In addition to providing biologically available nitrogen, nitrogen-fixing nodules represent an important site of thiol biosynthesis, which reprograms sulfur partitioning and metabolism in the plant.
Abstract
We combined transcriptomic and biochemical approaches to study rhizobial and plant sulfur (S) metabolism in nitrogen (N) fixing nodules (Fix+) of Lotus japonicus, as well as the link of S-metabolism to symbiotic nitrogen fixation and the effect of nodules on whole-plant S-partitioning and metabolism. Our data reveal that N-fixing nodules are thiol-rich organs. Their high adenosine 5′-phosphosulfate reductase activity and strong 35S-flux into cysteine and its metabolites, in combination with the transcriptional upregulation of several rhizobial and plant genes involved in S-assimilation, highlight the function of nodules as an important site of S-assimilation. The higher thiol content observed in nonsymbiotic organs of N-fixing plants in comparison to uninoculated plants could not be attributed to local biosynthesis, indicating that nodules are an important source of reduced S for the plant, which triggers whole-plant reprogramming of S-metabolism. Enhanced thiol biosynthesis in nodules and their impact on the whole-plant S-economy are dampened in plants nodulated by Fix− mutant rhizobia, which in most respects metabolically resemble uninoculated plants, indicating a strong interdependency between N-fixation and S-assimilation.
INTRODUCTION
Symbiotic nitrogen fixation (SNF) by rhizobia bacteria in legume plants is a major source of biologically available nitrogen (N) in agricultural and natural ecosystems (Vance, 2001). Rhizobia-legume interaction results in the formation of a specialized plant organ, the root nodule (Oldroyd et al., 2005), within which the symbiotic form of rhizobia, called the bacteroid, can reduce atmospheric N2 to ammonia (NH3). Both partners benefit from this symbiosis, since the plant is supplied with reduced N and amino acids from the rhizobia, which in turn receive organic carbon derived from photosynthesis and other nutrients from the plant (Udvardi and Day, 1997; Prell and Poole, 2006; Udvardi and Poole, 2013). In N-fixing nodules, plant cells and their microsymbionts interact to provide the proper biochemical environment for reduction and assimilation of N, as well as the efficient exchange of metabolites. The establishment of this environment is underpinned by coordinated changes in gene expression in both the plant and rhizobia, which reprograms many biochemical and molecular processes in both symbionts (Colebatch et al., 2002, 2004; Fedorova et al., 2002; Ampe et al., 2003; Becker et al., 2004; Kouchi et al., 2004; Küster et al., 2004; Uchiumi et al., 2004; Benedito et al., 2008). Complete elucidation of the physiological and biochemical changes and their coordination during nodule development is a major goal of ongoing research.
Sulfur (S) is an essential element for SNF, although it has not been studied much in this context. Nitrogenase, the bacterial enzyme responsible for the reduction of N, is a complex [Fe-S] enzyme (Rees and Howard, 2000) that is produced in relatively high amounts by N-fixing bacteroids (Gaude et al., 2004). NifS is a cysteine desulfurase, which uses l-cysteine for the specific mobilization of S for maturation of nitrogenase, and accumulates only under N-fixing conditions (Zheng et al., 1993; Johnson et al., 2005). The Lotus japonicus Sst1 gene, which is expressed in a nodule-specific manner, encodes a sulfate transporter that is essential for SNF (Krusell et al., 2005). The SST1 protein appears to reside on the symbiosome membrane (Wienkoop and Saalbach, 2003) and is thought to transport sulfate from the plant cell cytoplasm to the bacteroids (Krusell et al., 2005). In legume nodules, the content of (homo)glutathione, a S-containing antioxidant, correlates strongly with nitrogenase activity and is able to modulate the efficiency of SNF (Dalton et al., 1993; Matamoros et al., 2003; Groten et al., 2005, 2006; El Msehli et al., 2011).
In plants and many microorganisms growing with sulfate as S-source, sulfate is taken up by specific transporters and subsequently reduced to sulfide for the biosynthesis of cysteine, methionine, coenzymes, and iron sulfur clusters of enzymes (Schmidt and Jager, 1992; Brunold, 1993; Leustek and Saito, 1999; Höfgen et al., 2001; Kopriva, 2006; Takahashi et al., 2011). The first step of sulfate reduction is its activation to adenosine 5′-phosphosulfate (APS), catalyzed by ATP sulfurylase (ATPS) (Schmidt, 1972; Brunold, 1993; Leustek and Saito, 1999; Suter et al., 2000). APS is converted to sulfite, SO32−, by APS reductase (APR) and subsequently reduced to sulfide, S2−, by sulfite reductase (Suter et al., 2000). Cysteine is formed by incorporation of S2− into O-acetyl-l-serine (OAS) via O-acetylserine(thiol)lyase. OAS is synthesized from l-serine and acetyl-CoA by serine acetyltransferase. Cysteine serves as the precursor for other reduced S-compounds, including primary metabolites such as methionine, GSH, homoglutathione (hGSH), coenzymes, and cofactors, as well as secondary compounds like phytochelatins, glucosinolates, and others (Hassinen et al., 2011; Takahashi et al., 2011; Noctor et al., 2012; Van der Weerden and Anderson, 2013).
Previous studies have highlighted the interaction between S- and N-metabolism (Koprivova et al., 2000; Hesse et al., 2004; Kopriva and Rennenberg, 2004). Nodules are the main source of assimilated N for legumes during SNF. However, little is known about the molecular and biochemical mechanisms governing sulfate uptake and assimilation during SNF, either in nodules or nonsymbiotic plant organs. In this article, we present a detailed, whole-plant map of the changes in S-uptake, assimilation, and metabolism triggered by the establishment of SNF, based on transcript profiling and biochemical data from L. japonicus. Our results highlight the function of N-fixing nodules as a strong source of assimilated S for the plant and reveal whole-plant reprogramming of S-metabolism upon the establishment of effective SNF.
RESULTS
Effects of SNF on S-Metabolite Contents and APR Activity
Changes in whole-plant S-metabolism associated with nodule formation and SNF were studied by comparing sulfate and thiol content and APR activity of nodules, roots, stems, and leaves of L. japonicus plants inoculated with wild-type or ΔnifA and ΔnifH mutants of Mesorhizobium loti, and of uninoculated plants. Both mutant strains of M. loti form ineffective (Fix−) nodules with no nitrogenase activity, in contrast to Fix+ nodules containing wild-type rhizobia. In nodules harboring the ΔnifA strain, infected cells contain undifferentiated rhizobia, whereas ΔnifH nodules contain well-differentiated, albeit ineffective bacteroids (Fotelli et al., 2011).
Sulfate levels varied significantly within and between organs, depending on the symbiotic and N-fixing status of plants (Figure 1; Supplemental Data Set 1). Sulfate levels were lowest in nodules of N-fixing plants containing wild-type bacteria and significantly higher in nodules containing either of the nif mutants. Sulfate levels were severalfold higher in roots, stems, and leaves than in nodules of plants inoculated with wild-type bacteria. Sulfate levels were significantly higher in roots of inoculated than of uninoculated plants, whereas sulfate levels in stems of inoculated plants were lower than of uninoculated plants. Interestingly, sulfate levels in each organ were highest for nodulated plants that were unable to fix N due to the presence of Fix− bacteria (Figure 1).
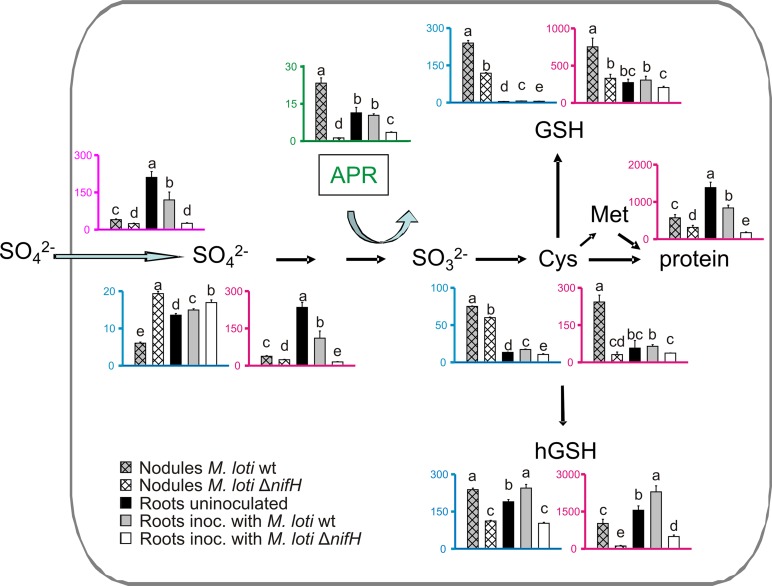
Figure 1.
S-Metabolite Levels and APR Activity in Nodules, Roots, Stems, and Leaves.
Sulfate content (SO42−), APR activity, cysteine (Cys), γEC, GSH, and hGSH content in nodules, roots, stems, and leaves of uninoculated L. japonicus plants and plants inoculated with M. loti wild-type, ΔnifA, or ΔnifH strain are shown. Bars represent means (±se) of five biological replicates. Significant differences at P ≤ 0.05 are indicated by different letters.
APR catalyzes the key step in sulfate reduction (Vauclare et al., 2002), so the activity of this enzyme was measured to gain insight into changes in this pathway in response to symbiosis. In non-nodulated plants, APR activity was highest in leaves, followed by stems and roots (Figure 1). In N-fixing plants, APR activity was highest in leaves, followed by nodules, stems, and roots. Interestingly, APR activity was lower in leaves and stems of N-fixing plants than in those of non-nodulated plants (Figure 1). APR activity in nodules appeared to be linked to N-fixation, as both types of Fix− nodules exhibited very low levels of APR activity. However, the presence of ineffective nodules did not significantly affect APR activity in leaves, in comparison to the leaves of N-fixing plants. These data indicate that N-fixing nodules are an important site of sulfate reduction in plants.
Consistent with the idea that N-fixing nodules are active in sulfate reduction, N-fixing nodules accumulated the highest levels of thiols, with the exception of hGSH, which was most abundant in leaves (Figure 1). Cysteine content in wild-type nodules was 88.4 ± 1.9 nmol g−1 fresh weight (FW), almost 5-fold higher than that in roots of the same plants. Similarly, N-fixing nodules exhibited higher γ-glutamylcysteine (γEC) contents than roots inoculated with wild-type M. loti. GSH accumulated almost exclusively in nodules. Nodule GSH content was found to be 422 ± 15 nmol g−1 FW, more than 24-fold higher than the corresponding roots. hGSH was also abundant in N-fixing nodules reaching 333 ± 53 nmol g−1 FW, a value similar to that of the corresponding roots. In Fix− nodules formed by both M. loti nif mutant strains, levels of all thiols analyzed were substantially lower than in N-fixing nodules.
The presence of N-fixing nodules significantly affected thiol levels in nonsymbiotic organs (Figure 1). Roots of N-fixing plants accumulated higher levels of all thiols than those of uninoculated plants. Roots inoculated with Fix− rhizobia contained lower levels of thiols than roots harboring wild-type rhizobia. The presence of N-fixing nodules increased the level of γEC, but not other thiols, in stems (Figure 1). Nodulation by both nif mutants resulted in lower accumulation of cysteine, γEC, and hGSH in stems. Thiol accumulation, with the exception of GSH, was also higher in leaves of plants harboring N-fixing nodules than in leaves of uninoculated plants (Figure 1). Leaves of Fix− plants accumulated significantly lower levels of cysteine, γEC, and hGSH than leaves of N-fixing plants. Taken together, these results revealed that N-fixing nodules are active in S-reduction and assimilation and possibly serve as a source of S-metabolites for other organs.
