Abstract
Goats and sheep are versatile domesticates that have been integrated into diverse environments and production systems. Natural and artificial selection have shaped the variation in the two species, but natural selection has played the major role among indigenous flocks. To investigate signals of natural selection, we analyzed genotype data generated using the caprine and ovine 50K SNP BeadChips from Barki goats and sheep that are indigenous to a hot arid environment in Egypt's Coastal Zone of the Western Desert. We identify several candidate regions under selection that spanned 119 genes. A majority of the genes were involved in multiple signaling and signal transduction pathways in a wide variety of cellular and biochemical processes. In particular, selection signatures spanning several genes that directly or indirectly influenced traits for adaptation to hot arid environments, such as thermo-tolerance (melanogenesis) (FGF2, GNAI3, PLCB1), body size and development (BMP2, BMP4, GJA3, GJB2), energy and digestive metabolism (MYH, TRHDE, ALDH1A3), and nervous and autoimmune response (GRIA1, IL2, IL7, IL21, IL1R1) were identified. We also identified eight common candidate genes under selection in the two species and a shared selection signature that spanned a conserved syntenic segment to bovine chromosome 12 on caprine and ovine chromosomes 12 and 10, respectively, providing, most likely, the evidence for selection in a common environment in two different but closely related species. Our study highlights the importance of indigenous livestock as model organisms for investigating selection sweeps and genome-wide association mapping.
Introduction
In terms of numbers, geographic distribution and socioeconomic importance, goats and sheep are two of the most important global livestock species. They were the first ruminants to be domesticated around 11 000 years ago in the Fertile Crescent (Zeder, 2008). They were subsequently dispersed across the globe, adapting themselves to diverse biophysical and production environments. Since the early eighteenth century, continuous anthropogenic selection for production, reproduction and physical traits has been practiced in the quest to standardize most breeds of livestock in the temperate regions. However, strong artificial selection for such traits has not occurred, in general, for their indigenous (local) counterparts in the tropics and sub-tropics. The latter are most often exposed to natural selection for adaptation to diverse environments. Differences in the intensity and objectives of natural and artificial selection must have resulted in detectable differences within their genomes. This is evidenced in the phenotypic uniformity of temperate breeds that contrasts with the wide diversity that is observed among indigenous tropical livestock (Hanotte et al., 2010).
The detection of genomic differences can shed light on the genetic basis of adaptation to diverse environments and provide insights into functionally important genetic variants (Andersson and Georges, 2004). The availability of genomic data that has been generated using genome-wide single nucleotide polymorphism (SNP) analyses and the comparison of genomic patterns of SNP variation between divergent breeds has successfully identified putative genomic regions and genes associated with differences in selection pressure in various livestock species including sheep (Kijas et al., 2012; Lv et al., 2014), Yak (Qiu et al., 2012), cattle (Xu et al., 2015) and chicken (Elferink et al., 2012).
The indigenous Barki goats and sheep are known to be well adapted to Egypt's Coastal Zone of the Western Desert (Haider, 1982) and represent the primary livestock species and breed raised in the area. The defining feature of the Coastal Zone of the Western Desert, including most of the arid and semiarid regions of the world, is intense solar radiation, high diurnal temperatures, low precipitation, and water and feed scarcity almost throughout the year. It therefore can be hypothesized that the Barki goats and sheep have evolved to survive and reproduce under such conditions and their genomes should harbor footprints for adaptation to such an environment. If this is the case, they represent a unique genetic resource with the potential to identify genetic markers, molecular pathways and selection sweep regions that can provide insights into the genome biology and the mechanisms of adaptation to extreme environments.
Here, we investigated possible genomic signatures of natural selection for adaptation to a hot arid environment using SNP data of Barki goats and sheep and five exotic breeds of goats and three of sheep that were generated with Illumina's caprine and ovine 50K SNP BeadChips. We applied two selection sweep approaches (FST and integrated Haplotype Score (iHS)) and identified for the first time a set of candidate genes and genomic regions under direct or indirect selection in a hot arid environment including a common candidate region under selection in the two species. Our results are of importance for their potential application in functional genomics and in the design of breeding programs in view of the current and future predicted global effects of climate change.
Materials and methods
Sample preparation, genotyping and quality control
Blood samples were collected from 68 Barki goats and 59 Barki sheep from Borg Arab Research Farm and from farmers flocks in Egypt's Coastal Zone of the Western Desert (Latitude: 31° 31' 12" N; Longitude: 30° 10' 12" E; Altitude: 54 meters above sea level; Mean annual rainfall: <150 mm (range=0–199 mm); Mean annual temperature: 28 °C (range=9–35 °C)). The animals were selected at random as representatives of farmers' flocks that have grazed on poor quality pastures under direct solar radiation for several generations. Whole blood was collected via jugular venipuncture into EDTA-coated vacutainer tubes, and DNA was extracted with the GenElute Blood Genomic DNA extraction kit (Sigma, St Louis, MO, USA).
Genotyping was performed using Illumina's caprine and ovine 50K SNP BeadChips (Illumina Inc., San Diego, CA, USA) with standard procedures at GeneSeek Inc. Lincoln, NE, USA. Basic information on the 53 347 and 54 241 SNPs available on the two chips, including SNP name, chromosome and map position were based on the Caprine v2.0 and Ovine v3.1 genome assemblies. We included in our analysis SNP genotypes from 194 individuals from five exotic breeds of goats (Boer (n=34), Myotonic (n=15), Spanish (n=20), Kiko (n=11), LaMancha (n=13)), and three exotic breeds of sheep (Romney/Romney-Finn (n=50), Texel (n=26) and Corriedale crosses (n=25)). The data for goats came from individuals that were sampled from various private farms and ranches in the USA, while the sheep data were from experimental populations that had been genotyped and described in a previous study (Zhao et al., 2012). Compared with Barki goats and sheep, the exotic breeds (see http://www.ansi.okstate.edu/breeds) have been artificially selected for different production and phenotypic traits under temperate conditions. These two groups of animals, therefore provide appropriate genetic material to analyze genomic differences arising from different evolutionary and breeding histories.
We assessed SNP genotype data quality with PLINK v1.07 (Purcell et al., 2007). We pruned out all individuals that failed the inclusion criterion of successful genotypes of ⩾90%. Only mapped autosomal SNPs were screened and those with call rates of <80% and minor allele frequency (MAF) of <0.03 were discarded. In total, 2621 and 5893 SNPs from goats and sheep, respectively, failed the inclusion criterion, leaving 50 726 and 48 348 SNPs for downstream analysis.
Assessing genetic diversity and structure
Two statistics, observed (Ho) and expected (He) heterozygosity calculated with PLINK v1.07 (Purcell et al., 2007), were used to provide estimates of genetic diversity. Private allele richness calculated with ADZE (Szpiech et al., 2008) was used as a measure of genetic distinctiveness. We also estimated Froh (the individual autozygosity) from runs of homozygosity (ROH) using default values and Fgeno from values of observed and expected homozygosity of each SNP, as measures of inbreeding using PLINK v1.07. We also estimated for each breed the effective population sizes (Ne) at 5 and 70 generations ago using the SNeP package (Barbato et al., 2015) to summarize the extent of linkage disequilibrium.
The SNP genotypes that passed quality control were used to investigate genetic relationships between individuals. We performed the principal component analysis (PCA) using ADEGENET v1.4-2 (Jombart and Ahmed, 2011) executed in R (R Development Core Team, 2013). Model-based clustering, with no prior knowledge on breed origins, was also performed using ADMIXTURE 1.2.2 (Alexander et al., 2009) to investigate individual admixture proportions. For these analyses, 1 in every 10 SNPs was sampled to minimize possible confounding effects of linkage disequilibrium on the underlying patterns of genetic structure. We performed 10 runs with different random seed numbers for each value of 1⩽K⩽12 and selected the optimum number of ancestral populations (K) on the basis of the lowest value of the cross-validation error.
