Abstract
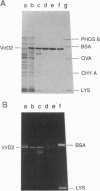
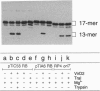
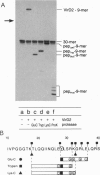
As an early stage of plant transformation by Agrobacterium tumefaciens, the Ti plasmid is nicked at the border sequences that delimit the T-DNA. Cleavage results in covalent attachment of VirD2 to the 5' terminal of the nicked strand by a process resembling initiation of DNA transfer that occurs in the donor cell during bacterial conjugation. We demonstrate that this cleavage can be reproduced in vitro: VirD2 protein, the border-cleaving enzyme, was overproduced and purified. Cleavage assays were performed with single-stranded oligodeoxyribonucleotides encompassing the Ti plasmid border region or the transfer origin's nick region of the conjugative plasmid RP4. VirD2 of pTiC58 cleaves both border- and nick region-containing oligonucleotides. However, the relaxase TraI of RP4 can cut only the cognate nick regions. The respective proteins remain covalently bound to the 5' end of the cleavage sites, leaving the 3' termini unmodified. VirD2-mediated oligonucleotide cleavage was demonstrated to be an equilibrium reaction that allows specific joining of cleavage products restoring border and nick regions, respectively. The possible role of VirD2 in T-DNA integration into the plant cell's genome is discussed in terms of less stringent target-sequence requirements.
Full text
PDF

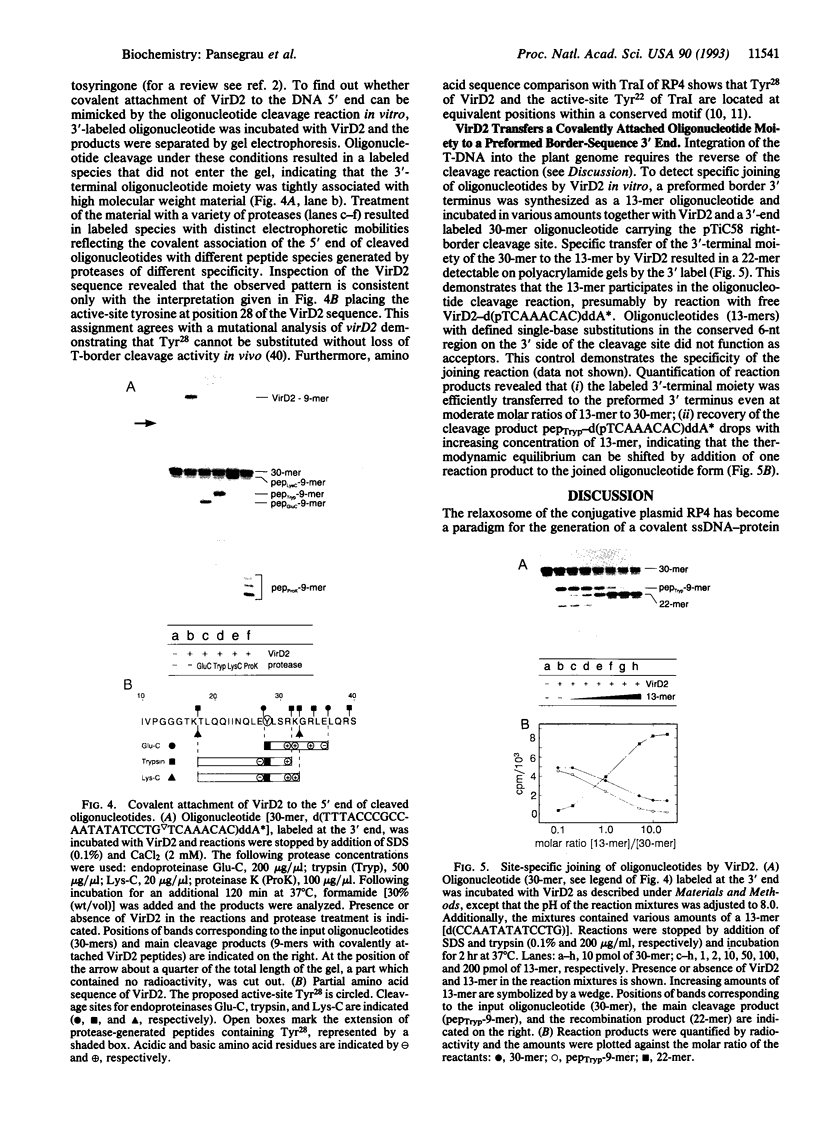

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Albright L. M., Yanofsky M. F., Leroux B., Ma D. Q., Nester E. W. Processing of the T-DNA of Agrobacterium tumefaciens generates border nicks and linear, single-stranded T-DNA. J Bacteriol. 1987 Mar;169(3):1046–1055. doi: 10.1128/jb.169.3.1046-1055.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alt-Moerbe J., Rak B., Schröder J. A 3.6-kbp segment from the vir region of Ti plasmids contains genes responsible for border-sequence-directed production of T region circles in E. coli. EMBO J. 1986 Jun;5(6):1129–1135. doi: 10.1002/j.1460-2075.1986.tb04337.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balzer D., Ziegelin G., Pansegrau W., Kruft V., Lanka E. KorB protein of promiscuous plasmid RP4 recognizes inverted sequence repetitions in regions essential for conjugative plasmid transfer. Nucleic Acids Res. 1992 Apr 25;20(8):1851–1858. doi: 10.1093/nar/20.8.1851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dürrenberger F., Crameri A., Hohn B., Koukolíková-Nicola Z. Covalently bound VirD2 protein of Agrobacterium tumefaciens protects the T-DNA from exonucleolytic degradation. Proc Natl Acad Sci U S A. 1989 Dec;86(23):9154–9158. doi: 10.1073/pnas.86.23.9154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Filichkin S. A., Gelvin S. B. Formation of a putative relaxation intermediate during T-DNA processing directed by the Agrobacterium tumefaciens VirD1,D2 endonuclease. Mol Microbiol. 1993 May;8(5):915–926. doi: 10.1111/j.1365-2958.1993.tb01637.x. [DOI] [PubMed] [Google Scholar]
