Abstract
Transcription by RNA polymerase II (Pol II) is a complex process that requires general transcription factors and Pol II to assemble on DNA into preinitiation complexes that can begin RNA synthesis upon binding of NTPs (nucleoside triphosphate). The pathways by which preinitiation complexes form, and how this impacts transcriptional activity are not completely clear. To address these issues, we developed a single molecule system using TIRF (total internal reflection fluorescence) microscopy and purified human transcription factors, which allows us to visualize transcriptional activity at individual template molecules. We see that stable interactions between polymerase II (Pol II) and a heteroduplex DNA template do not depend on general transcription factors; however, transcriptional activity is highly dependent upon TATA-binding protein, TFIIB and TFIIF. We also found that subsets of general transcription factors and Pol II can form stable complexes that are precursors for functional transcription complexes upon addition of the remaining factors and DNA. Ultimately we found that Pol II, TATA-binding protein, TFIIB and TFIIF can form a quaternary complex in the absence of promoter DNA, indicating that a stable network of interactions exists between these proteins independent of promoter DNA. Single molecule studies can be used to learn how different modes of preinitiation complex assembly impact transcriptional activity.
INTRODUCTION
Transcription is the first step in gene expression and requires the regulated assembly of numerous protein factors and DNA elements. Eukaryotic protein coding genes are transcribed by RNA polymerase II (Pol II), which requires a set of general transcription factors (GTFs), including TFIID, TFIIA, TFIIB, TFIIF, TFIIE and TFIIH to achieve promoter specific transcription (1). During transcription initiation, the GTFs function to properly recruit and position Pol II at the core promoters of genes; this protein assembly on promoter DNA is referred to as a preinitiation complex (PIC). Within cells, regulated PIC assembly and transcription also require the coordination of multiple other components such as activators, repressors, coregulators and chromatin remodeling complexes (2,3).
Biochemical experiments in highly purified transcription systems have shown that not all GTFs are required to observe basal level transcriptional activity. For example, functional transcription complexes can be assembled with the TATA-binding protein (TBP) subunit of TFIID, TFIIB, TFIIF and Pol II (4–7). In this system lacking TFIIE and the TFIIH helicase, studies of transcriptional activity require that the DNA be underwound, either by negative supercoiling or by the presence of a heteroduplex that mimics a melted transcription bubble (5,6,8,9). Minimal transcription systems have enabled studies to systematically determine the specific roles that each factor plays in PIC assembly and activity, as well as elucidate the mechanisms of specific steps early in transcription. Additional GTFs or other factors can then be added to minimal systems to directly assess the impact that they have on transcription.
Regulation of PIC assembly is essential for setting transcription levels, thus it is important to understand specific molecular mechanisms regarding how active PICs form (10). Extensive biochemical and cell based experiments have established two primary models describing how PICs form: the sequential assembly model and the Pol II holoenzyme model. The sequential assembly model suggests that PICs form via the ordered binding of specific GTFs to promoter DNA (as reviewed in (1,11,12)). The holoenzyme model suggests that complexes of Pol II and GTFs assemble off of promoter DNA, and then bind DNA as complexes (13–15). The sequential assembly model was supported by early biochemical data demonstrating that transcriptional activity required that chromatographic fractions from HeLa cells, which contained different GTFs, be added to transcription reactions in a specific order (16,17). DNase I footprinting analysis and electrophoretic mobility shift assays (EMSAs) with recombinant and highly purified factors later established the order in which GTFs bind to promoter DNA to facilitate formation of transcription complexes (6,18,19). Specifically, these studies found that TFIID initially binds promoter DNA, nucleating the PIC. This is followed by binding of TFIIA and TFIIB, which stabilize the interactions between TFIID and DNA. This platform then assists in recruitment of the Pol II/TFIIF complex. Following Pol II/TFIIF binding, TFIIE and TFIIH then associate with the complex. Data supporting the Pol II holoenzyme pathway also suggest that TFIID binds to the promoter initially to facilitate recruitment of ‘holoenzyme’ complexes containing GTFs and Pol II (13–15). The compositions of Pol II holoenzyme complexes that have been purified from cells differ between studies (20,21); however, they all contain many of the GTFs, Pol II and coregulatory factors.
While each assembly mechanism (i.e. ordered binding or holoenzyme binding) is supported by biochemical data, molecular details pertaining to the pathway of PIC formation in cells remain unclear. It is likely that both modes of assembly exist, are not mutually exclusive, and different pathways may serve as points of regulation for transcription at different promoters. Additionally, it is possible that assembly pathways other than these two classical models occur. Interestingly, PICs form with high efficiency in vitro, however only a small percentage (∼2–20%) of these complexes is transcriptionally active (22–25). Similar results were found using a live-cell imaging approach; only ∼1% of the interactions between Pol II and promoter DNA resulted in productive transcription (26). This indicates that formation of active complexes is likely a key regulatory point for transcription. It is possible that the pathway by which PICs form dictates their transcriptional activity.
To study the formation of active transcription complexes, we developed a single molecule platform using total internal reflection fluorescence (TIRF) microscopy. Our experimental setup allows visualization of assembled transcription complexes and their activity at individual DNA molecules, thereby overcoming limitations of studying transcription in ensemble systems in which only a relatively small fraction of assembled complexes are active. The single molecule assay exhibits levels of activity similar to, if not better than, ensemble transcription assays. Interestingly, levels of activity are the same when either promoter DNA or Pol II is immobilized on the surface of the microscope slide. Our single molecule data reveal that assembly of complexes containing Pol II and heteroduplex DNA does not depend on GTFs, but transcriptional activity is highly dependent upon GTFs. Additionally, stable subcomplexes of Pol II/GTFs can assemble and act as precursors to transcriptionally active PICs. The single molecule system provides a highly sensitive method to evaluate how activity changes when PICs are assembled via different pathways, thereby yielding new insight into how active transcription complexes form.
MATERIALS AND METHODS
Oligonucleotides
For the three piece heteroduplex DNA construct the sequence of the template strand was 5′CCTGAGGTTAGTGTGAGTAGTGATTAAAGATAGTGATGAGGACGAACGCGCCCCCACCCCCTTTTATAGCCCCCCTT (−40 to +37) and the sequences of the two nontemplate strand pieces were 5′AAGGGGGGCTATAAAAGGGGGTGGGGGCGCGAAGCAGGAGTAGACTATCTTTAATCACTA (−40 to +20) and 5′CTCACACTAACCTCAGG (+21 to +37). The underlined region (−9 to +3) indicates the mismatches that created a premelted heteroduplex region. For experiments in which the DNA was immobilized, the nontemplate strand (−40 to +20 oligo) was functionalized with a biotin molecule at the 5′ end and contained a linker sequence (5′TACGAGGAAT) that preceded the AdML promoter region (Integrated DNA Technologies). Colocalization experiments were performed with a template strand functionalized with Cy5 and Cy3 molecules on the 5′ and 3′ ends, respectively (Eurofins Scientific). For transcriptional activity assays, the short nontemplate strand oligonucleotide (+21 to +37) was functionalized with the Iowa Black RQ-Sp quencher on the 3′ end (Integrated DNA Technologies). The unlabeled template strand and the long nontemplate strand (−40 to +20) oligos were gel purified using 7 M urea/8 or 13% polyacrylamide gels. The oligos were cut out of the gels, crushed slices were nutated overnight in 10 mM Tris (pH 7.9), 0.1 mM EDTA at 4°C, and then ethanol precipitated. The doubly labeled template strand was not purified in order to preserve the integrity of the fluorophores. Three-piece DNA constructs were assembled as follows: oligos were heated to 95°C for 5 min, cooled to 60°C at a rate of 0.1°C/s, incubated at 60°C for 60 min, cooled to 45°C at a rate of 0.1°C/s, incubated at 45°C for 60 min, then cooled to 4°C at a rate of 0.1°C/s. The sequence of the template stand of the 18 bp TATA DNA construct was as follows: 5′ CGTCCCTTTTATAGCCTG. The nontemplate stand was the reverse complement. The 5′ ends of the template and nontemplate strands were functionalized with Alexa647 and Alexa555, respectively (Integrated DNA Technologies). The 18 bp TATA construct was annealed as described for the three-piece construct, but was incubated at 57°C for 60 min in place of the 60 and 45°C incubations.
