Abstract
Background:
The clinical and biological effects of metabolic alterations in cancer are not fully understood.
Methods:
In high-grade serous ovarian cancer (HGSOC) samples (n = 101), over 170 metabolites were profiled and compared with normal ovarian tissues (n = 15). To determine NAT8L gene expression across different cancer types, we analyzed the RNA expression of cancer types using RNASeqV2 data available from the open access The Cancer Genome Atlas (TCGA) website (http://www.cbioportal.org/public-portal/). Using NAT8L siRNA, molecular techniques and histological analysis, we determined cancer cell viability, proliferation, apoptosis, and tumor growth in in vitro and in vivo (n = 6–10 mice/group) settings. Data were analyzed with the Student’s t test and Kaplan-Meier analysis. Statistical tests were two-sided.
Results:
Patients with high levels of tumoral NAA and its biosynthetic enzyme, aspartate N-acetyltransferase (NAT8L), had worse overall survival than patients with low levels of NAA and NAT8L. The overall survival duration of patients with higher-than-median NAA levels (3.6 years) was lower than that of patients with lower-than-median NAA levels (5.1 years, P = .03). High NAT8L gene expression in other cancers (melanoma, renal cell, breast, colon, and uterine cancers) was associated with worse overall survival. NAT8L silencing reduced cancer cell viability (HEYA8: control siRNA 90.61%±2.53, NAT8L siRNA 39.43%±3.00, P < .001; A2780: control siRNA 90.59%±2.53, NAT8L siRNA 7.44%±1.71, P < .001) and proliferation (HEYA8: control siRNA 74.83%±0.92, NAT8L siRNA 55.70%±1.54, P < .001; A2780: control siRNA 50.17%±4.13, NAT8L siRNA 26.52%±3.70, P < .001), which was rescued by addition of NAA. In orthotopic mouse models (ovarian cancer and melanoma), NAT8L silencing reduced tumor growth statistically significantly (A2780: control siRNA 0.52 g±0.15, NAT8L siRNA 0.08 g±0.17, P < .001; HEYA8: control siRNA 0.79 g±0.42, NAT8L siRNA 0.24 g±0.18, P = .008, A375-SM: control siRNA 0.55 g±0.22, NAT8L siRNA 0.21 g±0.17g, P = .001). NAT8L silencing downregulated the anti-apoptotic pathway, which was mediated through FOXM1.
Conclusion:
These findings indicate that the NAA pathway has a prominent role in promoting tumor growth and represents a valuable target for anticancer therapy.
Altered energy metabolism is a hallmark of cancer (1). Proliferating cancer cells have much greater metabolic requirements than nonproliferating differentiated cells (2,3). Moreover, altered cancer metabolism elevates unique metabolic intermediates, which can promote cancer survival and progression (4,5). Furthermore, emerging evidence suggests that proliferating cancer cells exploit alternative metabolic pathways to meet their high demand for energy and to accumulate biomass (6–8).
The fundamental diversity and connectedness of the metabolic pathways make the characterization of the metabolome within a heterogeneous disease process, such as cancer, complex. Although recent research suggests that metabolic changes occur at a molecular level and genetic alterations contribute to this shift (9,10), little is known about how metabolites outside of glycolysis are affected. A better understanding and elucidation of these key metabolic pathways in cancer cell growth may lead to novel targets for cancer therapy and/or the development of reliable biomarkers. Using a global metabolic profile of normal and epithelial ovarian cancer tissues, we set out to identify profoundly altered metabolites in ovarian cancer that promote tumor growth.
Methods
For full descriptions of the following experiments, please see the Supplementary Methods (available online).
Patient Specimens
This study was approved by the Institutional Research Boards of The University of Texas MD Anderson Cancer Center and the University of Iowa. Human high-grade serous ovarian cancer samples were obtained from chemotherapy-naïve patients. Also, normal ovarian samples were obtained from patients undergoing surgery for a benign condition. Clinical data were also collected on these patients.
Metabolic Profiling and Validation
For metabolic profiling, human ovarian tumor (n = 101) and normal ovarian tissue (n = 15) samples were prepared, as described previously (11,12). The untargeted metabolic profiling (Metabolon, Inc., Durham, NC) employed three independent platforms: ultrahigh performance liquid chromatography/tandem mass spectrometry (UHLC/MS/MS2) optimized for basic species, UHLC/MS/MS2 optimized for acidic species, and gas chromatography/mass spectrometry (GC/MS) (13,14). For the metabolic quantification of NAA, human ovarian tumor samples were analyzed using nuclear magnetic resonance (NMR) spectroscopy. To assess N-acetyltransferase (NAT8L) expression in ovarian cancer, total RNA was extracted from 135 HGSOC and 15 normal ovary specimens and subjected to quantitative reverse-transcriptase polymerase chain reaction (qRT-PCR) analysis as previously described (15,16).