35S-Sulfate Uptake and Distribution within L. japonicus Plants Uninoculated or Inoculated with Different M. loti Strains
35S-sulfate was supplied to the root system of intact 6-week-old plants uninoculated or inoculated with the M. loti wild-type or the ΔnifH mutant strain. Total 35S-sulfate uptake per plant, uptake per root system FW, and organ uptake rate were lower in L. japonicus plants inoculated with M. loti wild type than in uninoculated plants (Figures 2 and 3). Surprisingly, the corresponding values for plants nodulated by the ΔnifH strain were in the same range as observed for uninoculated plants, although the FW of all types of plants (Figure 2) and of their organs (Figure 3) were similar, with the exception of nodules of Fix− plants. Relative partitioning of 35S between different plant organs was determined by setting total 35S-sulfate found within the whole plant to 100%. Relative 35S abundance was higher in N-fixing nodules than in ineffective, Fix− nodules harboring the ΔnifH strain (Figure 3). The roots of both types of nodulated plants exhibited lower relative 35S abundance than uninoculated roots. This was accompanied by higher relative 35S abundance in leaves of plants inoculated either with M. loti wild type or the ΔnifH strain than in leaves of uninoculated plants (Figure 3).
Figure 2.
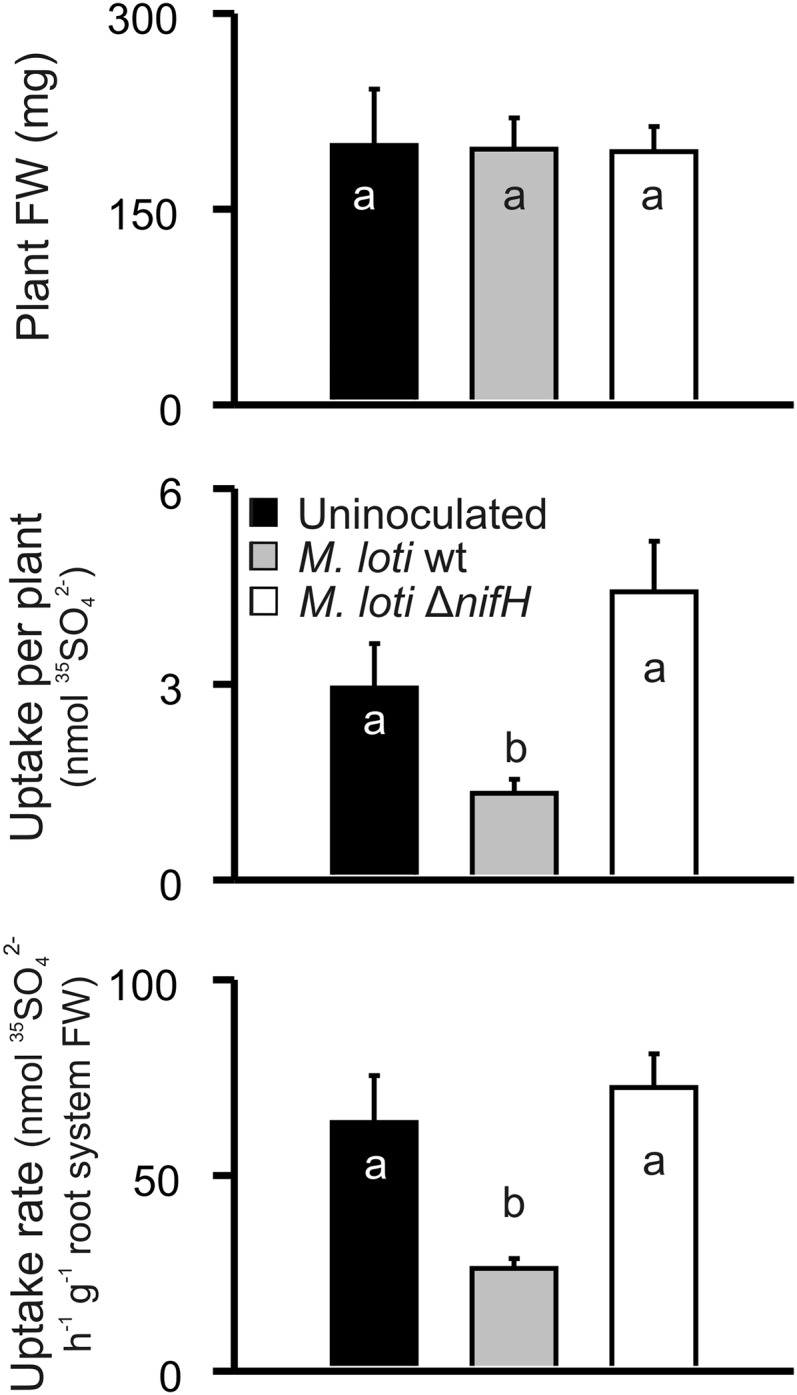
Plant Fresh Weight and 35S-Sulfate Uptake of L. japonicus Plants.
Root systems from intact plants uninoculated and inoculated with M. loti wild-type and ΔnifH strain were exposed to 35SO42− for 1 h. Plant fresh weight, 35S-sulfate uptake per plant, and uptake rate are shown. Mean values (±se) of seven biological replicates are presented. Different letters indicate significantly different values at P ≤ 0.05.
Figure 3.
Organ Fresh Weight and 35S-Contents of Nodules and Nonsymbiotic Organs of L. japonicus Plants.
Root systems of intact plants uninoculated or inoculated with M. loti wild-type or ΔnifH strain were exposed to 35SO42−. After 1 h, nodules, roots, stems, and leaves were separated to determine fresh weight and the radioactivity in each organ. Relative organ FW was calculated from total plant FW (100%). Organ uptake rate was calculated from the amount of 35S and the organ FW. The relative 35S-partitioning was calculated from the amount of 35S within the whole plant (100%). Mean values (±se) of seven biological replicates are presented. Different letters indicate significantly different values at P ≤ 0.05.
35S-Flux into the Sulfate, Thiol, and Protein Pools of Roots and Nodules Exposed to 35S-Sulfate
To further study the potential of N-fixing nodules as S-assimilators, S-flux through the assimilatory sulfate reduction pathway was determined in nodules and roots of 6-week-old L. japonicus plants uninoculated or inoculated with M. loti wild-type or ΔnifH strains. For this purpose, excised nodules and roots were fed with 35S-sulfate for 4 h. Sulfate uptake rate and 35S-fluxes into the sulfate, thiol, and protein pools were measured. In addition, total sulfate and thiol contents as well as APR activity were determined (Figure 4).
Figure 4.
S-Metabolite Content and 35S-Flux into Different Metabolite Pools.
Roots from uninoculated L. japonicus plants and nodules and roots from plants inoculated with wild-type M. loti or with ΔnifH strain were excised and exposed to 35SO42− for 4 h to measure sulfate uptake and the 35S-flux into different metabolite pools. Sulfate uptake (nmol 35S 4 h−1 g−1 FW) is indicated on the graph with purple axes and APR activity (nmol min−1 g−1 FW) on the graph with green axes. Sulfate (μmol g−1 FW) and thiol content (nmol g−1 FW) were determined (blue axes). 35S-flux into internal sulfate (nmol 35S 4 h−1 g−1 FW), thiols (pmol 35S 4 h−1 g−1 FW), and protein (pmol 35S 4 h−1 g−1 FW) are given on graphs with pink axes. Mean values (±se) of eight biological replicates are presented. Bars with different letters indicate significant differences (P ≤ 0.05).
In N-fixing plants, 35S-sulfate uptake and 35S-flux into internal sulfate pools was 65% lower in nodules than in roots. However, 35S-flux into cysteine in N-fixing nodules was 243 ± 28 pmol 35S 4 h−1g−1 FW, representing an almost 4-fold higher flux than in roots of the same plants (Figure 4). This was accompanied by a higher nodule APR activity in N-fixing nodules compared with associated roots. N-fixing nodules exhibited significantly higher 35S-flux into GSH but lower flux into hGSH than did associated roots. 35S-flux into protein was ∼30% lower in N-fixing nodules than in the associated roots (Figure 4). Fix− nodules formed by the ΔnifH strain exhibited lower 35S-sulfate uptake and lower 35S-flux into internal sulfate and cysteine pools compared with wild-type nodules. 35S-flux into cysteine in Fix− nodules was only ∼10% of that in Fix+ nodules and coincided with a 95% reduction in APR activity in the nonfixing nodules. Likewise, 35S-flux into GSH, hGSH, and protein was significantly lower in Fix− than Fix+ nodules (Figure 4).
Uptake of 35S-sulfate was lower in roots of plants inoculated with wild-type M. loti than in uninoculated roots (Figure 4). This result is consistent with a lower 35S-flux into the internal sulfate pool observed in roots inoculated with wild-type M. loti. However, APR activity was similar in these two types of roots, as was 35S-flux into cysteine and GSH. On the other hand, 35S-flux into hGSH was higher and that into protein was lower in roots of N-fixing plants compared with roots of uninoculated plants. 35S-sulfate uptake was lower in roots of plants inoculated with the ΔnifH strain than in roots inoculated with the wild-type rhizobia, which was reflected in a smaller internal 35S-sulfate pool (Figure 4). 35S-flux into cysteine and GSH was slightly lower in Fix− roots than in roots of Fix+ plants, but similar to roots of uninoculated plants. 35S-flux into hGSH was substantially lower (80% lower) in roots inoculated with the M. loti ΔnifH strain than in roots of Fix+ plants. Also, 35S-flux into protein was much lower in roots of Fix− plants than of Fix+ plants. These results show that effective (Fix+) nodulation shifts sulfate assimilation in roots toward hGSH biosynthesis.
Transcript Profiling of Rhizobial Genes Involved in S-Metabolism during SNF
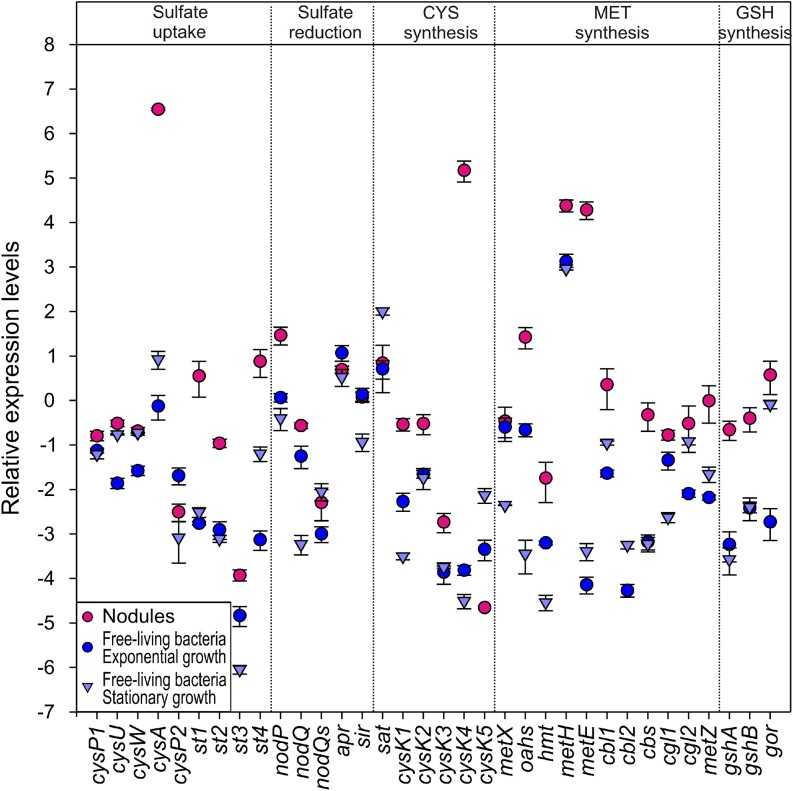
To assess whether bacteria contribute to nodule S-assimilation, transcript profiling of rhizobial genes involved in S-metabolism was performed. Relative transcript levels of M. loti genes involved in S-metabolism of N-fixing bacterioids and free-living bacteria at exponential and stationary phases of growth are presented in Figure 5 and Supplemental Data Set 1. Changes in relative transcript level between bacteroids and free-living M. loti wild type at exponential and stationary phases of growth, and the respective P values, are presented in Supplemental Data Sets 2 and 3, respectively. The qRT-PCR analyses revealed that expression of most of the rhizobial genes involved in S-uptake, reduction, assimilation, and subsequent metabolism were enhanced during SNF compared with both exponential and stationary phase rhizobia. Transcript levels of all sulfate transporters analyzed, except CysP2 and CysW, were higher in bacteroids from nodules than in to free-living bacteria. The cysA sulfate transporter gene showed the greatest upregulation compared with exponential (∼800-fold change) and stationary phase bacteria (∼300-fold change). Within the bacterial sulfate reduction pathway, transcript levels of nodP, coding for the bacterial ATPS, and nodQ, coding for a bifunctional enzyme with ATPS and APS kinase activity, were significantly higher in bacteroids than in free-living bacteria. Relative transcript levels of sir, encoding sulfite reductase, were higher in bacteroids than in stationary phase rhizobia. Cysteine biosynthesis transcript levels were also significantly higher in bacteroids than in free-living bacteria. Transcript levels of the cysteine synthase gene, cysK4, which is located in the symbiosis island, a chromosomally integrated element containing all the genes likely to be required for nitrogen fixation (Sullivan et al., 2002), were ∼8000-fold and ∼16,000-fold higher in bacteroids than in exponential- and stationary-phase rhizobia, respectively. Transcript levels for all enzymes involved in methionine and GSH biosynthesis, except cbl2 which was not detected in bacteroids, were the same or higher in bacterioids than in free-living rhizobia. Transcript levels for the methionine synthase, metE gene, also located in the symbiosis island (Sullivan et al., 2002), were more than 4500- and 2000-fold higher in bacteroids than in exponential- and stationary-phase rhizobia, respectively. Thus, sulfate acquisition and S-assimilation genes were strongly upregulated at the transcriptional level in bacteroids compared with free-living rhizobia.
