Abstract
A variety of stress signals stimulate cardiac myocytes to undergo hypertrophy. Persistent cardiac hypertrophy is associated with elevated risk for the development of heart failure. Recently, we showed that class II histone deacetylases (HDACs) suppress cardiac hypertrophy and that stress signals neutralize this repressive function by triggering phosphorylation- and CRM1-dependent nuclear export of these chromatin-modifying enzymes. However, the identities of cardiac HDAC kinases have remained unclear. Here, we demonstrate that signaling by protein kinase C (PKC) is sufficient and, in some cases, necessary to drive nuclear export of class II HDAC5 in cardiomyocytes. Inhibition of PKC prevents nucleocytoplasmic shuttling of HDAC5 in response to a subset of hypertrophic agonists. Moreover, a nonphosphorylatable HDAC5 mutant is refractory to PKC signaling and blocks cardiomyocyte hypertrophy mediated by pharmacological activators of PKC. We also demonstrate that protein kinase D (PKD), a downstream effector of PKC, directly phosphorylates HDAC5 and stimulates its nuclear export. These findings reveal a novel function for the PKC/PKD axis in coupling extracellular cues to chromatin modifications that control cellular growth, and they suggest potential utility for small-molecule inhibitors of this pathway in the treatment of pathological cardiac gene expression.
Coordinated changes in gene transcription during cell growth and differentiation require mechanisms for coupling intracellular signaling pathways with the genome. The acetylation of nucleosomal histones has emerged as a central mechanism in the control of gene transcription during such cellular transitions (20). Acetylation of histones by histone acetyltransferases promotes transcription by relaxing chromatin structure, whereas histone deacetylation by histone deacetylases (HDACs) reverses this process, resulting in transcriptional repression. How these chromatin-modifying enzymes are linked to, and controlled by, intracellular signaling is only beginning to be understood.
There are two classes of HDACs that can be distinguished by their structures and expression patterns. Class I HDACs (HDAC1, HDAC2, and HDAC3) are expressed ubiquitously and are composed mainly of a catalytic domain (13). In contrast, class II HDACs (HDAC4, HDAC5, HDAC7, and HDAC9) display more restricted expression patterns and contain an N-terminal extension, which mediates interactions with other transcriptional cofactors and confers responsiveness to calcium-dependent signaling (12, 25, 33). Signaling by calcium/calmodulin-dependent protein kinase (CaMK) results in phosphorylation of the N termini of class II HDACs, which govern their intracellular localization and interactions with other factors (29, 32). Phosphorylation of signal-responsive serine residues creates docking sites for the 14-3-3 family of chaperone proteins, which promote shuttling of HDACs from the nucleus to the cytoplasm in a CRM1-dependent fashion (14, 21, 30, 31, 48).
CaMK signaling to class II HDACs governs the activity of the myocyte enhancer factor-2 (MEF2) transcription factor, which plays central roles in the control of muscle-specific and stress-responsive gene expression (32). Class II HDACs interact with MEF2 through a short motif near their N termini; this interaction represses the expression of MEF2 target genes. Phosphorylation of class II HDACs, in response to CaMK signaling, results in their dissociation from MEF2 with consequent potentiation of MEF2 activity. Thus, class II HDACs provide a calcium-sensitive switch to control large sets of genes regulated by MEF2.
Recently, we reported that class II HDACs act as signal-responsive repressors of cardiac hypertrophy, which is triggered by calcium-sensitive signals (28, 49). Hypertrophy of cardiomyocytes is accompanied by an increase in cell size, assembly of sarcomeres, and activation of a fetal gene program (8, 27). We have shown that signal-resistant HDAC mutants block cardiomyocyte hypertrophy in response to diverse agonists and that mice lacking HDAC9 are sensitized to hypertrophic stimuli (6, 49). These findings suggest that HDAC phosphorylation is an essential step in coupling stress signals to the hypertrophic gene program. Induction of cardiac hypertrophy is accompanied by the posttranslational activation of MEF2, which is presumed to occur, at least in part, as a consequence of the dissociation and nuclear export of class II HDACs (38). CaMK can also promote skeletal myogenesis by relieving HDAC repression of MEF2 activity (26, 29).
Numerous signaling pathways have been implicated in cardiac hypertrophy (11, 27). Because of the critical role of HDAC phosphorylation in regulating myocyte differentiation and hypertrophy, there has been intense interest in identifying the kinase(s) responsible for class II HDAC nuclear export and inactivation. To further define the signaling pathways leading to the phosphorylation of class II HDACs, we examined the potential of multiple kinase pathways to stimulate HDAC5 nuclear export. Here we show that the protein kinase C (PKC) pathway promotes nuclear export of HDAC5 by stimulating phosphorylation of the 14-3-3 docking sites. Signal-resistant HDAC5 blocks cardiomyocyte hypertrophy stimulated by PKC activators. Conversely, PKC inhibition selectively blocks HDAC5 export and hypertrophy in response to a subset of hypertrophic agonists. In addition, PKCμ/protein kinase D (PKD), which acts as a downstream effector kinase of PKC, stimulates the nuclear export of HDAC5. These findings establish class II HDACs as critical effectors in the pathway whereby PKC signaling leads to cardiac hypertrophy.
MATERIALS AND METHODS
Chemical reagents and plasmids.
Phorbol 12-myristate 13-acetate (PMA), 8-Br-cAMP, pCPT-cGMP, and anisomycin were obtained from Sigma Chemical Co. (St. Louis, Mo.). The following kinase inhibitors were purchased from the indicated vendors: bisindolylmaleimide I (Bis I) and Gö6976 (A.G. Scientific, San Diego, Calif.); KN93, SB216763, and wortmannin (BIOMOL, Plymouth Meeting, Pa.); Gö6983, staurosporine, PD98059, U1026, Y-27632, rapamycin, and DAG kinase inhibitor II (Calbiochem). KN93, wortmannin, and staurosporine were used at 1 μM. U1026, HA1077, Y-27632, DAG kinase inhibitor II, SB216763, Bis I, and Gö6976 were used at 10 μM. Rapamycin was used at 30 ng/ml. Phenylephrine (PE) and endothelin-1 (ET-1) were purchased from Sigma. Mammalian expression vectors encoding PKD isoforms were provided by A. Toker (Harvard University) and have been described elsewhere (45). Plasmids encoding the catalytic domains of PKCα or PKCδ were provided by J. W. Son (Inha University, Inchon, Korea). A plasmid encoding constitutively active PKCɛ was provided by P. Ping (University of California Los Angels), and P. Goldspink (University of Illinois, Chicago, Ill.) and S. Shaw (National Cancer Institute, Bethesda, Md.) provided vectors for PKCβII and PKCθ, respectively.
Generation of phospho-HDAC-specific antibodies.
Research Genetics, Huntsville, Ala., generated antibodies against human HDAC5 phosphorylated at serine 259. Rabbits were immunized with a keyhole limpet hemocyanin-conjugated peptide corresponding to amino acids 251 to 269 of HDAC5 and containing a phosphate group at position 259, as indicated by [p] in the following sequence: (DFPLRKTAS[p]EPNLKVRSRI). Phosphospecific antibodies were purified from crude rabbit serum by sequential negative and positive affinity purification with nonphosphorylated and phosphorylated peptide, respectively.
Cell culture and transfection assays.
COS cells were maintained in Dulbecco minimal essential medium (DMEM) with 10% fetal bovine serum (FBS), 2 mM l-glutamine, and penicillin-streptomycin. COS cells were transfected with Fugene 6 (Roche Molecular Biochemicals) as specified by the manufacturer. For HDAC localization experiments, cells were treated 16 to 24 h after transfection with 100 nM PMA, 1 μM ionomycin, 1 μM 8-Br-cAMP, 1 μM pCPT-GMP, or 1 μM anisomycin. Where noted, specific protein kinase inhibitors were added 30 min prior to the addition of any chemical stimulus. Green fluorescent protein (GFP)-HDAC5 was visualized by standard fluorescent microscopic techniques. For indirect immunofluorescence of FLAG-HDAC5, COS cells were seeded on glass coverslips, transfected, and treated as above. After specific treatment, cells were fixed with 10% buffered formalin and exposed to FLAG M2 antibody (1:200 dilution; Sigma) in phosphate-buffered saline (PBS) containing 3% bovine serum albumin and 0.1% Nonidet P-40. Secondary fluorescein-conjugated antibody (Vector Laboratories) was also used at a concentration of 1:200. Cardiomyocytes were stained for sarcomeres and atrial natriuretic factor (ANF) by indirect immunofluorescence detection as above, with antibodies directed against sarcomeric α-actinin (Sigma) and ANF (Peninsula Laboratories), respectively.
Cardiac myocyte culture and adenovirus infection.
