein S10) factors (Figure 3), as well as ribosomal protein S4 and perhaps other cellular factors (127), which form a proteinaceous extension of the RNA exit channel (115).
Figure 3.
NusA and the antitermination complex. (a) Functions and distribution of NusA domains. In archaeal and eukaryotic RNAPs, the S1 domain is part of a dissociable heterodimeric stalk module that, like NusA, is located near the RNA exit channel. Although S1 domains are found in many RNA-binding proteins, those of NusA and stalk subunits—RpoE (archaea), A43 (RNAP I), Rpb7 (RNAP II), and Rpc25 (RNAP III)—are more similar to each other than to translation-associated S1 domains and thus may have similar functions. (b) NusABEG assembly (individual domains depicted as colored circles) facilitates the transcription and folding of rRNA. (c) rRNA operon structure in prokaryotes. Abbreviations: CTD, C-terminal domain; NTD, N-terminal domain; RNAP, RNA polymerase.
NusA, the largest Nus factor and the centerpiece of the antitermination complex, consists of several domains with distinct functionalities and conservation patterns. The nusA gene is invariably present as a single copy and lacks paralogs. The four-domain core, composed of the RNAP-binding N-terminal domain (NTD) and the nascent RNA-binding module (S1-KH-KH domains), is ubiquitous in bacteria (Figure 3a). NusA is presumed to have at least three major cellular functions. It (a) enhances sensitivity of RNAP to RNA hairpins that mediate pausing and intrinsic termination (35, 47, 117), (b) participates in the assembly of antitermination complexes (119), and (c) facilitates cotranscriptional folding of structural RNAs (12, 91). The first function can be attributed largely to the NTD (49), whereas the last two functions are mediated presumably by the RNA-binding domains.
The NTD tethers NusA to RNAP and is thus at least indirectly involved in all NusA activities. It binds to the mouth of, and remodels, the RNA exit channel via interactions with the nascent RNA hairpin, β flap, and possibly β′ dock domains (49, 129, 149). Although the exact structural rearrangements upon NusA binding to RNAP are not yet fully understood, it is conceivable that interactions with a flexible wall (β flap domain) of the RNA exit channel may affect the formation of RNA secondary structures just upstream of the RNA-DNA hybrid. The NusA NTD is restricted to bacteria, probably reflecting the prevalence of hairpin-dependent termination in bacteria but not in archaea and eukaryotes (3, 109).
The NusA RNA-binding module is connected to the NTD via a flexible linker and consists of one S1 domain (named after ribosomal protein S1 and commonly found in translation-related proteins) and two KH domains, which interact extensively with each other to form a continuous RNA-binding surface (43, 146). The S1 domain is predicted to reside near the RNA exit channel and closely resembles bacterial cold-shock proteins that function as general RNA chaperones (59). Hence, the S1 domain may loosen unwanted, and assist the formation of desired, secondary structures to facilitate folding of structural RNAs and recruitment of ribosome to mRNA. The chaperoning action of S1 may also contribute to rRNA antitermination, as RNA chaperones typically display antitermination activity (7, 103, 127).
The NusABEG complex (Figure 3b) is held together by relatively weak protein-protein and protein-RNA interactions and assembles in a highly cooperative manner (75). The NTDs of NusA and NusG anchor the ends of the assembly to β flap (49, 115) and β′ clamp helices (10), respectively. The NusA KH domains and NusB do not interact directly but are instead linked via binding to the adjacent RNA sequences in the upstream regions of rRNA genes (44, 101). NusE simultaneously binds to NusB and NusG (14, 75), blocking the Rho-binding site on the NusG CTD (14) and thereby inhibiting Rho-mediated termination.
Recent studies suggest that the key role of the NusABEG complex is to mediate cotranscriptional folding of rRNA by stabilizing pre-rRNA in a looped conformation via contacts of NusA S1 and KH domains with RNA emerging from the exit channel and contacts of NusB/E and KH domains with the 5′ region (12). This idea is supported by rRNA genes structures in bacteria and archaea (150), in which the 5′ and 3′ flanking regions of pre-16S and pre-23S rRNAs form giant stem-loop structures with ~50-bp stems (Figure 3c). These stems presumably maintain the looped conformation to provide time for rRNA folding to complete before processing by dedicated RNases (12). The association of the KH domain with rRNA biogenesis is consistent with its distribution pattern: Archaea, but not eukaryotes, harbor a factor that displays high sequence identity to KH-KH repeats of bacterial NusA and similar spacerless head-to-tail packing (116). Intriguingly, the presumed loss of KH domains in eukaryotes coincides with the evolution of a specialized RNAP for rRNA synthesis and, perhaps more importantly, a principally different rRNA folding mechanism (124). NusABEG proteins are absolutely conserved in bacteria whereas Rho is absent in many species, suggesting a conserved NusABEG function distinct from antitermination (28).
TRANSCRIPTION AND TRANSLATION
In bacteria, transcription and translation are considered coupled: They occur in the same compartment at highly correlated rates (102, 152). However, ribosomes outnumber RNAPs at least 10:1 (74) and the bulk of translation occurs on preexisting mRNAs (8), governed by the stability and translation efficiency of mRNAs. Coupling is limited to the first round of translation, which plays several unique roles. During this pioneer round, the ribosome could push RNAP forward to prevent backtracking and maintain processive RNA synthesis (102); inhibit premature release of mRNA by Rho (104); and alter folding of the nascent mRNA, thereby controlling the termination/antitermination decision during attenuation (54). Coupling between transcription and translation allows for coordinated regulation of gene expression in response to physiological cues. Slowing down translation by adding antibiotics or rare codons leads to a concomitant slowdown of transcription to maintain the protective link (102), whereas starvation-induced ribosome arrest (154) and ribosome release at a stop codon would trigger uncoupling (104), leading to arrest and termination. Thus, coupling acts as a surveillance mechanism to ensure that mRNA is translatable and that cellular conditions are conducive.
In E. coli, NusG plays a key role in this quality-control mechanism. The NusG NTD binds to RNAP transcribing most genes (83) to reduce pausing (55), whereas the CTD can bind either Rho to facilitate RNA release (84) or small ribosomal subunit protein S10 to enable coupling (14). The CTD contacts with Rho and S10 are mutually exclusive; thus, translation would be doubly protective against termination, hindering Rho from binding to either RNA or NusG. The NusG-S10 interaction is well suited to maintain the tight link between the two machines (14) and may also facilitate ribosome recruitment to mRNAs, perhaps aided by RNAP pausing at translation start sites (71). This mechanism has been proposed for the NusG paralog RfaH (13), which dramatically activates expression of genes otherwise silenced by Rho (114).
TRANSCRIPTION AND REPAIR
Mfd: The First Bridge Between Transcription and Repair
Many types of lesions in the template DNA strand stall, but do not dissociate, the RNAP (126). The stalled RNAP occludes the site of damage, blocking the repair enzymes from accessing it (Figure 4). However, active transcription of a gene facilitates preferential repair of the template strand in vivo (41). Mfd was identified as a key transcription-coupled repair (TCR) factor in E. coli over two decades ago (111). Mfd binds to DNA upstream of the stalled RNAP and, utilizing ATP hydrolysis, moves along the double-stranded DNA, pushing the enzyme forward and displacing it from the DNA template (92, 111). Following RNAP release, Mfd recruits UvrA, a key nucleotide excision repair (NER) factor, to the unmasked lesion (111, 112), initiating the repair pathway composed of lesion excision and gap filling (79, 111). Mfd is homologous to TCR factors in other systems and has been characterized extensively (25, 110).
