Abstract
Oxidative stress is an important mechanism of chemical toxicity, contributing to developmental toxicity and teratogenesis as well as to cardiovascular and neurodegenerative diseases and diabetic embryopathy. Developing animals are especially sensitive to effects of chemicals that disrupt the balance of processes generating reactive species and oxidative stress, and those anti-oxidant defenses that protect against oxidative stress. The expression and inducibility of anti-oxidant defenses through activation of NFE2-related factor 2 (Nrf2) and related proteins is an essential process affecting the susceptibility to oxidants, but the complex interactions of Nrf2 in determining embryonic response to oxidants and oxidative stress are only beginning to be understood. The zebrafish (Danio rerio) is an established model in developmental biology and now also in developmental toxicology and redox signaling. Here we review the regulation of genes involved in protection against oxidative stress in developing vertebrates, with a focus on Nrf2 and related cap’n’collar (CNC)-basic-leucine zipper (bZIP) transcription factors. Vertebrate animals including zebrafish share Nfe2, Nrf1, Nrf2, and Nrf3 as well as a core set of genes that respond to oxidative stress, contributing to the value of zebrafish as a model system with which to investigate the mechanisms involved in regulation of redox signaling and the response to oxidative stress during embryolarval development. Moreover, studies in zebrafish have revealed nrf and keap1 gene duplications that provide an opportunity to dissect multiple functions of vertebrate NRF genes, including multiple sensing mechanisms involved in chemical-specific effects.
Keywords: oxidative stress, NRF2, NFE2L2, embryo, zebrafish, development
Introduction1
Oxidative stress occurs when the cellular redox balance is altered, disrupting redox signaling and regulation [1-3]. It can occur through the generation and action of reactive species such as superoxide anion, hydrogen peroxide, peroxynitrite, and other reactive products that can alter thiol redox circuits [4-6]. Oxidative stress is increasingly recognized as a significant mechanism of chemical toxicity and as a contributing factor in a wide range of pathological conditions and diseases in adults, and in developing animals. The roles of reactive species and redox status in normal physiology, and the mechanisms underlying adverse effects of oxidative stress, have been studied extensively in adult animals and cells. However, less is known about the role of these processes in developmental disorders and chemical toxicity during embryonic and fetal development.
Developing animals are uniquely sensitive to oxidative stress because of the rapid changes in cell proliferation and differentiation that occur at this time, and because enzymatic systems that protect against or repair toxic damage may not be fully mature early in development. The expression and inducibility of anti-oxidant defenses are critical factors affecting susceptibility to oxidative stress, but the ontogenic development of antioxidant defenses and their regulation in at early life stages are only beginning to be understood. The zebrafish (Danio rerio), an established model in developmental biology, has emerged recently as a valuable system for studying the expression and regulation of antioxidant defenses during development, and in particular the role of Nfe2-related transcription factors. Here we provide a review of the development and regulation of genes involved in protection against oxidative stress, with a focus on insights obtained from studies in zebrafish early life stages (embryos and larvae). For an additional perspective on the use of zebrafish as a model to study oxidative mechanisms, please see a recent review [7].
Oxidant signaling and oxidative stress in development and developmental toxicology
Importance of redox status and oxidant signaling during development
Embryonic development involves precisely orchestrated processes--including proliferation, differentiation, apoptosis, establishment of left-right asymmetry, gene expression, and epigenetic modifications--that increasingly are found to depend on redox signaling and intracellular redox potentials (Eh) [8-16]. Because endogenous oxidants and cellular redox potential play such important and fundamental roles in normal embryonic development, tight regulation of redox balance is critical. Toxicant exposures or other environmental stressors that increase oxidant levels or disrupt redox balance can fundamentally alter cell fate decisions. This can result in functional or structural changes, some of which could have long-term consequences that may only become apparent with subsequent stress or age [17, 18].
Role of oxidative stress in developmental toxicity
Oxidative stress contributes to a variety of human diseases [19-23] and has been implicated in the mechanism of action of human teratogens including valproic acid, phenytoin, ethanol, and thalidomide [16, 24-29], as well as in congenital malformations associated with PAH exposure [30, 31] and diabetic embryopathy [27, 32]. A variety of chemicals can generate reactive species or disrupt redox balance in other ways, through redox cycling, following biotransformation to reactive intermediates, or through induction or uncoupling of enzymes that can generate oxidants [33-36].
Studies of oxidative stress in adult organisms and cultured cells dominate the literature. However, developing animals (vertebrates including mammals and fish) are often more sensitive than young adults to chemical toxicity [37-40] and embryos appear to be especially sensitive to oxidative damage [24-27, 40, 41]. The risk of embryotoxicity and teratogenicity following exposure to pro-oxidant chemicals depends on the balance between reactions generating oxidative stress and reactions that are protective or repair the damage [24, 25, 41-45]. Consistent with this, supplementation with small molecular anti-oxidants or stimulation of anti-oxidant enzymes protects embryos against the developmental toxicity of a variety of chemicals [46-55] and embryos with reduced antioxidant capacity are at increased risk for developmental toxicity and teratogenicity [24, 30, 44, 56-58].
Constitutive and inducible antioxidant defenses in embryos
Anti-oxidant defenses can be constitutive, protecting against and balancing redox potential under normal physiological conditions, and inducible defenses, which protect against toxicant-stimulated levels of oxidants or redox imbalance. Embryos express relatively low basal (constitutive) levels of most anti-oxidants and anti-oxidant enzymes [59-67], which can put them at increased risk for oxidative toxicity [24, 25, 27]. Less is known about the inducible component of the anti-oxidant defense in embryos. A number of studies have examined inducible anti-oxidant defenses and the oxidative stress response in mammalian development [16, 17, 65, 68-70]. These studies have demonstrated the capacity of vertebrate embryos to respond to oxidative stress with increased expression of anti-oxidant genes, and in some cases have demonstrated a role for NFE2-related factor 2 (Nrf2)2 [70, 71]. Several groups including ours have begun to examine the oxidative stress response in zebrafish embryos (see below). These reports suggest that the responsiveness of embryos to oxidative stress can vary dramatically with developmental stage, and that this involves Nrf proteins. However, there is much yet to learn concerning the oxidative stress response in embryos and how it is regulated. Before discussing what has been learned from studies in zebrafish embryos, we briefly review what is known about the oxidative stress response and its regulation by NRF proteins in mammals. Additional details can be found in other reviews in this volume.
The oxidative stress response – regulation by Nfe2-related factors
Vertebrate animals possess an inducible enzymatic defense system that acts to detoxify reactive species and replenish the supply of small molecule anti-oxidants. Oxidants, electrophiles, and some so-called anti-oxidant chemicals activate an oxidative stress response via a family of related, cap’n’collar (CNC)-basic-leucine zipper (bZIP) transcription factors that interact with an anti-oxidant response element (ARE; a.k.a. electrophile response element or EpRE: 5’-TGACnnnGC-3’) in the promoters of target genes [72-76]. Known target genes in mammals include GSTA1, GSTP, NQO1, GCL, SOD, heme oxygenase (HO-1), and others.
The best-characterized oxidative stress response mechanism is that involving NRF2. NRF2 is normally found in the cytosol as an inactive complex with Kelch-like-ECH-associated protein (Keap1, also called iNRF2). Keap1 both retains NRF2 in the cytoplasm and enhances its proteasomal degradation [77-80]. Oxidative stress disrupts the interaction between NRF2 and Keap1 through a mechanism that may involve phosphorylation and/or disruption of sulfhydryl interactions [72, 81-87] affecting the nucleocytoplasmic shuttling of NRF2 [88, 89]. Once free of the repressor Keap1, NRF2 protein accumulates in the cell, enters the nucleus and forms a heterodimer with one of several small Maf proteins (MafF, MafG, MafK), which also contain bZIP motifs [90-92]. The NRF2-Maf dimer binds to AREs and activates transcription. A few studies suggest that Nrf2 may not dissociate from Keap1, but that the disruption of ubiquitination maintains Keap1 in a Nrf2-saturated state, permitting newly synthesized Nrf2 to travel unimpeded to the nucleus (e.g. reviewed in [93]). However, this remains controversial and has not yet been examined in zebrafish. Irrespective of whether Nrf2 dissociates from Keap1, or whether the de novo/Keap1 saturation model proves correct, accumulation of Nrf2 in the nucleus results in ARE-mediated gene regulation. An additional layer of regulation is provided by the related proteins BACH1 (BTB and CNC homology 1) and BACH2, which can also act as repressors of NRF2 [94, 95].
An oxidative stress response involving CNC proteins appears to be evolutionarily conserved in animals, with NRF homologs in Drosophila melanogaster [96, 97] and C. elegans [98, 99]. However, most of our current understanding of the oxidative stress response has been obtained from studies of the four mammalian CNC-bZIP proteins.
Nuclear factor erythroid-2 (NFE2)
NFE2 (also called p45) is a hematopoietic cell-specific transcription factor controlling globin gene expression as a complex with one of several small Maf proteins (p18) [100, 101]. NFE2 null mice are deficient in platelets and exhibit hemorrhaging and high mortality [102].
NFE2-related factor-1 (NRF1; also called NFE2L1, TCF11 and LCR-F1)
NRF1 is expressed in a variety of tissues [103] and as multiple isoforms [104]. Mice in which the Nrf1 gene has been disrupted die in utero either by day 7.5 or day 17.5 and exhibit a defect in definitive erythropoiesis [105, 106]. Studies in Nrf1−/−/Nrf1+/+ chimeric mice show that NRF1 is required for the development of the liver but not other tissues, that the effect is specific to hepatocytes, and that Nrf1−/− hepatocytes exhibit increased oxidative stress and undergo apoptosis late in pre-natal development [107]. NRF1 interacts with the ARE and regulates the basal expression and inducibility of GCL [108] and NQO1 [109, 110]; it is also important for glutathione homeostasis [111]. NRF1 can also act as a repressor of gene expression [104, 112]. Unlike NRF2 (below), NRF1 is localized to the endoplasmic reticulum and may play a role in the response to ER stress [113]. NRF1 is involved in regulating lipid metabolism [112, 114] and the expression of proteasome subunits in response to oxidant-damaged proteins [115, 116].
NRF2 (NFE2L2)
NRF2 was initially cloned from humans [117] and chickens [118] (called ECH). NRF2 binds to ARE sequences in vitro [119] and experiments in NRF2-deficient mice show that it regulates the basal and inducible expression of genes in the oxidative stress response, including NQO1, GSTP, GSTA, GSTM, GCL, ferritin, HO-1 [120-123] and other genes [124, 125]. Unlike targeted disruption of NRF1, which is embryo-lethal, NRF2 knock-out mice develop and reproduce normally [126]. Interestingly, constitutive activation of NRF2 in Keap1-deficient mice leads to postnatal lethality [127]. Moreover, disruption of both NRF2 and NRF1 results in pronounced oxidative stress, apoptosis, and embryolethality at an earlier stage than seen in the Nrf1−/− mice, demonstrating that NRF1 and NRF2 both have developmentally important, partially overlapping roles [128]. In addition, although the Nrf2−/− single knock-out mice are viable, as adults they are much more sensitive to chemicals that cause oxidative stress [129-132], including benzo[a]pyrene (BaP) [133, 134]. Thus, NRF2 has a key role in mediating the oxidative stress response in adult animals.
NRF3 (NFE2L3)
NRF3 is expressed in several tissues, most notably in the placenta and in B-cell lineages [135, 136], and like NRF1 exists as several isoforms and is localized to the endoplasmic reticulum [137, 138]. NRF3-null mice develop normally and compound NRF3/NRF2 and NRF3/NF-E2 mutants exhibit no additional phenotypes [139]. However, NRF3-null mice have increased susceptibility to benzo[a]pyrene-induced T-cell lymphoblastic lymphoma [140]. NRF3 can activate or repress gene expression and may act as a negative regulator of NRF2 [138, 141]. The role of NRF3 in protection against oxidative stress is currently unclear.
Most information about the regulation of the oxidative stress response by NRF proteins has been obtained in adult mammals and mammalian cells. What is the role of NRF-related transcription factors in regulating the response to oxidative stress in developing animals?
Zebrafish as a model for studying development roles of Nrf2 and related proteins
The zebrafish has emerged as a powerful system in which to examine mechanisms involved in the regulation of the oxidative stress response by Nrf2 and related proteins in developing animals. The zebrafish embryo is an important vertebrate model (or “tool” [142]) in embryology and developmental biology [143-153]. Because zebrafish are vertebrates, results obtained in this system have more direct relevance for humans as compared to studies in invertebrates. The fundamental features of developmental signaling pathways are conserved between fish and mammals, facilitating extrapolation of results from zebrafish to humans [151, 154-156].
A key technical advantage of zebrafish is the external development of nearly transparent embryos, facilitating direct observation of all stages of embryonic development, something not possible in mammals. The small size, rapid development (~48-72 hr from fertilization to hatch), and short generation time (2-3 mo) of zebrafish allow for large numbers of animals to be produced and housed [157]. Transgenic technologies are well developed in zebrafish [158]; transient and stable (germline) expression of transgenes can be used to visualize cell lineages [159], test promoter function [160, 161], and map regulatory elements [162-164], all in vivo. Heterologous promoters and proteins have been shown to function faithfully in zebrafish, recapitulating native expression patterns [162] or rescuing mutant phenotypes [165]. The coupling of green fluorescent protein (GFP)-based reporters [166-168] and transparent zebrafish embryos provides a powerful system for visualizing in vivo gene expression. Saturation mutagenesis screens have been performed using chemical or insertional mutagenesis [169-172] and many of the mutants reproduce human genetic diseases [173, 174].
Powerful loss-of-function approaches contribute to the utility of zebrafish in studying gene function. An anti-sense approach using morpholino-modified oligonucleotides (MOs) has been widely used to produce targeted gene “knock-downs” in developing zebrafish [175-177]. This approach has been useful for identifying gene function during development [175, 178-183] and to understand the role of specific genes in mechanisms of toxicity [184-191]. More recently, a trio of gene targeting methods have been developed and applied to zebrafish, making it possible to generate null alleles and homozygous null mutants at any locus. These methods include DNA-based targeted endonuclease approaches involving zinc-finger nucleases (ZFN) and transcription activator-like effector nuclease (TALENS) [192-195], as well as the RNA-guided clustered regularly interspaced short palindromic repeats (CRISPR)-Cas9 approach [196, 197]. Recent results comparing mutant and morphant phenotypes have highlighted the strengths and limitations of each approach [198-200].
A whole genome sequence supports a variety of genomic tools for studies in zebrafish [201, 202], and affords explicit comparisons with other species. Thus, zebrafish chromosomes exhibit large regions of conserved synteny with human chromosomes, facilitating studies of genome evolution and the identification of orthologous genes [203, 204]. An important advantage of zebrafish as a model is the existence of multiple copies of some human genes.
In general, zebrafish share with humans most features of key biochemical pathways (including signaling pathways). However, because of genome duplication in the teleost fish lineage shortly after its divergence from the tetrapod lineage [205-208], teleost fish have retained extra copies (paralogs) of some transcription factors, not found in humans. The existence of paralogs in fish provides an opportunity for new mechanistic insights, because duplicated fish genes might be exploited to obtain new information about the function of their single human counterpart [205-207, 209, 210]. This might apply especially to human genes with multiple functions or expression domains. The duplication, degeneration, complementation model of gene evolution [209, 210] predicts that the multiple functions or complex expression patterns of such a gene in humans may be partitioned between its fish “co-orthologs,” so that each of the duplicates retains a subset of the original functions. By studying each paralog individually (e.g. by loss-of-function approaches), their distinct roles (subsets of the original roles or expression patterns) can be examined in isolation and thereby elucidated. This approach may be especially informative in studying pleiotropic transcription factors such as Nrf2, because it can help to reveal some of the subtle functions of the mammalian homolog.
Because of these advantages, zebrafish were prominent among a group of alternative models recommended by the National Research Council for research in developmental toxicology [20]. More recently, the zebrafish model has been adopted and embraced by regulatory agencies and scientists in the U.S. and Europe [211-216] and by the pharmaceutical industry [217-224]. The number of studies in which this model system has been used to investigate mechanisms underlying the oxidative stress response was initially small [225-229], but has been rapidly growing. The emerging picture is that the fundamental features of the oxidative stress response are conserved in vertebrates, and thus that zebrafish is a powerful model with which to investigate oxidative phenomena during development.
Although there are many strengths of zebrafish as a model in which to study developmental processes, it is important to keep in mind that early in development (prior to the onset of circulation at ~26 hpf [230]), zebrafish embryos may experience a somewhat different oxygen environment than that experienced by mammalian embryos. Zebrafish embryos are typically cultured and develop at ambient [O2] [231, 232], whereas at early stages mammalian embryos experience lower oxygen tension, and higher concentrations can be toxic [233]. After the onset of circulation, however, zebrafish embryos are likely to experience oxygen environments similar to those of mammalian embryos. Thus, while the zebrafish embryo may be a valuable system for investigating fundamental mechanisms underlying the developmental roles of Nrf2 and related proteins, as with any model there may also be unique features that must be considered in interpreting the results.
Diversity of CNC-bZip transcription factors and related genes in zebrafish
Studies over the past 15 years have begun to establish the molecular mechanisms underlying the regulation of the oxidative stress response in fish and their similarity to those in mammals. Early studies [225, 226] demonstrated that reporter gene constructs containing a luciferase gene under control of mammalian ARE sequences, when transfected into fish cells, could be induced by exposure to tert-butylhydroquinone (tBHQ), suggesting that fish oxidant-responsive transcription factors are able to recognize mammalian ARE sequences. Subsequently, it was shown that exposure of zebrafish embryos or larvae to tBHQ or tert-butylhydroperoxide (tBOOH) induces an oxidative stress response [227, 234-237]. The first direct evidence for an oxidative stress response in fish and its mechanistic similarity to that in mammals was from Kobayashi et al. [227, 229], who cloned the cDNAs for zebrafish Nrf2 and Keap1 and showed that these proteins mediated the induction of gstp1, nqo1, and gclc by tBHQ in zebrafish embryos. Zebrafish also possess small Maf proteins (including a novel form, MafT) that form heterodimers with Nrf2 [238]. An evolutionarily conserved ARE 50-bp upstream of the gstp1 transcriptional start site was shown to interact with Nrf2 to regulate gstp1 expression [228].