Ensemble in vitro transcription reactions
Recombinant human TBP, TFIIB and TFIIF were expressed in E. coli and purified as previously described (22,27,28). Human Pol II was purified from HeLa cells as previously described (29). Ensemble in vitro transcription reactions (Figure 1A) were performed in 10% glycerol, 10 mM Tris (pH 7.9), 50 mM KCl, 1 mM DTT, 0.05 mg/ml BSA, 10 mM HEPES (pH 7.9) and 4 mM MgCl2. The three-piece DNA construct lacked modifications on the DNA. DNA and proteins were added to the reactions at approximate final concentrations of 2 nM DNA, 3.5 nM TBP, 10 nM TFIIB, 2 nM TFIIF and 2 nM Pol II. DNA was incubated with TBP, TFIIB, TFIIF, and Pol II for 20 min at room temperature (∼23°C) to allow PICs to form. Nucleotides (final concentration of 625 μM ATP (adenosine triphosphate), 625 μM UTP (uridine triphosphate), 25 μM [α-32P] CTP (cytidine triphosphate), 5 μCi per reaction) were then added to the reactions and transcription was allowed to proceed for 30 min at room temperature. Transcription was stopped and RNA products were ethanol precipitated as previously described (22). RNAs were resolved on a 20% polyacrylamide/7 M urea gel and imaged using a PhosphorImager.
Figure 1.
Transcriptional activity can be monitored by the release of an oligo containing a quencher molecule. (A) Transcription from a 3-piece chimeric heteroduplex template. Reactions were performed in the absence of GTP, which halted Pol II after synthesis of a 35 nt RNA due to the presence of the first C in the template strand at +36. 32P-labeled RNA transcripts were resolved on a 20% denaturing polyacrylamide gel. (B) Schematic of the three-piece quenched construct used to visualize transcription. (C) PICs migrate slower than Pol II/DNA complexes in native gels. Migration of the fluorescent three-piece heteroduplex template was monitored with Pol II and the GTFs or with Pol II alone. Shown is a 4% native gel scanned for Cy3 fluorescence. (D) Release of the quencher oligo in the three piece DNA construct pictured in panel B is NTP dependent. Shown is a 4% native gel scanned for both Cy3 (left panel) and Cy5 (right panel) fluorescence. The positions at which free DNA and PICs migrate are indicated.
Electrophoretic mobility shift transcriptional activity assays
EMSAs (Figure 1C and D) were performed using the three-piece quenched DNA construct pictured in Figure 1B. Reactions were assembled and performed as described for the ensemble transcription assays, except final NTP (ATP, CTP, UTP) concentrations were 625 μM each and 0.5 ng/μl poly-dGdC was added as a competitor DNA. After allowing transcription to occur, reactions were run on a 4% native polyacrylamide gel containing 0.5X TBE and 5% glycerol and imaged by scanning with both a 532 nm and 640 nm laser and reading fluorescence emission using 580 and 670 nm emission filters, respectively, on a Typhoon Imager 9400 (GE biosciences).
Single molecule surface functionalization
Glass coverslips and microscope slides were cleaned and hydroxylated by sonicating slides in 1% Alconox for 20 min, ethanol for 60 min and 1 M KOH for 20 min. Coverslips and slides were then silanated by incubating for 20 min in a solution of 2% N-(2-Aminoethyl)-3-Aminopropyltrimethoxysilane (United Chemical Technologies) in methanol, with a 1 min sonication after 10 min. Following silanation, slides were rinsed in methanol and baked at 110°C for 10 min. Slides and coverslips were PEGylated using a mixture of 0.38% w/v biotin-PEG-SC (MW 5000, Laysan Bio, Inc.) and 20% w/v mPEG-SVA (MW 5000, Laysan Bio, Inc.) in 0.1 M sodium bicarbonate. The PEG mixture was incubated on the glass for at least 3 h. Following PEGylation, slides and coverslips were rinsed with MilliQ water, dried with ultrapure nitrogen gas and flow chambers were assembled by adhering a coverslip to a microscope slide using double-sided tape (30).
Prior to all single molecule experiments, flow-chambers were washed thoroughly with reaction buffer (10% glycerol, 10 mM Tris (pH 7.9), 10 mM HEPES (pH 7.9), 50 mM KCl, 1 mM DTT, 0.05 mg/ml BSA, 4 mM MgCl2). Streptavidin (Sigma) was resuspended to a final concentration of 1 mg/ml in resuspension buffer (10 mM Tris (pH 7.9), 10% glycerol, 50 mM KCl and 5 mM MgCl2) and stored at −20°C. Surfaces were coated with streptavidin by flowing in a solution of 0.2 mg/ml streptavidin and incubating for 5 min, followed by thorough washing to remove excess protein. For experiments in which DNA was immobilized on the surface, biotinylated DNA was flowed into the chamber at a concentration of 0.125–0.2 nM, then surfaces were washed thoroughly with reaction buffer.
Single molecule assays
Single molecule transcription assays were performed using the same conditions and protein concentrations as described for the electrophoretic mobility shift transcription activity assays. For experiments in which Pol II was immobilized on the surface, 10 nM biotinylated Protein A (Millipore) was flowed into the chamber in reaction buffer, incubated for 10 min and then the chamber was washed. The surface was then coated with a Pol II-specific antibody (8WG16) by flowing in a solution of 0.05 mg/ml 8WG16 in reaction buffer, incubating for 10 min and then washing out excess antibody. Pol II was added to the chamber at a final concentration of 2 nM in reaction buffer, incubated for 20 min and unbound Pol II was washed out of the chamber with reaction buffer. For both binding and transcription assays, DNA was flowed into the imaging chamber at a final concentration of 0.25 nM for assays in which Pol II was immobilized and 0.125–0.2 nM for assays in which DNA was directly immobilized on the surface via biotin.
Prior to any fluorescence imaging, imaging buffer containing an oxygen scavenging system was flowed into the chamber. Imaging buffer contained reaction buffer plus 0.04 mg/ml catalase, 1.02 mg/ml glucose oxidase, 0.83% w/v D-glucose and 3 mM trolox. Stock solutions of catalase and glucose oxidase (Sigma) were resuspended to final concentrations of 3.4 mg/ml and 44 mg/ml in storage buffer (50% glycerol, 100 mM Tris (pH 7.9), 50 mM KCl) and stored at −20°C. D-glucose (Sigma) was resuspended at 10% w/v in 10 mM Tris (pH7.9), 50 mM NaCl, and stored at −20°C. Trolox (Sigma) was dissolved in 18 MΩ water by adding 4M NaOH to a stock concentration of 100 mM.
Data collection and analysis
All single molecule data were collected using an objective-based TIRF microscope (Nikon TE-2000U) equipped with a 1.49 NA immersion objective and both a 532 and a 640 nm continuous wave laser. Fluorescence emission from the 532 nm laser was acquired using an Evolve CCD and emission from the 640 nm laser was collected using a Cascade II CCD. Data were collected using NIS-Elements software using a 100 ms exposure time.
Fluorescence emission movies were analyzed for colocalization using an in-house mapping software program. This analysis involved first localizing all spots in both the green and red movies separately and then mapping the locations at which green and red signal overlapped, indicating a colocalized spot pair. For binding experiments, colocalized spot pairs were compared as a metric for the number of binding events. For transcriptional activity experiments, template usage was calculated from the percentage of total green spots that colocalized with a red spot (template usage = (colocalized spot pairs)/(total green spots) × 100). As a control, additional colocalization analyses were always performed with the Cy3 image rotated by 90° to ensure that colocalization was not due to random overlap of spots.