Selection sweep, gene annotation and functional analysis
We calculated locus-specific divergence in allele frequencies between Barki goats and sheep with their exotic counterparts, based on the unbiased estimates of pairwise FST, following Akey et al. (2010) to identify, most likely, old and fixed signatures of selection. For each SNP in a pairwise comparison, we calculated the expected value of FST with the ADEGENET v1.4-2 in R. Individual FST values derived from pairwise comparisons between breeds were then combined and averaged across all SNPs using a custom R script to obtain the standardized summary ‘di' statistic (Akey et al., 2010). Candidate genomic regions under selection were defined by SNP outliers falling in the 99th percentile of the empirical distribution of ‘di' (Akey et al., 2010), which is equivalent to a cutoff FST value of greater than 0.3.
To infer recent and generally segregating selection sweeps, we examined the decay of haplotypes using the iHS (Voight et al., 2006) approach. For this analysis, the ancestral and derived alleles are required; these, however, remain unknown in sheep and goats. In this case, prior to the analysis, we used two approaches to define the ancestral/derived state of the alleles. First, we assigned these at random, and second, we permutated (100 times) the ancestral/descendant allelic states. We phased the haplotypes using Beagle (Browning and Browning, 2007), and then calculated iHS scores for each SNP/haplotype within a breed/population. Haplotype frequencies were calculated using sliding windows of 10 SNPs that overlapped by 5 SNPs. For each locus, the |iHS| score was computed using the REHH package (Gautier and Vitalis, 2012) in R and an |iHS| score >3.0 was used to infer the candidate genomic regions under selection.
Genes found within the intervals spanning the candidate genome regions were retrieved from the Ensembl genome browser (http://www.ensembl.org/) using the bovine UMD3.1 reference genome assembly and these were considered as candidate genes. Enrichment and functional annotation of the candidate genes were defined using the Enrichr program (Chen et al., 2013). Phenotypes that are known to be affected by the identified candidate genes were inferred from the literature.
Results
Genetic diversity and structure
We investigated genetic diversity using filtered SNPs within each breed (Table 1). The results indicated differences in genetic diversity between the indigenous and exotic breeds. For instance, the average Ho in Barki goats was lower than that of the four other exotic breeds combined but higher than that of the Boer goat. However, when Ho was calculated for each exotic goat breed, the value for Barki was similar to that of LaMancha but higher than that of Myotonic and lower than that of Kiko and Spanish (Supplementary Table S1). The average He was lower in Barki goats than in the other exotic breeds and the Boer goat, whereas the private allele richness was higher in Barki goats than in the other exotic breeds and Boer goat. However, the He value of Barki goats was similar to that of Kiko but slightly lower than that of Spanish and higher than that of LaMancha and Myotonic (Supplementary Table S1). The average values of Ho, He and private allele richness were higher in Barki sheep compared with the three exotic breeds. The inbreeding coefficients (Froh, Fgeno) and the average lengths of ROH were higher in all the exotic breeds of goats and sheep with the exception of Fgeno in the Corriedale crosses. The Barki goats and sheep generally had a higher proportion of individuals lacking long stretches of ROH and their estimated Ne at 5 and 70 generations ago were also higher than those of the exotic breeds.
Table 1. Average coefficients of inbreeding, runs of homozygosity and genetic diversity in sheep and goats.
|
Goats |
Sheep |
||||||
|---|---|---|---|---|---|---|---|
| Barki | Other exoticsa | Boer | Barki | Romney | Texel | Corriedale | |
| Sample sizes | 68 | 59 | 34 | 59 | 50 | 26 | 25 |
| Froh | 0.02 (0.07)b | 0.03 (0.06) | 0.09 (0.03) | 0.02 (0.03) | 0.05 (0.01) | 0.10 (0.02) | 0.03 (0.03) |
| Fgeno | 0.01 (0.05) | 0.09 (0.07) | 0.19 (0.05) | −0.00 (0.04) | 0.08 (0.01) | 0.25 (0.02) | −0.05 (0.03) |
| r c | 0.80 | 0.85 | 0.84 | 0.76 | 0.77 | 0.65 | 0.83 |
| Mean length of ROH (Mb) | 1.48 (2.8) | 4.19 (4.5) | 10.61 (8.4) | 4.40 (7.6) | 12.49 (8.3) | 16.48 (9.9) | 14.25 (4.0) |
| Individuals lacking ROH | 72% | 6% | 0% | 58% | 2% | 0% | 17% |
| Ho | 0.40 (0.12) | 0.42 (0.08) | 0.35 (0.02) | 0.37 (0.14) | 0.33 (0.03) | 0.22 (0.03) | 0.32 (0.03) |
| He | 0.40 (0.11) | 0.44 (0.07) | 0.43 (0.11) | 0.37 (0.13) | 0.33 (0.03) | 0.31 (0.03) | 0.30 (0.03) |
| Ne (5)d | 127 | 87 | 66 | 130 | 45 | 21 | 30 |
| Ne (70)d | 1531 | 729 | 230 | 1331 | 374 | 161 | 244 |
| Private allelic richness (pAr) | 0.016 | 0.007 | 0.003 | 0.017 | 0.008 | 0.002 | 0.005 |
Abbreviations: Ho, Observed homozygosity; He, Expected homozygosity; ROH, runs of homozygosity.
Other exotics=Myotonic, Spanish, Kiko, LaMancha.
Numbers in parenthesis represent standard deviations.
r=Correlation coefficient between ROH-based inbreeding coefficient (Froh) and genotype-based inbreeding coefficient (Fgeno).
Ne=Effective population size (5 or 70 generations ago).
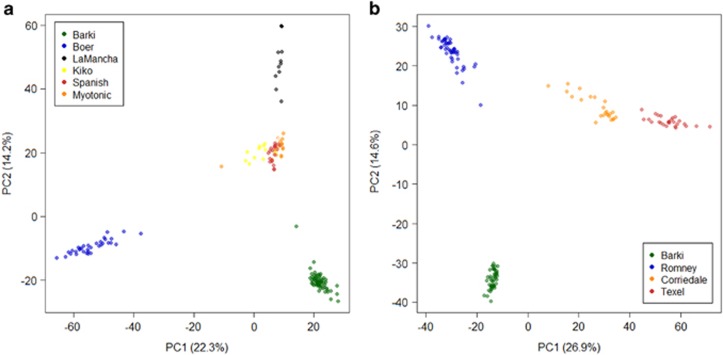
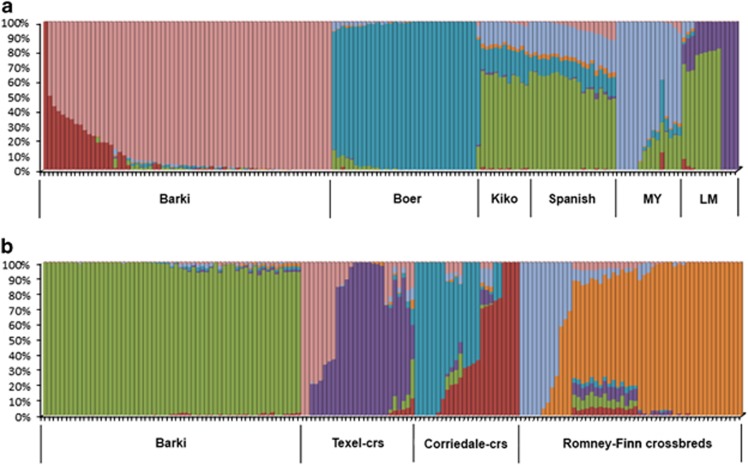
Population substructure was investigated using PCA and Bayesian model-based clustering. The first and second principal components (PC1 and PC2) of the PCA differentiated Barki goats and sheep from the exotic breeds (Figure 1), as well as the Boer goat from the other four exotic breeds of goats (Figure 1a). The three breeds of exotic sheep, Romney/Romney-Finn crosses, Texel crosses and Corriedale crosses, were separated from each other by PC1 (Figure 1b). In general, the first two PCs explained 36.5% (PC1=22.3%, PC2=14.2%) and 41.5% (PC1=26.9%, PC2=14.6%) of the total genetic variation in goats and sheep, respectively. Higher PCs (>2) did not result in any observable genetic clusters. Model-based clustering implemented in Admixture 1.2.2 was used to determine the number of genetic backgrounds (K) that explain the observed total sum of within-population genetic variation. The lowest cross-validation error was obtained at K=6 and 7, respectively, in goats and sheep (Supplementary Figure S1) indicating the presence of at least six and seven genetic backgrounds. Similar to the PCA, the individual assignment probabilities revealed that the genetic background of Barki goats and sheep differed from that of the exotic breeds (Figure 2) suggesting clear genetic divergence, and this was evident across all the values of K tested (2⩽K⩽9). To investigate whether there is any genetic structure within the Barki goats and sheep, we ran Admixture 1.2.2 separately for these populations and as expected, the lowest cross-validation error was at K=1 (Supplementary Figure S2), further confirming the genetic uniformity of Barki goats and sheep. The clear genetic divergence between the indigenous Barki and the exotic breeds indicates that the chosen individuals were adequate to explore their genomic characteristics.
