Abstract
The brain’s ability to associate different stimuli is vital to long-term memory, but how neural ensembles encode associative memories is unknown. Here we studied how cell ensembles in the basal and lateral amygdala (BLA) encode associations between conditioned and unconditioned stimuli (CS, US). Using a miniature fluorescence microscope, we tracked BLA ensemble neural Ca2+ dynamics during fear learning and extinction over six days in behaving mice. Fear conditioning induced both up- and down-regulation of individual cells’ CS-evoked responses. This bi-directional plasticity mainly occurred after conditioning and reshaped the CS ensemble neural representation to gain similarity to the US-representation. During extinction training with repetitive CS presentations, the CS-representation became more distinctive without reverting to its original form. Throughout, the strength of the ensemble-encoded CS-US association predicted each mouse’s level of behavioral conditioning. These findings support a supervised learning model in which activation of the US-representation guides the transformation of the CS-representation.
Associative fear conditioning induces a long-term memory that requires BLA1–3 but not hippocampal4 activity. Past work found BLA neurons with potentiated responses to a CS, such as an auditory tone, after associative conditioning with an aversive US1–3. This prompted a Hebbian model in which ‘fear cells’ with co-active inputs conveying the paired CS–US presentations potentiate their responses to subsequent CS presentations1,3,5. However, the dynamics of individual fear cells seem too stochastic to support reliable memory storage1. Neural ensembles might allow more robust storage, but how cell ensembles encode associative memories and whether this fits the Hebbian model remain unknown.
To track BLA neural ensemble activity in behaving mice, we combined time-lapse microendoscopy, a head-mounted microscope6,7 and expression of the GCaMP6m8 Ca2+-indicator in excitatory neurons (Fig. 1a,b; Extended Data Fig. 1; Methods). This differs from past electrophysiological studies of BLA, which lacked access to ensemble activity patterns and had limited recording durations1, and studies of immediate early gene (IEG) activation9,10, which poorly reports declines, temporal patterns and gradations of electrical activity.
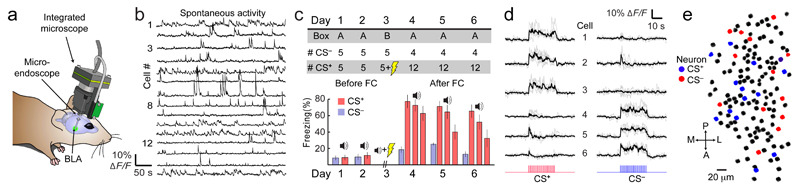
Fig. 1. Ca2+ imaging of BLA neural activity across a six-day fear conditioning protocol.
(a) A miniature microscope and implanted microendoscope allowed large-scale neural Ca2+ imaging.
(b) Traces of spontaneous Ca2+ activity from 15 BLA neurons.
(c) Upper, Conditioning protocol, with numbers of stimuli. Lower, Mean ± s.e.m. percentages of time 12 mice froze during CS+ and CS– presentations. Values are respectively averaged over 5 and 4 stimulus presentations, before and after conditioning.
(d) Activity traces of cells responsive to CS+ or CS– presentations before conditioning.
(e) Map of BLA cells in one mouse. Colored cells responded to CS+ or CS– tones.
Traces in b and d were down-sampled to 200 ms time bins.
We first examined neural responses to tones and electric shocks in awake mice (Extended Data Fig. 2). The cells that responded to these stimuli were sparse and interspersed across BLA10,11. This intermingling may help the BLA to link temporally associated signals of different types via local circuit interactions10–14.
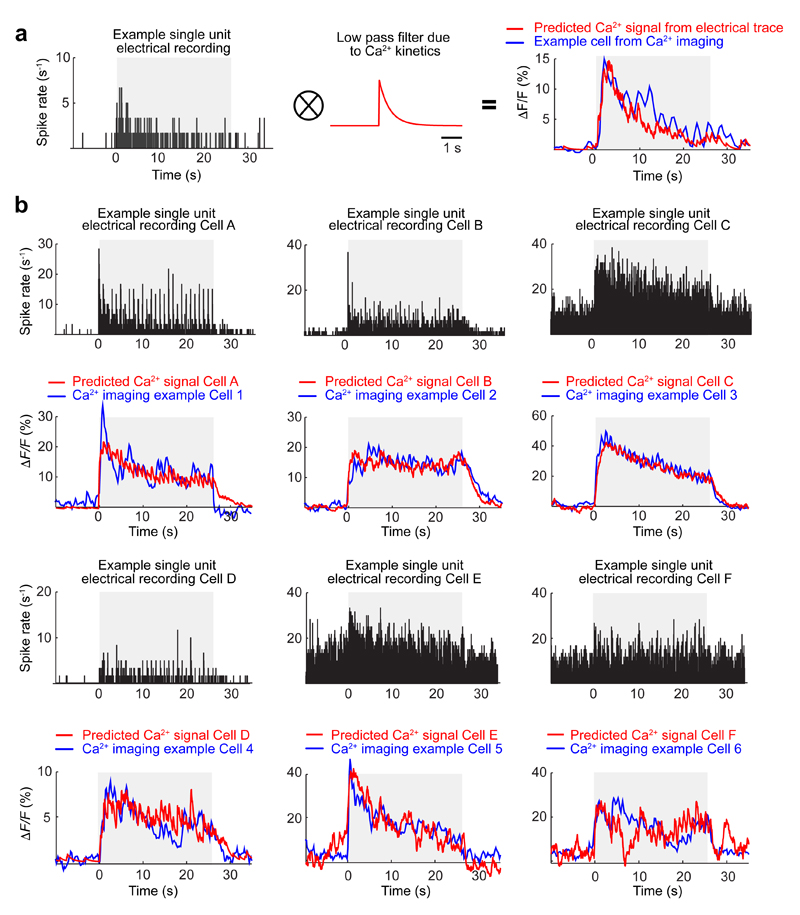
To study associative memory1,12,14–16, we repeatedly paired an auditory cue (CS+; 25 × 200-ms-tone-pulses per presentation) with a foot-shock US. As a control, we repeatedly presented another tone (CS–) without the US (Fig. 1c). Mice with and without implanted microendoscopes had comparable CS+-evoked fear expression, visible as conditioned freezing15,17 (Extended Data Fig. 3). Across a six-day protocol, cells responding to the CS+ or CS– (P ≤ 0.01; evoked signals vs. baseline; rank-sum test) were sparse, interspersed and largely distinct (Fig. 1c–e). CS-evoked Ca2+ transients closely resembled those expected from past electrical recordings12 (Extended Data Fig. 4).