Figure 5.
Transcript Profiling of M. loti Genes Involved in Sulfate Uptake and Metabolism during SNF.
Relative transcript levels of M. loti genes in nodules formed by M. loti wild-type and free-living rhizobia at the exponential and stationary growth phases. Measurements were conducted on three biological replicates, and transcript levels were normalized using polyribonucleotide nucleotidyltransferase-encoding transcripts as reference. Mean values (±se) are presented in a base e logarithmic scale.
Changes in Transcript Levels of Plant Genes Involved in S-Metabolism during SNF
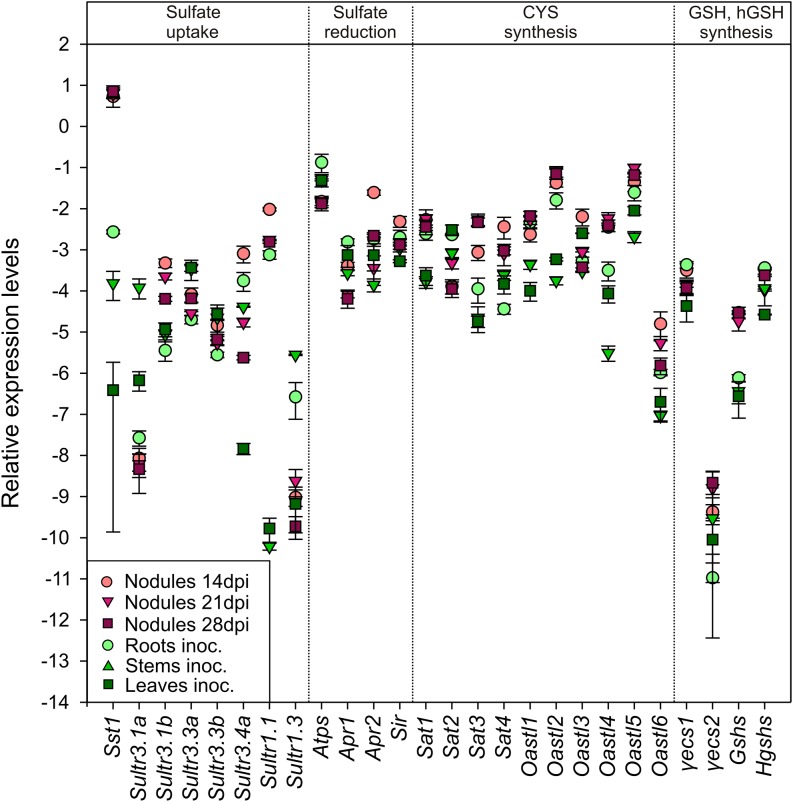
Transcript levels of plant genes involved in S-metabolism in nodules and other organs of nodulated and non-nodulated plants were measured and compared (Figure 6; Supplemental Data Sets 1 and 2). Several plant genes involved in S-metabolism were significantly upregulated during nodule development, compared with roots and other organs (Figure 6). As expected, the nodule-specific sulfate transporter gene Sst1, necessary for SNF (Krusell et al., 2005), was specifically expressed in nodules at all developmental stages. In addition, transcripts of the sulfate transporter genes Sultr3.1b and Sultr1.1 were more abundant in N-fixing nodules, especially in young nodules at 14 d postinoculation (dpi), than in other organs. Within the sulfate reduction pathway, Atps was expressed in nodules at all developmental stages measured, although highest mRNA levels were found in roots of N-fixing plants. Transcripts of Apr2, the main Apr isoform in nodules, were 3- to 9-fold higher in nodules at 14 dpi than in nonsymbiotic organs. Sir showed enhanced expression levels in young nodules (14 dpi) compared with other organs. Transcript levels of all cysteine biosynthesis genes were more abundant in nodules than in other organs, with the exception of Sat2 whose expression levels were lowest in nodules (Figure 6). Sat3 and Sat4 transcripts were found predominantly in nodules. Sat1 and Oastl1 transcript levels were enhanced during nodule development in comparison to the aboveground organs. Oastl3 and Oastl6 transcripts were most abundant in early stage nodules, although Oastl6 expression levels were low compared with the other Oastl isoforms. All the remaining Oastl isoforms were consistently upregulated in N-fixing nodules, with Oastl2 and Oastl5 being the predominant Oastl isoforms in nodules. Likewise, genes involved in GSH biosynthesis tended to be more highly expressed in nodules than in other organs. Transcript accumulation of γecs1, the main γ-glutamylcysteine synthetase isoform since its relative expression levels were over 100-fold higher than γecs2, was highest in roots and in young nodules at 14 dpi. In contrast, transcript levels of γecs2 were 5- to 10-fold higher in nodules than in roots, regardless of the nodule developmental stage. Transcripts of glutathione synthetase Gshs were 4- to 8-fold more abundant in nodules at all developmental stages than in nonsymbiotic organs. In contrast, relative expression levels of homoglutathione synthetase Hgshs were highest in roots and in early stage nodules at 14 dpi. In summary, most of the plant genes involved in sulfate transport and assimilation were more highly expressed in nodules than in other organs during SNF.
Figure 6.
Transcript Profiling of L. japonicus Genes Involved in Sulfate Uptake and Metabolism during SNF.
Relative transcript levels of L. japonicus genes in nodules formed by M. loti wild-type strain at various stages of development, namely at 14, 21, and 28 dpi, as well as in nonsymbiotic organs of the inoculated L. japonicus plants. Measurements were conducted on three biological replicates and transcript levels were normalized using ubiquitin-encoding transcripts as reference. Mean values (±se) are presented in a base e logarithmic scale.
Profiling of Bacterial and Plant Transcripts Involved in S-Metabolism in Fix− Nodules Formed by ΔnifH and ΔnifA Strains
To determine whether the transcriptional changes associated with S-metabolism in nodules are linked to active N-fixation, transcript profiling was also performed on Fix− nodules formed by either ΔnifA or ΔnifH rhizobia. Relative transcript levels of M. loti and L. japonicus genes in ineffective and N-fixing nodules are presented in Figure 7 and Supplemental Data Sets 1, 2 and 3. Focusing first on bacterial gene expression, transcripts of the cysA sulfate transporter were ∼50% less abundant in Fix− nodules formed by the ΔnifA and ΔnifH strains than in Fix+ nodules containing wild-type rhizobia, while transcript levels of the sulfate transporters cysP1, cysU, cysW, st1, and st2 were higher in Fix− nodules than in Fix+ nodules (Figure 7). However, the relative expression levels of these transporters were low in comparison to cysA. Transcript levels of the cysteine synthase gene, cysK4, encoding the main isoform expressed in N-fixing bacteroids, were approx. 85-fold lower in the mutant bacteria. The relative transcript levels of γ-glutamylcysteine synthetase, gshA, were also lower in nodules harboring ΔnifA and ΔnifH rhizobia. In both types of Fix− nodules, transcript levels of methionine synthetase metH and of methionine synthetase metE, which was found to be highly upregulated in N-fixing nodules, were much lower in comparison to N-fixing nodules. In summary, many of the rhizobial genes required for sulfate uptake and assimilation were expressed at lower levels in Fix− nodules than in Fix+ nodules.
Figure 7.
Heat Map of Transcriptional Changes in L. japonicus and M. loti Genes Involved in Nodule Sulfate Uptake, Reduction, and Assimilation during SNF.
(A) Heat map of fold changes in relative transcript levels of L. japonicus genes in nodules formed by M. loti wild-type in comparison to the transcript levels in roots of the same plants.
(B) and (C) Heat map of fold changes in relative transcript levels of L. japonicus genes in Fix− nodules formed by either ΔnifA or ΔnifH strain, respectively, in comparison to the transcript levels in nodules formed by the wild-type M. loti.
(D) Heat map of fold changes in relative transcript levels of M. loti genes in N-fixing nodules in comparison to the transcript levels in free-living rhizobia at the exponential growth phase.
(E) and (F) Heat map of fold changes in relative transcript levels of M. loti genes in Fix− nodules formed by ΔnifA or ΔnifH strain, respectively, in comparison to the transcript levels in N-fixing nodules. Measurements were conducted on three biological replicates, and transcript levels of bacterial and plant genes were normalized using polyribonucleotide nucleotidyltransferase- and ubiquitin-encoding transcripts as references, respectively. Statistically significant differences are indicated by asterisks. The whole data set of fold changes and the statistical analysis are available in Supplemental Data Sets 2 and 3, respectively.
Most of the plant genes involved in sulfate uptake, reduction, assimilation, and metabolism were found to be expressed at significantly lower levels in Fix− nodules than in Fix+ nodules (Figure 7; Supplemental Data Sets 1 and 2). This included all sulfate transporter genes, except for Sultr1.3. For the nodule-specific Sst1 transporter, transcript levels were substantially lower in nodules formed by ΔnifA and ΔnifH strains compared with the wild-type strain. For the plant sulfate reduction pathway, Atps transcript levels were lower in nodules harboring ΔnifH rhizobia, while Apr2 and Sir transcript levels were lower in both types of Fix− nodules than in Fix+ nodules. Transcript levels of Sat1, Sat4, and of all the Oastl isoforms were significantly lower in both Fix− than in Fix+ nodules. Beyond cysteine biosynthesis, mRNA levels of γecs1 and Hgshs, responsible for γEC and hGSH biosynthesis, respectively, were lower in both Fix− than Fix+ nodules. The expression level of Gshs, responsible for the GSH biosynthesis, was lower in the nodules formed by the ΔnifA strain. The expression of other plant genes remained unaffected by mutations in the rhizobial nif genes (Supplemental Data Sets 2 and 3).