RESULTS
A fluorescence based Pol II transcriptional activity assay
To develop a single molecule transcription assay, we optimized a DNA template consisting of a core promoter and early transcribed region that was transcribed by Pol II with reasonably high template usage. We arrived at a 77 bp chimeric DNA construct consisting of the adenovirus major late promoter (AdMLP) sequence from positions −40 to −1 and a modified early transcribed region of the interleukin-2 (IL-2) gene from positions +1 to +37. We did not use AdMLP sequence downstream of the start site because it is prone to transcript slippage, which decreases productive RNA synthesis (31–33); rather, we knew the IL-2 sequence to be transcribed efficiently through the early steps of transcription (34). Three additional point mutations in the transcribed sequence were made: a C to A at position +7 in the nontemplate strand which enhances the rate of promoter escape (27) and two C to G mutations at positions +36 and +37 in the nontemplate strand to cause transcription to halt in the absence of GTP. The DNA also contained a heteroduplex region with base mismatches from −9 to +3 that generated a bubble (8). This alleviated the requirement for TFIIE and the helicase activity of TFIIH, thereby allowing specific initiation to occur in a minimal system consisting only of highly purified TBP, TFIIB, TFIIF and Pol II. Ensemble in vitro transcription from this promoter DNA was highly TBP-dependent (Figure 1A).
To detect transcription from this promoter at the single molecule level, we designed the three-piece DNA construct illustrated in Figure 1B. The template strand was labeled with fluorophores on both the 3′ and the 5′ ends (Cy3 and Cy5, respectively). The nontemplate strand consisted of two pieces (−40 to +20 and +21 to +37), resulting in a nick in the backbone between bases +20 and +21. The nontemplate strand oligo from +21 to +37 contained a small-molecule quencher on the 3′ end, which quenched fluorescence from the Cy5 molecule on the 5′ end of the template strand. Due to the nick in the backbone, transcription caused the quencher oligo (nontemplate strand) to be released, thereby revealing fluorescent signal from the Cy5 dye on the template strand. Therefore, DNAs that were transcribed could be identified at the single molecule level by visualizing colocalization of the Cy3 and Cy5 dye molecules after addition of NTPs.
We first confirmed this approach was feasible using ensemble transcription reactions coupled to EMSAs. This allowed us to use the Cy3 fluorescent signal to monitor PIC formation via the DNA, and the appearance of Cy5 fluorescent signal to monitor transcriptional activity. Figure 1C shows that addition of Pol II and GTFs to DNA resulted in formation of PICs and a complete shift of the Cy3 signal to a slower migrating band (compare lanes 1 and 2 to lanes 3 and 4). The DNA migrated at a distinctly different position in the absence of the general transcription factors (lanes 5 and 6), allowing us to distinguish PICs from Pol II alone bound to the heteroduplex DNA template. To observe both PIC formation and transcriptional activity, we performed transcription reactions using the 3-piece quenched template shown in Figure 1B and subsequently scanned the gel for both Cy3 and Cy5 signal. Figure 1D shows an EMSA that was scanned for Cy3 (left panel) and then Cy5 (right panel), and the images were aligned. Lanes 1 and 2 in the Cy5 channel show that the Cy5 fluorophore was efficiently quenched in the double stranded DNA. After addition of NTPs (G-less), Cy5 signal was observed (lane 3), showing that transcription leads to release of the quencher oligo. The Cy5 signal migrated at the same position as PICs in the Cy3 channel, indicating that Pol II remained on the template after transcription was complete, which is consistent with transcription halting at +35 in the absence of GTP.
Template usage is the same from PICs assembled under ensemble and single molecule conditions
Performing this assay at the single molecule level required that we be able to specifically colocalize Cy3 and Cy5 on the surface of a coverslip in our TIRF system. To optimize this, we utilized a three-piece DNA construct that omitted the quencher at the 3′ end of the nontemplate strand and contained a biotin molecule at the 5′ end of the nontemplate strand. We immobilized this construct on a streptavidin-coated surface and imaged the DNA using TIRF microscopy. The Cy3 and the Cy5 molecules were excited using 532 and 640 nm lasers, respectively; emissions from the two dyes were recorded on separate CCD cameras. We analyzed each set of movies using in-house software to identify red and green spots separately and then mapped the locations where the signals from both channels aligned. We found that 71.1% of Cy3 molecules colocalized with a Cy5 molecule (Table 1), and 65.5% of Cy5 molecules colocalized with a Cy3 molecule. Because both dyes were present on a single strand of DNA, these numbers suggest that ∼30–35% of the templates contained only one photoactive dye. As a control we performed the same analysis after rotating the Cy3 image 90°. This resulted in only a nominal number of colocalized spot pairs, demonstrating that colocalization does not result from random overlap of green and red spots. Lastly, to assess how well the quencher oligo worked at eliminating Cy5 fluorescence at the single molecule level, we immobilized the construct illustrated in Figure 1B (via a biotin on the 5′ end of the nontemplate strand) and used TIRF to colocalize red and green spots. As shown in Table 1, far fewer red spots were observed and only 2.2% of the Cy3 spots colocalized with a Cy5 spot, compared to 71.1% in the absence of the quencher; therefore background colocalization due to incomplete annealing of the quencher oligo was low.
Table 1. Colocalization of green (Cy3) and red (Cy5) dyes in the presence and absence of a quencher molecule.
| Green spots | Red spots | Colocalized spot pairs | % of green spots that have a colocalized red | |
|---|---|---|---|---|
| DNA without quencher | 4647 | 5041 | 3305 | 71.1 |
| DNA with quencher | 5440 | 436 | 118 | 2.2 |
To measure transcriptional activity using single molecule TIRF, we first performed ensemble transcription using a biotinylated three-piece construct containing the quencher oligo, and then flowed the reaction onto a streptavidin-coated slide. We evaluated the amount of transcription that occurred by determining the template usage, defined as the percentage of immobilized DNA templates (i.e. Cy3 spots) that were transcribed (i.e. had a colocalized Cy5 spot after the addition of NTPs). We found that 12.2% of the DNAs in the ensemble reaction contained a Cy3 spot that colocalized with a Cy5 spot (Table 2). We had determined that 2.2% of Cy3 spots colocalized with a Cy5 spot in the absence of NTPs (Table 1). Correcting for this, we conclude that under ensemble conditions, we observed a template usage value of 10.0%. The template usage values reported in the text were corrected for NTP-independent colocalization observed in each experiment. Our template usage value of 10.0% is on par or greater than previous reports of template usage from model core promoters (22–24). This template usage value does not account for the DNA templates that do not contain a photoactive red dye. Accounting for DNAs containing a ‘dark’ red dye, we calculate a template usage of 14.1%. Because we are primarily interested in comparing template usage values under different conditions, and not with the absolute template usage values, the numbers we report are not corrected for the percentage of inactive red dyes.
Table 2. Template usage under ensemble conditions is comparable to template usage from complexes formed on surface-immobilized DNA.
| Green spots | Red spots | Colocalized spot pairs | % of green spots that have a colocalized red | |
|---|---|---|---|---|
| Ensemble transcription | 3932 | 1086 | 479 | 12.2 |
| Transcription on surface (-NTPs) | 5561 | 901 | 172 | 3.1 |
| Transcription on surface (+NTPs) | 5523 | 2713 | 682 | 12.4 |
We then determined template usage when both PIC assembly and transcription occurred on the surface of the slide. We flowed TBP, TFIIB, TFIIF and Pol II into the chamber containing immobilized DNA and allowed PICs to form. We then imaged the DNA to confirm that the formation of complexes did not disrupt the quencher oligo; indeed, only 3.1% of Cy3 spots had a colocalized Cy5 spot (Table 2), which is similar to the value obtained in the absence of proteins (Table 1). We then added NTPs to the slide, allowed transcription to proceed, and subsequently imaged the dyes. After addition of NTPs, 12.4% of Cy3 spots had a colocalized Cy5 spot, resulting in a template usage value of 9.3% (Table 2). These data show that PIC assembly and transcription occurred on the surface of the slide as efficiently as under ensemble conditions.