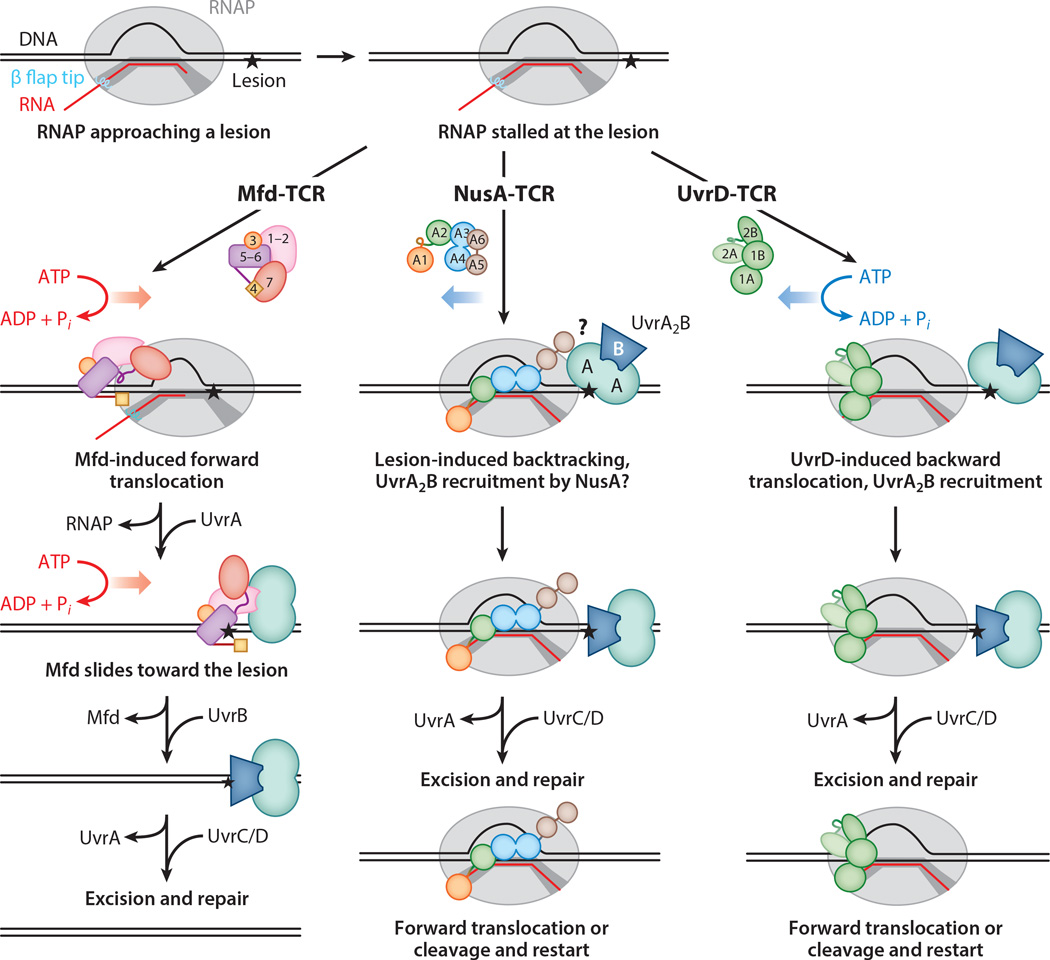
Figure 4.
Three pathways of TCR. DNA repair requires removal of the RNAP that occludes a DNA lesion (black star) in the template DNA strand. Mfd pushes RNAP forward, releasing it from DNA, and then recruits UvrA. UvrD pushes RNAP backward, exposing the lesion to Uvr proteins. NusA may stabilize a backtracked state induced by RNAP recognition of certain lesions and recruit UvrA. Following recruitment of UvrB, the repair pathway follows the canonical NER mechanism. Structural domains of Mfd, NusA, and UvrD are indicated by numbers; only Mfd has been studied in sufficient detail to assign its structural domains individual roles in TCR. Abbreviations: NER, nucleotide excision repair; RNAP, RNA polymerase; TCR, transcription-coupled repair.
E. coli Mfd is a large, dynamic multidomain protein (26). Domains 1 and 2 are structurally similar to the NER protein UvrB, which also binds UvrA; domain 4 interacts with the β subunit; and domains 5 and 6 are necessary for ATP-dependent translocation along the DNA (25). Domain 7 plays a critical autoinhibitory role. In free Mfd, it interacts with domains 1 and 2 to mask the UvrA-binding surface (27) and restrains translocase domains 5 and 6 to inhibit ATPase function and translocation on DNA (85). These interactions must be disrupted to enable Mfd-induced termination and subsequent NER recruitment.
The simplest model posits that a single conformational switch upon Mfd binding to stalled RNAP simultaneously enables RNAP release and UvrA recruitment (85). However, a single-step activation appears unlikely. Mfd is recruited to a paused RNAP in the absence of a lesion (50, 92) and fortuitous recruitment of UvrA could prove deleterious, diverting resources from global NER. Indeed, recent studies provide evidence that different Mfd activities can be uncoupled in a multistep activation pathway. First, substitutions at the D2–D7 interface elevate ATPase but not DNA translocation activity, and Mfd lacking the inhibitory domain 7 is defective in repair, despite efficient RNAP displacement activity (79). Second, a substitution designed to disrupt communications between domains 4, 5/6, and 7 results in a hyperactive Mfd able to displace any transcription complex (118), including those formed by RNAPs lacking the Mfd-binding site or Mfd-resistant initiation complexes. Third, cross-linking of domains 2 and 7 impairs Mfd binding to DNA in the absence of RNAP but not termination (27). Finally, single-molecule experiments revealed two sequential ATP-dependent steps: collapse of the transcription bubble followed by formation of a long-lived intermediate in which Mfd remained bound to DNA (56).
The point of RNAP release from Mfd remains unclear but likely coincides with transcription bubble collapse. During productive engagement with the stalled TEC (56), Mfd likely undergoes conformational changes (27) that convert it into an ATP-driven plow that displaces nonspecific obstacles, such as triplex oligonucleotides, from its path (118). The continual association with RNAP following termination could be difficult to maintain via a small interaction surface (144) without stabilizing contacts with DNA (56) and may not be required to maintain Mfd in an activated state (118). The timing of UvrA loading is also unclear. Repositioning of domain 7 is needed to expose the UvrA-binding site on Mfd but is dispensable for termination (27). It was proposed that, following RNAP release, Mfd slides forward and, upon recognition of a lesion, undergoes conformational changes that unmask the UvrA interface (25, 27). In support of this idea, activated Mfd can slide along the DNA for hundreds of base pairs in search of a lesion (50).
These studies revealed an intricate Mfd-TCR pathway that couples recognition of a DNA lesion by transcribing RNAP with NER. However, they did not explain why E. coli cells that lack Mfd were not hypersensitive to UV-induced DNA damage (111), the primary target for NER(41). Recent reports posited that NusA and UvrD, the key players in transcription and repair, contribute to TCR in E. coli (21, 33). These leads, although lacking the depth of the Mfd studies, reveal a common strategy: Proteins with well-established regulatory roles have additional moonlighting activities that may be equally important for cell viability.
NusA: DNA Repair and Damage Tolerance
Observations that plasmids overexpressing E. coli dinB or umuDC, SOS-induced translesion synthesis (TLS) DNA polymerases, could suppress the nusA11 strain phenotypes and that NusA and DinB interact directly with each other suggest NusA’s involvement in DNA repair (20). TLS polymerases repair gaps in DNA at the cost of lower fidelity to enable survival in the face of massive DNA damage that will block progression of replisomes (38). NusA, bound to the RNAP stalled by a gap in the transcribed strand, could recruit a TLS polymerase to fill this gap (20). However, that the nusA11 mutant is 100 times more defective in DinB-dependent stress-induced mutagenesis than dinB− (22) suggested that NusA plays other roles in DNA repair.
Further analysis demonstrated that strains lacking functional nusA were sensitive to nitrofurazone (NFZ), UV radiation, and methylmethane sulfonate, while ruling out NusA-mediated changes in expression of DNA repair genes (21). NusA did not help RNAP bypass a large gap or an NFZ-induced adduct in vitro, prompting a search for a missing link that culminated in the identification of UvrA as an interaction partner (21); NFZ-induced DNA damage is processed by the Uvr pathway (89). Genetic analysis revealed that NusA and Mfd function in parallel TCR pathways, with double deletion strains dramatically more sensitive to UV-induced killing.
The proposed NusA-TCR mechanism invokes RNAP backtracking upon running into a lesion, followed by NusA-mediated recruitment of NER. In support of this model, β subunit substitutions that confer NusA-dependent sensitivity or resistance to NFZ have been identified (21). These residues are located at the leading edge of RNAP and can directly sense the damaged DNA, prompting RNAP to retreat. It remains unclear how NusA reaches from the RNA exit channel to the downstream DNA to recruit UvrA to the lesion.
UvrD: Pushing RNAP Backward
UvrD is an integral late player in NER—it is a helicase that removes the damaged DNA fragment following processing by UvrABC (148). However, UvrD also interacts with RNAP in E. coli (33) and Bacillus subtilis (48), suggestive of a TCR model in which UvrD makes the stalled RNAP retreat from the lesion, exposing it for repair (33). In support of this model, UvrD induced RNAP backtracking in vivo and in vitro and relieved an RNAP-imposed block to UvrABC excision of a thymine dimer in a reconstituted NER system. UvrD cross-linked to the β flap tip and the nontemplate DNA strand at the upstream part of the transcription bubble. Upon engaging the single DNA strand, UvrD could slide on the DNA and push RNAP backward, a mechanism consistent with displacement of DNA-binding proteins by UvrD (148). Sensitivity of the uvrD strain to genotoxic agents was alleviated partially by deletions in greA/B or by the presence of chloramphenicol, an inhibitor of translation. Because Gre activity and trailing ribosomes inhibit backtracking (88), these results suggest that if RNAP could retreat on its own, then repair would be possible even in the absence of UvrD.