SNF Triggers Changes in Expression of Plant S-Metabolism Genes at the Whole-Plant Level
To explore the impact that sulfate assimilation in N-fixing nodules has on whole-plant S-metabolism, we measured and compared transcript levels of S-transport and assimilation genes in various organs of Fix+ and Fix− plants. Several plant genes involved in S-metabolism were expressed at lower levels in uninoculated roots and roots of Fix− plants than in roots of Fix+ plants, while no genes were expressed at higher levels in roots of non N-fixing plants (Figure 8). Expression of the sulfate transporter gene Sultr1.1 was lower in uninoculated roots and roots of Fix− plants than in roots of Fix+ plants. Systemic transcriptional changes were also observed in the sulfate reduction pathway. Transcript levels of Apr1 in uninoculated roots and of Apr2 in roots of Fix− plants were lower than in roots of Fix+ plants. Within the cysteine biosynthesis pathway, transcript accumulation of Sat1, Sat2, Oastl1, Oastl3, and Oastl5 was significantly lower in uninoculated roots and roots of Fix− plants than in roots of Fix+ plants. Genes encoding three additional Oastl isoforms, Oastl2, Oastl4, and Oastl6, were also expressed at lower levels in uninoculated roots than in roots of N-fixing plants. Transcript levels of γecs1 were lower in uninoculated roots and roots of Fix− plants than in roots of Fix+ plants. Finally, mRNA levels of Hgshs were lower in uninoculated roots and roots inoculated with the ΔnifH strain than in roots of Fix+ plants inoculated with the wild-type M. loti.
Figure 8.
Heat Map of Transcriptional Changes in L. japonicus Genes Involved in Sulfate Uptake, Reduction, and Assimilation at the Whole-Plant Level during SNF.
Heat map of fold changes in relative transcript levels in nonsymbiotic organs, including roots (R), stems (S), and leaves (L) of uninoculated plants (A) and plants inoculated with M. loti ΔnifA (B) and ΔnifH (C) strains in comparison to the transcript levels in plants inoculated with wild-type M. loti. Measurements were conducted in three biological replicates, and transcript levels were normalized using ubiquitin as a housekeeping gene. Statistically significant differences are indicated by asterisks. The whole data set of fold changes and the statistical analysis are available in Supplemental Data Sets 2 and 3, respectively.
In stems, mRNA levels of the sulfate transporter Sultr3.4a were lower in uninoculated and Fix− plants than in Fix+ plants. Transcript levels of the sulfate transporters Sultr1.3 and Sultr3.1b were also lower in stems of uninoculated plants than in N-fixing plants. In contrast, transcripts of Sultr3.3a were higher in stems of Fix− than of Fix+ plants. In stems of uninoculated plants, lower transcript levels were found for all of the genes of the sulfate reduction pathway (Atps, Apr1, Apr2, and Sir) and for several genes involved in cysteine and GSH and hGSH [(h)GSH] biosynthesis, including Sat1, Sat2, Sat4, Oastl1, Oastl4, Oastl5, and γecs1, compared with stems of Fix+ plants. Likewise, expression of Apr2, Sat4, and Oastl3 was lower in the stems of Fix− compared with Fix+ plants. In contrast, stems of Fix− plants had higher transcript levels of Sat1, Oastl2, and Oastl5 than did stems of Fix+ plants.
In leaves, transcript levels of the sulfate transporter gene Sultr3.3a were lower in uninoculated and Fix− plants than in Fix+. Transcript levels of the mitochondrial serine acetyltransferase gene Sat4 and the cytosolic O-acetylserine(thiol)lyase gene Oastl2 were higher in leaves of uninoculated and Fix− plants than in leaves of N-fixing plants. Transcript levels of Hgshs were lower in leaves of uninoculated and Fix− plants than in leaves of Fix+ plants.
In summary, the presence of N-fixing nodules with substantial sulfate assimilation capacity had a significant impact on the expression of many genes involved in sulfate transport and assimilation throughout the plant. In general, genes involved in sulfate uptake were found to be upregulated, while numerous genes for sulfate reduction and assimilation were expressed at lower levels in other (non-nodule) organs of N-fixing plants than in those of uninoculated or Fix− plants (Figure 8). In other words, N-fixation and sulfate assimilation in nodules was accompanied by transcriptional reprogramming of many of the plant genes involved in S-assimilation in other organs.
DISCUSSION
It is well established that uptake, assimilation, and metabolism of N and S are tightly connected in plants (Smith, 1980; Clarkson et al., 1989; Koprivova et al., 2000; Ho and Saito, 2001; Vauclare et al., 2002; Hesse et al., 2004; Kopriva and Rennenberg, 2004; Davidian and Kopriva, 2010). In this context, several studies demonstrated the importance of sulfate uptake and metabolism for the establishment and function of SNF (Matamoros et al., 1999, 2003; Abbas et al., 2002; Harrison et al., 2005; Krusell et al., 2005; Bianucci et al., 2008; Muglia et al., 2008; Chang et al., 2009; Becana et al., 2010; El Msehli et al., 2011; Frendo et al., 2013). However, little is known about the molecular and biochemical mechanisms governing S-reduction and metabolism during SNF and the impact of nodulation on the whole-plant S-metabolism. This study revealed that nodules are a strong source of assimilated S for the whole plant and trigger whole-plant reprogramming of S-partitioning and metabolism.
N-Fixing Nodules Are a Strong Source of Assimilated S
Our results from 35S-flux analysis, APR activity, and S-metabolite measurements combined with transcriptomic analysis in nodules clearly indicate that S-uptake, assimilation, and metabolism are enhanced in both symbionts during SNF. To test whether the enhancement of S-metabolism in nodules is linked to active SNF or whether it is just related to nodule development, we experimented with M. loti ΔnifA and ΔnifH mutant strains. The ΔnifA strain triggers aberrant nodule development, fails to differentiate into bacteroids, and has no detectable nitrogenase activity, whereas the ΔnifH strain triggers normal nodule development but with no detectable nitrogenase activity (Fotelli et al., 2011). The results from these experiments with Fix− nodules revealed that enhanced S-assimilation and metabolism is strongly linked to active SNF. The major steps of S-uptake and metabolism upregulated in nodules and regulated by SNF are summarized in Figure 9.
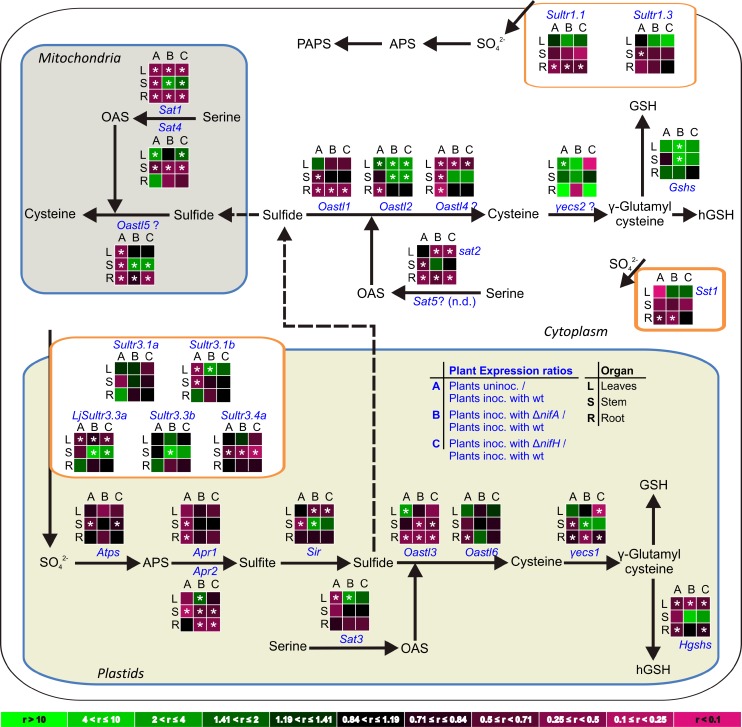
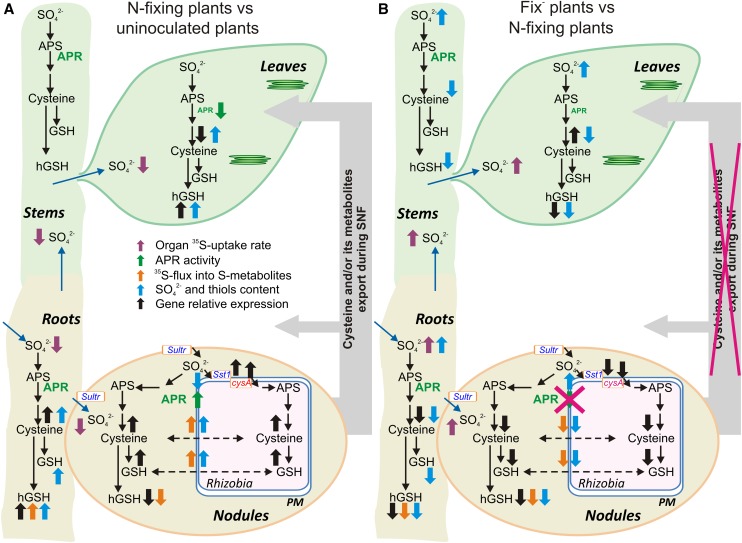
Figure 9.
Schematic Presentation of Sulfate Uptake, Reduction, and Assimilation at the Whole-Plant Level during SNF Indicating That N-Fixing Nodules Represent a Significant Source of Assimilated S for the Plant and Trigger Whole-Plant Reprogramming of S-Metabolism upon the Establishment of Effective SNF.
(A) Differences in S-uptake, reduction, and assimilation observed in N-fixing plants compared with uninoculated plants are presented. Nonsymbiotic organs of N-fixing plants are compared with the respective organs of uninoculated plants. Nodules are compared with roots of N-fixing plants and bacteroids to the free-living rhizobia.
(B) Differences in S-uptake, reduction, and assimilation observed in Fix− plants compared with N-fixing plants are presented. Nonsymbiotic organs of Fix− plants are compared with the respective organs of N-fixing plants. Fix− nodules are compared with N-fixing nodules and Fix− rhizobia to N-fixing bacteroids.
Differences in organ 35S-uptake rate, APR activity, 35S-flux into S-metabolites, sulfate and thiols content, and transcriptional regulation are indicated by purple, green, orange, blue, and black arrows, respectively.
The first clear hint that N-fixing nodules are an important site of S-assimilation in L. japonicus came from the observation that they are the main site of cysteine accumulation in these plants (Figure 1). This observation was consistent with those of Matamoros et al. (1999), who reported that nodules from pea (Pisum sativum), broad bean (Vicia faba), alfalfa (Medicago sativa), soybean (Glycine max), mung bean (Vigna radiata), and cowpea (Vigna unguiculata) plants are also rich in cysteine. In addition, our data suggest that cysteine is synthesized in situ in N-fixing nodules and not imported from nonsymbiotic organs. This conclusion is supported by the observation that N-fixing nodules appeared to be, together with leaves, a major site of sulfate reduction and assimilation, exhibiting high APR activity and 35S-flux into cysteine (Figures 1, 4, and 9). The high APR activity and cysteine content in nodules were found to be linked to active SNF, as APR activity and 35S-flux into cysteine were largely abolished in Fix− nodules, resulting in significantly lower cysteine content than in Fix+ nodules. The link between S-assimilation and active SNF in nodules presumably reflects the regulation of sulfate assimilation by N-availability (Clarkson et al., 1989; Kopriva and Rennenberg, 2004; Davidian and Kopriva, 2010). Both ammonium and amino acid levels have been implicated as significant factors in the regulation of APR activity and subsequent S-assimilation (Brunold and Suter, 1984; Koprivova et al., 2000; Davidian and Kopriva, 2010). In line with these reports, our data suggest that nodules function as strong sulfate assimilators only in the presence of active SNF, indicating that the immediate products of N-fixation act as positive regulators of S-assimilation and metabolism in nodules.