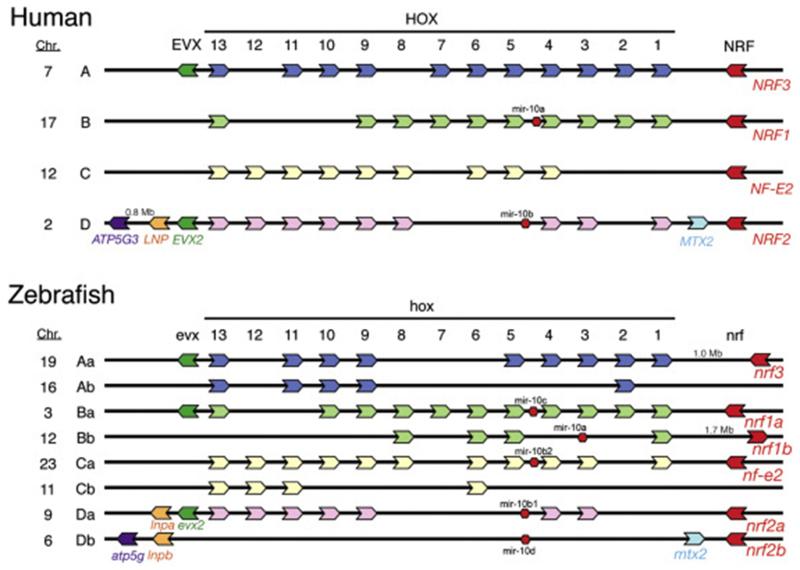
Since the initial studies in zebrafish cells and embryos, the number and identity of Nrf genes and their partners has become clear, revealing in some cases enhanced diversity in zebrafish as compared to humans. As noted above, four CNC-bZIP family members exist in humans and other mammals: NFE2, NRF1 (NFE2L1), NRF2 (NFE2L2), and NRF3 (NFE2L3). Zebrafish have a single nfe2 gene that is expressed in erythroid cells and in the developing ear [239]. The function of zebrafish nfe2 is not well understood, but loss-of-function studies involving knockdown of Nfe2 by morpholino oligonucleotides suggested that nfe2 may be required for proper cellular organization in the pneumatic duct and subsequent swim bladder function, as well as for proper formation of the otic vesicles [240]. In contrast to the single NRF1 gene in humans, zebrafish have two nrf1 paralogs (nrf1a, nrf1b)3 [234, 241], whose functions are not yet known. In addition to the original nrf2 gene [227] (now known as nrf2a), a second nrf2 gene (nrf2b [234]) has been identified. Overall, Timme-Laragy et al. [234] found that zebrafish (Danio rerio) possess six nrf genes, including nfe2, nrf3, and duplicated nrf1 and nrf2 genes. Comparative genomic analysis demonstrated extensive conserved synteny involving human and zebrafish nrf2 and hox genes (Fig. 1), indicating that nrf2a and nrf2b are co-orthologs of human NRF2, and nrf1a and nrf1b are co-orthologs of NRF1. The conserved synteny suggests that the nrf1 and nrf2 duplicates—like the hox clusters to which they are linked—arose as part of the fish-specific whole-genome duplication that occurred after divergence of fish and mammalian lineages [207, 242]. In general, the comparative genomic analyses demonstrated strong support for the hypothesis that the zebrafish nrf genes are orthologs (nfe2; nrf3) or co-orthologs (nrf1a and nrf1b; nrf2a and nrf2b) of the corresponding human NRF genes. Zebrafish nrf genes are summarized in Table 1.
Fig. 1. Comparative genomics of NFE2-related genes in zebrafish.
Location and shared synteny involving NFE2-related genes and HOX genes in the human genome (top) and zebrafish genome (bottom). This research was originally published in The Journal of Biological Chemistry. Timme-Laragy et al., Nrf2b, Novel Zebrafish Paralog of Oxidant-responsive Transcription Factor NF-E2-related Factor 2 (NRF2). The Journal of Biological Chemistry. 2012; Vol. 287: 4609-4627. [234] © the American Society for Biochemistry and Molecular Biology.
Table 1. nfe2-related genes in zebrafish.
| genea | protein | Expression during development |
Morphant phenotype |
Mutants available |
References |
|---|---|---|---|---|---|
| nfe2 | Nfe2 | yes; localized | swim bladder, otic vesicles |
no | [234, 239, 240] |
| nrf1a (nfe2l1a) | Nrf1a | yes | noneb | no | [234, 240] |
| nrf1b (nfe2l1b) | Nrf1b | yes | none | no | [234, 240] |
| nrf2a (nfe2l2a) | Nf2a | yes, localized; increasing 0-5 dpf |
gene expression |
nfe2l2fh318 (R479L) |
[227, 234, 240, 243, 244] |
| nrf2b (nfe2l2b) | Nrf2b | yes | gene expression |
no | [234, 240, 244] |
| nrf3 (nfe2l3) | Nrf3 | high | none | no | [234, 240] |
Notes:
Commonly used gene name is given. When different from the common name, the official gene name is provided in parentheses.
None = no phenotype detected so far, but phenotypes may be revealed upon further analysis.
The HOX genes have important and well-established roles in embryonic development [245]. The conservation of the syntenic relationship between the HOX clusters and all of the NFE2-related genes suggest that there has been strong selective pressure to preserve this arrangement over 500 million years of evolution. This has been seen also for other HOX-associated genes and in comparisons of human and pufferfish (Takifugu rubripes) genomes [246, 247]. An intriguing possibility is that there may be coordinated regulation of these genes during development—a possible example of “genomic regulatory blocks”—large regions of conserved synteny and conserved gene order with overlapping regulatory features that resist recombination [248, 249]. Notably, all six of the NFE2-related genes are expressed in embryos [234, 240] (see below), supporting a plausible regulatory association with the HOX developmental patterning genes.
Similar to the findings with nrf1 and nrf2, zebrafish also possess duplicated keap1 genes (keap1a and keap1b [227, 229]) and duplicated bach1 genes (bach1a, bach1b [250, 251]). Together, these results demonstrate that duplicated zebrafish genes are common in the Nrf pathways, as they are in some other pathways involved in regulation of response to environmental stressors [252, 253].
Ontogeny of glutathione redox dynamics and constitutive antioxidant defenses in developing embryos
In addition to the oxidative stress response and developmental events regulated by the Nrf-family of transcription factors, other endogenous antioxidant defenses play important roles in embryonic development. Glutathione (GSH) is the most abundant cellular antioxidant, present in mM concentrations, and is required for successful embryonic development [254, 255]. Importantly, numerous studies have demonstrated a strong relationship between concentrations and nuclear localization of GSH and cell cycle progression [256-259]. Cell fate decisions have also been shown to be closely related to GSH Eh, with more oxidized Eh associated with differentiation, and more reduced Eh with proliferation [260-262].
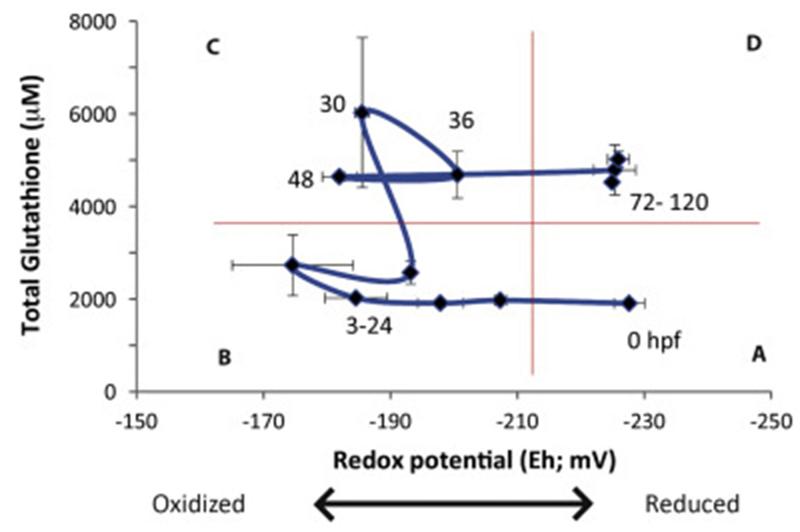
Timme-Laragy et al. [263] assessed GSH homeostasis during embryonic development by measuring GSH redox dynamics over 0-120 hours of zebrafish development. They used high-performance liquid chromatography (HPLC) to measure reduced and oxidized glutathione (GSH, GSSG) and calculated the whole embryo total glutathione (GSHT) concentrations and redox potentials (Eh) at each of twelve stages. The results (Fig. 2) revealed distinct windows of GSH concentration and redox status that correspond to major developmental events. Initially (at fertilization and shortly after), embryos have relatively low concentrations of GSHT and reducing conditions. Subsequently (3-24 hpf) concentrations of GSHT remain low (2-3 mM) but the redox potential becomes more oxidized. From 30-48 hpf, concentrations of GSHT increase while the redox potential remains oxidized. This is a period of cellular differentiation, which would be promoted at a cellular level by an oxidized redox status [260-262]. After hatching, GSH concentrations remain high and the redox potential becomes more reduced, conditions that promote cellular proliferation [261]. These results suggest that embryonic development involves a highly regulated trajectory of glutathione status involving changes in both the amount and oxidation state of this important regulator of cellular redox status. Studies in other fish demonstrate similar cycles of GSH regulation during development [264, 265].
Fig. 2. Concentrations of GSHT and Eh during the first 120 hours of zebrafish development.
Four “windows” of dynamic glutathione conditions during embryonic development are observed. A) reduced Eh and low GSHT, observed in mature oocytes; B) oxidized Eh and low GSHT, observed in embryos from the mid-blastula transition through somitogenesis (3-24 hpf); C) oxidized Eh and high GSHT, observed in embryos undergoing organ differentiation (30-48 hpf), and D) reduced Eh and high GSHT, observed in post-hatch eleutheroembryos. Reproduced, with permission, from Timme-Laragy et al. [263].
Timme-Laragy et al. [263] also identified the set of zebrafish orthologs of mammalian genes involved in glutathione synthesis, recycling, and utilization and measured their expression during development. They identified in the zebrafish genome extensive duplication of genes encoding proteins involved in GSH synthesis and recycling, and these genes exhibited complex patterns of gene expression in developing embryos. A number of the genes involved in GSH synthesis were maternally loaded and/or expressed highly in early embryos. Overall, ontogenic changes in the expression of GSH-related genes supported the hypothesis that GSH redox status is tightly regulated during development. However, the role of Nrf proteins and other transcription factors in regulating the constitutive expression of these genes in embryos is not yet well understood.
Developmental expression of Nfe2-related genes and inducible antioxidant defenses
The constitutive expression of genes involved in antioxidant defense may help to ensure that redox conditions are appropriate for normal embryonic development, and may provide some immediate protection against challenges posed by exposure to chemicals or other stressors. In addition, embryos possess the ability to up-regulate antioxidant defenses in response to challenges, but how these inducible responses vary by cell type and developmental stage and how they are regulated during development are poorly known.
Kobayashi et al. [227] found that exposure of zebrafish larvae to tBHQ (30 μM) for 6 hr at 4- or 7-days post fertilization (dpf) caused the induction of gstp1, nqo1, and gclc. However, exposure of 24-hpf embryos to tBHQ for 6 hours did not induce gstp1, suggesting that 24-hr old zebrafish embryos are less capable of mounting an oxidative stress response. Timme-Laragy et al. [235] found induction of gstp1, gpx1, and gclc in zebrafish embryos exposed to tBOOH or ß-naphthoflavone for 24 hr beginning at 24-hpf, suggesting that at some point during the 24-48-hpf period the embryos are capable of responding to these chemicals. A recent study [237] showed that tBHQ exposure for 6 hr caused strong induction of gstp1 in 1- and 2-dpf embryos. tBHQ also induced gclc (1-, 2-, and 4-dpf), but not sod1 or nqo1. Together, these reports suggest that there are stage- or chemical-specific differences in responsiveness to oxidative stress during development.
Gene expression profiles
A few studies have gene expression profiles in embryos exposed to oxidant chemicals. In a comparison of responses of zebrafish larvae to eleven different chemicals using microarrays, Yang et al. [236] found that tBHQ at 5-dpf produced a distinct profile that included a number of genes known from studies in mammals to be involved in the oxidative stress response. Nakajima et al. [266] reported that exposure of 4-dpf larvae to the sulfhydryl-reactive chemical diethylmaleate (DEM) induced 42 genes by more than 2-fold, including gclc, txnrd1 (thioredoxin reductase 1), peroxiredoxin 1, and several gst genes. In another study [237], microarray analysis of gene expression in 4-dpf larvae acutely exposed to tBHQ showed a robust response, with both up-regulated genes (220 genes induced by more than 2-fold) and down-regulated genes (109 genes). The induced genes included several involved in the synthesis and utilization of GSH and other sulfhydryl-reactive anti-oxidants (thioredoxin) and their regulation, including gclc, gclm, glutathione synthase (gss), GSH reductase (gr1), gamma-glutamyl transferase (ggt1a), and cystathionine beta-synthase (cbsb). There was substantial overlap (10/15) between genes induced in these larvae and a set of marker genes induced by oxidative stress in mammals (Table 2), demonstrating the evolutionary conservation in the oxidative stress response.
Table 2. Genes induced by exposure to tBHQ: Comparison of mammals and zebrafish.
| Marker gene in mammals | Zebrafish (co)-orthologs |
tBHQ / DMSO |
|---|---|---|
|
| ||
| HSP70 1A, 1B, 6 | hsp70 (Chr.3) | 52.3 |
| hsp70l (Chr.8) | 15.5 | |
|
| ||
| HSP70 9B | hspa9 (Chr.14) | 1.2 |
|
| ||
| HSP90 1 alpha | hsp90a (Chr.20) | 1.2 |
| hsp90a2 (Chr.20) | 10.4 | |
|
| ||
| DnaJ(Hsp40) B1 | dnajb1a (Chr.3) | 1.4 |
| dnajb1b (Chr.1) | 10.5 | |
|
| ||
| NAD(P)H quinone oxidoreductase-1 (NQO1) |
nqo1 | 0.7 |
|
| ||
| Glutamate-cysteine ligase, modifier subunit |
gclm | 4.1 |
|
| ||
| Thioredoxin (TXN) | txn1 (Chr.7) | 5.5 |
| txn2 (Chr.1) | 0.8 | |
|
| ||
| Thioredoxin reductase-1 | txnrd1 | 1.6 |
|
| ||
| Glutathione reductase | glutathione reductase 1 | 3.8 |
|
| ||
| Ferritin, heavy polypeptide-1 | fth1 (Chr.7) | 1.0 |
| ferritin-like (Chr.3) | 6.8 | |
| ferritin-like (Chr.25) | 0.7 | |
|
| ||
| Ferritin light polypeptide | ferritin L | 0.9 |
|
| ||
| Carbonyl reductase-1 | cbr1 | 1.1 |
|
| ||
| Phosphogluconate dehydrogenase |
pgd | 2.5 |
|
| ||
| Sequestosome-1 | sqstm1 | 7.6 |
|
| ||
| Ubiquitin thioesterase | usp4 | 1.0 |
A set of 20 candidate genes, based on the multiple data sets of genes responding to oxidative stress in mammalian cells, was compiled by Johnson and colleagues, as listed in Table 2 of Li et al. [267]. Here, the 20 mammalian candidate genes have been collapsed into 15 sets based on orthologous or co-orthologous relationships with zebrafish genes and including only those genes with orthologs represented on the Agilent zebrafish microarray. Of these 15 genes, 10 have at least one co-ortholog induced by tBHQ in 4-dpf zebrafish (indicated in bold type and green shading), whereas 5 are not induced under these conditions. This table is modified from Table 1 of Hahn et al. [237]; use is permitted by the Creative Commons Attribution license.
Developmental expression of nrf genes
How the expression of nrf genes and Nrf proteins varies during development is not fully understood, but current data suggest that there are substantial differences in developmental patterns (ontogeny), levels, and localization. Expression of nrf2a mRNA as assessed by RT-PCR is low in early embryos (≤24 hpf), but increases substantially from 2- to 7-dpf [268]. A similar pattern was seen using quantitative RT-PCR [234] and microarray [240]. Interestingly, microarray data suggest that nrf3 mRNA is maternally loaded and that levels of nrf3 expression through the first two days of embryonic development are much greater (by several orders of magnitude) than those of any other nrf gene [240].
Localization of nrf gene expression also varies by gene and developmental stage. As noted earlier, zebrafish nfe2 is expressed in erythroid cells and in the developing ear [239]. The expression of nrf2a mRNA at 24 hpf is restricted to the olfactory system, but by 5-dpf this gene is more widely expressed, appearing also in gill, liver, and intestine [266]. Expression of nrf2a mRNA is also seen in the exocrine pancreas at 84 hpf [250]. The expression of nrf3 mRNA is localized to fore, mid- and hindbrain and pectoral fin buds between 24- and 36-hpf [240]. The localization of nrf1 paralogs and nrf2b is not known. These patterns of tissue-, cell- and developmental stage-specific expression suggest that there are specific and varying needs for regulating redox status, and also that there are target cells, tissues, or stages that may be more susceptible to non-physiological oxidant challenge; neither of these is well understood.
Functional assessment of Nfe2-related genes in development and developmental toxicity
Loss-of-function approaches to define the roles of Nrf proteins
Measurements of gene expression—whether of nrf genes themselves or their putative target genes—provide intriguing associations but do not reveal regulatory relationships or mechanisms. In contrast, loss-of-function experiments in which the expression or function of a protein is reduced or eliminated can provide valuable mechanistic information about the role of that protein. The primary approach used so far to elucidate the functional roles of Nrf proteins in zebrafish embryos has been knock-down of protein expression using morpholino-modified anti-sense oligonucleotides (MOs) [175]. Although questions have arisen recently about the use of the MO approach to identify functions of novel genes [198-200], MO-based knock-down remains a valuable method for testing specific hypotheses about the role of transcription factors in regulating putative target genes.