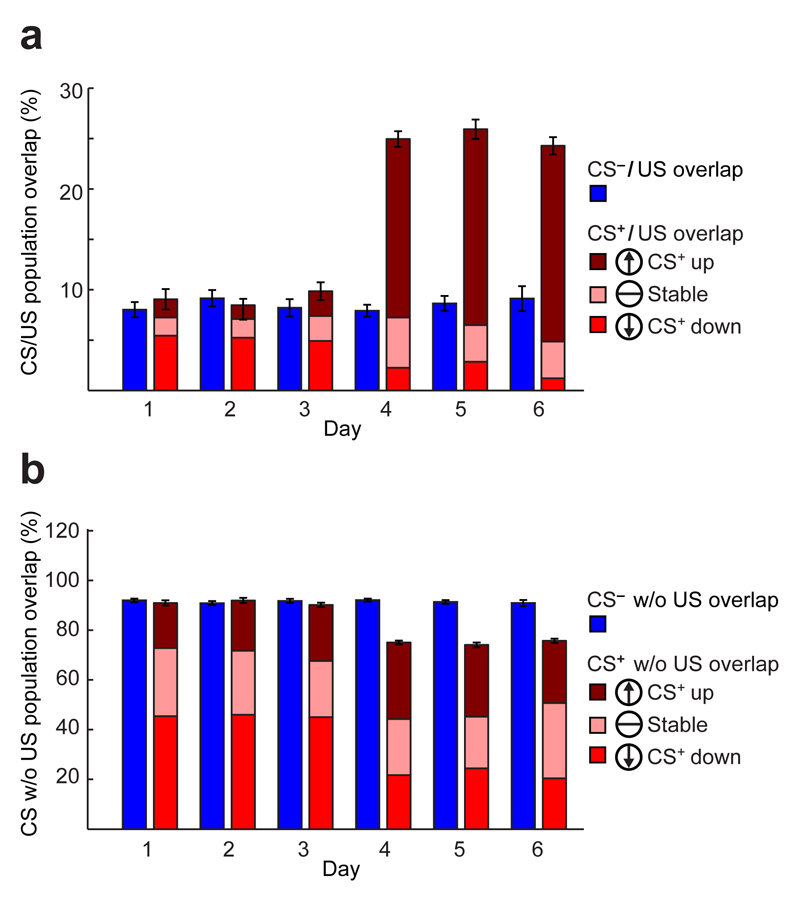
Across all six days, the number of active cells stayed constant [152 ± 14 (s.e.m.) cells per day per mouse; Friedman Test; 12 mice; see Supplementary Table 1 for P- and χ2-values], but after conditioning ~45% more responded1,10 to the CS+ (Fig. 2), [before training, 9 ± 1% (s.e.m) cells were CS+-responsive vs. 14 ± 1% afterward; P ≤ 0.01; rank-sum test; 2 pre- and 3 post-training sessions]. Percentages of CS–-responsive cells also rose, paralleling the small rise in CS–-evoked freezing above baseline levels and suggesting the CS– was not a learned safety signal18 (Figs. 1c, 2a; Extended Data Fig. 3g; Supplementary Note). During conditioning (Day 3), 14 ± 3% of active cells responded to the US; within this subset a minority up- (7 ± 3%) or down-regulated (13 ± 5%) these responses during training (Fig. 2c,d).
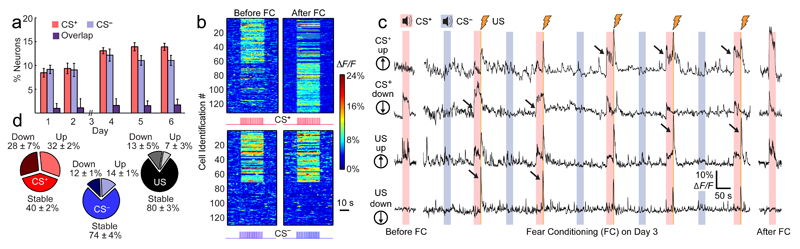
Fig. 2. Fear conditioning induces bi-directional changes in BLA signaling.
(a) Percentages (± s.e.m.) of cells responding to the CS+, CS–, or both stimuli.
(b) Ca2+ signals, showing changes in CS+ encoding and stable CS– encoding for two sets of 125 cells detected throughout the study. Top, Cells responsive to the CS+ on at least one day. Bottom, Cells that either responded to the CS– on one or more days, or lacked responses to both CS types. Colors show each cell’s Ca2+ response averaged over 5 CS presentations on the day the cell responded maximally, for days before and after fear conditioning (FC). Cells are arranged by whether they responded maximally before or after conditioning.
(c) Ca2+ signals from four cells, before (left, mean over 5 CS+ presentations), during (middle, single trial), and after (right, mean over 5 CS+ presentations) conditioning, illustrating altered responses to the CS+ (top two traces) or US (bottom two traces).
(d) Percentages (± s.e.m.) of cells after conditioning with stable, increased or decreased responses to the CS+ (red), CS– (blue) and US (black), respectively based on 231, 362 and 261 neurons. Cells in the former two charts responded to the CS on at least one day before or after conditioning. Cells in the latter chart responded significantly to the US on Day 3.
Traces in b and c were down-sampled to 200 ms time bins. a, b, d are from N=12 mice.