Active cysteine biosynthesis in nodules was underpinned by SNF-dependent transcriptional upregulation of many bacterial and plant genes for S-uptake and assimilation (Figure 9). As a starting point, several plant and rhizobial sulfate transporter genes were found to be induced during nodule development, in an SNF-dependent manner (Figures 7 and 9). These included the plant genes Sst1 and Sultr1;1. SST1 is located on the symbiosome membrane of L. japonicus nodules (Wienkoop and Saalbach, 2003) and is believed to transport sulfate from the host cell cytoplasm to the bacteroids (Krusell et al., 2005). SST1 is crucial for SNF in L. japonicus, as sst1 mutant plants develop Fix− nodules, which indicates that sulfate is a primary source of S for bacteroids (Krusell et al., 2005). Transcriptional upregulation of the rhizobial cysA gene, which encodes one of the five subunits of a sulfate transport complex (Laudenbach and Grossman, 1991; Kredich, 1996; Piłsyk and Paszewski, 2009), may ensure sulfate uptake from the periplasmic space into the bacteroids. In Gram-negative bacteria, CysA is known to form a dimeric periplasmic bridge between the outer membrane and cytoplasmic membrane subunits of the sulfate transport complex (Scheffel et al., 2005; Piłsyk and Paszewski, 2009). In the bacteria Salmonella typhimurium, Escherichia coli (Kredich, 1992), and Klebsiella aerogenes (Lynch et al., 1994), activation of genes involved in sulfate uptake, activation, reduction, and assimilation is dependent on high levels of acetylserine, linking S-assimilation to N availability. Several bacterial and plant genes involved in cysteine biosynthesis were also found to be transcriptionally upregulated in nodules in response to SNF. The M. loti cysteine synthase gene cysK4, residing on the symbiosis island of the genome (Sullivan et al., 2002), was strongly induced in N-fixing bacteroids, but not in Fix− nodules harboring rhizobial nif mutants (Figures 5, 7, and 9). Most plant Sat and Oastl genes were also induced during nodule development in an SNF-dependent manner, including mitochondrial Sat4 and Oastl5, and cytosolic Oastl2 and Oastl4 (Figures 6 and 9). In Arabidopsis thaliana, mitochondrial serine acetyltransferase plays the predominant role in cellular OAS formation (Heeg et al., 2008; Haas et al., 2008; Watanabe et al., 2008; Krueger et al., 2009; Takahashi et al., 2011; Rennenberg and Herschbach, 2014), while the main site of cysteine biosynthesis is considered to be the cytosol (Heeg et al., 2008; Watanabe et al., 2008; Krueger et al., 2009; Rennenberg and Herschbach, 2014). All of these transcriptional changes that promote cysteine biosynthesis were largely abolished in Fix− nodules (Figures 7 and 9), indicating that SNF positively affects this process in nodules. Downregulation of cysteine biosynthesis in Fix− nodules was reflected in the relatively high levels of sulfate that accumulated in ineffective nodules (Figures 1 and 9).
Cysteine is a precursor of other important S-compounds in nodules, including γ-glutamylcysteine, (h)GSH, and methionine, production of which was also reduced in Fix− compared with Fix+ nodules (Figures 1, 4, and 9). Biosynthesis of these thiols appeared to be transcriptionally regulated since several of the rhizobial and plant genes involved were upregulated in N-fixing nodules, in contrast to Fix− nodules (Figures 5, 6, 7, and 9). The importance of these thiols in legume-rhizobia symbiosis is well documented. (h)GSH plays a crucial role in nodule development and SNF efficiency, since Mt-γECS RNAi-silenced lines of M. truncatula developed smaller nodules with decreased (h)GSH content and lower acetylene reduction activity per nodule than control plants (El Msehli et al., 2011). With respect to methionine, previous studies have demonstrated that all Sinorhizobium meliloti Rmd20I methionine auxotrophs (metA/metZ, metE, and metF mutants) formed ineffective nodules on alfalfa plants, a phenotype related mainly to the reduced number of infected nodule cells and incomplete bacterial differentiation into bacteroids (Abbas et al., 2002). This Fix− phenotype was completely reverted by addition of methionine to the plant nutrient medium, suggesting that infected plant cells can but normally do not provide sufficient methionine for bacteroid development and SNF. Although relevant literature is not available for M. loti, our results indicate that M. loti bacteroids are geared to produce methionine via induction of genes for methionine biosynthesis. These include metE, a gene located in the M. loti symbiosis island (Sullivan et al., 2002), whose expression was several thousand-fold induced in N-fixing bacteroids (Figures 5 and 9). The majority of bacterial genomes encode more than one enzymatic mechanism for the methylation of homocysteine to generate methionine. However, methionine production is dependent not only on the genetic/metabolic potential of the bacteria, but also on the availability of the precursors for each pathway (Barra et al., 2006). In M. loti, the contribution of alternative pathways of methionine biosynthesis in free-living bacteria and N-fixing bacteroids could differ due to differences in the availability of methionine precursors acting as methyl-donors, namely, 5-methyltetrahydropteroyl-tri-l-glutamate for the cobalamin-independent MetE and 5-methyltetrahydrofolate for the cobalamin-dependent MetH (Mordukhova and Pan, 2013). Further studies are needed in order to elucidate the role of methionine in N-fixing bacteroids, and the construction of M. loti methionine auxotrophs is underway.
Overall, our data suggest that sulfate reduction, assimilation, and metabolism are activated in N-fixing nodules, although 35S-sulfate uptake was lower in nodules than in roots (Figure 9). This result might reflect the specialized role of roots in sulfate uptake from the nutrient medium. Irrespective of lower sulfate uptake compared with roots, N-fixing nodules exhibited significantly higher APR activity and fluxes into cysteine and GSH compared with roots. Surprisingly, this was observed despite the high nodule thiol content (Figures 1, 4, and 9). This is in contrast to previous studies where an excess of thiols repressed APR activity and S-assimilation through reduction of 35S-flux into thiols (Herschbach and Rennenberg, 1994; Zhao et al., 1999; Vauclare et al., 2002; Hesse et al., 2004). This apparent discrepancy could reflect different biochemical mechanisms regulating S-uptake and assimilation and/or compartmentalization of thiol biosynthesis and thiol accumulation within the infected nodule cells. N-fixing bacteroids could actively participate in the increased efficiency of S-assimilation in nodules by the compartmentalization of S-metabolism in the infected cell. Thus, they appear to be significant players in nodule sulfate assimilation. Most of the rhizobial genes for sulfate uptake and metabolism were induced in N-fixing nodules compared with either free-living bacteria or Fix− nodules (Figures 5 and 7), making them potential sinks for sulfate and sources of thiols. Bacteroids are able to maintain oxidative phosphorylation to supply energy for N-fixation (De Visser and Lambers, 1983), despite the very low free-oxygen concentrations that are a prerequisite for nitrogenase activity (Bergersen, 1982; Kuzma et al., 1993; Udvardi and Poole, 2013), via the involvement of leghemoglobin (Appleby, 1984; Ott et al., 2005). Thus, they are well equipped to supply energy for S-assimilation under the microaerobic conditions of nodules. Although more experiments are needed to determine the relative contribution of N-fixing bacteroids to nodule sulfate assimilation and metabolism, our data indicate that they play a significant role in this regard.
S-Assimilation in N-Fixing Nodules Influences Whole-Plant S-Partitioning and Metabolism
Although S is a crucial element in many molecules involved in SNF, such as proteins and protein cofactors (Zheng et al., 1993; Dos Santos et al., 2004; Johnson et al., 2005), our results suggest that thiols synthesized in nodules do not only fulfill the needs of SNF, but also contribute to the S-economy of the whole plant (Figure 9). N-fixing nodules, as a likely source of reduced-S, triggered global reprogramming of sulfate uptake, reduction, and assimilation at both the transcriptional and metabolite levels (Supplemental Data Sets 1 and 2). Interestingly, many of these changes are dependent on SNF because they were not observed in Fix− plants, which in most respects resembled, metabolically, uninoculated plants.
Our findings indicate that thiols synthesized in N-fixing nodules could be subjected to long-distance transport contributing to the S-budget of other organs. Indeed, reduced S-compounds have been detected in the xylem sap of several species (Rennenberg et al., 1994a, 1994b; Schneider et al., 1994b, 1994a; Schupp et al., 1991; Herschbach and Rennenberg, 2001a, 2001b). Composition and contents of S-compounds in the xylem sap vary between species, season, S-nutrition, developmental stage, part of the trunk, as well as with the presence of mycorrhization (Herschbach and Rennenberg, 2001a, 2001b; Herschbach et al., 2012). GSH is considered to be the predominant reduced-S compound subjected to long-distance transport in plants (Schneider et al., 1994a, 1994b; Herschbach and Rennenberg, 1995; Herschbach et al., 2012; Gigolashvili and Kopriva, 2014). However, many plants also contain cysteine and γEC in appreciable amounts in the xylem sap (Schupp et al., 1991; Rennenberg et al., 1994a, 1994b; Blaschke et al., 1996). In xylem sap of beech trees, cysteine is the predominant low molecular weight thiol, whereas GSH is present in minute amounts (Schupp et al., 1991; Rennenberg et al., 1994a;Herschbach et al., 2012). It has been well documented that transport of thiols in the xylem sap contributes to the whole plant sulfur cycling and nutrition (Rennenberg and Herschbach, 2014).
The higher cysteine content found in leaves of N-fixing plants in comparison to uninoculated and Fix− plants is unlikely to have resulted from enhanced local biosynthesis because APR activity in leaves of N-fixing plants was found to be significantly lower than in uninoculated plants (Figures 1 and 9). Interestingly, transcript levels of both Apr genes in leaves were not affected by the presence of either nodules or of SNF. Thus, the decrease of APR activity in nodulated plants may reflect posttranscriptional or posttranslational regulation of APR, which is known to occur in plants (Bick et al., 2001; Kopriva and Koprivova, 2004; Koprivova et al., 2008; Davidian and Kopriva, 2010; Takahashi et al., 2011). The low APR activity and cysteine levels in the leaves of Fix− plants could account for both the accumulation of sulfate and the lower thiol levels throughout the plant. A similar observation was made in Arabidopsis, where reduced APR activity correlated with high foliar sulfate levels (Loudet et al., 2007; Davidian and Kopriva, 2010).
Downregulation of cysteine biosynthesis in the leaves of N-fixing plants was also apparent at the transcriptional level. Several genes involved in this process, including mitochondrial Sat4 and cytoplasmic Oastl2, exhibited significantly lower transcript levels in leaves of N-fixing plants than of uninoculated and Fix− plants (Figures 8 and 9). The higher cysteine content found in roots of Fix+ plants compared with uninoculated and Fix− plants also points to nodules as a source of cysteine for other organs, since root APR activity and 35S-flux into cysteine were similar in uninoculated, Fix+, and Fix− plants (Figures 1, 4, and 9).
Our data indicate that GSH may also be transported from the N-fixing nodules to the adjacent roots. Similar Gshs transcript levels and 35S-flux into the GSH pool in roots of uninoculated, Fix+, and Fix− plants do not suggest higher GSH biosynthesis in roots of Fix+ plants and cannot account for their higher GSH content (Figures 1, 4, 8, and 9). On the other hand, N-fixing nodules appeared to be the main site of GSH biosynthesis, exhibiting relatively high transcript levels of Gshs and 35S-flux into GSH (Figures 4, 6, and 9). Supporting the notion that nodules are a source of GSH for roots and other organs, GSH is known to be transported over long distances in plants (Schneider et al., 1994a, 1994b; Herschbach and Rennenberg, 1995; Herschbach et al., 2012; Gigolashvili and Kopriva, 2014). In contrast, our results do not point to long-distance transport of hGSH from nodules to other organs, as the elevated levels of hGSH in roots and leaves of N-fixing plants, compared with uninoculated and Fix− plants, correlated well with upregulation of the Hgshs gene in both organs (Figures 1, 8, and 9). Furthermore, greater 35S-flux into hGSH in the roots of Fix+ plants compared with uninoculated and Fix− plants (Figures 4 and 9) is indicative of the local biosynthesis of hGSH.
The intriguing hypothesis that nodules serve as a substantial source of thiols for other organs is further supported by the relatively low 35S-flux into protein observed in nodules compared with roots of Fix+ plants (Figure 4). This indicates that the biosynthesis of nodule S-containing proteins, including nitrogenase subunits, generates a lower demand for S-flux into protein than protein biosynthesis in roots. Thus, the demands for protein biosynthesis in N-fixing nodules cannot account for the high levels of S-assimilation and cysteine biosynthesis.