Transcriptionally active PICs can assemble on immobilized Pol II
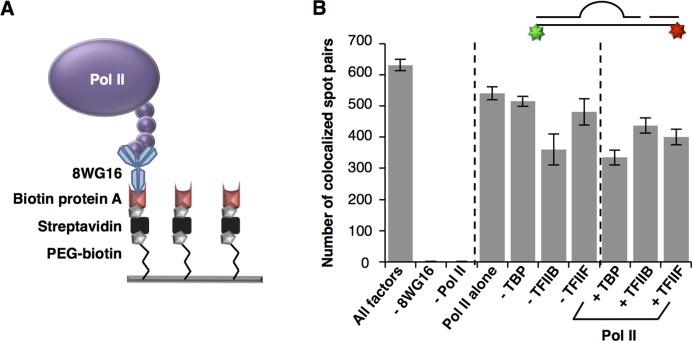
Although we were able to effectively measure template usage, this approach did not allow us to determine whether DNA molecules that were not transcribed still formed complexes containing Pol II, or if these DNAs were simply not bound by the polymerase. To begin to investigate this, we immobilized Pol II on the surface and visualized fluorescent DNA bound to the surface in the presence of the GTFs. We did so by immobilizing biotinylated protein A on a streptavidin-coated surface and then adding an antibody (8WG16) that binds the C-terminal domain of Pol II (Figure 2A). We subsequently flowed in TBP, TFIIB, TFIIF and a three-piece DNA construct lacking the quencher molecule and lacking biotin, (Figure 2B). We determined the number of captured DNA molecules per imaged region by counting the number of colocalized spot pairs on the surface. As seen in the plot in Figure 2B (left bar) on average, hundreds of DNA molecules were captured in each imaged region in the presence of all factors. Importantly, this was highly dependent on both the antibody and Pol II; leaving these components out resulted in almost no colocalized spot pairs.
Figure 2.
Pol II binding to heteroduplex DNA is not highly dependent upon GTFs. (A) Schematic of the surface used to immobilize Pol II on a glass coverslip. (B) The level of binding heteroduplex DNA by surface-immobilized Pol II did not significantly change with addition of TBP, TFIIB or TFIIF. Each GTF was individually omitted from reactions, and individually added to Pol II, as indicated. Binding events were measured as the number of colocalized (Cy3 and Cy5) spot pairs on the surface. The bars represent the average of five separate movies and errors bars represent one standard deviation. The average number of spot pairs in the absence of 8WG16 (1.2 ± 0.6) and Pol II (0.8 ± 0.7) are not visible on this scale.
We used this assay to investigate how the presence of different GTFs impacts the number of DNA molecules bound to immobilized Pol II (Figure 2B). Our data show that the heteroduplex template bound to immobilized Pol II in the absence of GTFs nearly as efficiently as when all the GTFs were present (compare the Pol II alone bar to the All factors bar). We also individually omitted each GTF, and observed only small differences in the number of colocalized spot pairs. Moreover, the number of DNA/Pol II binding events did not vary greatly when each GTF was individually added to Pol II (right three bars). These data are consistent with previous studies showing that Pol II can bind heteroduplex templates in the absence of GTFs (8,9).
Our single molecule binding data indicate that GTFs are not required to observe association of heteroduplex DNA with Pol II. This raised the question of whether the complexes that formed under each condition were transcriptionally active. To test this we performed experiments using immobilized Pol II and the three-piece DNA template containing a quencher molecule on the nontemplate strand, which enables us to monitor transcriptional activity (see the construct in Figure 1B). We flowed GTFs and DNA onto surface-immobilized Pol II and imaged the bound DNA before and after addition of NTPs. As shown in the left two bars in Figure 3, prior to NTP addition, 2.4% of green spots had a colocalized red spot (gray bar), and after NTP addition this number increased to 17.5% (black bar). This confirmed that immobilized Pol II retains transcriptional activity, and moreover, in the presence of all the GTFs, template usage on immobilized Pol II (15.1%) was found to be higher than from immobilized DNA (9.3%).
Figure 3.
Transcriptional activity at the single molecule level is dependent upon GTFs. Pol II was immobilized and the three-piece quenched construct was used. The percentage of Cy3 molecules that colocalized with Cy5 was determined before and after the addition of NTPs (gray versus black bars). Each GTF was individually omitted from reactions, and individually added to Pol II, as indicated. The bars represent the average of five separate movies and errors bars represent one standard deviation.
To understand whether GTFs were required for transcriptional activity at the single molecule level, we performed transcription assays with immobilized Pol II under conditions lacking one or more of the GTFs. The percentage of green spots that colocalized with a red spot was determined before and after addition of NTPs, as indicated by the gray and black bars, respectively (Figure 3). Transcription by Pol II alone was not detected over the minus NTP background. Removal of either TBP or TFIIB also resulted in no transcriptional activity. Removal of TFIIF caused approximately a 2-fold decrease in template usage, indicating that TFIIF facilitates transcription under these conditions, but is not absolutely required. Adding individual GTFs to Pol II and DNA did not recover transcription (right three bars), although addition of TFIIF caused a small increase. This small increase could be due to TFIIF enhancing the ability of Pol II to initiate at the start site; however, it is also possible that TFIIF facilitates Pol II binding at the nontemplate strand nick, allowing transcription from this position to displace the quencher. The data in Figure 3, in conjunction with the binding data in Figure 2, suggest that Pol II/heteroduplex DNA interactions form in a manner that is independent of GTFs; however, the activity of the complexes that form is highly dependent on TBP and TFIIB, and partially dependent on TFIIF.
TFIIF can activate partially assembled complexes whereas TFIIB can only do so in the absence of TFIIF
The single molecule system allows us to wash the slide surface between protein additions, providing a means to determine how different pathways of PIC formation influence transcriptional activity. We asked whether adding TFIIF or TFIIB to partially assembled complexes would result in full transcriptional activity. We first incubated TBP, TFIIB and DNA with surface-immobilized Pol II, and subsequently washed free protein and DNA from the chamber. We then flowed in TFIIF and allowed it to incubate on the slide. Finally, we washed out unbound TFIIF, and imaged the slide before and after the addition of NTPs. We found that 13.5% of templates in the sample were transcribed (Figure 4A, left two bars). As a point of comparison, data from Figure 3 in which all factors were incubated together or when TFIIF was omitted are re-plotted here (right four bars). These data indicate that Pol II can form stable complexes with TBP/TFIIB/DNA, and that TFIIF can efficiently incorporate into and activate these complexes.
Figure 4.
The order of assembly of PICs affects their activity. (A) TFIIF can incorporate into complexes last and yield transcriptionally active PICs. The bars labeled ‘All factors’ and ‘-TFIIF’ were taken from Figure 3, as a point of comparison. The bars represent the average of five separate movies and errors bars represent one standard deviation. (B) TFIIB is unable to incorporate into and activate complexes containing immobilized Pol II/TFIIF/TBP/DNA. Bars labeled ‘All factors’ and ‘-TFIIB’ were taken from Figure 3, as a point of comparison. The bars represent the average of five separate movies and errors bars represent one standard deviation. (C) TFIIB can activate Pol II/TBP/DNA complexes in the absence of TFIIF. Bars labeled ‘-TFIIF’ and ‘-TFIIF, -TFIIB’ were taken from Figure 3. The bars represent the average of five separate movies and errors bars represent one standard deviation.
We next tested whether TFIIB was similarly capable of incorporating into pre-assembled partial complexes. We flowed TBP, TFIIF and DNA onto a Pol II-coated surface and allowed complexes to form. The chamber was then washed extensively prior to adding TFIIB. Unbound TFIIB was washed out of the chamber, and the surface was imaged before and after the addition of NTPs to initiate transcription. Only 2.6% of templates were transcriptionally active (Figure 4B, left two bars). This is similar to the level of activity observed in the absence of TFIIB (right two bars) and far less than the activity observed when all the factors assemble together (middle two bars). These data suggest that TFIIB is not able to build into and activate partial Pol II/TBP/TFIIF/DNA complexes, although it is also possible that stable partial complexes did not form prior to the addition of TFIIB. When considered together, the data in Figure 4A and B suggest that binding of TFIIF prior to TFIIB could occlude key contacts between TFIIB and the rest of the complex that are required for the formation of active PICs. To further probe this, we omitted TFIIF and again asked whether TFIIB could activate partial complexes. We formed a Pol II/TBP/DNA complex on the surface, washed out excess protein and DNA, and then added TFIIB to the slide. We then washed out excess TFIIB, added NTPs and allowed transcription to proceed. We observed that in the absence of TFIIF, TFIIB could activate partial Pol II/TBP/DNA complexes (Figure 4C). Hence, TFIIB can incorporate into and activate partially assembled complexes in the absence of TFIIF, but cannot do so after TFIIF has assembled into partial complexes.
