Abstract
Small regulatory RNAs guide Argonaute (Ago) proteins in a sequence-specific manner to their targets and thereby play important roles in eukaryotic gene silencing1. Of the three small RNA classes, microRNAs and siRNAs are processed from double-stranded precursors into defined 21- to 23-mers by Dicer, an endoribonuclease with intrinsic ruler function. piRNAs—the 22-30 nt long guides for PIWI-clade Ago proteins that silence transposons in animal gonads—are generated Dicer-independently from single-stranded precursors2,3. piRNA 5' ends are defined either by Zucchini, a mitochondria-anchored endonuclease4,5, or by piRNA-guided target cleavage6,7. Formation of piRNA 3' ends is poorly understood. Here, we find that two genetically and mechanistically distinct pathways generate piRNA 3' ends in Drosophila. The initiating nucleases are either Zucchini or the PIWI-clade proteins Aubergine (Aub)/Ago3. While Zucchini-mediated cleavages directly define mature piRNA 3' ends8,9, Aub/Ago3-mediated cleavages liberate pre-piRNAs that require extensive resection by the 3'-to-5' exoribonuclease Nibbler/Mut-710–13. The relative activity of these two pathways dictates the extent to which piRNAs are fueled into cytoplasmic or nuclear PIWI-clade proteins and thereby sets the balance between post-transcriptional and transcriptional silencing. Strikingly, loss of both Zucchini and Nibbler reveals a minimal, Argonaute-driven small RNA biogenesis pathway where piRNA 5' and 3' ends are directly produced by closely spaced Aub/Ago3-mediated cleavage events. Our data establish a coherent blueprint for piRNA biogenesis, and set the stage for the mechanistic dissection of the processes that govern piRNA 3' end formation.
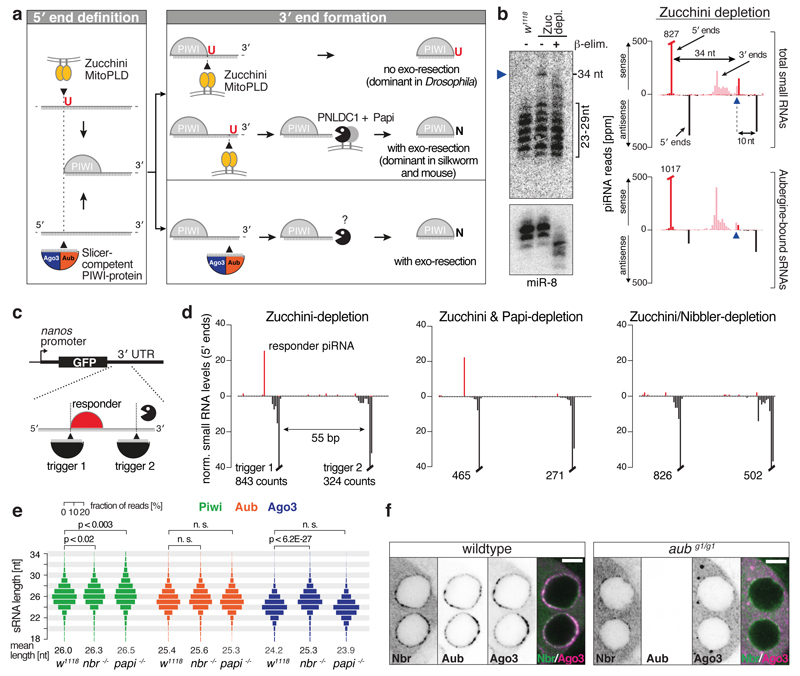
piRNA biogenesis is initiated by endonucleolytic definition of piRNA 5' ends (Fig. 1a). Based on the nuclease involved, this defines primary (processed by Zucchini)4,5 and secondary (processed by Aub/Ago3)6,7 piRNAs. Attributes of these two endo-cleavages are a 5' Uridine signature for primary piRNAs, and a 10-nt 5' sense/antisense offset for secondary piRNAs (ping-pong signature). Following 5' end cleavage, piRNA intermediates are anchored with their 5' ends in PIWI proteins before their 3' ends are matured14,15 (we refer to these as pre-piRNAs). Zucchini also liberates 3' ends of primary and secondary piRNAs8,9. In zucchini mutants, however, Aub/Ago3-bound secondary piRNAs are still abundant16, indicative of alternative 3' end formation pathways (Fig. ED1a). The 3' ends of these piRNAs lack signs for endonucleolytic processing, such as a coupling signature stemming from Zucchini-mediated, phased piRNA biogenesis8,9, or a 3'/5' ping-pong signature indicative of slicer-mediated 3' end formation (Fig. ED1b). This supports the hypothesis that Zucchini-independent piRNA 3' biogenesis involves exonucleolytic resection of pre-piRNAs that have been generated by Aub/Ago3 (Fig. 1a)9.
Figure 1. The 3'-to-5' exonuclease Nibbler matures piRNA 3' ends from slicer-cleaved pre-piRNAs.
a) Schematic illustration of piRNA 5' and 3' biogenesis.
b) Northern blot against individual piRNA 5' species (mature piRNAs: 23-29 nt) detects a 34-nt pre-piRNA (blue arrowhead). 3' end methylation probed by β-elimination (miR-8 serves as control). To the right, sequencing counts of the corresponding piRNAs (normalized to 1 million miRNA reads: ppm) from total small RNAs or from an Aub-IP are shown.
c) Schematic of the dual-site piRNA biogenesis reporter.
d) Levels of small RNAs (ppm) mapping to the biogenesis reporter (5' ends only) in indicated genetic backgrounds.
e) Length profiles of TE-mapping small RNAs isolated from Piwi/Aub/Ago3-immuno-precipitates (genetic background indicated; p-values: two-sided t-test).
f) Confocal images showing localization of GFP-Nibbler, Aub and Ago3 in w1118 or aubergine mutant egg chambers (scale bar: 5 μm; individual channels as inverted gray scale images).
We identified piRNA 5' species (piRNAs with the same 5' end) that—besides piRNAs in the 23-29 nt range—exhibit one abnormally long isoform that extends to the cleavage position of a complementary piRNA (Fig. 1b). These isoforms are also found in libraries from immuno-purified PIWI proteins (Fig. 1b), indicating that they represent Aub/Ago3-loaded pre-piRNAs whose 3' ends have been formed by slicing and await trimming. Consistent with this, the long isoforms lack 2'-O-methylation at their 3' ends (Fig. 1b).
To identify the 3' exonuclease involved we utilized a piRNA biogenesis reporter that recapitulates Zucchini-independent piRNA 3' end formation9; expression of a reporter with two target sites for cellular piRNAs forces the generation of responder piRNAs in Zucchini-depleted ovaries (Fig. 1c,d). We combined this reporter with a double-shRNA expression cassette to co-deplete Zucchini and a gene of interest (Fig. ED2a-c). A strong candidate for the exonuclease is the PARN-like nuclease PNLDC1, which trims pre-piRNAs in silkworm17. As PARN-family nucleases are absent in Drosophila, we tested instead the mitochondria-anchored Tudor/KH-domain protein Papi, an essential PNLDC1 co-factor in silkworm (Fig. ED2d,e)17,18. Co-depletion of Zucchini/Papi does not impair piRNA generation from two independent reporters (Fig. 1d, ED2b,f), and global piRNA levels are comparable between Zucchini- versus Zucchini/Papi-depleted ovaries (Fig. ED2g). Interestingly, Piwi-bound piRNAs increase by ~0.5 nt in length in papi mutants (Fig. 1e, ED3a)8. As 3' ends of Piwi-bound piRNAs are generated predominantly by Zucchini8,9, we conclude that Papi-assisted piRNA trimming—if conserved in flies—occurs downstream of Zucchini, consistent with its role in mouse and silkworm8,9,17,18.
We next tested the 3'-to-5' exoribonuclease Nibbler/Mut-7, which trims some miRNAs after their loading into Ago110,11, and which has been reported to modulate piRNA lengths12,13. Co-depletion of Zucchini and Nibbler (Fig. ED2c) ablates piRNA production from both reporters despite trigger piRNAs remaining abundant and silencing-competent (Fig. 1d, ED2f,h).
Consistent with Nibbler acting on slicer-generated pre-piRNAs, it is enriched in perinuclear nuage together with Aub/Ago3, while Papi co-localizes with Zucchini at mitochondria (Fig. 1f, ED2d,e). In aubergine mutants, Nibbler’s nuage localization is reduced, yet Nibbler does not enrich in Krimper foci where unloaded Ago3 accumulates (Fig. 1f)16,19,20. Nibbler’s co-localization with Aub/Ago3 therefore probably depends on these factors being loaded with pre-piRNAs. We did not detect robust interactions between Nibbler and Aub/Ago3 by co-IP (weak interactions between Nibbler and Piwi were detected13), hinting at a transient interaction (Fig. ED2i,j).
To characterise Nibbler’s role in piRNA biogenesis we generated flies that express no detectable Nibbler protein (Fig. ED3b,c). As reported12,13, nibbler mutants are viable and fertile, but defective in mir-34 maturation (Fig. ED3d). Also as reported, localization and abundance of PIWI proteins, overall piRNA levels, and transposon silencing are not impacted (Fig. ED3e-h). Average piRNA length, however, is mildly increased (Fig. ED3i; our sequencing libraries span 18-40 nt: Fig. ED4). Notably, this originates primarily from Ago3-bound piRNAs, which increase >1 nt in length (Fig. 1e). This supports a specific role for Nibbler in resecting Aub/Ago3-generated pre-piRNAs. Indeed, somatic Piwi-bound piRNAs, whose 3' ends are generated by Zucchini, show no length change in nibbler mutants, in contrast to papi mutants (Fig. ED3j)12,13. These results indicate that Nibbler does not fine-tune piRNA length as proposed12,13, but instead represents the central exonuclease of a distinct piRNA 3' end pathway that resects slicer-generated pre-piRNAs to mature piRNAs.
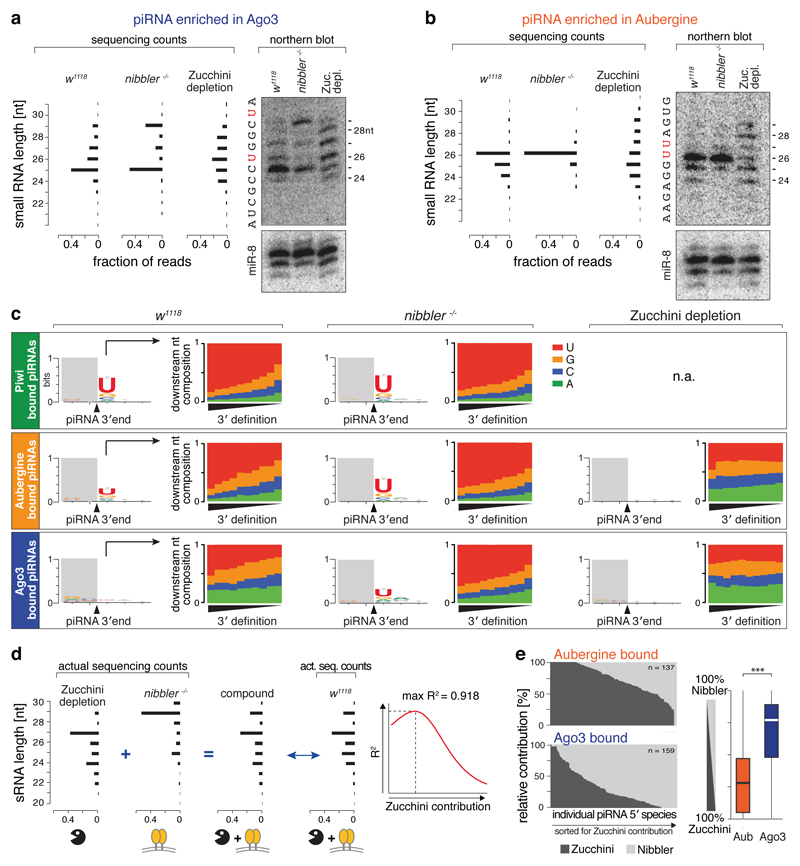
If Zucchini endonuclease and Nibbler exonuclease act in separate pathways to generate piRNA populations with similar overall length, the 3' profiles of piRNA 5' species should differ in single mutant ovaries. We inspected individual Aub/Ago3-bound piRNA 5' species by northern blot analysis and sequencing (Fig. 2a,b). While piRNAs in Zucchini-depleted ovaries display a broad length profile (consistent with exonucleolytic resection), piRNAs in nibbler mutants display discrete length patterns with major isoforms typically being followed by Uridine (downstream-U signature), a hallmark of Zucchini cleavages (Fig. 2a,b)8,9.