Several groups have used MO-mediated knock down of Nrf2a to assess the role of this protein in the regulation of gene expression in zebrafish embryos in response to oxidant exposure. Some of these results are summarized in Table 3. These studies show that a number of oxidant-inducible genes (and at least one repressed gene) are under control of Nrf2a; the genes include some well-known Nrf2 targets such as gstp1, gclc, hmox1, and prdx1. However, other oxidant-inducible genes do not appear to be regulated via Nrf2a (Table 3), suggesting that other transcription factors may be involved in controlling their induction.
Table 3. Use of MO-mediated knock-down to determine Nrf-dependencea of altered gene expression in oxidant-exposed zebrafish embryos.
| Gene | Inducer | Age | Nrf2a- dependence? |
Reference |
|---|---|---|---|---|
| bcat1 | DEM | 4 dpf | Yes | [266] |
|
ferric-chelate
reductase 1 |
DEM | 4 dpf | Yes | [266] |
| ferritin heavy chain | DEM | 4 dpf | Yes | [266] |
| gclc | tBOOH, ANF + BNF | 2 dpf | Yes | [235] |
| gclc | CdCl2 | 4 dpf | Yes | [271] |
| gpx1 | tBOOH, ANF + BNF | 2 dpf | Yes | [235] |
| gsta1 | DEM | 4 dpf | Yes | [266] |
| gstp | tBHQ | 4 dpf | Yes | [227] |
| gstp1/gstp2 | tBOOH, ANF + BNF | 2 dpf | Yes | [235] |
| gstp | CdCl2 | 4 dpf | Yes | [271] |
| gstp1 | DEM | 4 dpf | Yes | [266] |
| gstp1 | tBHQ | 2 dpf | Yes | [234] |
| hmox1 | CdCl2 | 4 dpf | Yes | [271] |
| hmox1 | PFOS | 4 dpf | Yes | [272] |
| mitfa | tBHQ | 2 dpf | Yesb | [234] |
| peroxiredoxin 1 | DEM | 4 dpf | Yes | [266] |
| prdx1 | CdCl2 | 4 dpf | Yes | [271] |
| prdx1 | GSNORi | 54 hpf | Yes | [273] |
| sepw2b | DEM | 4 dpf | Yes | [266] |
| sod1 | ANF + BNF | 2 dpf | Yes | [235] |
| sod2 | ANF + BNF | 2 dpf | Yes | [235] |
| abcc2 | DEM | 4 dpf | No | [266] |
| atf3 | tBHQ | 2 dpf | No | [234] |
| bc2 | DEM | 4 dpf | No | [266] |
| cx32.3 | DEM | 4 dpf | No | [266] |
| hsp70 | tBHQ | 2 dpf | No | [234] |
| txnrd1 | DEM | 4 dpf | No | [266] |
| ugdh | DEM | 4 dpf | No | [266] |
Nrf2a-dependence is defined as genes showing a reduction of induction (or repression) by chemical in embryos injected with Nrf2a-MO as compared to a control MO.
Gene is repressed, rather than induced, by chemical treatment.
Abbreviations used: ANF: alpha-naphthoflavone; BNF: beta-naphthoflavone; DEM: diethylmaleate; GSNORi: S-nitrosoglutathione reductase (GSNOR) inhibitor N6547; PFOS: perfluorooctane sulfonate; tBHQ: tert-butylhydroquinone; tBOOH: tert-butylhydroperoxide.
A recent study [244] compared the tBHQ-induced gene expression profiles in 2-dpf zebrafish embryos in which either Nrf2a or Nrf2b had been knocked down. Knockdown of Nrf2a or Nrf2b blocked some but not all of the tBHQ-induced changes in gene expression and the effects of Nrf2a-MO and Nrf2b-MO were distinct. The results indicated that Nrf2 paralogs regulate distinct gene sets, with some overlapping targets, in response to oxidative stress in embryos. This study also highlighted the importance of gene down-regulation as a component of the oxidative stress response during embryonic development.
Much less is known about the role of other Nrf proteins, including the Nrf1 paralogs and Nrf3, in regulating specific genes in response to oxidative stress. Studies in mammals (e.g. [269, 270]) suggest that much will be learned by defining the sets of genes regulated by each Nrf protein in embryos. In addition, other redox-sensitive transcription factors may be involved in regulating the response to oxidant exposure (see below).
In addition to regulating the induction (or repression) of oxidant-responsive genes, Nrf proteins are likely to be involved in the regulation of constitutive (basal) expression of some genes. For example, Timme-Laragy et al. [234] performed gene expression profiling of Nrf2a-MO-injected embryos and Nrf2b-MO-injected embryos and found that Nrf2a and Nrf2b regulate distinct but partially overlapping gene sets even in the absence of chemical exposure. In Nrf2a-morphants, 198 probes were up-regulated, whereas 310 were down-regulated as compared with embryos injected with the control-MO. In contrast, Nrf2b-morphants had more up-regulated probes (280) than down-regulated probes (254), suggesting that Nrf2b can act as a repressor of constitutive gene transcription during development.
The loss-of-function caused by MO-based knock-down is partial and temporary. Thus, while MO experiments can help to identify functions during early development, they may miss functions occurring later or that can proceed in the presence of reduced Nrf protein expression. A more powerful approach to defining gene function is to generate null mutants in which the expression of a gene or function of its encoded protein is completely eliminated. While such an approach is common in mice and has been applied extensively to understand the role of murine Nrf proteins (see above), application of this approach to understand Nrf function in zebrafish is in its infancy.
Two point mutants of zebrafish nrf2a have been generated using the approach known as TILLING (Targeting Induced Local Lesions in Genomes [274, 275]). Each of these mutants (nfe2l2fh318 (R485L) and nfe2l2fh319 (N494I)) possesses a single amino acid change in a conserved residue in the basic region near the C-terminal end of the Nrf2 protein [243]. Kobayashi and colleagues [243] found that the protein encoded by the nfe2l2fh319 mutant retained activity, but that the transcriptional activity of the protein encoded by the nfe2l2fh318 allele was greatly reduced. Homozygous nfe2l2fh318 mutant fish were viable and fertile, similar to what is seen in Nrf2-knock-out mice [126]. However, mutant fish were more sensitive to the toxicity of H2O2 and other peroxides and certain electrophilic compounds, but not to some other oxidants [243]. The mechanism responsible for this increased sensitivity may involve a reduction in the ability to mount an oxidative stress response. Consistent with this, the induction of gstp1, prdx1, txn1, and gclc by H2O2 was greatly reduced in homozygous mutant larvae as compared to wild-type larvae [243].
The nfe2l2fh318 fish are also being used by other groups [273, 276, 277] to explore the role of nrf2a in protection against chemicals that cause oxidative stress.
Chemical specificity: patterns and mechanisms
A variety of chemicals can disrupt redox balance by generating oxidants or reacting with sulfhydryl groups, and thereby activate an oxidative stress response. There are questions about the mechanisms by which these chemicals activate this response in embryos. For example, do they act through Nrf2 or other Nrf proteins? Do chemicals that act by different mechanisms produce the same gene expression response?
A number of genes that are induced by oxidant exposure are shared in zebrafish and humans, and in a given species exposed to different oxidants [236, 237, 266, 267]. A recent study [278] examined the set of oxidant responsive genes that is induced by structurally distinct activators of the oxidative stress response in developing zebrafish. Zebrafish larvae (4 dpf) were exposed to tBHQ, tBOOH, diquat (DQ), or sulforaphane (SFN)) and gene expression was measured 6 hr later by microarray and Q-RT-PCR. The compounds caused overlapping but distinct patterns of altered gene expression. A core set of genes responded to all oxidants. However, other genes exhibited oxidant-specific changes in expression. Principal components analysis revealed that the changes in gene expression caused by SFN, a sulfhydryl-reactive agent, were distinct from those produced by the other oxidants. The results demonstrate that the oxidative stress response in developing animals is dependent upon the nature of the oxidative stress.
Kobayashi and colleagues [268, 279, 280] have begun to elucidate some of the mechanisms by which distinct responses might be produced for different chemicals. Their studies of this “multiple sensing mechanism” have identified six classes of inducers based in part upon which cysteines in Keap1 are targeted for oxidation. Additional mechanisms may involve activation of different Nrf paralogs. For example, MO knockdown studies in zebrafish embryos have shown that Nrf2a and Nrf2b regulate distinct gene sets, with only a small number of overlapping target genes [244]. The roles of other Nrf proteins—including Nrf1a, Nrf1b, and Nrf3—in regulating the response of developing animals to oxidative stress are completely unknown. Beyond the Nrf gene family, there are other redox-sensitive transcription factors such as AP-1, NFκB, p53, and HIF [281, 282] that may be involved in generating chemical-specific effects.
Conclusions and Future Directions
Regulation of constitutive and inducible defenses against oxidative stress by Nfe2-related factors is an evolutionarily conserved feature of animals. Conservation is especially evident among vertebrate animals, which share Nfe2, Nrf1, Nrf2, and Nrf3, as well as a core set of genes that respond to oxidative stress [227, 234, 237]. This conservation contributes to the value of zebrafish as a model system with which to investigate the mechanisms involved in regulation of redox signaling and the response to oxidative stress during embryo-larval development. Studies in zebrafish have revealed nrf and keap1 gene duplications that provide an opportunity to dissect multiple functions of vertebrate NRF genes, including multiple sensing mechanisms involved in chemical-specific effects.
Important questions about the regulation of the oxidative stress response in developing animals remain to be answered. We highlight three areas where substantial progress can be expected. One concerns the role of different Nrf homologs and paralogs in such regulation, and how it may vary by cell type and stage of development. Understanding the roles of Nrf proteins will be facilitated by gene-targeting and genome-editing approaches such as CRISPR-Cas9 [196, 197]. This method, first used for gene inactivation in 2013, has now exploded and is being used for a variety of applications, including single-nucleotide genome editing [283, 284]. This is a transformative technology that is dramatically enhancing our ability to manipulate systems to better understand gene function. It is possible, for example, to introduce known human polymorphisms into zebrafish nrf or keap1 genes to better understand how they affect Nrf protein function during development.
Another important goal is to understand the localized generation of oxidants and oxidative stress and the consequent responses. Use of transgenic technologies [158, 285] will facilitate new insight into the cell and tissue specificity of the oxidative stress response in embryos. Initial studies have begun to address such questions, using molecular probes to localize oxidative stress [7] and transient [228, 286] and germ-line transgenics [279, 287] to measure the response.
A third topic of great interest and where progress should be substantial in the near future is cross-talk between Nrf signaling pathways and other stress-response and developmental signaling pathways. One example is the aryl hydrocarbon receptor (AHR) signaling pathway, involved in both developmental processes and the response to endogenous and exogenous chemicals. Interactions between AHR and NRF signaling occur in mammals [288-291] and recent studies suggest that such interactions also occur in zebrafish embryos [234, 277, 292]. In addition to the AHR pathway, there are several other signaling pathways that engage in cross-talk with Nrf signaling in mammalian systems. Some of these pathways include those involving Hsf1, Notch, NFκB, PPARγ/RXR, and the unfolded protein response [93, 293-298]. Whether any of these interact with Nrf signaling during development remains to be determined.
Although substantial progress has been made in using the zebrafish embryo system to understand the role of Nrf proteins in regulating the oxidative stress response in developing animals, much remains to be done. The next few years will be an exciting time as new approaches are applied in novel ways to address longstanding questions about Nrf function during development.
Highlights.
The zebrafish is a valuable model for studying developmental roles of Nrf proteins.
Zebrafish and mammals share a core set of oxidative stress response genes.
Zebrafish and mammals share four types of NRF family transcription factors.
Zebrafish nrf gene duplicates can help elucidate pleotropic roles of Nrf proteins.
Loss-of-function approaches can reveal Nrf target genes in embryos.
Acknowledgments
We thank Dr. Makoto Kobayashi (University of Tsukuba, Japan) for his generosity in providing plasmids and information, and for his groundbreaking work to establish the zebrafish embryo model for studying the oxidative stress response. We thank Dr. Elwood Linney (Duke University) for inspiration and many helpful discussions on the use of the zebrafish model. This work was supported in part by National Institutes of Health grants R01ES016366 (MEH), R01ES015912 (JJS), and F32ES017585 (ART-L). The U.S. Government is authorized to produce and distribute reprints for governmental purposes notwithstanding any copyright notation that may appear hereon.
Footnotes
Abbreviations used: ARE: anti-oxidant response element; CNC-bZIP: cap’n’collar basic-leucine zipper; CRISPR: clustered regularly interspaced short palindromic repeats; DEM: diethylmaleate; DQ: diquat; GFP: green fluorescent protein; GSH: glutathione; MO: morpholino oligonucleotide; NRF: NFE2-related factor; RT-PCR: reverse transcription-polymerase chain reaction; SFN: sulforaphane; tBHQ: tert-butylhydroquinone; tBOOH: tert-butylhydroperoxide; TILLING: Targeting Induced Local Lesions IN Genomes
Nomenclature: NRF is a commonly used notation for NFE2-Related Factor genes, which are officially designated as NFE2L (NFE2-Like). For example, NRF2 is officially designated as NFE2L2, and NRF1 as NFE2L1. Throughout the paper, we use the more common NRF designation. Otherwise, we utilize the approved format for designating genes and proteins (https://wiki.zfin.org/display/general/ZFIN+Zebrafish+Nomenclature+Guidelines). In particular, human genes and proteins are designated using all capitals (NRF2 and NRF2, respectively), whereas zebrafish genes are designated nrf2 and Nrf2 for genes and proteins, respectively. When not referring to a specific species, we have used the human notation as a default format.
The designation of duplicated genes (and their encoded proteins) as “a” (as in nrf1a) or “b” (nrf1b) is according to the approved zebrafish nomenclature for duplicates resulting from the fish-specific whole-genome duplication (https://wiki.zfin.org/display/general/ZFIN+Zebrafish+Nomenclature+Guidelines).