Previous studies have shown repression of sulfate uptake by an excess of cysteine or other reduced S-compounds (Herschbach and Rennenberg, 1994; Zhao et al., 1999; Vauclare et al., 2002; Hesse et al., 2004). Such feedback repression could account for the significantly lower total 35S-sulfate uptake per plant, sulfate uptake per root system FW, and organ uptake rate in Fix+ compared with uninoculated and Fix− plants (Figures 2, 3, and 9).
In conclusion, our data show that in the L. japonicus-M. loti symbiosis, nodules are not only the main site of plant N-assimilation but are also a major site of S-assimilation and thiol biosynthesis. Furthermore, the data indicate that nodules may be a significant source of assimilated S for other organs; certainly, the presence of Fix+ nodules leads to reprogramming of S-metabolism genes and altered S-metabolism and S-partitioning throughout the plant. The impact of nodules on whole-plant S-economy is dampened in the absence of active SNF in nodules, a situation that forces the plant to switch back to its nonsymbiotic N- and S-assimilation patterns. Future work, employing both plant and rhizobial mutants, will help to clarify the relative contribution of rhizobia and plant cells to nodule sulfate assimilation and the contribution of nodules to the overall plant S-metabolism.
METHODS
Plant Material, Bacterial Strains, Culture Conditions, and Plant Growth
Mesorhizobium loti wild-type strain R7A and the Fix− mutant strains ΔnifA and ΔnifH were kindly provided by Clive Ronson (University of Otago, Otago, New Zealand). Both mutant strains form ineffective small white nodules with no nitrogenase activity (Fotelli et al., 2011). The main difference between the two types of ineffective nodules is that ΔnifH nodules contain well differentiated, although ineffective, bacteroids, in contrast to nodules harboring ΔnifA strain where infected cells are colonized by undifferentiated rhizobia. All strains were grown for 72 h at 28°C on solid yeast mannitol broth supplemented with the appropriate antibiotic (30 μg/mL kanamycin for ΔnifA; 50 μg/mL gentamycin for ΔnifH). For the liquid cultures, rhizobial minimal medium was used (per 1 liter: 0.25 g MgSO4⋅7H2O, 0.1 g CaCl2⋅2H2O, 1 g K2HPO4, 0.53 g NH4Cl, 1 g KH2PO4, 10 μg FeCl3⋅6H2O, 20 μg CoCl2, 20 μg Na2MoO4, 20 μg MnSO4⋅H2O, 20 μg ZnSO4⋅7H2O, 20 μg CuSO4⋅5H2O, 10 g glucose, 20 μg biotin, 20 μg thiamine, and 20 μg vitamin B5).
Lotus japonicus (Gifu B-129) seeds were kindly provided by Jens Stougaard (University of Aarhus, Aarhus, Denmark). The seeds were scarified for 5 min with H2SO4, sterilized for 20 min in 2% (v/v) NaOCl-0.02% (v/v) Tween 20, pregerminated at 18°C in the dark for 72 h, and spot-inoculated with a suspension culture of the respective M. loti strain at an OD600 of 0.1. For each independent experiment, sets of plants were grown at an 18-h-day/6-h-night cycle, a 22°C day/18°C night temperature regime, and 70% relative air humidity (Handberg and Stougaard, 1992). Uninoculated plants were watered with nitrate containing Hoagland solution (Hoagland and Arnon, 1950), plants harboring effective nodules were watered with N-free Hoagland solution, and plants harboring ineffective nodules were watered alternately with N-free and nitrate-containing Hoagland solution.
Thiol Analysis
A total of 30 to 100 mg homogenized frozen plant material was suspended in 0.75 mL 0.1 n HCl containing 50 mg of insoluble polyvinylpolypyrrolidone. The samples were centrifuged for 30 min at 14,000g and 4°C. For reduction of thiols, 120 μL of the supernatant was added to 180 μL 200 mM CHES buffer, pH 9.3, and 30 μL 15 mM DTT. After incubation at room temperature for 60 min, thiols were derivatized in the dark after adding 20 μL 30 mM monobromobimane for 15 min. Bimane derivatives were stabilized with 250 μL 10% acetic acid and separated with a HPLC system (Beckman Gold System), creating a gradient with increasing methanol on a C-18 column (ODS-Hypersil 250 × 4.6 mm i.d., 5-μm particle size; Bischoff Chromotography). Thiol derivatives were determined by fluorescence detection with a Shimadzu RF-535-Fluorescence HPLC detector (Shimadzu Europe) and quantified by external standards (Schupp and Rennenberg, 1988).
Sulfate Analysis
Fifty milligrams of homogenized frozen plant material was suspended in 1 mL of distilled water containing 100 mg of insoluble polyvinylpolypyrrolidone to remove phenolic compounds. After shaking for 1 h at 4°C, samples were boiled for 15 min and centrifuged for 15 min at 14,000g and 4°C. The clear supernatant was used for sulfate analysis by anion exchange chromatography. Anions were separated on an IonPac column (AS9-SC, 250 × 4 mm; Dionex) eluted with a mixture of 2 mM Na2CO3 and 0.75 mM NaHCO3 at a flow rate of 0.5 mL min−1. Sulfate was detected by a conductivity detector module (CDM; Dionex).
APR Activity and Protein Determination
APS reductase activity was measured as production of 35S-sulfite, assayed as acid volatile radioactivity, formed in the presence of 75 μM AP35S and 4 mM dithioerythritol, according to Brunold and Suter (1990).
Radioactivity in proteins was determined in 30 mg of frozen samples. Powdered plant samples were suspended in 1 mL of extraction buffer (125 mM Tris-HCl, pH 6.8, and 6% SDS) by vigorous vortexing. Protein was precipitated in 10% trichloroacetic acid for 30 min on ice, washed, and redissolved in 0.2 M NaOH. The amount of 35S incorporated into protein was determined in the redissolved protein by liquid scintillation counting after adding 5 mL of liquid scintillation fluid (HiSafe 2; Perkin-Elmer).
35S-Sulfate Uptake of Intact L. japonicus Plants
Root systems of intact 6-week-old L. japonicus plants were transferred into 2 mL N-free Hoagland solution adjusted to 0.1 mM SO42– containing 10 μCi (3.7 × 105 Bq) of carrier-free 35SO42− (Hartmann Analytic). Incubation was performed at room temperature (20 to 25°C) and 180 ± 30 µE (Osram L 58W/12 Lumilux de luxe Daylight and Osram L 58W/77 Fluora). After 1 h, sulfate uptake was terminated by washing the root system three times with nonradioactive N-free Hoagland solution. Nodules, roots, stems, and leaves were separated, weighed, frozen in liquid N, and stored at −20°C until analysis of 35S.
35S-Sulfate Flux Analyses
35S-flux analyses were conducted according to Scheerer et al. (2010). Nodules or roots were excised from 6-week-old L. japonicus plants. The excised organs were then transferred into 10 mL N-free Hoagland solution adjusted to 0.1 mM SO42– and supplemented with 3% sucrose containing 150 μCi (5.5 × 106 Bq) of carrier-free 35SO42− (Hartmann Analytic). After 4 h of incubation at room temperature, sulfate uptake was stopped by washing roots or nodules three times with nonradioactive N-free Hoagland solution. Samples were frozen and powdered in liquid N and stored at –20°C until analysis of the 35S-content and the 35S-flux into sulfate, thiols, and protein.
35S Analyses
The 35S-content of the samples was determined by measurement of radioactivity in 20 mg of each sample, powdered under liquid N. After solubilization with a tissue solubilizer (Soluene 350; Perkin-Elmer), the dissolved samples were bleached with 200 μL of H2O2 (30%) overnight. After adding 5 mL of scintillation fluid (HiSafe 2; Perkin-Elmer), radioactivity was determined by scintillation counting (Wallac System 1409) (Herschbach and Rennenberg, 1996).
35S-Metabolite Analyses
Sulfate was quantified by anion exchange chromatography and specific radioactivity of the sulfate was determined by liquid scintillation counting of 1.2-mL fractions of the eluate after anion exchange chromatography (see above). The amount of 35S in thiols was determined in the eluate after HPLC analysis (see above). A total of 4 mL of scintillation fluid (HiSafe 3; Perkin-Elmer) was added to each 1-mL fraction, and radioactivity was determined by liquid scintillation counting. Radioactivity in thiols of the different fractions was correlated to the respective peaks after fluorescent detection (Scheerer et al., 2010).
Transcript Analysis Using qRT-PCR
Total RNA was isolated from free-living bacteria harvested during exponential and stationary growth, as well as nodules and nonsymbiotic organs of L. japonicus plants either uninoculated or inoculated with M. loti wild-type, ΔnifA, or ΔnifH stains according to Brusslan and Tobin (1992) and quantified by spectrophotometry and agarose gel electrophoresis. RNA was treated with DNase I (Promega) at 37°C for 45 min to eliminate contaminating genomic DNA. First-strand bacterial or plant cDNA was reverse transcribed using Super-Script II (Invitrogen) and random hexanucleotides or oligo(dT)12–18mer (Invitrogen) primers, respectively. qRT-PCR was performed on the Stratagene MX3005P using iTaq fast SYBR Green Supermix with ROX (Bio-Rad Laboratories) and gene-specific primers. Detailed information for all genes in this study, including names, gene symbols, accession numbers, predicted topology, and gene-specific primers, is presented in Supplemental Data Set 4. PCR cycling started at 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min. The primer specificity and the formation of primer dimers were monitored by dissociation curves and agarose gel electrophoresis on a 4% (w/v) gel. The expression levels of the M. loti polyribonucleotide nucleotidyltransferase gene (mlr5562) and a L. japonicus ubiquitin gene were used as internal standards for the bacterial and plant cDNAs, respectively. Relative transcript levels of the gene of interest (X) were calculated as a ratio to the internal standard gene transcripts (H), as (1+E)–ΔCt, where ΔCt was calculated as (CtX–CtH). PCR efficiency (E) for each amplicon was calculated by employing the linear regression method on the log (fluorescence) per cycle number data, using LinRegPCR software (Ramakers et al., 2003).
Statistical Analysis
All statistical analyses were performed using SigmaStat 3.5. Planned comparisons were performed by applying pairwise t tests. When the normality test failed, the Mann-Whitney Rank Sum test was used. Statistical significance was postulated for P ≤ 0.05. The statistical analysis was performed on the values from independent plant sets used as biological replicates within the same organ, while wild-type nodules were also compared with the inoculated roots. For sulfate and thiol content, APR activity, and 35S-flux analysis, two independent experiments were performed at different time periods under controlled conditions, including controls and all genotypes, comprising independent plant sets used as biological replicates. Transcript analysis was performed on three independently grown plant sets and bacterial cultures used as biological replicates.
Accession Numbers
The accession numbers of the sequences used in this article are provided in Supplemental Data Set 4.
Supplemental Data
Supplemental Data Set 1. Relative transcript levels of genes involved in S-metabolism.
Supplemental Data Set 2. Fold changes in transcript levels of genes involved in S-metabolism.
Supplemental Data Set 3. Statistical analysis of results presented in Supplemental Data Set 2.
Supplemental Data Set 4. List of genes involved in S-metabolism.
Supplementary Material
Acknowledgments
We thank Ursula Scheerer for technical assistance provided during experiments employing 35S and Susanne Mult for preparing 35S-APS. This work was funded by a joint DAAD and Greek State Scholarship Foundation project.
AUTHOR CONTRIBUTIONS
E.F., H.R., C.H., and C.K. designed the research. C.K. performed the experiments and analyzed the data. P.K. and G.K. performed the experiments for the bacterial gene expression analysis. C.K., P.K., and G.K. carried out bioinformatics analyses. C.K., E.F., C.H., H.R., and M.K.U. contributed to experimental design and interpretation. C.K. and E.F. wrote the article. C.H., H.R., and M.K.U. critically reviewed the article and contributed to the preparation of the final version.