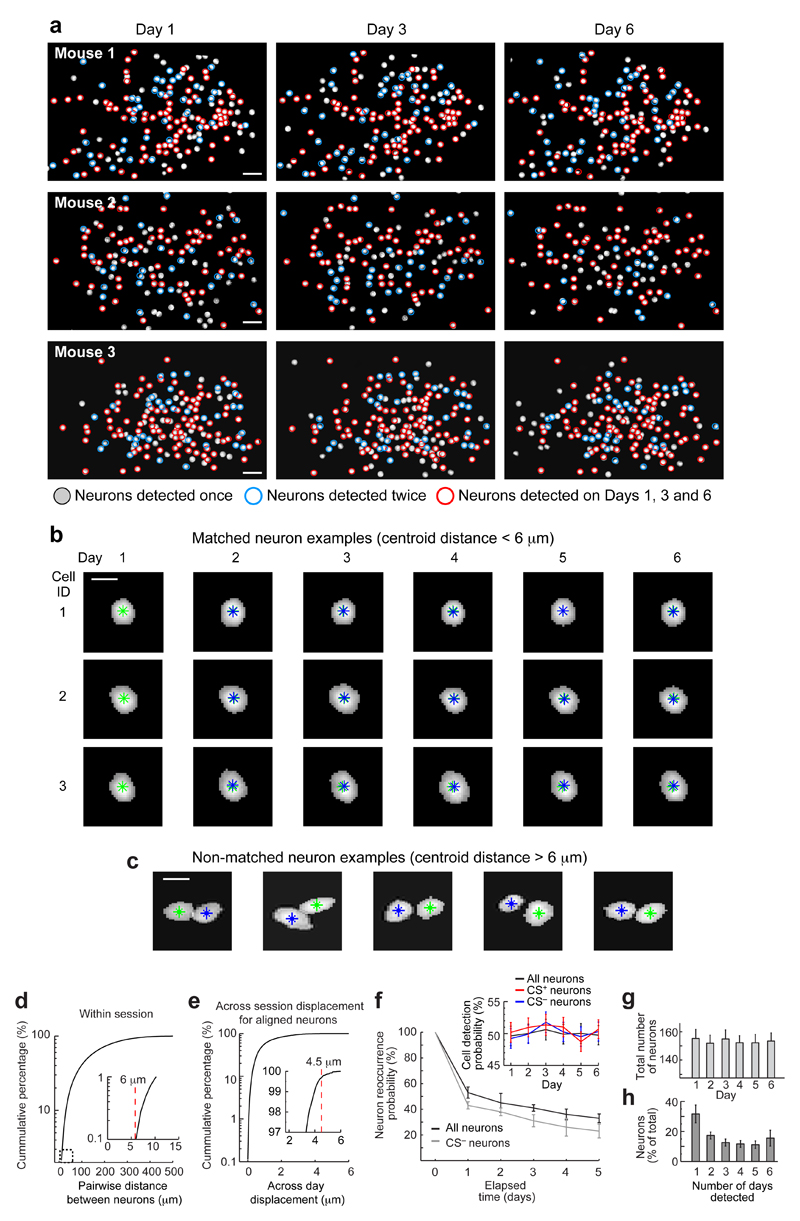
Using image alignment we registered cell identities over the six days (171–438 cells per mouse; 3655 total; 12 mice). Similar percentages of cells were active each day [49 ± 2% (s.e.m.); Extended Data Fig. 5; Supplementary Table 1]. A plurality of cells was active on 1–2 days (49 ± 3%) and a minority on all days (16 ± 2%). Individual cells came in and out of the active ensemble day-to-day; there were ~55% cells in common for consecutive sessions and ~35% for 5 d apart. This turnover resembles that seen in long-term studies of hippocampus7,19 and might be a general phenomenon in brain areas processing long-term memories.
We next studied the encoding of the CS+–US association and tested the Hebbian model20. Surprisingly, only 38 ± 5% of cells with heightened CS+-evoked responses after training responded to the US during training, whereas 65 ± 6% of cells that were initially responsive to both the CS+ and US were less CS+-responsive after training (Extended Data Fig. 6). Of cells with significant responses to the CS+ on at least one day, 32 ± 2% potentiated these responses after training, whereas 28 ± 7% reduced them (Fig. 2d; rank-sum test; P ≤ 0.05; 125 CS+ tone-pulses per day before training, 300 afterward). CS–-responsive cells underwent analogous changes, to a lesser extent (Fig. 2d). Overall, this bi-directional plasticity was unpredicted from Hebbian potentiation20.