Subcomplexes of Pol II and GTFs remain stably associated in the absence of a DNA template
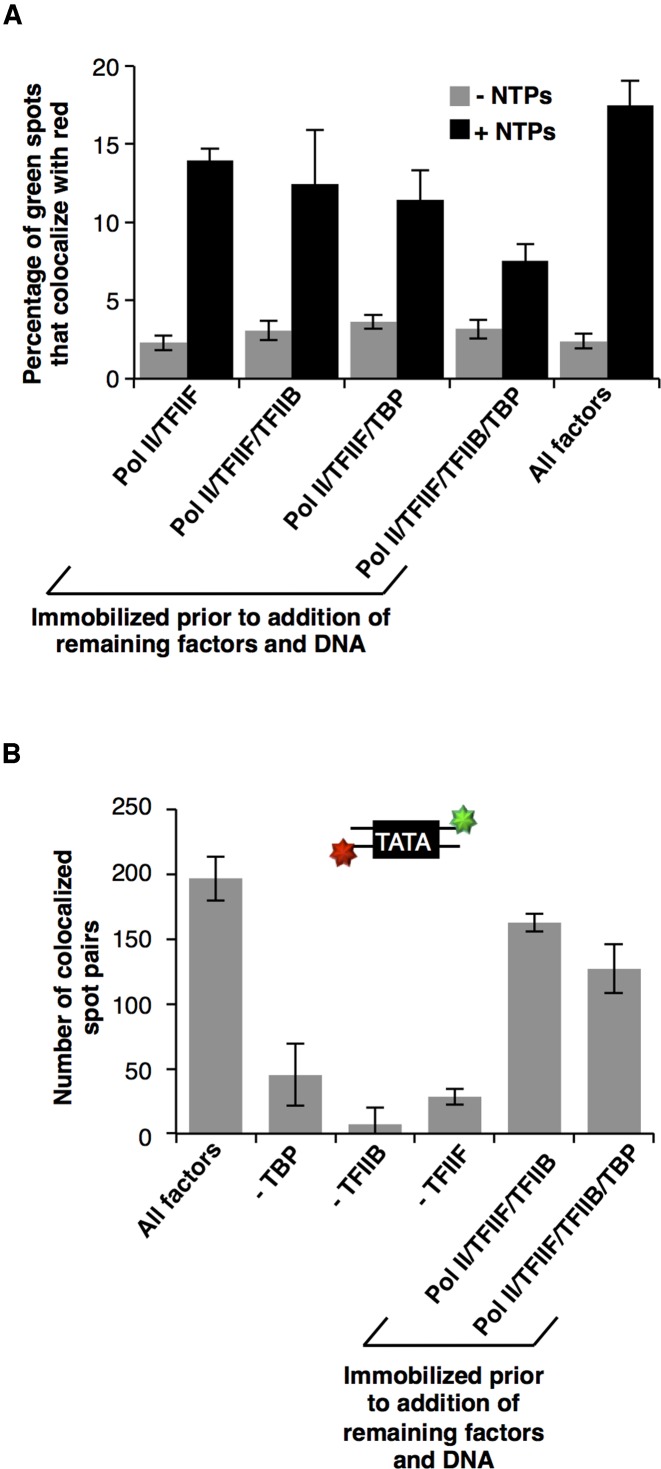
We next tested whether different combinations of GTFs could stably associate with immobilized Pol II in the absence of DNA, then subsequently bind a promoter and remaining GTFs to generate active complexes. We began with Pol II and TFIIF, which we anticipated would stably assemble off of DNA (35). TFIIF was incubated with surface-immobilized Pol II, and the slide was thoroughly washed before addition of TBP, TFIIB and DNA. After addition of NTPs, analysis of colocalized dyes revealed that 11.6% of templates were transcribed (Figure 5A, left two bars), which is comparable to that observed when all factors and DNA were incubated together (15.1%, right two bars). We next asked if stable Pol II/TFIIF/TFIIB ternary complexes could assemble on the surface of the slide and subsequently bind TBP and DNA. Upon addition of NTPs, we found that 9.4% of templates were transcribed (Figure 5A), indicating that both TFIIB and TFIIF remained stably bound to Pol II to allow active transcription complexes to form upon addition of TBP and DNA.
Figure 5.
Combinations of GTFs and Pol II form stable complexes in the absence of DNA. (A) The indicated complexes of Pol II and GTFs were pre-assembled on the surface. NTP-dependent transcriptional activity was evaluated after the remaining GTFs and promoter DNA were added. The bar labeled ‘All factors’ was taken from Figure 3, as a point of comparison. The bars represent the average of five separate movies and errors bars represent one standard deviation. (B) Pol II, TFIIB, TFIIF and TBP are capable of forming a stable complex in the absence of DNA. The schematic shows the 18 bp Alexa555/Alexa647 labeled piece of TATA DNA used to probe the presence of TBP in complexes assembled on surface-immobilized Pol II. With the exception of bar 2 (-TBP) and bar 6, TBP and DNA were pre-incubated prior to addition to the slide. The bars represent the average of five separate movies and errors bars represent one standard deviation.
To determine whether TBP could establish stable contacts with a Pol II/TFIIF complex in the absence of TFIIB and DNA, we prebound TFIIF to immobilized Pol II, washed the slide thoroughly and then added TBP. We then washed out excess TBP and flowed in TFIIB and DNA. We found that after the addition of NTPs, 7.9% of the templates were transcribed, similar to the level of activity observed with other preformed complexes. This indicates that TBP can form stable contacts with Pol II/TFIIF that allow active PICs to form upon addition of TFIIB and DNA.
Finally, we tested whether TBP could build into the stable Pol II/TFIIF/TFIIB complexes to form quaternary complexes capable of recruiting DNA and becoming transcriptionally active. We formed Pol II/TFIIF/TFIIB complexes as described above, washed the chamber and then added TBP. We then washed excess TBP out of the flow chamber and added DNA. Upon addition of NTPs, 4.4% of templates were transcribed (Figure 5A). Although there was a reduction in activity when reactions were assembled in this manner, transcriptional activity was still observed at approximately half the level of the other conditions in Figure 5A.
The lower activity when DNA was added to quaternary protein complexes could be attributable to TBP not stably binding all the Pol II/TFIIB/TFIIF complexes, or the DNA construct not assembling into the quaternary complex in a transcriptionally active conformation. To determine whether TBP is able to stably associate with Pol II/TFIIB/TFIIF complexes, we performed a series of binding experiments in which we used an 18 bp fluorescent DNA construct containing a consensus TATA box to mark the presence of TBP in immobilized complexes. The 18 bp TATA construct was labeled on each 5′ end with Alexa555 or Alexa647; therefore, binding could be assessed via colocalization on the Pol II-immobilized surface. As seen in the first four bars of Figure 5B, the TATA DNA associated with surface-immobilized Pol II in a manner that depended on all of the GTFs. Given that the TATA DNA is a tag for TBP rather than an assembly platform, the loss of colocalized spot pairs when TFIIB or TFIIF was omitted indicates that TBP requires TFIIB and TFIIF to stably associate with the immobilized Pol II. We also formed a Pol II/TFIIB/TFIIF ternary complex on the surface, washed the slide, and then added TBP and the TATA DNA. As seen in Figure 5B, this yielded a comparable number binding events to when all GTFs were added simultaneously.