Figure 1.
PCA plots showing the relationship between the breeds analyzed for each species. (a) Barki goats and exotic breeds. (b) Barki sheep and Exotic breeds.
Figure 2.
Individual assignment probabilities generated with ADMIXTURE (1⩽K⩽12) in goats (a) and sheep (b). Each color represents a cluster, and the ratio of colored bars is proportional to the assignment probability of an individual to each cluster. MY and LM represent Myotonic and LaMancha goats, respectively.
Signatures of selection—FST approach
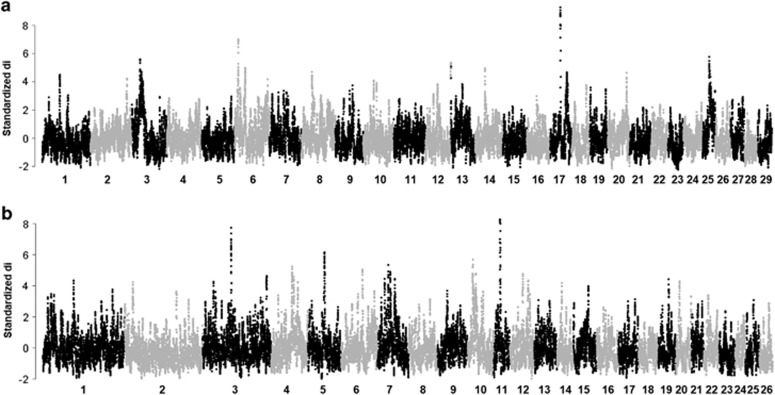
On the basis of the results of PCA and admixture, we used selection mapping to explore differences among groups of breeds/populations that are exposed to different selection pressures and from different geographic zones (cold humid temperate versus tropical dry arid regions) by calculating the average ‘di' values between the following breed pairs: Barki goats versus the Boer, Barki goats versus the other four exotic breeds (Myotonic, Spanish, Kiko, LaMancha) grouped together and Barki sheep versus Romney/Romney-Finn crosses, Texel crosses and Corriedale crosses. For this analysis, the Myotonic, Spanish, Kiko and LaMancha breeds were grouped together on the basis of their clustering pattern on the PCA and their small sample sizes. The genome-wide distribution of the standardized ‘di' values is shown in Figure 3a (goats) and Figure 3b (sheep). For each pairwise comparison, candidate regions under selection were defined as outliers falling within the 99th percentile of the empirical distribution of the ‘di' values (equivalent to having an FST value of greater than 0.3). Using this strategy, seven candidate selection sweep regions were identified in the goat comparisons, in particular the one involving Barki and Boer goats. The regions were observed on chromosomes CHI3, 6, 13, 14, 17 and 25 (Figure 3a) with 33 genes distributed across them (Table 2; Supplementary Table S2). We further identified two regions at the highest values of FST on CHI6 (FST=0.59; 11.4–12.7 Mb) and CHI17 (FST=0.70; 34.0–35.0 Mb), respectively. The region on CHI6 spanned one gene, UGT8, while the one on CHI17 spanned four genes, FGF2, IL2, IL7 and IL21. Similarly, nine candidate selection sweep regions found across four chromosomes OAR3, 5, 10 and 11 (Figure 3b), and which spanned 54 genes (Table 3; Supplementary Table S3) were identified in the sheep comparisons. Among these, we identified three regions at the highest selection signals (Figure 3b; Table 3) on OAR3 (98.1–99.6 Mb), OAR5 (62.1–64.1 Mb) and OAR11 (26.8–29.9 Mb). The region on OAR3 spanned one gene (IL1R1), the one on OAR5, two genes (GRIA1 and CNOT8) and the one on OAR11, seven genes (FGF11, PIK3R5, MYH1, MYH3, MYH8, MYH10 and MYH13), respectively.
Figure 3.
Genome-wide differential selection (di) in goats (a) and sheep (b). The standardized FST value (di) calculated from pairwise breed comparisons are plotted for each SNP across the genome on the basis of an overlapping (5 SNP) window size of 20 SNPs for goats (a) and sheep (b).
Table 2. Pairwise F ST and iHS scores in goats.
| Chromosome | Regiona | Barki-Boerb | Barki-multibreedsb,c | iHSb (Boer) | Synteny to cattle | Gened |
|---|---|---|---|---|---|---|
| 1 | 55.9–56.3 | 0.44 | 0.19 | 1.86 | BTA 1: 57.4–58.1 | ATP6V1H, ATG3 |
| 3 | 30.6–31.4 | 0.43 | 0.14 | 2.66 | BTA 3: 32.8–33.7 | GNAI3, GTM1, |
| 6 | 11.4–12.7 | 0.59 | 0.25 | 1.82 | BTA 6: 14.0–15.4 | UGT8 |
| 6 | 35.1–36.1 | 0.31 | 0.52 | 1.54 | BTA 6: 38.3–39.4 | IBSP, SPP1 |
| 13 | 0.0–0.5 | 0.64 | 0.08 | 0.52 | BTA 13: 0.0–1.1 | PLCB1 |
| 14 | 32.2–32.4 | 0.56 | 0.06 | 1.03 | BTA 14: 23.4–23.6 | POLR2K |
| 17 | 34.0–35.0 | 0.70 | 0.46 | 1.88 | BTA 17: 34.7–36.1 | FGF2, IL2, IL21 |
| 17 | 55.2–56.1 | 0.47 | 0.08 | 1.17 | BTA 17: 57.9–58.8 | PRKAB1 |
| 25 | 22.1–23.4 | 0.58 | 0.27 | 2.62 | BTA 25: 22.3–23.6 | LCMT1 |
Abbreviation: iHS, integrated Haplotype Score.
Region is defined by FST >0.3.
Highest FST and iHS value in the candidate selection region is shown.
Multibreeds=Myotonic, Spanish, Kiko, LaMancha as one group.
Gene(s) located nearest the highest FST value is listed.
Table 3. Pairwise F ST and iHS scores in sheep.
| Chromosome | Regionb | FSTa |
iHSa |
|||
|---|---|---|---|---|---|---|
| Barki-Romney/Romney-Finn crosses | Barki-Texel crosses | Barki-Corridale crosses | Romney/Romney-Finn crosses | Genec | ||
| 3 | 98.1–99.6 | 0.53 | 0.42 | 0.61 | 2.10 | IL1R1 |
| 4 | 72.4–73.3 | 0.47 | 0.54 | 0.66 | 1.84 | ZNF804B |
| 5 | 62.1–64.1 | 0.63 | 0.71 | 0.55 | 2.25 | GRIA1, CNOT8 |
| 6 | 75.9–76.3 | 0.48 | 0.41 | 0.56 | 1.21 | LPHN3 |
| 7 | 44.5–44.8 | 0.54 | 0.51 | 0.60 | 3.30 | MAP2K4, TLN2 |
| 10 | 18.3–19.3 | 0.34 | 0.33 | 0.49 | 2.21 | Rb1 |
| 11 | 26.8–29.9 | 0.52 | 0.73 | 0.53 | 2.09 | FGF11, PIK3R5, MYH1, MYH10 |
Abbreviation: iHS, integrated Haplotype Score.