In this model, UvrD facilitates repair by removing RNAP from the lesion, just as Mfd does, but with opposite polarity and dramatically different consequences for transcription. Mfd TCR entails RNAP release, leaving behind the naked DNA, whereas UvrD TCR would produce an extensively backtracked RNAP. This arrested complex could be reactivated by GreA/B-assisted cleavage or released by Mfd or Rho. Triple deletion of greA/B and mfd decreases the sensitivity of uvrD+ cells to DNA damage (33), suggesting that Rho-mediated termination could be essential (142). This hypothesis raises the question of UvrD interplay with NusA and NusG, which bind to the β flap (49) and nontemplate DNA (125), respectively, and are associated with transcribing RNAP throughout the genome (83). NusG is required for efficient Rho-dependent termination (16), but it could likely act on TEC only after UvrD has left. NusA cooperates with UvrD in vitro (33), suggesting that the two proteins could cooperate in Mfd-independent repair pathway (21), but both proteins interact with the β flap. Unraveling the details of the two-step action of UvrD in NER awaits extensive functional analysis and would be greatly facilitated by the identification of UvrD variants in which only one of its activities is compromised.
TRANSCRIPTION AND REPLICATION
Bacterial RNAP moves along the template 20-fold slower than the replisome, and sometimes in the opposite direction (81). Thus, the conflicts between the replication and transcription complexes are inevitable, particularly in actively growing cells, in which the number of actively transcribing RNAPs exceeds that of moving replisomes by at least two orders of magnitude, and in heavily transcribed regions. Although unusual DNA structures and DNA-bound proteins can impede the replisome, TECs present the most formidable barrier in E. coli (46). Under most conditions, replication is not strongly affected by codirectional transcription, whereas head-on transcription severely inhibits the progression of replication forks in vivo (37), most likely via direct physical interaction with the TEC (80). This bias is reflected in genome organization, with highly expressed, long, and essential genes preferentially located on the leading strand (106). Inversion of rRNA loci leads to disruption of DNA replication, loss of genome integrity, and cell death, underscoring the importance of co-orientation of replication and transcription (120).
These conflicts are exacerbated when RNAP encounters a roadblock. Although progression of both polymerases is blocked by the same types of DNA lesions (126), transcription is also slowed down by sequence-specific interactions of RNAP (11, 64) or transcription factors (5, 105, 147) with nucleic acid chains, nascent RNA hairpins (18), or DNA-bound proteins (45). Many of these delays are temporary and underlie regulatory decisions by RNAP (68), but they can also lead to backtracking that generates arrested complexes (88). Whereas an actively transcribing RNAP can be readily bypassed by a replisome moving in the same direction (100) and, in some cases, even during a head-on collision (30), a stable arrested complex would block replication in both directions. In E. coli, even codirectional collisions may lead to replisome reloading in vitro (100) and double-stranded DNA breaks (DSBs) in vivo (29), events that can trigger mutagenesis and genome instability.
The cell uses several redundant mechanisms to resolve replication-transcription conflicts (29, 123, 132, 142). Preventing backtracking may be the most effective way to inhibit DSB formation upon codirectional collisions. On coding RNAs, ribosomes disfavor RNAP backtracking (102); inhibition of translation increases DSBs (29). In the highly transcribed rRNA operons, the rRNA antitermination complex (127) and cooperating RNAPs (34) likely act jointly to prevent backtracking. Once formed, backtracked complexes can be reactivated by Gre factors (128), giving RNAP another chance to bypass the roadblock, if not permanent. Formation of DSBs is augmented in the absence of greB (29). As a last resort, the stalled RNAP can be removed from the replisome’s path by the enzymatic action of Mfd or Rho (133, 142); sublethal concentrations of the Rho inhibitor bicyclomycin favor DSBs (29), an effect augmented by deletion of mfd (142).
Destabilization of TECs by ppGpp, its cofactor DksA, and some ppGpp-mimicking substitutions in RNAP has been hypothesized to underlie their effects on genome stability (133). This idea is supported by destabilization of promoter complexes (94) and elongating RNAP arrays (132) conferred by these factors. DksA protects cells against UV damage (132) and prevents replication arrest in amino acid–starved cells via its effects on transcription elongation (123). Because DksA (39) and rpo*35 substitution (29) do not affect the stability of isolated TECs, an alternative explanation for their effects could be resistance to backtracking observed in vitro (29, 96). This interpretation is supported by observations that DksA reduces stalling of the elongating RNAP in vivo, particularly in promoter-proximal regions and under conditions that trigger ribosome stalling and uncoupling of transcription and translation (155).
Recent data suggest that NusA may also contribute to transcription-replication conflicts. A screen for conditional suppressors of recG, a DNA translocase that helps maintain genomic integrity, identified rpo alleles that could impair interactions with NusA (77).
None of these mechanisms appears to be absolutely essential. Under optimal growth conditions, a parallel pathway must be compromised to observe the effects of each factor. Gre-deficient cells exhibit more DSBs only when translation or Rho-dependent termination is inhibited (29);mild UV sensitivity of mfd deletion is augmented when greA or dksA is deleted (132); rpo*35 can suppress the lack of ppGpp, dksA, mfd, and greA (132); and mutations in genes affecting replication fork repair and translation increase dependence on Rho (131). However, under environmental conditions encountered by bacteria, the joint action of all partners could be required for survival.
While active transcription may provoke genome instability, it could also accelerate evolution. Transcription generates R-loops (extended RNA-DNA hybrids), which can prime replication forks; formation of excessive R-loops upon RNAP backtracking is lethal, and their control is an essential function of E. coli Rho (73). However, R-loops can serve as intermediates in stress-induced mutagenesis and gene amplification, targeting rearrangements to transcribed regions and thus promoting adaptation (145). Transcription-associated mutagenesis, in which mutations preferentially arise in the exposed nontemplate strand of a transcribed gene, has been argued to foster survival during stress by accelerating adaptation (60). Even in organisms with strong strand bias, many essential genes are located in the lagging strand. In these genes, nonsynonymous mutations accumulate at a faster rate and are positively correlated with gene length and transcriptional activity (95), suggesting that head-on collisions promote evolution of specific genes.
DISTRIBUTION OF RNAP AND ACCESSORY FACTORS
In the cell, RNAP moves along the template bound by architectural proteins, synthesizing RNAs that become targets for ribosomes, RNA-processing enzymes, and RNA-binding proteins. Interactions of RNAP with factors that organize, replicate, and repair DNA and process the newly synthesized RNA are complex, and their regulatory outcomes cannot always be anticipated from in vitro experiments. Although defined coupled systems provide important insights into RNAP cross talk with other processes (17, 79), they cannot identify new players or include all cellular components, necessitating the use of genome-wide analyses. Recent methodological developments have enabled monitoring of transcription in a cellular context with precision and scale unimaginable just a few years ago. Chromatin immunoprecipitation in conjunction with DNA microarray analysis (ChIp-on-chip) or high-throughput sequencing (ChIp-seq) maps positions of RNAP and transcription factors along the genomic DNA at a resolution of approximately 25 bp (58, 83, 136). Nascent transcript sequencing (NET-seq), although limited to transcribing RNAPs, visualizes RNAP position at single-nucleotide resolution (71, 138). Finally, super-resolution microscopy reveals the overall three-dimensional organization of transcription, translation, and replication systems within the bacterial cell (8, 15,31, 108, 141).
These studies produced many expected results. RNAP occupancies are highest at highly expressed genes and steadily decrease toward the 3′ end (83); this decline may be attributed to premature termination by Rho or to RNAP acceleration toward the end of genes. The σ factor is replaced by NusA soon after promoter escape, as shown in vitro (42). Nus factors are recruited to the 5′ region of rRNA operons and remain associated with RNAP throughout transcription (83). NusB is selectively enriched on rRNA operons (13), consistent with its specialized role in rRNA synthesis; NusE is associated with both protein-coding and rRNA operons, presumably as part of the ribosome and the NusABEG complex, respectively. NusG is present on all genes, except those regulated by its paralog RfaH (9).
New, sometimes perplexing, data have also emerged. Many transcription units are characterized by large promoter-proximal peaks of RNAP in E. coli (83) and Synechococcus elongatus (136), but not in B. subtilis (58). These peaks contain Rho and NusA, representing TECs that escaped the promoter, and presumably result from inefficient elongation prior to the recruitment of the leading ribosome. Consistent with their ability to rescue stalled RNAPs, DksA and GreA associate with elongating RNAP and reduce promoter-proximal accumulation of RNAP (67, 154). Curiously, DksA appears to be excluded from structural RNA genes, whereas GreA is more evenly distributed.