Neonatal rat cardiac myocytes (NRVMs) were isolated from 1- to 2-day-old Sprague-Dawley rats as previously described (2, 35). For adenovirus production, cDNAs encoding LacZ or FLAG-tagged HDAC5 containing alanines in place of serines 259 and 498 (S259/498A) were subcloned into the pACCMV vector and cotransfected with pJM17 into 293 cells. Primary lysates were used to reinfect 293 cells, and viral plaques were obtained using the agar overlay method. Complementary DNA for full-length human HDAC5 (encoding 1,122 amino acids) was fused to sequences encoding enhanced GFP (Clontech) in pcDNA3.1+ (Invitrogen). The resultant construct encodes GFP fused in frame to the amino terminus of HDAC5. A construct encoding GFP fused to HDAC5 (S259/498A) was generated in the same manner. GFP-HDAC5 cDNAs were subcloned into pACCMV for adenovirus production. Clonal populations of adenoviruses were amplified and subjected to titer determination.
Coimmunoprecipitation assays.
FLAG-HDAC5 expression plasmid was transfected into COS cells treated as described above. Treated cells were harvested in 50 mM Tris (pH 7.4)-150 mM NaCl--1 mM EDTA—1% Triton X-100 and further disrupted by passage through a 22-gauge needle, and cell debris removed by centrifugation. FLAG-HDAC5 was immunoprecipitated with M2-agarose conjugate (Sigma) and thoroughly washed with lysis buffer. Bound proteins were resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and Western blot analysis was performed using FLAG M2 (Sigma) or 14-3-3 antibody (Santa Cruz Biotechnology). For studies with NRVMs, whole-cell protein extracts were prepared from cells expressing GFP-HDAC5 by using the same buffer supplemented with protease inhibitor cocktail (Complete; Roche), 1 mM phenylmethylsulfonyl fluoride, and phosphatase inhibitors (1 mM sodium pyrophosphate, 2 mM sodium fluoride, 10 mM β-glycerol phosphate, 1 mM sodium molybdate, 1 mM sodium orthovanadate). Lysates were sonicated briefly and clarified by centrifugation. For immunoprecipitation, protein lysates were exposed to HDAC5-specific antiserum (29) and protein G-Sepharose beads (Amersham Biosciences). Immunoprecipitates were washed five times with lysis buffer, resolved by SDS-PAGE, and immunoblotted with mouse monoclonal antibodies specific for either GFP (BD Biosciences; 1:2,500 dilution) or 14-3-3 (Santa Cruz [H-8]; 1:1,000 dilution). Associated PKD was detected by immunoblotting with rabbit polyclonal antibodies against either PKD-1 or PKD-1 phosphorylated at serine 916 (Cell Signaling Technologies), employed at 1:1,000 dilutions.
In vitro kinase assays.
FLAG-HDAC5 was immunoprecipitated with anti-FLAG M2 antibody, as described above. Bound FLAG-HDAC5 was washed and equilibrated with kinase buffer (25 mM Tris [pH 7.4], 10 mM MgCl2, 1 mM dithiothreitol). Following equilibration, kinase reaction mix was added (kinase buffer plus 0.1 mM ATP and 50 μCi of [γ-32P]-ATP). Kinase reactions were carried out at 30°C for 30 min and terminated by the addition of an equal volume of 2× SDS-PAGE loading buffer. Phosphorylated proteins were resolved by SDS-PAGE and visualized by autoradiography.
GFP-HDAC5 localization studies.
For analysis of GFP-HDAC5 in NRVMs, cells were plated in the presence of adenovirus (multiplicity of infection, ∼50 to 100) on gelatin-coated 96-well dishes (Costar; 104 cells/well) containing DMEM plus 10% FBS, 2 mM l-glutamine, and penicillin-streptomycin. After overnight culture, the cells were washed with serum-free medium and maintained in 100 μl of DMEM supplemented with 0.1% Neutridoma-SP (Roche Applied Science), which contains albumin, insulin, transferrin, and other defined organic and inorganic compounds. Following culture in serum-free medium (3 h), the cells were exposed to kinase inhibitors (for 30 min) prior to stimulation with the agonist for 2.5 h. The cells were washed with PBS and fixed with 10% formalin in PBS containing Hoechst dye 33342 (H-3570; Molecular Probes). Images were captured at a magnification of ×40, using a fluorescence microscope (Nikon Eclipse TS100) equipped with a digital camera (Photometrics CoolSNAP HQ) and MetaMorph imaging software. The relative abundance of GFP-HDAC5 in the nucleus versus the cytoplasm was quantified by using the high-content imaging system (Cellomics, Inc., Pittsburgh, Pa.), which demarcates nuclei based on Hoechst fluorescence and defines a cytoplasmic ring based on these nuclear dimensions. Values for HDAC5 localization represent averages from a minimum of 400 randomly captured cells per experimental condition.
RNA analysis.
NRVMs were plated on gelatin-coated 10-cm dishes (2 × 106 cells/dish). Following the indicated treatments, RNA was isolated from cardiomyocytes by using Trizol reagent (Gibco/BRL). Total RNA (2 μg) was vacuum blotted onto nitrocellulose membranes (Bio-Rad) by using a 96-well format dot blotter (Bio-Rad). Membranes were blocked in 4× SSC (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate) containing 1% SDS, 5× Denhardt's reagent, 0.05% sodium pyrophosphate, and 100 μg of sonicated salmon sperm DNA per ml (4 h at 50°C) and incubated with 32P-end-labeled oligonucleotide probes (106 cpm/ml; 14 h at 50°C). The sequences of the oligonucleotides were as follows: ANF, 5′-AATGTGACCAAGCTGCGTGACACACCACAAGGGCTTAGGATCTTTTGCGATCTGCTCAAG-3; brain natriuretic protein (BNP), 5′-TGAACTATGTGCCATCTTGGAATTTCGAAGTCTCTCCT-3′; α-SK-actin 5′-TGGAGCAAAACAGAATGGCTGGCTTTAATGCTTCAAGTTTTCCATTTCCTTTCCACAGGG-3′; glyceraldehyde-3-phosphate dehydrogenase (GAPDH), 5′-GGAACATGTAGACCATGTAGTTGAGGTCAATGAAG-3′. Blots were washed twice with 0.5× SSC containing 0.1% SDS (10 min at 50°C) and analyzed by autoradiography.
Mammalian two-hybrid analysis.
A mammalian expression vector encoding the GAL4 DNA binding domain fused to the amino terminus of human HDAC5 (amino acids 2 to 664) was generated in the pM1 expression vector (42). GAL4-HDAC5 fusions harboring alanine in place of serine 259 and/or 498 were constructed in an analogous manner. A construct encoding the herpesvirus VP16 transcriptional activation domain fused to the amino terminus of 14-3-3 sigma was generated by employing pVP16 (Clontech). COS cells were transiently transfected with vectors for GAL4-HDAC5, VP16-14-3-3, and a luciferase reporter gene under the control of five copies of a GAL4 DNA binding site (5XUAS-luciferase) in the absence or presence of a construct for constitutively active PKD-1. At 48 h posttransfection, the cells were harvested and luciferase levels were quantified by using the luciferase assay kit (Promega).
RESULTS
A PKC-dependent pathway stimulates nuclear export of HDAC5.
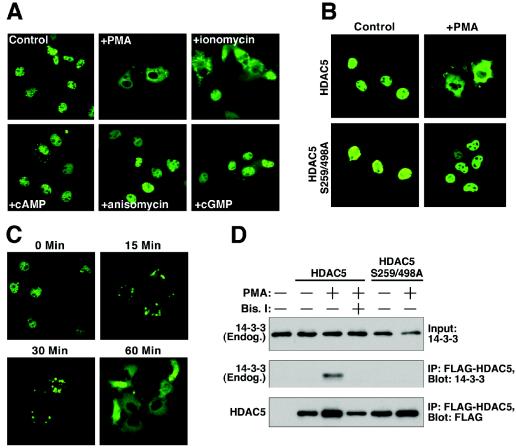
To further define the signaling pathways leading to phosphorylation and nuclear export of class II HDACs, we tested a variety of activators of protein kinase pathways for their ability to stimulate nuclear export of HDAC5 in COS cells. HDAC5 is located primarily in the nuclei of COS cells, allowing for a convenient system to assess nuclear export. Activators of PKA (8-Br-cAMP), PKG (pCPT-GMP), PKC (PMA), CaMK (ionomycin), and Jun N-terminal kinase (anisomycin) were tested for their ability to trigger nuclear export of HDAC5 fused to GFP. Among these compounds, only ionomycin and PMA stimulated nuclear export of GFP-HDAC5 (Fig. 1A). PMA was a more potent stimulator of export than ionomycin at the concentrations tested.
FIG. 1.
PKC-dependent nuclear export of HDAC5. (A) COS cells were cultured in six-well dishes, transfected with a GFP-HDAC5 expression vector (1 μg), and stimulated with the indicated compounds, as described in Materials and Methods. At 1 h after addition of compounds, GFP-HDAC5 distribution was determined by fluorescence microscopy. PMA stimulation resulted in complete relocalization of GFP-HDAC5 from the nucleus to the cytoplasm, while ionomycin triggered a partial response. (B) COS cells were transfected with expression vectors encoding FLAG-tagged versions of either HDAC5 or an HDAC5 mutant harboring alanines in place of serines 259 and 498 (HDAC5 S259/498A) (1 μg each). The cells were stimulated with PMA for 1 h, and HDAC5 distribution was determined by indirect immunofluorescence with anti-FLAG primary antibody and fluorecein-conjugated secondary antibody. HDAC5 S259/498A is refractory to PMA stimulation. (C) COS cells were transfected with GFP-HDAC5 encoding expression vector (1 μg) and stimulated with PMA for the indicated times. (D) COS cells were transiently transfected with expression vectors encoding FLAG-tagged versions of HDAC5 or HDAC5 S259/498A (1 μg each). The cells were pretreated with the PKC inhibitor Bis I at 10 μM for 30 min and stimulated with PMA for 30 min, as indicated. Association of FLAG-HDAC5 with endogenous 14-3-3 was detected by sequential immunoprecipitation (IP) and immunoblotting (Blot).