Figure 2. Two genetically independent pathways generate piRNA 3' ends.
a, b) Northern blot analysis (loading control: miR-8) and sequencing counts for an Ago3- (a) or an Aub- (b) enriched piRNAs in indicated genetic backgrounds. Bar charts display fraction of reads with indicated length. Respective RNA sequences are shown.
c) Sequence logos display nucleotide bias around dominant 3' ends of Piwi/Aub/Ago3-bound piRNAs isolated from ovaries of indicated genetic background. Bar plots display the nucleotide composition at the position following dominant 3' ends (piRNAs are split into 10 equally sized bins sorted for their precision index).
d) Shown is how the Nibbler/Zucchini contribution to 3' end formation of individual piRNA 5' species was determined.
e) Shown are the Zucchini/Nibbler contributions for 3' end formation for Aub- and Ago3-bound piRNAs (left: as stacked bar diagram; right: as boxplot; Tukey definition; *** p < 0.001 after Wilcoxon rank-sum test).
To generalize these findings, we determined the downstream-U signature and the 3' end precision index for thousands of piRNA 5' species bound to Piwi/Aub/Ago3. This allows several conclusions: (1) In agreement with the two-pathway model, the downstream-U signature increases in nibbler mutant ovaries, yet is ablated in Zucchini-depleted ovaries (Fig. 2c; Piwi-piRNAs are lost in the absence of Zucchini). (2) In wild-type ovaries, the downstream-U signature is strong for Piwi-bound piRNAs, intermediate for Aub, and very weak for Ago3 (Fig. 2c), indicating that Zucchini acts predominantly on Piwi and Aub. (3) The downstream-U signature correlates with the 3' end precision index of piRNA populations (Fig. 2c). (4) In nibbler mutants, the 3' end precision index increases for Piwi-, Aub-, and Ago3-bound piRNAs, indicating that Nibbler acts on all three PIWI proteins (Fig. ED5a). (5) Characteristics of piRNA 5' ends do not correlate with the 3' end precision index (Fig. ED5b), arguing that 5' end generation does not dictate the mode of 3' end formation.
In agreement with Nibbler and Zucchini acting in parallel pathways, the length profiles of wild-type piRNAs appear to be a composite of the two respective single-pathway profiles (Fig. 2a,b). Inspired by this observation, we computed the relative contribution of Nibbler and Zucchini for Aubergine/Ago3-bound piRNAs. For each of ~300 analyzable piRNA 5' species, we determined the Zucchini/Nibbler contribution at which the combined length profile best mimics the wild-type profile (Fig. 2d). For both, Aub- and Ago3-bound species the wild-type profiles can be accurately modeled from the single pathway profiles (Fig. ED5c). This results in a median ~70:30 dominance of Zucchini over Nibbler for Aub-bound piRNAs and an opposite ratio for Ago3-bound piRNAs (Fig. 2e), in agreement with the 3' end characteristics of the respective piRNA populations in wild-type ovaries (Fig. 2c). Our data demonstrate that two parallel pathways with varying contributions form 3' ends of Aub/Ago3-bound piRNAs: Zucchini generates most Aub-bound piRNAs, while Nibbler generates most Ago3-bound piRNAs.
Zucchini-mediated piRNA 3' end formation results in processing of the downstream precursor RNA into phased piRNAs bound to Piwi (referred to as triggering)8,9. 3' end formation by Nibbler instead prevents triggering due to degradation of the downstream precursor (Fig. 3a). As Zucchini compensates for 3' end formation in the absence of Nibbler, triggering levels should increase in nibbler mutants. Indeed, the occurrence of Piwi-bound piRNA 5' ends immediately downstream of Aub/Ago3-bound piRNA 3' ends increases in nibbler mutant compared to wild-type ovaries (Fig. 3b). As expected, this increase is more pronounced for Ago3/Piwi linkages compared to Aub/Piwi linkages. We conclude that in wild-type ovaries, downstream slicing and subsequent Nibbler-catalysed pre-piRNA resection limits the extent of triggering, especially for Ago3-bound pre-piRNAs.
Figure 3. Nibbler and Zucchini set the balance between primary and secondary piRNA biogenesis.
a) Schematic of the two-pathway model for piRNA 3' end formation.
Panels b-c show scatter plots based on piRNAs mapping to germline-dominant TEs (n=63).
b) Plotted are Z-scores of piRNA 3'/5' coupling in w1118 versus nibbler-/- ovaries (orange: Aub-bound antisense piRNA 3' ends to Piwi-bound piRNA 5' ends; blue: Ago3-bound sense piRNA 3' ends to Piwi-bound piRNA 5' ends).
c) Plotted are normalized contributions of Piwi/Aub/Ago3-bound piRNAs to total TE piRNA profiles in w1118 versus nibbler-/- ovaries. Stacked bar plots show sum for all analyzed TEs.
In agreement with elevated triggering, the levels of Piwi-bound piRNAs increase at the expense of ping-pong piRNAs in nibbler mutants (Fig. 3c). It is unclear why Aub-bound, but not Ago3-bound piRNAs are reduced in the absence of Nibbler. Possibly this is due to Ago3 incorporating abnormally high levels of antisense piRNAs (Fig. ED6a,b). As a consequence of the shifts in piRNA populations, ping-pong signatures for nearly all TEs decrease (Fig. ED6c). This indicates a competitive relationship between two piRNA 3' end formation pathways with consequences for piRNA loading into PIWI proteins: Nibbler limits the extent at which slicer-induced piRNA biogenesis propagates into Zucchini-mediated downstream piRNA biogenesis that fuels nuclear Piwi. Zucchini on the other hand consumes piRNA precursors, reducing their participation in ping-pong during post-transcriptional regulation.
These findings prompted us to re-examine the long-standing question why secondary piRNA populations from some TEs remain abundant in Zucchini-depleted ovaries (‘robust TEs’; e.g. Doc), while others collapse (‘sensitive TEs’; e.g. I-element; Fig. ED6d)16. We reasoned that only TEs with a minimal abundance of Nibbler substrates (reflected by abundant ping-pong piRNAs) could maintain piRNA biogenesis in the absence of Zucchini. That is because Zucchini generates piRNA 3' ends independent of additional precursor cleavages, while Nibbler-mediated 3' end formation requires a second piRNA-guided cleavage event close by (Fig. 3a). Indeed, ping-pong piRNA levels are substantially higher for robust compared to sensitive TEs in wild-type ovaries (Fig. ED6d). TEs with ping-pong piRNAs below a threshold level therefore cannot compensate for Zucchini loss, as the production of Nibbler substrates is too inefficient. As ping-pong is a feed-forward loop, this results in the collapse of piRNA biogenesis.
As the most direct test for two separate piRNA 3' end pathways, we co-depleted Zucchini and Nibbler. As expected from the piRNA biogenesis reporter experiments (Fig. 1d), this results in piRNA loss for some TEs, which generate abundant piRNAs in Zucchini-depleted ovaries (Fig. ED1a, ED7a). Surprisingly, however, piRNAs mapping to several other TEs are only mildly affected (Fig. ED7a,b), total germline piRNA levels are reduced less than two-fold compared to Zucchini-depleted ovaries (Fig. ED7c), and TE derepression is similar to Zucchini-depleted ovaries (Fig. ED7d). The remaining germline piRNAs in Zucchini/Nibbler- depleted ovaries populate Aub/Ago3 (Piwi is largely lost; Fig. ED8a) and exhibit less defined size profiles with many piRNAs being abnormally long or short (Fig. ED8b).
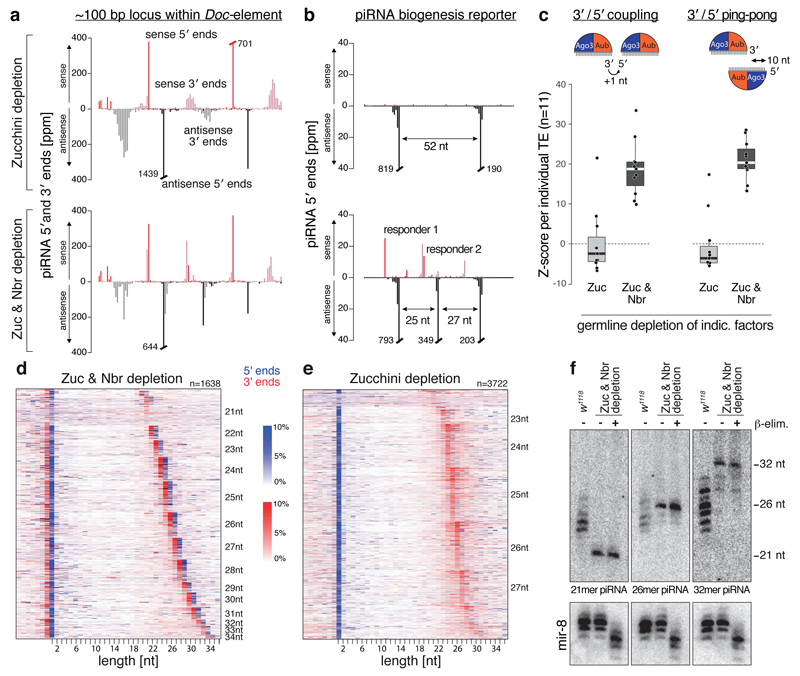
Based on a strong ping-pong signature, Aub/Ago3-mediated slicing defines the 5' ends of Zucchini/Nibbler-independent piRNAs (Fig. ED8c). A closer look at their mappings provides an explanation of how their 3' ends are generated (Fig. 4a): In double-depleted ovaries, novel ping-pong pairs emerge between two distantly spaced ping-pong pairs, thereby reducing the cleavage intervals to ~20-30 nt. Also, piRNA 3' ends change from a bell-shaped profile (consistent with Nibbler-mediated exo-resection) to discrete profiles where a single dominating 3' end precedes the 5' end of a flanking piRNA by 1 nt (Fig. 4a). This suggests that two slicer events, spaced by one piRNA length, directly generate 5' and 3' ends of piRNAs. To test this prediction we turned to the piRNA biogenesis reporter with two cleavage sites spaced by 52 nt that is incompatible with piRNA biogenesis in Zucchini/Nibbler-depleted ovaries (Fig. 4b). Introducing a third central target site re-installs biogenesis for two responder piRNAs, whose 3' ends map precisely to the downstream slicer sites (Fig. 4b).