Laboratory Measures
Ovarian cancer cell lines (SKOV3, HeyA8, and A2780) were obtained from the M. D. Anderson Characterized Cell Line Core Facility, which supplies authenticated cell lines. The melanoma cell lines (UACC 62, UACC 257, M14, and A375-SM) were a kind gift from Menashe Bar-Eli and Suhendan Ekmekcioglu, U.T. M. D. Anderson Cancer Center (Houston, TX) and maintained in modified MEM medium (17). For in vitro functional assays, all cell lines were transfected with Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA) using gene-specific or nontargeting siRNA (Sigma-Aldrich, St. Louis, MO) according to the manufacturer’s protocol. Apoptosis was assessed in vitro using Annexin V and 7-amino-actinomycin-D staining, and cell proliferation was assessed in vitro using the Click-iT EdU flow cytometry kit (Invitrogen, Carlsbad, CA). For cell-cycle analysis, propidium iodide fluorescence was assessed by flow cytometry. Formalin-fixed, paraffin-embedded samples were assembled as a tissue microarray and subjected to immunohistochemical analyses, as previously described (18).
Orthotopic Mouse Models of Ovarian Cancer and Melanoma
Orthotopic mouse models of ovarian cancer and melanoma were used as described previously (16,19,20). All in vivo experiments were approved by the Institutional Animal Care and Use Committee. Additional details are included in the Supplementary Methods (available online).
Microarray and Pathway Enrichment Analysis
A2780 cells were treated with siNAT8L, and total RNA was extracted 48 hours following transfection using mirVana RNA isolation labeling kit (Ambion). Five hundred nanograms of total RNA were used for labeling and hybridization on a Human HT-12 v4 Beadchip (Illumina) according to the manufacturer’s protocols. After the bead chips were scanned with an Illumina BeadArray Reader (Illumina), the microarray data were normalized using the quantile normalization method in the Linear Models for Microarray Data (LIMMA) package in the R language environment. The expression level of each gene was transformed into a log2 base before further analysis.
The top pathways perturbed by siNAT8L treatment were identified using NetWalker software. Each interaction in the global network of biological relationships was scored based on combined assessment of the network connectivity and the input data. Upstream analysis was subsequently performed using the Ingenuity Pathway Analysis (IPA) software.
Statistical Analyses
For identifying metabolites statistically significantly altered between HGSOC and normal ovarian samples, Welch’s two-sample t tests were used using Array Studio software (OmicSoft) following log transformation of imputed values. Multiple comparisons were accounted for by estimating the false discovery rate (FDR) using q-values (21). For determining the association between aspartate N-acetyltransferase (NAT8L gene expression) and clinical outcomes, RNA expression was analyzed using RNASeqV2 data from The Cancer Genome Atlas (TCGA) (www.cbioportal.org; accessed January 28, 2013). The overall survival durations of patients in each tumor group queried were analyzed using the best cutoff method on provisional databases. For additional cohorts, CEL (Affymetrix U133Plus2) array files from Tothill et al. (22) (GSE9899), from Bogunovic et al. (23) (GSE19234), and from Jonsson et al. (24) (GSE22155) Illumina arrays were also analyzed. Correlation between continuous variables was assessed by computing the Pearson’s product-moment correlation coefficient and the corresponding test of statistical significance of the null hypothesis of zero correlation.
For the survival analysis, patients were grouped according to NAT8L expression. Samples were dichotomized into two groups after determining the best cutoff level for gene expression. The log-rank test was used to determine if there was a statistically significant association between NAT8L expression and overall survival. The Kaplan-Meier method was employed to generate survival curves. The Cox proportional hazards model was used to determine the association of each prognostic factor with disease-specific survival duration, while a multivariable analysis was performed to analyze the contribution of independent risk factors. The proportionality assumption was verified using the regression approach in Grambsh and Thernau (25). For all analyses where dichotomized variables were used, we considered the median values of the continuous measurements. We also investigated the choice of and the optimal cutoff values, defined as the value inducing the most statistically significant association with the survival outcome, as determined by the log-rank test, that led to similar conclusions. Hazard ratios (HRs) including 95% confidence intervals (CIs) were calculated. All testing was two-sided, and a P value of less than .05 was considered statistically significant. All analyses were completed with R or with IBM SPSS.
Results
Metabolic Profile of High-Grade Epithelial Ovarian Cancer
We first examined global metabolic profiling differences between high-grade serous ovarian cancer (HGSOC) samples (n = 101) and noncancerous ovarian tissues (n = 15), and a number of differences were identified. Specifically, 172 metabolites were statistically significantly altered between HGSOC and normal ovarian samples (Supplementary Figure 1A, available online). Of those, 142 metabolites were statistically significantly elevated while 30 were statistically significantly decreased (FDR q-values < 0.05) (Figure 1A). Of the metabolites that were statistically significantly upregulated, the majority involved lipid (53.5%) or amino acid (19.0%) metabolism (Supplementary Figure 1B, available online). In lipid metabolism (Supplementary Table 1, available online), the phosphatidylcholine intermediate cytidine 5’-diphosphocholine had the highest fold-change in levels between normal ovarian tissues and HGSOC (30.38-fold, P < .001). In amino acid metabolism, the neuron-specific metabolite NAA had the highest fold-change in level between normal ovarian tissue and HGSOC (28.38-fold, P < .001) (Supplementary Table 2, available online).
Figure 1.

Relative metabolite levels in high-grade serous ovarian cancer (HGSOC) and metabolic profiling. A) The hierarchical clustering of 172 metabolites in HGSOC and normal ovaries. B) A comparison of the median levels of all metabolites in HGSOC and in normal ovary. C) NAT8L is the rate-limiting enzyme in NAA biosynthesis; NAA is metabolized by aspartoacyclase (ASPA). D) Spearman correlation (r) of relative NAA levels in 101 HGSOC samples and expression levels of genes in the NAA pathway (NAT8L and ASPA). All statistical tests were two-sided.