Nuclear export of HDAC5 and other class II HDACs in response to CaMK signaling requires two serines located in the N-terminal regions of the HDAC proteins (29, 30). An HDAC5 mutant (HDAC5 S259/498A) containing alanine substitutions of these serine residues (residues 259 and 498) was not exported in response to PMA treatment (Fig. 1B), verifying the requisite role of these sites for responsiveness to PKC activation. Of note, FLAG- and GFP-tagged versions of HDAC5 were equally responsive to PMA stimulation (compare Fig. 1A and B). Nuclear export of HDAC5 was initiated within 15 min following addition of PMA and was complete by 30 min (Fig. 1C).
Phosphorylation of serines 259 and 498 in HDAC5 creates docking sites for 14-3-3 proteins, which escort HDAC5 to the cytoplasm (14, 30). To further confirm that PMA promoted the phosphorylation of these sites, we analyzed the interaction of HDAC5 with 14-3-3 in coimmunoprecipitation assays. As shown in Fig. 1D, the association of 14-3-3 with HDAC5 was enhanced in the presence of PMA. In contrast, HDAC5 S259/498A failed to respond to PMA and did not associate with 14-3-3. We conclude that PKC signaling leads to the phosphorylation of serines 259 and 498 of HDAC5 and consequent nuclear export through a 14-3-3-dependent mechanism.
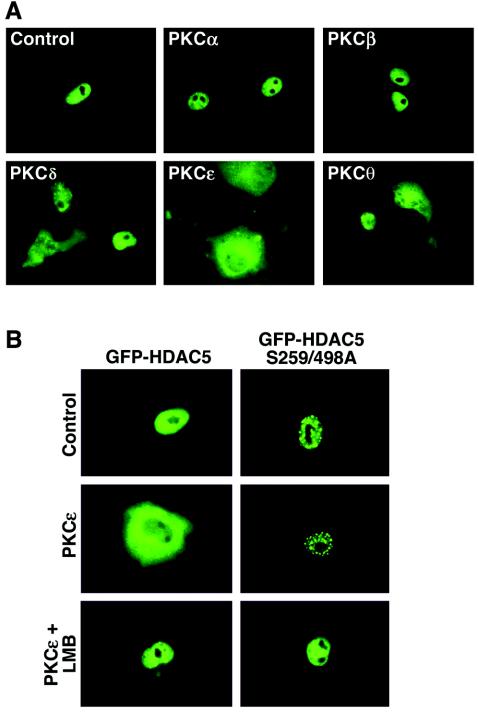
Since non-PKC targets of PMA have been identified, we used a panel of expression vectors encoding distinct PKCs to confirm that PKC signaling is sufficient to trigger nuclear export of HDAC5. As shown in Fig. 2A, overexpression of the calcium-dependent PKCα and PKCβ had no effect on the subcellular localization of HDAC5. In contrast, calcium-independent forms of PKC, PKCδ, PKCɛ, and PKCθ, each triggered nuclear export of HDAC5, albeit to different degrees. In these experiments with CV-1 cells, PKCɛ was the most efficacious HDAC5 export kinase. PKCɛ-mediated nuclear export of HDAC5 was dependent on phosphorylation of the 14-3-3 target sites at serines 259 and 498 and was blocked by the CRM1 antagonist, leptomycin B (Fig. 2B).
FIG. 2.
Calcium-independent PKCs trigger the nuclear export of HDAC5. (A) CV-1 cells were transiently transfected with an expression vector encoding GFP-HDAC5 and constructs for PKCα, PKCβ, PKCδ, PKCɛ, or PKCθ (0.5 μg each). The effects of PKC overexpression were assessed by fluorescence microscopy 24 h post-transfection. (B) CV-1 cells were transfected with vectors encoding either GFP-HDAC5 or GFP-HDAC5 S259/498A in the absence or presence of a vector for constitutively active PKCɛ (0.5 μg each). Prior to analysis, some cells received 18.5 nM leptomycin B (LMB) for 1 h.
PKC-dependent nuclear export of HDAC5 in cardiac myocytes.
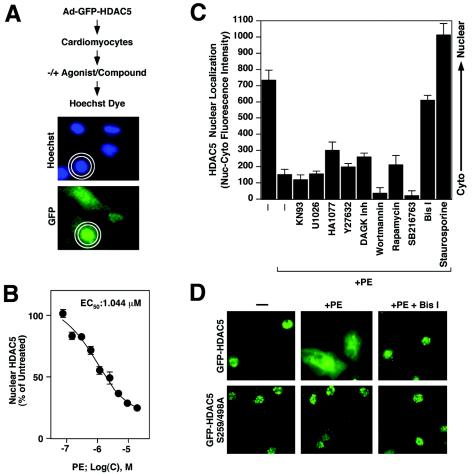
To begin to address the role of PKC signaling in control of HDAC5 trafficking during cardiac hypertrophy, we developed a quantitative assay to measure agonist-dependent nuclear export of HDAC5 in primary cardiomyocytes. This assay employs the Cellomics high-content imaging system, which randomly captures images of fluorescent cells and quantifies nuclear and cytoplasmic GFP fluorescence intensity, providing an objective readout of nuclear export (Fig. 3A). To validate the assay, NRVMs were infected with an adenovirus expressing GFP-HDAC5 (Ad-GFP-HDAC5) and stimulated with increasing doses of the α1-adrenergic agonist PE, a hypertrophic agonist that promotes nuclear export of HDAC5 (6). In unstimulated cells, ∼65 to 80% of HDAC5 was found in the nucleus, reflecting a low level of basal nuclear export of the protein. Conversely, HDAC5 S259/498A was found exclusively in the nuclear compartment. As shown in Fig. 3B, PE triggered the nuclear export of HDAC5 in a concentration-dependent manner. HDAC5 S259/498A was resistant to PE-mediated nuclear export (Fig. 3D and data not shown). The results of Cellomics analyses were confirmed by visual inspection of the cells (Fig. 3D).
FIG. 3.
PKC inhibition blocks PE-mediated nuclear export of HDAC5 in cardiomyocytes. (A) Schematic representation of a quantitative assay for HDAC5 nuclear export. NVRMs are cultured in 96-well dishes and infected with adenovirus encoding GFP-HDAC5. Cells are serum starved, subjected to agonists and inhibitors, fixed, and stained with Hoechst dye. The relative abundance of GFP-HDAC5 in the nucleus versus the cytoplasm is quantified by using the Cellomics high-content imaging system, which demarcates nuclei based on Hoechst fluorescence and defines a cytoplasmic ring based on these nuclear dimensions. Values represent the mean of nuclear minus cytoplasmic fluorescence intensity. (B) Assay validation. NRVMs were infected with adenovirus encoding GFP-HDAC5 and exposed to PE at concentrations ranging from 0.1 to 20 μM. Cells were prepared for Cellomics analysis following 2 h of stimulation. Mean nuclear minus cytoplasmic fluorescence intensity was determined for at least 50 cells/well in eight wells per condition (400 cells total). The value for untreated cells was set to 100%. PE triggered the dose-dependent nuclear export of HDAC5. EC50, 50% effective concentration. (C) NRVMs were infected with adenovirus encoding GFP-HDAC5 and pretreated with kinase inhibitors (the concentrations of the inhibitors are given in Materials and Methods). The subcellular distribution of HDAC5 was quantified following stimulation with PE (20 μM) for 2 h. Mean nuclear minus cytoplasmic fluorescence intensity was determined for at least 50 cells/well in eight wells per condition (400 cells total). Higher values indicate greater abundance of HDAC5 in the nucleus. Well-to-well standard deviations are shown. Only staurosporine and the PKC inhibitor Bis I effectively blocked HDAC5 nuclear export. (D) Representative images of GFP-HDAC5 and GFP-HDAC5 S259/498A in the absence and presence of Bis I.
Having established the validity of the quantitative nuclear export assay, we tested an array of inhibitors of different kinases for their abilities to block PE-induced translocation of HDAC5 out of the nucleus. The general serine/threonine protein kinase inhibitor staurosporine and the PKC inhibitor Bis I were effective in blocking the PE-dependent export of HDAC5 (Fig. 3C and D). In contrast, inhibitors of CaMK (KN93), MEK1 (U1026), Rho kinase (Y-27632), diacylglycerol kinase (diacylglycerol kinase inhibitor II), phosphatidylinositol 3-kinase (wortmannin), S6 kinase (rapamycin), glycogen synthase kinase (SB216763), or an inhibitor of protein kinase G, myosin light-chain kinase, and protein kinase A (HA1077) did not significantly affect PE-induced nuclear export of HDAC5. Bis I and staurosporine also blocked the low level of HDAC5 nuclear export observed in unstimulated cells (data not shown).