Figure 4. An Argonaute-only pathway generates piRNAs in the absence of Zucchini and Nibbler.
a) Normalized 5' and 3' end counts (ppm) of piRNAs isolated from ovaries with indicated genetic background mapping to a ~100-nt sequence of Doc.
b) Normalized 5' and 3' end counts (ppm) of piRNAs from Zucchini/Nibbler-depleted ovaries mapping to a biogenesis reporter with two (top) or three (bottom) piRNA target sites.
c) Boxplots displaying Z-score distributions of 3'/5' coupling and 3'/5' ping-pong for piRNAs from ovaries of indicated genotype (n=11 germline TEs that maintain piRNA production in Zucchini/Nibbler-depleted ovaries).
d, e) Displayed are 5' and 3' ends of piRNAs—isolated from Zucchini/Nibbler-depleted (d) or from Zucchini-depleted (e) ovaries—that map within a 40-nt window surrounding ping-pong responder piRNA 5' ends (position 0). Heat maps sorted first by dominant responder piRNA length and then by abundance. The sum of all 3' ends and all 5' ends per line is 100%.
f) Northern blot analysis of piRNAs with dominant length of 21, 26, or 32 nt in Zucchini/Nibbler-depleted ovaries.
We systematically analysed Zucchini/Nibbler-independent piRNAs for two characteristic signatures, namely 3'/5' coupling (nucleotide-precision phasing), and 3'/5' ping-pong (presence of complementary piRNA 5' ends 10 nt downstream of piRNA 3' ends). Both signatures—while absent in Zucchini-depleted ovaries—are pronounced in double-depleted ovaries, indicative of tightly spaced ping-pong pairs (Fig. 4c). When piRNAs are grouped into length cohorts, coupling of flanking piRNAs is apparent for all size classes (Fig. 4d). No piRNA coupling is observed in Zucchini-depleted ovaries, as here Nibbler allows ping-pong pairs to be spaced in a larger window (Fig. 4e, ED8d).
Zucchini/Nibbler-independent piRNAs bound to Aub or Ago3 retain their respective nucleotide bias of 1U and 10A (Fig. ED8e). Given the precise piRNA coupling, this explains that slicer/slicer-generated piRNAs display downstream-1U and downstream-10A signatures (Fig. ED8e). As Uridine residues are not spaced in pre-fixed patterns, this requires flexibility on the Argonaute-side to accommodate piRNAs with different lengths. Indeed, while ~80% of piRNA species in Zucchini-depleted ovaries are 23-27 nt, this is only ~50% in Zucchini/Nibbler-depleted ovaries (Fig. 4d,e, ED8b). Nucleotide-resolution northern blots confirm the existence of piRNAs as short as 21 nt and as long as 32 nt (Fig. 4f, ED9a). While these piRNAs are similarly abundant as the corresponding species in wild-type ovaries, their length is restricted to essentially a single isoform.
When examining slicer/slicer-generated piRNAs for their 2'-O-methylation status, a hallmark of mature piRNA 3' ends21,22, even 32 nt long piRNAs are methylated (Fig. 4f). This extends to all piRNA sizes (Fig. ED9b), indicating that 2'-O-methylation can occur independently of Zucchini- or Nibbler-mediated 3' end formation. As pre-piRNAs with similar length are not methylated in wild-type ovaries (Fig. 1b), Nibbler probably acts faster than Hen1.
Together with recent findings8,9,17, our data provide a blueprint for piRNA 3' end formation: Two separate exonucleolytic pathways—initiated by endonucleolytic cleavages—are dedicated to pre-piRNA trimming: the Papi/PNLDC1 pathway17,18, and the Nibbler/Mut-7 pathway. This is probably an ancient pathway architecture as similar to ping-pong23, all involved nucleases—Zucchini, PNDLC1, and Nibbler/Mut-7—are conserved from sponges to mammals (Fig. ED10). There are, however, interesting exceptions: (1) Nematodes have lost Zucchini and an entirely different small RNA biogenesis system fuels their PIWI proteins24. In C. elegans, PARN trims PIWI-bound small RNAs25, while Nibbler/Mut-7 is required for 22G siRNA biogenesis26. (2) While PARN or PNDLC1 have been sporadically lost in several lineages (e.g. fish), only flies lost both enzymes. We postulate that this central pre-piRNA exonuclease is dispensable in flies as here Zucchini directly forms mature piRNA 3' ends. (3) Only two groups have lost Nibbler: all Anopheles species, and several mammals including rodents. Whether this indicates a lesser importance of efficient ping-pong in these species is currently unclear.
The balance between the two 3' end-generating pathways defines to which extent precursors are processed into Piwi-bound versus Aub/Ago3-bound piRNAs. Ultimately, this determines the ratio between transcriptional (Piwi) and post-transcriptional silencing (Aub/Ago3). Aub-bound piRNAs, which are antisense-biased, are more abundant than Ago3-bound sense piRNAs. Piwi-bound piRNAs, which are generated in response to Aub/Ago3-initiated triggering, are, however, antisense biased27,28. Ago3-generated cleavage intermediates must therefore be transferred more efficiently to mitochondria for Zucchini-mediated 3' end formation than Aub-generated intermediates8,9. Also, Nibbler matures Ago3-bound pre-piRNAs probably more frequently compared to Aub-bound pre-piRNAs, because Aub-bound piRNAs are more abundant than Ago3-bound piRNAs. Consequently, two closely spaced cleavages will be more frequent downstream of Aub than of Ago3.
In the absence of Zucchini and Nibbler, PIWI proteins accommodate 2'-O-methylated piRNAs ranging in length from 20-34 nt. We speculate that the slicer-only pathway represents potentially an ancient small RNA-generating unit, onto which dedicated endo- and exonucleases were added during evolution in order to efficiently generate piRNAs of optimal length.
Methods
Fly husbandry and strains
Flies were kept at 25° C. short hairpin RNA (shRNA) fly lines were crossed to the maternal triple driver (MTD)-GAL4 line (#31777; Bloomington stock center) to drive expression of the shRNA in the germline. shRNA constructs for double depletion of Zucchini-Nibbler and Zucchini-Papi were inserted into attP229. TRiP.GL00111 line was used (#35227; Bloomington) for depleting Zucchini alone. GFP reporter constructs and GFP-tagged BAC rescue constructs were inserted into the attP40 landing site. A nibbler and a papi allele with frame-shift mutations were generated by CRISPR/Cas9 using guide RNAs TGACGCCACCTTGGACGCAA and CGAGCCGCCTTAACCGCATC, respectively, as previously described30. N-terminally FLAG tagged nibbler allele was generated using a guide RNA CACGGGAAACCCGTGAGAAA. The resulting allele has an insertion of IDYKDHDGDYKDHDIDYKDDD after the start codon. w1118 strain was used as a wild-type control throughout the study except the analysis of TE expression from ovaries using RNA sequencing where an shRNA line against white crossed to MTD-GAL4 was used as a control27. Flies were aged for 6 days and kept on apple juice agar plates supplemented with yeast paste to ensure consistent ovarian morphology.
Construction of shRNA expression vectors
To achieve simultaneous knockdown of two genes with a single sh-construct, we modified the Valium20 vector31 and inserted two tandem shRNA sequences. The two hairpins are separated by the sequence that spans the miR-6-3 and miR-6-2 hairpins in the genome in order to maximize efficient processing (referred to as miR-6 backbone in Extended Data Figure 2a). NheI/EcoRI and AgeI/SphI sites were used to clone the two shRNAs oligos. The modified shRNA expression cassette (restriction sites used for cloning of shRNA oligos are underlined and the miR-6 spacer sequence is in bold):
TTCAGCCGCTAGCATGGACGTTCATGGATTCTAAAACGGATTACCAGGGATTTCAGTCGATGTGAATTCAGGCGAGCACGGCCAATTCCAACGATTTGTCATTTGTGGCACGCATTTGTGTCACCTCAGTGCGAAAATTGAAAATTGTATTCAGCCACCGGTTGCATAAGGCTATGAAGAGATACGCCCTGCATGCAGGCGAG
The shRNA oligo sequences are:
Zucchini sh-RNA fwd (guide miR sequence is underlined):
CTAGCAGTCACGAACTTGATGCACAACAATAGTTATATTCAAGCATATTGTTGTGCATCAAGTTCGTGGCG
Zucchini sh-RNA rev:
AATTCGCCACGAACTTGATGCACAACAATATGCTTGAATATAACTATTGTTGTGCATCAAGTTCGTGACTG
Nibbler sh-RNA fwd (guide miR sequence is underlined):
CCGGTAGTATGGTCAGTGATCTCAGTGTATAGTTATATTCAAGCATATACACTGAGATCACTGACCATGCGCATG
Nibbler sh-RNA rev:
CGCATGGTCAGTGATCTCAGTGTATATGCTTGAATATAACTATACACTGAGATCACTGACCATACTA
Papi sh-RNA fwd (guide miR sequence is underlined):
CCGGTAGTTCGACATATCCTAGATCCTAATAGTTATATTCAAGCATATTAGGATCTAGGATATGTCGAGCGCATG
Papi sh-RNA rev:
CGCTCGACATATCCTAGATCCTAATATGCTTGAATATAACTATTAGGATCTAGGATATGTCGAACTA
GFP-tagged Pacman rescue constructs
Recombineering of Pacman32 rescue constructs was as described33. Papi (Pacman clone CH322-41G09) and Zucchini (CH322-41M17) were tagged with GFP-FLAG tags C-terminally, and Nibbler (CH322-18I04) was tagged N-terminally.
Generation of piRNA reporter constructs
All piRNA reporter constructs were generated as described previously9 using the following sequences (target sites for endogenous piRNAs are marked in bold):
Reporter with 2 target sites (Figure 1d):
GGACGAGCTGTACAAGTAATGATCGAGTCAGATAAGCCGTATCAATCAAAGGTTATCAAAGAGCAACTGAAGTACAAGAAAATCGCCTCGGTTCAATGAAATAACTCCAAGAATGCTCATTGAAATGATACTGTCAAAATGTTTCACAGGAAACTACTCAAATATAACCAAAAATTGAAGCAGTCATCAAGTGAACGAAAGAAATGAAAACATGACACGTGATCAACTCGAGCGCATAGAAATTAATTTATAAATTGCAAAACCTAATTACGTAGCTAAACAAAAAACCCAAAAGAATAATGTAAGCAAAAAGAAAATTTAATCGTCATAATCGTCACCAAACCTTTTGATTGGCGAGCATTTCATAGATGTTAAATTTTCCTAATTCGAGAATCCCAGGAGGATAAGCGATAGGGATGATCAGAA
Reporter with 2 target sites (Extended Data Figure 2f):
GGACGAGCTGTACAAGTAATGATCGAGTCAGATAAGCCGTATCAAATTTCAAAGGTTATCAAAGAGCAACTGAAaACCACCGTATAGTGACTATACAGCAAcTCCCACTTAAAAAAGGAAATATTCATGAaCctGctttAgcCtAGGAAACTACTCAAATATAACCAAAAATTGAAGCAGTCATCAAGTGAACGAAAGAAATGAAAACATGACACGTGATCAACTCGAGCGCATAGAAATTAATTTATAAATTGCAAAACCTAATTACGTAGCTAAACAAAAAACCCAAAAGAATAATGTAAGCAAAAAGAAAATTTAATCGTCATAATCGTCACCAAACCTTTTGATTGGCGAGCATTTCATAGATGTTAAATTTTCCTAATTCGAGAATCCCAGGAGGATAAGCGATAGGGATGATCAGAA
Reporter with 3 target sites (Figure 4b):
GGACGAGCTGTACAAGTAATGATCGAGTCAGATAAGCCGTATCAAATTTCAAAGGTTATCAAAGAGCAACTGAAaACCACCGTATAGTGACTAAATACCTAcTCCCACTTAAAAAAGGAAATATTCATGAaCctGctttAgcCTAGGAAACTACTCAAATATAACCAAAAATTGAAGCAGTCATCAAGTGAACGAAAGAAATGAAAACATGACACGTGATCAACTCGAGCGCATAGAAATTAATTTATAAATTGCAAAACCTAATTACGTAGCTAAACAAAAAACCCAAAAGAATAATGTAAGCAAAAAGAAAATTTAATCGTCATAATCGTCACCAAACCTTTTGATTGGCGAGCATTTCATAGATGTTAAATTTTCCTAATTCGAGAATCCCAGGAGGATAAGCGATAGGGATGATCAGAA
Generation of Nibbler antibody
Purified His-tagged full-length Nibbler protein was used to generate the mouse anti-Nibbler monoclonal antibody.