Of the metabolites that were upregulated in HGSOC, NAA had the highest relative median level in ovarian cancer (Figure 1B). Metabolic profiling did not reveal a consistent pattern of broadly elevated N-acetyl amino acid levels in ovarian cancer (data not shown). This finding suggests that elevated NAA is not a result of global N-acetylation processes, but rathert aspartate is selectively N-acetylated in ovarian cancer.
According to the Kyoto Encyclopedia of Genes and Genomes database, the synthesis and breakdown of NAA require the enzymes NAT8L and aspartoacyclase (ASPA), respectively (Figure 1C). Analysis of NAT8L and ASPA gene expression in the ovarian cancer samples revealed that although NAT8L expression was strongly correlated with tumor NAA levels (r = 0.52, P < .001), ASPA expression was not (r = -0.11, P = .30) (Figure 1D). Furthermore, levels of aspartate, the NAA substrate, were statistically significantly lower in ovarian tumors than in normal ovarian tissues (data not shown). Overall, these findings suggest that the elevated levels of NAA in ovarian cancer are because of its increased biosynthesis.
Clinical Implications of Elevated NAA Levels
Because NAA levels were substantially elevated in ovarian cancer, we next sought to determine whether NAA levels are associated with clinical outcomes. Indeed, the overall survival duration of patients with higher-than-median NAA levels (3.6 years) was lower than that of patients with lower-than-median NAA levels (5.1 years) (Figure 2A). Analysis using a Cox proportional hazards model revealed a statistically significant association between NAA levels dichotomized at the median and disease-specific survival duration (HR = 3.58, 95% CI = 2.51 to 4.66, P = .03).
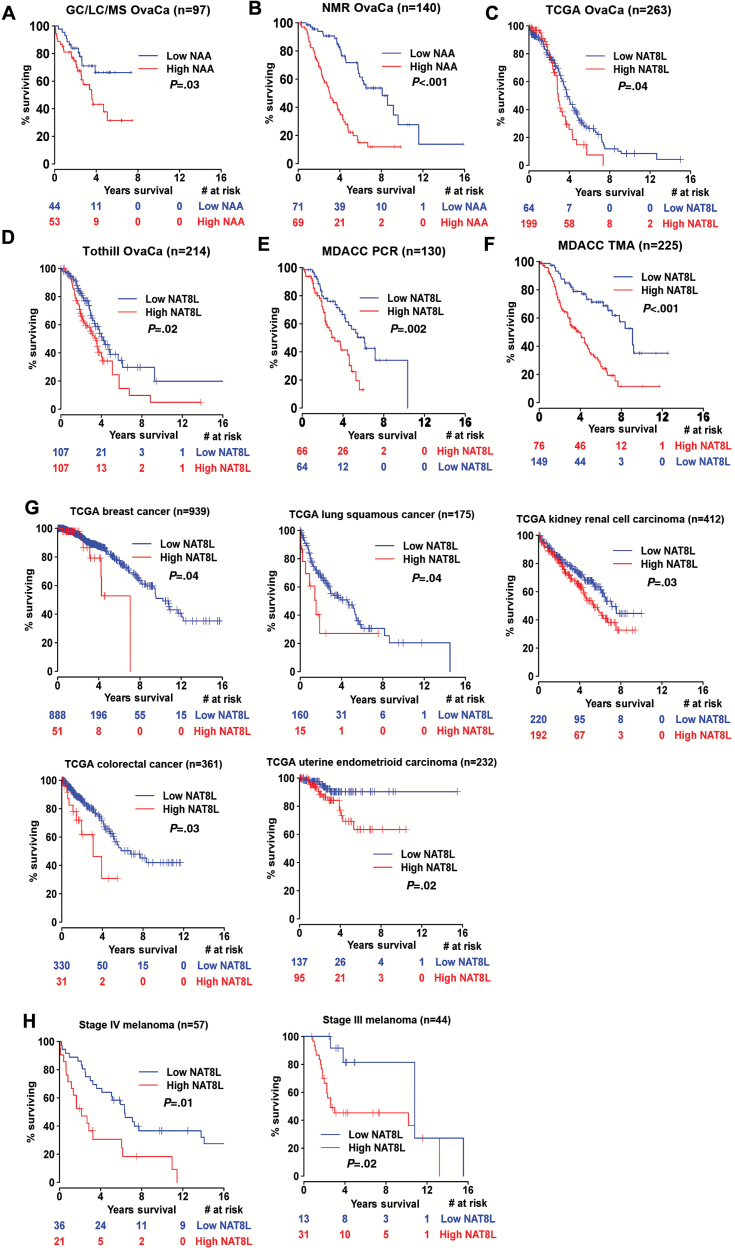
Figure 2.
Kaplan-Meier survival curves for patients according to NAA metabolism. A) Overall survival rates according to NAA levels in high-grade serous ovarian cancer (HGSOC) samples used for metabolic profiling. B) NAA levels in HGSOC samples used for nuclear magnetic resonance (NMR) spectroscopy. C) NAT8L gene expression in HGSOC samples from The Cancer Genome Atlas (TCGA). D) NAT8L gene expression in HGSOC samples from the Tothill database. E) Polymerase chain reaction NAT8L gene expression in HGSOC samples. F) NAT8L protein expression in HGSOC samples used for tissue microarray. G) NAT8L gene expression in breast adenocarcinoma, lung squamous cancer, kidney renal cell carcinoma, uterine endometrioid carcinoma, and colorectal adenocarcinoma from TCGA. H) NAT8L gene expression in melanoma from two different cohorts are shown. All statistical tests were two-sided.