PKC signaling induces cardiac hypertrophy via HDAC phosphorylation.
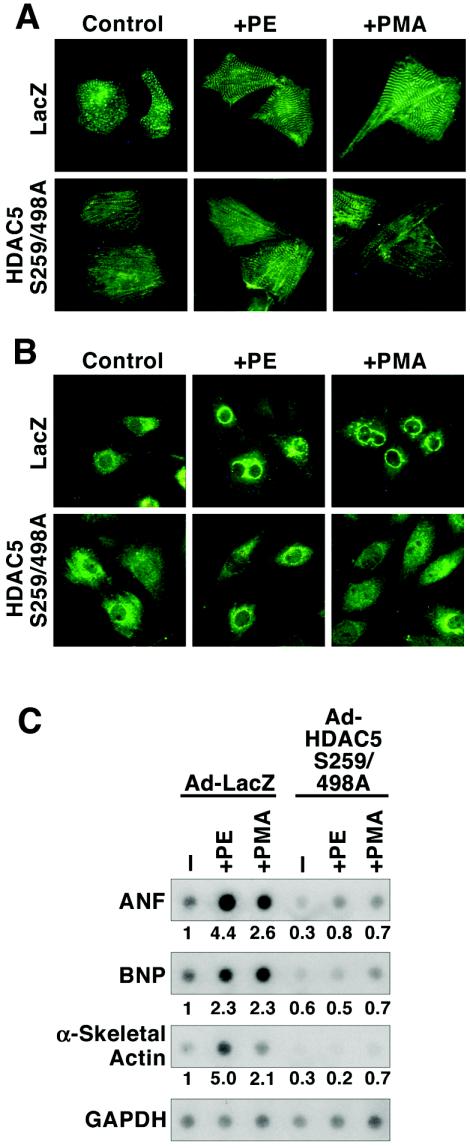
PKC activation is sufficient, and in some cases necessary, for cardiomyocyte hypertrophy (see Discussion). The above results implicate PKC-dependent nuclear export of HDAC5 or other class II HDACs in the development of cardiomyocyte hypertrophy. To address this possibility, we examined whether hypertrophy in response to PKC activation required phosphorylation and nuclear export of class II HDACs. NRVMs were infected with adenoviruses encoding the signal-resistant HDAC5 S259/498A mutant protein or LacZ as a control. As shown in Fig. 4A, expression of the HDAC5 S259/498A mutant in primary cardiomyocytes prevented sarcomere assembly and cell enlargement in response to PE or PMA.
FIG. 4.
Inhibition of PKC-mediated cardiac hypertrophy by signal-resistant HDAC5. NRVMs were cultured on six-well dishes and infected with adenoviruses (multiplicity of infection, 10) encoding a LacZ control (Ad-LacZ) or FLAG-tagged HDAC5 harboring alanines in place of serines 259 and 498 (Ad-HDAC5 S259/498A), which are required for 14-3-3-mediated nuclear export. Cells were treated with 20 μM PE or 100 nM PMA for 24 h prior to analysis. (A) Cells were fixed, and sarcomeres were visualized by indirect immunofluorescence with primary antibody specific for α-actinin and fluorescein-conjugated secondary antibody. (B) ANF protein was detected by indirect immunofluorescence with anti-ANF primary antibody. (C) Total RNA was harvested from cells and subjected to dot blot analysis with radiolabeled oligonucleotides specific for the indicated transcripts. RNA levels were quantified using a phosphorimager and are depicted as the fold change relative to amounts in unstimulated cells infected with Ad-LacZ. Values were normalized to GAPDH controls.
Cardiac hypertrophy is associated with reactivation of a pathological “fetal” gene program, which includes the genes encoding ANF, BNP, and α-skeletal actin. Agonist-dependent elevation of ANF expression can be examined by immunostaining cardiomyocytes with ANF-specific antibodies. As shown in Fig. 4B, prominent perinuclear ANF protein expression was observed in NRVMs treated with PE or PMA. Agonist-dependent induction of ANF expression was unaffected by ectopic expression of LacZ but was markedly reduced in the presence of signal-resistant HDAC5. In addition, nonphosphorylatable HDAC5 blocked PE- and PMA-mediated induction of ANF transcripts, as well as those for BNP and α-skeletal actin (Fig. 4C). Together, these results suggest that PKC signaling triggers cardiac hypertrophy in part by stimulating the nuclear export of class II HDACs.
Differential sensitivity of HDAC5 nuclear export to PKC inhibitors.
We next examined whether nuclear export of HDAC5 in response to other hypertrophic signals was also dependent on PKC signaling. ET-1 and FBS, which stimulate hypertrophy, also effectively promote nuclear export of HDAC5 in cardiac myocytes (Fig. 5A) (B. C. Harrison and T. A. McKinsey, submitted for publication). However, in contrast to its inhibitory effect on PE-dependent HDAC5 nuclear export, Bis I had no effect on the nuclear export of HDAC5 in response to ET-1 or FBS (Fig. 5B). These findings suggested that PE triggers different kinase pathways than ET-1 and FBS to promote the nuclear export of HDAC5.
FIG. 5.
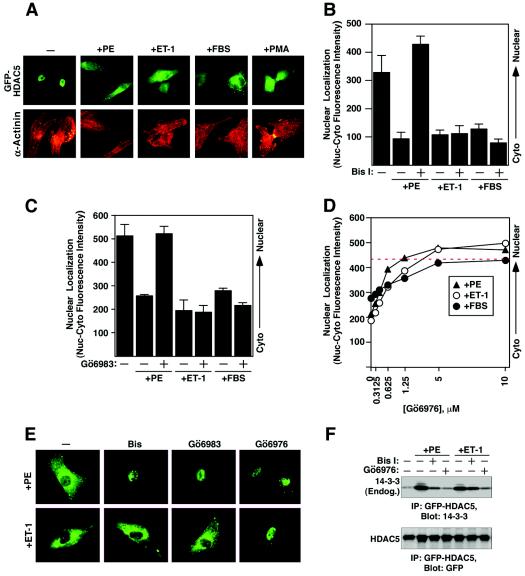
Differential repression of agonist-mediated nuclear export of HDAC5 by PKC inhibitors. (A) NRVMs were cultured in six-well dishes and infected with adenovirus encoding GFP-HDAC5. The cells were serum starved for 4 h prior to stimulation with 20 μM PE, 50 nM ET-1, 10% FBS, or 50 nM PMA for 2 h. The cells were fixed and stained with antibodies against sarcomeric α-actinin (red) to confirm that HDAC5 was being visualized in cardiomyocytes. (B) NRVMs were cultured in 96-well dishes and infected with adenovirus encoding GFP-HDAC5. Following serum starvation, infected cells were pretreated with 10 μM Bis I for 30 min and stimulated with the indicated agonists for 2 h. Nuclear export of HDAC5 was quantified using the Cellomics imaging system. Higher values indicate greater abundance of HDAC5 in the nucleus. (C) The experiment was performed as described for panel B, except that the cells received 10 μM Gö6983 before being treated with the agonists. (D) The experiment was performed as described for panel B, except that the cells received increasing doses of Gö6976 before being treated with the agonists. (E) Representative images from each treatment group were captured using a fluorescence microscope equipped with a digital camera. (F) NRVMs were infected with adenovirus encoding GFP-HDAC5 and cultured on 10-cm dishes. At 24 h postinfection, the cells were serum starved for 4 h and pretreated with 10 μM Bis I or 10 μM Gö6976 for 1 h before being stimulated with 20 μM PE or 50 nM ET-1 for 1 h. Whole-cell protein lysates were prepared and subjected to sequential immunoprecipitation (IP) and immunoblotting (Blot) as indicated.
To further examine the nature of the protein kinase effectors of the above hypertrophic agonists, we tested additional PKC inhibitors for their possible effects on HDAC5 nuclear export. The activity of Gö6983, another general inhibitor of PKCs, paralleled that of Bis I, inhibiting HDAC5 nuclear export in response to PE but not to ET-1 or FBS (Fig. 5C and E). In contrast, Gö6976, a specific inhibitor of the calcium-dependent PKCα and PKCβ isozymes, efficiently blocked the nuclear export of HDAC5 triggered by PE, ET-1 or FBS (Fig. 5D and E). The differential effects of the above inhibitors on HDAC5 nuclear export were paralleled by their effects on the association of 14-3-3 with HDAC5, an indicator of HDAC5 phosphorylation. Gö6976, but not Bis I, blocked the association of HDAC5 and 14-3-3 in response to both PE and ET-1 (Fig. 5F).
The ability of Gö6976, but not Bis I or Gö6983, to block HDAC5 nuclear export in response to multiple agonists was seemingly paradoxical, since the last two compounds block PKCα and PKCβ as effectively as Gö6976 does. However, this inhibitor profile was similar to that used by others to distinguish the actions of PKCα or PKCβ from PKD/PKCμ (51), which is sensitive to Gö6976 but not to Bis I or Gö6983 (Table 1) (15).