E. coli NusG acts on short RNAs in vitro (4) but associates with protein-coding genes slowly in vivo (83). This delay may signify the requirement for simultaneous binding to RNAP and the leading ribosome or preferential recruitment to specific sequences in the nontemplate DNA, as observed for RfaH (9); sequence preference has been reported for B. subtilis NusG (147). An increase in NusG occupancy toward the end of the genes, compared with its presumably stoichiometric levels on rRNA genes (83), suggests that NusG could be present in several copies, perhaps recruited to RNA by NusBE or uncoupled ribosomes.
NET-seq data revealed that, in addition to frequent pausing along the genes (71, 138), E. coli and B. subtilis RNAPs ubiquitously pause at translation initiation codons provided that the Shine-Dalgarno sequence is also present approximately 10 bp upstream (71). The paused RNAP covers the Shine-Dalgarno sequence and initiating codon, preventing inhibitory RNA structures from forming but also blocking the ribosome binding. Initial NusG-NusE interaction may position the ribosome near the RNA, facilitating the formation of the translation initiation complex following the escape from the pause.
Rho appears to bind RNAP soon after promoter escape and closely follows the RNAP occupancy profile on both protein-coding and structural RNA operons (83). This finding is consistent with the proposal that Rho binds to RNAP (32) but not with the presumption that structured RNAs can resist Rho (104). Rho-dependent termination of tRNA genes (98) further supports this pattern. In E. coli, inhibition of Rho derepresses synthesis of many RNAs, notably surprisingly abundant antisense transcripts (97, 136), but also leads to the depletion of H-NS across the genome (19). The latter effect could trigger large-scale chromosomal rearrangements (141) and contribute to silencing of foreign DNA by Rho (16), because H-NS is also a potent silencer (1).
Super-resolution imaging revealed that in fast-growing cells, where the potential for transcription-replication conflicts is highest, the two machineries are spatially separated, with most RNA synthesis confined to a few foci (15, 31), in which RNAP is bound to NusA and NusB (15). Division of the chromosome into functional clusters would help prevent damaging collisions between the replisome and RNAP. The transcription and translation machineries are also spatially segregated (8), but localization of isolated ribosome subunits in the nucleoid (108) is consistent with the cotranscriptional pioneer round of translation.
CONCLUSIONS
In the last decade, we have witnessed impressive progress in studies of bacterial transcription. Complementary approaches, ranging from genomics to single-molecule fluorescence, revealed fine details of the molecular mechanism and regulation of RNAP at all stages of transcription and produced a comprehensive, even though very complex, view of the transcription cycle. Many key questions have been answered, and others will likely be addressed soon. However, the forthcoming answers may not be universal, particularly with respect to regulatory features. Although all bacterial RNAPs share the catalytic core and overall structure, they have many species-specific domains that serve as targets for accessory factors (69). In addition, requirements for transcription regulators vary among phyla (28, 40, 57) with some diversity, such as the absence of Gre factor homologues in Cyanobacteria, Dictyoglomi, Fusobacteria, and Aquificae, only evident from genome databases. Historically, most functional studies were carried out with just one RNAP, that from E. coli, with only a few focusing on B. subtilis. Even a cursory comparison between these two model systems revealed differential response to nucleic acid signals, NusG, and Rho (6, 57, 147), as well as the different distribution of RNAP along the transcription units (58, 83). A recent study revealed that Mycobacterium tuberculosis RNAP recognizes noncanonical intrinsic termination signals, which are ignored by the E. coli RNAP, and that RNA release is strongly potentiated by NusG, an effect opposite of that seen in E. coli (23). These examples demonstrate that even the mechanisms that are thought to be universal need not be and question the utility of functional predictions based on the results obtained in model systems. The astonishing diversity of bacterial species and the dearth of data on transcriptional regulation therein all but guarantee that studies of other divergent RNAP systems will bring new insights into their transcription control strategies.
ACKNOWLEDGMENTS
The work was supported by Academy of Finland grants 130581 and 263713 to GAB and NIH grant GM067153 to IA.
Footnotes
DISCLOSURE STATEMENT
The authors are not aware of any affiliations, memberships, funding, or financial holdings that might be perceived as affecting the objectivity of this review.
LITERATURE CITED
- 1.Ali SS, Xia B, Liu J, Navarre WW. Silencing of foreign DNA in bacteria. Curr. Opin. Microbiol. 2012;15:175–181. doi: 10.1016/j.mib.2011.12.014. [DOI] [PubMed] [Google Scholar]
- 2.Andrecka J, Treutlein B, Arcusa MA, Muschielok A, Lewis R, et al. Nano positioning system reveals the course of upstream and nontemplate DNA within the RNA polymerase II elongation complex. Nucleic Acids Res. 2009;37:5803–5809. doi: 10.1093/nar/gkp601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Arimbasseri AG, Kassavetis GA, Maraia RJ. Transcription. Comment on “Mechanism of eukaryotic RNA polymerase III transcription termination”. Science. 2014;345:524. doi: 10.1126/science.1253783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Artsimovitch I, Landick R. Pausing by bacterial RNA polymerase is mediated by mechanistically distinct classes of signals. PNAS. 2000;97:7090–7095. doi: 10.1073/pnas.97.13.7090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Artsimovitch I, Landick R. The transcriptional regulator RfaH stimulates RNA chain synthesis after recruitment to elongation complexes by the exposed nontemplate DNA strand. Cell. 2002;109:193–203. doi: 10.1016/s0092-8674(02)00724-9. [DOI] [PubMed] [Google Scholar]
- 6.Artsimovitch I, Svetlov V, Anthony L, Burgess RR, Landick R. RNA polymerases from Bacillus subtilis and Escherichia coli differ in recognition of regulatory signals in vitro. J. Bacteriol. 2000;182:6027–6035. doi: 10.1128/jb.182.21.6027-6035.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bae W, Xia B, Inouye M, Severinov K. Escherichia coli CspA-family RNA chaperones are transcription antiterminators. PNAS. 2000;97:7784–7789. doi: 10.1073/pnas.97.14.7784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bakshi S, Siryaporn A, Goulian M, Weisshaar JC. Superresolution imaging of ribosomes and RNA polymerase in live Escherichia coli cells. Mol. Microbiol. 2012;85:21–38. doi: 10.1111/j.1365-2958.2012.08081.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Belogurov GA, Mooney RA, Svetlov V, Landick R, Artsimovitch I. Functional specialization of transcription elongation factors. EMBO J. 2009;28:112–122. doi: 10.1038/emboj.2008.268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Belogurov GA, Vassylyeva MN, Svetlov V, Klyuyev S, Grishin NV, et al. Structural basis for converting a general transcription factor into an operon-specific virulence regulator. Mol. Cell. 2007;26:117–129. doi: 10.1016/j.molcel.2007.02.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bochkareva A, Yuzenkova Y, Tadigotla VR, Zenkin N. Factor-independent transcription pausing caused by recognition of the RNA-DNA hybrid sequence. EMBO J. 2012;31:630–639. doi: 10.1038/emboj.2011.432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bubunenko M, Court DL, Al Refaii A, Saxena S, Korepanov A, et al. Nus transcription elongation factors and RNase III modulate small ribosome subunit biogenesis in Escherichia coli. Mol. Microbiol. 2013;87:382–393. doi: 10.1111/mmi.12105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Burmann BM, Knauer SH, Sevostyanova A, Schweimer K, Mooney RA, et al. An α helix to β barrel domain switch transforms the transcription factor RfaH into a translation factor. Cell. 2012;150:291–303. doi: 10.1016/j.cell.2012.05.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Burmann BM, Schweimer K, Luo X, Wahl MC, Stitt BL, et al. A NusE:NusG complex links transcription and translation. Science. 2010;328:501–504. doi: 10.1126/science.1184953. [DOI] [PubMed] [Google Scholar]