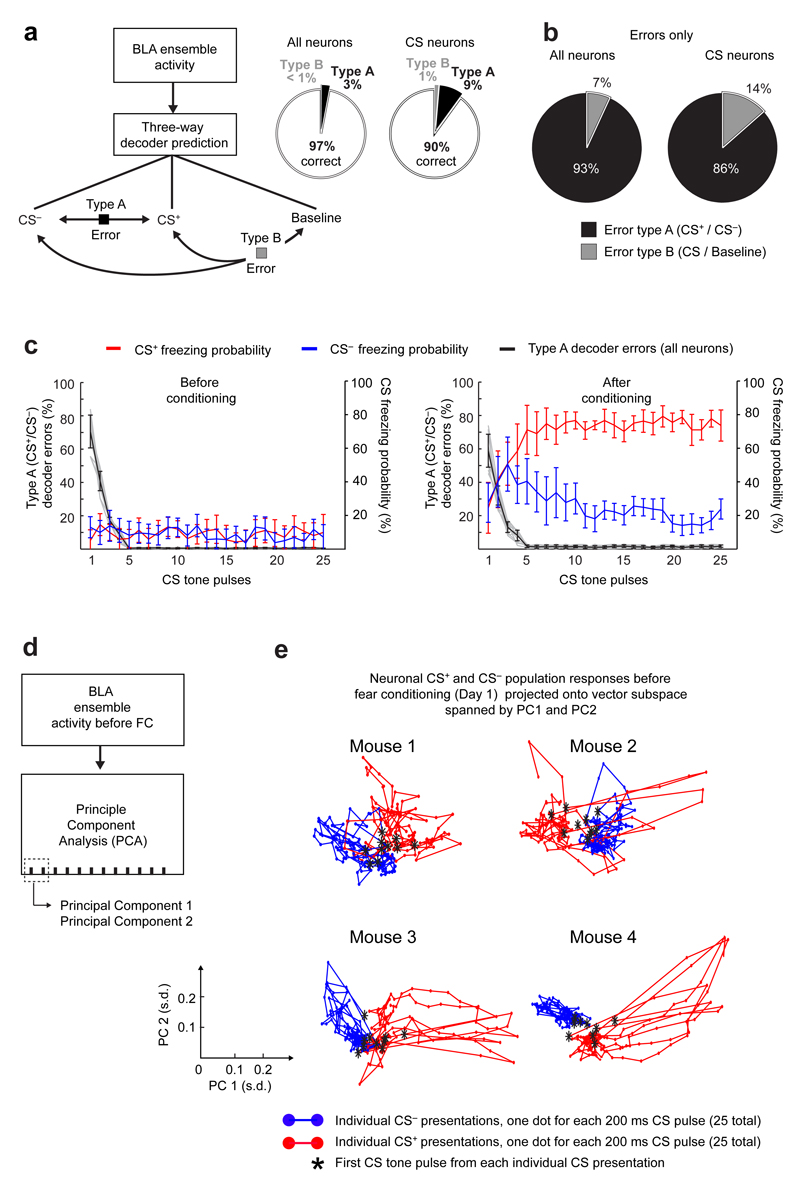
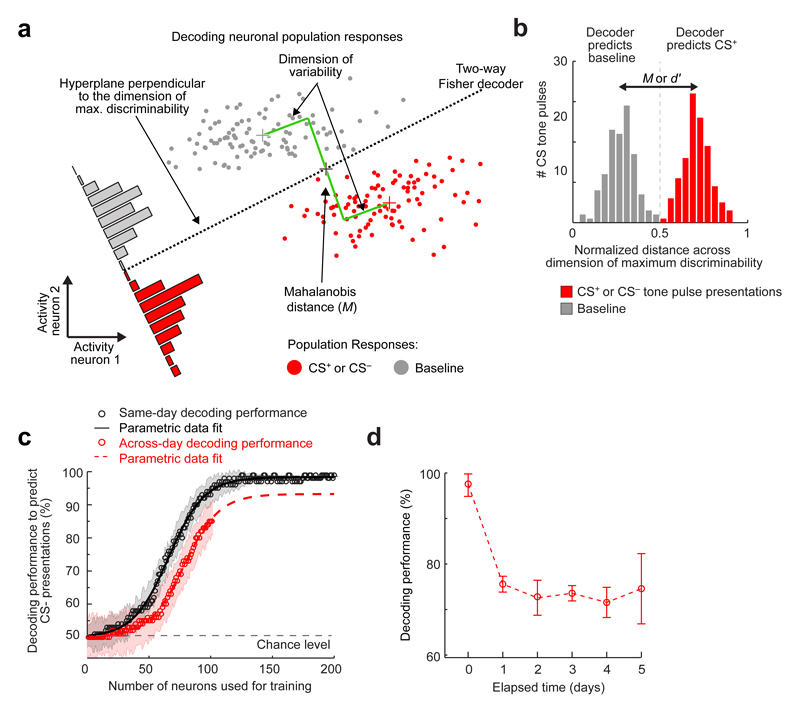
To study ensemble coding, we tested if CS+ and CS– presentations were identifiable from their evoked activity patterns. We trained three-way, Fisher linear decoders to distinguish baseline conditions from CS+ and CS– presentations on each day. These decoders classified the three conditions accurately [97 ± 0.3% (s.e.m.) of 1 s time-segments] for all 6 days (Fig. 3a). Accuracy fell modestly using only CS+- and CS–-responsive cells (90 ± 3% accuracy), but substantially when we omitted all CS-responsive cells (61 ± 2% accuracy). Across the first five tone-pulses of each CS presentation, decoding accuracy and conditioned freezing both rose to an asymptote (Extended Data Fig. 7), suggesting BLA coding fidelity improved as the tones repeated within a CS presentation.
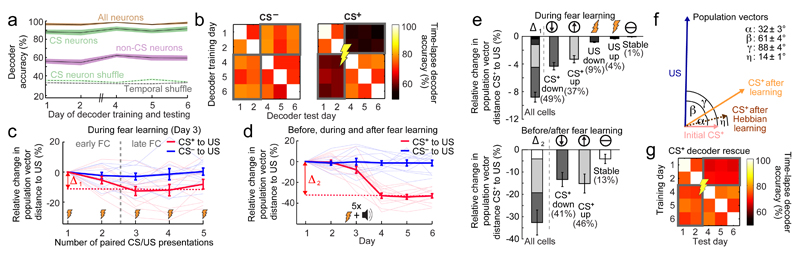
Fig. 3. Learning increases the similarity of the CS+ and US representations.
(a) Accuracies of same-day, three-way decoders discriminating baseline, CS+ and CS– presentations. Decoders based on CS-responsive cells (green curve) nearly matched those using all cells (brown). Decoders based on cells un-responsive to the CS (purple) performed poorly, but better than decoders given temporally shuffled Ca2+ traces (gray dotted line) or shuffled cell identities (green dotted line). 152 ± 14 (s.e.m.) cells per day per mouse (3655 cells total; 12 mice). Shading denotes s.e.m.
(b) Accuracies of inter-day, binary decoders distinguishing CS– (left) or CS+ (right) presentations from baseline conditions.
(c) Population vector distances (PVD) between US-evoked ensemble activity and that evoked by the CS– (blue) or CS+ (red) during conditioning (Day 3). CS+–US PVDs declined by an amount Δ1 as responses to the two stimuli gained similarity. Dashed vertical line separates early and late CS+–US pairings; to calculate Δ1 we compared these two portions of the session. 155 ± 11 (s.e.m.) cells per mouse on Day 3 (1860 cells total; 12 mice).
(d) CS+–US PVDs declined during and after training, indicating increased similarity of the two representations. Δ2 is the difference in PVD values before vs. after training (3655 cells).
