Abstract
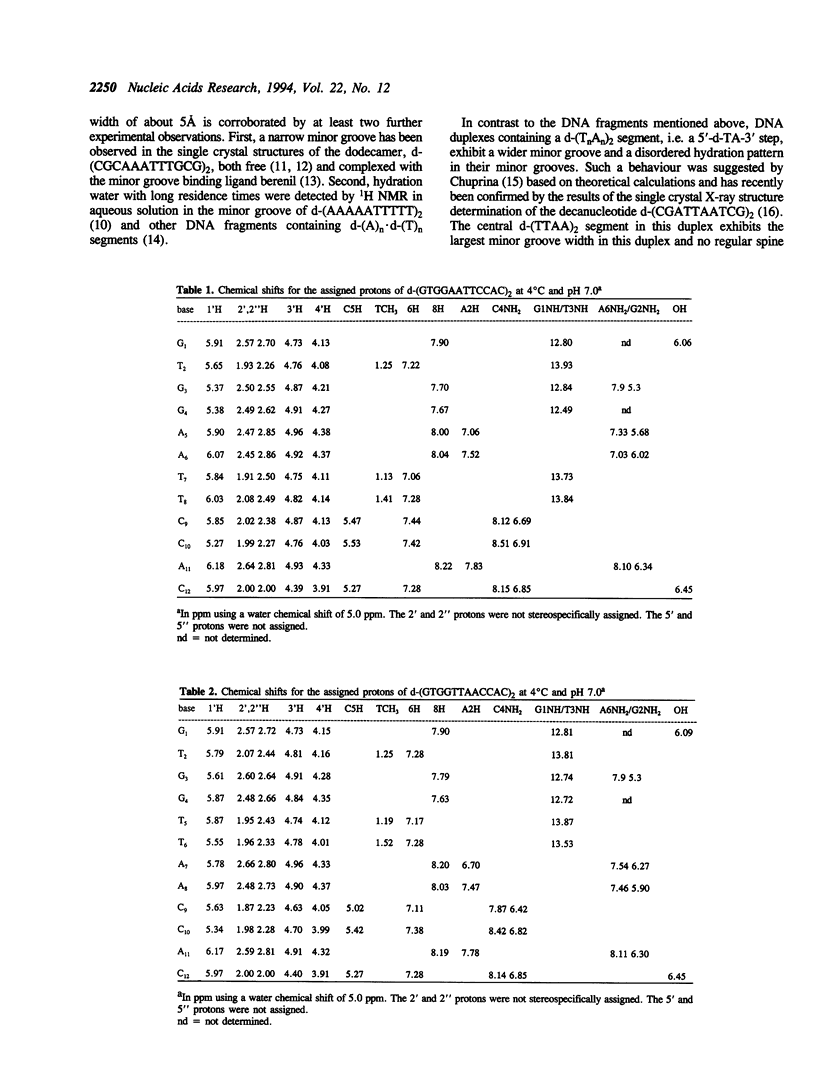
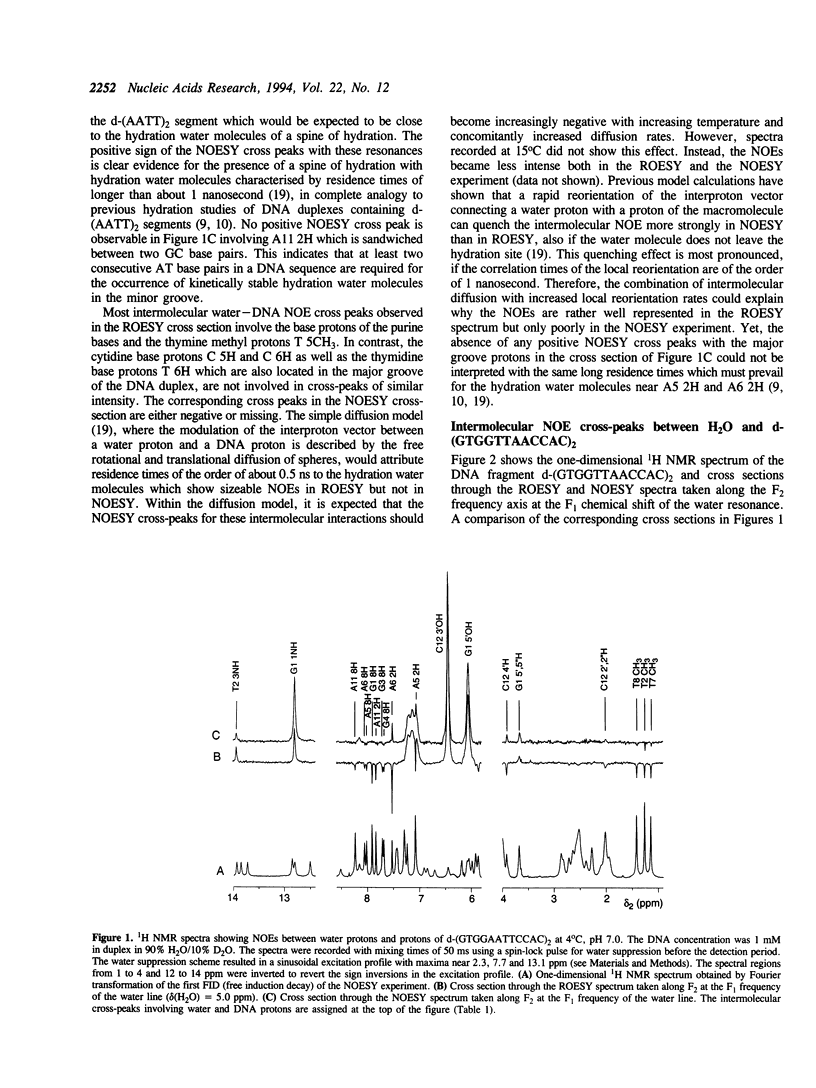
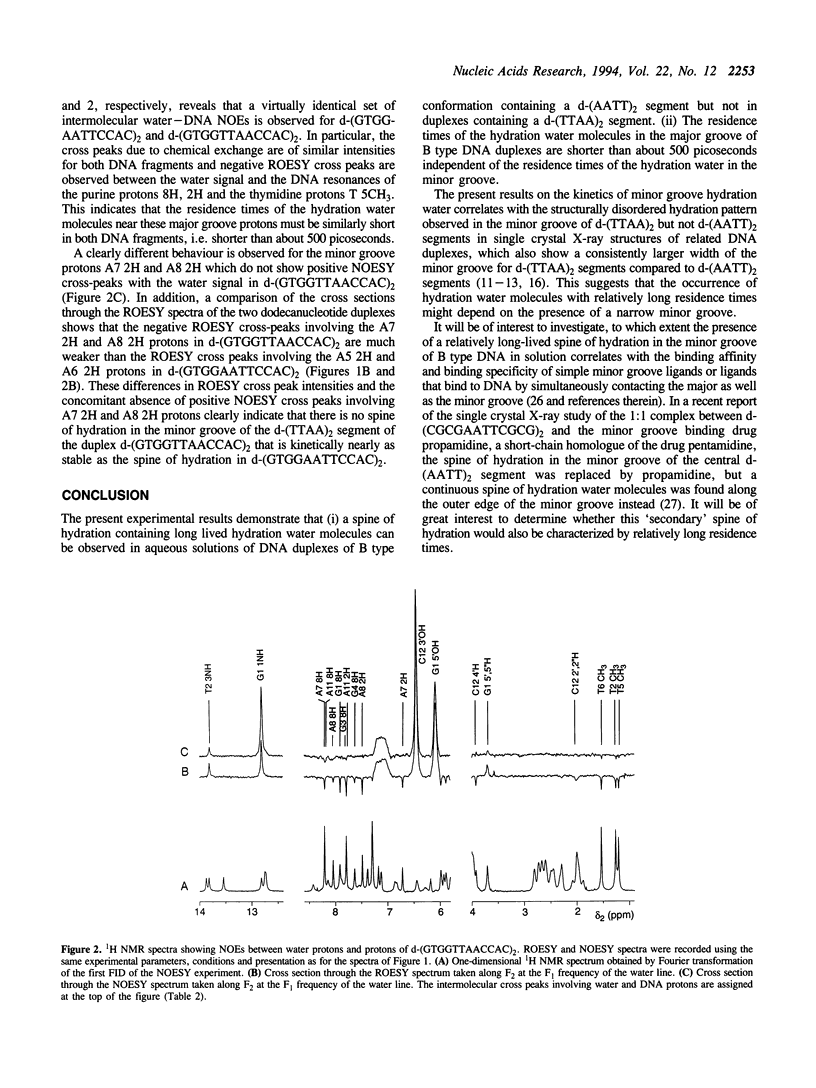
The residence times of the hydration water molecules near the base protons of d-(GTGGAATTCCAC)2 and d-(GTGGTTAACCAC)2 were investigated by nuclear magnetic resonance (NMR) spectroscopy. Nuclear Overhauser effects (NOE) were observed between base protons of the DNA and hydration water in NOESY and ROESY experiments. Large positive NOESY cross peaks observed between the resonances of the water and the adenine 2H protons of the central d-(AATT)2 segment in the duplex d-(GTGGAATTCCAC)2 indicate the presence of a 'spine of hydration' with water molecules exhibiting residence times on the DNA longer than 1 nanosecond. In contrast, no positive intermolecular NOESY cross peaks were detected in the d-(TTAA)2 segment of the duplex d-(GTGGTTAACCAC)2, indicating that no water molecules bound with similarly long residence times occur in the minor groove of this fragment. These results can be correlated with the larger width of the minor groove in d-(TTAA)2 segments as compared to that in d-(AATT)2 segments, as observed previously in single crystal structures of related oligonucleotide duplexes in B type conformation. The present experiments confirm earlier experimental results from single crystal studies and theoretical predictions that a 5'-dTA-3' step in the nucleotide sequence interrupts the spine of hydration in the minor groove.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brown D. G., Sanderson M. R., Garman E., Neidle S. Crystal structure of a berenil-d(CGCAAATTTGCG) complex. An example of drug-DNA recognition based on sequence-dependent structural features. J Mol Biol. 1992 Jul 20;226(2):481–490. doi: 10.1016/0022-2836(92)90962-j. [DOI] [PubMed] [Google Scholar]