- 15.Cagliero C, Zhou YN, Jin DJ. Spatial organization of transcription machinery and its segregation from the replisome in fast-growing bacterial cells. Nucleic Acids Res. 2014;42:13696–13705. doi: 10.1093/nar/gku1103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cardinale CJ, Washburn RS, Tadigotla VR, Brown LM, Gottesman ME, Nudler E. Termination factor Rho and its cofactors NusA and NusG silence foreign DNA in E. coli. Science. 2008;320:935–938. doi: 10.1126/science.1152763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Castro-Roa D, Zenkin N. In vitro experimental system for analysis of transcription-translation coupling. Nucleic Acids Res. 2012;40:e45. doi: 10.1093/nar/gkr1262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chan CL, Wang D, Landick R. Multiple interactions stabilize a single paused transcription intermediate in which hairpin to 3′ end spacing distinguishes pause and termination pathways. J. Mol. Biol. 1997;268:54–68. doi: 10.1006/jmbi.1997.0935. [DOI] [PubMed] [Google Scholar]
- 19.Chandraprakash D, Seshasayee AS. Inhibition of factor-dependent transcription termination in Escherichia coli might relieve xenogene silencing by abrogating H-NS-DNA interactions in vivo. J. Biosci. 2014;39:53–61. doi: 10.1007/s12038-014-9413-4. [DOI] [PubMed] [Google Scholar]
- 20.Cohen SE, Godoy VG, Walker GC. Transcriptional modulator NusA interacts with translesion DNA polymerases in Escherichia coli. J. Bacteriol. 2009;191:665–672. doi: 10.1128/JB.00941-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cohen SE, Lewis CA, Mooney RA, Kohanski MA, Collins JJ, et al. Roles for the transcription elongation factor NusA in both DNA repair and damage tolerance pathways in Escherichia coli. PNAS. 2010;107:15517–15522. doi: 10.1073/pnas.1005203107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cohen SE, Walker GC. The transcription elongation factor NusA is required for stress-induced mutagenesis in Escherichia coli. Curr. Biol. 2010;20:80–85. doi: 10.1016/j.cub.2009.11.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Czyz A, Mooney RA, Iaconi A, Landick R. Mycobacterial RNA polymerase requires a U-tract at intrinsic terminators and is aided by NusG at suboptimal terminators. mBio. 2014;5:e00931. doi: 10.1128/mBio.00931-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dangkulwanich M, Ishibashi T, Liu S, Kireeva ML, Lubkowska L, et al. Complete dissection of transcription elongation reveals slow translocation of RNA polymerase II in a linear ratchet mechanism. Elife. 2013;2:e00971. doi: 10.7554/eLife.00971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Deaconescu AM, Artsimovitch I, Grigorieff N. Interplay of DNA repair with transcription: from structures to mechanisms. Trends Biochem. Sci. 2012;37:543–552. doi: 10.1016/j.tibs.2012.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Deaconescu AM, Chambers AL, Smith AJ, Nickels BE, Hochschild A, et al. Structural basis for bacterial transcription-coupled DNA repair. Cell. 2006;124:507–520. doi: 10.1016/j.cell.2005.11.045. [DOI] [PubMed] [Google Scholar]
- 27.Deaconescu AM, Sevostyanova A, Artsimovitch I, Grigorieff N. Nucleotide excision repair (NER) machinery recruitment by the transcription-repair coupling factor involves unmasking of a conserved intramolecular interface. PNAS. 2012;109:3353–3358. doi: 10.1073/pnas.1115105109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.D’Heygere F, Rabhi M, Boudvillain M. Phyletic distribution and conservation of the bacterial transcription termination factor Rho. Microbiology. 2013;159:1423–1436. doi: 10.1099/mic.0.067462-0. [DOI] [PubMed] [Google Scholar]
- 29.Dutta D, Shatalin K, Epshtein V, Gottesman ME, Nudler E. Linking RNA polymerase backtracking to genome instability in E. coli. Cell. 2011;146:533–543. doi: 10.1016/j.cell.2011.07.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Elias-Arnanz M, Salas M. Resolution of head-on collisions between the transcription machinery and bacteriophage phi29 DNA polymerase is dependent on RNA polymerase translocation. EMBO J. 1999;18:5675–5682. doi: 10.1093/emboj/18.20.5675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Endesfelder U, Finan K, Holden SJ, Cook PR, Kapanidis AN, Heilemann M. Multiscale spatial organization of RNA polymerase in Escherichia coli. Biophys. J. 2013;105:172–181. doi: 10.1016/j.bpj.2013.05.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Epshtein V, Dutta D, Wade J, Nudler E. An allosteric mechanism of Rho-dependent transcription termination. Nature. 2010;463:245–249. doi: 10.1038/nature08669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Epshtein V, Kamarthapu V, McGary K, Svetlov V, Ueberheide B, et al. UvrD facilitates DNA repair by pulling RNA polymerase backwards. Nature. 2014;505:372–377. doi: 10.1038/nature12928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Epshtein V, Nudler E. Cooperation between RNA polymerase molecules in transcription elongation. Science. 2003;300:801–805. doi: 10.1126/science.1083219. [DOI] [PubMed] [Google Scholar]
- 35.Farnham PJ, Greenblatt J, Platt T. Effects of NusA protein on transcription termination in the tryptophan operon of Escherichia coli. Cell. 1982;29:945–951. doi: 10.1016/0092-8674(82)90457-3. [DOI] [PubMed] [Google Scholar]
- 36.Feig M, Burton ZF. RNA polymerase II with open and closed trigger loops: active site dynamics and nucleic acid translocation. Biophys. J. 2010;99:2577–2586. doi: 10.1016/j.bpj.2010.08.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.French S. Consequences of replication fork movement through transcription units in vivo. Science. 1992;258:1362–1365. doi: 10.1126/science.1455232. [DOI] [PubMed] [Google Scholar]
- 38.Fuchs RP, Fujii S. Translesion DNA synthesis and mutagenesis in prokaryotes. Cold Spring Harb. Perspect. Biol. 2013;5:a012682. doi: 10.1101/cshperspect.a012682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Furman R, Sevostyanova A, Artsimovitch I. Transcription initiation factor DksA has diverse effects on RNA chain elongation. Nucleic Acids Res. 2012;40:3392–3402. doi: 10.1093/nar/gkr1273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Furman R, Tsodikov OV, Wolf YI, Artsimovitch I. An insertion in the catalytic trigger loop gates the secondary channel of RNA polymerase. J. Mol. Biol. 2013;425:82–93. doi: 10.1016/j.jmb.2012.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ganesan A, Spivak G, Hanawalt PC. Transcription-coupled DNA repair in prokaryotes. Prog. Mol. Biol. Transl. Sci. 2012;110:25–40. doi: 10.1016/B978-0-12-387665-2.00002-X. [DOI] [PubMed] [Google Scholar]
- 42.Gill SC, Weitzel SE, von Hippel PH. Escherichia coli σ70 and NusA proteins. I. Binding interactions with core RNA polymerase in solution and within the transcription complex. J. Mol. Biol. 1991;220:307–324. doi: 10.1016/0022-2836(91)90015-x. [DOI] [PubMed] [Google Scholar]
- 43.Gopal B, Haire LF, Gamblin SJ, Dodson EJ, Lane AN, et al. Crystal structure of the transcription elongation/anti-termination factor NusA from Mycobacterium tuberculosis at 1.7 Å resolution. J. Mol. Biol. 2001;314:1087–1095. doi: 10.1006/jmbi.2000.5144. [DOI] [PubMed] [Google Scholar]
- 44.Greive SJ, Lins AF, von Hippel PH. Assembly of an RNA-protein complex. Binding of NusB and NusE (S10) proteins to boxA RNA nucleates the formation of the antitermination complex involved in controlling rRNA transcription in Escherichia coli. J. Biol. Chem. 2005;280:36397–3408. doi: 10.1074/jbc.M507146200. [DOI] [PubMed] [Google Scholar]
- 45.Guerin M, Leng M, Rahmouni AR. High resolution mapping of E. coli transcription elongation complex in situ reveals protein interactions with the non-transcribed strand. EMBO J. 1996;15:5397–5407. [PMC free article] [PubMed] [Google Scholar]
- 46.Gupta MK, Guy CP, Yeeles JT, Atkinson J, Bell H, et al. Protein-DNA complexes are the primary sources of replication fork pausing in Escherichia coli. PNAS. 2013;110:7252–7257. doi: 10.1073/pnas.1303890110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gusarov I, Nudler E. Control of intrinsic transcription termination by N and NusA: the basic mechanisms. Cell. 2001;107:437–449. doi: 10.1016/s0092-8674(01)00582-7. [DOI] [PubMed] [Google Scholar]