TABLE 1.
Effects of kinase inhibitors on agonist-dependent nuclear export of HDAC5
| Inhibitor | Inhibition of agonist:
|
Kinases inhibited | ||
|---|---|---|---|---|
| PE | ET-1 | FBS | ||
| Staurosporine | + | + | + | General Ser/Thr kinases |
| Bis I | + | − | − | General PKCs |
| Gö-6983 | + | − | − | General PKCs |
| Gö-6976 | + | + | + | Ca2+-dependent PKC-α and -β, PKD |
PKD stimulates the nuclear export of HDAC5.
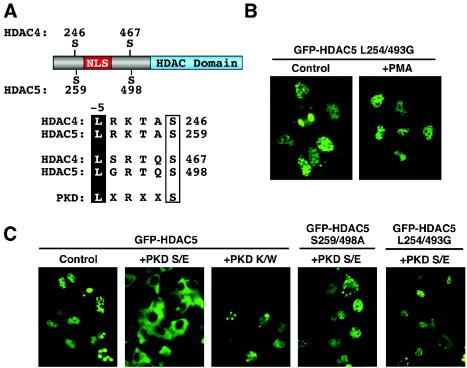
In light of the above results, which suggested the possible involvement of PKD in agonist-dependent nuclear export of HDAC5, we examined the amino acid sequence surrounding the signal-responsive serines in HDAC5 for a potential PKD consensus phosphorylation site. PKD has a strong preference for a leucine residue at the −5 position relative to the phosphorylated serine (37). HDAC5 contains a leucine at this position relative to both signal-responsive serine residues (Fig. 6A). Interestingly, the class II HDAC4, HDAC7, and HDAC9 also contain leucine at these positions.
FIG. 6.
PKD stimulates the nuclear export of HDAC5. (A) Amino acid sequences surrounding the regulatory phosphorylation sites of class II HDACs. NLS, nuclear localization signal; HDAC domain, deacetylase catalytic domain. The consensus target site for PKD is shown. Leucine at position −5 relative to the phosphorylation site is required for optimal PKD-directed phosphorylation of other proteins. (B) COS cells were transfected with an expression vector encoding GFP fused to HDAC5 harboring glycines in place of leucines 254 and 493 (L254/493G). At 24 h posttransfection, the cells were left untreated (control) or stimulated with PMA for 30 min. (C) COS cells were cotransfected with expression vectors (1 μg each) encoding GFP-HDAC5 or GFP-HDAC5 S259/498A and constitutively active (S/E) or catalytically inactive (K/W) forms of PKD. HDAC5 localization was determined at 24 h posttransfection.
To assess the importance of the leucine at position −5, we converted leucines 254 and 493 in HDAC5 to glycines, leaving the actual phosphorylation sites intact. This HDAC5 mutant (L254/493G) was constitutively localized to the nucleus and was completely refractory to PMA (Fig. 6B). Further support for the involvement of PKD in HDAC5 nuclear export was provided by transfection assays in which an activated form of PKD (PKD S/E), but not a catalytically inactive mutant (PKD K/W), effectively stimulated nuclear export of the protein (Fig. 6C). Mutation of the signal-responsive serine residues or the leucines at positions 254 and 493 of HDAC5 abolished nuclear export in response to PKD (Fig. 6C).
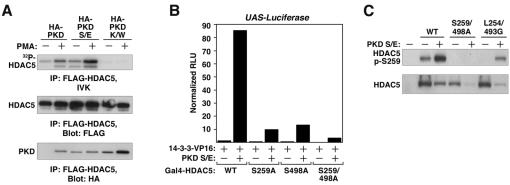
To further explore the potential role of PKD as an HDAC5 nuclear export kinase, we performed coimmunoprecipitation and in vitro kinase assays. Coimmunoprecipitation of HDAC5 and PKD followed by an in vitro kinase assay confirmed that PKD directly phosphorylated HDAC5. As shown in Fig. 7A, cotransfection of wild-type PKD resulted in little phosphorylation of HDAC5, and treatment of cells with PMA increased the degree of HDAC5 phosphorylation coincident with PKD binding. Binding and phosphorylation of HDAC5 by activated PKD S/E did not require PMA, although PMA treatment enhanced the phosphorylation of HDAC5, perhaps owing to the presence of endogenous PKD in immune complexes. No phosphorylation of HDAC5 was observed with the catalytically inactive mutant PKD K/W. Interestingly, however, PKD K/W bound to HDAC5 even in the absence of PMA.
FIG. 7.
PKD is an HDAC5 kinase. (A) COS cells were cotransfected with expression vectors (1 μg each) encoding FLAG-HDAC5 and HA-tagged versions of either wild-type, constitutively active (S/E), or catalytically inactive (K/W) PKD. At 24 h posttransfection, the cells were treated with PMA or vehicle control for 30 min. FLAG-HDAC5 was immunoprecipitated (IP) from whole-cell protein lysates and either incorporated into an in vitro kinase assay (IVK) or resolved by SDS-PAGE for Western blot analysis (Blot) to detect associated PKD, as indicated. Phosphorylated HDAC5 was resolved by SDS-PAGE and detected by autoradiography. (B) Mammalian two-hybrid assay. Expression vectors encoding the GAL4 DNA binding domain fused to HDAC5 (Gal4-HDAC5) or the indicated HDAC5 alanine substitution mutants were cotransfected into COS cells with a plasmid encoding 14-3-3 fused to the VP16 transcriptional activation domain (14-3-3-VP16), a Gal4-dependent luciferase reporter (UAS-Luciferase), and a vector encoding constitutively active PKD (S/E). PKD stimulates the association between HDAC5 and 14-3-3, which is dependent on the phospho-acceptors at positions 259 and 498. WT, wild type. (C) COS cells were left untransfected or transfected with expression vectors encoding FLAG-tagged versions of wild-type HDAC5, HDAC5 S259/498A, or HDAC5 L254/493G (1 μg) in the absence or presence of 1 μg of constitutively active PKD S/E. After 24 h of transfection, protein lysates were prepared and subjected to immunoblotting with antibodies directed against either HDAC5 phosphorylated at serine 259 (P-S259) or FLAG, to reveal the total levels of ectopic HDAC5.
PKD also increased the interaction between HDAC5 and 14-3-3, as assessed by a mammalian two-hybrid assay in which HDAC5 was fused to the GAL4 DNA binding domain and 14-3-3 was fused to the VP16 transcription activation domain (Fig. 7B). Mutation of either signal-responsive serine in HDAC5 markedly decreased the interaction between HDAC5 and 14-3-3, and mutation of both serines completely abolished the binding of HDAC5 to 14-3-3.
Finally, we assessed the ability of PKD to phosphorylate HDAC5 in cells by employing an antibody that specifically recognizes HDAC5 that has been phosphorylated at serine 259. HDAC5 was basally phosphorylated at serine 259, and phosphorylation at this site was dramatically increased by coexpression of constitutively active PKD S/E (Fig. 7C). HDAC5 S259/498A was not recognized by the phosphoantibody, establishing the specificity of the reagent. The L254/493G substitution abolished basal phosphorylation at serine 259 and reduced PKD-directed phosphorylation of this site. We predict that this level of HDAC5 L254/493G phosphorylation is insufficient to trigger HDAC5 nuclear export. In addition, it should be noted that the antibody does not detect phosphorylation of serine 498 in HDAC5, which is also required for efficient nuclear export of the protein. Together, these data suggest that PKD is capable of directly phosphorylating HDAC5.
PKD is a cardiac HDAC5 kinase.
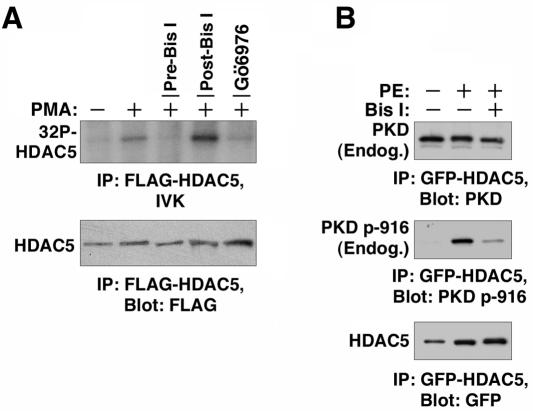
We next examined whether PKD could serve as an HDAC kinase in cardiomyocytes. Cells were infected with adenovirus encoding FLAG-HDAC5 and treated with PMA. Increased HDAC5 phosphorylation was observed in an in vitro kinase assay performed with FLAG-HDAC5 immunoprecipitated from PMA-treated cells (Fig. 8A). Incubation of the cells with Bis I before the addition of PMA blocked the phosphorylation of HDAC5. However, addition of Bis I directly to the kinase reaction mixture had no effect, while Gö6976 blocked the phosphorylation of HDAC5. These results suggest that PKD binds HDAC5 in cardiac myocytes and that Bis I blocks the PMA-induced activation of the kinase, while Gö6976 is able to directly inhibit HDAC5-bound PKD.
FIG. 8.
