Abstract
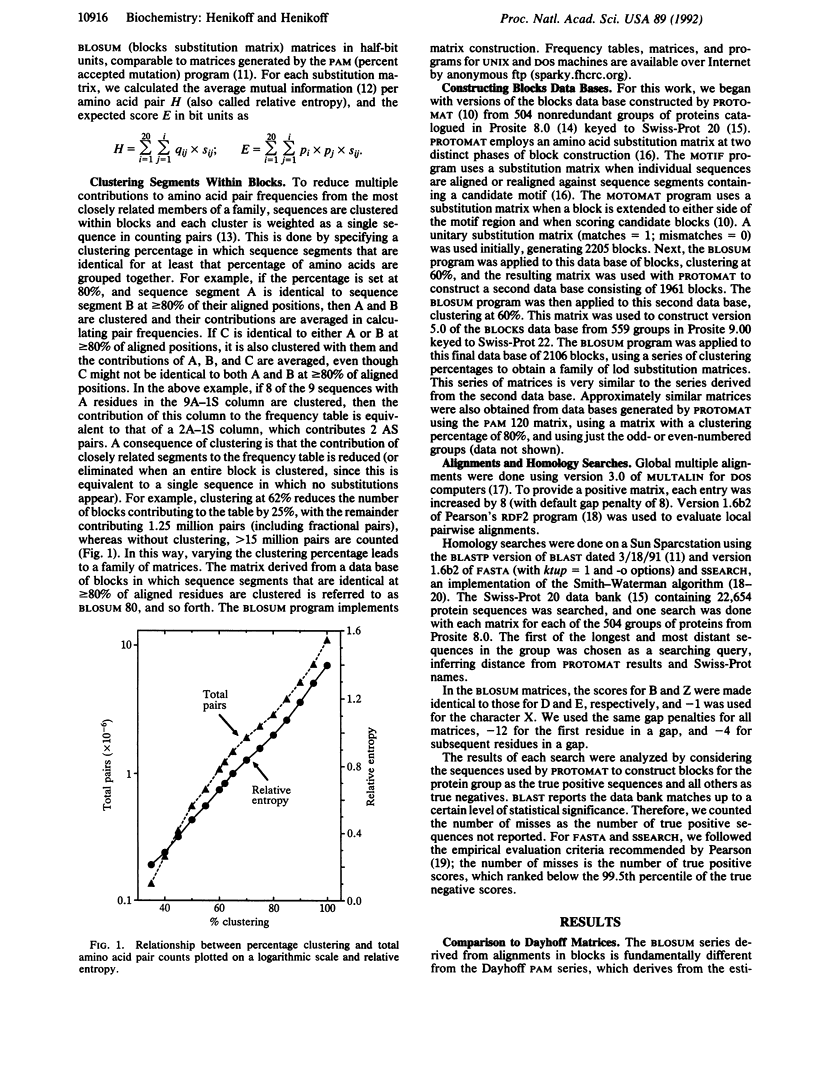
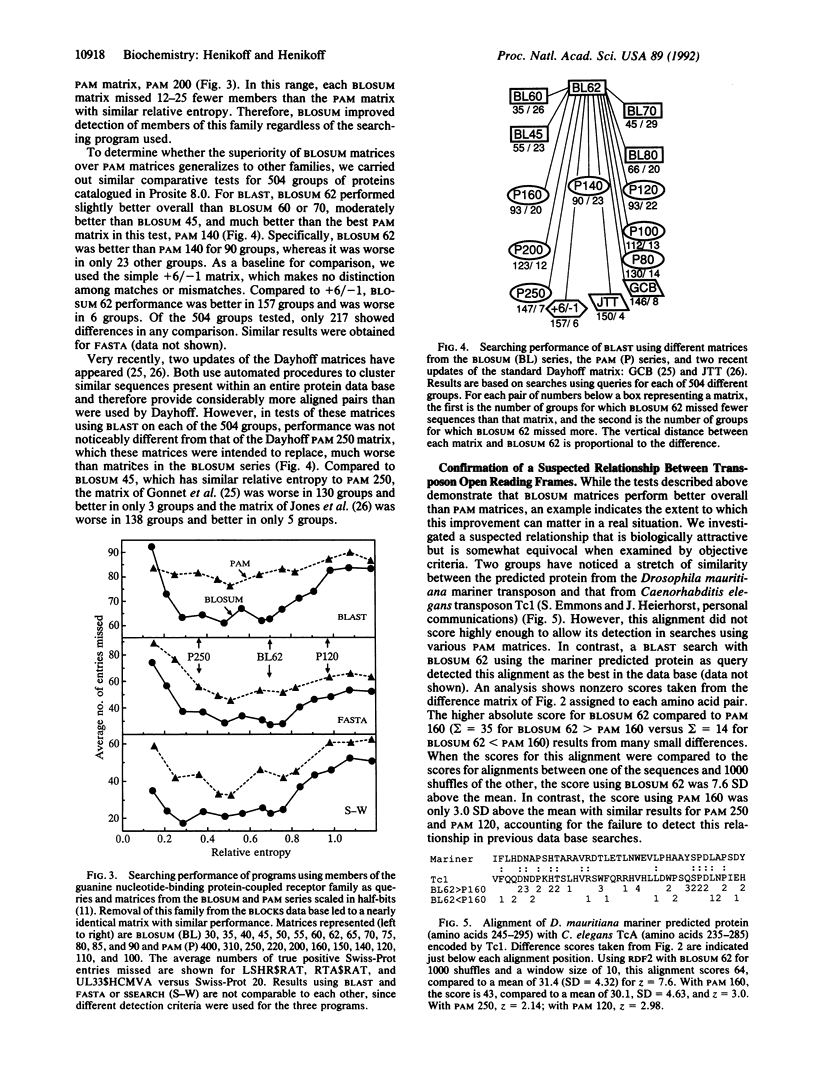
Methods for alignment of protein sequences typically measure similarity by using a substitution matrix with scores for all possible exchanges of one amino acid with another. The most widely used matrices are based on the Dayhoff model of evolutionary rates. Using a different approach, we have derived substitution matrices from about 2000 blocks of aligned sequence segments characterizing more than 500 groups of related proteins. This led to marked improvements in alignments and in searches using queries from each of the groups.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altschul S. F. Amino acid substitution matrices from an information theoretic perspective. J Mol Biol. 1991 Jun 5;219(3):555–565. doi: 10.1016/0022-2836(91)90193-A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Attwood T. K., Eliopoulos E. E., Findlay J. B. Multiple sequence alignment of protein families showing low sequence homology: a methodological approach using database pattern-matching discriminators for G-protein-linked receptors. Gene. 1991 Feb 15;98(2):153–159. doi: 10.1016/0378-1119(91)90168-b. [DOI] [PubMed] [Google Scholar]
- Bairoch A., Boeckmann B. The SWISS-PROT protein sequence data bank. Nucleic Acids Res. 1991 Apr 25;19 (Suppl):2247–2249. doi: 10.1093/nar/19.suppl.2247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bairoch A. PROSITE: a dictionary of sites and patterns in proteins. Nucleic Acids Res. 1991 Apr 25;19 (Suppl):2241–2245. doi: 10.1093/nar/19.suppl.2241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corpet F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988 Nov 25;16(22):10881–10890. doi: 10.1093/nar/16.22.10881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doolittle R. F. Searching through sequence databases. Methods Enzymol. 1990;183:99–110. doi: 10.1016/0076-6879(90)83008-w. [DOI] [PubMed] [Google Scholar]