- 48.Gwynn EJ, Smith AJ, Guy CP, Savery NJ, McGlynn P, Dillingham MS. The conserved C-terminus of the PcrA/UvrD helicase interacts directly with RNA polymerase. PLOS ONE. 2013;8:e78141. doi: 10.1371/journal.pone.0078141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ha KS, Toulokhonov I, Vassylyev DG, Landick R. The NusA N-terminal domain is necessary and sufficient for enhancement of transcriptional pausing via interaction with the RNA exit channel of RNA polymerase. J. Mol. Biol. 2010;401:708–725. doi: 10.1016/j.jmb.2010.06.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Haines NM, Kim YI, Smith AJ, Savery NJ. Stalled transcription complexes promote DNA repair at a distance. PNAS. 2014;111:4037–4042. doi: 10.1073/pnas.1322350111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hein PP, Kolb KE, Windgassen T, Bellecourt MJ, Darst SA, et al. RNA polymerase pausing and nascent-RNA structure formation are linked through clamp-domain movement. Nat. Struct. Mol. Biol. 2014;21:794–802. doi: 10.1038/nsmb.2867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Hein PP, Landick R. The bridge helix coordinates movements of modules in RNA polymerase. BMC Biol. 2010;8:141. doi: 10.1186/1741-7007-8-141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hein PP, Palangat M, Landick R. RNA transcript 3′-proximal sequence affects translocation bias of RNA polymerase. Biochemistry. 2011;50:7002–7014. doi: 10.1021/bi200437q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Henkin TM, Yanofsky C. Regulation by transcription attenuation in bacteria: how RNA provides instructions for transcription termination/antitermination decisions. Bioessays. 2002;24:700–707. doi: 10.1002/bies.10125. [DOI] [PubMed] [Google Scholar]
- 55.Herbert KM, Zhou J, Mooney RA, Porta AL, Landick R, Block SM. E. coli NusG inhibits backtracking and accelerates pause-free transcription by promoting forward translocation of RNA polymerase. J. Mol. Biol. 2010;399:17–30. doi: 10.1016/j.jmb.2010.03.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Howan K, Smith AJ, Westblade LF, Joly N, Grange W, et al. Initiation of transcription-coupled repair characterized at single-molecule resolution. Nature. 2012;490:431–434. doi: 10.1038/nature11430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Ingham CJ, Dennis J, Furneaux PA. Autogenous regulation of transcription termination factor Rho and the requirement for Nus factors in Bacillus subtilis. Mol. Microbiol. 1999;31:651–663. doi: 10.1046/j.1365-2958.1999.01205.x. [DOI] [PubMed] [Google Scholar]
- 58.Ishikawa S, Oshima T, Kurokawa K, Kusuya Y, Ogasawara N. RNA polymerase trafficking in Bacillus subtilis cells. J. Bacteriol. 2010;192:5778–5787. doi: 10.1128/JB.00489-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Jiang W, Hou Y, Inouye M. CspA, the major cold-shock protein of Escherichia coli is an RNA chaperone. J. Biol. Chem. 1997;272:196–202. doi: 10.1074/jbc.272.1.196. [DOI] [PubMed] [Google Scholar]
- 60.Jinks-Robertson S, Bhagwat AS. Transcription-associated mutagenesis. Annu. Rev. Genet. 2014;48:341–359. doi: 10.1146/annurev-genet-120213-092015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Kaplan CD, Larsson K-M, Kornberg RD. The RNA polymerase II trigger loop functions in substrate selection and is directly targeted by α-amanitin. Mol. Cell. 2008;30:547–556. doi: 10.1016/j.molcel.2008.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Kireeva M, Kashlev M, Burton ZF. Translocation by multi-subunit RNA polymerases. Biochim. Biophys. Acta. 2010;1799:389–401. doi: 10.1016/j.bbagrm.2010.01.007. [DOI] [PubMed] [Google Scholar]
- 63.Kireeva ML, Domecq C, Coulombe B, Burton ZF, Kashlev M. Interaction of RNA polymerase II fork loop 2 with downstream non-template DNA regulates transcription elongation. J. Biol. Chem. 2011;286:30898–30910. doi: 10.1074/jbc.M111.260844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kireeva ML, Kashlev M. Mechanism of sequence-specific pausing of bacterial RNA polymerase. PNAS. 2009;106:8900–8905. doi: 10.1073/pnas.0900407106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kireeva ML, Nedialkov YA, Cremona GH, Purtov YA, Lubkowska L, et al. Transient reversal of RNA polymerase II active site closing controls fidelity of transcription elongation. Mol. Cell. 2008;30:557–566. doi: 10.1016/j.molcel.2008.04.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Komissarova N, Kashlev M. Transcriptional arrest: Escherichia coli RNA polymerase translocates backward, leaving the 3′ end of the RNA intact and extruded. PNAS. 1997;94:1755–1760. doi: 10.1073/pnas.94.5.1755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Kusuya Y, Kurokawa K, Ishikawa S, Ogasawara N, Oshima T. Transcription factor GreA contributes to resolving promoter-proximal pausing of RNA polymerase in Bacillus subtilis cells. J. Bacteriol. 2011;193:3090–3099. doi: 10.1128/JB.00086-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Landick R. The regulatory roles and mechanism of transcriptional pausing. Biochem. Soc. Trans. 2006;34:1062–1066. doi: 10.1042/BST0341062. [DOI] [PubMed] [Google Scholar]
- 69.Lane WJ, Darst SA. Molecular evolution of multisubunit RNA polymerases: sequence analysis. J. Mol. Biol. 2010;395:671–685. doi: 10.1016/j.jmb.2009.10.062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Laptenko O, Lee J, Lomakin I, Borukhov S. Transcript cleavage factors GreA and GreB act as transient catalytic components of RNA polymerase. EMBO J. 2003;22:6322–6334. doi: 10.1093/emboj/cdg610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Larson MH, Mooney RA, Peters JM, Windgassen T, Nayak D, et al. A pause sequence enriched at translation start sites drives transcription dynamics in vivo. Science. 2014;344:1042–1047. doi: 10.1126/science.1251871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Larson MH, Zhou J, Kaplan CD, Palangat M, Kornberg RD, et al. Trigger loop dynamics mediate the balance between the transcriptional fidelity and speed of RNA polymerase II. PNAS. 2012;109:6555–6560. doi: 10.1073/pnas.1200939109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Leela JK, Syeda AH, Anupama K, Gowrishankar J. Rho-dependent transcription termination is essential to prevent excessive genome-wide R-loops in Escherichia coli. PNAS. 2013;110:258–263. doi: 10.1073/pnas.1213123110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Li GW, Burkhardt D, Gross C, Weissman JS. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell. 2014;157:624–635. doi: 10.1016/j.cell.2014.02.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Luo X, Hsiao HH, Bubunenko M, Weber G, Court DL, et al. Structural and functional analysis of the E. coli NusB-S10 transcription antitermination complex. Mol. Cell. 2008;32:791–802. doi: 10.1016/j.molcel.2008.10.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Mah TF, Kuznedelov K, Mushegian A, Severinov K, Greenblatt J. The αsubunit of E. coli RNA polymerase activates RNA binding by NusA. Genes Dev. 2000;14:2664–2675. doi: 10.1101/gad.822900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Mahdi AA, Briggs GS, Lloyd RG. Modulation of DNA damage tolerance in Escherichia coli recG and ruv strains by mutations affecting PriB, the ribosome and RNA polymerase. Mol. Microbiol. 2012;86:675–691. doi: 10.1111/mmi.12010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Malinen AM, Turtola M, Parthiban M, Vainonen L, Johnson MS, Belogurov GA. Active site opening and closure control translocation of multisubunit RNA polymerase. Nucleic Acids Res. 2012;40:7442–7451. doi: 10.1093/nar/gks383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Manelyte L, Kim YI, Smith AJ, Smith RM, Savery NJ. Regulation and rate enhancement during transcription-coupled DNA repair. Mol. Cell. 2010;40:714–724. doi: 10.1016/j.molcel.2010.11.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Mirkin EV, Mirkin SM. Mechanisms of transcription-replication collisions in bacteria. Mol. Cell. Biol. 2005;25:888–895. doi: 10.1128/MCB.25.3.888-895.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Mirkin EV, Mirkin SM. Replication fork stalling at natural impediments. Microbiol. Mol. Biol. Rev. 2007;71:13–35. doi: 10.1128/MMBR.00030-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Miropolskaya N, Esyunina D, Klimasauskas S, Nikiforov V, Artsimovitch I, Kulbachinskiy A. Interplay between the trigger loop and the F loop during RNA polymerase catalysis. Nucleic Acids Res. 2014;42:544–552. doi: 10.1093/nar/gkt877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Mooney RA, Davis SE, Peters JM, Rowland JL, Ansari AZ, Landick R. Regulator trafficking on bacterial transcription units in vivo. Mol. Cell. 2009;33:97–108. doi: 10.1016/j.molcel.2008.12.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Mooney RA, Schweimer K, Rosch P, Gottesman M, Landick R. Two structurally independent domains of E. coli NusG create regulatory plasticity via distinct interactions with RNA polymerase and regulators. J. Mol. Biol. 2009;391:341–358. doi: 10.1016/j.jmb.2009.05.078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Murphy MN, Gong P, Ralto K, Manelyte L, Savery NJ, Theis K. An N-terminal clamp restrains the motor domains of the bacterial transcription-repair coupling factor Mfd. Nucleic Acids Res. 2009;37:6042–6053. doi: 10.1093/nar/gkp680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Nedialkov YA, Nudler E, Burton ZF. RNA polymerase stalls in a post-translocated register and can hyper-translocate. Transcription. 2012;3:260–269. doi: 10.4161/trns.22307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Neuman KC, Abbondanzieri EA, Landick R, Gelles J, Block SM. Ubiquitous transcriptional pausing is independent of RNA polymerase backtracking. Cell. 2003;115:437–447. doi: 10.1016/s0092-8674(03)00845-6. [DOI] [PubMed] [Google Scholar]