Antibodies used
The following primary antibodies were used: Mouse monoclonal anti-Piwi (8C-E4)27 (western: 1/1000; used for IP), rabbit anti-Piwi6 (IF: 1/500), mouse monoclonal anti-Ago3 (7B4-C2)*(western: 1/1000; used for IP), mouse monoclonal anti-Ago3 (5H12-G12)27 (IF: 1/30), mouse monoclonal anti-Nibbler (IF: 1/400; western: 1/2000), mouse monoclonal anti-Aub (8A3-D7)27 (IF: 1:500; western: 1/1000; used for IP), mouse anti-ATP-synthase 5A (abcam 14748; IF: 1/2000; western: 1/20,000).
Secondary antibodies used were: goat-anti-mouse HRP (Invitrogen), Alexa Fluor 488 anti-mouse (Thermo Fisher), Alexa Fluor 568 anti-mouse (Thermo Fisher), Alexa Fluor 647anti-mouse (Thermo Fisher), Alexa Fluor 568 anti-rabbit (Thermo Fisher).
Immunofluorescence staining
Ovaries were dissected into 300 μl PBS containing 4% PFA and fixed for 25 min at room temperature (RT). Tissue was permeabilized 3 times with PBX (1x PBS, 0.3% TritonX-100) and blocked with BBX (1x PBS, 0.3% TritonX-100, 0.1% BSA) for 30 min. 200 μl of primary antibodies diluted in BBX were added and ovaries were incubated while rotating over night at 4 °C. After washing 3 times with PBX, the ovaries were incubated with the respective secondary antibodies diluted in BBX rotating overnight at 4 °C. Subsequently, ovaries were washed three times with PBX before mounting. To the second washing step DAPI was added. Images were taken on a Zeiss LSM-780 Axio Imager confocal-microscope using a 40x/1.3 EC plan-neofluar Oil DIC objective. All images were processed using Image J.
Immunoprecipitation of PIWI-family proteins for piRNA sequencing
For each genotype, 600 μl of ovaries were dissected into 1x PBS on ice. 1.5 ml of 1x RIPA buffer were added (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% TritonX-100, 0.1% SDS, 0.1% Na-deoxycholate, 1 mM EDTA, 0.1 mM Pefabloc (Roche)) and the tissue was homogenized, using a glass tissue homogenizer, on ice. The lysate was cleared by centrifugation and diluted with 3 ml of IP dilution buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl). Antibodies were coupled to M280 sheep anti-mouse IgG Dynabeads (Life Technologies). For Piwi and Aub IP, 150 μl of beads coupled to the respective antibody were mixed with 1.5 ml of lysate. For Ago3 IP, 300 μl of beads coupled to Ago3 antibody were mixed with 3 ml of lysate. Antibodies used are indicated above. Lysates were incubated rotating at 4 °C overnight. Subsequently, the beads were captured and washed seven times with IP wash buffer (50 mM Tris-HCl pH 7.5, 500 mM NaCl, 2 mM MgCl2, 10% glycerol, 1% Empigen). For Piwi IP, 150 mM NaCl was used instead of 500 mM NaCl. The bound RNA was extracted using acid-phenol:chloroform (Ambion) followed by ethanol precipitation. To visualize the extracted RNA during the cloning process 10% of it were labeled with [γ-32P]-ATP.
Co-immunoprecipitation of FLAG-Nibbler
100 μl each of freshly dissected ovaries from 1-2 days old w1118 and flag-nibbler homozygous females were homogenized in lysis buffer (30 mM HEPES-KOH pH 7.4, 500 mM NaCl, 2 mM MgCl2, 0.5% v/v Triton-X, 0.2 mM DTT, and 1x cOmplete, EDTA-free protease inhibitor cocktail (Roche)) using a glass tissue homogenizer, on ice. The lysates were cleared by centrifugation and incubated with 30 μl of anti-FLAG M2 agarose beads (sigma) for 4 hr at 4 °C. Beads were captured and washed 5 times with the lysis buffer. IP fraction was eluted by incubating the beads with 30 μl of lysis buffer containing 0.5 μg/μl 3 x FLAG peptides for 15 min at RT with gentle shaking.
2S rRNA depletion from total RNA
For the depletion of 2S rRNA from 10 μg of total ovarian RNA, 100 μl slurry of Myone Streptavidin C1 beads (Invitrogen) were used. The beads were washed twice with 500 μl 0.5x SSC. After washing, the beads were re-suspended in 300 μl 0.5x SSC and 1 μl of 100 μM 2s-rRNA-as-oligo (Bio-AGTCTTACAACCCTCAACCATATGTAGTCCAAGCAGCACT) were added and mixed. This mixture was incubated on ice for 30 min, followed by a wash with 500 μl of 0.5x SSC to remove unbound 2s-rRNA-as-oligo. After resuspension of the beads in 100 μl, they were incubated at 65 °C for 5 min. The RNA was denatured for 5 min at 80 °C for 5 min and mixed with the beads, followed by incubation at 50 °C for 1 hr. Unbound fraction to the beads was collected and the RNA was ethanol precipitated and served as input for the small RNA cloning procedure.
Small RNA cloning and sequencing
Small RNA libraries were prepared as described previously34. In brief, total RNA from respective ovaries was isolated using TRIzol, and was subjected to 2S-depletion. Oxidation was done by incubating 2S-depleted total RNA in borate buffer (pH 8.6) containing 25 mM NaIO4 at room temperature for 30 min. RNA samples from total RNA and from IP experiments were resolved on a denaturing polyacrylamide gel and small RNAs ranging from 18-40 nt were excised and recovered. These were subsequently ligated to 3' and 5' adapters containing four random nucleotides at both ends to reduce ligation biases. Ligated RNA was reverse-transcribed, PCR amplified and the libraries were sequenced on an Illumina HiSeq 2500 machine in single-read 50 mode. IP libraries of the Zucchini/Nibbler-depleted flies were sequenced in single-read 100 mode.
RNA sequencing
PolyA+ RNA-seq was performed as described in Mohn et al. 2014 using NEBNext Ultra directional RNA library prep kit for Illumina (NEB) and libraries were sequenced on Illumina HiSeq 2500 in paired-end 125 mode35. Six bases at each end of the reads were trimmed and the remaining part was split into three reads (37, 38, 38 nt). Reads were mapped against TE consensus sequences using Bowtie (0. 12. 9) allowing up to three mismatches27. RPKM for each TE was calculated as reads per million genome mapped reads per kb. TEs that were expressed more than 1 RPKM in at least one of the libraries in the comparison (group1: w1118 and nibbler-/-, group2: control depletion, zucchini depletion and zucchini and nibbler depletion) were included for the analysis.
Northern blot
Small RNAs were enriched from 50 μg of total RNA using size selection on a 12% denaturing urea-polyacrylamide gel. Oxidation was performed as mentioned above. β-elimination was done after the oxidation by adding 50 mM f.c. NaOH and incubating for 90 min at 45 °C. The protocol for northern blotting was modified from Pall et al.36.
In brief, a 15% denaturing urea-polyacrylamide gel (0.4 mm thick) was used to run the size selected RNA samples. The gel was blotted to a Hybond-NX membrane (Amersham) in a semi-dry blotting apparatus. This was followed by chemical crosslinking using 0.373 g 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) (Sigma) powder in 12 ml 1x methylimidazole (Sigma) at 60 °C for 1 hr. Afterwards the membrane was pre-hybridized in 10-15 ml (depending on membrane size) church buffer (1 mM EDTA, 0.5 M Na2HPO4/NaH2PO4 pH 7.2, 7% SDS) at 65 °C for at least 20 min. DNA oligonucleotide probes were labeled using [γ-32P]-ATP (PerkinElmer). The radioactively-labeled probe was added to the church buffer and incubated overnight at 32 °C. The membrane was subsequently washed 3 times with 15 ml of 1x SSC + 0.1% SDS. Finally, a phosphor-storage screen was exposed to the membrane and developed using a phosphor imager. All images were processed using Image J.
The following DNA oligos were used as probes:
miR-34 (Extended Data Figure 3d):
AACCAGCTAACCACACTGCCA
miR-8 (Fig. 1b, 2a, 2b and 4f):
GACATCTTTACCTGACAGTATTA
piRNA enriched in Ago3 (Fig. 2a):
GCGATTTTCTTGGGTTCAGTTGCT
piRNA enriched in Aubergine (Fig. 2b):
TCTTGGAGCTAACTTCTTTCGTA
21nt long piRNA (Fig. 4f):
TCTAGATTGGCTGCTATTAAA
26nt long piRNA (Fig. 4f):
AAGCTACTGAAGTCATACCTATA
32nt long piRNA (Fig. 4f):
AGAACAAACTGGCCAAGGTATCAA
piRNA precursor (Fig. 1b):
AGTCTGGAGTTCAAAGCTCTTCTA
Computational Analysis
Processing of small RNA libraries
Processing and mapping of sequenced small RNA reads was carried out as described in Mohn et al. In brief, raw reads were trimmed off the 3' linker sequence (AGATCGGAAGAGCACACGTCT) and the four random nucleotides at each end were removed. Processed reads were mapped to the Drosophila genome (dm3, release 5.55) using Bowtie (0. 12. 9) allowing zero mismatches. Reads that mapped to genomic regions annotated as TE were used for the subsequent analyses (filtered reads). Libraries from total RNA were normalized to 1 million miRNA reads. Normalization of PIWI-IP libraries was as described previously9. R (version 2.15.3) was used for statistical and graphical analyses unless otherwise indicated.
TEs that were included in the analyses
63 TEs that fulfilled the following criteria were defined as germline-enriched TEs: 1. TEs that produced more than 1000 ppm of piRNAs (sum of sense and anti-sense piRNAs) in wild-type ovaries. 2. Germline-derived Piwi-bound piRNAs constitute more than 85% of total Piwi-bound piRNAs27. 63 TEs were grouped into two groups based on the ratio of summed TE mapping reads of normalized Piwi/Aub/Ago3 IP libraries in Zucchini-depleted ovaries relative to the control depletion (group1: >0.2, n=19, group2: 0.2>, n=44, used in Fig. ED1b and 3e). 11 out of 63 TEs that fulfilled the following criteria were used for the coupling/ping-pong analyses in Fig. 4c, ED8c: 1. TEs that produced more than 10% of piRNAs in Zucchini and Nibbler-depleted ovaries compared to the control depletion. 2. TEs that scored the Z-score of canonical ping-pong (see below) greater than 15.
Statistical analysis of size profiles
1000 TE-mapping reads were randomly sampled 100 times. The size distribution of these 1000 reads was compared between genotypes using a t-test. The mean p-value of 100 comparisons was calculated to determine the significance.
Linkage analysis
Linkage calculation was adapted from Webster et al.20. First, filtered small RNA reads were mapped to TE consensus sequences27 allowing three mismatches. Second, 5' and 3' ends of mapped reads were counted at each nucleotide position on both strands. For each linkage analysis, the statistical significance of observing a certain linkage was calculated as follows:
For example, for the calculation of canonical ping-pong, piRNA 5' ends were randomly subsampled for a fixed number of times from anti-sense and sense mapped piRNAs. We sampled for the number of a fifth of TE length (bp). For example, if the size of a TE is 5000 bp, we sampled 5' ends for 1000 times. A fixed number of samplings allows a fair comparison between libraries of different sequencing depths. shuf function in bash 4.2.37(1) was used to obtain random numbers.