Glossary
- SNF
symbiotic nitrogen fixation
- ATPS
ATP sulfurylase
- APS
adenosine 5′-phosphosulfate
- APR
APS reductase
- OAS
O-acetyl-l-serine
- hGSH
homoglutathione
- FW
fresh weight
- γEC
γ-glutamylcysteine
- dpi
days postinoculation
References
- Abbas B.A., Vineetha K.E., Prasad C.K., Vij N., Hassani R., Randhawa G.S. (2002). Symbiotic characteristics of cysteine and methionine auxotrophs of Sinorhizobium meliloti. Indian J. Exp. Biol. 40: 1121–1130. [PubMed] [Google Scholar]
- Ampe F., Kiss E., Sabourdy F., Batut J. (2003). Transcriptome analysis of Sinorhizobium meliloti during symbiosis. Genome Biol. 4: R15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Appleby C.A. (1984). Leghemoglobin and Rhizobium respiration. Annu. Rev. Plant Physiol. Plant Mol. Biol. 35: 443–478. [Google Scholar]
- Barra L., Fontenelle C., Ermel G., Trautwetter A., Walker G.C., Blanco C. (2006). Interrelations between glycine betaine catabolism and methionine biosynthesis in Sinorhizobium meliloti strain 102F34. J. Bacteriol. 188: 7195–7204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becana M., Matamoros M.A., Udvardi M., Dalton D.A. (2010). Recent insights into antioxidant defenses of legume root nodules. New Phytol. 188: 960–976. [DOI] [PubMed] [Google Scholar]
- Becker A., et al. (2004). Global changes in gene expression in Sinorhizobium meliloti 1021 under microoxic and symbiotic conditions. Mol. Plant Microbe Interact. 17: 292–303. [DOI] [PubMed] [Google Scholar]
- Benedito V.A., et al. (2008). A gene expression atlas of the model legume Medicago truncatula. Plant J. 55: 504–513. [DOI] [PubMed] [Google Scholar]
- Bergersen, F.J. (1982). Root Nodules of Legumes: Structure and Functions. (Letchworth, UK: Research Studies Press). [Google Scholar]
- Bianucci E., Tordable Mdel.C., Fabra A., Castro S. (2008). Importance of glutathione in the nodulation process of peanut (Arachis hypogaea). Physiol. Plant. 134: 342–347. [DOI] [PubMed] [Google Scholar]
- Bick J.-A., Setterdahl A.T., Knaff D.B., Chen Y., Pitcher L.H., Zilinskas B.A., Leustek T. (2001). Regulation of the plant-type 5′-adenylyl sulfate reductase by oxidative stress. Biochemistry 40: 9040–9048. [DOI] [PubMed] [Google Scholar]
- Blaschke L., Schneider A., Herschbach C., Rennenberg H. (1996). Reduced sulphur allocation from three-year-old needles of Norway spruce (Picea abies [Karst] L.). J. Exp. Bot. 47: 1025–1032. [Google Scholar]
- Brunold C. (1993). Regulatory interactions between sulphate and nitrate assimilation. In Sulphur Nutrition and Sulphur Assimilation in Higher Plants, De Kok L.J., Stulen I., Rennenberg H., Brunold C., Rauser W.E., eds (The Hague, The Netherlands: SPB Academic Publishing; ), pp. 61–75. [Google Scholar]
- Brunold C., Suter M. (1990). Adenosine 5′-phosphosulphate sulfotransferase. In Methods in Plant Bichemistry, P. Lea, ed (London: Academic Press; ), pp. 339–343. [Google Scholar]
- Brunold C., Suter M. (1984). Regulation of sulfate assimilation by nitrogen nutrition in the duckweed Lemna minor L. Plant Physiol. 76: 579–583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brusslan J.A., Tobin E.M. (1992). Light-independent developmental regulation of cab gene expression in Arabidopsis thaliana seedlings. Proc. Natl. Acad. Sci. USA 89: 7791–7795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang C., Damiani I., Puppo A., Frendo P. (2009). Redox changes during the legume-rhizobium symbiosis. Mol. Plant 2: 370–377. [DOI] [PubMed] [Google Scholar]
- Clarkson D.T., Saker L.R., Purves J.V. (1989). Depression of nitrate and ammonium transport in barley plants with diminished sulphate status: evidence of co-regulation of nitrogen and sulphate intake. J. Exp. Bot. 40: 953–963. [Google Scholar]
- Colebatch G., Desbrosses G., Ott T., Krusell L., Montanari O., Kloska S., Kopka J., Udvardi M.K. (2004). Global changes in transcription orchestrate metabolic differentiation during symbiotic nitrogen fixation in Lotus japonicus. Plant J. 39: 487–512. [DOI] [PubMed] [Google Scholar]
- Colebatch G., Kloska S., Trevaskis B., Freund S., Altmann T., Udvardi M.K. (2002). Novel aspects of symbiotic nitrogen fixation uncovered by transcript profiling with cDNA arrays. Mol. Plant Microbe Interact. 15: 411–420. [DOI] [PubMed] [Google Scholar]
- Dalton D., Langeberg L., Treneman N.C. (1993). Correlations between the ascorbate-glutathione pathway and effectiveness in legume root nodules. Physiol. Plant. 87: 365–370. [Google Scholar]
- Davidian J.-C., Kopriva S. (2010). Regulation of sulfate uptake and assimilation--the same or not the same? Mol. Plant 3: 314–325. [DOI] [PubMed] [Google Scholar]
- De Visser R., Lambers H. (1983). Growth and the efficiency of root respiration of Pisum sativum as dependent on the source of nitrogen. Physiol. Plant. 58: 533–543. [Google Scholar]
- Dos Santos P.C., Dean D.R., Hu Y., Ribbe M.W. (2004). Formation and insertion of the nitrogenase iron-molybdenum cofactor. Chem. Rev. 104: 1159–1173. [DOI] [PubMed] [Google Scholar]
- Fedorova M., Van De Mortel J., Matsumoto P.A., Cho J., Town C.D., Vandenbosch K.A., Gantt J.S., Vance C.P. (2002). Genome-wide identification of nodule-specific transcripts in the model legume Medicago truncatula. Plant Physiol. 130: 519–537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fotelli M.N., Tsikou D., Kolliopoulou A., Aivalakis G., Katinakis P., Udvardi M.K., Rennenberg H., Flemetakis E. (2011). Nodulation enhances dark CO₂ fixation and recycling in the model legume Lotus japonicus. J. Exp. Bot. 62: 2959–2971. [DOI] [PubMed] [Google Scholar]
- Frendo P., Matamoros M.A., Alloing G., Becana M. (2013). Thiol-based redox signaling in the nitrogen-fixing symbiosis. Front. Plant Sci. 4: 376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaude N., Tippmann H., Flemetakis E., Katinakis P., Udvardi M., Dörmann P. (2004). The galactolipid digalactosyldiacylglycerol accumulates in the peribacteroid membrane of nitrogen-fixing nodules of soybean and Lotus. J. Biol. Chem. 279: 34624–34630. [DOI] [PubMed] [Google Scholar]
- Gigolashvili T., Kopriva S. (2014). Transporters in plant sulfur metabolism. Front. Plant Sci. 5: 442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groten K., Dutilleul C., van Heerden P.D.R., Vanacker H., Bernard S., Finkemeier I., Dietz K.-J., Foyer C.H. (2006). Redox regulation of peroxiredoxin and proteinases by ascorbate and thiols during pea root nodule senescence. FEBS Lett. 580: 1269–1276. [DOI] [PubMed] [Google Scholar]
- Groten K., Vanacker H., Dutilleul C., Bastian F., Bernard S., Carzaniga R., Foyer C.H. (2005). The roles of redox processes in pea nodule development and senescence. Plant Cell Environ. 28: 1293–1304. [Google Scholar]
- Haas F.H., Heeg C., Queiroz R., Bauer A., Wirtz M., Hell R. (2008). Mitochondrial serine acetyltransferase functions as a pacemaker of cysteine synthesis in plant cells. Plant Physiol. 148: 1055–1067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Handberg, K., and Stougaard, J. (1992). Lotus japonicus, an autogamous, diploid legume species for classical and molecular genetics. Plant J. 2: 487–496.
- Harrison, J., Jamet, A., Muglia, C.I., Van de Sype, G., Aguilar, O.M., Puppo, A., and Frendo, P. (2005). Glutathione plays a fundamental role in growth and symbiotic capacity of Sinorhizobium meliloti. J. Bacteriol. 187: 168–174. [DOI] [PMC free article] [PubMed]
- Hassinen V.H., Tervahauta A.I., Schat H., Kärenlampi S.O. (2011). Plant metallothioneins--metal chelators with ROS scavenging activity? Plant Biol (Stuttg) 13: 225–232. [DOI] [PubMed] [Google Scholar]
- Heeg C., Kruse C., Jost R., Gutensohn M., Ruppert T., Wirtz M., Hell R. (2008). Analysis of the Arabidopsis O-acetylserine(thiol)lyase gene family demonstrates compartment-specific differences in the regulation of cysteine synthesis. Plant Cell 20: 168–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herschbach, C., Gessler, A., and Rennenberg, H. (2012). Long-distance transport and plant internal cycling of N- and S-compounds. In Progress in Botany, U. Lüttge, W. Beyschlag, B. Büdel, and D. Francis, eds (Berlin, Heidelberg: Springer Berlin Heidelberg), pp. 161–188. [Google Scholar]
- Herschbach C., Rennenberg H. (1994). Influence of glutathione (GSH) on net uptake of sulfate and sulfate transport in tobacco plants. J. Exp. Bot. 45: 1069–1076. [Google Scholar]
- Herschbach C., Rennenberg H. (1995). Long-distance transport of 35S-sulphur in 3-year-old beech trees (Fagus sylvatica). Physiol. Plant. 95: 379–386. [Google Scholar]
- Herschbach C., Rennenberg H. (2001a). Significance of phloem-translocated organic sulfur compounds for the regulation of sulfur nutrition. Prog. Bot. 62: 177–193. [Google Scholar]
- Herschbach C., Rennenberg H. (1996). Storage and remobilization of sulphur in beech trees (Fagus sylvatica). Physiol. Plant. 98: 125–132. [Google Scholar]
- Herschbach C., Rennenberg H. (2001b). Sulfur nutrition of deciduous trees. Naturwissenschaften 88: 25–36. [DOI] [PubMed] [Google Scholar]
- Hesse H., Nikiforova V., Gakière B., Hoefgen R. (2004). Molecular analysis and control of cysteine biosynthesis: integration of nitrogen and sulphur metabolism. J. Exp. Bot. 55: 1283–1292. [DOI] [PubMed] [Google Scholar]
- Ho C.-L., Saito K. (2001). Molecular biology of the plastidic phosphorylated serine biosynthetic pathway in Arabidopsis thaliana. Amino Acids 20: 243–259. [DOI] [PubMed] [Google Scholar]
- Hoagland, D.R., and Arnon, D.I. (1950). The water-culture method for growing plants without soil. Circular. (California Agricultural Experiment Station 347).