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- [1].Fridovich I. Oxygen toxicity: a radical explanation. The Journal of experimental biology. 1998;201(Pt 8):1203–1209. doi: 10.1242/jeb.201.8.1203. [DOI] [PubMed] [Google Scholar]
- [2].Kohen R, Nyska A. Oxidation of biological systems: oxidative stress phenomena, antioxidants, redox reactions, and methods for their quantification. Toxicologic pathology. 2002;30:620–650. doi: 10.1080/01926230290166724. [DOI] [PubMed] [Google Scholar]
- [3].Hansen JM, Go YM, Jones DP. Nuclear and mitochondrial compartmentation of oxidative stress and redox signaling. Annual review of pharmacology and toxicology. 2006;46:215–234. doi: 10.1146/annurev.pharmtox.46.120604.141122. [DOI] [PubMed] [Google Scholar]
- [4].Jones DP. Redefining oxidative stress. Antioxidants & redox signaling. 2006;8:1865–1879. doi: 10.1089/ars.2006.8.1865. [DOI] [PubMed] [Google Scholar]
- [5].Sies H. Oxidative stress: a concept in redox biology and medicine. Redox biology. 2015;4:180–183. doi: 10.1016/j.redox.2015.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Jones DP. Radical-free biology of oxidative stress. American journal of physiology. Cell physiology. 2008;295:C849–868. doi: 10.1152/ajpcell.00283.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [7].Fang L, Miller YI. Emerging applications for zebrafish as a model organism to study oxidative mechanisms and their roles in inflammation and vascular accumulation of oxidized lipids. Free radical biology & medicine. 2012;53:1411–1420. doi: 10.1016/j.freeradbiomed.2012.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [8].Coffman JA, Coluccio A, Planchart A, Robertson AJ. Oral-aboral axis specification in the sea urchin embryo III. Role of mitochondrial redox signaling via H2O2. Developmental biology. 2009;330:123–130. doi: 10.1016/j.ydbio.2009.03.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Coffman JA, Davidson EH. Oral-aboral axis specification in the sea urchin embryo. I. Axis entrainment by respiratory asymmetry. Developmental biology. 2001;230:18–28. doi: 10.1006/dbio.2000.9996. [DOI] [PubMed] [Google Scholar]
- [10].Hernandez-Garcia D, Wood CD, Castro-Obregon S, Covarrubias L. Reactive oxygen species: A radical role in development? Free radical biology & medicine. 2010;49:130–143. doi: 10.1016/j.freeradbiomed.2010.03.020. [DOI] [PubMed] [Google Scholar]
- [11].Covarrubias L, Hernandez-Garcia D, Schnabel D, Salas-Vidal E, Castro-Obregon S. Function of reactive oxygen species during animal development: passive or active? Developmental biology. 2008;320:1–11. doi: 10.1016/j.ydbio.2008.04.041. [DOI] [PubMed] [Google Scholar]
- [12].Coffman JA, Denegre JM. Mitochondria, redox signaling and axis specification in metazoan embryos. Developmental biology. 2007;308:266–280. doi: 10.1016/j.ydbio.2007.05.042. [DOI] [PubMed] [Google Scholar]
- [13].Ufer C, Wang CC, Borchert A, Heydeck D, Kuhn H. Redox control in mammalian embryo development. Antioxidants & redox signaling. 2010;13:833–875. doi: 10.1089/ars.2009.3044. [DOI] [PubMed] [Google Scholar]
- [14].Ufer C, Wang CC. The Roles of Glutathione Peroxidases during Embryo Development. Frontiers in molecular neuroscience. 2011;4:12. doi: 10.3389/fnmol.2011.00012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Dennery PA. Role of redox in fetal development and neonatal diseases. Antioxidants & redox signaling. 2004;6:147–153. doi: 10.1089/152308604771978453. [DOI] [PubMed] [Google Scholar]
- [16].Dennery PA. Effects of oxidative stress on embryonic development. Birth Defects Res C Embryo Today. 2007;81:155–162. doi: 10.1002/bdrc.20098. [DOI] [PubMed] [Google Scholar]
- [17].Dennery PA. Oxidative stress in development: nature or nurture? Free radical biology & medicine. 2010;49:1147–1151. doi: 10.1016/j.freeradbiomed.2010.07.011. [DOI] [PubMed] [Google Scholar]
- [18].Wells PG, McCallum GP, Chen CS, Henderson JT, Lee CJ, Perstin J, Preston TJ, Wiley MJ, Wong AW. Oxidative stress in developmental origins of disease: teratogenesis, neurodevelopmental deficits, and cancer. Toxicol Sci. 2009;108:4–18. doi: 10.1093/toxsci/kfn263. [DOI] [PubMed] [Google Scholar]
- [19].Cooke MS, Evans MD, Dizdaroglu M, Lunec J. Oxidative DNA damage: mechanisms, mutation, and disease. Faseb J. 2003;17:1195–1214. doi: 10.1096/fj.02-0752rev. [DOI] [PubMed] [Google Scholar]
- [20].National Research Council. National Academy of Sciences . Scientific frontiers in developmental toxicology and risk assessment. National Academy Press; 2000. [PubMed] [Google Scholar]
- [21].Rangasamy T, Cho CY, Thimmulappa RK, Zhen L, Srisuma SS, Kensler TW, Yamamoto M, Petrache I, Tuder RM, Biswal S. Genetic ablation of Nrf2 enhances susceptibility to cigarette smoke-induced emphysema in mice. The Journal of clinical investigation. 2004;114:1248–1259. doi: 10.1172/JCI21146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Andersen JK. Oxidative stress in neurodegeneration: cause or consequence? Nature medicine. 2004;10(Suppl):S18–25. doi: 10.1038/nrn1434. [DOI] [PubMed] [Google Scholar]
- [23].Bossy-Wetzel E, Schwarzenbacher R, Lipton SA. Molecular pathways to neurodegeneration. Nature medicine. 2004;10(Suppl):S2–9. doi: 10.1038/nm1067. [DOI] [PubMed] [Google Scholar]
- [24].Wells PG, Bhuller Y, Chen CS, Jeng W, Kasapinovic S, Kennedy JC, Kim PM, Laposa RR, McCallum GP, Nicol CJ, Parman T, Wiley MJ, Wong AW. Molecular and biochemical mechanisms in teratogenesis involving reactive oxygen species. Toxicol Appl Pharmacol. 2005;207:354–366. doi: 10.1016/j.taap.2005.01.061. [DOI] [PubMed] [Google Scholar]
- [25].Wells PG, Kim PM, Laposa RR, Nicol CJ, Parman T, Winn LM. Oxidative damage in chemical teratogenesis. Mutation research. 1997;396:65–78. doi: 10.1016/s0027-5107(97)00175-9. [DOI] [PubMed] [Google Scholar]
- [26].Hansen JM. Oxidative stress as a mechanism of teratogenesis. Birth Defects Res C Embryo Today. 2006;78:293–307. doi: 10.1002/bdrc.20085. [DOI] [PubMed] [Google Scholar]
- [27].Ornoy A. Embryonic oxidative stress as a mechanism of teratogenesis with special emphasis on diabetic embryopathy. Reproductive toxicology (Elmsford, N.Y. 2007;24:31–41. doi: 10.1016/j.reprotox.2007.04.004. [DOI] [PubMed] [Google Scholar]
- [28].De Santis M, Di Gianantonio E, Straface G, Cavaliere AF, Caruso A, Schiavon F, Berletti R, Clementi M. Ionizing radiations in pregnancy and teratogenesis: a review of literature. Reproductive toxicology (Elmsford, N.Y. 2005;20:323–329. doi: 10.1016/j.reprotox.2005.04.004. [DOI] [PubMed] [Google Scholar]
- [29].De Santis M, Cesari E, Nobili E, Straface G, Cavaliere AF, Caruso A. Radiation effects on development. Birth Defects Res C Embryo Today. 2007;81:177–182. doi: 10.1002/bdrc.20099. [DOI] [PubMed] [Google Scholar]
- [30].Lammer EJ, Shaw GM, Iovannisci DM, Finnell RH. Maternal smoking, genetic variation of glutathione s-transferases, and risk for orofacial clefts. Epidemiology. 2005;16:698–701. doi: 10.1097/01.ede.0000172136.26733.4b. [DOI] [PubMed] [Google Scholar]
- [31].Parman T, Wells PG. Embryonic prostaglandin H synthase-2 (PHS-2) expression and benzo[a]pyrene teratogenicity in PHS-2 knockout mice. Faseb J. 2002;16:1001–1009. doi: 10.1096/fj.01-0140com. [DOI] [PubMed] [Google Scholar]
- [32].Loeken MR. Advances in understanding the molecular causes of diabetes-induced birth defects. J Soc Gynecol Investig. 2006;13:2–10. doi: 10.1016/j.jsgi.2005.09.007. [DOI] [PubMed] [Google Scholar]
- [33].Bolton JL, Trush MA, Penning TM, Dryhurst G, Monks TJ. Role of quinones in toxicology. Chemical research in toxicology. 2000;13:135–160. doi: 10.1021/tx9902082. [DOI] [PubMed] [Google Scholar]
- [34].Kuthan H, Ullrich V. Oxidase and oxygenase function of the microsomal cytochrome P450 monooxygenase system. European journal of biochemistry / FEBS. 1982;126:583–588. doi: 10.1111/j.1432-1033.1982.tb06820.x. [DOI] [PubMed] [Google Scholar]
- [35].Schlezinger JJ, White RD, Stegeman JJ. Oxidative inactivation of cytochrome P450 1A (CYP1A) stimulated by 3,3’,4,4’-tetrachlorobiphenyl: Production of reactive oxygen by vertebrate CYP1As. Mol Pharmacol. 1999;56:588–597. doi: 10.1124/mol.56.3.588. [DOI] [PubMed] [Google Scholar]
- [36].Park J-YK, Shigenaga MK, Ames BN. Induction of cytochrome P4501A1 by 2,3,7,8-tetrachlorodibenzo-p-dioxin or indolo(3,2-b)carbazole is associated with oxidative DNA damage. Proceedings of the National Academy of Sciences U.S.A. 1996;93:2322–2327. doi: 10.1073/pnas.93.6.2322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [37].Brent RL. Utilization of animal studies to determine the effects and human risks of environmental toxicants (drugs, chemicals, and physical agents) Pediatrics. 2004;113:984–995. [PubMed] [Google Scholar]
- [38].Brent RL, Tanski S, Weitzman M. A pediatric perspective on the unique vulnerability and resilience of the embryo and the child to environmental toxicants: the importance of rigorous research concerning age and agent. Pediatrics. 2004;113:935–944. [PubMed] [Google Scholar]
- [39].Barton HA, Cogliano VJ, Flowers L, Valcovic L, Setzer RW, Woodruff TJ. Assessing susceptibility from early-life exposure to carcinogens. Environmental health perspectives. 2005;113:1125–1133. doi: 10.1289/ehp.7667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Perera FP, Tang D, Tu YH, Cruz LA, Borjas M, Bernert T, Whyatt RM. Biomarkers in maternal and newborn blood indicate heightened fetal susceptibility to procarcinogenic DNA damage. Environmental health perspectives. 2004;112:1133–1136. doi: 10.1289/ehp.6833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Wells PG, Winn LM. Biochemical toxicology of chemical teratogenesis. Crit Rev Biochem Mol Biol. 1996;31:1–40. doi: 10.3109/10409239609110574. [DOI] [PubMed] [Google Scholar]
- [42].Vinson RK, Hales BF. DNA repair during organogenesis. Mutation research. 2002;509:79–91. doi: 10.1016/s0027-5107(02)00223-3. [DOI] [PubMed] [Google Scholar]
- [43].Vinson RK, Hales BF. Genotoxic stress response gene expression in the mid-organogenesis rat conceptus. Toxicol Sci. 2003;74:157–164. doi: 10.1093/toxsci/kfg097. [DOI] [PubMed] [Google Scholar]
- [44].Nicol CJ, Zielenski J, Tsui LC, Wells PG. An embryoprotective role for glucose-6-phosphate dehydrogenase in developmental oxidative stress and chemical teratogenesis. Faseb J. 2000;14:111–127. doi: 10.1096/fasebj.14.1.111. [DOI] [PubMed] [Google Scholar]
- [45].Nicol CJ, Harrison ML, Laposa RR, Gimelshtein IL, Wells PG. A teratologic suppressor role for p53 in benzo[a]pyrene-treated transgenic p53-deficient mice. Nature genetics. 1995;10:181–187. doi: 10.1038/ng0695-181. [DOI] [PubMed] [Google Scholar]
- [46].Cantrell SM, Lutz LH, Tillitt DE, Hannink M. Embryotoxicity of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD): The embryonic vasculature is a physiological target for TCDD-induced DNA damage and apoptotic cell death in Medaka (Orizias latipes) Toxicol Appl Pharmacol. 1996;141:23–34. doi: 10.1006/taap.1996.0256. [DOI] [PubMed] [Google Scholar]
- [47].Dong W, Teraoka H, Yamazaki K, Tsukiyama S, Imani S, Imagawa T, Stegeman JJ, Peterson RE, Hiraga T. 2,3,7,8-tetrachlorodibenzo-p-dioxin toxicity in the zebrafish embryo: local circulation failure in the dorsal midbrain is associated with increased apoptosis. Toxicol Sci. 2002;69:191–201. doi: 10.1093/toxsci/69.1.191. [DOI] [PubMed] [Google Scholar]
- [48].Reimers MJ, La Du JK, Periera CB, Giovanini J, Tanguay RL. Ethanol-dependent toxicity in zebrafish is partially attenuated by antioxidants. Neurotoxicology and teratology. 2006;28:497–508. doi: 10.1016/j.ntt.2006.05.007. [DOI] [PubMed] [Google Scholar]
- [49].Kotch LE, Chen SY, Sulik KK. Ethanol-induced teratogenesis: free radical damage as a possible mechanism. Teratology. 1995;52:128–136. doi: 10.1002/tera.1420520304. [DOI] [PubMed] [Google Scholar]
- [50].Wentzel P, Rydberg U, Eriksson UJ. Antioxidative treatment diminishes ethanol-induced congenital malformations in the rat. Alcoholism, clinical and experimental research. 2006;30:1752–1760. doi: 10.1111/j.1530-0277.2006.00208.x. [DOI] [PubMed] [Google Scholar]
- [51].Usenko CY, Harper SL, Tanguay RL. Fullerene C60 exposure elicits an oxidative stress response in embryonic zebrafish. Toxicol Appl Pharmacol. 2008;229:44–55. doi: 10.1016/j.taap.2007.12.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [52].Al Deeb S, Al Moutaery K, Arshaduddin M, Tariq M. Vitamin E decreases valproic acid induced neural tube defects in mice. Neuroscience letters. 2000;292:179–182. doi: 10.1016/s0304-3940(00)01457-9. [DOI] [PubMed] [Google Scholar]
- [53].Sahambi SK, Hales BF. Exposure to 5-bromo-2’-deoxyuridine induces oxidative stress and activator protein-1 DNA binding activity in the embryo. Birth defects research. 2006;76:580–591. doi: 10.1002/bdra.20284. [DOI] [PubMed] [Google Scholar]
- [54].Winn LM, Wells PG. Evidence for embryonic prostaglandin H synthase-catalyzed bioactivation and reactive oxygen species-mediated oxidation of cellular macromolecules in phenytoin and benzo[a]pyrene teratogenesis. Free radical biology & medicine. 1997;22:607–621. doi: 10.1016/s0891-5849(96)00340-1. [DOI] [PubMed] [Google Scholar]
- [55].Parng C, Ton C, Lin YX, Roy NM, McGrath P. A zebrafish assay for identifying neuroprotectants in vivo. Neurotoxicology and teratology. 2006;28:509–516. doi: 10.1016/j.ntt.2006.04.003. [DOI] [PubMed] [Google Scholar]
- [56].Yan J, Hales BF. Depletion of glutathione induces 4-hydroxynonenal protein adducts and hydroxyurea teratogenicity in the organogenesis stage mouse embryo. The Journal of pharmacology and experimental therapeutics. 2006;319:613–621. doi: 10.1124/jpet.106.109850. [DOI] [PubMed] [Google Scholar]
- [57].Ozolins TR, Harrouk W, Doerksen T, Trasler JM, Hales BF. Buthionine sulfoximine embryotoxicity is associated with prolonged AP-1 activation. Teratology. 2002;66:192–200. doi: 10.1002/tera.10084. [DOI] [PubMed] [Google Scholar]
- [58].Shi ZZ, Osei-Frimpong J, Kala G, Kala SV, Barrios RJ, Habib GM, Lukin DJ, Danney CM, Matzuk MM, Lieberman MW. Glutathione synthesis is essential for mouse development but not for cell growth in culture. Proc Natl Acad Sci U S A. 2000;97:5101–5106. doi: 10.1073/pnas.97.10.5101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [59].Khan JY, Black SM. Developmental changes in murine brain antioxidant enzymes. Pediatric research. 2003;54:77–82. doi: 10.1203/01.PDR.0000065736.69214.20. [DOI] [PubMed] [Google Scholar]
- [60].el-Hage S, Singh SM. Temporal expression of genes encoding free radical-metabolizing enzymes is associated with higher mRNA levels during in utero development in mice. Developmental genetics. 1990;11:149–159. doi: 10.1002/dvg.1020110205. [DOI] [PubMed] [Google Scholar]
- [61].Zaken V, Kohen R, Ornoy A. The development of antioxidant defense mechanism in young rat embryos in vivo and in vitro. Early pregnancy (Online) 2000;4:110–123. [PubMed] [Google Scholar]
- [62].Ozolins TR, Siksay DL, Wells PG. Modulation of embryonic glutathione peroxidase activity and phenytoin teratogenicity by dietary deprivation of selenium in CD-1 mice. The Journal of pharmacology and experimental therapeutics. 1996;277:945–953. [PubMed] [Google Scholar]
- [63].Choe H, Hansen JM, Harris C. Spatial and temporal ontogenies of glutathione peroxidase and glutathione disulfide reductase during development of the prenatal rat. Journal of biochemical and molecular toxicology. 2001;15:197–206. doi: 10.1002/jbt.17. [DOI] [PubMed] [Google Scholar]
- [64].Stuart RO, Bush KT, Nigam SK. Changes in global gene expression patterns during development and maturation of the rat kidney. Proc Natl Acad Sci U S A. 2001;98:5649–5654. doi: 10.1073/pnas.091110798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [65].Stover SK, Gushansky GA, Salmen JJ, Gardiner CS. Regulation of gamma-glutamate-cysteine ligase expression by oxidative stress in the mouse preimplantation embryo. Toxicol Appl Pharmacol. 2000;168:153–159. doi: 10.1006/taap.2000.9030. [DOI] [PubMed] [Google Scholar]
- [66].Gardiner CS, Reed DJ. Synthesis of glutathione in the preimplantation mouse embryo. Arch Biochem Biophys. 1995;318:30–36. doi: 10.1006/abbi.1995.1200. [DOI] [PubMed] [Google Scholar]
- [67].Gardiner CS, Reed DJ. Status of glutathione during oxidant-induced oxidative stress in the preimplantation mouse embryo. Biology of reproduction. 1994;51:1307–1314. doi: 10.1095/biolreprod51.6.1307. [DOI] [PubMed] [Google Scholar]