For validation of the NAA findings, nuclear magnetic resonance (NMR) spectroscopy was used to analyze an independent cohort of ovarian cancer samples (n = 145) and normal ovarian tissue samples (n = 13). NMR spectroscopy revealed a distinct peak at 2.005±0.002ppm, consistent with the chemical shift of NAA in ovarian cancer samples (Supplementary Figure 2A, available online). The resonance for NAA was verified with two-dimensional proton correlated spectroscopy (1H COESY). The ratio of the amplitude of the peak at 2.005ppm and the NMR standard was used to determine the concentration of NAA in each sample. Resonances for other n-acetylated amino acids were not present (Chenomx Software, Edmonton, Alberta, Canada). This further suggests that elevated NAA is not a result of global N-acetylation processes; rather, there is a selection for N-acetylation of aspartate to form NAA in ovarian cancer. Overall, NAA levels in the ovarian cancer samples were statistically significantly higher than those in the normal ovary samples. The mean (± 95 % confidence interval) NAA levels were 63 µM (± 12 µM) in ovarian cancer and 20 µM (± 4 µM) in normal ovary samples (Supplementary Figure 2A, available online). Analysis using a Cox proportional hazards model revealed a statistically significant association between NAA levels dichotomized at the median and disease-specific survival duration (HR = 3.67, 95% CI = 2.27 to 5.94, P < .001) (Figure 2B). A multivariable analysis from NMR that included patient age, disease stage, extent of cytoreduction, and NAA values revealed that disease-specific survival duration was statistically significantly associated with NAA level (P < .001) and optimal cytoreduction to no gross residual disease (P = .02) (Supplementary Table 3, available online). The association between continuous, nondichotomized NAA values and disease-specific survival duration using a Cox proportional hazards model was also statistically significant (P < .001).
Next, we asked whether gene expression levels of NAT8L, the rate-limiting enzyme in NAA metabolism, were also related to patient outcome. To address this question, we first used genomic data from The Cancer Genome Atlas (TCGA) and Tothill databases analyzed through cBio portal and Tothill’s data. Analysis of TCGA ovarian cancer samples using the RNA Seq platform (RNASeqv2) revealed that high NAT8L expression (>0.5 standard deviations above the mean expression in diploid samples for NAT8L) was associated with worse overall survival duration (Figure 2C). The median overall survival duration of patients with high NAT8L expression was statistically significantly shorter than that of patients with low NAT8L expression (P = .04). Similarly, in the Tothill cohort, high NAT8L gene expression (>0.5 SDs above the mean) was associated with worse overall survival duration (Figure 2D). The median overall survival duration of patients with high NAT8L expression was statistically significantly lower than that of patients with low NAT8L expression (P = .02).
To validate the findings related to the clinical significance of NAT8L in ovarian cancer, we tested a separate cohort of HGSOC samples using qRT-PCR. Consistent with the findings described above, NAT8L expression levels in HGSOC (n = 135) were much higher than those in normal ovarian tissue (n = 15) (Supplementary Figure 2B, available online). Furthermore, above-the-median NAT8L levels were associated with worse clinical outcomes than below-the-median NAT8L levels were (Figure 2E). Analysis using a Cox proportional hazards model revealed a statistically significant association between NAT8L values dichotomized at the median and disease-specific survival duration (HR = 2.32, 95% CI = 1.37 to 3.95, P = .002). A multivariable analysis that included patient age, dichotomized disease stage, extent of cytoreduction, and NAT8L level dichotomized at the median revealed a statistically significant association between disease-specific survival duration and NAT8L level (P = .002) (Supplementary Table 4, available online).
To determine whether NAT8L protein levels are also elevated, we evaluated NAT8L protein expression in 225 HGSOC samples. NAT8L protein levels in ovarian cancer were statistically significantly higher than those in the normal ovary (n = 10) (Supplementary Figure 2C, available online). Increased NAT8L expression level in ovarian cancer was associated with worse overall survival (Figure 2F). Analysis using a Cox proportional hazards model revealed a statistically significant association between dichotomized NAT8L measurements and disease-specific survival duration (HR = 3.17, 95% CI = 2.03 to 4.94, P < .001). A multivariable analysis that included age, disease stage and grade, extent of cytoreduction, and NAT8L expression levels revealed a statistically significant association between disease-specific survival duration and NAT8L level (HR = 4.89, 95% CI = 2.54 to 9.43, P < .001) (Supplementary Table 5, available online). The association between continuous, nondichotomized NAT8L values and disease-specific survival duration using a Cox proportional hazards model was also statistically significant (P < .001).