We counted the number of 5' ends of anti-sense piRNAs that find at least one 5' end of sense piRNA at 10-nt off-set (N).
We computationally repeated 1 and 2 for 500 times and calculated the mean of Nmean.
Suppose two sets of random 5' ends, the probability of an anti-sense 5' end to find a ping-pong partner p is 1 - (1-1/L)^L where L is the length of TE. The expected number of N from a random set (Nexpected) is L/5 *p.
Z-score was calculated as (Nmean-Nexpected)/σ where the standard deviation σ is (L/5*p*(1-p))^0.5.
We used the following sets of sequencing reads for calculation. Canonical ping-pong: 5' ends of anti-sense and sense mapped reads from total libraries (10-nt off-set, Extended Data Figures 1b and 8c). 3' / 5' coupling: 3' ends and 5' ends of anti-sense mapped reads from total libraries (1-nt off-set, Extended Data Figures 1b and Figure 4c). 3' / 5' ping-pong: 3' ends of anti-sense mapped reads and 5' ends of sense mapped reads (10-nt off-set,Extended Data Figures 1b and Figure 4c).
Mapping small RNA reads to reporter constructs
Responder and trigger piRNA reads were determined by mapping the total small RNA reads including genome-unmapped reads to reporter construct sequences using bowtie allowing zero mismatches. To determine trigger piRNA levels, only the first 20 nt of sequencing reads were used in order to account for 3' heterogeneity. Mapped reads were normalized to 1 million miRNA reads.
Measure the definition of 3'ends
The analysis was adapted from Mohn et al.9. In brief, piRNA 5' end positions where the downstream 20–35 nt window had equal number of mappings in the reference genome were selected as unambiguous 5' end positions. We only included the reads whose 5' ends are defined as unambiguous (“analyzable” reads). These “analyzable” reads were collapsed on 5' ends yielding the counts of each length per given 5' end. 5' ends that have more than 20 raw counts as well as more than 2 counts normalized to one million TE-mapping reads (ppm) were included in the analysis. These cutoffs allowed the comparison of different sequencing depths as well as the confident assessment of 3' end variants. Finally, the fraction of the counts representing the dominant length per 5' end was calculated in percent (definition). For the analysis of somatic piRNAs, piRNAs that mapped to soma-enriched 1 kb tiles were used with a cutoff of soma-index greater than 835.
Nucleotide analysis
Only piRNA reads that passed the requirements for 3' end analysis (see above) were used for the analysis. The entire analysis is sequence-based, not read-based (cloning frequency is therefore not considered). Nucleotide windows surrounding the respective positions were extracted using get-fasta from the fastX-toolkit (Hannon lab). For each 5' end, the dominant 3' end position was used to extract the surrounding sequences. When indicated, the 5' ends were binned into 10 groups containing an equal number of 5' ends according to the definition of their 3' ends. The nucleotide signatures were generated using weblogo 3.4 and Prism 6 was used for visualization.
Contribution Analysis for Nibbler and Zucchini
The relative contribution of Nibbler and Zucchini in forming the 3' ends of Aub- or Ago3-bound piRNAs was modelled as follows:
“Analyzable” piRNA 5' ends (see above) with at least 20 ppm in the respective IP libraries of all three genotypes (w1118, nibbler -/- , and Zucchini depletion) were included in the analysis. First, the 3' end composition of individual 5' ends in a window of 20-30 nt of length was determined. The 3' end profile of piRNAs from nibbler -/- and Zucchini depletion libraries were summed in different ratios (0-100% and the reciprocal values) to generate a combined profile. Second, the combined profile was compared to the w1118 profile using a linear regression model. The R2 value was used to assess the best fit. Finally, the ratios of Nibbler vs. Zucchini contribution were extracted at the maximal R2 value.
Heat maps for 3'-to-5' coupling
Heat maps were generated as previously described9. Briefly, filtered reads (see above) from total small RNA libraries were mapped to TE consensus sequences allowing up to three mismatches. We selected pairs of complementary reads with 10-nt 5' overlap (ping-pong piRNA pairs). Ping-pong piRNAs that were cloned greater than 10 ppm and more than 2% of the most abundant piRNA per individual TEs were selected for the analysis. We extracted the counts of 5' and 3' ends mapped in the window of +/- 100 nt for each ping-pong piRNA (two instances per pair) using annotatePeaks (Homer v4.5) and transformed the counts at each position to a percentage value (read count table). The read count tables per ping-pong piRNA were sorted by the length of the dominant ping-pong responder piRNA and by the relative abundance of the ping-pong responder piRNA in the window, and were assembled into heat maps. The distribution of 5'-to-5' distance of flanking ping-pong piRNAs was calculated by summing the read count table of all ping-pong piRNAs in the respective genotypes (Extended Data Figure 8d).
Ortholog search
Orthologs for Zucchini/MitoPLD, PARN, PNLDC1, and Nibbler/Mut7 were first searched using orthoDB v9 against metazoan species (http://www.orthodb.org/)37. Ortholog groups (EOG091G0FI5: Zucchini/MitoPLD, EOG091G04CM: PARN/PNLDC1, EOG091G04KJ: Nibbler/Mut7) were analysed. Protein entries that were not annotated with the following pfam domains were excluded: Zucchini/MitoPLD: PLDc_2 (PF13091), PARN/PNLDC1: CAF1 (PF04857), Nibbler/Mut7: DNA_pol_A_exo1 (PF01612). Pfamscan (http://www.ebi.ac.uk/Tools/pfa/pfamscan/) was used to search for the domains (pfam-A, E-value: 0.1). We defined PARN as proteins that have RNA_bind (PF08675) and/or RRM_7 (PF16367) and PNLDC1 as proteins that do not have these annotations. Species that did not have orthologs in orthoDB were further searched for orthologs firstly by BLASTp (NCBI) against non-redundant protein sequences and secondly by tBLASTn (NCBI) against whole genome shotgun sequences using the default cutoffs. Hits from BLASTp and tBLASTn were reciprocally surveyed for predicting the query proteins as the most confident homolog. All hits were combined and displayed in a phylogenetic tree (either presence or absence) using iTOL (http://itol.embl.de/)38.
Code availability
The code that underlies the computational analyses in this manuscript is available at: https://gitlab.com/groups/Genetic_and_mechanistic_diversity_of_piRNA_3prime_end_formation
Data availability statement
All sequence data that support the findings of this study (see SI) have been deposited in NCBI GEO with the accession code https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE83698. All fly lines used in this study are available from VDRC (http://stockcenter.vdrc.at/control/main). Source data for all gel images are provided with the paper (see SI).
Extended Data
Figure ED1. 3' ends of Zucchini-independent ping-pong piRNAs are formed by an exonuclease.
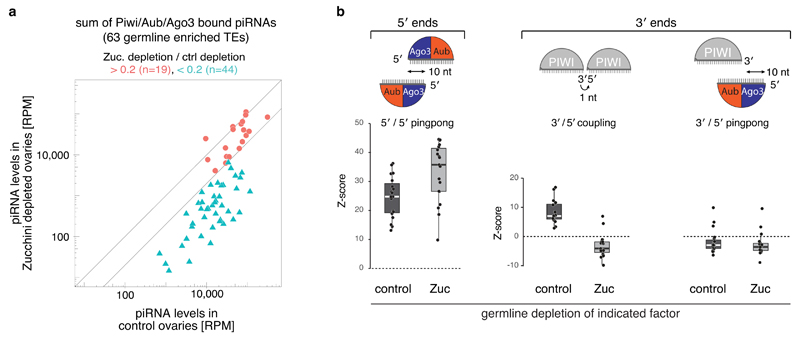
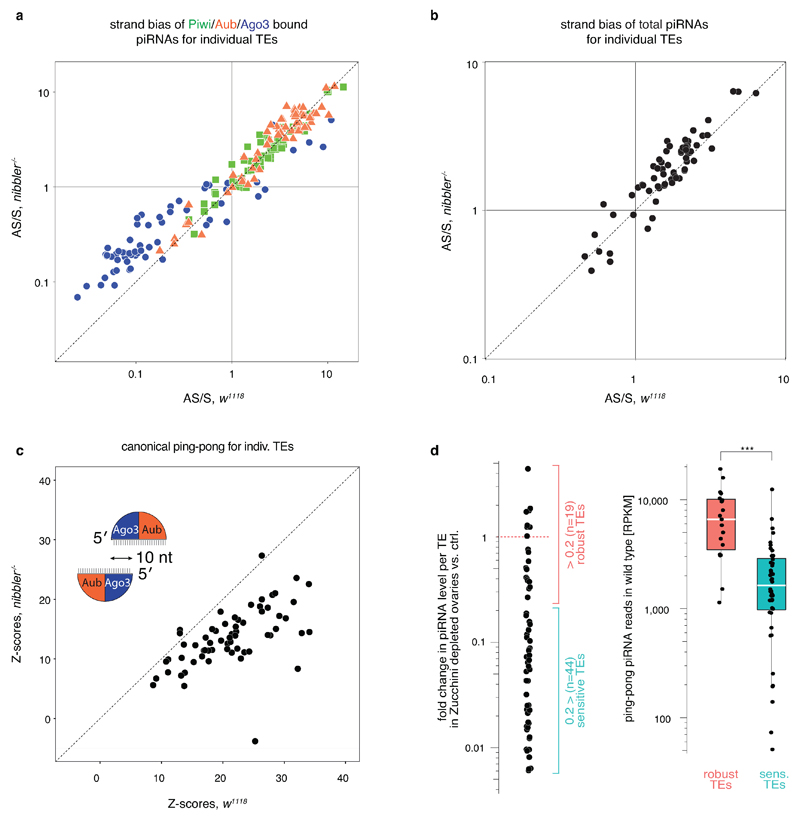
a) Scatter plot showing the -fold change in piRNA levels (for 63 germline-dominant TEs) in Zucchini-depleted compared to control ovaries (calculated as sum of normalized Piwi/Aub/Ago3-bound piRNAs). TEs were grouped into robust (red) and sensitive (blue) based on the piRNA loss (threshold = 5x loss).
b) Boxplots displaying the Z-scores of canonical 5'/5' ping-pong, 3'/5' coupling, and 3'/5' ping-pong for piRNAs isolated from ovaries of indicated genotype (for the 19 robust germline-enriched TEs that maintain piRNA production in Zucchini-depleted ovaries; defined in panel a).
Figure ED2. Nibbler, but not Papi, is required for the generation of piRNAs in the absence of Zucchini.