- [68].Harris C, Hansen JM. Nrf2-mediated resistance to oxidant-induced redox disruption in embryos. Birth defects research. Part B, Developmental and reproductive toxicology. 2012;95:213–218. doi: 10.1002/bdrb.21005. [DOI] [PubMed] [Google Scholar]
- [69].Hansen JM, Harris C. Redox Control of Teratogenesis. Reproductive toxicology. 2013;35:165–179. doi: 10.1016/j.reprotox.2012.09.004. [DOI] [PubMed] [Google Scholar]
- [70].Dong J, Sulik KK, Chen SY. Nrf2-mediated transcriptional induction of antioxidant response in mouse embryos exposed to ethanol in vivo: implications for the prevention of fetal alcohol spectrum disorders. Antioxidants & redox signaling. 2008;10:2023–2033. doi: 10.1089/ars.2007.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [71].Ramkissoon A, Wells PG. Developmental role of nuclear factor E2-related factor 2 in mitigating methamphetamine fetal toxicity and postnatal neurodevelopmental deficits. Free radical biology & medicine. 2013;65:620–631. doi: 10.1016/j.freeradbiomed.2013.07.043. [DOI] [PubMed] [Google Scholar]
- [72].Nguyen T, Sherratt PJ, Pickett CB. Regulatory mechanisms controlling gene expression mediated by the antioxidant response element. Annual review of pharmacology and toxicology. 2003;43:233–260. doi: 10.1146/annurev.pharmtox.43.100901.140229. [DOI] [PubMed] [Google Scholar]
- [73].Friling RS, Bensimon A, Tichauer Y, Daniel V. Xenobiotic-inducible expression of murine glutathione S-transferase Ya subunit gene is controlled by an electrophile-responsive element. Proc Natl Acad Sci U S A. 1990;87:6258–6262. doi: 10.1073/pnas.87.16.6258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [74].Telakowski-Hopkins CA, King RG, Pickett CB. Glutathione S-transferase Ya subunit gene: identification of regulatory elements required for basal level and inducible expression. Proc Natl Acad Sci U S A. 1988;85:1000–1004. doi: 10.1073/pnas.85.4.1000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [75].Kensler TW, Wakabayashi N, Biswal S. Cell Survival Responses to Environmental Stresses Via the Keap1-Nrf2-ARE Pathway. Annual review of pharmacology and toxicology. 2007;47:89–116. doi: 10.1146/annurev.pharmtox.46.120604.141046. [DOI] [PubMed] [Google Scholar]
- [76].Ma Q, He X. Molecular basis of electrophilic and oxidative defense: promises and perils of Nrf2. Pharmacological reviews. 2012;64:1055–1081. doi: 10.1124/pr.110.004333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [77].Nguyen T, Sherratt PJ, Huang HC, Yang CS, Pickett CB. Increased protein stability as a mechanism that enhances Nrf2-mediated transcriptional activation of the antioxidant response element. Degradation of Nrf2 by the 26 S proteasome. The Journal of biological chemistry. 2003;278:4536–4541. doi: 10.1074/jbc.M207293200. [DOI] [PubMed] [Google Scholar]
- [78].McMahon M, Itoh K, Yamamoto M, Hayes JD. Keap1-dependent proteasomal degradation of transcription factor Nrf2 contributes to the negative regulation of antioxidant response element-driven gene expression. The Journal of biological chemistry. 2003;278:21592–21600. doi: 10.1074/jbc.M300931200. [DOI] [PubMed] [Google Scholar]
- [79].Dhakshinamoorthy S, Jaiswal AK. Functional characterization and role of INrf2 in antioxidant response element-mediated expression and antioxidant induction of NAD(P)H:quinone oxidoreductase1 gene. Oncogene. 2001;20:3906–3917. doi: 10.1038/sj.onc.1204506. [DOI] [PubMed] [Google Scholar]
- [80].Itoh K, Wakabayashi N, Katoh Y, Ishii T, O’Connor T, Yamamoto M. Keap1 regulates both cytoplasmic-nuclear shuttling and degradation of Nrf2 in response to electrophiles. Genes Cells. 2003;8:379–391. doi: 10.1046/j.1365-2443.2003.00640.x. [DOI] [PubMed] [Google Scholar]
- [81].Dinkova-Kostova AT, Holtzclaw WD, Cole RN, Itoh K, Wakabayashi N, Katoh Y, Yamamoto M, Talalay P. Direct evidence that sulfhydryl groups of Keap1 are the sensors regulating induction of phase 2 enzymes that protect against carcinogens and oxidants. Proc Natl Acad Sci U S A. 2002;99:11908–11913. doi: 10.1073/pnas.172398899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [82].Bloom DA, Jaiswal AK. Phosphorylation of Nrf2S40 by PKC in response to antioxidants leads to the release of Nrf2 from INrf2 but not required for Nrf2 stabilization/accumulation in the nucleus and transcriptional activation of ARE-mediated NQO1 gene expression. The Journal of biological chemistry. 2003;278:44675–44682. doi: 10.1074/jbc.M307633200. [DOI] [PubMed] [Google Scholar]
- [83].Kang MI, Kobayashi A, Wakabayashi N, Kim SG, Yamamoto M. Scaffolding of Keap1 to the actin cytoskeleton controls the function of Nrf2 as key regulator of cytoprotective phase 2 genes. Proc Natl Acad Sci U S A. 2004;101:2046–2051. doi: 10.1073/pnas.0308347100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [84].Wakabayashi N, Dinkova-Kostova AT, Holtzclaw WD, Kang MI, Kobayashi A, Yamamoto M, Kensler TW, Talalay P. Protection against electrophile and oxidant stress by induction of the phase 2 response: fate of cysteines of the Keap1 sensor modified by inducers. Proc Natl Acad Sci U S A. 2004;101:2040–2045. doi: 10.1073/pnas.0307301101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [85].Kobayashi A, Kang MI, Okawa H, Ohtsuji M, Zenke Y, Chiba T, Igarashi K, Yamamoto M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Molecular and cellular biology. 2004;24:7130–7139. doi: 10.1128/MCB.24.16.7130-7139.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [86].Cullinan SB, Gordan JD, Jin J, Harper JW, Diehl JA. The Keap1-BTB protein is an adaptor that bridges Nrf2 to a Cul3-based E3 ligase: oxidative stress sensing by a Cul3-Keap1 ligase. Molecular and cellular biology. 2004;24:8477–8486. doi: 10.1128/MCB.24.19.8477-8486.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [87].Eggler AL, Liu G, Pezzuto JM, van Breemen RB, Mesecar AD. Modifying specific cysteines of the electrophile-sensing human Keap1 protein is insufficient to disrupt binding to the Nrf2 domain Neh2. Proc Natl Acad Sci U S A. 2005;102:10070–10075. doi: 10.1073/pnas.0502402102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [88].Jain AK, Bloom DA, Jaiswal AK. Nuclear import and export signals in control of Nrf2. The Journal of biological chemistry. 2005;280:29158–29168. doi: 10.1074/jbc.M502083200. [DOI] [PubMed] [Google Scholar]
- [89].Nguyen T, Sherratt PJ, Nioi P, Yang CS, Pickett CB. Nrf2 controls constitutive and inducible expression of ARE-driven genes through a dynamic pathway involving nucleocytoplasmic shuttling by Keap1. The Journal of biological chemistry. 2005;280:32485–32492. doi: 10.1074/jbc.M503074200. [DOI] [PubMed] [Google Scholar]
- [90].Marini MG, Chan K, Casula L, Kan YW, Cao A, Moi P. hMAF, a small human transcription factor that heterodimerizes specifically with Nrf1 and Nrf2. The Journal of biological chemistry. 1997;272:16490–16497. doi: 10.1074/jbc.272.26.16490. [DOI] [PubMed] [Google Scholar]
- [91].Johnsen O, Skammelsrud N, Luna L, Nishizawa M, Prydz H, Kolsto AB. Small Maf proteins interact with the human transcription factor TCF11/Nrf1/LCR-F1. Nucleic acids research. 1996;24:4289–4297. doi: 10.1093/nar/24.21.4289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [92].Motohashi H, Katsuoka F, Engel JD, Yamamoto M. Small Maf proteins serve as transcriptional cofactors for keratinocyte differentiation in the Keap1-Nrf2 regulatory pathway. Proc Natl Acad Sci U S A. 2004;101:6379–6384. doi: 10.1073/pnas.0305902101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [93].Hayes JD, Dinkova-Kostova AT. The Nrf2 regulatory network provides an interface between redox and intermediary metabolism. Trends in biochemical sciences. 2014;39:199–218. doi: 10.1016/j.tibs.2014.02.002. [DOI] [PubMed] [Google Scholar]
- [94].Igarashi K, Sun J. The heme-Bach1 pathway in the regulation of oxidative stress response and erythroid differentiation. Antioxidants & redox signaling. 2006;8:107–118. doi: 10.1089/ars.2006.8.107. [DOI] [PubMed] [Google Scholar]
- [95].Igarashi K, Watanabe-Matsui M. Wearing red for signaling: the heme-bach axis in heme metabolism, oxidative stress response and iron immunology. The Tohoku journal of experimental medicine. 2014;232:229–253. doi: 10.1620/tjem.232.229. [DOI] [PubMed] [Google Scholar]
- [96].Sykiotis GP, Bohmann D. Stress-activated cap’n’collar transcription factors in aging and human disease. Sci Signal. 2010;3:re3. doi: 10.1126/scisignal.3112re3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [97].Sykiotis GP, Bohmann D. Keap1/Nrf2 signaling regulates oxidative stress tolerance and lifespan in Drosophila. Developmental cell. 2008;14:76–85. doi: 10.1016/j.devcel.2007.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [98].Walker AK, See R, Batchelder C, Kophengnavong T, Gronniger JT, Shi Y, Blackwell TK. A conserved transcription motif suggesting functional parallels between Caenorhabditis elegans SKN-1 and Cap’n’Collar-related basic leucine zipper proteins. The Journal of biological chemistry. 2000;275:22166–22171. doi: 10.1074/jbc.M001746200. [DOI] [PubMed] [Google Scholar]
- [99].An JH, Blackwell TK. SKN-1 links C. elegans mesendodermal specification to a conserved oxidative stress response. Genes & development. 2003;17:1882–1893. doi: 10.1101/gad.1107803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [100].Ney PA, Andrews NC, Jane SM, Safer B, Purucker ME, Weremowicz S, Morton CC, Goff SC, Orkin SH, Nienhuis AW. Purification of the human NF-E2 complex: cDNA cloning of the hematopoietic cell-specific subunit and evidence for an associated partner. Molecular and cellular biology. 1993;13:5604–5612. doi: 10.1128/mcb.13.9.5604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [101].Andrews NC, Erdjument-Bromage H, Davidson MB, Tempst P, Orkin SH. Erythroid transcription factor NF-E2 is a haematopoietic-specific basic-leucine zipper protein. Nature. 1993;362:722–728. doi: 10.1038/362722a0. [DOI] [PubMed] [Google Scholar]
- [102].Shivdasani RA, Rosenblatt MF, Zucker-Franklin D, Jackson CW, Hunt P, Saris CJ, Orkin SH. Transcription factor NF-E2 is required for platelet formation independent of the actions of thrombopoietin/MGDF in megakaryocyte development. Cell. 1995;81:695–704. doi: 10.1016/0092-8674(95)90531-6. [DOI] [PubMed] [Google Scholar]
- [103].Chan JY, Han XL, Kan YW. Cloning of Nrf1, an NF-E2-related transcription factor, by genetic selection in yeast. Proc Natl Acad Sci U S A. 1993;90:11371–11375. doi: 10.1073/pnas.90.23.11371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [104].Biswas M, Chan JY. Role of Nrf1 in antioxidant response element-mediated gene expression and beyond. Toxicol Appl Pharmacol. 2010;244:16–20. doi: 10.1016/j.taap.2009.07.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [105].Chan JY, Kwong M, Lu R, Chang J, Wang B, Yen TS, Kan YW. Targeted disruption of the ubiquitous CNC-bZIP transcription factor, Nrf-1, results in anemia and embryonic lethality in mice. The EMBO journal. 1998;17:1779–1787. doi: 10.1093/emboj/17.6.1779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [106].Farmer SC, Sun CW, Winnier GE, Hogan BL, Townes TM. The bZIP transcription factor LCR-F1 is essential for mesoderm formation in mouse development. Genes & development. 1997;11:786–798. doi: 10.1101/gad.11.6.786. [DOI] [PubMed] [Google Scholar]
- [107].Chen L, Kwong M, Lu R, Ginzinger D, Lee C, Leung L, Chan JY. Nrf1 is critical for redox balance and survival of liver cells during development. Molecular and cellular biology. 2003;23:4673–4686. doi: 10.1128/MCB.23.13.4673-4686.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [108].Kwong M, Kan YW, Chan JY. The CNC basic leucine zipper factor, Nrf1, is essential for cell survival in response to oxidative stress-inducing agents. Role for Nrf1 in gamma-gcs(l) and gss expression in mouse fibroblasts. The Journal of biological chemistry. 1999;274:37491–37498. doi: 10.1074/jbc.274.52.37491. [DOI] [PubMed] [Google Scholar]
- [109].Venugopal R, Jaiswal AK. Nrf1 and Nrf2 positively and c-Fos and Fra1 negatively regulate the human antioxidant response element-mediated expression of NAD(P)H:quinone oxidoreductase1 gene. Proc Natl Acad Sci U S A. 1996;93:14960–14965. doi: 10.1073/pnas.93.25.14960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [110].Venugopal R, Jaiswal AK. Nrf2 and Nrf1 in association with Jun proteins regulate antioxidant response element-mediated expression and coordinated induction of genes encoding detoxifying enzymes. Oncogene. 1998;17:3145–3156. doi: 10.1038/sj.onc.1202237. [DOI] [PubMed] [Google Scholar]
- [111].Han W, Ming M, Zhao R, Pi J, Wu C, He YY. Nrf1 CNC-bZIP protein promotes cell survival and nucleotide excision repair through maintaining glutathione homeostasis. The Journal of biological chemistry. 2012;287:18788–18795. doi: 10.1074/jbc.M112.363614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [112].Tsujita T, Peirce V, Baird L, Matsuyama Y, Takaku M, Walsh SV, Griffin JL, Uruno A, Yamamoto M, Hayes JD. Transcription factor Nrf1 negatively regulates the cystine/glutamate transporter and lipid-metabolizing enzymes. Molecular and cellular biology. 2014;34:3800–3816. doi: 10.1128/MCB.00110-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [113].Wang W, Chan JY. Nrf1 is targeted to the endoplasmic reticulum membrane by an N-terminal transmembrane domain. Inhibition of nuclear translocation and transacting function. The Journal of biological chemistry. 2006;281:19676–19687. doi: 10.1074/jbc.M602802200. [DOI] [PubMed] [Google Scholar]
- [114].Hirotsu Y, Hataya N, Katsuoka F, Yamamoto M. NF-E2-related factor 1 (Nrf1) serves as a novel regulator of hepatic lipid metabolism through regulation of the Lipin1 and PGC-1beta genes. Molecular and cellular biology. 2012;32:2760–2770. doi: 10.1128/MCB.06706-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [115].Radhakrishnan SK, Lee CS, Young P, Beskow A, Chan JY, Deshaies RJ. Transcription factor Nrf1 mediates the proteasome recovery pathway after proteasome inhibition in mammalian cells. Molecular cell. 2010;38:17–28. doi: 10.1016/j.molcel.2010.02.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [116].Steffen J, Seeger M, Koch A, Kruger E. Proteasomal degradation is transcriptionally controlled by TCF11 via an ERAD-dependent feedback loop. Molecular cell. 2010;40:147–158. doi: 10.1016/j.molcel.2010.09.012. [DOI] [PubMed] [Google Scholar]
- [117].Moi P, Chan K, Asunis I, Cao A, Kan YW. Isolation of NF-E2-related factor 2 (Nrf2), a NF-E2-like basic leucine zipper transcriptional activator that binds to the tandem NF-E2/AP1 repeat of the beta-globin locus control region. Proc Natl Acad Sci U S A. 1994;91:9926–9930. doi: 10.1073/pnas.91.21.9926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [118].Itoh K, Igarashi K, Hayashi N, Nishizawa M, Yamamoto M. Cloning and characterization of a novel erythroid cell-derived CNC family transcription factor heterodimerizing with the small Maf family proteins. Molecular and cellular biology. 1995;15:4184–4193. doi: 10.1128/mcb.15.8.4184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [119].Nguyen T, Huang HC, Pickett CB. Transcriptional regulation of the antioxidant response element. Activation by Nrf2 and repression by MafK. The Journal of biological chemistry. 2000;275:15466–15473. doi: 10.1074/jbc.M000361200. [DOI] [PubMed] [Google Scholar]
- [120].Noda S, Harada N, Hida A, Fujii-Kuriyama Y, Motohashi H, Yamamoto M. Gene expression of detoxifying enzymes in AhR and Nrf2 compound null mutant mouse. Biochem Biophys Res Commun. 2003;303:105–111. doi: 10.1016/s0006-291x(03)00306-1. [DOI] [PubMed] [Google Scholar]
- [121].Alam J, Stewart D, Touchard C, Boinapally S, Choi AM, Cook JL. Nrf2, a Cap’n’Collar transcription factor, regulates induction of the heme oxygenase-1 gene. The Journal of biological chemistry. 1999;274:26071–26078. doi: 10.1074/jbc.274.37.26071. [DOI] [PubMed] [Google Scholar]
- [122].Pietsch EC, Chan JY, Torti FM, Torti SV. Nrf2 mediates the induction of ferritin H in response to xenobiotics and cancer chemopreventive dithiolethiones. The Journal of biological chemistry. 2003;278:2361–2369. doi: 10.1074/jbc.M210664200. [DOI] [PubMed] [Google Scholar]