To examine whether these findings apply to other cancer types, we analyzed relative NAT8L gene expression across various cancer types using the TCGA datasets. We evaluated the relative gene expression levels and copy number alterations of NAT8L across 22 cancer types (Supplementary Table 6, available online) and found relatively high levels of NAT8L expression in HGSOC, melanoma, glioblastoma multiforme, chromophobe kidney cancer, and low-grade brain glioma, as well as in breast, uterine, and thyroid cancers (Supplementary Figure 2D, available online). Interestingly, HGSOC had the greatest degree of NAT8L copy number alterations (Supplementary Figure 2E, available online). High NAT8L gene expression was associated with worse overall survival duration in patients with invasive breast, renal, colon, squamous cell lung, and uterine cancers (Figure 2G). Furthermore, given that melanoma had one of the highest NAT8L expression levels, we analyzed two independent, previously published melanoma gene expression datasets and found that in both cohorts melanoma patients with above-the-median NAT8L gene expression levels had worse survival (Figure 2H).
Role of NAT8L Metabolism
We next investigated the direct effect of NAT8L on glucose and glutamine metabolism by performing 13C-based isotope tracer analysis using U-13C6 glucose and C-13C5 glutamine (26,27). Indeed, NAT8L knockdown increased glucose-derived aspartate and total aspartate (Figure 3, A and H). This increase in the aspartate level upon NAT8L knockdown is consistent with the reduced aspartate levels observed in ovarian cancer tissues compared with normal ovary (Interestingly, our results also suggest an increase in the contribution of glucose to the TCA cycle metabolites such as fumarate, citrate, and α-ketoglutarate with NAT8L knockdown. Furthermore, with glutamine-tracing experiments (Figure 3I), we found that while NAT8L knockdown leads to reduced glutamine contribution to the replenishment of TCA cycle metabolites (Figure 3J), there was no change in the total levels of these metabolites (Figure 3K).
Figure 3.
Effect of NAT8L on glucose and glutamine metabolism. 13C-based isotope tracer analysis using gas chromatography/mass spectrometry (GC/MS). A) Relative metabolite levels compared between NAT8L knockdown and control (n = 8). B) U-13C6 glucose isotope labeling map of glycolysis and TCA cycle. Mass isotopomer distribution of different metabolites compared between NAT8L knockdown and control (n = 4): (C) aspartate, (D) citrate, (E) fumarate, (F) alpha-ketoglutarate, (G) glutamate. H) The percentage of U-13C6 glucose contribution to TCA cycle metabolites. I) U-13C5 glutamine isotope labeling map of glycolysis and TCA cycle. J) Mass isotopomer distribution of M5 glutamate, alpha-ketoglutarate, M4 fumarate, asparate, citrate, and M5 citrate. K) The percentage of U-13C5 glutamine contribution to TCA cycle metabolites. Data are represented as mean ± SD (*P < .001). All statistical tests were two-sided (n = 4 from two independent experiments).
Biological Effects of NAA
To evaluate the potential biological roles of NAA in tumor growth, we used NAT8L siRNA to diminish NAA in ovarian cancer cells and confirmed the knockdown of both NAT8L and NAA (Supplementary Figure 2, F and G, available online). NAT8L silencing statistically significantly reduced cell proliferation (HEYA8: control siRNA 74.83%±0.92, NAT8L siRNA 55.70%±1.54, P < .001; A2780: control siRNA 50.17%±4.13, NAT8L siRNA 26.52%±3.70, P < .001) (Figure 4A) and induced G1 arrest (control siRNA 44.74%±0.15, NAT8L siRNA 53.14%±0.90, P < .001) and S arrest (control siRNA 44.18% ±1.05, NAT8L siRNA 32.53%±1.57, P < .001) (Figure 4B). Moreover, NAT8L silencing statistically significantly diminished cell viability (HEYA8: control siRNA 90.61%±2.53, NAT8L siRNA 39.43%±3.00, P < .001; A2780: control siRNA 90.59%±2.53, NAT8L siRNA 7.44%±1.71, P < .001) (Figure 4C) and increased apoptosis (HEYA8: control siRNA 19.31%±2.19, NAT8L siRNA 57.84%±0.12, P < .001; A2780: control siRNA 21.70%±1.64, NAT8L siRNA 59.55%±3.17, P < .001) (Figure 4D). In addition, increased apoptosis was observed with paclitaxel plus NAT8L silencing compared with treatment with paclitaxel alone (HEYA8: control siRNA 48.22%±2.63, NAT8L siRNA 78.05%±9.55, P < .001) (Figure 4E). A2780 cells transfected with 50 and 100nM of NAT8L siRNA for 48 and 72 hours were treated with NAA and demonstrated both partial and complete rescue from apoptosis (Figure 4F). Similar increases in apoptosis occurred in melanoma cell lines subjected to NAT8L siRNA treatment (Supplementary Figure 3A, available online).
Figure 4.
Silencing NAT8L in ovarian cancer cell lines. Comparison of treatment with control (NT) siRNA and NAT8L siRNA in (A) cell proliferation, (B) cell cycle progression, (C) cell viability, and (D) apoptosis in ovarian cancer cell lines (HEYA8 AND A2780). F) Administration of 100nM NAA in A2780 cells transfected with NAT8L siRNA (*P < .001). Data are presented as mean ± SD. Unpaired t test was used. All tests were two-sided. Graphs represent replicates of multiple repeated experiments. ptx = paclitaxel.