Highest value of FST and iHS in the interval is shown.
Region is defined by FST>0.3.
Gene(s) located nearest the highest FST value is listed.
Signatures of selection—iHS approach
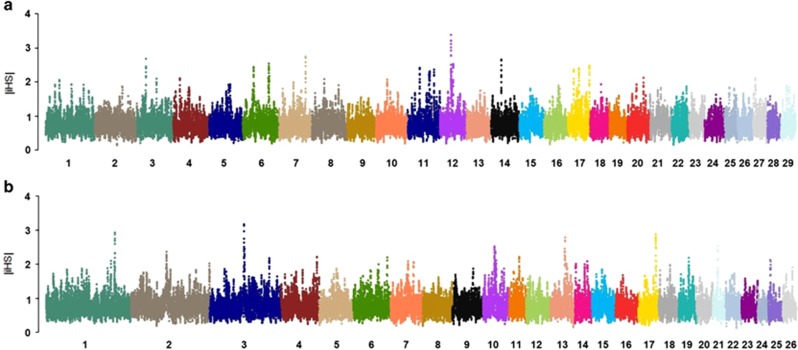
The two approaches we used to assign the ancestral/derived state of the alleles returned similar results. Eight candidate sweep regions (|iHS| >3.0) distributed across seven chromosomes (CHI3, 6, 7, 11, 12, 14 and 17) were identified in Barki goats (Figure 4a, Supplementary Figure S3). Eighteen genes were located in these eight regions (Table 4; Supplementary Figure S4). We also identified two regions at the highest |iHS| on CHI6 (|iHS|=5.10; 82.6–83.2 Mb) and CHI12 (|iHS|=5.17; 39.8–42.6 Mb), each spanning one gene, CSN3 and PCDH9, respectively (Table 4; Supplementary Figure S5). We identified seven candidate regions with |iHS| >3.0 which were distributed across nine chromosomes, OAR1, 3, 7, 10, 12, 13, 17, 19 and 21 (Table 4; Figure 4b; Supplementary Figure S4), in Barki sheep. Twenty-five genes were located in these regions (Table 4; Supplementary Table S5). We observed two regions at the highest |iHS| on OAR3 (|iHS|=5.18; 112.7–117.9 Mb) and OAR13 (|iHS|=5.24; 51.4–52.6 Mb); and each spanned one gene, TRHDE and BMP2 respectively. The approach further identified eight common genes (GJB2, GJB6, GJA3 and ATP12A on CHI12 and OAR12; FGF9, IFG88 and SAP18 on CHI12 and OAR10, and POLR2K on CHI14 and OAR3, respectively) that were located on candidate regions that were most likely under selection in Barki goats and sheep found in a common environment.
Figure 4.
Genome-wide |iHS| scores in Barki goats (a) and sheep (b). The standardized |iHS| scores generated using an overlapping (5 SNPs) sliding window of 10 SNPs are plotted across the genome. Details of |iHS| for each SNP on each chromosome is shown in Supplementary Figures S3 and S4 for goats and sheep, respectively.
Table 4. Candidate region under selection based on iHS scores in Egyptian Barki sheep and goats.
| Species | Chromosome | Region | Maximum iHS (position)a | Synteny to cattleb (UMD3.1) | Genesc |
|---|---|---|---|---|---|
| Goat | 3 | 27.3–28.6 | 4.04 (27.8) | BTA 3: 29.5–30.9 | MAGI3 |
| 6 | 34.7–35.6 | 4.16 (35.1) | BTA 6: 37.9–38.8 | LAP3 | |
| 82.6–83.2 | 5.10 (82.9) | BTA 6 : 87.0–87.7 | CSN3 | ||
| 7 | 87.0–87.6 | 3.46 (87.2) | BTA 7: 93.3–94.0 | ARRDC3 | |
| 11 | 37.8–38.6 | 3.66 (38.0) | BTA 11: 38.2–39.1 | MIR217 | |
| 68.7–70.6 | 4.22 (69.0) | BTA 11: 69.7–71.7 | ALK | ||
| 85.5–85.8 | 4.00 (85.6) | BTA 11: 86.7–87.1 | PDIA6 | ||
| 12 | 34.2–38.0 | 3.59 (37.7) | BTA 12: 35.5–39.4 | LMLN | |
| 39.8–42.6 | 5.17 (41.3) | BTA 12: 41.2–44.0 | PCDH9 | ||
| 14 | 32.6–32.8 | 4.16 (32.7) | BTA 14: 23.6–23.9 | LYPLA1 | |
| 17 | 18.1–18.3 | 3.97 (18.2) | BTA 17: 18.7–18.9 | ELF2 | |
| 27.1–29.4 | 3.68 (29.3) | BTA 17: 27.6–29.9 | PGRMC2 | ||
| 34.2–34.8 | 4.41 (34.7) | BTA 17:34.8–35.4 | BBS12 | ||
| Sheep | 1 | 237.6–241.2 | 4.66 (240.9) | BTA 1: 121.1–121.8 | KNSTRN |
| 2 | 117.7–123.4 | 4.54 (121.5) | BTA 2: 7.8–9.8 | PLEKHB2 | |
| 3 | 112.7–117.9 | 5.18 (115.2) | BTA 5: 91.5–105.6 | TRHDE | |
| 10 | 36.0–40.3 | 4.18 (37.8) | BTA 12: 36.0–39.7 | ATP12A | |
| 41.5–45.6 | 4.80 (42.9) | BTA 12: 40.7–44.8 | PCDH9 | ||
| 13 | 51.4–52.6 | 5.24 (51.5) | BTA 13: 52.0–53.1 | BMP2 | |
| 17 | 66.9–67.6 | 4.05 (67.1) | BTA 17: ~69 | RPL6 | |
| 19 | 35.3–36.7 | 3.37 (35.3) | BTA 22: ~35 | SUCLG2 | |
| 21 | 19.7–20.5 | 4.57 (20.3) | BTA 29: 22.3–22.9 | PAK1 |
Abbreviation: iHS, integrated Haplotype Score.
Genomic position (Mb) with the highest iHS.
Range defined by the position of conserved genes in cattle.
Gene located at the position with the highest iHS score.
Two genes, SPP1 (CHI6) and FGF2 (CHI17), were identified by both FST and iHS approaches to be in a candidate selection sweep region in Barki goats only.
The iHS analyses also identified candidate selection sweep regions in the exotic breeds of goats and sheep that were, however, too broad (exceeded 5 Mb) to be resolved. The candidate regions with |iHS| >3.0 in the exotic goats, and in particular, the Boer goat, were on CHI3, 4, 7, 8, 13, 15 and 20 (Supplementary Figure S5a). Those in the exotic sheep, were mainly observed in the Romney/Romney-Finn crosses on OAR4, 6, 8, 10, 15 and 22 (Supplementary Figure S5b). None of these candidate regions overlapped with the ones that were identified in the Barki goats and sheep, further confirming their genomic differences.