- Feng D. F., Johnson M. S., Doolittle R. F. Aligning amino acid sequences: comparison of commonly used methods. J Mol Evol. 1984;21(2):112–125. doi: 10.1007/BF02100085. [DOI] [PubMed] [Google Scholar]
- George D. G., Barker W. C., Hunt L. T. Mutation data matrix and its uses. Methods Enzymol. 1990;183:333–351. doi: 10.1016/0076-6879(90)83022-2. [DOI] [PubMed] [Google Scholar]
- Gonnet G. H., Cohen M. A., Benner S. A. Exhaustive matching of the entire protein sequence database. Science. 1992 Jun 5;256(5062):1443–1445. doi: 10.1126/science.1604319. [DOI] [PubMed] [Google Scholar]
- Greer J. Comparative model-building of the mammalian serine proteases. J Mol Biol. 1981 Dec 25;153(4):1027–1042. doi: 10.1016/0022-2836(81)90465-4. [DOI] [PubMed] [Google Scholar]
- Henikoff S., Henikoff J. G. Automated assembly of protein blocks for database searching. Nucleic Acids Res. 1991 Dec 11;19(23):6565–6572. doi: 10.1093/nar/19.23.6565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S., Wallace J. C., Brown J. P. Finding protein similarities with nucleotide sequence databases. Methods Enzymol. 1990;183:111–132. doi: 10.1016/0076-6879(90)83009-x. [DOI] [PubMed] [Google Scholar]
- Jones D. T., Taylor W. R., Thornton J. M. The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci. 1992 Jun;8(3):275–282. doi: 10.1093/bioinformatics/8.3.275. [DOI] [PubMed] [Google Scholar]
- Lipman D. J., Altschul S. F., Kececioglu J. D. A tool for multiple sequence alignment. Proc Natl Acad Sci U S A. 1989 Jun;86(12):4412–4415. doi: 10.1073/pnas.86.12.4412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McLachlan A. D. Tests for comparing related amino-acid sequences. Cytochrome c and cytochrome c 551 . J Mol Biol. 1971 Oct 28;61(2):409–424. doi: 10.1016/0022-2836(71)90390-1. [DOI] [PubMed] [Google Scholar]