(e) Upper, Composition of the changes, Δ1, in CS+–US PVDs between early and late phases of training, defined in c. Lower, Analogous graph for Δ2, showing how CS+–US PVDs changed from before (Days 1,2) to after (Days 4–6) conditioning. To decompose Δ1 and Δ2 we examined cells with stable (white), up- (light gray) or down-regulated (dark gray) responses to the CS+, and cells with up- or down-regulated responses to the US (black). Error bars: s.e.m.
(f) Before conditioning, population vector representations of the US (blue) and CS+ (pink) were orthogonal [88° ± 4° (s.e.m.); 12 mice]. Afterward, the CS+ population vector (orange) was 210 ± 20% longer, rotated 32° ± 3° from its initial orientation, and had a 61° ± 4° angle to the US representation, indicating the rotation was in the plane defined by the US-representation and that of the initial CS+. These changes differed from predictions of Hebbian potentiation (maroon) [angle and length changes each P < 10-4 ; rank-sum test].
(g) Mean accuracies of time-lapse decoders after computational rescue of their ability to distinguish CS+ presentations from baseline. For each pairing of one pre- and one post-training day (pairs inside gray rectangles), we rescued population vectors from the testing day by applying the optimal transformation, determined as in Extended Data Fig. 10c.
How did conditioning affect ensemble coding? To separately investigate CS+ and CS– encoding, we trained two sets of binary decoders, which discriminated either CS+ or CS– presentations from baseline conditions. We trained each decoder on data from one day and tested it on data from other days. Despite cells’ day-to-day fluctuations, CS– decoders had up to 85% accuracy across days [74 ± 1% (s.e.m.)] (Extended Data Fig. 8). CS+ decoders performed similarly, provided the training and testing days were both before or both after conditioning (74 ± 1% accuracy), but if they spanned the conditioning session, accuracy fell to chance levels (55 ± 1%) (Fig. 3b). Hence, representations of the CS+, but not the CS–, changed significantly during memory formation, consistent with the CS+-responsive cells’ bi-directional plasticity.
To further study plasticity, we constructed multi-dimensional population vectors (one dimension per cell) for each response to a CS or US. To assess the responses’ differentiability, we used the Mahalanobis population vector distance21 (PVD). This resembles an Euclidean distance, but like the discriminability index (d´) from statistics it accounts for mean differences and trial-to-trial variability21, using the correlations in the cells’ responses (Extended Data Figs. 8, 9). To examine how training changed the CS+-representation, we divided Day 3 into early and late training phases and computed the mean PVDs between US- and CS+-evoked responses in each phase (Fig. 3c). Notably, training increased the similarity and decreased the discriminability of the US and CS+ representations. Across five CS+–US pairings, PVDs declined by a significant amount [Δ1: –8 ± 2% (s.e.m.); P = 0.02; signed-rank test; 12 mice, early vs. late mean PVDs; 3655 cells], owing to increased similarity of the mean responses to CS+ and US, not decreases in their variability (Extended Data Fig. 9). CS–-representations remained invariant (Δ1 = –0.2 ± 0.3%; P = 0.3).
Even larger coding changes occurred after training. By Day 4, CS+- and US-representations were 32% less differentiable than before training (Fig. 3d,e), unforeseen from studies of consolidation that suggested a stabilization of neural coding22. 75% of the total change in CS+–US PVD (Δ2) first appeared on Day 4 (Fig. 3d) [CS+: Δ2 = –32 ± 6% relative to Day 1 PVDs; CS–: Δ2 = 0.5 ± 5%; 2 PVDs before training and 3 afterward, in each of 12 mice]. On Days 4–6 the CS+ population vector had increased 210 ± 20% (s.e.m.; 12 mice) in amplitude and rotated (32° ± 3°) nearly directly towards the US population vector (Fig. 3f). The re-scaling reflected increased CS+-evoked responses of many cells that never responded to the US, tempered by the decreased CS+-evoked responses of other cells. The rotation toward the US-representation reflected newfound CS+-evoked responses in cells previously lacking them. These changes differed from the predictions of Hebbian potentiation (changes in vector length and angle, each P < 10-4; rank-sum test; 12 mice).
Cells with decreased CS+-evoked responses and those with increased responses were equally important for the re-coding (Fig. 3e), during training [P = 0.2; signed-rank test, comparing contributions to Δ1 of cells with up- (37 ± 2%) vs. down-regulated (49 ± 2%) CS+ responses], and during consolidation [P= 0.9; contributions to Δ2 of cells with up- (46 ± 2%) vs. down-regulated (41 ± 2%) CS+ responses]. Changes in US encoding made modest (13 ± 1%) but significant (P = 0.008) contributions to the similarity increase between CS+ and US representations.
How did the CS+ encoding changes during consolidation relate to those from training? We hypothesized that consolidation proportionally accentuates changes from training. To test this, we linearly extrapolated the changes to the CS+-representation from conditioning (ΔA) and examined how well this captured the consolidated responses (Extended Data Fig. 10). Successful extrapolations should rescue the unsuccessful time-lapse CS+ decoders trained and tested on days spanning conditioning. With extrapolations 4–5× ΔA in amplitude, CS+-decoding reached 72 ± 3% accuracy, nearing that of time-lapse CS–-decoders (74 ± 1%) (Fig. 3g). If we limited extrapolation to cells with only up- or only down-regulated CS+-evoked responses, the rescue of CS+ decoding badly degraded, highlighting the importance of bi-directional plasticity during consolidation.
On Days 4–6, mice underwent partial behavioral extinction, comprising acute (within session) and consolidated (across session) effects (Figs. 1c, 4, 5). Did this reflect a change in the encoded CS+–US association? As found previously1, individual cells up- or down-regulated their CS+-evoked responses during acute extinction (Fig. 4a). We assessed how this affected CS–US PVDs across 4 CS– and 12 un-reinforced CS+ presentations. Between the first four and last four CS+ presentations, the CS+- and US-representations became significantly more differentiable (Δ3 = 20 ± 1%, normalized to the mean PVD on Day 1; signed-rank test; P < 10-3; 144 early vs. 144 late CS+ presentations on Days 4–6; Fig. 4b). This acute change reflected an 18 ± 5% (s.e.m.; 12 mice) reduction in CS+ population vector amplitude and a 8° ± 3° rotation away from the US vector. These changes were absent for the CS– (Δ3 = –3 ± 3%; P = 0.3).