Signatures of selection that are common to Barki goats and sheep
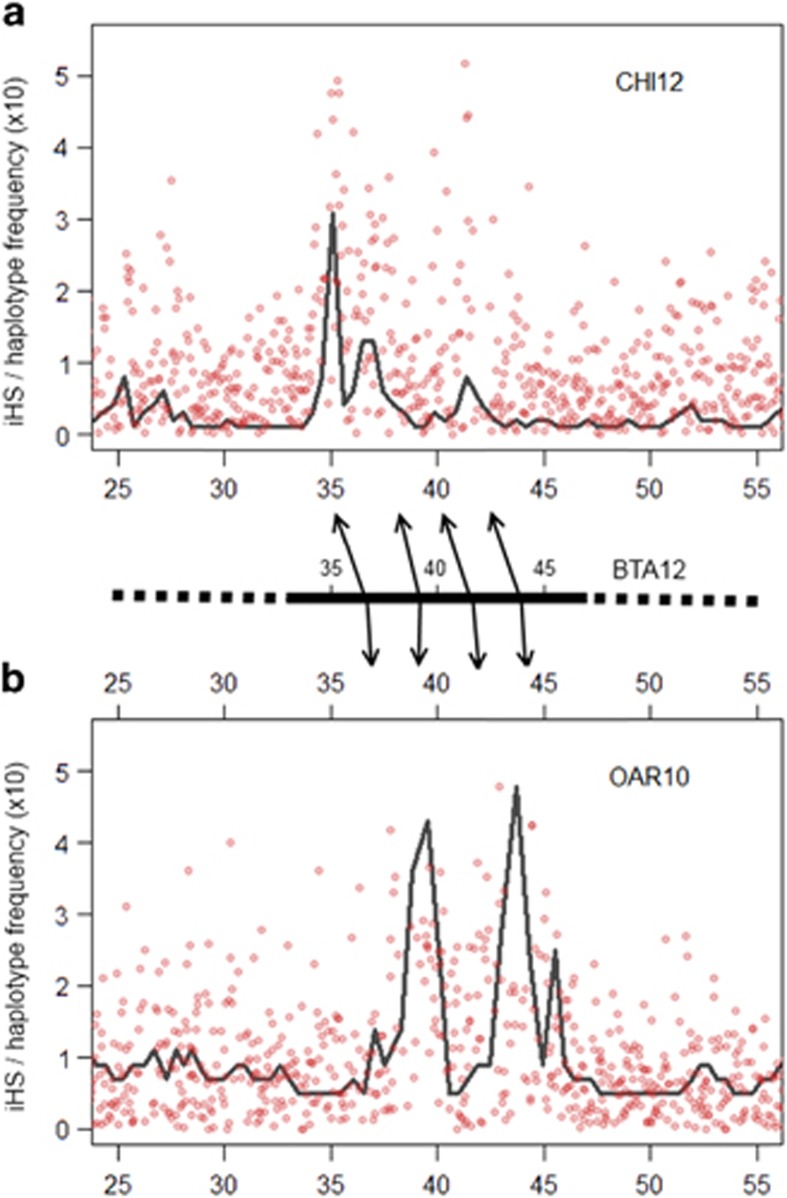
We investigated the genome of Barki goats and sheep to reveal possible candidate regions under selection in a common environment in the two species. For this purpose, we used a comparative genomic map of ruminants and the position of orthologous genes to investigate common regions of conserved synteny among goats, sheep and cattle. We found a candidate selection sweep region on CHI12 and OAR10, respectively, with synteny to bovine chromosome 12 (BTA12) where the presence of a long extended haplotype reveals a region most likely under selection (Figure 5). To fine map this region, we generated |iHS| scores spanning the candidate region in Barki goats (Figure 5a) and sheep (Figure 5b) and also estimated the frequency of the most common haplotype in the two species using 20-SNP sliding windows that overlapped with more than 5 SNPs having iHS scores >3.0. We found that the most common haplotype also had the highest frequency in the two species (Supplementary Figure S6). In spite of the attempt at fine mapping, the region was still too broad (>5 Mb) and spanned more than 20 genes. Five of these genes (FGF9, IFG88, GJB2, GJB6 and GJA3) were among the eight that were identified by iHS to be under selection in both Barki goats and sheep. Although the pattern of the individual haplotypes in each species suggested a narrow region of about 2–3 Mb, the interval spanning the commonest haplotype in Barki goats did not overlap perfectly with its synteny in the Barki sheep (Figure 5).
Figure 5.
Comparison of iHS scores between Barki sheep (OAR10) and Barki goat (CHI12). The absolute value of iHS (red circle) was plotted for each locus with the most frequent haplotype using 20-SNP window (black line) for each SNP position in Barki goats (a) or Barki sheep (b). The map position conserved in cattle is displayed with reference to the bovine genome assembly UMD3.1 in between (a) and (b). CHI, OAR and BTA represent notations for goat, sheep and cattle chromosomes, respectively.
Functional annotation of the candidate genes
The 4 and 15 candidate genes that were found within the four and five genomic regions at the highest iHS and FST values, respectively, and the 8 common candidate genes that were identified by iHS to be under selection in both Barki goats and sheep were evaluated for functional enrichment in specific biological and molecular processes and functions, using the KEGG (http://www.genome.jp/kegg/pathway.html) and Wiki pathways (http://www.wikipathways.org/index.php/WikiPathways) databases. In Supplementary Tables S2 (FST in goats), S3 (FST in sheep), S4 (iHS in Barki goats) and S5 (iHS in Barki sheep) are listed the 119 candidate genes that were identified in this study and their functions. Most of the candidate genes were associated with multiple signaling and signal transduction pathways involving various cellular and biochemical processes; a finding that has also been observed in camels from the analysis of full genome sequences (Jirimutu et al., 2012). This hints that adaptation to environmental stressors most likely is mediated through posttranslational modifications. If this is substantiated through experimental approaches, notwithstanding the fact that signaling genes are overrepresented in the mammalian genome, it would present a huge contribution to our understanding of the genetic mechanisms to environmental stressor adaptation.
The MYH1, MYH3, MYH8, MYH10 and MYH13 genes on OAR11 have a direct role in energy metabolism that facilitates diverse functions including muscular contraction, cytokinesis and phagocytosis (Miyagi et al., 1999). The UGT8 (CHI6), GRIA1 (OAR5), PCDH9 (CHI12) and FGF11 (OAR11) genes are associated with the development and function of the nervous and endocrine systems (Monaghan et al., 1989; Barnard and Henley, 1990; Bosio et al., 1996; Smallwood et al., 1996; Halbleib and Nelson, 2006), while the BMP2 (OAR3), FGF2 (CHI17), FGF9 (CHI12, OAR10), GJB2 (CHI12), GJB6 (CHI12), GJA3 (CHI12, OAR12) and SPP1 (CHI6) genes encode proteins that influence body size, skeletal and embryonic development, and testicular embryogenesis (Abraham et al., 1986; Burgess and Maciag, 1989; Shimoyama et al., 1991; Ornitz et al., 1996; Ortega et al., 1998; Miyagi et al., 1999; Bandyopadyay et al., 2006). The BMP2 gene has also been reported to influence body size in cattle and sheep (Kijas et al., 2012; Xu et al., 2015) and the SPP1 gene was found to be located on a candidate region under selection in sheep (Lv et al., 2014), but the authors concluded that it had no role in the adaptation to climate-mediated selection pressure. The IL2, IL7, IL21 (all on CHI17) and IL1R1 (OAR3) genes encode cytokine receptors that participate in host defense mechanisms, including immune and inflammatory responses (Parrish-Novak et al., 2000). The PIKR5 (OAR11), CNOT8 (OAR5), POLR2K (CHI14) and SAP18 (CHI12) genes are associated with various cellular functions and processes (http://www.genecards.org/). Although coat color and pattern are important in adaptation, we only found FGF2 and FGF9, which encode proteins that regulate melanoma angiogenesis and metastasis (Reiland et al., 2006), in a strong candidate selection sweep region. Two other genes, GNAI3 (CHI3) and PLCB1 (CHI13), although not found in any of the regions at the highest FST and iHS scores, also play a role in melanogenesis. The TRHDE (OAR3) gene encodes an extracellular peptidase that cleaves and inactivates thyrotropin-releasing hormone which is critical in regulating appetite and metabolism (Friedman et al., 1995). Apart from the candidate region that spanned BMP2 (OAR13) and SPP1 (CHI6), none of our candidate regions overlapped with those identified in previous studies in sheep (Kijas et al., 2012; Lv et al., 2014), cattle (Xu et al., 2015) and horses (Petersen et al., 2013), most likely owing to different study design, samples (breeds under different selection pressures) and statistical methods. Lv et al. (2014) found TBC1D12 on OAR22 (15.3–15.4 Mb) and proposed it to be a strong candidate gene for selection in response to environmental stress. The authors found that the frequency of the causative SNP and its haplotype were higher in cold humid regions but low in hot dry regions. Our results appear to support this finding; none of our candidate regions occurred in OAR22 within the region spanning the TBC1D12 gene.