To systematically examine the molecular mechanisms by which NAT8L mediates it anti-apoptotic effect, a microarray was performed following NAT8L siRNA treatment in A2780 cells. Pathway enrichment analysis using Netwalker (28) showed that NAT8L siRNA treatment statistically significantly downregulates the anti-apoptotic pathway (P < .005). Upstream analysis was subsequently performed using Ingenuity Pathway Analysis (IPA) software to identify common upstream regulators of these genes. Out of all the potential upstream regulators identified, LEF1, POLR2A, CREBBP, and FOXM1 were statistically significantly downregulated after NAT8L siRNA treatment (Figure 5A). We next silenced NAT8L in A2780 cells in the presence or absence of NAA and assessed the expression levels of these genes. FOXM1 was statistically significantly downregulated after NAT8L silencing (7.7-fold, P = .03), and this effect was completely rescued after addition of NAA (13.8-fold, P < .001) (Figure 5B; Supplementary Figure 3B, available online). This demonstrated the critical role of FOXM1 in mediating the anti-apoptotic effects of NAT8L.
Figure 5.
Computational Pathway Analysis and in vitro validation of the effect of NAT8L silencing on expression of anti-apoptotic genes. A) Pathway analysis using IPA software. B) The effect of NAT8L silencing on FOXM1 expression level in A2780 cells. Cells were transfected with NAT8L siRNA (100nM) for six hours before exposing to complete media with or without NAA (50nM). FOXM1 levels were assessed at 24 hours post-transfection. Data are presented as mean ± SD (*P < .001). Unpaired t test was used. All tests were two-sided. Graphs represent replicates of a single experiment.
Next, we evaluated the effect of NAA biosynthesis on tumor growth in orthotopic ovarian cancer mouse models using 1,2-dioleoyl-sn-glycero-3-phosphatidylcholine (DOPC) nanoliposomes containing NAT8L siRNA. Tumor growth was statistically significantly decreased in mice treated with NAT8L siRNA–DOPC compared with controls (A2780: control siRNA 0.52 g±0.15, NAT8L siRNA 0.08 g±0.17, P < .001; HEYA8: control siRNA 0.79 g±0.42, NAT8L siRNA 0.24 g±0.18, P = .008) (Figure 6A). Immunohistochemical analysis of the tumors using Ki67 and caspase-3 revealed that cell proliferation was statistically significantly decreased and apoptosis statistically significantly increased in the samples treated with NAT8L siRNA–DOPC (Supplementary Figure 3, C and D, available online). In addition, compared with treatment with control siRNA or monotherapy (NAT8L siRNA or paclitaxel), treatment with NAT8L siRNA–DOPC plus paclitaxel (0.06 g±0.06) resulted in statistically significantly decreased tumor burden (control siRNA 1.19 g±1.18, P = .03; paclitaxel 0.37 g±0.19, P = .004; NAT8L siRNA 0.16 g±0.09, P = .03) (Figure 6B). Similarly, A375-SM orthotopic melanoma mouse models treated with NAT8L siRNA–DOPC had a statistically significantly lower tumor burden than the control mice (control siRNA 0.55 g±0.22, NAT8L siRNA 0.21 g±0.17, P = .001) (Figure 6C).
Figure 6.
Silencing NAT8L in orthotopic mouse models of ovarian cancer and melanoma. A) A2780: siRNA NT (n = 6), siRNA NAT8L siRNA NT (n = 6); HEYA8: siRNA NT (n = 8), siRNA NAT8L siRNA NT (n = 7). B) Combination = siRNA NAT8L + paclitaxel. HEYA8: siRNA NT (n = 8), paclitaxel (n = 7), siRNA NAT8L (n = 7), siRNA combination (n = 7). C) A375-SM: siRNA NT (n = 10), siRNA NAT8L siRNA NT (n = 10). siRNA NT = control siRNA. Data are presented as mean ± SD. Unpaired t test was used. All tests were two-sided. Graphs represent replicates of a single experiment.
Discussion
The ability of tumor cells to reprogram their metabolomic profile, compared with normal cells, has been recognized as a hallmark of cancer (1). Using nonbiased, global approaches to characterize the metabolic differences between normal ovarian tissue and ovarian cancer, we identified NAA as one of the most highly upregulated metabolites in ovarian cancer. NAA, a metabolite synthesized by acetyl coenzyme A and the nonessential amino acid aspartic acid, is the second most abundant amino acid derivative in the brain that is found equally in the gray and white matter (29,30). Its synthesis occurs within neuronal mitochondria and is transported to astrocytes via a shuttle with release into circulation or to oligodendrocytes, where its catabolism occurs (31,32). NAA functions to provide a source of metabolic acetate necessary for oligodendrocyte myelination, but other functions are thought to include mitochondrial metabolism, neuronal osmoregulation, and axon-glial signaling (33).
Because of its prevalence within the central nervous systems, past work has primarily focused on neurodegenerative disorders such as Canavan disease, an autosomal recessive degenerative disorder that results from a missense mutation in ASPA. This group of patients has abnormally high levels of NAA. Within oncology, NAA metabolism has been exploited for novel imaging modalities (34–36). However, its clinical and biological significance were not known.
Here, we found that ovarian cancer patients with elevated levels of NAA have worse clinical outcomes and survival. Previous work has found NAA to be more abundant in tumors compared with noncancer tissues (4,37,38). However, to our knowledge, no other study to date has investigated the clinical and biological roles of NAA in cancer. We explored the tumor cell’s reliance on NAA and found that the abrogation of NAA biosynthesis can impede tumor growth and survival. Administration of NAA to cells transfected with NAT8L siRNA rescued cells from apoptosis, suggesting the cellular dependence on NAA for growth and survival. This reprogramming of the tumor metabolism using siRNA combined with traditional chemotherapeutics led to even greater reductions in tumor growth in vitro and in vivo. Furthermore, we identified FOXM1, a transcription factor, as a regulator of the mechanism by which NAT8L promotes survival. FOXM1 is statistically significantly altered in a substantial proportion of high-grade serous ovarian cancers (39). Hence, tumor metabolic reprogramming by silencing NAT8L disrupts NAA production and could block FOXM1-mediated tumor cell survival.