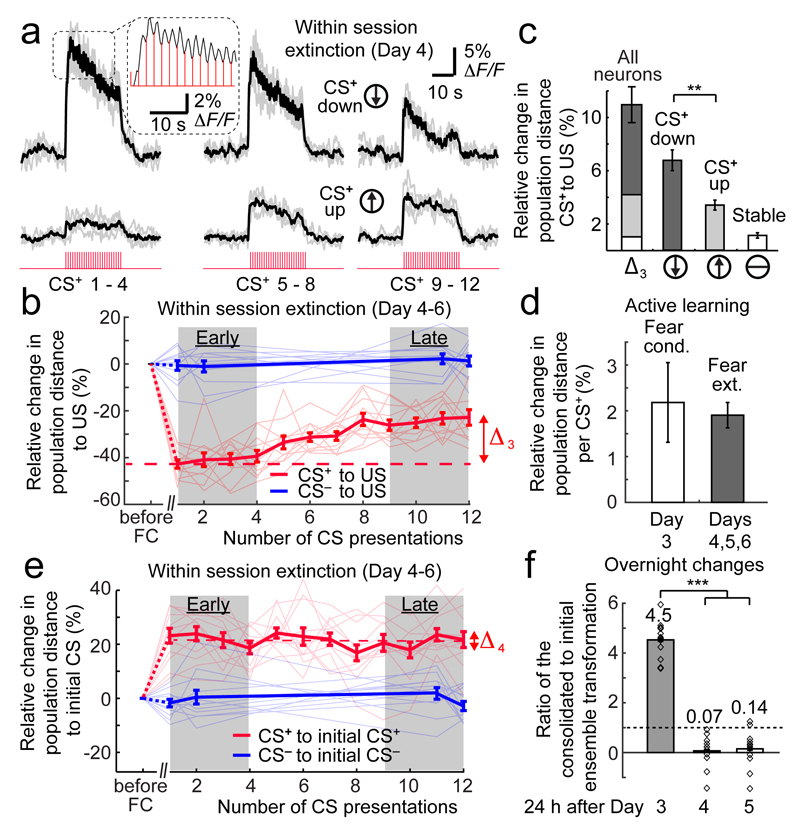
Fig. 4. During behavioral extinction, the CS+ representation becomes more distinguishable from the US representation but does not revert to its initial form.
(a) Ca2+ signals from two neurons, illustrating bi-directional plasticity of CS+-evoked responses in early (left), middle (center), and late (right) phases of fear extinction on Day 4. Gray lines: Individual CS+ presentations (4 per set). Black lines: Mean responses. Inset: Magnified view of responses to individual CS+ tone pulses.
(b) Population vector distances (PVDs) between CS- and US-evoked activity during extinction (Days 4–6), for individual mice (thin lines) and averaged across 12 mice (thick lines) for 12 CS+ and 4 CS– presentations per day (2181 total cells). CS+–US PVDs (red lines) increased by an amount, Δ3, within the sessions after fear conditioning (FC). CS––US PVDs (blue lines) were stable.
(c) Composition of the change in PVD, Δ3, in b, from cells with stable (white), increased (light gray), or decreased (dark gray) responses to the CS+ after training. **: P < 0.001 (signed-rank test; 12 mice).
(d) Within individual sessions, the CS+ representation changed at similar rates during learning and extinction (quantified by the change in mean CS+–US PVD per CS+ presentation; P = 0.6; signed-rank test; 12 mice).
(e) During extinction sessions (Days 4–6), there was little change (Δ4) in the mean PVDs (thick lines) between CS+ and CS– representations and their initial forms before conditioning (averaged over Days 1, 2). Thin lines: data from individual mice.
(f) Overnight consolidation induced long-term changes in the CS+ representation 24 h after conditioning (Day 3) but not after extinction training (Days 4, 5). Horizontal line marks where coding changes from training are neither amplified nor diminished in consolidation. Numbers above each dataset denote mean coding changes occurring overnight after each day, i.e. a 450% increase after Day 3, and reductions to 7% and 14% of their values after Ca2+ imaging on Days 4 and 5. Open diamonds: values from 12 individual mice. ***: P < 0.001 (signed-rank test; Day 3 vs. Days 4 or 5).
All error bars are s.e.m.
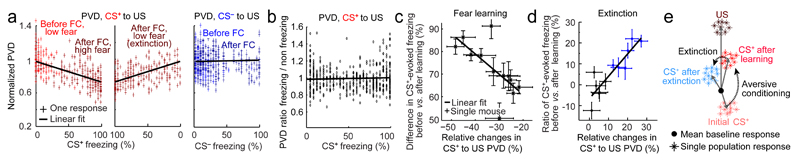
Fig. 5. The similarity of the CS+ and US representations encodes the CS+–US association strength.
(a) CS+–US PVDs for each CS+ presentation, normalized to the CS+–US PVD for the mouse’s first CS+ presentation, are predictive of the freezing level that each CS+ evoked before and during conditioning (left), and during extinction (middle). CS––US PVDs (right) lack this relationship. a and b are based on 3655 neurons in 12 mice. Black lines: linear fits.
(b) Within each 25-s CS+, the 1 s time bins with and without freezing had a near unity ratio between their CS+–US PVD values, irrespective of the evoked freezing level.
(c) How much each mouse (individual data points) exhibited post-training changes in the CS+–US PVD was predictive of its learned, CS+-evoked freezing behavior. Black line: linear fit (r = 0.7; P <10-3). Error bars: s.e.m. across 6 pair-wise comparisons of one day before (Days, 1, 2) and one day after training (Days 4–6) for each mouse.