- Chuprina V. P. Anomalous structure and properties of poly (dA).poly(dT). Computer simulation of the polynucleotide structure with the spine of hydration in the minor groove. Nucleic Acids Res. 1987 Jan 12;15(1):293–311. doi: 10.1093/nar/15.1.293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chuprina V. P., Lipanov A. A., Fedoroff OYu, Kim S. G., Kintanar A., Reid B. R. Sequence effects on local DNA topology. Proc Natl Acad Sci U S A. 1991 Oct 15;88(20):9087–9091. doi: 10.1073/pnas.88.20.9087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chuprina V. P., Sletten E., Fedoroff OYu Investigation of solution structure of d(GAATTTAAATTC)2 by 1H NMR, molecular dynamics, mechanics, refinement by back-calculation of the NOESY spectrum and analysis of this structure using X-ray data. J Biomol Struct Dyn. 1993 Feb;10(4):693–707. doi: 10.1080/07391102.1993.10508001. [DOI] [PubMed] [Google Scholar]
- Coll M., Frederick C. A., Wang A. H., Rich A. A bifurcated hydrogen-bonded conformation in the d(A.T) base pairs of the DNA dodecamer d(CGCAAATTTGCG) and its complex with distamycin. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8385–8389. doi: 10.1073/pnas.84.23.8385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickerson R. E., Drew H. R. Structure of a B-DNA dodecamer. II. Influence of base sequence on helix structure. J Mol Biol. 1981 Jul 15;149(4):761–786. doi: 10.1016/0022-2836(81)90357-0. [DOI] [PubMed] [Google Scholar]
- Drew H. R., Dickerson R. E. Structure of a B-DNA dodecamer. III. Geometry of hydration. J Mol Biol. 1981 Sep 25;151(3):535–556. doi: 10.1016/0022-2836(81)90009-7. [DOI] [PubMed] [Google Scholar]
- Edwards K. J., Brown D. G., Spink N., Skelly J. V., Neidle S. Molecular structure of the B-DNA dodecamer d(CGCAAATTTGCG)2. An examination of propeller twist and minor-groove water structure at 2.2 A resolution. J Mol Biol. 1992 Aug 20;226(4):1161–1173. doi: 10.1016/0022-2836(92)91059-x. [DOI] [PubMed] [Google Scholar]