- Furuya N., Komano T. Determination of the nick site at oriT of IncI1 plasmid R64: global similarity of oriT structures of IncI1 and IncP plasmids. J Bacteriol. 1991 Oct;173(20):6612–6617. doi: 10.1128/jb.173.20.6612-6617.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furuya N., Nisioka T., Komano T. Nucleotide sequence and functions of the oriT operon in IncI1 plasmid R64. J Bacteriol. 1991 Apr;173(7):2231–2237. doi: 10.1128/jb.173.7.2231-2237.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghai J., Das A. The virD operon of Agrobacterium tumefaciens Ti plasmid encodes a DNA-relaxing enzyme. Proc Natl Acad Sci U S A. 1989 May;86(9):3109–3113. doi: 10.1073/pnas.86.9.3109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gheysen G., Villarroel R., Van Montagu M. Illegitimate recombination in plants: a model for T-DNA integration. Genes Dev. 1991 Feb;5(2):287–297. doi: 10.1101/gad.5.2.287. [DOI] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Herrera-Estrella A., Chen Z. M., Van Montagu M., Wang K. VirD proteins of Agrobacterium tumefaciens are required for the formation of a covalent DNA--protein complex at the 5' terminus of T-strand molecules. EMBO J. 1988 Dec 20;7(13):4055–4062. doi: 10.1002/j.1460-2075.1988.tb03299.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hewick R. M., Hunkapiller M. W., Hood L. E., Dreyer W. J. A gas-liquid solid phase peptide and protein sequenator. J Biol Chem. 1981 Aug 10;256(15):7990–7997. [PubMed] [Google Scholar]
- Howard E. A., Winsor B. A., De Vos G., Zambryski P. Activation of the T-DNA transfer process in Agrobacterium results in the generation of a T-strand-protein complex: Tight association of VirD2 with the 5' ends of T-strands. Proc Natl Acad Sci U S A. 1989 Jun;86(11):4017–4021. doi: 10.1073/pnas.86.11.4017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howard E. A., Zupan J. R., Citovsky V., Zambryski P. C. The VirD2 protein of A. tumefaciens contains a C-terminal bipartite nuclear localization signal: implications for nuclear uptake of DNA in plant cells. Cell. 1992 Jan 10;68(1):109–118. doi: 10.1016/0092-8674(92)90210-4. [DOI] [PubMed] [Google Scholar]
- Ilyina T. V., Koonin E. V. Conserved sequence motifs in the initiator proteins for rolling circle DNA replication encoded by diverse replicons from eubacteria, eucaryotes and archaebacteria. Nucleic Acids Res. 1992 Jul 11;20(13):3279–3285. doi: 10.1093/nar/20.13.3279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston R. F., Pickett S. C., Barker D. L. Autoradiography using storage phosphor technology. Electrophoresis. 1990 May;11(5):355–360. doi: 10.1002/elps.1150110503. [DOI] [PubMed] [Google Scholar]
- Lessl M., Balzer D., Pansegrau W., Lanka E. Sequence similarities between the RP4 Tra2 and the Ti VirB region strongly support the conjugation model for T-DNA transfer. J Biol Chem. 1992 Oct 5;267(28):20471–20480. [PubMed] [Google Scholar]