(d) How much each mouse (data points) exhibited a changed CS+–US PVD between the first 6 CS+ on Day 4 versus the first 6 CS+ on Day 6 was predictive of its loss of CS+-evoked freezing. Black line: linear fit (r = 0.9; P <10-3). Blue points denote mice with significant consolidated extinction (signed-rank test comparing time spent freezing between the two sets of CS+ presentations; P < 0.05). Error bars: s.e.m. across the 6 pair-wise comparisons of one CS+ from among the first six presentations on Day 4 and the corresponding CS+ from the first six presented on Day 6.
(e) Schematic of CS+ population vector changes during learning and extinction. During learning, this vector doubles in length. It also rotates directly toward and becomes less differentiable from the US population vector, supporting a model in which the US representation provides a learning supervision signal. During acute extinction, the CS+ population vector shrinks ~20% and rotates ~5–8° out of the plane defined by the US and the initial CS+.
Unlike acute fear learning, during acute extinction cells with decreased CS+-evoked responses contributed more to the CS+-representation changes than cells with increased responses (Fig. 4c). However, the rates at which ensemble coding changed during acute learning (Day 3) and extinction were equivalent (P = 0.6; Fig. 4d), suggesting a common process for initial storage of a memory and its acute extinction. During within-session extinction the CS+-representation did not revert and gained no more similarity to its initial representation before learning (Fig. 4e) (Δ4 = –2 ± 2%; Friedman Test; P = 0.37). Instead, the CS+ population vector rotated out of the plane defined by the US and the initial CS+ (Fig. 5e), maintaining a 28° ± 3° angle to its initial form that differed little from the 32° ± 3° at the end of learning.
Hence, BLA ensembles explicitly encode extinction training as new learning1. We did not find overt signals of US omission, but sub-threshold signals might drive plasticity in an extinction-specific subset of cells1 (Fig. 4a). Extinction engages hippocampus, thalamus and neocortex23, and their inputs to BLA might signal US omission. Unlike learning consolidation, most coding changes that accumulated in each extinction session reversed before the next session (Fig. 4f), consistent with the modest behavioral extinction that persisted overnight (Figs. 1c, 5d).
We examined how encoding of the CS+–US association related to conditioned behavior. The differentiability of the two representations predicted the overall extent of freezing behavior, throughout learning and extinction (r = 0.7; P < 10-14; Fig. 5a). Yet, on the seconds time-scale, mean CS+–US PVDs were no different between freezing and non-freezing epochs (Fig. 5b). Thus, resemblance of the CS+- and US-encodings predicts the general acquisition, not the instantaneous performance, of learned freezing24. How much the CS+-encoding increased in similarity to the US-encoding strongly predicted the behavior of individual mice during learning and extinction (Fig. 5c,d).
Discussion
Based on recordings of >3600 BLA cells across six days, the analyses here show how neural ensembles represent associative information. The sets of active and CS–-responsive neurons exhibited day-to-day turnover, but the neural ensembles encoded information far more reliably than individual cells7,19,25,26. It is unclear what mechanisms preserve information despite cellular turnover, which might reflect variations in IEG expression that help time-stamp individual memories26–29.
Single cell recordings have shown that neurons in several amygdalar areas can individually depress or potentiate their response properties under various conditions, leading to the impression that depression and potentiation may result from opposing influences on memory storage1,30,31. The recordings here show that learning simultaneously induces potentiation and depression of cells’ CS+-evoked responses in an equally balanced manner (Fig. 2a, 3e). This coordinated bi-directional plasticity was crucial to transforming the ensemble level CS+-representation to increase its similarity to the US-representation (Fig. 3f), was undetectable in past studies using IEG10 or pharmacologic inactivation methods15,17, and mainly occurred during consolidation (Fig. 3d–g).
Notably, our results diverge from the predictions of Hebbian fear-learning1,2,27,32, which invokes a bi-conditional rule requiring coincident CS+ and US signals and posits that among cells receiving CS+ signals, those activated by the US will potentiate their CS+-evoked responses20. Mechanisms associated with this rule, such as NMDA-receptor-dependent synaptic potentiation32, likely contribute to transforming the CS+-representation, but the basic Hebb rule alone does not predict all the observed plasticity.
First, up- and down-regulation of stimulus-evoked responses were equally prevalent and important for transforming coding during learning (Fig. 3e). Second, most cells with potentiated CS+ responses were unresponsive to the US (Fig. 3f; Extended Data Fig. 6). Third, a majority of cells that were CS+- and US-responsive before training had reduced CS+-evoked responses afterward. Fourth, bi-conditional plasticity rules have difficulty explaining why many CS+-responsive cells depress their responses but CS–-responsive cells generally do not (Fig. 2a,d). A mere lack of US-related input cannot explain this difference. Hebbian models require coincident CS+ and US inputs to induce potentiation20, but, in reality, amygdala-dependent fear learning does not require coincidence4,33. Explaining this temporal permissiveness, and the differences in plasticity between CS+- and CS–-responsive cells likely requires a modified Hebb rule.
One possibility is a tri-conditional rule that refers not only to CS and US presentations but also a third factor, such as a neuromodulator or network-wide US-evoked inhibition14,34–36, to explain the plasticity differences between CS+- and CS–-responsive cells (Supplementary Table 2; Supplementary Note). Theorists have studied such ‘neo-Hebbian’ tri-conditional rules37, and both inhibitory signaling and neuromodulator release are crucial for fear learning-induced changes to occur in BLA at normal rates34,35. Our data suggest these network-wide factors might aid ensemble encoding by promoting bi-directional plasticity for CS+–US pairings in close but not strict concurrence38. Nevertheless, different cells might follow different plasticity rules, and some might follow the simple Hebb rule.