- [123].McMahon M, Itoh K, Yamamoto M, Chanas SA, Henderson CJ, McLellan LI, Wolf CR, Cavin C, Hayes JD. The Cap’n’Collar basic leucine zipper transcription factor Nrf2 (NF-E2 p45-related factor 2) controls both constitutive and inducible expression of intestinal detoxification and glutathione biosynthetic enzymes. Cancer research. 2001;61:3299–3307. [PubMed] [Google Scholar]
- [124].Lee JM, Calkins MJ, Chan K, Kan YW, Johnson JA. Identification of the NF-E2-related factor-2-dependent genes conferring protection against oxidative stress in primary cortical astrocytes using oligonucleotide microarray analysis. The Journal of biological chemistry. 2003;278:12029–12038. doi: 10.1074/jbc.M211558200. [DOI] [PubMed] [Google Scholar]
- [125].Thimmulappa RK, Mai KH, Srisuma S, Kensler TW, Yamamoto M, Biswal S. Identification of Nrf2-regulated genes induced by the chemopreventive agent sulforaphane by oligonucleotide microarray. Cancer research. 2002;62:5196–5203. [PubMed] [Google Scholar]
- [126].Chan K, Lu R, Chang JC, Kan YW. NRF2, a member of the NFE2 family of transcription factors, is not essential for murine erythropoiesis, growth, and development. Proc Natl Acad Sci U S A. 1996;93:13943–13948. doi: 10.1073/pnas.93.24.13943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [127].Wakabayashi N, Itoh K, Wakabayashi J, Motohashi H, Noda S, Takahashi S, Imakado S, Kotsuji T, Otsuka F, Roop DR, Harada T, Engel JD, Yamamoto M. Keap1-null mutation leads to postnatal lethality due to constitutive Nrf2 activation. Nature genetics. 2003;35:238–245. doi: 10.1038/ng1248. [DOI] [PubMed] [Google Scholar]
- [128].Leung L, Kwong M, Hou S, Lee C, Chan JY. Deficiency of the Nrf1 and Nrf2 transcription factors results in early embryonic lethality and severe oxidative stress. J Biol Chem. 2003;278:48021–48029. doi: 10.1074/jbc.M308439200. [DOI] [PubMed] [Google Scholar]
- [129].Chan K, Kan YW. Nrf2 is essential for protection against acute pulmonary injury in mice. Proc Natl Acad Sci U S A. 1999;96:12731–12736. doi: 10.1073/pnas.96.22.12731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [130].Chan K, Han XD, Kan YW. An important function of Nrf2 in combating oxidative stress: detoxification of acetaminophen. Proc Natl Acad Sci U S A. 2001;98:4611–4616. doi: 10.1073/pnas.081082098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [131].Enomoto A, Itoh K, Nagayoshi E, Haruta J, Kimura T, ’Connor T, Harada T, Yamamoto M. High sensitivity of Nrf2 knockout mice to acetaminophen hepatotoxicity associated with decreased expression of ARE-regulated drug metabolizing enzymes and antioxidant genes. Toxicol Sci. 2001;59:169–177. doi: 10.1093/toxsci/59.1.169. [DOI] [PubMed] [Google Scholar]
- [132].Aoki Y, Sato H, Nishimura N, Takahashi S, Itoh K, Yamamoto M. Accelerated DNA adduct formation in the lung of the Nrf2 knockout mouse exposed to diesel exhaust. Toxicol Appl Pharmacol. 2001;173:154–160. doi: 10.1006/taap.2001.9176. [DOI] [PubMed] [Google Scholar]
- [133].Ramos-Gomez M, Kwak MK, Dolan PM, Itoh K, Yamamoto M, Talalay P, Kensler TW. Sensitivity to carcinogenesis is increased and chemoprotective efficacy of enzyme inducers is lost in nrf2 transcription factor-deficient mice. Proc Natl Acad Sci U S A. 2001;98:3410–3415. doi: 10.1073/pnas.051618798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [134].Ramos-Gomez M, Dolan PM, Itoh K, Yamamoto M, Kensler TW. Interactive effects of nrf2 genotype and oltipraz on benzo[a]pyrene-DNA adducts and tumor yield in mice. Carcinogenesis. 2003;24:461–467. doi: 10.1093/carcin/24.3.461. [DOI] [PubMed] [Google Scholar]
- [135].Kobayashi A, Ito E, Toki T, Kogame K, Takahashi S, Igarashi K, Hayashi N, Yamamoto M. Molecular cloning and functional characterization of a new Cap’n’ collar family transcription factor Nrf3. The Journal of biological chemistry. 1999;274:6443–6452. doi: 10.1074/jbc.274.10.6443. [DOI] [PubMed] [Google Scholar]
- [136].Chenais B, Derjuga A, Massrieh W, Red-Horse K, Bellingard V, Fisher SJ, Blank V. Functional and placental expression analysis of the human NRF3 transcription factor. Mol Endocrinol. 2005;19:125–137. doi: 10.1210/me.2003-0379. [DOI] [PubMed] [Google Scholar]
- [137].Zhang Y, Kobayashi A, Yamamoto M, Hayes JD. The Nrf3 transcription factor is a membrane-bound glycoprotein targeted to the endoplasmic reticulum through its N-terminal homology box 1 sequence. The Journal of biological chemistry. 2009;284:3195–3210. doi: 10.1074/jbc.M805337200. [DOI] [PubMed] [Google Scholar]
- [138].Chevillard G, Blank V. NFE2L3 (NRF3): the Cinderella of the Cap’n’Collar transcription factors. Cell Mol Life Sci. 2011;68:3337–3348. doi: 10.1007/s00018-011-0747-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [139].Derjuga A, Gourley TS, Holm TM, Heng HH, Shivdasani RA, Ahmed R, Andrews NC, Blank V. Complexity of CNC transcription factors as revealed by gene targeting of the Nrf3 locus. Molecular and cellular biology. 2004;24:3286–3294. doi: 10.1128/MCB.24.8.3286-3294.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [140].Chevillard G, Paquet M, Blank V. Nfe2l3 (Nrf3) deficiency predisposes mice to T-cell lymphoblastic lymphoma. Blood. 2011;117:2005–2008. doi: 10.1182/blood-2010-02-271460. [DOI] [PubMed] [Google Scholar]
- [141].Sankaranarayanan K, Jaiswal AK. Nrf3 negatively regulates antioxidant-response element-mediated expression and antioxidant induction of NAD(P)H:quinone oxidoreductase1 gene. The Journal of biological chemistry. 2004;279:50810–50817. doi: 10.1074/jbc.M404984200. [DOI] [PubMed] [Google Scholar]
- [142].Sive H. ‘Model’ or ‘tool’? New definitions for translational research. Dis Model Mech. 2011;4:137–138. doi: 10.1242/dmm.007666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [143].Brown KS. Making a splash with zebrafish. Bioscience. 1997;47:68–71. [Google Scholar]
- [144].Eisen JS. Zebrafish make a big splash. Cell. 1996;87:969–977. doi: 10.1016/s0092-8674(00)81792-4. [DOI] [PubMed] [Google Scholar]
- [145].Roush W. Zebrafish embryology builds a better model vertebrate. Science. 1996;272:1103. doi: 10.1126/science.272.5265.1103. [DOI] [PubMed] [Google Scholar]
- [146].Briggs JP. The zebrafish: a new model organism for integrative physiology. Am J Physiol Regul Integr Comp Physiol. 2002;282:R3–9. doi: 10.1152/ajpregu.00589.2001. [DOI] [PubMed] [Google Scholar]
- [147].Dodd A, Curtis PM, Williams LC, Love DR. Zebrafish: bridging the gap between development and disease. Human molecular genetics. 2000;9:2443–2449. doi: 10.1093/hmg/9.16.2443. [DOI] [PubMed] [Google Scholar]
- [148].Dooley K, Zon LI. Zebrafish: a model system for the study of human disease. Current opinion in genetics & development. 2000;10:252–256. doi: 10.1016/s0959-437x(00)00074-5. [DOI] [PubMed] [Google Scholar]
- [149].Paw BH, Zon LI. Zebrafish: a genetic approach in studying hematopoiesis. Curr Opin Hematol. 2000;7:79–84. doi: 10.1097/00062752-200003000-00002. [DOI] [PubMed] [Google Scholar]
- [150].Barut BA, Zon LI. Realizing the potential of zebrafish as a model for human disease. Physiological genomics. 2000;2:49–51. doi: 10.1152/physiolgenomics.2000.2.2.49. [DOI] [PubMed] [Google Scholar]
- [151].Thisse C, Zon LI. Organogenesis--heart and blood formation from the zebrafish point of view. Science. 2002;295:457–462. doi: 10.1126/science.1063654. [DOI] [PubMed] [Google Scholar]
- [152].Talbot WS, Hopkins N. Zebrafish mutations and functional analysis of the vertebrate genome. Genes & development. 2000;14:755–762. [PubMed] [Google Scholar]
- [153].Fishman MC. Genomics. Zebrafish--the canonical vertebrate. Science. 2001;294:1290–1291. doi: 10.1126/science.1066652. [DOI] [PubMed] [Google Scholar]
- [154].Ward AC, Lieschke GJ. The zebrafish as a model system for human disease. Front Biosci. 2002;7:d827–833. doi: 10.2741/A814. [DOI] [PubMed] [Google Scholar]
- [155].Amsterdam A, Hopkins N. Mutagenesis strategies in zebrafish for identifying genes involved in development and disease. Trends Genet. 2006;22:473–478. doi: 10.1016/j.tig.2006.06.011. [DOI] [PubMed] [Google Scholar]
- [156].Shin JT, Fishman MC. From Zebrafish to human: modular medical models. Annu Rev Genomics Hum Genet. 2002;3:311–340. doi: 10.1146/annurev.genom.3.031402.131506. [DOI] [PubMed] [Google Scholar]
- [157].Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of embryonic development of the zebrafish. Developmental Dynamics. 1995;203:253–310. doi: 10.1002/aja.1002030302. [DOI] [PubMed] [Google Scholar]
- [158].Udvadia AJ, Linney E. Windows into development: historic, current, and future perspectives on transgenic zebrafish. Developmental biology. 2003;256:1–17. doi: 10.1016/s0012-1606(02)00083-0. [DOI] [PubMed] [Google Scholar]
- [159].Motoike T, Loughna S, Perens E, Roman BL, Liao W, Chau TC, Richardson CD, Kawate T, Kuno J, Weinstein BM, Stainier DY, Sato TN. Universal GFP reporter for the study of vascular development. Genesis. 2000;28:75–81. doi: 10.1002/1526-968x(200010)28:2<75::aid-gene50>3.0.co;2-s. [DOI] [PubMed] [Google Scholar]
- [160].Long Q, Meng A, Wang H, Jessen JR, Farrell MJ, Lin S. GATA-1 expression pattern can be recapitulated in living transgenic zebrafish using GFP reporter gene. Development (Cambridge, England) 1997;124:4105–4111. doi: 10.1242/dev.124.20.4105. [DOI] [PubMed] [Google Scholar]
- [161].Jessen JR, Meng A, McFarlane RJ, Paw BH, Zon LI, Smith GR, Lin S. Modification of bacterial artificial chromosomes through chi-stimulated homologous recombination and its application in zebrafish transgenesis. Proc Natl Acad Sci U S A. 1998;95:5121–5126. doi: 10.1073/pnas.95.9.5121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [162].Barton LM, Gottgens B, Gering M, Gilbert JG, Grafham D, Rogers J, Bentley D, Patient R, Green AR. Regulation of the stem cell leukemia (SCL) gene: a tale of two fishes. Proc Natl Acad Sci U S A. 2001;98:6747–6752. doi: 10.1073/pnas.101532998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [163].Meng A, Tang H, Ong BA, Farrell MJ, Lin S. Promoter analysis in living zebrafish embryos identifies a cis-acting motif required for neuronal expression of GATA-2. Proc Natl Acad Sci U S A. 1997;94:6267–6272. doi: 10.1073/pnas.94.12.6267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [164].Meng A, Tang H, Yuan B, Ong BA, Long Q, Lin S. Positive and negative cis-acting elements are required for hematopoietic expression of zebrafish GATA-1. Blood. 1999;93:500–508. [PubMed] [Google Scholar]
- [165].Porcher C, Liao EC, Fujiwara Y, Zon LI, Orkin SH. Specification of hematopoietic and vascular development by the bHLH transcription factor SCL without direct DNA binding. Development (Cambridge, England) 1999;126:4603–4615. doi: 10.1242/dev.126.20.4603. [DOI] [PubMed] [Google Scholar]
- [166].Amsterdam A, Lin S, Hopkins N. The Aequorea victoria green fluorescent protein can be used as a reporter in live zebrafish embryos. Developmental biology. 1995;171:123–129. doi: 10.1006/dbio.1995.1265. [DOI] [PubMed] [Google Scholar]
- [167].Finley KR, Davidson AE, Ekker SC. Three-color imaging using fluorescent proteins in living zebrafish embryos. BioTechniques. 2001;31:66–70. 72. doi: 10.2144/01311st02. [DOI] [PubMed] [Google Scholar]
- [168].Gibbs PDL, Schmale MC. GFP as a Genetic Marker Scorable Throughout the Life Cycle of Transgenic Zebra Fish. Mar. Biotechnol. 2000;2:107–125. doi: 10.1007/s101269900014. [DOI] [PubMed] [Google Scholar]
- [169].Gaiano N, Hopkins N. Introducing genes into zebrafish. Biochim Biophys Acta. 1996;1288:O11–O14. doi: 10.1016/0304-419x(96)00017-0. [DOI] [PubMed] [Google Scholar]
- [170].Amsterdam A, Burgess S, Golling G, Chen W, Sun Z, Townsend K, Farrington S, Haldi M, Hopkins N. A large-scale insertional mutagenesis screen in zebrafish. Genes & development. 1999;13:2713–2724. doi: 10.1101/gad.13.20.2713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [171].Driever W, Solnica-Krezel L, Schier AF, Neuhauss SC, Malicki J, Stemple DL, Stainier DY, Zwartkruis F, Abdelilah S, Rangini Z, Belak J, Boggs C. A genetic screen for mutations affecting embryogenesis in zebrafish. Development (Cambridge, England) 1996;123:37–46. doi: 10.1242/dev.123.1.37. [DOI] [PubMed] [Google Scholar]
- [172].Haffter P, Granato M, Brand M, Mullins MC, Hammerschmidt M, Kane DA, Odenthal J, van Eeden FJ, Jiang YJ, Heisenberg CP, Kelsh RN, Furutani-Seiki M, Vogelsang E, Beuchle D, Schach U, Fabian C, Nusslein-Volhard C. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development (Cambridge, England) 1996;123:1–36. doi: 10.1242/dev.123.1.1. [DOI] [PubMed] [Google Scholar]
- [173].Wang H, Long Q, Marty SD, Sassa S, Lin S. A zebrafish model for hepatoerythropoietic porphyria. Nature genetics. 1998;20:239–243. doi: 10.1038/3041. [DOI] [PubMed] [Google Scholar]
- [174].Childs S, Weinstein BM, Mohideen MA, Donohue S, Bonkovsky H, Fishman MC. Zebrafish dracula encodes ferrochelatase and its mutation provides a model for erythropoietic protoporphyria. Curr Biol. 2000;10:1001–1004. doi: 10.1016/s0960-9822(00)00653-9. [DOI] [PubMed] [Google Scholar]
- [175].Nasevicius A, Ekker SC. Effective targeted gene ‘knockdown’ in zebrafish. Nature genetics. 2000;26:216–220. doi: 10.1038/79951. [DOI] [PubMed] [Google Scholar]
- [176].Anonymous Targeting zebrafish. Nature genetics. 2000;26:129–130. doi: 10.1038/79826. [DOI] [PubMed] [Google Scholar]
- [177].Sumanas S, Larson JD. Morpholino phosphorodiamidate oligonucleotides in zebrafish: a recipe for functional genomics? Briefings in functional genomics & proteomics. 2002;1:239–256. doi: 10.1093/bfgp/1.3.239. [DOI] [PubMed] [Google Scholar]
- [178].Nasevicius A, Larson J, Ekker SC. Distinct requirements for zebrafish angiogenesis revealed by a VEGF-A morphant. Yeast. 2000;17:294–301. doi: 10.1002/1097-0061(200012)17:4<294::AID-YEA54>3.0.CO;2-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [179].Zhong TP, Childs S, Leu JP, Fishman MC. Gridlock signalling pathway fashions the first embryonic artery. Nature. 2001;414:216–220. doi: 10.1038/35102599. [DOI] [PubMed] [Google Scholar]
- [180].Scholpp S, Brand M. Morpholino-induced knockdown of zebrafish engrailed genes eng2 and eng3 reveals redundant and unique functions in midbrain--hindbrain boundary development. Genesis. 2001;30:129–133. doi: 10.1002/gene.1047. [DOI] [PubMed] [Google Scholar]
- [181].Bauer H, Lele Z, Rauch GJ, Geisler R, Hammerschmidt M. The type I serine/threonine kinase receptor Alk8/Lost-a-fin is required for Bmp2b/7 signal transduction during dorsoventral patterning of the zebrafish embryo. Development (Cambridge, England) 2001;128:849–858. doi: 10.1242/dev.128.6.849. [DOI] [PubMed] [Google Scholar]
- [182].Ekker SC, Larson JD. Morphant technology in model developmental systems. Genesis. 2001;30:89–93. doi: 10.1002/gene.1038. [DOI] [PubMed] [Google Scholar]
- [183].Lele Z, Bakkers J, Hammerschmidt M. Morpholino phenocopies of the swirl, snailhouse, somitabun, minifin, silberblick, and pipetail mutations. Genesis. 2001;30:190–194. doi: 10.1002/gene.1063. [DOI] [PubMed] [Google Scholar]
- [184].Mathew LK, Andreasen EA, Tanguay RL. Aryl hydrocarbon receptor activation inhibits regenerative growth. Mol Pharmacol. 2006;69:257–265. doi: 10.1124/mol.105.018044. [DOI] [PubMed] [Google Scholar]
- [185].Carney SA, Peterson RE, Heideman W. 2,3,7,8-Tetrachlorodibenzo-p-dioxin activation of the aryl hydrocarbon receptor/aryl hydrocarbon receptor nuclear translocator pathway causes developmental toxicity through a CYP1A-independent mechanism in zebrafish. Mol Pharmacol. 2004;66:512–521. doi: 10.1124/mol.66.3.. [DOI] [PubMed] [Google Scholar]
- [186].Antkiewicz DS, Peterson RE, Heideman W. Blocking Expression of AHR2 and ARNT1 in Zebrafish Larvae Protects Against Cardiac Toxicity of 2,3,7,8-Tetrachlorodibenzo-p-dioxin. Toxicol Sci. 2006;94:175–182. doi: 10.1093/toxsci/kfl093. [DOI] [PubMed] [Google Scholar]
- [187].Prasch AL, Tanguay RL, Mehta V, Heideman W, Peterson RE. Identification of zebrafish ARNT1 homologs: 2,3,7,8-tetrachlorodibenzo-p-dioxin toxicity in the developing zebrafish requires ARNT1. Mol Pharmacol. 2006;69:776–787. doi: 10.1124/mol.105.016873. [DOI] [PubMed] [Google Scholar]
- [188].Tseng HP, Hseu TH, Buhler DR, Wang WD, Hu CH. Constitutive and xenobiotics-induced expression of a novel CYP3A gene from zebrafish larva. Toxicol Appl Pharmacol. 2005;205:247–258. doi: 10.1016/j.taap.2004.10.019. [DOI] [PubMed] [Google Scholar]