Association of endogenous PKD with HDAC5 in cardiomyocytes. (A) NRVMs were cultured on 10-cm dishes and infected with adenovirus encoding FLAG-HDAC5. At 24 h posttransfection, the cells were stimulated with PMA for 30 min and whole-cell protein lysates were prepared. Some cells were pretreated with 10 μM Bis I (pre-Bis I) for 30 min prior to PMA stimulation. FLAG-HDAC5 was immunoprecipitated (IP) and incorporated into in vitro kinase reaction mixtures supplemented with 10 μM Bis I (post-Bis I) or 10 μM Gö6976 (post-Gö6976), as indicated. Phosphorylation of HDAC5 was blocked when the cells were pretreated with Bis I (pre-Bis I). Gö6976 but not Bis I blocked phosphoryl transfer to HDAC5 when added directly to kinase reaction mixtures. (B) NRVMs were infected with adenovirus encoding GFP-HDAC5 and cultured on 10-cm dishes. At 24 h postinfection, the cells were serum starved for 4 h and pretreated with 10 μM Bis I for 30 min before being stimulated with 20 μM PE for 1 h. HDAC5 was immunoprecipitated from whole-cell lysates, and associated total PKD or PKD autophosphorylated at serine 916 (p-916) were detected by immunoblotting. Blots were reprobed with GFP-specific antibodies to determine the total amounts of immunoprecipitated HDAC5.
The ability of PKD to interact with HDAC5 in cardiac myocytes was further addressed by sequential immunoprecipitation and immunoblotting. NRVMs were infected with a GFP-HDAC5-encoding adenovirus and treated with PE in the absence or presence of Bis I. As shown in Fig. 8B, endogenous PKD efficiently coimmunoprecipitated with HDAC5. PKD was associated with HDAC5 in the absence of agonist and became activated in a PKC-dependent manner following PE treatment. The results further suggest that PKD is a cardiac class II HDAC kinase.
DISCUSSION
We have recently shown that class II HDACs act as signal-responsive repressors of cardiac hypertrophy (49). The antihypertrophic activity of class II HDACs is neutralized through phosphorylation of specific serine residues in the N-terminal portions of the proteins. Phosphorylation of these serines leads to binding of 14-3-3 proteins and subsequent export from the nucleus (see Fig. 9). HDACs with these serine residues converted to alanines are unresponsive to kinase-mediated inactivating signals. Such signal-unresponsive HDAC mutants block the activation of the fetal gene program characteristic of hypertrophy in cardiac myocytes, while genetic inactivation of HDAC9 or HDAC5 in mice leads to an increased sensitivity to hypertrophic stimuli (49; S. Chang, T. A. McKinsey and E. N. Olson, submitted for publication). Taken together, these observations suggest that the nuclear export and inactivation of class II HDACs is a key regulatory step in the development of cardiac hypertrophy.
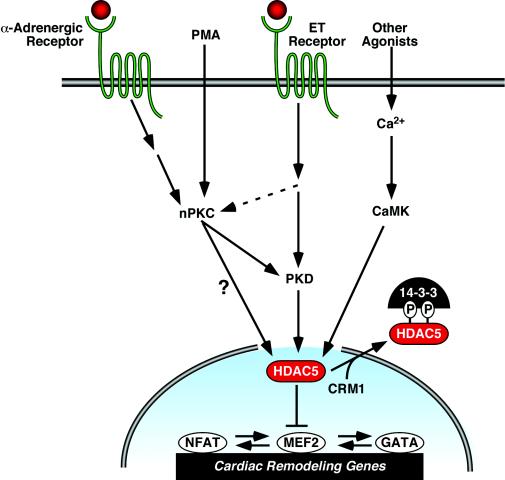
FIG. 9.
Model of kinase-dependent signaling pathways that regulate the nuclear export of class II HDACs and cardiac hypertrophy. HDAC5 represses pathological cardiac gene expression and remodeling via interactions with MEF2. MEF2 associates with other prohypertrophic transcription factors, including NFAT and GATA4, and thus HDAC5 also indirectly represses genes under the control of these factors. The repressive effects of HDAC5 are neutralized by signals that culminate in phosphorylation of the protein. Phospho-HDAC5 binds 14-3-3 proteins, resulting in nuclear export of HDAC5 through a CRM1-dependent mechanism. Phosphorylation of HDAC5 can be triggered by CaMK and, as shown in the present study, by signaling via calcium-independent PKCs, also referred to as novel (nPKCs). Hypertrophic signaling cascades, including those elicited by the α-adrenergic and ET receptors, stimulate the nuclear export of HDAC5 via activation of nPKCs and their downstream effector PKD. However, ET receptor signaling also appears to activate PKD through a PKC-independent mechanism. PMA directly activates PKC. It remains possible that PKCs bypass PKD and directly phosphorylate HDAC5.
The regulatory sites in class II HDACs are efficiently phosphorylated by CaMK, which has been implicated in hypertrophic growth of the myocardium in vivo and in vitro (32, 38, 50). However, an important issue that has not been previously resolved is whether CaMK is the sole kinase responsible for regulating HDAC nuclear export and hypertrophy or whether multiple kinases might converge on the regulatory HDAC phosphorylation sites, such that different HDAC kinases might be activated in response to different stimuli.
Convergence of multiple protein kinases on class II HDACs.
The results of this study show that PKC signaling leads to the phosphorylation of the same sites in HDAC5 that were previously shown to be phosphorylated by CaMK (29). Thus, the PKC and CaMK pathways converge at the same regulatory sites on HDAC5 (and presumably other class II HDACs). Both CaMK and PKC have been implicated in cardiac hypertrophy. Cardiac-specific expression of CaMK induces hypertrophy, at least in part through direct phosphorylation of class II HDACs, leading to their dissociation from and subsequent activation of MEF2 (25, 38). However, there is also evidence suggesting that kinases other than CaMK can serve as HDAC kinases. We have observed, for example, an HDAC kinase activity from cardiac extracts that is enhanced by hypertrophic stimuli but is not inhibited by inhibitors of CaMK or a variety of other kinases (49). In addition, agonist-dependent nuclear export of HDAC5 in cardiac myocytes is resistant to the CaMK inhibitors KN93 (Fig. 3C) and KN62 (data not shown).
While CaMK and some isoforms of PKC are calcium dependent, they each display distinct cofactor requirements and calcium requirements, as well as different subcellular distributions (17, 43). Thus, some hypertrophic stimuli may selectively activate one or the other of these kinases while others could activate both. In addition, it is possible that CaMK cross talks with the PKC-PKD pathway.
PKC and cardiac hypertrophy.
The PKC family includes at least 12 different isoforms, many but not all of which are expressed at appreciable levels in the myocardium. PKC family members can be subdivided based on their activation profiles. Conventional PKCs including PKCα, PKCβI, PKCβII, and PKCγ require calcium in addition to phospholipids for activation, while the atypical PKCs including PKCδ, PKCɛ, PKCη, and PKCθ are calcium independent.
Numerous studies have implicated PKC signaling in cardiac hypertrophy. PMA activates the fetal gene program and increases cardiomyocyte size (1, 10). PKCα, the most predominant PKC isoform in the heart, is sufficient to promote cardiac hypertrophy when overexpressed in neonatal rat cardiac myocytes (4). PKCα activation has also been shown to precede hypertrophy in a mouse model of L-type voltage-dependent calcium channel overexpression (36). Recently, it was shown that cardiac function was maintained in PKCα-null mice following aortic banding, a surgical model of pressure overload induced heart failure (5). Cardiac-specific overexpression of PKCβ in mice also leads to cardiac hypertrophy and sudden death (3), although genetic ablation of PKCβ does not abolish hypertrophy in response to pressure overload or PE infusion (41). In addition, a peptide modulator of PKCɛ which leads to membrane translocation and activation of PKCɛ elicits cardiac hypertrophy in vivo while maintaining normal heart function (34).
Our results demonstrate that direct activation of PKC by PMA is sufficient to induce the nuclear export of HDAC5 and that hypertrophic agonists such as PE stimulate the nuclear export of HDAC5 in cardiac myocytes through a signaling pathway that depends on PKC activation. Signal-resistant forms of HDAC5 are also capable of blocking hypertrophy caused by PE and PMA. These findings demonstrate that HDAC5 is a target for PKC signaling in cardiomyocytes and demonstrate that the nuclear export of HDAC5 or other class II HDACs by PKC is a critical step in the signaling pathway leading to cardiac hypertrophy.
PKD regulation of class II HDACs during cardiac hypertrophy.
We also show here that PKD can serve as an HDAC kinase. The amino acid sequence surrounding the signal-responsive serines of HDAC5 conforms to the PKD consensus phosphorylation site (29, 37). PKD is activated through direct phosphorylation by PKC on two serine residues in its activation loop (18). PKD activation can be inhibited by Bis I or Gö6983, which block PKC activity (18, 24). Calcium-independent PKCs have been most strongly implicated in the control of PKD (39). Thus, our demonstration that calcium-independent, but not calcium-dependent, PKCs trigger the nuclear export of HDAC5 (Fig. 2) further suggests a role for PKD in the regulation of HDAC5. According to the model (Fig. 9), PKD-directed phosphorylation of HDAC5 occurs in the nucleus. While we have not established this to be the case in the present study, it should be noted that PKD has previously been shown to translocate to the nucleus in response to G-protein-coupled receptor signaling (40).