- Lessl M., Balzer D., Weyrauch K., Lanka E. The mating pair formation system of plasmid RP4 defined by RSF1010 mobilization and donor-specific phage propagation. J Bacteriol. 1993 Oct;175(20):6415–6425. doi: 10.1128/jb.175.20.6415-6425.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lessl M., Pansegrau W., Lanka E. Relationship of DNA-transfer-systems: essential transfer factors of plasmids RP4, Ti and F share common sequences. Nucleic Acids Res. 1992 Nov 25;20(22):6099–6100. doi: 10.1093/nar/20.22.6099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayerhofer R., Koncz-Kalman Z., Nawrath C., Bakkeren G., Crameri A., Angelis K., Redei G. P., Schell J., Hohn B., Koncz C. T-DNA integration: a mode of illegitimate recombination in plants. EMBO J. 1991 Mar;10(3):697–704. doi: 10.1002/j.1460-2075.1991.tb07999.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morton T. M., Eaton D. M., Johnston J. L., Archer G. L. DNA sequence and units of transcription of the conjugative transfer gene complex (trs) of Staphylococcus aureus plasmid pGO1. J Bacteriol. 1993 Jul;175(14):4436–4447. doi: 10.1128/jb.175.14.4436-4447.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moshiri F., Chawla A., Maier R. J. Cloning, characterization, and expression in Escherichia coli of the genes encoding the cytochrome d oxidase complex from Azotobacter vinelandii. J Bacteriol. 1991 Oct;173(19):6230–6241. doi: 10.1128/jb.173.19.6230-6241.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motallebi-Veshareh M., Balzer D., Lanka E., Jagura-Burdzy G., Thomas C. M. Conjugative transfer functions of broad-host-range plasmid RK2 are coregulated with vegetative replication. Mol Microbiol. 1992 Apr;6(7):907–920. doi: 10.1111/j.1365-2958.1992.tb01541.x. [DOI] [PubMed] [Google Scholar]
- Pansegrau W., Balzer D., Kruft V., Lurz R., Lanka E. In vitro assembly of relaxosomes at the transfer origin of plasmid RP4. Proc Natl Acad Sci U S A. 1990 Sep;87(17):6555–6559. doi: 10.1073/pnas.87.17.6555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pansegrau W., Lanka E. Common sequence motifs in DNA relaxases and nick regions from a variety of DNA transfer systems. Nucleic Acids Res. 1991 Jun 25;19(12):3455–3455. doi: 10.1093/nar/19.12.3455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pansegrau W., Schröder W., Lanka E. Relaxase (TraI) of IncP alpha plasmid RP4 catalyzes a site-specific cleaving-joining reaction of single-stranded DNA. Proc Natl Acad Sci U S A. 1993 Apr 1;90(7):2925–2929. doi: 10.1073/pnas.90.7.2925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pansegrau W., Ziegelin G., Lanka E. Covalent association of the traI gene product of plasmid RP4 with the 5'-terminal nucleotide at the relaxation nick site. J Biol Chem. 1990 Jun 25;265(18):10637–10644. [PubMed] [Google Scholar]