Discussion
By assessing patterns of SNP variation, we present results of a genome-wide scan that were used to detect signatures of selection in indigenous Barki goats and sheep raised in a hot arid environment vis a vis those developed in temperate regions following artificial selection. Our analysis revealed clear genetic differentiation between the two groups of animals which can be attributed to differences in their evolutionary, breeding and management history. We observed longer stretches of ROH in the exotic breeds, possibly the outcome of stringent artificial selection on a few preferred individuals (narrow genetic base) and which is supported by their comparatively higher levels of inbreeding and low effective population sizes. In contrast, the indigenous breeds, which utilize mainly communal grazing resources where individuals from different flocks mix and interbreed, hence their low levels of inbreeding, are exposed mainly to natural selection for survival in their natural environments. Although it can be argued that natural selection should also be able to retain long stretches of ROH among the indigenous breeds, it is likely that it is not as strong as artificial selection and random mating between individuals from different flocks results in high effective population sizes and decay in linkage disequilibrium. Indeed, our selection signature scans show that the candidate regions under selection in the indigenous Barki goats and sheep are narrow and distributed across various chromosomes. It suggests at least, a partial role of natural selection and random mating in shaping the genome of indigenous livestock while maintaining high within- and between-population genetic variability.
Our genome-wide scans revealed strong putative selection sweep regions that span 27 candidate genes and minor ones that spanned 92 candidate genes, with diverse functions. This suggests that adaptation to hot arid environments may be mediated by a complex network of genes that act in tandem rather than the action of single candidate genes. This agrees with the findings of Lv et al. (2014) and Kemper et al. (2014). The former observed that adaptation results from the interaction of several complex traits that most often are controlled by several genes, while the latter noted that selection for complex traits leaves little or no classic signatures of selection owing to weak selection acting on several sites across the genome. It is therefore not surprising that our strongest candidate selection signatures spanned several candidate genes which directly or indirectly influence several traits that are critical for survival in hot arid environments; these included energy and digestive metabolism, muscular function, autoimmune response function and regulation, thermo-tolerance (melanogenesis), and proper embryonic development and reproduction. This appears to be common among species that are adapted to desert and desert-like environments. For instance, in a comparative genomic analysis, Wu et al. (2014) revealed complex features, such as fat and water metabolism, stress response to heat and aridity, intense ultraviolet radiation and chocking dust, in the genome architecture of camelids that are related to adaptation to desert environments.
The biological pathways and functions of candidate genes identified within the putative selection sweep regions indicates that the selection forces shaping the genome architecture of the indigenous Barki goats and sheep might be associated with survival in a hot arid environment. For instance, reduced metabolism and feed intake in cattle under heat stress have been associated with reduced secretion of thyroid hormone and gut motility (Lu, 1989). Ambient temperatures above 30 °C are associated with rapid declines in feed intake and metabolic rates, and at 40 °C, dietary intake may decline by as much as 40% (National Research Council, 1989). Variations in body size and shape have been shown to be closely related to thermoregulation capacity and/or efficiency and this explains why several breeds in the temperate regions where good quality feeds are abundant and temperatures are low have large body sizes, whereas majority of the breeds in the tropics where pastures are low in nutritive value and temperatures are high have small body sizes (McManus et al., 2011). Changes in the patterns of myosin heavy chain isoform expression in response to different physical exercises (strength, endurance and so on) have been reported in humans and represent important adaptations that underlie changes in skeletal muscle strength, resilience and power output (Bottinelli et al., 1999; Ahtiainen et al., 2009). We hypothesize here that the putative selection signatures that span several MYH genes most likely are a response to the need for animals to develop the resilience and endurance to travel long distances to access pastures and watering points. The KIT locus, and its interaction with the MITF locus, has been shown to confer resistance to melanosis and the ability to withstand UV radiation (Satzger et al., 2008; Pausch et al., 2012). Although we detected no obvious signals of selection around the KIT or MITF genes, we still identified selection signatures around genes that are involved in melanogenesis and melanoma angiogenesis. The epidermis, the outermost layer of the skin, protects an organism from the extremes of the external environment, reduces water and heat loss and pathogen entry (Feingold, 2007). In view of recent findings (Zhu et al., 2014) that have implicated the BMP-FGF signaling axis in mediating Wnt-induced epidermis stratification and regeneration, the candidate signatures that flanked the BMP2 and FGF genes may be of interest in understanding the mechanisms of adaptation to hot arid environments.
Although our candidate selection signatures were located within regions enriched with genes of adaptive significance, it can be argued that they could also arise from other factors. Differences in demographic history (genetic drift, effective population sizes, bottlenecks/expansion, inbreeding and so on) have been implicated but are unlikely to explain our results because demographic history shapes variations across the whole genome, whereas the effects of selection tend to be localized in a genomic region (the selected locus and linked genetic markers) (Akey, 2009). Owing to this, outlier loci tend to be strong candidates for natural selection and they can lead to many false positives/negatives (Lachance and Tishkoff, 2013), particularly when an ascertained subset of SNPs is assayed. Although it would be unusual not to have false-positive SNPs on each array, we discount false positives because we performed an extra layer of more stringent quality filtering following the initial SNP genotype calling. Ascertainment bias, a major drawback of SNP arrays (Lachance and Tishkoff, 2013), can affect selection scans. Among our study breeds, only the Boer goat, Texel and Romney sheep were used in the development of the two chips. We doubt that ascertainment bias influenced our results because athough the exotic breeds showed almost similar levels of diversity as the indigenous breeds, especially for goats, as we had expected their Ne and private allele richness values were lower and their inbreeding levels were higher. The strategy used in the development of the chips included divergent breeds in the discovery panel, and prior to their release, they were tested and validated on a large panel of divergent breeds and populations (Kijas et al., 2009; 2012; Tosser-Klopp et al., 2014). Ascertainment bias has also not been reported as an issue in studies that have used the two chips (for example, Kijas et al., 2009; 2012; Lv et al., 2014; Lashmar et al., 2015), and results from these studies and ours show that the two chips are highly polymorphic.
A high degree of similarity in full genome sequences (Dong et al., 2013) and a significant overlap in regions harboring copy number variations (Fontanesi et al., 2011) has been observed among cattle, sheep and goats, indicating minor genomic differences since their divergence from a common ancestor around 19–40 million years ago. If one therefore assumes that goats and sheep share a common genome architecture associated with similar phenotypes, then the expectation is that they should share consensus signatures of natural selection when exposed to a common environment. We, however, detected only one common candidate region under selection in a similar environment in the two species and were unable to delineate its precise boundaries. Genomic evolutionary rate variability has been reported within the family Bovidae (Kijas et al., 2006). Differences in evolutionary rates among goats and sheep may have contributed to our observation and most likely great differences in the mapping location of SNPs on the two chips, which would complicate the identification of common candidate regions. The different stages of the reference genome assemblies of the two species may also have played a role. The sheep genome has been the subject of several improvements and refinements (Maddox et al., 2001; Goldammer et al., 2009), including the development of a virtual genome map based on bovine, dog and human genome assemblies (Dalrymple et al., 2007). These efforts culminated in the first draft of the ovine reference genome (Archibald et al., 2010), which has been improved further and the current released reference assembly, Oar v3.1, has a contig N50 length of ~40 Kb and a total assembled length of 2.61 Gb, with ~99% anchored onto the 26 autosomes and the X chromosome (Jiang et al., 2014). In contrast, similar efforts at developing and refining the goat genome have lagged behind, but recent years have witnessed a slow increase in gene mapping data since the first genetic and cytogenetic maps were produced (Vaiman et al., 1996; Schibler et al., 1998) and ~550 loci have been mapped to the goat genome using linkage maps of low resolution (Schibler et al., 2009). The initial release of the goat genome sequence (Dong et al., 2013) was based entirely on short-read de novo sequencing that yielded an ~ 2.66 Gb assembly with a contig N50 length of 3.06 Mb. Du et al. (2014) have attempted to improve the quality of this genome using additional illumina new-generation short-read sequences and the optical mapping technology of large DNA molecules that have permitted the assembly of 349 super-scaffolds with N50 length of 18.2 Mb. The current assembled base pairs are about 91% of the estimated goat genome (~2.92 Gb). However, it still lacks long-range continuity and its current state only gives a fragmented view of the caprine genome. Our results further demonstrate that improvements in the assembly and annotation of the goat and sheep genomes are necessary to make it possible to compare common conserved regions under selection and investigate their evolutionary history and significance.