- 88.Nudler E. RNA polymerase backtracking in gene regulation and genome instability. Cell. 2012;149:1438–1445. doi: 10.1016/j.cell.2012.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Ona KR, Courcelle CT, Courcelle J. Nucleotide excision repair is a predominant mechanism for processing nitrofurazone-induced DNA damage in Escherichia coli. J. Bacteriol. 2009;191:4959–4965. doi: 10.1128/JB.00495-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Opalka N, Chlenov M, Chacon P, Rice WJ, Wriggers W, Darst SA. Structure and function of the transcription elongation factor GreB bound to bacterial RNA polymerase. Cell. 2003;114:335–345. doi: 10.1016/s0092-8674(03)00600-7. [DOI] [PubMed] [Google Scholar]
- 91.Pan T, Artsimovitch I, Fang XW, Landick R, Sosnick TR. Folding of a large ribozyme during transcription and the effect of the elongation factor NusA. PNAS. 1999;96:9545–9550. doi: 10.1073/pnas.96.17.9545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Park JS, Marr MT, Roberts JW. E. coli transcription repair coupling factor (Mfd protein) rescues arrested complexes by promoting forward translocation. Cell. 2002;109:757–767. doi: 10.1016/s0092-8674(02)00769-9. [DOI] [PubMed] [Google Scholar]
- 93.Park JS, Roberts JW. Role of DNA bubble rewinding in enzymatic transcription termination. PNAS. 2006;103:4870–4875. doi: 10.1073/pnas.0600145103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Paul BJ, Ross W, Gaal T, Gourse RL. rRNA transcription in Escherichia coli. Annu. Rev. Genet. 2004;38:749–770. doi: 10.1146/annurev.genet.38.072902.091347. [DOI] [PubMed] [Google Scholar]
- 95.Paul S, Million-Weaver S, Chattopadhyay S, Sokurenko E, Merrikh H. Accelerated gene evolution through replication-transcription conflicts. Nature. 2013;495:512–515. doi: 10.1038/nature11989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Perederina A, Svetlov V, Vassylyeva MN, Tahirov TH, Yokoyama S, et al. Regulation through the secondary channel–structural framework for ppGpp-DksA synergism during transcription. Cell. 2004;118:297–309. doi: 10.1016/j.cell.2004.06.030. [DOI] [PubMed] [Google Scholar]
- 97.Peters JM, Mooney RA, Grass JA, Jessen ED, Tran F, Landick R. Rho and NusG suppress pervasive antisense transcription in Escherichia coli. Genes Dev. 2012;26:2621–2633. doi: 10.1101/gad.196741.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Peters JM, Mooney RA, Kuan PF, Rowland JL, Keles S, Landick R. Rho directs widespread termination of intragenic and stable RNA transcription. PNAS. 2009;106:15406–15411. doi: 10.1073/pnas.0903846106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Peters JM, Vangeloff AD, Landick R. Bacterial transcription terminators: the RNA 3′-end chronicles. J. Mol. Biol. 2011;412:793–813. doi: 10.1016/j.jmb.2011.03.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Pomerantz RT, O’Donnell M. The replisome uses mRNA as a primer after colliding with RNA polymerase. Nature. 2008;456:762–766. doi: 10.1038/nature07527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Prasch S, Jurk M, Washburn RS, Gottesman ME, Wohrl BM, Rosch P. RNA-binding specificity of E. coli NusA. Nucleic Acids Res. 2009;37:4736–4742. doi: 10.1093/nar/gkp452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Proshkin S, Rahmouni AR, Mironov A, Nudler E. Cooperation between translating ribosomes and RNA polymerase in transcription elongation. Science. 2010;328:504–508. doi: 10.1126/science.1184939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Rabhi M, Espéli O, Schwartz A, Cayrol B, Rahmouni AR, et al. The Sm-like RNA chaperone Hfq mediates transcription antitermination at Rho-dependent terminators. EMBO J. 2011;30:2805–2816. doi: 10.1038/emboj.2011.192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Richardson JP. Rho-dependent termination and ATPases in transcript termination. Biochim. Biophys. Acta. 2002;1577:251–260. doi: 10.1016/s0167-4781(02)00456-6. [DOI] [PubMed] [Google Scholar]
- 105.Ring BZ, Yarnell WS, Roberts JW. Function of E. coli RNA polymerase σfactor-σ70 in promoter-proximal pausing. Cell. 1996;86:485–493. doi: 10.1016/s0092-8674(00)80121-x. [DOI] [PubMed] [Google Scholar]
- 106.Rocha EP. The organization of the bacterial genome. Annu. Rev. Genet. 2008;42:211–233. doi: 10.1146/annurev.genet.42.110807.091653. [DOI] [PubMed] [Google Scholar]
- 107.Ross W, Schneider DA, Paul BJ, Mertens A, Gourse RL. An intersubunit contact stimulating transcription initiation by E. coli RNA polymerase: interaction of the αC-terminal domain and ζregion 4. Genes Dev. 2003;17:1293–1307. doi: 10.1101/gad.1079403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Sanamrad A, Persson F, Lundius EG, Fange D, Gynna AH, Elf J. Single-particle tracking reveals that free ribosomal subunits are not excluded from the Escherichia coli nucleoid. PNAS. 2014;111:11413–11418. doi: 10.1073/pnas.1411558111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Santangelo TJ, Cubonova L, Skinner KM, Reeve JN. Archaeal intrinsic transcription termination in vivo. J. Bacteriol. 2009;191:7102–7108. doi: 10.1128/JB.00982-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Savery N. Prioritizing the repair of DNA damage that is encountered by RNA polymerase. Transcription. 2011;2:168–172. doi: 10.4161/trns.2.4.16146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Selby CP, Sancar A. Molecular mechanism of transcription-repair coupling. Science. 1993;260:53–58. doi: 10.1126/science.8465200. [DOI] [PubMed] [Google Scholar]
- 112.Selby CP, Sancar A. Structure and function of transcription-repair coupling factor. II. Catalytic properties. J. Biol. Chem. 1995;270:4890–4895. doi: 10.1074/jbc.270.9.4890. [DOI] [PubMed] [Google Scholar]
- 113.Sevostyanova A, Artsimovitch I. Functional analysis of Thermus thermophilus transcription factor NusG. Nucleic Acids Res. 2010;38:7432–7445. doi: 10.1093/nar/gkq623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Sevostyanova A, Belogurov GA, Mooney RA, Landick R, Artsimovitch I. The βsubunit gate loop is required for RNA polymerase modification by RfaH and NusG. Mol. Cell. 2011;43:253–262. doi: 10.1016/j.molcel.2011.05.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Shankar S, Hatoum A, Roberts JW. A transcription antiterminator constructs a NusA-dependent shield to the emerging transcript. Mol. Cell. 2007;27:914–927. doi: 10.1016/j.molcel.2007.07.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Shibata R, Bessho Y, Shinkai A, Nishimoto M, Fusatomi E, et al. Crystal structure and RNA-binding analysis of the archaeal transcription factor NusA. Biochem. Biophys. Res. Commun. 2007;355:122–128. doi: 10.1016/j.bbrc.2007.01.119. [DOI] [PubMed] [Google Scholar]
- 117.Sigmund CD, Morgan EA. Effects of Escherichia coli Nus A protein on transcription termination in vitro are not increased or decreased by DNA sequences sufficient for antitermination in vivo. Biochemistry. 1988;27:5628–5835. doi: 10.1021/bi00415a035. [DOI] [PubMed] [Google Scholar]
- 118.Smith AJ, Pernstich C, Savery NJ. Multipartite control of the DNA translocase, Mfd. Nucleic Acids Res. 2012;40:10408–10416. doi: 10.1093/nar/gks775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Squires CL, Greenblatt J, Li J, Condon C, Squires CL. Ribosomal RNA antitermination in vitro: requirement for Nus factors and one or more unidentified cellular components. PNAS. 1993;90:970–974. doi: 10.1073/pnas.90.3.970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Srivatsan A, Tehranchi A, MacAlpine DM, Wang JD. Co-orientation of replication and transcription preserves genome integrity. PLOS Genet. 2010;6:e1000810. doi: 10.1371/journal.pgen.1000810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Steitz TA. A mechanism for all polymerases. Nature. 1998;391:231–232. doi: 10.1038/34542. [DOI] [PubMed] [Google Scholar]