A potential limitation of our study is that NAT8L enzyme activity was not directly assessed with regard to levels of NAA in tumor cells. However, prior publications have shown that the key NAA biosynthesis enzyme is NAT8L (31,32). Furthermore, tumor gene expression of NAT8L and ASPA in our study showed a statistically significant correlation (Figure 1D) with only NAT8L with respect to tumor NAA levels.
In summary, our study provides a new understanding of metabolic alterations in ovarian cancer. Furthermore, the role of NAA is likely to extend to other malignancies, which suggests its role as a key oncometabolite. Targeting the NAA pathway could have relevance for the development of future therapies or biomarkers for ovarian and other malignancies.
Funding
This work was supported, in part, by the National Cancer Institute (NCI) Network on Biobehavioral Pathways in Cancer (contract number HHSN261200800001E), National Institutes of Health (NIH) grants (P50CA083639, CA109298, P50CA098258, U54CA151668, UH2TR000943, CA016672, U54CA96300, and U54CA96297), CPRIT (RP110595 and RP120214), an Ovarian Cancer Research Fund Program Project Development Grant, the Betty Ann Asche Murray Distinguished Professorship, the RGK Foundation, the Gilder Foundation, the Judi A. Rees Ovarian Cancer Research Fund, the Chapman Foundation, the Meyer and Ida Gordon Foundation, and the Blanton-Davis Ovarian Cancer Research Program. SYW is supported by Ovarian Cancer Research Fund, Inc., Foundation for Women’s Cancer, and Cancer Prevention Research Institute Texas (CPRIT) training grants (RP101502 and RP101489). BZ, RAP, and HJD are supported by NIH T32 Training Grant CA101642. We also acknowledge support from the Small Animal Imaging Facility. MD Anderson’s Nuclear Magnetic Resonance Facility is supported in part by the NIH through MD Anderson Cancer Center Support Grant CA016672.
Supplementary Material
The study funders had no role in design of the study; the collection, analysis, or interpretation of the data; the writing of the manuscript; or the decision to submit the manuscript for publication.
The authors thank Dr. Robert Langley for helpful discussions.
None of the authors have any conflicts of interest related to the contents of this manuscript.
References
- 1. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. [DOI] [PubMed] [Google Scholar]
- 2. Warburg O. On the origin of cancer cells. Science. 1956;123(3191):309–314. [DOI] [PubMed] [Google Scholar]
- 3. DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metabolism. 2008;7(1):11–20. [DOI] [PubMed] [Google Scholar]
- 4. Sreekumar A, Poisson LM, Rajendiran TM, et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009;457(7231):910–914. [DOI] [PMC free article] [PubMed] [Google Scholar] [Research Misconduct Found]
- 5. Losman JA, Looper RE, Koivunen P, et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science. 2013;339(6127):1621–1625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Vander Heiden MG, Locasale JW, Swanson KD, et al. Evidence for an alternative glycolytic pathway in rapidly proliferating cells. Science. 2010;329(5998):1492–1499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Possemato R, Marks KM, Shaul YD, et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature. 2011;476(7360):346–350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Salimian Rizi B, Caneba C, Nowicka A, et al. Nitric oxide mediates metabolic coupling of omentum-derived adipose stroma to ovarian and endometrial cancer cells. Cancer Res. 2015;75(2):456–471. [DOI] [PubMed] [Google Scholar]
- 9. Dang CV, Le A, Gao P. MYC-induced cancer cell energy metabolism and therapeutic opportunities. Clin Cancer Res. 2009;15(21):6479–6483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Son J, Lyssiotis CA, Ying H, et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature. 2013;496(7443):101–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Evans AM, DeHaven CD, Barrett T, Mitchell M, Milgram E. Integrated, nontargeted ultrahigh performance liquid chromatography/electrospray ionization tandem mass spectrometry platform for the identification and relative quantification of the small-molecule complement of biological systems. Anal Chem. 2009;81(16):6656–6667. [DOI] [PubMed] [Google Scholar]
- 12. Weiner J, 3rd, Parida SK, Maertzdorf J, et al. Biomarkers of inflammation, immunosuppression and stress with active disease are revealed by metabolomic profiling of tuberculosis patients. PloS One. 2012;7(7):e40221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Lawton KA, Berger A, Mitchell M, et al. Analysis of the adult human plasma metabolome. Pharmacogenomics. 2008;9(4):383–397. [DOI] [PubMed] [Google Scholar]
- 14. Dehaven CD, Evans AM, Dai H, Lawton KA. Organization of GC/MS and LC/MS metabolomics data into chemical libraries. J Cheminform. 2010;2(1):9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Merritt WM, Lin YG, Han LY, et al. Dicer, Drosha, and Outcomes in Patients with Ovarian Cancer. New Engl J Med. 2008;359(25):2641–2650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Thaker PH, Han LY, Kamat AA, et al. Chronic stress promotes tumor growth and angiogenesis in a mouse model of ovarian carcinoma. Nat Med. 2006;12(8):939–944. [DOI] [PubMed] [Google Scholar]