- Höfgen R., Kreft O., Willmitzer L., Hesse H. (2001). Manipulation of thiol contents in plants. Amino Acids 20: 291–299. [DOI] [PubMed] [Google Scholar]
- Johnson D.C., Dean D.R., Smith A.D., Johnson M.K. (2005). Structure, function, and formation of biological iron-sulfur clusters. Annu. Rev. Biochem. 74: 247–281. [DOI] [PubMed] [Google Scholar]
- Kopriva S. (2006). Regulation of sulfate assimilation in Arabidopsis and beyond. Ann. Bot. (Lond.) 97: 479–495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kopriva S., Koprivova A. (2004). Plant adenosine 5′-phosphosulphate reductase: the past, the present, and the future. J. Exp. Bot. 55: 1775–1783. [DOI] [PubMed] [Google Scholar]
- Kopriva S., Rennenberg H. (2004). Control of sulphate assimilation and glutathione synthesis: interaction with N and C metabolism. J. Exp. Bot. 55: 1831–1842. [DOI] [PubMed] [Google Scholar]
- Koprivova A., North K.A., Kopriva S. (2008). Complex signaling network in regulation of adenosine 5′-phosphosulfate reductase by salt stress in Arabidopsis roots. Plant Physiol. 146: 1408–1420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koprivova A., Suter M., den Camp R.O., Brunold C., Kopriva S. (2000). Regulation of sulfate assimilation by nitrogen in Arabidopsis. Plant Physiol. 122: 737–746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kouchi H., et al. (2004). Large-scale analysis of gene expression profiles during early stages of root nodule formation in a model legume, Lotus japonicus. DNA Res. 11: 263–274. [DOI] [PubMed] [Google Scholar]
- Kredich N.M. (1996). Biosynthesis of cysteine. In Escherichia coli and Salmonella: Cellular and Molecular Biology, Neidhardt F., ed (Washington, D.C.: ASM Press; ), pp. 514–527. [Google Scholar]
- Kredich N.M. (1992). The molecular basis for positive regulation of cys promoters in Salmonella typhimurium and Escherichia coli. Mol. Microbiol. 6: 2747–2753. [DOI] [PubMed] [Google Scholar]
- Krueger S., Niehl A., Lopez Martin M.C., Steinhauser D., Donath A., Hildebrandt T., Romero L.C., Hoefgen R., Gotor C., Hesse H. (2009). Analysis of cytosolic and plastidic serine acetyltransferase mutants and subcellular metabolite distributions suggests interplay of the cellular compartments for cysteine biosynthesis in Arabidopsis. Plant Cell Environ. 32: 349–367. [DOI] [PubMed] [Google Scholar]
- Krusell L., et al. (2005). The sulfate transporter SST1 is crucial for symbiotic nitrogen fixation in Lotus japonicus root nodules. Plant Cell 17: 1625–1636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Küster H., et al. (2004). Construction and validation of cDNA-based Mt6k-RIT macro- and microarrays to explore root endosymbioses in the model legume Medicago truncatula. J. Biotechnol. 108: 95–113. [DOI] [PubMed] [Google Scholar]
- Kuzma M.M., Hunt S., Layzell D.B. (1993). Role of oxygen in the limitation and inhibition of nitrogenase activity and respiration rate in individual soybean nodules. Plant Physiol. 101: 161–169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laudenbach D.E., Grossman A.R. (1991). Characterization and mutagenesis of sulfur-regulated genes in a cyanobacterium: evidence for function in sulfate transport. J. Bacteriol. 173: 2739–2750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leustek T., Saito K. (1999). Sulfate transport and assimilation in plants. Plant Physiol. 120: 637–644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loudet O., Saliba-Colombani V., Camilleri C., Calenge F., Gaudon V., Koprivova A., North K.A., Kopriva S., Daniel-Vedele F. (2007). Natural variation for sulfate content in Arabidopsis thaliana is highly controlled by APR2. Nat. Genet. 39: 896–900. [DOI] [PubMed] [Google Scholar]
- Lynch A.S., Tyrrell R., Smerdon S.J., Briggs G.S., Wilkinson A.J. (1994). Characterization of the CysB protein of Klebsiella aerogenes: direct evidence that N-acetylserine rather than O-acetylserine serves as the inducer of the cysteine regulon. Biochem. J. 299: 129–136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matamoros M.A., Clemente M.R., Sato S., Asamizu E., Tabata S., Ramos J., Moran J.F., Stiller J., Gresshoff P.M., Becana M. (2003). Molecular analysis of the pathway for the synthesis of thiol tripeptides in the model legume Lotus japonicus. Mol. Plant Microbe Interact. 16: 1039–1046. [DOI] [PubMed] [Google Scholar]
- Matamoros M.A., Moran J.F., Iturbe-Ormaetxe I., Rubio M.C., Becana M. (1999). Glutathione and homoglutathione synthesis in legume root nodules. Plant Physiol. 121: 879–888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mordukhova E.A., Pan J.-G. (2013). Evolved cobalamin-independent methionine synthase (MetE) improves the acetate and thermal tolerance of Escherichia coli. Appl. Environ. Microbiol. 79: 7905–7915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- El Msehli S., Lambert A., Baldacci-Cresp F., Hopkins J., Boncompagni E., Smiti S.A., Hérouart D., Frendo P. (2011). Crucial role of (homo)glutathione in nitrogen fixation in Medicago truncatula nodules. New Phytol. 192: 496–506. [DOI] [PubMed] [Google Scholar]
- Muglia C., Comai G., Spegazzini E., Riccillo P.M., Aguilar O.M. (2008). Glutathione produced by Rhizobium tropici is important to prevent early senescence in common bean nodules. FEMS Microbiol. Lett. 286: 191–198. [DOI] [PubMed] [Google Scholar]
- Noctor G., Mhamdi A., Chaouch S., Han Y., Neukermans J., Marquez-Garcia B., Queval G., Foyer C.H. (2012). Glutathione in plants: an integrated overview. Plant Cell Environ. 35: 454–484. [DOI] [PubMed] [Google Scholar]
- Oldroyd G.E.D., Harrison M.J., Udvardi M. (2005). Peace talks and trade deals. Keys to long-term harmony in legume-microbe symbioses. Plant Physiol. 137: 1205–1210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ott T., van Dongen J.T., Günther C., Krusell L., Desbrosses G., Vigeolas H., Bock V., Czechowski T., Geigenberger P., Udvardi M.K. (2005). Symbiotic leghemoglobins are crucial for nitrogen fixation in legume root nodules but not for general plant growth and development. Curr. Biol. 15: 531–535. [DOI] [PubMed] [Google Scholar]
- Piłsyk S., Paszewski A. (2009). Sulfate permeasesphylogenetic diversity of sulfate transport. Acta Biochim. Pol. 56: 375–384. [PubMed] [Google Scholar]
- Prell J., Poole P. (2006). Metabolic changes of rhizobia in legume nodules. Trends Microbiol. 14: 161–168. [DOI] [PubMed] [Google Scholar]
- Ramakers C., Ruijter J.M., Deprez R.H., Moorman A.F.M. (2003). Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 339: 62–66. [DOI] [PubMed] [Google Scholar]
- Rees D.C., Howard J.B. (2000). Nitrogenase: standing at the crossroads. Curr. Opin. Chem. Biol. 4: 559–566. [DOI] [PubMed] [Google Scholar]
- Rennenberg H., Herschbach C. (2014). A detailed view on sulphur metabolism at the cellular and whole-plant level illustrates challenges in metabolite flux analyses. J. Exp. Bot. 65: 5711–5724. [DOI] [PubMed] [Google Scholar]
- Rennenberg H., Schupp R., Glavac V., Jochheim H. (1994a). Xylem sap composition of beech (Fagus sylvatica L.) trees: seasonal changes in the axial distribution of sulfur compounds. Tree Physiol. 14: 541–548. [DOI] [PubMed] [Google Scholar]
- Rennenberg H., Schupp R., Schneider A. (1994b). Thiol composition of a xylem-tapping mistletoe and the xylem sap of its hosts. Phytochemistry 37: 975–977. [Google Scholar]
- Scheerer U., Haensch R., Mendel R.R., Kopriva S., Rennenberg H., Herschbach C. (2010). Sulphur flux through the sulphate assimilation pathway is differently controlled by adenosine 5′-phosphosulphate reductase under stress and in transgenic poplar plants overexpressing γ-ECS, SO, or APR. J. Exp. Bot. 61: 609–622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scheffel F., Demmer U., Warkentin E., Hülsmann A., Schneider E., Ermler U. (2005). Structure of the ATPase subunit CysA of the putative sulfate ATP-binding cassette (ABC) transporter from Alicyclobacillus acidocaldarius. FEBS Lett. 579: 2953–2958. [DOI] [PubMed] [Google Scholar]
- Schmidt A. (1972). On the mechanism of photosynthetic sulfate reduction. An APS-sulfotransferase from Chlorella. Arch. Mikrobiol. 84: 77–86. [DOI] [PubMed] [Google Scholar]
- Schmidt A., Jager K. (1992). Open questions about sulfur metabolism in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 43: 325–349. [Google Scholar]
- Schneider A., Kreuzwieser J., Schupp R., Sauter J.J., Rennenberg H. (1994a). Thiol and amino acid composition of the xylem sap of poplar trees (Populus×canadensis “robusta”). Can. J. Bot. 72: 347–351. [Google Scholar]
- Schneider A., Schatten T., Rennenberg H. (1994b). Exchange between phloem and xylem during long distance transport of glutathione in spruce trees (Picea abies [Karst.] L.). J. Exp. Bot. 45: 457–462. [DOI] [PubMed] [Google Scholar]
- Schupp R., Glavac V., Rennenberg H. (1991). Thiol composition of xylem sap of beech trees. Phytochemistry 30: 1415–1418. [Google Scholar]
- Schupp R., Rennenberg H. (1988). Diurnal changes in the glutathione concentration of spruce needles (Picea abies L.). Plant Sci. 57: 113–117. [Google Scholar]
- Smith I.K. (1980). Regulation of sulfate assimilation in tobacco cells. Effect of nitrogen and sulfur nutrition on sulfate permease and O-acetylserine sulfhydrylase. Plant Physiol. 66: 877–883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sullivan J.T., et al. (2002). Comparative sequence analysis of the symbiosis island of Mesorhizobium loti strain R7A. J. Bacteriol. 184: 3086–3095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suter M., von Ballmoos P., Kopriva S., den Camp R.O., Schaller J., Kuhlemeier C., Schürmann P., Brunold C. (2000). Adenosine 5′-phosphosulfate sulfotransferase and adenosine 5′-phosphosulfate reductase are identical enzymes. J. Biol. Chem. 275: 930–936. [DOI] [PubMed] [Google Scholar]
- Takahashi H., Kopriva S., Giordano M., Saito K., Hell R. (2011). Sulfur assimilation in photosynthetic organisms: molecular functions and regulations of transporters and assimilatory enzymes. Annu. Rev. Plant Biol. 62: 157–184. [DOI] [PubMed] [Google Scholar]
- Uchiumi T., et al. (2004). Expression islands clustered on the symbiosis island of the Mesorhizobium loti genome. J. Bacteriol. 186: 2439–2448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Udvardi M., Poole P.S. (2013). Transport and metabolism in legume-rhizobia symbioses. Annu. Rev. Plant Biol. 64: 781–805. [DOI] [PubMed] [Google Scholar]
- Udvardi M.K., Day D.A. (1997). Metabolite transport across symbiotic membranes of legume nodules. Annu. Rev. Plant Physiol. Plant Mol. Biol. 48: 493–523. [DOI] [PubMed] [Google Scholar]
- Vance C.P. (2001). Symbiotic nitrogen fixation and phosphorus acquisition. Plant nutrition in a world of declining renewable resources. Plant Physiol. 127: 390–397. [PMC free article] [PubMed] [Google Scholar]
- Van der Weerden N.L., Anderson M.A. (2013). Plant defensins: Common fold, multiple functions. Fungal Biol. Rev. 26: 121–131. [Google Scholar]
- Vauclare P., Kopriva S., Fell D., Suter M., Sticher L., von Ballmoos P., Krähenbühl U., den Camp R.O., Brunold C. (2002). Flux control of sulphate assimilation in Arabidopsis thaliana: adenosine 5′-phosphosulphate reductase is more susceptible than ATP sulphurylase to negative control by thiols. Plant J. 31: 729–740. [DOI] [PubMed] [Google Scholar]
- Watanabe M., Mochida K., Kato T., Tabata S., Yoshimoto N., Noji M., Saito K. (2008). Comparative genomics and reverse genetics analysis reveal indispensable functions of the serine acetyltransferase gene family in Arabidopsis. Plant Cell 20: 2484–2496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wienkoop S., Saalbach G. (2003). Proteome analysis. Novel proteins identified at the peribacteroid membrane from Lotus japonicus root nodules. Plant Physiol. 131: 1080–1090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao F.J., Hawkesford M.J., McGrath S.P. (1999). Sulphur assimilation and effects on yield and quality of wheat. J. Cereal Sci. 30: 1–17. [Google Scholar]
- Zheng L., White R.H., Cash V.L., Jack R.F., Dean D.R. (1993). Cysteine desulfurase activity indicates a role for NIFS in metallocluster biosynthesis. Proc. Natl. Acad. Sci. USA 90: 2754–2758. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.