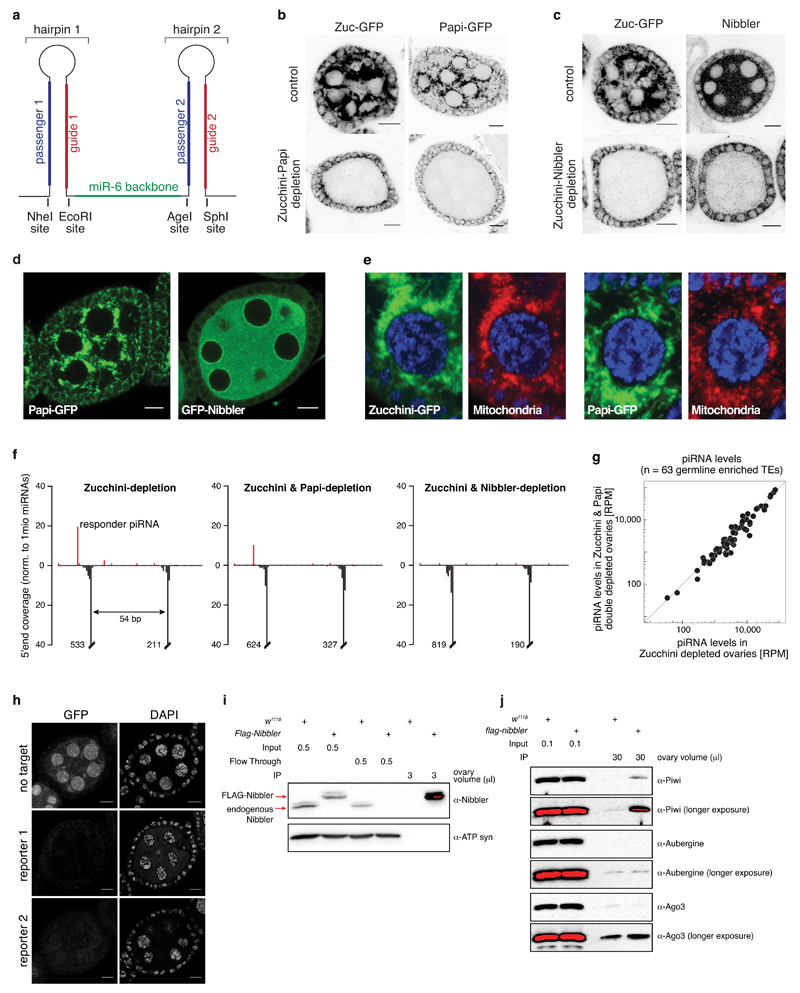
a) Design of the short hairpin expression cassette that allows the simultaneous RNAi-mediated knockdown of two genes in a tissue specific manner. shRNAs are separated by the miR-6 backbone and can be cloned via indicated restriction sites.
b, c) Confocal sections of egg chambers (scale bars: 10 μm) of indicated genotype expressing GFP-tagged Zucchini (b, c) and GFP-tagged Papi (b), showing the efficient shRNA-mediated knockdown of Zucchini and Papi (b), or Zucchini and Nibbler (c) in the germline. Nibbler was detected using a monoclonal antibody.
d) Confocal images showing the localization of GFP-tagged Papi and Nibbler in Drosophila egg-chambers (scale bars: 10 μm). Functionality of GFP-Nibbler is demonstrated in Fig. ED3d.
e) Confocal sections through single nurse cell nuclei of egg chambers expressing GFP-tagged Zucchini (left) or Papi (right) and stained for mitochondria (immuno-staining of ATP synthase).
f) Shown are mappings of piRNAs (5' ends only; red: sense; black: antisense) from ovaries of indicated genotype to a second reporter construct (as in Fig. 1d). Values are normalized to 1 million sequenced miRNA reads.
g) Scatter plot displaying miRNA-normalized piRNA levels mapping to 63 germline-dominant TEs in Zucchini-depleted versus Zucchini/Papi-depleted ovaries.
h) Shown are confocal sections of egg chambers expressing indicated piRNA biogenesis reporters (GFP-fluorescence; DNA stained with DAPI) in wild-type (top) or Zucchini and Nibbler-depleted ovaries (middle and bottom). top: reporter with no target site; middle: reporter used in Fig. 1d; bottom: reporter used in Fig. ED2f.
i) Ovary lysates expressing N-terminally FLAG tagged wild-type Nibbler was immunoprecipitated (IPed) using M2 magnetic beads. Wild-type ovary lysates were used as a control. Red color in the blot indicates a saturated signal. ATP synthase 5A (ATP syn) serves as loading control. Ovary volume indicates the amount of loaded lysate/IP fraction.
j) Eluates from IP were blotted with indicated antibodies. Piwi is slightly enriched in the FLAG-Nibbler IP fraction, while there is no detectable enrichment of Aub or Ago3 in the IP fraction.
Figure ED3. Molecular characterization of the piRNA pathway in nibbler and papi mutant flies.
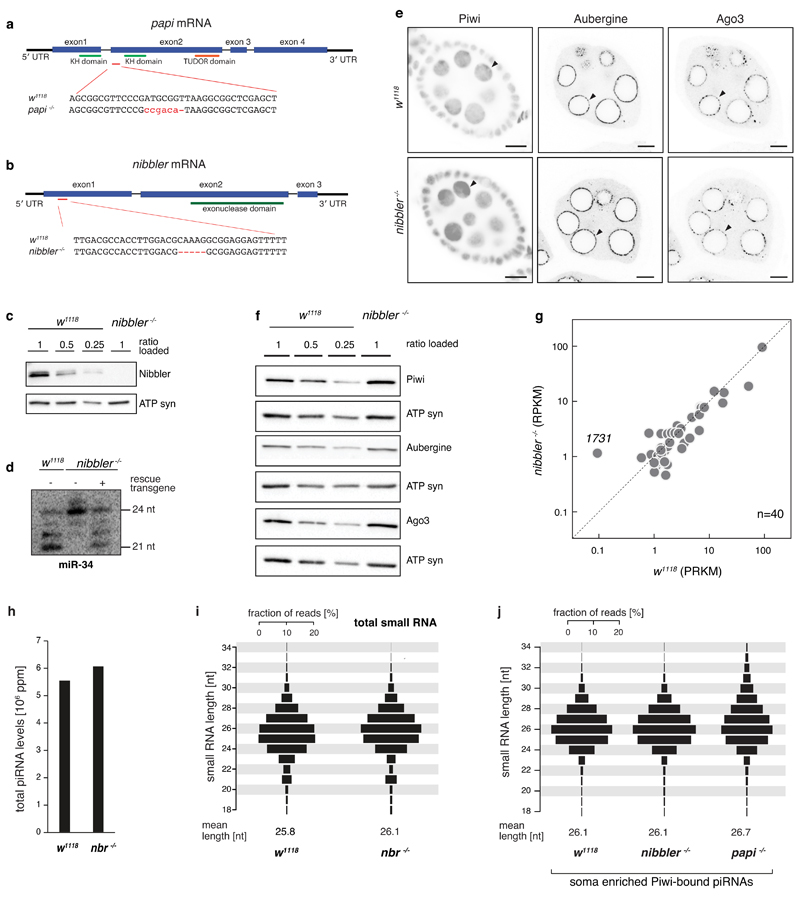
a) papi gene locus indicating the position of the Cas9-induced frameshift allele.
b) nibbler gene locus indicating the position of the Cas9-induced frameshift allele.
c) Western blot analysis showing the loss of detectable Nibbler protein in nibbler -/- ovaries. ATP-synthase 5A antibody is used as loading control. Loaded amounts of ovary lysates are indicated.
d) Northern blot analysis of the Nibbler-substrate miR-34 comparing small RNAs obtained from ovaries of w1118 or nibbler -/- flies. The GFP-Nibbler rescue transgene (used in Fig. 1f) restores miR-34 processing.
e, f) Immunostainings (e) and western blot analysis (f) of Piwi, Aub and Ago3 in w1118 or in nibbler -/- ovaries, showing that localization and expression of the three PIWI-clade proteins are unperturbed (arrow heads; scale bars: 10 μm). ATP synthase 5A (ATP syn) served as loading control.
g) Scatter plot displaying the steady-state RNA level of TEs in indicated genetic background (only TEs with RPKM > 1 in either background; n = 40).
h) Bar chart displaying TE mapping piRNA levels in w1118 or in nibbler -/- ovaries (values normalized to 1 million sequenced miRNA reads).
i, j) Length profiles of TE mapping small RNA reads obtained from ovaries of indicated genotypes. Shown are all ovarian small RNAs (i) or Piwi-bound piRNAs defined as soma-enriched (j; see Methods). Displayed are fractions of reads of indicated length in percent (mean lengths are indicated below).
Figure ED4. A small RNA library cloning approach that allows the recovery of longer piRNA species.
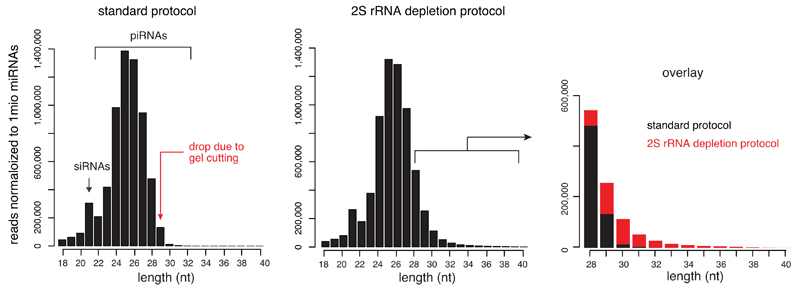
Drosophila total RNA contains large amounts of the 31-nt long 2S rRNA. Previous cloning approaches therefore typically restrict small RNA cloning to the 18-29 nt window by cutting these small RNA populations from a gel. We used the 2S rRNA depletion method published by Seitz et al. (2008), followed by extracting small RNAs ranging from 18-40 nt in length for library preparation.
Shown are size distributions of TE mapping small RNAs (obtained from w1118 ovaries) comparing the standard small RNA cloning protocol (left) and the protocol using total RNA depleted of 2S rRNA (middle; see Methods). An overlay of the longer reads (>27 nt) is displayed to the right.
Figure ED5. Zucchini and Nibbler generate Aub- and Ago3-bound piRNA 3' ends independently of piRNA 5' end formation.
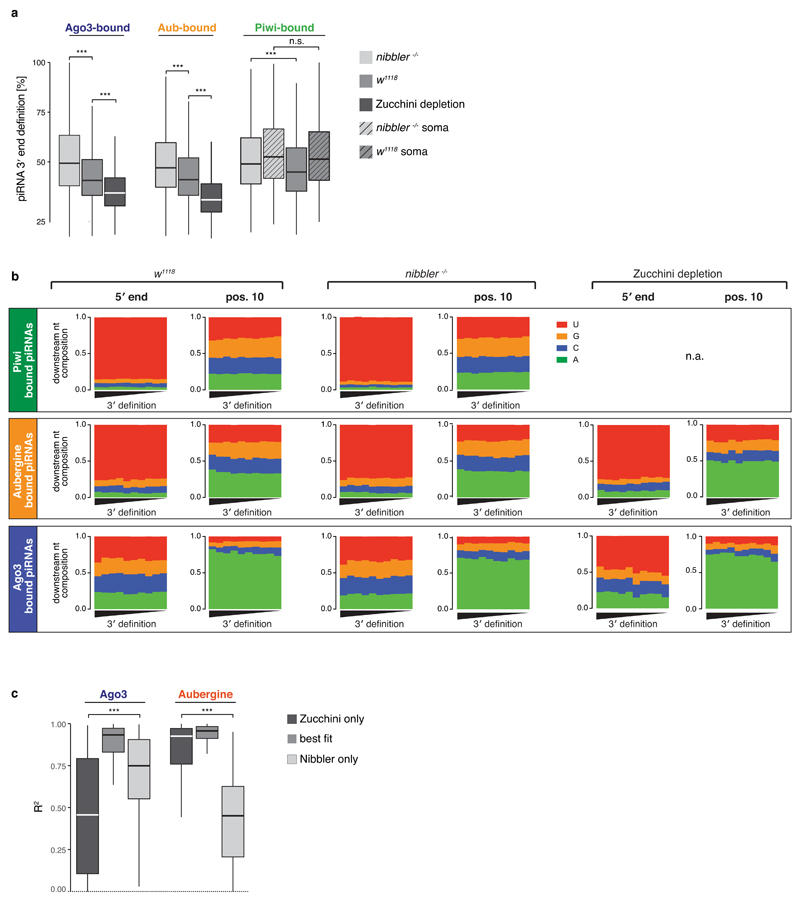
a) Boxplots (*** p < 0.001 by two-sided t-test) showing the 3' end definition (see Methods) of Ago3-, Aub-, and Piwi-bound piRNAs isolated from ovaries of the indicated genotypes. Soma-enriched Piwi-bound piRNAs (see Methods) are shown in boxes with diagonal lines.
b) Stacked bar plots displaying the nucleotide composition at the 5' end or position 10 of piRNAs bound to Aub-, Ago3-, or Piwi (isolated from ovaries of indicated genotypes). The plots show the composition within 10 equally sized bins, sorted for their 3' end precision index (see Methods).
c) Boxplots (*** p < 0.001 by Wilcoxon rank-sum test) showing the R2 values of the comparison between the 3' end profiles of Ago3- or Aub-bound piRNAs from w1118 ovaries and those of nibbler-/- ovaries (Zucchini only), those of ovaries depleted of Zucchini (Nibbler only), and those of the calculated composite that provides the highest R2 (best fit).