- Feng J. A., Johnson R. C., Dickerson R. E. Hin recombinase bound to DNA: the origin of specificity in major and minor groove interactions. Science. 1994 Jan 21;263(5145):348–355. doi: 10.1126/science.8278807. [DOI] [PubMed] [Google Scholar]
- Fratini A. V., Kopka M. L., Drew H. R., Dickerson R. E. Reversible bending and helix geometry in a B-DNA dodecamer: CGCGAATTBrCGCG. J Biol Chem. 1982 Dec 25;257(24):14686–14707. [PubMed] [Google Scholar]
- Fritzsche H., Kan L. S., Weller K., Scheiding W., Kast J. R., Ts'o P. O. Phosphate catalyzed exchange rates of NH-N protons of DNA. J Biomol Struct Dyn. 1988 Oct;6(2):383–390. doi: 10.1080/07391102.1988.10507720. [DOI] [PubMed] [Google Scholar]
- Guéron M., Kochoyan M., Leroy J. L. A single mode of DNA base-pair opening drives imino proton exchange. Nature. 1987 Jul 2;328(6125):89–92. doi: 10.1038/328089a0. [DOI] [PubMed] [Google Scholar]
- Leroy J. L., Broseta D., Guéron M. Proton exchange and base-pair kinetics of poly(rA).poly(rU) and poly(rI).poly(rC). J Mol Biol. 1985 Jul 5;184(1):165–178. doi: 10.1016/0022-2836(85)90050-6. [DOI] [PubMed] [Google Scholar]
- Liepinsh E., Otting G., Wüthrich K. NMR observation of individual molecules of hydration water bound to DNA duplexes: direct evidence for a spine of hydration water present in aqueous solution. Nucleic Acids Res. 1992 Dec 25;20(24):6549–6553. doi: 10.1093/nar/20.24.6549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maltseva T. V., Agback P., Chattopadhyaya J. How much hydration is necessary for the stabilisation of DNA-duplex? Nucleic Acids Res. 1993 Sep 11;21(18):4246–4252. doi: 10.1093/nar/21.18.4246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn C. M., Jenkins T. C., Neidle S. Crystal structure of d(CGCGAATTCGCG) complexed with propamidine, a short-chain homologue of the drug pentamidine. Biochemistry. 1993 Dec 21;32(50):13838–13843. doi: 10.1021/bi00213a012. [DOI] [PubMed] [Google Scholar]