- [189].Prasch AL, Teraoka H, Carney SA, Dong W, Hiraga T, Stegeman JJ, Heideman W, Peterson RE. Aryl Hydrocarbon Receptor 2 Mediates 2,3,7,8-Tetrachlorodibenzo-p-dioxin Developmental Toxicity in Zebrafish. Toxicol Sci. 2003;76:138–150. doi: 10.1093/toxsci/kfg202. [DOI] [PubMed] [Google Scholar]
- [190].Billiard SM, Timme-Laragy AR, Wassenberg DM, Cockman C, Di Giulio RT. The role of the aryl hydrocarbon receptor pathway in mediating synergistic developmental toxicity of polycyclic aromatic hydrocarbons to zebrafish. Toxicol Sci. 2006;92:526–536. doi: 10.1093/toxsci/kfl011. [DOI] [PubMed] [Google Scholar]
- [191].Incardona JP, Day HL, Collier TK, Scholz NL. Developmental toxicity of 4-ring polycyclic aromatic hydrocarbons in zebrafish is differentially dependent on AH receptor isoforms and hepatic cytochrome P4501A metabolism. Toxicol Appl Pharmacol. 2006;217:308–321. doi: 10.1016/j.taap.2006.09.018. [DOI] [PubMed] [Google Scholar]
- [192].Doyon Y, McCammon JM, Miller JC, Faraji F, Ngo C, Katibah GE, Amora R, Hocking TD, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Amacher SL. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nature biotechnology. 2008;26:702–708. doi: 10.1038/nbt1409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [193].Meng X, Noyes MB, Zhu LJ, Lawson ND, Wolfe SA. Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nature biotechnology. 2008;26:695–701. doi: 10.1038/nbt1398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [194].Foley JE, Yeh JR, Maeder ML, Reyon D, Sander JD, Peterson RT, Joung JK. Rapid mutation of endogenous zebrafish genes using zinc finger nucleases made by Oligomerized Pool ENgineering (OPEN) PLoS ONE. 2009;4:e4348. doi: 10.1371/journal.pone.0004348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [195].Joung JK, Sander JD. TALENs: a widely applicable technology for targeted genome editing. Nature reviews. Molecular cell biology. 2013;14:49–55. doi: 10.1038/nrm3486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [196].Sander JD, Joung JK. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature biotechnology. 2014;32:347–355. doi: 10.1038/nbt.2842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [197].Hruscha A, Krawitz P, Rechenberg A, Heinrich V, Hecht J, Haass C, Schmid B. Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development (Cambridge, England) 2013;140:4982–4987. doi: 10.1242/dev.099085. [DOI] [PubMed] [Google Scholar]
- [198].Schulte-Merker S, Stainier DY. Out with the old, in with the new: reassessing morpholino knockdowns in light of genome editing technology. Development (Cambridge, England) 2014;141:3103–3104. doi: 10.1242/dev.112003. [DOI] [PubMed] [Google Scholar]
- [199].Stainier DY, Kontarakis Z, Rossi A. Making sense of anti-sense data. Developmental cell. 2015;32:7–8. doi: 10.1016/j.devcel.2014.12.012. [DOI] [PubMed] [Google Scholar]
- [200].Kok FO, Shin M, Ni CW, Gupta A, Grosse AS, van Impel A, Kirchmaier BC, Peterson-Maduro J, Kourkoulis G, Male I, DeSantis DF, Sheppard-Tindell S, Ebarasi L, Betsholtz C, Schulte-Merker S, Wolfe SA, Lawson ND. Reverse genetic screening reveals poor correlation between morpholino-induced and mutant phenotypes in zebrafish. Developmental cell. 2015;32:97–108. doi: 10.1016/j.devcel.2014.11.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [201].Coverdale LE, Lean D, Martin CC. Not just a fishing trip - Environmental genomics using zebrafish. Current Genomics. 2004;5:299–308. [Google Scholar]
- [202].Howe K, Clark MD, Torroja CF, Torrance J, Berthelot C, Muffato M, Collins JE, Humphray S, McLaren K, Matthews L, McLaren S, Sealy I, Caccamo M, Churcher C, Scott C, Barrett JC, Koch R, Rauch GJ, White S, Chow W, Kilian B, Quintais LT, Guerra-Assuncao JA, Zhou Y, Gu Y, Yen J, Vogel JH, Eyre T, Redmond S, Banerjee R, Chi J, Fu B, Langley E, Maguire SF, Laird GK, Lloyd D, Kenyon E, Donaldson S, Sehra H, Almeida-King J, Loveland J, Trevanion S, Jones M, Quail M, Willey D, Hunt A, Burton J, Sims S, McLay K, Plumb B, Davis J, Clee C, Oliver K, Clark R, Riddle C, Elliot D, Threadgold G, Harden G, Ware D, Begum S, Mortimore B, Kerry G, Heath P, Phillimore B, Tracey A, Corby N, Dunn M, Johnson C, Wood J, Clark S, Pelan S, Griffiths G, Smith M, Glithero R, Howden P, Barker N, Lloyd C, Stevens C, Harley J, Holt K, Panagiotidis G, Lovell J, Beasley H, Henderson C, Gordon D, Auger K, Wright D, Collins J, Raisen C, Dyer L, Leung K, Robertson L, Ambridge K, Leongamornlert D, McGuire S, Gilderthorp R, Griffiths C, Manthravadi D, Nichol S, Barker G, Whitehead S, Kay M, Brown J, Murnane C, Gray E, Humphries M, Sycamore N, Barker D, Saunders D, Wallis J, Babbage A, Hammond S, Mashreghi-Mohammadi M, Barr L, Martin S, Wray P, Ellington A, Matthews N, Ellwood M, Woodmansey R, Clark G, Cooper J, Tromans A, Grafham D, Skuce C, Pandian R, Andrews R, Harrison E, Kimberley A, Garnett J, Fosker N, Hall R, Garner P, Kelly D, Bird C, Palmer S, Gehring I, Berger A, Dooley CM, Ersan-Urun Z, Eser C, Geiger H, Geisler M, Karotki L, Kirn A, Konantz J, Konantz M, Oberlander M, Rudolph-Geiger S, Teucke M, Lanz C, Raddatz G, Osoegawa K, Zhu B, Rapp A, Widaa S, Langford C, Yang F, Schuster SC, Carter NP, Harrow J, Ning Z, Herrero J, Searle SM, Enright A, Geisler R, Plasterk RH, Lee C, Westerfield M, de Jong PJ, Zon LI, Postlethwait JH, Nusslein-Volhard C, Hubbard TJ, Roest Crollius H, Rogers J, Stemple DL. The zebrafish reference genome sequence and its relationship to the human genome. Nature. 2013;496:498–503. doi: 10.1038/nature12111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [203].Woods IG, Wilson C, Friedlander B, Chang P, Reyes DK, Nix R, Kelly PD, Chu F, Postlethwait JH, Talbot WS. The zebrafish gene map defines ancestral vertebrate chromosomes. Genome research. 2005;15:1307–1314. doi: 10.1101/gr.4134305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [204].Phillips RB, Amores A, Morasch MR, Wilson C, Postlethwait JH. Assignment of zebrafish genetic linkage groups to chromosomes. Cytogenetic and genome research. 2006;114:155–162. doi: 10.1159/000093332. [DOI] [PubMed] [Google Scholar]
- [205].Amores A, Force A, Yan Y-L, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang Y-L, Westerfield M, Ekker M, Postlethwait JH. Zebrafish hox clusters and vertebrate genome evolution. Science. 1998;282:1711–1714. doi: 10.1126/science.282.5394.1711. [DOI] [PubMed] [Google Scholar]
- [206].Taylor JS, Van de Peer Y, Braasch I, Meyer A. Comparative genomics provides evidence for an ancient genome duplication event in fish. Phil. Trans. R. Soc. Lond. B. 2001;356:1661–1679. doi: 10.1098/rstb.2001.0975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [207].Postlethwait J, Amores A, Cresko W, Singer A, Yan YL. Subfunction partitioning, the teleost radiation and the annotation of the human genome. Trends Genet. 2004;20:481–490. doi: 10.1016/j.tig.2004.08.001. [DOI] [PubMed] [Google Scholar]
- [208].Christoffels A, Koh EG, Chia JM, Brenner S, Aparicio S, Venkatesh B. Fugu genome analysis provides evidence for a whole-genome duplication early during the evolution of ray-finned fishes. Molecular biology and evolution. 2004;21:1146–1151. doi: 10.1093/molbev/msh114. [DOI] [PubMed] [Google Scholar]
- [209].Force A, Lynch M, Pickett FB, Amores A, Yan Y-L, Postlethwait JH. Preservation of duplicate genes by complementary, degenerative mutations. Genetics. 1999;151:1531–1545. doi: 10.1093/genetics/151.4.1531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [210].Lynch M, Force A. The probability of duplicate gene preservation by subfunctionalization. Genetics. 2000;154:459–473. doi: 10.1093/genetics/154.1.459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [211].Sipes NS, Padilla S, Knudsen TB. Zebrafish: as an integrative model for twenty-first century toxicity testing. Birth Defects Res C Embryo Today. 2011;93:256–267. doi: 10.1002/bdrc.20214. [DOI] [PubMed] [Google Scholar]
- [212].Padilla S, Corum D, Padnos B, Hunter DL, Beam A, Houck KA, Sipes N, Kleinstreuer N, Knudsen T, Dix DJ, Reif DM. Zebrafish developmental screening of the ToxCast Phase I chemical library. Reproductive toxicology (Elmsford, N.Y. 2012;33:174–187. doi: 10.1016/j.reprotox.2011.10.018. [DOI] [PubMed] [Google Scholar]
- [213].Lammer E, Carr GJ, Wendler K, Rawlings JM, Belanger SE, Braunbeck T. Is the fish embryo toxicity test (FET) with the zebrafish (Danio rerio) a potential alternative for the fish acute toxicity test? Comp Biochem Physiol C Toxicol Pharmacol. 2009;149:196–209. doi: 10.1016/j.cbpc.2008.11.006. [DOI] [PubMed] [Google Scholar]
- [214].Embry MR, Belanger SE, Braunbeck TA, Galay-Burgos M, Halder M, Hinton DE, Leonard MA, Lillicrap A, Norberg-King T, Whale G. The fish embryo toxicity test as an animal alternative method in hazard and risk assessment and scientific research. Aquat Toxicol. 2010;97:79–87. doi: 10.1016/j.aquatox.2009.12.008. [DOI] [PubMed] [Google Scholar]
- [215].Selderslaghs IW, Van Rompay AR, De Coen W, Witters HE. Development of a screening assay to identify teratogenic and embryotoxic chemicals using the zebrafish embryo. Reproductive toxicology (Elmsford, N.Y. 2009;28:308–320. doi: 10.1016/j.reprotox.2009.05.004. [DOI] [PubMed] [Google Scholar]
- [216].Truong L, Reif DM, St Mary L, Geier MC, Truong HD, Tanguay RL. Multidimensional in vivo hazard assessment using zebrafish. Toxicol Sci. 2014;137:212–233. doi: 10.1093/toxsci/kft235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [217].Brannen KC, Panzica-Kelly JM, Danberry TL, Augustine-Rauch KA. Development of a zebrafish embryo teratogenicity assay and quantitative prediction model. Birth Defects Res B Dev Reprod Toxicol. 2010;89:66–77. doi: 10.1002/bdrb.20223. [DOI] [PubMed] [Google Scholar]
- [218].Panzica-Kelly JM, Zhang CX, Danberry TL, Flood A, DeLan JW, Brannen KC, Augustine-Rauch KA. Morphological score assignment guidelines for the dechorionated zebrafish teratogenicity assay. Birth Defects Res B Dev Reprod Toxicol. 2010;89:382–395. doi: 10.1002/bdrb.20260. [DOI] [PubMed] [Google Scholar]
- [219].McGrath P, Li CQ. Zebrafish: a predictive model for assessing drug-induced toxicity. Drug discovery today. 2008;13:394–401. doi: 10.1016/j.drudis.2008.03.002. [DOI] [PubMed] [Google Scholar]
- [220].Parng C, Seng WL, Semino C, McGrath P. Zebrafish: a preclinical model for drug screening. Assay and drug development technologies. 2002;1:41–48. doi: 10.1089/154065802761001293. [DOI] [PubMed] [Google Scholar]
- [221].Ball JS, Stedman DB, Hillegass JM, Zhang CX, Panzica-Kelly J, Coburn A, Enright BP, Tornesi B, Amouzadeh HR, Hetheridge M, Gustafson AL, Augustine-Rauch KA. Fishing for teratogens: a consortium effort for a harmonized zebrafish developmental toxicology assay. Toxicol Sci. 2014;139:210–219. doi: 10.1093/toxsci/kfu017. [DOI] [PubMed] [Google Scholar]
- [222].Hill A, Mesens N, Steemans M, Xu JJ, Aleo MD. Comparisons between in vitro whole cell imaging and in vivo zebrafish-based approaches for identifying potential human hepatotoxicants earlier in pharmaceutical development. Drug metabolism reviews. 2012;44:127–140. doi: 10.3109/03602532.2011.645578. [DOI] [PubMed] [Google Scholar]
- [223].Panzica-Kelly JM, Zhang CX, Augustine-Rauch K. Zebrafish embryo developmental toxicology assay. Methods in molecular biology (Clifton, N.J. 2012;889:25–50. doi: 10.1007/978-1-61779-867-2_4. [DOI] [PubMed] [Google Scholar]
- [224].Augustine-Rauch K, Zhang CX, Panzica-Kelly JM. In vitro developmental toxicology assays: A review of the state of the science of rodent and zebrafish whole embryo culture and embryonic stem cell assays. Birth Defects Res C Embryo Today. 2010;90:87–98. doi: 10.1002/bdrc.20175. [DOI] [PubMed] [Google Scholar]
- [225].Carvan MJ, III, Solis WA, Gedamu L, Nebert DW. Activation of transcription factors in zebrafish cell cultures by environmental pollutants. Arch Biochem Biophys. 2000;376:320–327. doi: 10.1006/abbi.2000.1727. [DOI] [PubMed] [Google Scholar]
- [226].Carvan MJ, III, Sonntag DM, Cmar CB, Cook RS, Curran MA, Miller GL. Oxidative stress in zebrafish cells: potential utility of transgenic zebrafish as a deployable sentinel for site hazard ranking. Science of the Total Environment. 2001;274:183–196. doi: 10.1016/s0048-9697(01)00742-2. [DOI] [PubMed] [Google Scholar]
- [227].Kobayashi M, Itoh K, Suzuki T, Osanai H, Nishikawa K, Katoh Y, Takagi Y, Yamamoto M. Identification of the interactive interface and phylogenic conservation of the Nrf2-Keap1 system. Genes Cells. 2002;7:807–820. doi: 10.1046/j.1365-2443.2002.00561.x. [DOI] [PubMed] [Google Scholar]
- [228].Suzuki T, Takagi Y, Osanai H, Li L, Takeuchi M, Katoh Y, Kobayashi M, Yamamoto M. Pi class glutathione S-transferase genes are regulated by Nrf 2 through an evolutionarily conserved regulatory element in zebrafish. The Biochemical journal. 2005;388:65–73. doi: 10.1042/BJ20041860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [229].Li L, Kobayashi M, Kaneko H, Nakajima-Takagi Y, Nakayama Y, Yamamoto M. Molecular Evolution of Keap1: Two Keap1 molecules with distinctive intervening region structures are conserved among fish. The Journal of biological chemistry. 2008;283:3248–3255. doi: 10.1074/jbc.M708702200. [DOI] [PubMed] [Google Scholar]
- [230].Iida A, Sakaguchi K, Sato K, Sakurai H, Nishimura D, Iwaki A, Takeuchi M, Kobayashi M, Misaki K, Yonemura S, Kawahara A, Sehara-Fujisawa A. Metalloprotease-dependent onset of blood circulation in zebrafish. Curr Biol. 2010;20:1110–1116. doi: 10.1016/j.cub.2010.04.052. [DOI] [PubMed] [Google Scholar]
- [231].Barrionuevo WR, Burggren WW. O2 consumption and heart rate in developing zebrafish (Danio rerio): influence of temperature and ambient O2. Am J Physiol. 1999;276:R505–513. doi: 10.1152/ajpregu.1999.276.2.R505. [DOI] [PubMed] [Google Scholar]
- [232].Strecker R, Seiler TB, Hollert H, Braunbeck T. Oxygen requirements of zebrafish (Danio rerio) embryos in embryo toxicity tests with environmental samples. Comp Biochem Physiol C Toxicol Pharmacol. 2011;153:318–327. doi: 10.1016/j.cbpc.2010.12.002. [DOI] [PubMed] [Google Scholar]
- [233].Catt JW, Henman M. Toxic effects of oxygen on human embryo development. Human reproduction (Oxford, England) 2000;15(Suppl 2):199–206. doi: 10.1093/humrep/15.suppl_2.199. [DOI] [PubMed] [Google Scholar]
- [234].Timme-Laragy AR, Karchner SI, Franks DG, Jenny MJ, Harbeitner RC, Goldstone JV, McArthur AG, Hahn ME. Nrf2b: a novel zebrafish paralog of the oxidant-responsive transcription factor NF-E2-related factor 2 (NRF2) J Biol Chem. 2012;287:4609–4627. doi: 10.1074/jbc.M111.260125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [235].Timme-Laragy AR, Van Tiem LA, Linney EA, Di Giulio RT. Antioxidant responses and NRF2 in synergistic developmental toxicity of PAHs in zebrafish. Toxicol Sci. 2009;109:217–227. doi: 10.1093/toxsci/kfp038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [236].Yang L, Kemadjou JR, Zinsmeister C, Legradi J, Bauer M, Muller F, Pankratz M, Jakel J, Straehle U. Transcriptional profiling reveals barcode-like toxicogenomic responses in the zebrafish embryo. Genome biology. 2007;8:R227. doi: 10.1186/gb-2007-8-10-r227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [237].Hahn ME, McArthur AG, Karchner SI, Franks DG, Jenny MJ, Timme-Laragy AR, Stegeman JJ, Woodin BR, Cipriano MJ, Linney E. The Transcriptional Response to Oxidative Stress During Vertebrate Development: Effects of tert-Butylhydroquinone and 2,3,7,8-Tetrachlorodibenzo-p-dioxin. PLoS ONE. 2014;9:e113158. doi: 10.1371/journal.pone.0113158. doi:113110.111371/journal.pone.0113158; [DOI] [PMC free article] [PubMed] [Google Scholar]
- [238].Takagi Y, Kobayashi M, Li L, Suzuki T, Nishikawa K, Yamamoto M. MafT, a new member of the small Maf protein family in zebrafish. Biochem Biophys Res Commun. 2004;320:62–69. doi: 10.1016/j.bbrc.2004.05.131. [DOI] [PubMed] [Google Scholar]
- [239].Pratt SJ, Drejer A, Foott H, Barut B, Brownlie A, Postlethwait J, Kato Y, Yamamoto M, Zon LI. Isolation and characterization of zebrafish NFE2. Physiological genomics. 2002;11:91–98. doi: 10.1152/physiolgenomics.00112.2001. [DOI] [PubMed] [Google Scholar]