Finally, we wanted to determine if TBP could remain stably bound to Pol II/TFIIB/TFIIF in the absence of TATA DNA. To do so, we incubated TBP with the preformed ternary complex on the surface, washed out excess TBP, and then added the TATA DNA to the sample. We found that the DNA was able to efficiently bind the surface, albeit at a slightly reduced level compared to assembly of the complex in the presence of the TATA DNA. These results indicate that the network of protein–protein interactions present in the Pol II/TFIIB/TFIIF/TBP quaternary complex is stable to washing and the conformation allows TATA DNA to build into the complex.
DISCUSSION
We developed a single molecule TIRF microscopy system to visualize transcriptional activity by human Pol II at individual template molecules, as well as monitor interactions between Pol II and promoter DNA. We found that interactions between Pol II and a heteroduplex DNA template were largely independent of the GTFs; however, transcriptional activity was strongly dependent on the presence of the GTFs. The single molecule system enabled us to ask whether pre-assembling different subsets of PIC components changed the fraction of DNA templates that were transcribed. We identified several stable subcomplexes that could assemble and subsequently recruit the remaining components to facilitate full transcriptional activity, and others that could not. Moreover, all the minimal system factors (Pol II, TFIIB, TFIIF and TBP) could stably assemble and subsequently bind an 18 bp piece of TATA DNA, suggesting that a promoter scaffold is not absolutely required for GTFs and Pol II to assemble. Together our data document a novel experimental system to investigate Pol II transcriptional activity at the single molecule level, which has provided insight into pathways by which active PICs can assemble.
The single molecule assay provided us the unique ability to monitor both the transcriptional activity of Pol II in the presence and absence of specific GTFs, as well as interactions between surface-immobilized Pol II and the heteroduplex DNA template. We found that transcriptional activity was highly dependent upon GTFs, and in particular TBP and TFIIB (Figure 3); however, Pol II/promoter binding did not strongly depend on the GTFs (Figure 2). The disparity between the dependence on GTFs for assembly versus activity suggests that subcomplexes of Pol II, DNA and GTFs can form, but in conformations that do not necessarily support productive transcription. We observed no transcriptional activity from Pol II alone, which is somewhat surprising given that we and others have reported in ensemble experiments that Pol II alone can transcribe heteroduplex templates, albeit somewhat inefficiently (9,33). It is possible that Pol II largely produced short abortive products from the heteroduplex DNA, and release of the quencher oligo in our system requires transcription to proceed past position +21. The addition of TFIIF to Pol II did allow some transcription to be observed. TFIIF has been shown to facilitate promoter escape (36), hence it is possible that TFIIF helped Pol II molecules transcribe far enough to allow release of the quencher oligo. We cannot distinguish between this model and TFIIF helping Pol II bind at the nontemplate strand nick, allowing transcription from this position to displace the quencher. Although TFIIF was the only GTF that itself facilitated transcriptional activity by Pol II, it was also the only GTF not required for transcription in the presence of the remaining factors (although it was stimulatory). Similar results have been reported on the IgH promoter (5); observing transcription in the absence of TFIIF on the chimeric AdMLP/IL-2 promoter used here could be facilitated by the heteroduplex (8).
We asked whether pre-assembling subsets of the PIC components altered the fraction of DNA templates that were ultimately transcribed. For example, the subcomplex containing Pol II/TBP/TFIIB/DNA assembled and recruited TFIIF to allow levels of transcription equal to that observed when all components were incubated together (Figure 4A). This result is consistent with models for ordered assembly in which TFIIF builds into complexes after TBP and TFIIB (18,37). It is also consistent with structural models of PICs obtained via crosslinking and cryo EM, which indicate that TFIIF interacts with an extended surface of Pol II and regions of the DNA that are unoccupied by other proteins (38–40). We also found that some subcomplexes cannot assemble and support transcriptional activity upon addition of the remaining factors. The preformed Pol II/TBP/TFIIF/DNA complex failed to yield transcriptionally active complexes after the addition of TFIIB (Figure 4B). In the absence of TFIIF, however, the addition of TFIIB to the partial Pol II/TBP/DNA complex did yield transcriptionally active complexes. This suggests that TFIIF occludes, either directly or via an altered conformation, the subsequent assembly of TFIIB into complexes. Structural work shows that a flexible region of TFIIB, termed the B-reader loop, makes specific contacts with Pol II that assist with start site selection, properly position the polymerase and stabilize the nascent RNA during the addition of the first nucleotides (41–43). It is possible that allowing Pol II to establish contacts with TFIIF in the absence of TFIIB resulted in a sub-complex that precluded the necessary contacts between the B-reader and Pol II that are necessary for active transcription.
Interestingly, our data support a model in which the network of protein–protein contacts within a PIC is a significant stabilizing force within the complex. Specifically, we observed that stable ternary Pol II/TFIIF/TFIIB and Pol II/TFIIF/TBP complexes can form, subsequently bind promoter DNA and the missing GTF to support active transcription (Figure 5A). This suggests that the protein–protein interaction interfaces in a PIC are either maintained in these subcomplexes, or exhibit plasticity depending on which factors are present. We also observed that a quaternary Pol II/TFIIF/TFIIB/TBP complex can form and subsequently bind DNA to support transcription, albeit at a reduced level. Moreover, this quaternary complex can also recruit a short TATA box DNA in a manner that depends on all the proteins (Figure 5B). Together these observations contrast with a model of PIC assembly in which the primary stabilizing factor is the promoter DNA acting as a scaffold upon which the PIC is built. These data also provide biochemical support for the model of a Pol II holoenzyme complex having the capacity to assemble off of promoter DNA. In unrelated studies, we found that several noncoding RNAs function as transcriptional repressors by binding Pol II and building into complexes at promoters (44–46). Interestingly, these complexes are inactive because Pol II has lost most of its contacts with promoter DNA, however, the complexes remain intact (47), further suggesting that Pol II/DNA contacts are not necessarily the primary force stabilizing complexes.
Together our single molecule data suggest that active PICs can form from pre-assembled subcomplexes containing Pol II; however, if fully active PICs are to form, there is a specificity to components of the subcomplexes and their order of assembly. Therefore, it is likely that both sequential binding and ‘holoenzyme’ modes of PIC assembly exist, and perhaps different pathways occur at different promoters and may serve as points of regulation. ChIP-chip studies in yeast probing the redistribution of Pol II and GTFs upon cell stress found that many promoters underwent partial PIC assembly, often lacking Pol II and TFIIH, and additional stress converted partial PICs into full PICs (48). The higher resolution ChIP-exo technique has mapped thousands of PICs in yeast, and generally found them compositionally homogeneous with orientations and positioning agreeing well with structural models (49). In addition, ChIP assays and RNA analyses in yeast have revealed that the dynamics with which TBP interacts with DNA can impact assembly of active complexes at promoters (50,51).
The system we developed to observe Pol II transcription at the single molecule level enables us to address unique mechanistic questions about Pol II transcription that are difficult to test using ensemble approaches. These studies complement other single molecule approaches to study Pol II transcription complexes that take advantage of FRET-based nanopositioning to observe structural changes (52–55), optical trapping to observe the movement of Pol II at multiple stages of transcription (56–61) and TIRF to observe transcriptional activity by more complex PICs (25). Single-molecule studies of transcription provide a powerful means to explore transcriptional mechanisms and together with structural, biochemical and cell-based approaches, will unravel the details of how regulation of Pol II transcription occurs.
Acknowledgments
We thank Andrey Revyakin, Zhengjian Zhang and Robert Tjian for help with studies of Pol II transcription using single molecule TIRF.
FUNDING
National Science Foundation [MCB-1244518]; National Institutes of Health [T32 GM-065103 to A.H.]. Funding for open access charge: National Science Foundation [MCB-1244518].
Conflict of interest statement. None declared.