- Otting G., Liepinsh E., Farmer B. T., 2nd, Wüthrich K. Protein hydration studied with homonuclear 3D 1H NMR experiments. J Biomol NMR. 1991 Jul;1(2):209–215. doi: 10.1007/BF01877232. [DOI] [PubMed] [Google Scholar]
- Otting G., Liepinsh E., Wüthrich K. Protein hydration in aqueous solution. Science. 1991 Nov 15;254(5034):974–980. doi: 10.1126/science.1948083. [DOI] [PubMed] [Google Scholar]
- Otwinowski Z., Schevitz R. W., Zhang R. G., Lawson C. L., Joachimiak A., Marmorstein R. Q., Luisi B. F., Sigler P. B. Crystal structure of trp repressor/operator complex at atomic resolution. Nature. 1988 Sep 22;335(6188):321–329. doi: 10.1038/335321a0. [DOI] [PubMed] [Google Scholar]
- Quintana J. R., Grzeskowiak K., Yanagi K., Dickerson R. E. Structure of a B-DNA decamer with a central T-A step: C-G-A-T-T-A-A-T-C-G. J Mol Biol. 1992 May 20;225(2):379–395. doi: 10.1016/0022-2836(92)90928-d. [DOI] [PubMed] [Google Scholar]
- Shakked Z., Guzikevich-Guerstein G., Frolow F., Rabinovich D., Joachimiak A., Sigler P. B. Determinants of repressor/operator recognition from the structure of the trp operator binding site. Nature. 1994 Mar 31;368(6470):469–473. doi: 10.1038/368469a0. [DOI] [PubMed] [Google Scholar]
- Westhof E. Re-refinement of the B-dodecamer d(CGCGAATTCGCG) with a comparative analysis of the solvent in it and in the Z-hexamer d(5BrCG5BrCG5BrCG). J Biomol Struct Dyn. 1987 Dec;5(3):581–600. doi: 10.1080/07391102.1987.10506414. [DOI] [PubMed] [Google Scholar]
- Wing R., Drew H., Takano T., Broka C., Tanaka S., Itakura K., Dickerson R. E. Crystal structure analysis of a complete turn of B-DNA. Nature. 1980 Oct 23;287(5784):755–758. doi: 10.1038/287755a0. [DOI] [PubMed] [Google Scholar]