- Sano T., Cantor C. R. Expression of a cloned streptavidin gene in Escherichia coli. Proc Natl Acad Sci U S A. 1990 Jan;87(1):142–146. doi: 10.1073/pnas.87.1.142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinland B., Koukolíková-Nicola Z., Hall M. N., Hohn B. The T-DNA-linked VirD2 protein contains two distinct functional nuclear localization signals. Proc Natl Acad Sci U S A. 1992 Aug 15;89(16):7442–7446. doi: 10.1073/pnas.89.16.7442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogel A. M., Das A. Mutational analysis of Agrobacterium tumefaciens virD2: tyrosine 29 is essential for endonuclease activity. J Bacteriol. 1992 Jan;174(1):303–308. doi: 10.1128/jb.174.1.303-308.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang K., Herrera-Estrella A., Van Montagu M. Overexpression of virD1 and virD2 genes in Agrobacterium tumefaciens enhances T-complex formation and plant transformation. J Bacteriol. 1990 Aug;172(8):4432–4440. doi: 10.1128/jb.172.8.4432-4440.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waters V. L., Hirata K. H., Pansegrau W., Lanka E., Guiney D. G. Sequence identity in the nick regions of IncP plasmid transfer origins and T-DNA borders of Agrobacterium Ti plasmids. Proc Natl Acad Sci U S A. 1991 Feb 15;88(4):1456–1460. doi: 10.1073/pnas.88.4.1456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss A. A., Johnson F. D., Burns D. L. Molecular characterization of an operon required for pertussis toxin secretion. Proc Natl Acad Sci U S A. 1993 Apr 1;90(7):2970–2974. doi: 10.1073/pnas.90.7.2970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winans S. C. Two-way chemical signaling in Agrobacterium-plant interactions. Microbiol Rev. 1992 Mar;56(1):12–31. doi: 10.1128/mr.56.1.12-31.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young C., Nester E. W. Association of the virD2 protein with the 5' end of T strands in Agrobacterium tumefaciens. J Bacteriol. 1988 Aug;170(8):3367–3374. doi: 10.1128/jb.170.8.3367-3374.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zambryski P. Basic processes underlying Agrobacterium-mediated DNA transfer to plant cells. Annu Rev Genet. 1988;22:1–30. doi: 10.1146/annurev.ge.22.120188.000245. [DOI] [PubMed] [Google Scholar]
- Ziegelin G., Fürste J. P., Lanka E. TraJ protein of plasmid RP4 binds to a 19-base pair invert sequence repetition within the transfer origin. J Biol Chem. 1989 Jul 15;264(20):11989–11994. [PubMed] [Google Scholar]
- Ziegelin G., Pansegrau W., Lurz R., Lanka E. TraK protein of conjugative plasmid RP4 forms a specialized nucleoprotein complex with the transfer origin. J Biol Chem. 1992 Aug 25;267(24):17279–17286. [PubMed] [Google Scholar]
- Ziegelin G., Pansegrau W., Strack B., Balzer D., Kröger M., Kruft V., Lanka E. Nucleotide sequence and organization of genes flanking the transfer origin of promiscuous plasmid RP4. DNA Seq. 1991;1(5):303–327. doi: 10.3109/10425179109020786. [DOI] [PubMed] [Google Scholar]
- van Haaren M. J., Sedee N. J., de Boer H. A., Schilperoort R. A., Hooykaas P. J. Mutational analysis of the conserved domains of a T-region border repeat of Agrobacterium tumefaciens. Plant Mol Biol. 1989 Nov;13(5):523–531. doi: 10.1007/BF00027312. [DOI] [PubMed] [Google Scholar]