Interestingly, HDAC5 nuclear export mediated by ET-1 or FBS is not blocked by Bis I or Gö6983 but is suppressed by Gö6976, a direct inhibitor of PKD. This suggests that there may be PKC-independent pathways stimulated by ET-1 and FBS that are responsible for PKD activation and HDAC5 export (Fig. 9). In this regard, PKD is also activated by direct binding of the βγ subunits of heterotrimeric G proteins to the pleckstrin homology domain (19). In addition, tyrosine phosphorylation of the pleckstrin homology domain by the Abl kinase precedes the phosphorylation of PKD by PKCδ in response to oxidative stress (44). PKC-independent activation of PKD was recently demonstrated in osteoblasts, although the alternative mechanism was not revealed (22).
Little is known about the role of PKD in cardiac hypertrophy. PKD is activated by PMA and PE in a Bis I-sensitive manner in NRVMs (16). Aldosterone also activates PKD in cardiomyocytes in culture and in mice (46). Furthermore, Gö6976 inhibits fetal gene expression in cardiomyocytes activated by aldosterone. In none of these cases were the intracellular targets of PKD responsible for hypertrophy identified. Our results demonstrate that PKD can serve as an HDAC kinase and suggest that PKD regulates cardiac hypertrophy, at least in part through phosphorylation and subsequent nuclear export of class II HDACs. However, this does not rule out the possible involvement of other kinases, including CaMK, in the regulation of the HDAC subcellular distribution and hypertrophy. It is interesting that although PKD is also referred to as PKCμ, its catalytic domain is most homologous to that of CaMK and is only distantly related to the catalytic domains of PKCs (24).
Implications for other cell types.
Given the expression of class II HDACs in a variety of cell types and the importance of calcium signaling in modulating cellular phenotypes, it is intriguing to speculate that class II HDACs have evolved to use the same regulatory phosphorylation sites to respond to different physiological stimuli and regulate distinct sets of genes in different cell types. In this regard, HDAC7 was recently shown to undergo nuclear export in T lymphocytes in response to PMA (9). Similarly, HDAC4 and HDAC5 are exported from the nucleus in hippocampal neurons in response to spontaneous activity and HDAC5 is similarly controlled in cerebellar granule neurons subjected to depolarizing potassium (7, 23). Although CaMK inhibition disrupts HDAC5 nuclear export in these cells, it has little effect on HDAC4 export, suggesting that CaMK-independent mechanisms regulate class II HDAC localization in neurons.
PKC signaling influences a wide range of cellular processes, including proliferation, differentiation, and oncogenesis. The specific response to PKC activation is dependent on cell identity and on other intracellular signals. In a postnatal cardiomyocyte, PKC signaling can evoke hypertrophy, while in other cell types it may control other cellular responses. The results of this study reveal a specific mechanism whereby PKC and PKD signaling can be coupled to transcriptional machinery via class II HDACs. Because these HDACs interact with a variety of positive and negative transcriptional regulators, they provide the potential to convey “generic” upstream signals from the cell membrane to cell-type-specific target genes. Pharmacological modulation of the upstream kinases and phosphatases that ultimately govern the phosphorylation of class II HDACs offers interesting possibilities for modulation of cellular phenotypes during development and disease.
Acknowledgments
We thank Nikos Pagratis for assistance with the Cellomics Imaging System; Chun Li Zhang for advice; and Ryan Plichta, Lisa Hollingsworth, and Matt Drietz for cardiomyocyte preparation. We are grateful to Alisha Tizenor for assistance with graphics.
This work was supported by grants from the NIH and the Donald W. Reynolds Center for Clinical Cardiovascular Research to E.N.O. and from the American Heart Association to R.V.
REFERENCES
- 1.Allo, S. N., L. L. Carl, and H. E. Morgan. 1992. Acceleration of growth of cultured cardiomyocytes and translocation of protein kinase C. Am. J. Physiol. 263:C319-C325. [DOI] [PubMed] [Google Scholar]
- 2.Antos, C. L., T. A. McKinsey, M. Dreitz, L. M. Hollingsworth, C. L. Zhang, K. Schreiber, H. Rindt, R. J. Gorczynski, and E. N. Olson. 2003. Dose-dependent blockade to cardiomyocyte hypertrophy by histone deacetylase inhibitors. J. Biol. Chem. 278:28930-28937. [DOI] [PubMed] [Google Scholar]
- 3.Bowman, J. C., S. F. Steinberg, T. Jiang, D. L. Geenen, G. I. Fishman, and P. M. Buttrick. 1997. Expression of protein kinase C beta in the heart causes hypertrophy in adult mice and sudden death in neonates. J. Clin. Investig. 100:2189-2195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Braz, J. C., O. F. Bueno, L. J. De Windt, and J. D. Molkentin. 2002. PKC alpha regulates the hypertrophic growth of cardiomyocytes through extracellular signal-regulated kinase1/2 (ERK1/2). J. Cell Biol. 156:905-919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Braz, J. C., K. Gregory, A. Pathak, W. Zhao, B. Sahin, R. Klevitsky, T. F. Kimball, J. N. Lorenz, A. C. Nairn, S. B. Liggett, I. Bodi, S. Wang, A. Schwartz, E. G. Lakatta, A. A. DePaoli-Roach, J. Robbins, T. E. Hewett, J. A. Bibb, M. V. Westfall, E. G. Kranias, and J. D. Molkentin. 2004. PKC-alpha regulates cardiac contractility and propensity toward heart failure. Nat. Med. 10:248-254. [DOI] [PubMed] [Google Scholar]
- 6.Bush, E., J. Fielitz, L. Melvin, M. Martinez-Arnold, T. A. McKinsey, R. Plichta, and E. N. Olson. 2004. A small molecular activator of cardiac hypertrophy uncovered in a chemical screen for modifiers of the calcineurin signaling pathway. Proc. Natl. Acad. Sci. USA 101:2870-2875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chawla, S., P. Vanhoutte, F. J. Arnold, C. L. Huang, and H. Bading. 2003. Neuronal activity-dependent nucleocytoplasmic shuttling of HDAC4 and HDAC5. J. Neurochem. 85:151-159. [DOI] [PubMed] [Google Scholar]
- 8.Chien, K. R. 1999. Stress pathways and heart failure. Cell 98:555-558. [DOI] [PubMed] [Google Scholar]
- 9.Dequiedt, F., H. Kasler, W. Fischle, V. Kiermer, M. Weinstein, B. G. Herndier, and E. Verdin. 2003. HDAC7, a thymus-specific class II histone deacetylase, regulates Nur77 transcription and TCR-mediated apoptosis. Immunity 18:687-698. [DOI] [PubMed] [Google Scholar]
- 10.Dunnmon, P. M., K. Iwaki, S. A. Henderson, A. Sen, and K. R. Chien. 1990. Phorbol esters induce immediate-early genes and activate cardiac gene transcription in neonatal rat myocardial cells. J. Mol. Cell. Cardiol. 22:901-910. [DOI] [PubMed] [Google Scholar]
- 11.Frey, N., and E. N. Olson. 2003. Cardiac hypertrophy: the good, the bad, and the ugly. Annu. Rev. Physiol. 65:45-79. [DOI] [PubMed] [Google Scholar]
- 12.Grozinger, C. M., C. A. Hassig, and S. L. Schreiber. 1999. Three proteins define a class of human histone deacetylases related to yeast Hda1p. Proc. Natl. Acad. Sci. USA 96:4868-4873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Grozinger, C. M., and S. L. Schreiber. 2002. Deacetylase enzymes: biological functions and the use of small-molecule inhibitors. Chem. Biol. 9:3-16. [DOI] [PubMed] [Google Scholar]
- 14.Grozinger, C. M., and S. L. Schreiber. 2000. Regulation of histone deacetylase 4 and 5 and transcriptional activity by 14-3-3-dependent cellular localization. Proc. Natl. Acad. Sci. USA 97:7835-7840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gschwendt, M., S. Dieterich, J. Rennecke, W. Kittstein, H. J. Mueller, and F. J. Johannes. 1996. Inhibition of protein kinase C mu by various inhibitors. Differentiation from protein kinase C isoenzymes. FEBS Lett. 392:77-80. [DOI] [PubMed] [Google Scholar]
- 16.Haworth, R. S., M. W. Goss, E. Rozengurt, and M. Avkiran. 2000. Expression and activity of protein kinase D/protein kinase C mu in myocardium: evidence for alpha1-adrenergic receptor- and protein kinase C-mediated regulation. J. Mol. Cell. Cardiol. 32:1013-1023. [DOI] [PubMed] [Google Scholar]
- 17.Hook, S. S., and A. R. Means. 2001. Ca2+/CaM-dependent kinases: from activation to function. Annu. Rev. Pharmacol. Toxicol. 41:471-505. [DOI] [PubMed] [Google Scholar]