REFERENCES
- 1.Thomas M.C., Chiang C.M. The general transcription machinery and general cofactors. Crit. Rev. Biochem. Mol. Biol. 2006;41:105–178. doi: 10.1080/10409230600648736. [DOI] [PubMed] [Google Scholar]
- 2.Swygert S.G., Peterson C.L. Chromatin dynamics: interplay between remodeling enzymes and histone modifications. Biochim. Biophys. Acta. 2014;1839:728–736. doi: 10.1016/j.bbagrm.2014.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ma J. Transcriptional activators and activation mechanisms. Protein Cell. 2011;2:879–888. doi: 10.1007/s13238-011-1101-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Parvin J.D., Timmers H.T.M., Sharp P.A. Promoter specificity of basal transcription factors. Cell. 1992;68:1135–1144. doi: 10.1016/0092-8674(92)90084-p. [DOI] [PubMed] [Google Scholar]
- 5.Parvin J.D., Sharp P.A. DNA topology and a minimal set of basal factors for transcription by RNA polymerase II. Cell. 1993;73:533–540. doi: 10.1016/0092-8674(93)90140-l. [DOI] [PubMed] [Google Scholar]
- 6.Tyree C.M., George C.P., Lira-DeVito L.M., Wampler S.L., Dahmus M.E., Zawel L., Kadonaga J.T. Identification of a minimal set of proteins that is sufficient for accurate initiation of transcription by RNA polymerase II. Genes Dev. 1993;7:1254–1265. doi: 10.1101/gad.7.7a.1254. [DOI] [PubMed] [Google Scholar]
- 7.Goodrich J.A., Tjian R. Transcription factors IIE and IIH and ATP hydrolysis direct promoter clearance by RNA polymerase II. Cell. 1994;77:145–156. doi: 10.1016/0092-8674(94)90242-9. [DOI] [PubMed] [Google Scholar]
- 8.Pan G., Greenblatt J. Initiation of transcription by RNA polymerase II is limited by melting of the promoter DNA in the region immediately upstream of the initiation site. J. Biol. Chem. 1994;269:30101–30104. [PubMed] [Google Scholar]
- 9.Keene R.G., Luse D.S. Initially transcribed sequences strongly affect the extent of abortive initiation by RNA polymerase II. J. Biol. Chem. 1999;274:11526–11534. doi: 10.1074/jbc.274.17.11526. [DOI] [PubMed] [Google Scholar]
- 10.Luse D.S. The RNA polymerase II preinitiation complex: Through what pathway is the complex assembled. Transcription. 2013;5:e27050. doi: 10.4161/trns.27050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Orphanides G., Lagrange T., Reinberg D. The general transcription factors of RNA polymerase II. Genes Dev. 1996;10:2657–2683. doi: 10.1101/gad.10.21.2657. [DOI] [PubMed] [Google Scholar]
- 12.Roeder R.G. The role of general initiation factors in transcription by RNA polymerase II. Trends Biochem. Sci. 1996;21:327–335. [PubMed] [Google Scholar]
- 13.Koleske A.J., Young R.A. The RNA polymerase II holoenzyme and its implications for gene regulation. Trends Biochem. Sci. 1995;20:113–116. doi: 10.1016/s0968-0004(00)88977-x. [DOI] [PubMed] [Google Scholar]
- 14.Koleske A.J., Young R.A. An RNA polymerase II holoenzyme responsive to activators. Nature. 1994;368:466–469. doi: 10.1038/368466a0. [DOI] [PubMed] [Google Scholar]
- 15.Kim Y.J., Bjorklund S., Li Y., Sayre M.H., Kornberg R.D. A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell. 1994;77:599–608. doi: 10.1016/0092-8674(94)90221-6. [DOI] [PubMed] [Google Scholar]
- 16.Fire A., Samuels M., Sharp P.A. Interactions between RNA polymerase II, factors, and template leading to accurate transcription. J. Biol. Chem. 1984;259:2509–2516. [PubMed] [Google Scholar]
- 17.Hawley D.K., Roeder R.G. Separation and partial characterization of three functional steps in transcription initiation by human RNA polymerase II. J. Biol. Chem. 1985;260:8163–8172. [PubMed] [Google Scholar]
- 18.Buratowski S., Hahn S., Guarente L., Sharp P.A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell. 1989;56:549–561. doi: 10.1016/0092-8674(89)90578-3. [DOI] [PubMed] [Google Scholar]
- 19.Ha I., Roberts S., Maldonada E., Sun X., Kim L.-U., Green M., Reinberg D. Multiple functional domains of human transcription factor IIB: distinct interactions with two general transcription factors and RNA polymerase II. Genes Dev. 1993;7:1021–1032. doi: 10.1101/gad.7.6.1021. [DOI] [PubMed] [Google Scholar]
- 20.Ossipow V., Tassan J.-P., Nigg E.A., Schibler U. A mammalian RNA polymerase II holoenzyme containing all components required for promoter-specific transcription initiation. Cell. 1995;83:137–146. doi: 10.1016/0092-8674(95)90242-2. [DOI] [PubMed] [Google Scholar]
- 21.Maldonado E., Ramin S., Sheldon M., Cho H., Drapkin R., Rickert P., Lees E., Anderson C.W., Linn S., Reinberg D. A human RNA polymerase II complex associated with SRB and DNA-repair proteins. Nature. 1996;381:86–89. doi: 10.1038/381086a0. [DOI] [PubMed] [Google Scholar]
- 22.Kugel J.F., Goodrich J.A. Promoter escape limits the rate of transcription from the adenovirus major late promoter on negatively supercoiled templates. Proc. Natl. Acad. Sci. U.S.A. 1998;95:9232–9237. doi: 10.1073/pnas.95.16.9232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kamakaka R.T., Tyree C.M., Kadonaga J.T. Accurate and efficient RNA polymerase II transcription with a soluble nuclear fraction derived from Drosophila embryos. Proc. Natl. Acad. Sci. U.S.A. 1991;88:1024–1028. doi: 10.1073/pnas.88.3.1024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Juven-Gershon T., Cheng S., Kadonaga J.T. Rational design of a super core promoter that enhances gene expression. Nat. Methods. 2006;3:917–922. doi: 10.1038/nmeth937. [DOI] [PubMed] [Google Scholar]
- 25.Revyakin A., Zhang Z., Coleman R.A., Li Y., Inouye C., Lucas J.K., Park S.R., Chu S., Tjian R. Transcription initiation by human RNA polymerase II visualized at single-molecule resolution. Genes Dev. 2012;26:1691–1702. doi: 10.1101/gad.194936.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Darzacq X., Shav-Tal Y., de Turris V., Brody Y., Shenoy S.M., Phair R.D., Singer R.H. In vivo dynamics of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 2007;14:796–806. doi: 10.1038/nsmb1280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Weaver J.R., Kugel J.F., Goodrich J.A. The sequence at specific positions in the early transcribed region sets the rate of transcript synthesis by RNA polymerase II in vitro. J. Biol. Chem. 2005;280:39860–39869. doi: 10.1074/jbc.M509376200. [DOI] [PubMed] [Google Scholar]
- 28.Ha I., Lane W.S., Reinberg D. Cloning of a human gene encoding the general transcription factor IIB. Nature. 1991;352:689–695. doi: 10.1038/352689a0. [DOI] [PubMed] [Google Scholar]
- 29.Lu H., Flores O., Weinmann R., Reinberg D. The nonphosphorylated form of RNA polymerase II preferentially associates with the preinitiation complex. Proc. Natl. Acad. Sci. U.S.A. 1991;88:10004–10008. doi: 10.1073/pnas.88.22.10004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Blair R.H., Goodrich J.A., Kugel J.F. Single-molecule fluorescence resonance energy transfer shows uniformity in TATA binding protein-induced DNA bending and heterogeneity in bending kinetics. Biochemistry. 2012;51:7444–7455. doi: 10.1021/bi300491j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pal M., Luse D.S. Strong natural pausing by RNA polymerase II within 10 bases of transcription start may result in repeated slippage and reextension of the nascent RNA. Mol. Cell. Biol. 2002;22:30–40. doi: 10.1128/MCB.22.1.30-40.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Pal M., Luse D.S. The initiation-elongation transition: lateral mobility of RNA in RNA polymerase II complexes is greatly reduced at +8/+9 and absent by +23. Proc. Natl. Acad. Sci. U.S.A. 2003;100:5700–5705. doi: 10.1073/pnas.1037057100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gilman B., Drullinger L.F., Kugel J.F., Goodrich J.A. TATA-binding protein and transcription factor IIB induce transcript slipping during early transcription by RNA polymerase II. J. Biol. Chem. 2009;284:9093–9098. doi: 10.1074/jbc.M900019200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ferguson H.A., Kugel J.F., Goodrich J.A. Kinetic and mechanistic analysis of the RNA polymerase II transcription reaction at the human interleukin-2 promoter. J. Mol. Biol. 2001;314:993–1006. doi: 10.1006/jmbi.2000.5215. [DOI] [PubMed] [Google Scholar]