- 17. Li L, Price JE, Fan D, Zhang RD, Bucana CD, Fidler IJ. Correlation of growth capacity of human tumor cells in hard agarose with their in vivo proliferative capacity at specific metastatic sites. J Natl Cancer Inst. 1989;81(18):1406–1412. [DOI] [PubMed] [Google Scholar]
- 18. Lu C, Han HD, Mangala LS, et al. Regulation of tumor angiogenesis by EZH2. Cancer Cell. 2010;18(2):185–197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Landen CN, Jr, Lu C, Han LY, et al. Efficacy and antivascular effects of EphA2 reduction with an agonistic antibody in ovarian cancer. J Natl Cancer Inst. 2006;98(21):1558–1570. [DOI] [PubMed] [Google Scholar]
- 20. Merritt WM, Lin YG, Spannuth WA, et al. Effect of interleukin-8 gene silencing with liposome-encapsulated small interfering RNA on ovarian cancer cell growth. J Natl Cancer Inst. 2008;100(5):359–372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Storey JD, Tibshirani R. Statistical significance for genomewide studies. Proc Natl Acad Sci U S A. 2003;100(16):9440–9445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Tothill RW, Tinker AV, George J, et al. Novel molecular subtypes of serous and endometrioid ovarian cancer linked to clinical outcome. Clin Cancer Res. 2008;14(16):5198–5208. [DOI] [PubMed] [Google Scholar]
- 23. Bogunovic D, O’Neill DW, Belitskaya-Levy I, et al. Immune profile and mitotic index of metastatic melanoma lesions enhance clinical staging in predicting patient survival. Proc Natl Acad Sci U S A. 2009;106(48):20429–20434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Jonsson G, Busch C, Knappskog S, et al. Gene expression profiling-based identification of molecular subtypes in stage IV melanomas with different clinical outcome. Clin Cancer Res. 2010;16(13):3356–3367. [DOI] [PubMed] [Google Scholar]
- 25. Grambsch PM, Therneau TM. Proportional Hazards Tests and Diagnostics Based on Weighted Residuals. Biometrika. 1994;81(3):515–526. [Google Scholar]
- 26. Caneba CA, Yang L, Baddour J, et al. Nitric oxide is a positive regulator of the Warburg effect in ovarian cancer cells. Cell Death Dis. 2014;5:e1302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Yang L, Moss T, Mangala LS, et al. Metabolic shifts toward glutamine regulate tumor growth, invasion and bioenergetics in ovarian cancer. Mol Syst Biol. 2014;10:728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Komurov K, Dursun S, Erdin S, Ram PT. NetWalker: a contextual network analysis tool for functional genomics. BMC Genomics. 2012;13:282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Tallan HH. Studies on the distribution of N-acetyl-L-aspartic acid in brain. J Biol Chem. 1957;224(1):41–45. [PubMed] [Google Scholar]
- 30. Tallan HH, Moore S, Stein WH. Studies on the free amino acids and related compounds in the tissues of the cat. J Biol Chem. 1954;211(2):927–939. [PubMed] [Google Scholar]
- 31. Ariyannur PS, Moffett JR, Manickam P, et al. Methamphetamine-induced neuronal protein NAT8L is the NAA biosynthetic enzyme: implications for specialized acetyl coenzyme A metabolism in the CNS. Brain Res. 2010;1335:1–13. [DOI] [PubMed] [Google Scholar]
- 32. Wiame E, Tyteca D, Pierrot N, et al. Molecular identification of aspartate N-acetyltransferase and its mutation in hypoacetylaspartia. Biochem J. 2010;425(1):127–136. [DOI] [PubMed] [Google Scholar]
- 33. Moffett JR, Ross B, Arun P, Madhavarao CN, Namboodiri AM. N-Acetylaspartate in the CNS: from neurodiagnostics to neurobiology. Prog Neurobiol. 2007;81(2):89–131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Stadlbauer A, Hammen T, Buchfelder M, et al. Differences in metabolism of fiber tract alterations in gliomas: a combined fiber density mapping and magnetic resonance spectroscopic imaging study. Neurosurgery. 2012;71(2):454–463. [DOI] [PubMed] [Google Scholar]
- 35. Li Y, Lupo JM, Parvataneni R, et al. Survival analysis in patients with newly diagnosed glioblastoma using pre- and postradiotherapy MR spectroscopic imaging. Neuro-oncology. 2013;15(5):607–617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Yao X, Zeng M, Wang H, Fei S, Rao S, Ji Y. Metabolite detection of pancreatic carcinoma by in vivo proton MR spectroscopy at 3T: initial results. Radiol Med. 2012;117(5):780–788. [DOI] [PubMed] [Google Scholar]
- 37. Fong MY, McDunn J, Kakar SS. Identification of metabolites in the normal ovary and their transformation in primary and metastatic ovarian cancer. PloS One. 2011;6(5):e19963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Ben Sellem D, Elbayed K, Neuville A, et al. Metabolomic Characterization of Ovarian Epithelial Carcinomas by HRMAS-NMR Spectroscopy. J Oncol. 2011;2011:174019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Cancer Genome Atlas Research N. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474(7353):609–615. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.