- 18.Iglesias, T., R. T. Waldron, and E. Rozengurt. 1998. Identification of in vivo phosphorylation sites required for protein kinase D activation. J. Biol. Chem. 273:27662-27667. [DOI] [PubMed] [Google Scholar]
- 19.Jamora, C., N. Yamanouye, J. Van Lint, J. Laudenslager, J. R. Vandenheede, D. J. Faulkner, and V. Malhotra. 1999. Gbetagamma-mediated regulation of Golgi organization is through the direct activation of protein kinase D. Cell 98:59-68. [DOI] [PubMed] [Google Scholar]
- 20.Jenuwein, T., and C. D. Allis. 2001. Translating the histone code. Science 293:1074-1080. [DOI] [PubMed] [Google Scholar]
- 21.Kao, H. Y., A. Verdel, C. C. Tsai, C. Simon, H. Juguilon, and S. Khochbin. 2001. Mechanism for nucleocytoplasmic shuttling of histone deacetylase 7. J. Biol. Chem. 276:47496-47507. [DOI] [PubMed] [Google Scholar]
- 22.Lemonnier, J., C. Ghayor, J. Guicheux, and J. Caverzasio. 2004. Protein kinase C-independent activation of protein kinase D is involved in BMP-2- induced activation of stress mitogen-activated protein kinases JNK and p38 and osteoblastic cell differentiation. J. Biol. Chem. 279:259-264. [DOI] [PubMed] [Google Scholar]
- 23.Linseman, D. A., C. M. Bartley, S. S. Le, T. A. Laessig, R. J. Bouchard, M. K. Meintzer, M. Li, and K. A. Heidenreich. 2003. Inactivation of the myocyte enhancer factor-2 repressor histone deacetylase-5 by endogenous Ca2+/calmodulin-dependent kinase II promotes depolarization-mediated cerebellar granule neuron survival. J. Biol. Chem. 278:41472-41481. [DOI] [PubMed] [Google Scholar]
- 24.Lint, J. V., A. Rykx, T. Vantus, and J. R. Vandenheede. 2002. Getting to know protein kinase D. Int. J. Biochem. Cell Biol. 34:577-581. [DOI] [PubMed] [Google Scholar]
- 25.Lu, J., T. A. McKinsey, R. L. Nicol, and E. N. Olson. 2000. Signal-dependent activation of the MEF2 transcription factor by dissociation from histone deacetylases. Proc. Natl. Acad. Sci. USA 97:4070-4075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lu, J., T. A. McKinsey, C. L. Zhang, and E. N. Olson. 2000. Regulation of skeletal myogenesis by association of the MEF2 transcription factor with class II histone deacetylases. Mol. Cell 6:233-244. [DOI] [PubMed] [Google Scholar]
- 27.MacLellan, W. R., and M. D. Schneider. 2000. Genetic dissection of cardiac growth control pathways. Annu. Rev. Physiol. 62:289-319. [DOI] [PubMed] [Google Scholar]
- 28.McKinsey, T. A., and E. N. Olson. 2004. Cardiac histone acetylation—therapeutic opportunities abound. Trends Genet. 20:206-213. [DOI] [PubMed] [Google Scholar]
- 29.McKinsey, T. A., C. L. Zhang, J. Lu, and E. N. Olson. 2000. Signal-dependent nuclear export of a histone deacetylase regulates muscle differentiation. Nature 408:106-111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.McKinsey, T. A., C. L. Zhang, and E. N. Olson. 2000. Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14-3-3 to histone deacetylase 5. Proc. Natl. Acad. Sci. USA 97:14400-14405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.McKinsey, T. A., C. L. Zhang, and E. N. Olson. 2001. Identification of a signal-responsive nuclear export sequence in class II histone deacetylases. Mol. Cell. Biol. 21:6312-6321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.McKinsey, T. A., C. L. Zhang, and E. N. Olson. 2002. MEF2: a calcium- dependent regulator of cell division, differentiation and death. Trends Biochem. Sci. 27:40-47. [DOI] [PubMed] [Google Scholar]
- 33.Miska, E. A., C. Karlsson, E. Langley, S. J. Nielsen, J. Pines, and T. Kouzarides. 1999. HDAC4 deacetylase associates with and represses the MEF2 transcription factor. EMBO J. 18:5099-5107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mochly-Rosen, D., G. Wu, H. Hahn, H. Osinska, T. Liron, J. N. Lorenz, A. Yatani, J. Robbins, and G. W. Dorn, Jr. 2000. Cardiotrophic effects of protein kinase C epsilon: analysis by in vivo modulation of PKCepsilon translocation. Circ. Res. 86:1173-1179. [DOI] [PubMed] [Google Scholar]
- 35.Molkentin, J. D., J. R. Lu, C. L. Antos, B. Markham, J. Richardson, J. Robbins, S. R. Grant, and E. N. Olson. 1998. A calcineurin-dependent transcriptional pathway for cardiac hypertrophy. Cell 93:215-228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Muth, J. N., I. Bodi, W. Lewis, G. Varadi, and A. Schwartz. 2001. A Ca2+-dependent transgenic model of cardiac hypertrophy: a role for protein kinase Calpha. Circulation 103:140-147. [DOI] [PubMed] [Google Scholar]
- 37.Nishikawa, K., A. Toker, F. J. Johannes, Z. Songyang, and L. C. Cantley. 1997. Determination of the specific substrate sequence motifs of protein kinase C isozymes. J. Biol. Chem. 272:952-960. [DOI] [PubMed] [Google Scholar]
- 38.Passier, R., H. Zeng, N. Frey, F. J. Naya, R. L. Nicol, T. A. McKinsey, P. Overbeek, J. A. Richardson, S. R. Grant, and E. N. Olson. 2000. CaM kinase signaling induces cardiac hypertrophy and activates the MEF2 transcription factor in vivo. J. Clin. Investig. 105:1395-1406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rey, O., J. R. Reeve, E. Zhukova, J. Sinnett-Smith, and E. Rozengurt. 2004. G protein-coupled receptor-mediated phosphorylation of the activation loop of protein kinase D: dependence on plasma membrane translocation and protein kinase c epsilon. J. Biol. Chem. 279:34361-34372. [DOI] [PubMed] [Google Scholar]
- 40.Rey, O., E. Zhukova, J. Sinnett-Smith, and E. Rozengurt. 2003. Vasopressin- induced intracellular redistribution of protein kinase D in intestinal epithelial cells. J. Cell. Physiol. 196:483-492. [DOI] [PubMed] [Google Scholar]
- 41.Roman, B. B., D. L. Geenen, M. Leitges, and P. M. Buttrick. 2001. PKC-beta is not necessary for cardiac hypertrophy. Am. J. Physiol. Ser. H 280:H2264-H2270. [DOI] [PubMed] [Google Scholar]
- 42.Sadowski, I., B. Bell, P. Broad, and M. Hollis. 1992. GAL4 fusion vectors for expression in yeast or mammalian cells. Gene 118:137-141. [DOI] [PubMed] [Google Scholar]
- 43.Shirai, Y., and N. Saito. 2002. Activation mechanisms of protein kinase C: maturation, catalytic activation, and targeting. J. Biochem. (Tokyo). 132:663-668. [DOI] [PubMed] [Google Scholar]
- 44.Storz, P., H. Doppler, and A. Toker. 2004. Protein kinase Cδ selectively regulates protein kinase D-dependent activation of NF-κB in oxidative stress signaling. Mol. Cell. Biol. 24:2614-2626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Storz, P., and A. Toker. 2003. Protein kinase D mediates a stress-induced NF- kappaB activation and survival pathway. EMBO J. 22:109-120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tsybouleva, N., L. Zhang, S. Chen, R. Patel, S. Lutucuta, S. Nemoto, G. DeFreitas, M. Entman, B. A. Carabello, R. Roberts, and A. J. Marian. 2004. Aldosterone, through novel signaling proteins, is a fundamental molecular bridge between the genetic defect and the cardiac phenotype of hypertrophic cardiomyopathy. Circulation 109:1284-1291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wang, L., M. Rolfe, and C. G. Proud. 2003. Ca2+-independent protein kinase C activity is required for alpha1-adrenergic-receptor-mediated regulation of ribosomal protein S6 kinases in adult cardiomyocytes. Biochem. J. 373:603-611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang, A. H., and X. J. Yang. 2001. Histone deacetylase 4 possesses intrinsic nuclear import and export signals. Mol. Cell. Biol. 21:5992-6005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang, C. L., T. A. McKinsey, S. Chang, C. L. Antos, J. A. Hill, and E. N. Olson. 2002. Class II histone deacetylases act as signal-responsive repressors of cardiac hypertrophy. Cell 110:479-488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zhang, L., S. Maier, N. D. Dalton, S. Miyamoto, J. Ross, Jr., D. M. Bers, and J. H. Brown. 2003. The deltaC isoform of CaMKII is activated in cardiac hypertrophy and induces dilated cardiomyopathy and heart failure. Circ. Res. 92:912-919. [DOI] [PubMed] [Google Scholar]
- 51.Zugaza, J. L., J. Sinnett-Smith, J. Van Lint, and E. Rozengurt. 1996. Protein kinase D (PKD) activation in intact cells through a protein kinase C-dependent signal transduction pathway. EMBO J. 15:6220-6230. [PMC free article] [PubMed] [Google Scholar]