Conclusions
Our study investigated signatures of selection in sheep and goats that are indigenous to a hot arid environment using genome-wide SNP scans. We identified several candidate genomic regions and genes underlying local adaptation including a common candidate region under selection in a similar environment in the two species. More detailed studies are, however, necessary to further confirm (for example, experimentally score phenotypic differences between animals) and refine our results by integrating comprehensive genomic data (for example, candidate gene sequencing, high density SNP genotyping, gene expression profiling) with environmental and physiological data. Our results, however, do provide a foundation to investigate ruminant evolution and functional genomics in different species coexisting in a similar environment.
Data archiving
All data analyzed in the present study have been deposited in Dryad. Provisional DOI: 10.5061/dryad.8m1v1.
Acknowledgments
This study was funded under the auspices of Project 9 of the ARC/ICARDA Small Ruminant Program and partially by the CGIAR research program on Livestock and Fish. The genotyping was supported by funding from the Ensminger Fund, the State of Iowa and Hatch funding. Provision of exotic sheep genotypes by D Garrick is appreciated. Support of A.R. Elbeltagy was provided by the ARC/ICARDA Small Ruminant Program, the Ensminger Fund and the State of Iowa funding.
The authors declare no conflict of interest.
Footnotes
Supplementary Information accompanies this paper on Heredity website (http://www.nature.com/hdy)
Supplementary Material
References
- Abraham JA, Whang JL, Tumolo A, Mergia A, Friedman J, Gospodarowicz D et al. (1986). Human basic fibroblast growth factor: nucleotide sequence and genomic organization. EMBO J 5: 2523–2528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahtiainen JP, Hulmi JJ, Kraemer WJ, Lohti M, Pakarinen A, Moro AA et al. (2009). Strength, endurance or combined training elicit diverse skeletal muscle myosin heavy chain isoform proportion but unaltered androgen receptor concentration in older men. Int J Sports Med 30: 879–887. [DOI] [PubMed] [Google Scholar]
- Akey JM. (2009). Constructing genomic maps of positive selection in humans: where do we go from here? Genome Res 19: 711–722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akey JM, Ruhe AL, Akey DT, Wong AK, Connelly CF, Madeoy J et al. (2010). Tracking footprints of artificial selection in the dog genome. Proc Natl Acad Sci USA 107: 1160–1165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alexander DH, Novembre J, Lange K. (2009). Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19: 1655–1664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersson L, Georges M. (2004). Domestic animal genomics: deciphering the genetics of complex traits. Nat Rev Genet 5: 202–212. [DOI] [PubMed] [Google Scholar]
- Archibald AL, Cockett NE, Dalrymple BP, Faraut T, Kijas JW, Maddox JF et al. (2010). The sheep genome reference sequence: a work in progress. Anim Genet 41: 449–453. [DOI] [PubMed] [Google Scholar]
- Bandyopadhyay A, Tsuji K, Cox K, Harfe BD, Rosen V, Tabin CJ. (2006). Genetic analysis of the roles of BMP2, BMP4, and BMP7 in limb patterning and skeletogenesis. PLos Genet 2: e216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbato M, Orozco-terWengel P, Tapio M, Bruford MW. (2015). SNeP: a tool to estimate trends in recent effective population size trajectories using genome-wide SNP data. Front Genet 6: 00109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barnard EA, Henley JM. (1990). The non-NMDA receptors: types, protein structure and molecular biology. Trends Pharmacol Sci 11: 500–507. [DOI] [PubMed] [Google Scholar]
- Bosio A, Binczek E, Lebeau MM, Fernald AA, Stoffel W. (1996). The human gene CGT encoding the UDP-galactose ceramide galactosyl transferase (cerebroside synthase): cloning, characterization, and assignment to human chromosome 4, band q26. Genomics 34: 69–75. [DOI] [PubMed] [Google Scholar]
- Bottinelli R, Pellegrino MA, Canepari M, Rossi R, Reggiani C. (1999). Specific contributions of various muscle fi brefiber types to human muscle performance: an in vitro study. J Electromyography Kinesiol 9: 87–95. [DOI] [PubMed] [Google Scholar]
- Browning SR, Browning BL. (2007). Rapid and accurate haplotype phasing and missing data inference for whole genome association studies using localized haplotype clustering. Am J Hum Genet 81: 1084–1097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burgess WH, Maciag T. (1989). The heparin-binding (fibroblast) growth factor family of proteins. Annu Rev Biochem 58: 575–606. [DOI] [PubMed] [Google Scholar]
- Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV et al. (2013). Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinfo 14: 129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dalrymple BP, Kirkness EF, Nefedov M, McWilliam S, Ratnakumar A, Barris W et al. (2007). Using comparative genomics to reorder the human genome sequence into a virtual sheep genome. Genome Biol 8: R152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong Y, Xie M, Jiang Y, Xiao N, Du X, Zhang W et al. (2013). Sequencing and automated whole-genome optical mapping of the genome of a domestic goat (Capra hircus). Nat Biotechnol 31: 135–141. [DOI] [PubMed] [Google Scholar]
- Du X, Servin B, Womack JE, Cao J, Yu M, Dong Y et al. (2014). An update of the goat genome assembly using dense radiation hybrid maps allows detailed analysis of evolutionary rearrangements in Bovidae. BMC Genomics 15: 625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elferink MG, Megens H, Vereijken A, Hu X, Crooijmans R, Groenen M. (2012). Signatures of selection in the genomes of commercial and non-commercial chicken breeds. Plos One 10: 1371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feingold KR. (2007). Thematic review series: skin lipids. The role of epidermal lipids in cutaneous permeability barrier homeostasis. J Lipid Res 48: 2531–2546. [DOI] [PubMed] [Google Scholar]
- Fontanesi L, Beretti F, Martelli PL, Colombo M, Dall'olio S, Occidente M et al. (2011). A first comparative map of copy number variations in the sheep genome. Genomics 97: 158–165. [DOI] [PubMed] [Google Scholar]
- Friedman TC, Yanovski JA, Jayasvasti V, Yanovski SZ, Koenig RJ, Wilk S. (1995). Pyroglutamyl peptidase-II ("thyroliberinase") activity in human serum: influence of weight and thyroid status. J Clin Endocrinol Metab 80: 1086–1089. [DOI] [PubMed] [Google Scholar]
- Gautier M, Vitalis R. (2012). rehh: an R package to detect footprints of selection in genome-wide SNP data from haplotype structure. Bioinformatics 28: 1176–1177. [DOI] [PubMed] [Google Scholar]
- Goldammer T, Di Meo GP, Lühken G, Drögemüller C, Wu CH, Kijas J et al. (2009). Molecular cytogenetics and gene mapping in sheep (Ovis aries, 2n=54). Cytogenet Genome Res 126: 63–76. [DOI] [PubMed] [Google Scholar]
- Haider AI. (1982). Studies of the performance of some breeds of goats and their crosses under desert condition in Egypt. PhD Thesis, Faculty of Agriculture, Alexandria University: Egypt. [Google Scholar]
- Halbleib JM, Nelson WJ. (2006). Cadherins in development: cell adhesion, sorting, and tissue morphogenesis. Genes Dev 20: 3199–3214. [DOI] [PubMed] [Google Scholar]
- Hanotte O, Dessie T, Kemp S. (2010). Time to tap Africa's livestock genomes. Science 328: 1640–1641. [DOI] [PubMed] [Google Scholar]