- [240].Williams LM, Timme-Laragy AR, Goldstone JV, McArthur AG, Stegeman JJ, Smolowitz RM, Hahn ME. Developmental expression of the Nfe2-related factor (Nrf) transcription factor family. PLoS ONE. 2013;8:e79574. doi: 10.1371/journal.pone.0079574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [241].Song HD, Sun XJ, Deng M, Zhang GW, Zhou Y, Wu XY, Sheng Y, Chen Y, Ruan Z, Jiang CL, Fan HY, Zon LI, Kanki JP, Liu TX, Look AT, Chen Z. Hematopoietic gene expression profile in zebrafish kidney marrow. Proc Natl Acad Sci U S A. 2004;101:16240–16245. doi: 10.1073/pnas.0407241101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [242].Meyer A, Van de Peer Y. From 2R to 3R: evidence for a fish-specific genome duplication (FSGD) Bioessays. 2005;27:937–945. doi: 10.1002/bies.20293. [DOI] [PubMed] [Google Scholar]
- [243].Mukaigasa K, Nguyen LT, Li L, Nakajima H, Yamamoto M, Kobayashi M. Genetic Evidence of an Evolutionarily Conserved Role for Nrf2 in the Protection against Oxidative Stress. Mol Cell Biol. 2012;32:4455–4461. doi: 10.1128/MCB.00481-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [244].Timme-Laragy AR, Karchner SI, Harbeitner RC, McArthur AG, Hahn ME. Nrf2 gene regulation during oxidative stress in embryonic development. Toxicological Sciences (The Toxicologist Supplement) 2013 Abstract #694. [Google Scholar]
- [245].Kmita M, Duboule D. Organizing axes in time and space; 25 years of colinear tinkering. Science. 2003;301:331–333. doi: 10.1126/science.1085753. [DOI] [PubMed] [Google Scholar]
- [246].Lee AP, Koh EG, Tay A, Brenner S, Venkatesh B. Highly conserved syntenic blocks at the vertebrate Hox loci and conserved regulatory elements within and outside Hox gene clusters. Proc Natl Acad Sci U S A. 2006;103:6994–6999. doi: 10.1073/pnas.0601492103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [247].Sundstrom G, Larsson TA, Larhammar D. Phylogenetic and chromosomal analyses of multiple gene families syntenic with vertebrate Hox clusters. BMC evolutionary biology. 2008;8:254. doi: 10.1186/1471-2148-8-254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [248].Kikuta H, Fredman D, Rinkwitz S, Lenhard B, Becker TS. Retroviral enhancer detection insertions in zebrafish combined with comparative genomics reveal genomic regulatory blocks - a fundamental feature of vertebrate genomes. Genome biology. 2007;8(Suppl 1):S4. doi: 10.1186/gb-2007-8-s1-s4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [249].Kikuta H, Laplante M, Navratilova P, Komisarczuk AZ, Engstrom PG, Fredman D, Akalin A, Caccamo M, Sealy I, Howe K, Ghislain J, Pezeron G, Mourrain P, Ellingsen S, Oates AC, Thisse C, Thisse B, Foucher I, Adolf B, Geling A, Lenhard B, Becker TS. Genomic regulatory blocks encompass multiple neighboring genes and maintain conserved synteny in vertebrates. Genome research. 2007;17:545–555. doi: 10.1101/gr.6086307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [250].Zhang S, Xu M, Huang J, Tang L, Zhang Y, Wu J, Lin S, Wang H. Heme acts through the Bach1b/Nrf2a-MafK pathway to regulate exocrine peptidase precursor genes in porphyric zebrafish. Dis Model Mech. 2014;7:837–845. doi: 10.1242/dmm.014951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [251].Fuse Y, Nakajima H, Nakajima-Takagi Y, Nakajima O, Kobayashi M. Heme-mediated inhibition of Bach1 regulates the liver specificity and transience of the Nrf2-dependent induction of zebrafish heme oxygenase 1. Genes Cells. 2015 doi: 10.1111/gtc.12249. [DOI] [PubMed] [Google Scholar]
- [252].Bardet PL, Horard B, Robinson-Rechavi M, Laudet V, Vanacker JM. Characterization of oestrogen receptors in zebrafish (Danio rerio) Journal of molecular endocrinology. 2002;28:153–163. doi: 10.1677/jme.0.0280153. [DOI] [PubMed] [Google Scholar]
- [253].Karchner SI, Franks DG, Hahn ME. AHR1B, a new functional aryl hydrocarbon receptor in zebrafish: tandem arrangement of ahr1b and ahr2 genes. Biochemical Journal. 2005;392:153–161. doi: 10.1042/BJ20050713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [254].Hansen JM, Harris C. Glutathione during embryonic development. Biochim Biophys Acta. 2014 doi: 10.1016/j.bbagen.2014.12.001. [DOI] [PubMed] [Google Scholar]
- [255].Dalton TP, Dieter MZ, Yang Y, Shertzer HG, Nebert DW. Knockout of the mouse glutamate cysteine ligase catalytic subunit (Gclc) gene: embryonic lethal when homozygous, and proposed model for moderate glutathione deficiency when heterozygous. Biochem Biophys Res Commun. 2000;279:324–329. doi: 10.1006/bbrc.2000.3930. [DOI] [PubMed] [Google Scholar]
- [256].Messina JP, Lawrence DA. Cell cycle progression of glutathione-depleted human peripheral blood mononuclear cells is inhibited at S phase. Journal of immunology. 1989;143:1974–1981. [PubMed] [Google Scholar]
- [257].Markovic J, Borras C, Ortega A, Sastre J, Vina J, Pallardo FV. Glutathione is recruited into the nucleus in early phases of cell proliferation. The Journal of biological chemistry. 2007;282:20416–20424. doi: 10.1074/jbc.M609582200. [DOI] [PubMed] [Google Scholar]
- [258].Pallardo FV, Markovic J, Garcia JL, Vina J. Role of nuclear glutathione as a key regulator of cell proliferation. Molecular aspects of medicine. 2009;30:77–85. doi: 10.1016/j.mam.2009.01.001. [DOI] [PubMed] [Google Scholar]
- [259].Garcia-Gimenez JL, Markovic J, Dasi F, Queval G, Schnaubelt D, Foyer CH, Pallardo FV. Nuclear glutathione. Biochimica et biophysica acta. 2013;1830:3304–3316. doi: 10.1016/j.bbagen.2012.10.005. [DOI] [PubMed] [Google Scholar]
- [260].Hansen JM, Carney EW, Harris C. Altered differentiation in rat and rabbit limb bud micromass cultures by glutathione modulating agents. Free radical biology & medicine. 2001;31:1582–1592. doi: 10.1016/s0891-5849(01)00751-1. [DOI] [PubMed] [Google Scholar]
- [261].Jones DP. Redox potential of GSH/GSSG couple: assay and biological significance. Methods in enzymology. 2002;348:93–112. doi: 10.1016/s0076-6879(02)48630-2. [DOI] [PubMed] [Google Scholar]
- [262].Kirlin WG, Cai J, Thompson SA, Diaz D, Kavanagh TJ, Jones DP. Glutathione redox potential in response to differentiation and enzyme inducers. Free radical biology & medicine. 1999;27:1208–1218. doi: 10.1016/s0891-5849(99)00145-8. [DOI] [PubMed] [Google Scholar]
- [263].Timme-Laragy AR, Goldstone JV, Imhoff BR, Stegeman JJ, Hahn ME, Hansen JM. Glutathione redox dynamics and expression of glutathione-related genes in the developing embryo. Free radical biology & medicine. 2013;65C:89–101. doi: 10.1016/j.freeradbiomed.2013.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [264].Hamre K, Penglase SJ, Rasinger JD, Skjaerven KH, Olsvik PA. Ontogeny of redox regulation in Atlantic cod (Gadus morhua) larvae. Free radical biology & medicine. 2014;73:337–348. doi: 10.1016/j.freeradbiomed.2014.05.017. [DOI] [PubMed] [Google Scholar]
- [265].Skjaerven KH, Penglase S, Olsvik PA, Hamre K. Redox regulation in Atlantic cod (Gadus morhua) embryos developing under normal and heat-stressed conditions. Free radical biology & medicine. 2013;57:29–38. doi: 10.1016/j.freeradbiomed.2012.11.022. [DOI] [PubMed] [Google Scholar]
- [266].Nakajima H, Nakajima-Takagi Y, Tsujita T, Akiyama S, Wakasa T, Mukaigasa K, Kaneko H, Tamaru Y, Yamamoto M, Kobayashi M. Tissue-restricted expression of Nrf2 and its target genes in zebrafish with gene-specific variations in the induction profiles. PLoS ONE. 2011;6:e26884. doi: 10.1371/journal.pone.0026884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [267].Li J, Spletter ML, Johnson JA. Dissecting tBHQ induced ARE-driven gene expression through long and short oligonucleotide arrays. Physiological genomics. 2005;21:43–58. doi: 10.1152/physiolgenomics.00214.2004. [DOI] [PubMed] [Google Scholar]
- [268].Kobayashi M, Li L, Iwamoto N, Nakajima-Takagi Y, Kaneko H, Nakayama Y, Eguchi M, Wada Y, Kumagai Y, Yamamoto M. The antioxidant defense system Keap1-Nrf2 comprises a multiple sensing mechanism for responding to a wide range of chemical compounds. Molecular and cellular biology. 2009;29:493–502. doi: 10.1128/MCB.01080-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [269].Ohtsuji M, Katsuoka F, Kobayashi A, Aburatani H, Hayes JD, Yamamoto M. Nrf1 and Nrf2 play distinct roles in activation of antioxidant response element-dependent genes. The Journal of biological chemistry. 2008;283:33554–33562. doi: 10.1074/jbc.M804597200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [270].Yang H, Magilnick N, Lee C, Kalmaz D, Ou X, Chan JY, Lu SC. Nrf1 and Nrf2 regulate rat glutamate-cysteine ligase catalytic subunit transcription indirectly via NF-kappaB and AP-1. Molecular and cellular biology. 2005;25:5933–5946. doi: 10.1128/MCB.25.14.5933-5946.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [271].Wang L, Gallagher EP. Role of Nrf2 antioxidant defense in mitigating cadmium-induced oxidative stress in the olfactory system of zebrafish. Toxicol Appl Pharmacol. 2013;266:177–186. doi: 10.1016/j.taap.2012.11.010. [DOI] [PubMed] [Google Scholar]
- [272].Shi X, Zhou B. The role of Nrf2 and MAPK pathways in PFOS-induced oxidative stress in zebrafish embryos. Toxicol Sci. 2010;115:391–400. doi: 10.1093/toxsci/kfq066. [DOI] [PubMed] [Google Scholar]
- [273].Cox AG, Saunders DC, Kelsey PB, Jr., Conway AA, Tesmenitsky Y, Marchini JF, Brown KK, Stamler JS, Colagiovanni DB, Rosenthal GJ, Croce KJ, North TE, Goessling W. S-nitrosothiol signaling regulates liver development and improves outcome following toxic liver injury. Cell reports. 2014;6:56–69. doi: 10.1016/j.celrep.2013.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [274].Wienholds E, van Eeden F, Kosters M, Mudde J, Plasterk RH, Cuppen E. Efficient target-selected mutagenesis in zebrafish. Genome research. 2003;13:2700–2707. doi: 10.1101/gr.1725103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [275].Moens CB, Donn TM, Wolf-Saxon ER, Ma TP. Reverse genetics in zebrafish by TILLING. Briefings in functional genomics & proteomics. 2008;7:454–459. doi: 10.1093/bfgp/eln046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [276].Gallagher EP, Wang L, Ramsden R, Mills M. Role of Nrf2 in regulating cellular antioxidant responses of fish. Toxicological Sciences (The Toxicologist Supplement) 2015:169. Abstract #794. [Google Scholar]
- [277].Rousseau ME, Sant KE, Borden LR, Franks DG, Hahn ME, Timme-Laragy AR. Crosstalk between Ahr and Nrf2 in PCB126 embryotoxicity in zebrafish (Danio rerio) Aquat Toxicol. 2015 doi: 10.1016/j.aquatox.2015.08.002. submitted. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [278].Hahn ME, Karchner SI, Franks DG, Timme-Laragy AR, McArthur AG. Chemical-Specific Oxidative Stress Response in Zebrafish Embryos. Toxicological Sciences (The Toxicologist Supplement) 2014:52. Abstract #209. [Google Scholar]
- [279].Tsujita T, Li L, Nakajima H, Iwamoto N, Nakajima-Takagi Y, Ohashi K, Kawakami K, Kumagai Y, Freeman BA, Yamamoto M, Kobayashi M. Nitro-fatty acids and cyclopentenone prostaglandins share strategies to activate the Keap1-Nrf2 system: a study using green fluorescent protein transgenic zebrafish. Genes Cells. 2011;16:46–57. doi: 10.1111/j.1365-2443.2010.01466.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [280].Takaya K, Suzuki T, Motohashi H, Onodera K, Satomi S, Kensler TW, Yamamoto M. Validation of the multiple sensor mechanism of the Keap1-Nrf2 system. Free radical biology & medicine. 2012;53:817–827. doi: 10.1016/j.freeradbiomed.2012.06.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [281].Brigelius-Flohe R, Flohe L. Basic principles and emerging concepts in the redox control of transcription factors. Antioxidants & redox signaling. 2011;15:2335–2381. doi: 10.1089/ars.2010.3534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [282].Lu SC. Glutathione synthesis. Biochim Biophys Acta. 2013;1830:3143–3153. doi: 10.1016/j.bbagen.2012.09.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [283].Hwang WY, Fu Y, Reyon D, Maeder ML, Kaini P, Sander JD, Joung JK, Peterson RT, Yeh JR. Heritable and precise zebrafish genome editing using a CRISPR-Cas system. PLoS ONE. 2013;8:e68708. doi: 10.1371/journal.pone.0068708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [284].Irion U, Krauss J, Nusslein-Volhard C. Precise and efficient genome editing in zebrafish using the CRISPR/Cas9 system. Development (Cambridge, England) 2014;141:4827–4830. doi: 10.1242/dev.115584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [285].Lee O, Green JM, Tyler CR. Transgenic fish systems and their application in ecotoxicology. Critical reviews in toxicology. 2015;45:124–141. doi: 10.3109/10408444.2014.965805. [DOI] [PubMed] [Google Scholar]
- [286].Kusik BW, Carvan MJ, 3rd, Udvadia AJ. Detection of mercury in aquatic environments using EPRE reporter zebrafish. Marine biotechnology (New York, N.Y. 2008;10:750–757. doi: 10.1007/s10126-008-9113-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [287].Priyadarshini M, Orosco LA, Panula PJ. Oxidative stress and regulation of Pink1 in zebrafish (Danio rerio) PLoS ONE. 2013;8:e81851. doi: 10.1371/journal.pone.0081851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [288].Ma Q, Kinneer K, Bi Y, Chan JY, Kan YW. Induction of murine NAD(P)H:quinone oxidoreductase by 2,3,7,8-tetrachlorodibenzo-p-dioxin requires the CNC (cap ‘n’ collar) basic leucine zipper transcription factor Nrf2 (nuclear factor erythroid 2-related factor 2): cross-interaction between AhR (aryl hydrocarbon receptor) and Nrf2 signal transduction. The Biochemical journal. 2004;377:205–213. doi: 10.1042/BJ20031123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [289].Miao W, Hu L, Scrivens PJ, Batist G. Transcriptional regulation of NF-E2 p45-related factor (NRF2) expression by the aryl hydrocarbon receptor-xenobiotic response element signaling pathway: direct cross-talk between phase I and II drug-metabolizing enzymes. The Journal of biological chemistry. 2005;280:20340–20348. doi: 10.1074/jbc.M412081200. [DOI] [PubMed] [Google Scholar]
- [290].Shin S, Wakabayashi N, Misra V, Biswal S, Lee GH, Agoston ES, Yamamoto M, Kensler TW. NRF2 modulates aryl hydrocarbon receptor signaling: influence on adipogenesis. Molecular and cellular biology. 2007;27:7188–7197. doi: 10.1128/MCB.00915-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [291].Yeager RL, Reisman SA, Aleksunes LM, Klaassen CD. Introducing the ‘TCDD Inducible AhR-Nrf2 Gene Battery’. Toxicol Sci. 2009;11:238–246. doi: 10.1093/toxsci/kfp115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [292].Rousseau ME, Hahn ME, Timme-Laragy AR. The Role of Nrf2a in the Transcriptional Response to PCB-126 in Zebrafish Embryos. Toxicological Sciences (The Toxicologist Supplement) 2015:376. Abstract #1757. [Google Scholar]
- [293].Chorley BN, Campbell MR, Wang X, Karaca M, Sambandan D, Bangura F, Xue P, Pi J, Kleeberger SR, Bell DA. Identification of novel NRF2-regulated genes by ChIP-Seq: influence on retinoid X receptor alpha. Nucleic acids research. 2012;40:7416–7429. doi: 10.1093/nar/gks409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [294].Wang H, Liu K, Geng M, Gao P, Wu X, Hai Y, Li Y, Li Y, Luo L, Hayes JD, Wang XJ, Tang X. RXRalpha inhibits the NRF2-ARE signaling pathway through a direct interaction with the Neh7 domain of NRF2. Cancer research. 2013;73:3097–3108. doi: 10.1158/0008-5472.CAN-12-3386. [DOI] [PubMed] [Google Scholar]
- [295].Wakabayashi N, Shin S, Slocum SL, Agoston ES, Wakabayashi J, Kwak MK, Misra V, Biswal S, Yamamoto M, Kensler TW. Regulation of notch1 signaling by nrf2: implications for tissue regeneration. Sci Signal. 2010;3:ra52. doi: 10.1126/scisignal.2000762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [296].Wakabayashi N, Chartoumpekis DV, Kensler TW. Crosstalk between Nrf2 and notch signaling. Crosstalk between Nrf2 and notch signaling. 2015 doi: 10.1016/j.freeradbiomed.2015.05.017. pii: S0891-5849(0815)00228-00222. doi: 00210.01016/j.freeradbiomed.02015.00205.00017. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- [297].Dayalan Naidu S, Kostov RV, Dinkova-Kostova AT. Transcription factors Hsf1 and Nrf2 engage in crosstalk for cytoprotection. Trends in pharmacological sciences. 2015;36:6–14. doi: 10.1016/j.tips.2014.10.011. [DOI] [PubMed] [Google Scholar]
- [298].Nakajima S, Kitamura M. Bidirectional regulation of NF-kappaB by reactive oxygen species: a role of unfolded protein response. Free radical biology & medicine. 2013;65:162–174. doi: 10.1016/j.freeradbiomed.2013.06.020. [DOI] [PubMed] [Google Scholar]