- 35.Sopta M., Carthew R.W., Greenblatt J. Isolation of three proteins that bind to mammalian RNA polymerase II. J. Biol. Chem. 1985;260:10353–10360. [PubMed] [Google Scholar]
- 36.Yan Q., Moreland R.J., Conaway J.W., Conaway R.C. Dual roles for transcription factor IIF in promoter escape by RNA polymerase II. J. Biol. Chem. 1999;274:35668–35675. doi: 10.1074/jbc.274.50.35668. [DOI] [PubMed] [Google Scholar]
- 37.Flores O., Lu H., Killeen M., Greenblatt J., Burton Z.F., Reinberg D. The small subunit of transcription factor IIF recruits RNA polymerase II into the preinitiation complex. Proc. Natl. Acad. Sci. U.S.A. 1991;88:9999–10003. doi: 10.1073/pnas.88.22.9999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.He Y., Fang J., Taatjes D.J., Nogales E. Structural visualization of key steps in human transcription initiation. Nature. 2013;495:481–486. doi: 10.1038/nature11991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chen H.T., Warfield L., Hahn S. The positions of TFIIF and TFIIE in the RNA polymerase II transcription preinitiation complex. Nat. Struct. Mol. Biol. 2007;14:696–703. doi: 10.1038/nsmb1272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Gaiser F., Tan S., Richmond T.J. Novel dimerization fold of RAP30/RAP74 in human TFIIF at 1.7 A resolution. J. Mol. Biol. 2000;302:1119–1127. doi: 10.1006/jmbi.2000.4110. [DOI] [PubMed] [Google Scholar]
- 41.Bushnell D.A., Westover K.D., Davis R.E., Kornberg R.D. Structural basis of transcription: an RNA polymerase II-TFIIB cocrystal at 4.5 Angstroms. Science. 2004;303:983–988. doi: 10.1126/science.1090838. [DOI] [PubMed] [Google Scholar]
- 42.Sainsbury S., Niesser J., Cramer P. Structure and function of the initially transcribing RNA polymerase II-TFIIB complex. Nature. 2013;493:437–440. doi: 10.1038/nature11715. [DOI] [PubMed] [Google Scholar]
- 43.Kostrewa D., Zeller M.E., Armache K.J., Seizl M., Leike K., Thomm M., Cramer P. RNA polymerase II-TFIIB structure and mechanism of transcription initiation. Nature. 2009;462:323–330. doi: 10.1038/nature08548. [DOI] [PubMed] [Google Scholar]
- 44.Espinoza C.A., Allen T.A., Hieb A.R., Kugel J.F., Goodrich J.A. B2 RNA binds directly to RNA polymerase II to repress transcript synthesis. Nat. Struct. Mol. Biol. 2004;11:822–829. doi: 10.1038/nsmb812. [DOI] [PubMed] [Google Scholar]
- 45.Espinoza C.A., Goodrich J.A., Kugel J.F. Characterization of the structure, function, and mechanism of B2 RNA, an ncRNA repressor of RNA polymerase II transcription. RNA. 2007;13:583–596. doi: 10.1261/rna.310307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Mariner P.D., Walters R.D., Espinoza C.A., Drullinger L.F., Wagner S.D., Kugel J.F., Goodrich J.A. Human Alu RNA is a modular transacting repressor of mRNA transcription during heat shock. Mol. Cell. 2008;29:499–509. doi: 10.1016/j.molcel.2007.12.013. [DOI] [PubMed] [Google Scholar]
- 47.Yakovchuk P., Goodrich J.A., Kugel J.F. B2 RNA and Alu RNA repress transcription by disrupting contacts between RNA polymerase II and promoter DNA within assembled complexes. Proc. Natl. Acad. Sci. U.S.A. 2009;106:5569–5574. doi: 10.1073/pnas.0810738106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zanton S.J., Pugh B.F. Full and partial genome-wide assembly and disassembly of the yeast transcription machinery in response to heat shock. Genes Dev. 2006;20:2250–2265. doi: 10.1101/gad.1437506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rhee H.S., Pugh B.F. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature. 2012;483:295–301. doi: 10.1038/nature10799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Poorey K., Sprouse R.O., Wells M.N., Viswanathan R., Bekiranov S., Auble D.T. RNA synthesis precision is regulated by preinitiation complex turnover. Genome Res. 2010;20:1679–1688. doi: 10.1101/gr.109504.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Poorey K., Viswanathan R., Carver M.N., Karpova T.S., Cirimotich S.M., McNally J.G., Bekiranov S., Auble D.T. Measuring chromatin interaction dynamics on the second time scale at single-copy genes. Science. 2013;342:369–372. doi: 10.1126/science.1242369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Muschielok A., Andrecka J., Jawhari A., Bruckner F., Cramer P., Michaelis J. A nano-positioning system for macromolecular structural analysis. Nat. Methods. 2008;5:965–971. doi: 10.1038/nmeth.1259. [DOI] [PubMed] [Google Scholar]
- 53.Treutlein B., Muschielok A., Andrecka J., Jawhari A., Buchen C., Kostrewa D., Hog F., Cramer P., Michaelis J. Dynamic architecture of a minimal RNA polymerase II open promoter complex. Mol. Cell. 2012;46:136–146. doi: 10.1016/j.molcel.2012.02.008. [DOI] [PubMed] [Google Scholar]
- 54.Andrecka J., Treutlein B., Arcusa M.A., Muschielok A., Lewis R., Cheung A.C., Cramer P., Michaelis J. Nano positioning system reveals the course of upstream and nontemplate DNA within the RNA polymerase II elongation complex. Nucleic Acids Res. 2009;37:5803–5809. doi: 10.1093/nar/gkp601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Andrecka J., Lewis R., Bruckner F., Lehmann E., Cramer P., Michaelis J. Single-molecule tracking of mRNA exiting from RNA polymerase II. Proc. Natl. Acad. Sci. U.S.A. 2008;105:135–140. doi: 10.1073/pnas.0703815105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hodges C., Bintu L., Lubkowska L., Kashlev M., Bustamante C. Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II. Science. 2009;325:626–628. doi: 10.1126/science.1172926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Palangat M., Larson M.H., Hu X., Gnatt A., Block S.M., Landick R. Efficient reconstitution of transcription elongation complexes for single-molecule studies of eukaryotic RNA polymerase II. Transcription. 2012;3:146–153. doi: 10.4161/trns.20269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Larson M.H., Zhou J., Kaplan C.D., Palangat M., Kornberg R.D., Landick R., Block S.M. Trigger loop dynamics mediate the balance between the transcriptional fidelity and speed of RNA polymerase II. Proc. Natl. Acad. Sci. U.S.A. 2012;109:6555–6560. doi: 10.1073/pnas.1200939109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Galburt E.A., Grill S.W., Wiedmann A., Lubkowska L., Choy J., Nogales E., Kashlev M., Bustamante C. Backtracking determines the force sensitivity of RNAP II in a factor-dependent manner. Nature. 2007;446:820–823. doi: 10.1038/nature05701. [DOI] [PubMed] [Google Scholar]
- 60.Zamft B., Bintu L., Ishibashi T., Bustamante C. Nascent RNA structure modulates the transcriptional dynamics of RNA polymerases. Proc. Natl. Acad. Sci. U.S.A. 2012;109:8948–8953. doi: 10.1073/pnas.1205063109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Fazal F.M., Meng C.A., Murakami K., Kornberg R.D., Block S.M. Real-time observation of the initiation of RNA polymerase II transcription. Nature. 2015;525:274–277. doi: 10.1038/nature14882. [DOI] [PMC free article] [PubMed] [Google Scholar]