- Jiang Y, Xie M, Chen W, Talbot R, Maddox JF, Faraut T et al. (2014). The sheep genome illuminates biology of the rumen and lipid metabolism. Science 344: 1168–1173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jirimutu WZ, Ding G, Chen G, Sun Y, Sun Z et al. (2012). Genome sequences of wild and domestic Bactrian camels. Nat Commun 3: 1202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jombart T, Ahmed I. (2011). adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinfo 27: 3070–3071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kemper KE, Saxton SJ, Bolormaa S, Hayes BJ, Goddard ME. (2014). Selection for complex traits leaves little or no classic signatures of selection. BMC Genomics 15: 246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kijas JW, Lenstra JA, Hayes B, Boitard S, Porto Neto LR, SanCristobal M et al. (2012). Genome-wide analysis of the World's sheep breeds reveals high levels of historic mixture and strong recent selection. Plos Biol 10: 31001258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kijas JW, Menzies M, Ingham A. (2006). Sequence diversity and rates of molecular evolution between sheep and cattle genes. Anim Genet 37: 171–174. [DOI] [PubMed] [Google Scholar]
- Kijas JW, Townley D, Dalrymple BP, Heaton MP, Maddox JF, McGrath A et al. (2009). A genome wide survey of SNP variation reveals the genetic structure of sheep breeds. PLoS One 4: e4668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lachance J, Tishkoff SA. (2013). SNP ascertainment bias in population genetic analyses: why it is important, and how to correct it. Bioessays 35: 780–786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lashmar SF, Visser C, van Marle-Köster E. (2015). Validation of the 50k Illumina goat SNP chip in the South African Angora goat. S Afr J Anim Sci 45: 56–59. [Google Scholar]
- Lu CD. (1989). Effect of heat stress on goat production. Small Rumin Res 2: 151–162. [Google Scholar]
- Lv F-H, Agha S, Kantanen J, Colli L, Stucki S, Kijas JW et al. (2014). Adaptations to climate-mediated selective pressures in sheep. Mol Biol Evol 31: 3324–3343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maddox JF, Davies KP, Crawford AM, Hulme DJ, Vaiman D, Cribiu EP et al. (2001). An enhanced linkage map of the sheep genome comprising more than 1000 loci. Genome Res 11: 1275–1289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McManus C, Louvandini H, Gugel R, Sasaki L, Bianchini E, Bernal F et al. (2011). Skin and coat traits in sheep in Brazil and their relation with heat tolerance. Trop Anim Health Prod 43: 121–126. [DOI] [PubMed] [Google Scholar]
- Miyagi N, Kato S, Terasaki M, Aoki T, Sugita Y, Yamaguchi M et al. (1999). Fibroblast growth factor-9 (glia-activating factor) stimulates proliferation and production of glial fibrillary acidic protein in human gliomas either in the presence or in the absence of the endogenous growth factor expression. Oncol Reprod 6: 87–92. [PubMed] [Google Scholar]
- Monaghan DT, Bridges RJ, Cotman CW. (1989). The excitatory amino acid receptors: their classes, pharmacology, and distinct properties in the function of the central nervous system. Annu Rev Pharmacol Toxicol 29: 365–402. [DOI] [PubMed] [Google Scholar]
- National Research Council. (1989). In: 6th Revised Edition Update Nutrient Requirements of Dairy Cattle. National Academy Press: Washington, DC. [Google Scholar]
- Ornitz DM, Xu J, Colvin JS, McEwen DG, MacArthur CA, Coulier F et al. (1996). Receptor specificity of the fibroblast growth factor family. J Biol Chem 271: 15292–15297. [DOI] [PubMed] [Google Scholar]
- Ortega S, Ittmann M, Tsang SH, Ehrlich M, Basilico C. (1998). Neuronal defects and delayed wound healing in mice lacking fibroblast growth factor 2. Proc Nat Acad Sci USA 95: 5672–5677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parrish-Novak J, Dillon SR, Nelson A, Hammond A, Sprecher C, Gross JA et al. (2000). Interleukin 21 and its receptor are involved in NK cell expansion and regulation of lymphocyte function. Nature 408: 57–63. [DOI] [PubMed] [Google Scholar]
- Pausch H, Wang X, Jung S, Krogmeier D, Edel C, Emmerling R et al. (2012). Identification of QTL for UV-protective eye area pigmentation in cattle by progeny phenotyping and genome-wide association analysis. PLos One 10: 1371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petersen JL, Mickelson JR, Rendahl AK, Valberg SJ, Andersson LS et al. (2013). Genome-wide analysis reveals selection for important traits in domestic horse breeds. PLOS Genet 9: e1003211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. (2007). PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81: 559–575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu Q, Zhang G, Ma T, Qian W, Wang J, Ye Z et al. (2012). The yak genome and adaptation to life at high altitude. Nat Genet 44: 946–949. [DOI] [PubMed] [Google Scholar]
- R Development Core Team. (2013) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria http://www.R-project.org. [Google Scholar]
- Reiland J, Kempf D, Roy M, Denkins Y, Marchetti D. (2006). FGF2 binding, signaling, and angiogenesis are modulated by heparanase in metastatic melanoma cells. Neoplasia 8: 596–606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Satzger I, Schaefer T, Kuettler U, Broecker V, Voelker B, Ostertag H et al. (2008). Analysis of c-KIT expression and KIT gene mutation in human mucosal melanomas. Br J Cancer 99: 2065–2069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schibler L, Di Meo GP, Cribiu EP, Iannuzzi L. (2009). Molecular cytogenetics and comparative mapping in goats (Capra hircus, 2n=60). Cytogenet Genome Res 126: 77–85. [DOI] [PubMed] [Google Scholar]
- Schibler L, Vaiman D, Oustry A, Giraud-Delville C, Cribiu EP. (1998). Comparative gene mapping a fine-scale survey of chromosome rearrangements between ruminants and humans. Genome Res 8: 901–915. [DOI] [PubMed] [Google Scholar]
- Shimoyama Y, Gotoh M, Ino Y, Sakamoto M, Kato K, Hirohashi S. (1991). Characterization of high-molecular-mass forms of basic fibroblast growth factor produced by hepatocellular carcinoma cells: possible involvement of basic fibroblast growth factor in hepatocarcinogenesis. Jpn J Cancer Res 82: 1263–1270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smallwood PM, Munoz-Sanjuan I, Tong P, Macke JP, Hendry SH, Gilbert DJ et al. (1996). Fibroblast growth factor (FGF) homologous factors: New members of the FGF family implicated in nervous system development. Proc Natl Acad Sci USA 93: 9850–9857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szpiech ZA, Jakobsson M, Rosenberg NA. (2008). ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24: 2498–2504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tosser-Klopp G, Bardou P, Bouchez O, Cabau C, Crooijmans R, Dong Y et al. (2014). Design and Characterization of a 52K SNP Chip for Goats. PLos One 9: e86227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vaiman D, Koutita O, Oustry A, Elsen JM, Manfredi E, Fellous M et al. (1996). Genetic mapping of the autosomal region involved in XX sex reversal and horn development in goats. Mamm Genome 7: 133–137. [DOI] [PubMed] [Google Scholar]
- Voight BF, Kudaravalli S, Wen X, Pritchard JK. (2006). A map of recent positive selection in the human genome. PLos Biol 4: e72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu HG, Guang XM, Al-Fageeh MB, Cao JW, Pan SK, Zhou HM et al. (2014). Camelid genomes reveal evolution and adaptation to desert environments. Nat Commun 5: 1–9. [DOI] [PubMed] [Google Scholar]
- Xu L, Bickhart DM, Cole JB, Schroeder SG, Song J, Tassell CP et al. (2015). Genomic signatures reveal new evidences for selection of important traits in domestic cattle. Mol Biol Evol 32: 711–725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeder MA. (2008). Domestication and early agriculture in the Mediterranean Basin: Origins, diffusion, and impact. Proc Natl Acad Sci USA 105: 11597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X, Onteru SK, Dittmer KE, Parton K, Blair HT, Rothschild MF et al. (2012). A missense mutation in AGTPBP1 was identified in sheep with a lower motor neuron disease. Heredity 109: 156–162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X-J, Liu YD, Dai Z-M, Zhang X, Yang XQ, Li Y et al. (2014). BMP-FGF signalling axis mediates Wnt-induced epidermal stratification in developing mammalian skin. PLos Genet 10: e1004687. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.