Figure ED6. Ago3 incorporates more TE antisense piRNAs in nibbler mutant flies.
a, b) Scatter plots displaying the strand bias (antisense divided by sense) of piRNA populations in w1118 versus nibbler -/- ovaries. (a) displays piRNAs bound to Piwi, Aubergine, or Ago3, while (b) displays piRNAs from total ovarian RNA.
c) Plotted are Z-scores of 5'/5' ping-pong levels per TE (63 germline dominant TEs) in w1118 versus nibbler-/- ovaries.
d) Grouping of TEs (63 germline dominant TEs) based on -fold change in piRNA levels between Zucchini-depleted and control ovaries (left). Boxplot indicates Aub/Ago3-bound piRNA levels in wild-type ovaries for defined TE groups (Tukey definition; *** p < 0.001 after Wilcoxon rank-sum test).
Figure ED7. piRNAs are abundantly produced in Zucchini/Nibbler double-depleted ovaries.
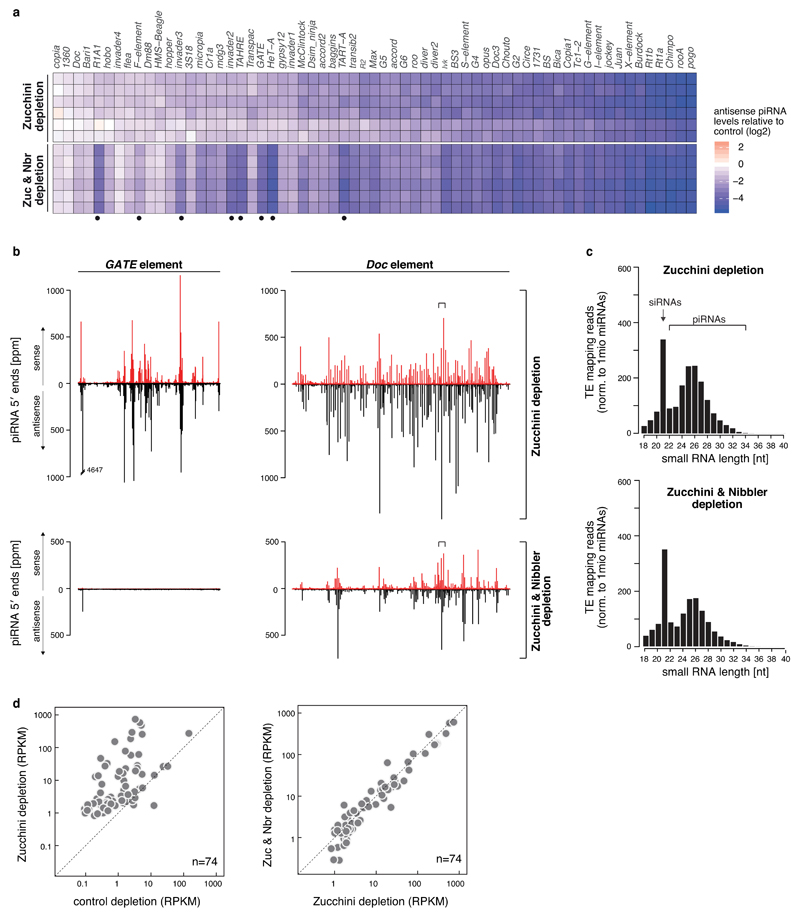
a) Heat map displaying -fold changes in antisense piRNA levels per TE (n=63) in indicated genotypes versus control (six biological replicates). Dots mark TEs, which loose piRNAs only in Zucchini/Nibbler-depleted ovaries.
b) Shown are mappings (5' ends only; red: sense; black: antisense) of piRNAs onto the GATE and Doc TE consensus sequences. The plots at the top are from a piRNA library obtained from Zucchini-depleted ovaries, and the ones at the bottom are from a piRNA library obtained from Zucchini/Nibbler double depleted ovaries. The ~100-bp window of Doc that is detailed in Fig. 4a is depicted by brackets.
c) Size distributions of TE mapping small RNAs obtained from total small RNA libraries from Zucchini-depleted (top) or Zucchini/Nibbler-depleted (bottom) ovaries. Shown are the average values from six biological replicates (reads were normalized to 1 million sequenced miRNA reads).
d) Scatter plots displaying the steady state RNA level of TEs in indicated genetic background (only TEs with RPKM > 1 in either background are shown; n = 74).
Figure ED8. Precise coupling of neighboring ping-pong piRNAs in Zucchini/Nibbler-depleted ovaries.
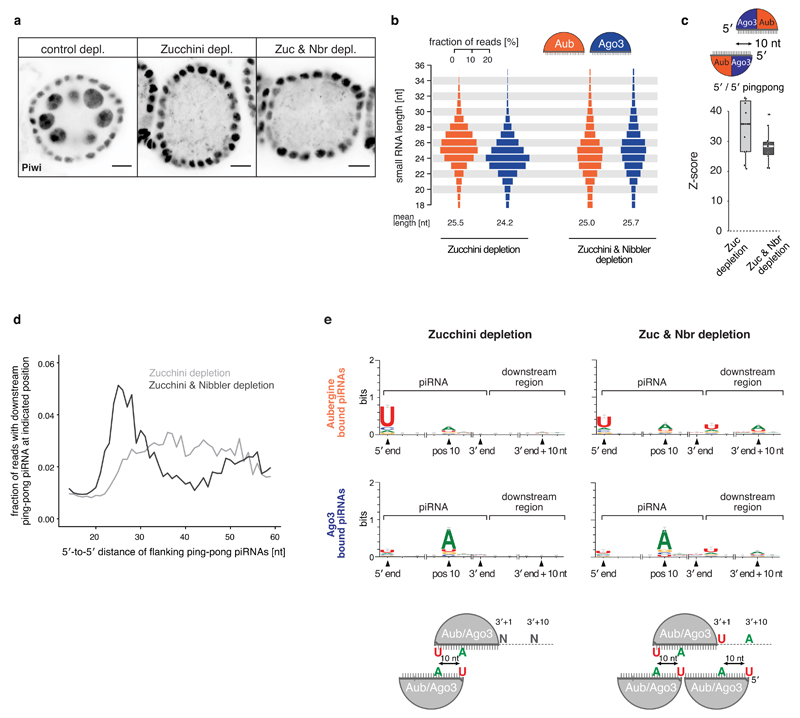
a) Immunofluorescence images (confocal sections) of egg chambers of indicated genotype stained for endogenous Piwi protein (scale bars: 10 μm). Note that the shRNA-mediated knockdown is specific for germline cells. Somatic follicle cells that also express Piwi serve therefore as control.
b) Length profiles (fractions of reads per indicated length in percent) of TE-mapping piRNAs bound to Aub/Ago3 in indicated genotypes.
c) Boxplot displaying the distribution of Z-scores for canonical 5'/5' ping-pong of piRNA populations isolated from ovaries of indicated genotypes. The analysis is restricted to the eleven germline enriched TEs that maintain piRNA production in Zucchini/Nibbler-depleted ovaries (see Methods).
d) Histogram showing the frequencies of a cloned ping-pong piRNA 5' end downstream of a responder piRNA 5' end at a certain distance on the same strand.
e) Sequence logos displaying the nucleotide composition within and downstream of Aub- and Ago3-bound piRNAs cloned from ovaries of indicated genotypes. 5' end position and position 10 are measured from the piRNA 5' end. 3' end position is the dominant 3’ end of a certain piRNA 5' species. Downstream positions are anchored by the dominant 3' end position. Nucleotide signatures with respect to the position of sense and antisense slicer piRNAs are depicted in the cartoons below.
Figure ED9. piRNAs generated by slicer/slicer are methylated.
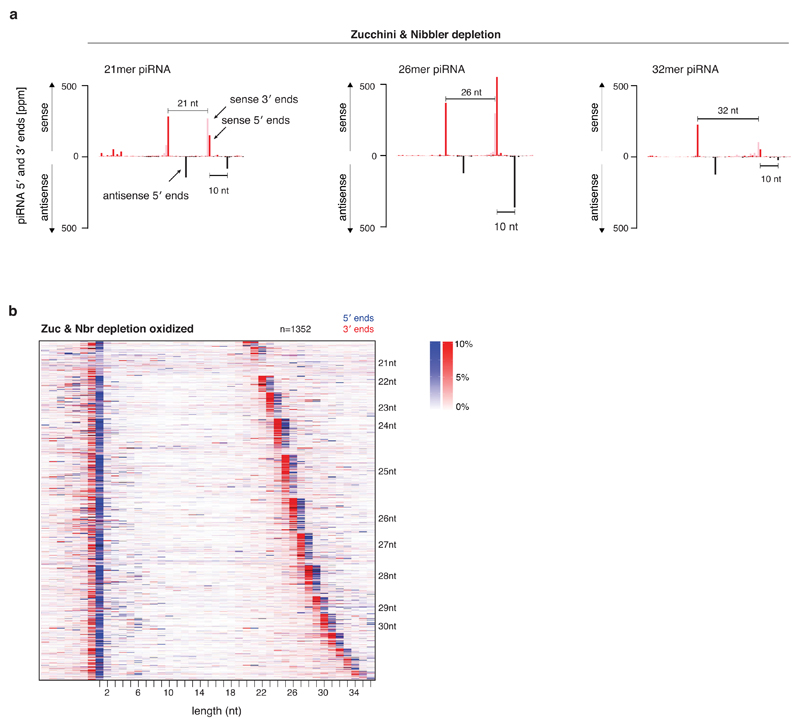
a) Shown are the mappings of those piRNAs (5' ends in black; 3' ends in gray) that were probed by Northern blots in Fig. 4f. The 5' ends of antisense piRNAs (red) map precisely 10nt downstream of the predicted slicer sites. Mappings were normalized to 1 million sequenced miRNA reads.
b) An oxidized library control of the heat map shown in Fig. 4d, showing the precise coupling of piRNA 5' ends (blue) and 3' ends (red) in Zucchini/Nibbler-depleted ovaries.