- Mohana Rao J. K. New scoring matrix for amino acid residue exchanges based on residue characteristic physical parameters. Int J Pept Protein Res. 1987 Feb;29(2):276–281. doi: 10.1111/j.1399-3011.1987.tb02254.x. [DOI] [PubMed] [Google Scholar]
- Pearson W. R. Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol. 1990;183:63–98. doi: 10.1016/0076-6879(90)83007-v. [DOI] [PubMed] [Google Scholar]
- Pearson W. R. Searching protein sequence libraries: comparison of the sensitivity and selectivity of the Smith-Waterman and FASTA algorithms. Genomics. 1991 Nov;11(3):635–650. doi: 10.1016/0888-7543(91)90071-l. [DOI] [PubMed] [Google Scholar]
- Risler J. L., Delorme M. O., Delacroix H., Henaut A. Amino acid substitutions in structurally related proteins. A pattern recognition approach. Determination of a new and efficient scoring matrix. J Mol Biol. 1988 Dec 20;204(4):1019–1029. doi: 10.1016/0022-2836(88)90058-7. [DOI] [PubMed] [Google Scholar]
- Smith H. O., Annau T. M., Chandrasegaran S. Finding sequence motifs in groups of functionally related proteins. Proc Natl Acad Sci U S A. 1990 Jan;87(2):826–830. doi: 10.1073/pnas.87.2.826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith R. F., Smith T. F. Automatic generation of primary sequence patterns from sets of related protein sequences. Proc Natl Acad Sci U S A. 1990 Jan;87(1):118–122. doi: 10.1073/pnas.87.1.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith T. F., Waterman M. S. Identification of common molecular subsequences. J Mol Biol. 1981 Mar 25;147(1):195–197. doi: 10.1016/0022-2836(81)90087-5. [DOI] [PubMed] [Google Scholar]