- 122.Svetlov V, Vassylyev DG, Artsimovitch I. Discrimination against deoxyribonucleotide substrates by bacterial RNA polymerase. J. Biol. Chem. 2004;279:38087–38090. doi: 10.1074/jbc.C400316200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Tehranchi AK, Blankschien MD, Zhang Y, Halliday JA, Srivatsan A, et al. The transcription factor DksA prevents conflicts between DNA replication and transcription machinery. Cell. 2010;141:595–605. doi: 10.1016/j.cell.2010.03.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Thomson E, Ferreira-Cerca S, Hurt E. Eukaryotic ribosome biogenesis at a glance. J. Cell Sci. 2013;126:4815–4821. doi: 10.1242/jcs.111948. [DOI] [PubMed] [Google Scholar]
- 125.Tomar SK, Artsimovitch I. NusG-Spt5 proteins - universal tools for transcription modification and communication. Chem. Rev. 2013;113:8604–8619. doi: 10.1021/cr400064k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Tornaletti S, Hanawalt PC. Effect of DNA lesions on transcription elongation. Biochimie. 1999;81:139–146. doi: 10.1016/s0300-9084(99)80046-7. [DOI] [PubMed] [Google Scholar]
- 127.Torres M, Condon C, Balada JM, Squires C, Squires CL. Ribosomal protein S4 is a transcription factor with properties remarkably similar to NusA, a protein involved in both non-ribosomal and ribosomal RNA antitermination. EMBO J. 2001;20:3811–3820. doi: 10.1093/emboj/20.14.3811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Toulme F, Mosrin-Huaman C, Sparkowski J, Das A, Leng M, Rahmouni AR. GreA and GreB proteins revive backtracked RNA polymerase in vivo by promoting transcript trimming. EMBO J. 2000;19:6853–6859. doi: 10.1093/emboj/19.24.6853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Toulokhonov I, Artsimovitch I, Landick R. Allosteric control of RNA polymerase by a site that contacts nascent RNA hairpins. Science. 2001;292:730–733. doi: 10.1126/science.1057738. [DOI] [PubMed] [Google Scholar]
- 130.Toulokhonov I, Zhang J, Palangat M, Landick R. A central role of the RNA polymerase trigger loop in active-site rearrangement during transcriptional pausing. Mol. Cell. 2007;27:406–419. doi: 10.1016/j.molcel.2007.06.008. [DOI] [PubMed] [Google Scholar]
- 131.Tran L, van Baarsel JA, Washburn RS, Gottesman ME, Miller JH. Single-gene deletion mutants of-Escherichia coli with altered sensitivity to bicyclomycin, an inhibitor of transcription termination factor Rho. J. Bacteriol. 2011;193:2229–2235. doi: 10.1128/JB.01463-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Trautinger BW, Jaktaji RP, Rusakova E, Lloyd RG. RNA polymerase modulators and DNA repair activities resolve conflicts between DNA replication and transcription. Mol. Cell. 2005;19:247–258. doi: 10.1016/j.molcel.2005.06.004. [DOI] [PubMed] [Google Scholar]
- 133.Trautinger BW, Lloyd RG. Modulation of DNA repair by mutations flanking the DNA channel through RNA polymerase. EMBO J. 2002;21:6944–6953. doi: 10.1093/emboj/cdf654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Vassylyev DG, Vassylyeva MN, Perederina A, Tahirov TH, Artsimovitch I. Structural basis for transcription elongation by bacterial RNA polymerase. Nature. 2007;448:157–162. doi: 10.1038/nature05932. [DOI] [PubMed] [Google Scholar]
- 135.Vassylyev DG, Vassylyeva MN, Zhang J, Palangat M, Artsimovitch I, Landick R. Structural basis for substrate loading in bacterial RNA polymerase. Nature. 2007;448:163–168. doi: 10.1038/nature05931. [DOI] [PubMed] [Google Scholar]
- 136.Vijayan V, Jain IH, O’Shea EK. A high resolution map of a cyanobacterial transcriptome. Genome Biol. 2011;12:R47. doi: 10.1186/gb-2011-12-5-r47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Vrentas CE, Gaal T, Ross W, Ebright RH, Gourse RL. Response of RNA polymerase to ppGpp: requirement for the ωsubunit and relief of this requirement by DksA. Genes Dev. 2005;19:2378–2387. doi: 10.1101/gad.1340305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Vvedenskaya IO, Vahedian-Movahed H, Bird JG, Knoblauch JG, Goldman SR, et al. Transcription. Interactions between RNA polymerase and the “core recognition element” counteract pausing. Science. 2014;344:1285–1289. doi: 10.1126/science.1253458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Wang D, Bushnell DA, Huang X, Westover KD, Levitt M, Kornberg RD. Structural basis of transcription: backtracked RNA polymerase II at 3.4 Å resolution. Science. 2009;324:1203–1206. doi: 10.1126/science.1168729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Wang D, Bushnell DA, Westover KD, Kaplan CD, Kornberg RD. Structural basis of transcription: role of the trigger loop in substrate specificity and catalysis. Cell. 2006;127:941–954. doi: 10.1016/j.cell.2006.11.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Wang W, Li GW, Chen C, Xie XS, Zhuang X. Chromosome organization by a nucleoid-associated protein in live bacteria. Science. 2011;333:1445–1449. doi: 10.1126/science.1204697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Washburn RS, Gottesman ME. Transcription termination maintains chromosome integrity. PNAS. 2011;108:792–797. doi: 10.1073/pnas.1009564108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Weixlbaumer A, Leon K, Landick R, Darst SA. Structural basis of transcriptional pausing in bacteria. Cell. 2013;152:431–441. doi: 10.1016/j.cell.2012.12.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Westblade LF, Campbell EA, Pukhrambam C, Padovan JC, Nickels BE, et al. Structural basis for the bacterial transcription-repair coupling factor/RNA polymerase interaction. Nucleic Acids Res. 2010;38:8357–8369. doi: 10.1093/nar/gkq692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Wimberly H, Shee C, Thornton PC, Sivaramakrishnan P, Rosenberg SM, Hastings PJ. R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli. Nat. Commun. 2013;4:2115. doi: 10.1038/ncomms3115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Worbs M, Bourenkov GP, Bartunik HD, Huber R, Wahl MC. An extended RNA binding surface through arrayed S1 and KH domains in transcription factor NusA. Mol. Cell. 2001;7:1177–1189. doi: 10.1016/s1097-2765(01)00262-3. [DOI] [PubMed] [Google Scholar]
- 147.Yakhnin AV, Babitzke P. Mechanism of NusG-stimulated pausing, hairpin-dependent pause site selection and intrinsic termination at overlapping pause and termination sites in the Bacillus subtilis trp leader. Mol. Microbiol. 2010;76:690–705. doi: 10.1111/j.1365-2958.2010.07126.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Yang W. Lessons learned from UvrD helicase: mechanism for directional movement. Annu. Rev. Biophys. 2010;39:367–385. doi: 10.1146/annurev.biophys.093008.131415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Yang X, Molimau S, Doherty GP, Johnston EB, Marles-Wright J, et al. The structure of bacterial RNA polymerase in complex with the essential transcription elongation factor NusA. EMBO Rep. 2009;10:997–1002. doi: 10.1038/embor.2009.155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Yip WS, Vincent NG, Baserga SJ. Ribonucleoproteins in archaeal pre-rRNA processing and modification. Archaea. 2013;2013:614735. doi: 10.1155/2013/614735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Yuzenkova Y, Zenkin N. Central role of the RNA polymerase trigger loop in intrinsic RNA hydrolysis. PNAS. 2010;107:10878–10883. doi: 10.1073/pnas.0914424107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Zellars M, Squires CL. Antiterminator-dependent modulation of transcription elongation rates by NusB and NusG. Mol. Microbiol. 1999;32:1296–1304. doi: 10.1046/j.1365-2958.1999.01442.x. [DOI] [PubMed] [Google Scholar]
- 153.Zhang G, Campbell EA, Minakhin L, Richter C, Severinov K, Darst SA. Crystal structure of Thermus aquaticus core RNA polymerase at 3.3 Å resolution. Cell. 1999;98:811–824. doi: 10.1016/s0092-8674(00)81515-9. [DOI] [PubMed] [Google Scholar]
- 154.Zhang J, Palangat M, Landick R. Role of the RNA polymerase trigger loop in catalysis and pausing. Nat. Struct. Mol. Biol. 2010;17:99–104. doi: 10.1038/nsmb.1732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Zhang Y, Mooney RA, Grass JA, Sivaramakrishnan P, Herman C, et al. DksA guards elongating RNA polymerase against ribosome-stalling-induced arrest. Mol. Cell. 2014;53:766–778. doi: 10.1016/j.molcel.2014.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Zuo Y, Wang Y, Steitz TA. The mechanism of E. coli RNA polymerase regulation by ppGpp is suggested by the structure of their complex. Mol. Cell. 2013;50:430–436. doi: 10.1016/j.molcel.2013.03.020. [DOI] [PMC free article] [PubMed] [Google Scholar]