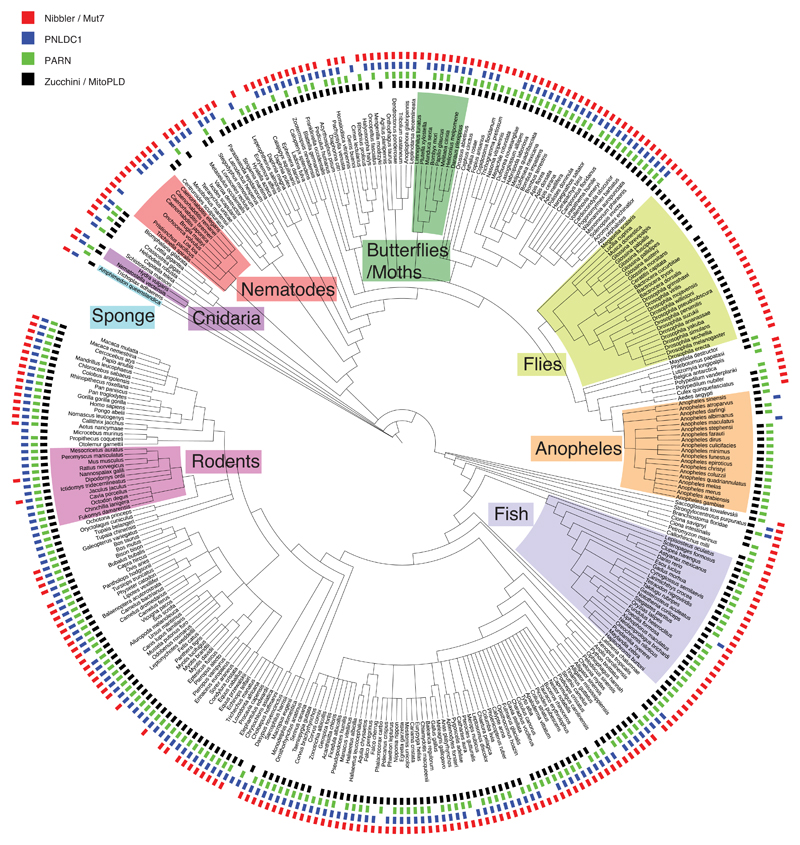
Figure ED10. Conservation of Zucchini/MitoPLD, PARN, PNLDC1, and Nibbler/Mut-7 across metazoa.
328 metazoan species are displayed in a phylogenetic tree (generated using iTOL). The presence of indicated orthologs in each species is marked (black: Zucchini/MitoPLD, green: PARN, blue: PNLDC1, red: Nibbler/Mut-7). Taxonomic groups mentioned in the text are highlighted (for details see Methods).
Supplementary Material
SUPPLEMENTARY INFORMATION is available in the online version of the paper.
Acknowledgements
We thank all lab members for help and discussions, P. Duchek, J. Gokcezade and K. Meixner for generating fly lines, M. Novatchkova for help on the conservation analysis of nucleases, the VBCF NGS facility for sequencing, and the MFPL monoclonal facility for Nibbler antibody. This work was supported by the Austrian Academy of Sciences, the European Community's 7th Framework Program (ERC-StG-260711; ERC-StG-338252), the Austrian Science Fund (Y510-B12; F4303-B09; W12-7-B09; Y733-B22), and an HFSP postdoctoral fellowship to F.M.
Footnotes
Author Contributions
F.M. made the initial observation that Nibbler trims Ago3-bound piRNAs, J.S. and R.H. did all experiments and did the computational analysis with the help of D.H. All authors designed the experiments and wrote the paper.
The authors declare that they have no conflict of interest.
References
- 1.Ghildiyal M, Zamore PD. Small silencing RNAs: an expanding universe. Nature Reviews Genetics. 2009;10:94–108. doi: 10.1038/nrg2504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Malone CD, Hannon GJ. Small RNAs as guardians of the genome. Cell. 2009;136:656–668. doi: 10.1016/j.cell.2009.01.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Iwasaki YW, Siomi MC, Siomi H. PIWI-Interacting RNA: Its Biogenesis and Functions. Annual Review of Biochemistry. 2015;84:405–433. doi: 10.1146/annurev-biochem-060614-034258. [DOI] [PubMed] [Google Scholar]
- 4.Nishimasu H, et al. Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature. 2012;491:284–287. doi: 10.1038/nature11509. [DOI] [PubMed] [Google Scholar]
- 5.Ipsaro JJ, Haase AD, Knott SR, Joshua-Tor L, Hannon GJ. The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature. 2012;491:279–283. doi: 10.1038/nature11502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Brennecke J, et al. Discrete Small RNA-Generating Loci as Master Regulators of Transposon Activity in Drosophila. Cell. 2007;128:1089–1103. doi: 10.1016/j.cell.2007.01.043. [DOI] [PubMed] [Google Scholar]
- 7.Gunawardane LS, et al. A slicer-mediated mechanism for repeat-associated siRNA 5' end formation in Drosophila. Science. 2007;315:1587–1590. doi: 10.1126/science.1140494. [DOI] [PubMed] [Google Scholar]
- 8.Han BW, Wang W, Li C, Weng Z, Zamore PD. Noncoding RNA. piRNA-guided transposon cleavage initiates Zucchini-dependent, phased piRNA production. Science. 2015;348:817–821. doi: 10.1126/science.aaa1264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mohn F, Handler D, Brennecke J. Noncoding RNA. piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science. 2015;348:812–817. doi: 10.1126/science.aaa1039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Han BW, Hung JH, Weng Z, Zamore PD, Ameres SL. The 3'-to-5' exoribonuclease Nibbler shapes the 3' ends of microRNAs bound to Drosophila Argonaute1. Current biology. 2011;21:1878–1887. doi: 10.1016/j.cub.2011.09.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu N, et al. The exoribonuclease Nibbler controls 3' end processing of microRNAs in Drosophila. Current Biology. 2011;21:1888–1893. doi: 10.1016/j.cub.2011.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Feltzin VL, et al. The exonuclease Nibbler regulates age-associated traits and modulates piRNA length in Drosophila. Aging Cell. 2015;14:443–452. doi: 10.1111/acel.12323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang H, et al. Antagonistic roles of Nibbler and Hen1 in modulating piRNA 3' ends in Drosophila. Development. 2016;143:530–539. doi: 10.1242/dev.128116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Vourekas A, et al. Mili and Miwi target RNA repertoire reveals piRNA biogenesis and function of Miwi in spermiogenesis. Nature Structural & Molecular Biology. 2012;19:773–781. doi: 10.1038/nsmb.2347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kawaoka S, Izumi N, Katsuma S, Tomari Y. 3' end formation of PIWI-interacting RNAs in vitro. Molecular Cell. 2011;43:1015–1022. doi: 10.1016/j.molcel.2011.07.029. [DOI] [PubMed] [Google Scholar]
- 16.Olivieri D, Senti KA, Subramanian S, Sachidanandam R, Brennecke J. The cochaperone shutdown defines a group of biogenesis factors essential for all piRNA populations in Drosophila. Molecular Cell. 2012;47:954–969. doi: 10.1016/j.molcel.2012.07.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Izumi N, et al. Identification and Functional Analysis of the Pre-piRNA 3' Trimmer in Silkworms. Cell. 2016;164:962–973. doi: 10.1016/j.cell.2016.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Saxe JP, Chen M, Zhao H, Lin H. Tdrkh is essential for spermatogenesis and participates in primary piRNA biogenesis in the germline. EMBO J. 2013;32:1869–1885. doi: 10.1038/emboj.2013.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sato K, et al. Krimper Enforces an Antisense Bias on piRNA Pools by Binding AGO3 in the Drosophila Germline. Molecular Cell. 2015;59:553–563. doi: 10.1016/j.molcel.2015.06.024. [DOI] [PubMed] [Google Scholar]
- 20.Webster A, et al. Aub and Ago3 Are Recruited to Nuage through Two Mechanisms to Form a Ping-Pong Complex Assembled by Krimper. Molecular Cell. 2015;59:564–575. doi: 10.1016/j.molcel.2015.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Horwich MD, et al. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Current Biology. 2007;17:1265–1272. doi: 10.1016/j.cub.2007.06.030. [DOI] [PubMed] [Google Scholar]
- 22.Saito K, Sakaguchi Y, Suzuki T, Siomi H, Siomi MC. Pimet, the Drosophila homolog of HEN1, mediates 2'-O-methylation of Piwi- interacting RNAs at their 3' ends. Genes Dev. 2007;21:1603–1608. doi: 10.1101/gad.1563607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Grimson A, et al. Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature. 2008;455:1193–1197. doi: 10.1038/nature07415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Sarkies P, et al. Ancient and novel small RNA pathways compensate for the loss of piRNAs in multiple independent nematode lineages. PLoS Biology. 2015;13:e1002061. doi: 10.1371/journal.pbio.1002061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Tang W, Tu S, Lee HC, Weng Z, Mello CC. The RNase PARN-1 Trims piRNA 3' Ends to Promote Transcriptome Surveillance in C. elegans. Cell. 2016;164:974–984. doi: 10.1016/j.cell.2016.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gu W, et al. Distinct argonaute-mediated 22G-RNA pathways direct genome surveillance in the C. elegans germline. Molecular Cell. 2009;36:231–244. doi: 10.1016/j.molcel.2009.09.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Senti KA, Jurczak D, Sachidanandam R, Brennecke J. piRNA-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear piRNA repertoire. Genes Dev. 2015;29:1747–1762. doi: 10.1101/gad.267252.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang W, et al. Slicing and Binding by Ago3 or Aub Trigger Piwi-Bound piRNA Production by Distinct Mechanisms. Molecular Cell. 2015;59:819–830. doi: 10.1016/j.molcel.2015.08.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Markstein M, Pitsouli C, Villalta C, Celniker SE, Perrimon N. Exploiting position effects and the gypsy retrovirus insulator to engineer precisely expressed transgenes. Nature Genetics. 2008;40:476–483. doi: 10.1038/ng.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gokcezade J, Sienski G, Duchek P. Efficient CRISPR/Cas9 plasmids for rapid and versatile genome editing in Drosophila. G3. 2014;4:2279–2282. doi: 10.1534/g3.114.014126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ni JQ, et al. A genome-scale shRNA resource for transgenic RNAi in Drosophila. Nature Methods. 2011;8:405–407. doi: 10.1038/nmeth.1592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Venken KJ, et al. Versatile P[acman] BAC libraries for transgenesis studies in Drosophila melanogaster. Nature Methods. 2009;6:431–434. doi: 10.1038/nmeth.1331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ejsmont RK, et al. Recombination-mediated genetic engineering of large genomic DNA transgenes. Methods in Molecular Biology. 2011;772:445–458. doi: 10.1007/978-1-61779-228-1_26. [DOI] [PubMed] [Google Scholar]
- 34.Jayaprakash AD, Jabado O, Brown BD, Sachidanandam R. Identification and remediation of biases in the activity of RNA ligases in small-RNA deep sequencing. Nucleic Acids Research. 2011;39:e141. doi: 10.1093/nar/gkr693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mohn F, Sienski G, Handler D, Brennecke J. The rhino-deadlock-cutoff complex licenses noncanonical transcription of dual-strand piRNA clusters in Drosophila. Cell. 2014;157:1364–1379. doi: 10.1016/j.cell.2014.04.031. [DOI] [PubMed] [Google Scholar]
- 36.Pall GS, Codony-Servat C, Byrne J, Ritchie L, Hamilton A. Carbodiimide-mediated cross-linking of RNA to nylon membranes improves the detection of siRNA, miRNA and piRNA by northern blot. Nucleic Acids Research. 2007;35:e60. doi: 10.1093/nar/gkm112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kriventseva EV, et al. OrthoDB v8: update of the hierarchical catalog of orthologs and the underlying free software. Nucleic Acids Research. 2015;43:D250–256. doi: 10.1093/nar/gku1220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Letunic I, Bork P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics. 2007;23:127–128. doi: 10.1093/bioinformatics/btl529. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All sequence data that support the findings of this study (see SI) have been deposited in NCBI GEO with the accession code https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE83698. All fly lines used in this study are available from VDRC (http://stockcenter.vdrc.at/control/main). Source data for all gel images are provided with the paper (see SI).