The data here naturally suggest an abstract interpretation of how associative information is stored and represented, namely that BLA ensembles implement a supervised learning algorithm39 to encode the CS–US association. Prior studies proposed the US acts as a cellular-level teaching signal20,40. Here, the plasticity of single cells was not strictly determined by US-evoked activity. Instead, US-driven activity seemed to provide an ensemble-level supervision signal, guiding rotation of the CS+ population vector directly toward the US representation (Figs. 3f, 5e), which would have been unapparent in smaller recordings1,40. An attraction of this account is its intrinsic measure of memory strength, the similarity of the US- and CS+-representations. Conditioned freezing closely tracked the US–CS+ PVD for each mouse, strongly supporting this interpretation. Principles of supervised learning might apply to brain areas beyond BLA, and future work should examine if coding transformations similar to those seen here occur in other limbic regions or neocortex.
Methods
Animals
We housed male C57BL6/J mice (Jackson Labs; 9–10 weeks old) under a normal 12 h light/dark cycle, and provided food and water ad libitum. Prior to fear conditioning experiments, we individually housed mice for ≥ 14 days. To habituate the mice to human handling, we handled them at least 7 times in 10 subsequent days. All animal procedures were approved and executed in accordance with institutional guidelines (Stanford Administrative Panel on Laboratory Animal Care). Mice were randomly assigned to different experimental groups in an informal manner, without regard to any of their characteristics.
Viral injection
We performed surgeries when mice were 9–10 weeks of age. We labeled excitatory neurons by injecting an adeno-associated virus (AAV, serotype 2/5) driving expression of GCaMP6m8 via the CaMK2a promoter. In brief, we anesthetized mice with isoflurane (induction, 2%; maintenance, 1–2%) in 95% O2 (Praxair) and fixed them in a stereotactic frame (Kopf Instruments). We stabilized the body temperature at 37° C using a temperature controller and a heating pad. We injected 500 nL of the AAV (injection coordinates relative to bregma: 1.7 mm posterior; 3.4 mm lateral; 4.7 mm ventral) via a borosilicate glass pipette with a 50-μm-diameter tip using short pressure pulses applied with a picospritzer (Parker).
Microendoscope implantation
7–12 days after AAV injection we performed a second surgery to implant either a small custom-designed 0.6-mm-diameter microendoscope probe (Grintech GmBH), or a stainless steel guide tube (1.2 mm diameter) with a custom glass cover slip glued to one end (0.125 mm thick BK7 glass, Electron Microscopy Science). To avoid damage of the internal capsule, we chose implantation coordinates for the tip of the microendoscope that were lateral to this structure (1.7 mm posterior; lateral 3.4 mm; 4.5 mm ventral, all relative to bregma). To perform the implantation, we first made a round craniotomy centered on the injection coordinates using a trephine drill (1.0–1.8 mm in diameter). To prevent increased intracranial pressure due to the insertion of the implant, we made a circular incision in the brain to a ventral depth of 4.5 mm by using a custom-made trephine (1 mm diameter). We aspirated all brain tissue inside the trephine. Next, we lowered either the microendoscope or a metal guide tube to the bottom of the incision. We fixed the implanted guide or microendoscope to the skull using ultraviolet-light curable glue (Loctite 4305). To ensure a stable attachment of the implant, once the cranium had dried we inserted two small screws into it above the contralateral cerebellum and contralateral sensory cortex (18-8 S/S, Component Supply). We then applied Metabond (Parkell) around both screws, the implant and the surrounding cranium. Lastly, we applied dental acrylic cement (Coltene, Whaledent) on top of the Metabond, for the joint purpose of attaching a metal head bar to the cranium and to further stabilize the implant. Mice recovered for 5–7 weeks, at which point we checked the level of GCaMP6m expression using a two-photon microscope and a 20× objective lens (LUCPlan FLN, 0.5 NA, Olympus). If expression was sufficiently bright, we considered the mouse ready for mounting of the miniature microscope (nVista HD, Inscopix Inc.).
Mouse behavior
For studies comparing a range of unconditioned stimuli (Extended Data Fig. 2), on the first day of testing we played eight 10 kHz tones (85 dB, 25× 200 ms duration tone pulses delivered at 1 Hz) while the mice were freely moving in an unfamiliar chamber. After one day of water restriction, we transferred mice to an experimental chamber where they received 30 μL of 4% sucrose water (500 ms reward delivery time). In the same session, after delivery of sucrose water we transferred mice to the conditioning chamber, where we delivered eight electric shocks above one eyelid (3 mA; 2 s duration) or to the paws (0.6 mA; 2 s duration) in a pseudo-random order.
Fear conditioning experiments involved a separate cohort of mice than that used for US comparisons, and took place in two different isolation chambers, chamber A (Day 1, 2, 4, 5 and 6) and chamber B (Day 3). The two chambers differed in their odors, shapes, lighting pattern, and textures of the walls and floor. Prior to each imaging session, we cleaned the chambers with a solution of 1% acetic acid (Chamber A) or 70% ethanol (Chamber B). For scoring of freezing behavior we used video-based freezing detection software (Freeze Frame, Actimetrics) that provided a binary time-trace of the mouse’s movement amplitude. The video frame rate was 20 Hz, but for behavioral analysis we down-sampled the resulting time trace to 5 Hz. Mice were scored as freezing if movement was below a minimum threshold for ≥1 s. To validate the semi-automated detection of freezing, we compared freezing values to a classical time-sampling procedure during which a human observer who was blind to the experimental conditions visually scored freezing behavior. Freezing values with both procedures were nearly identical (92 ± 3%, n = 12 mice).
Throughout habituation, training and extinction sessions, the CS+ and CS– comprised twenty-five, 200-ms-long tone pulses (4 kHz at 85 dB, or 10 kHz at 80 dB, with the twenty-five pulses delivered at 1 Hz). The acoustic frequencies of 4 kHz and 10 kHz were randomly assigned as the CS+ and CS– for the different mice, in a counterbalanced manner. During habituation (Days 1, 2) and conditioning (Day 3), mice received five CS+ and five CS– presentations in a pseudorandom order. During fear testing and extinction sessions (Days 4–6), mice received two CS– presentations before and two CS– after a block of 12 unreinforced CS+ presentations1,41. On all days, the inter-stimulus intervals between CS presentations were pseudo-randomly chosen between 20–180 s.
During conditioning on Day 3, at 800 ms after the termination of the last tone pulse in each CS+, the mouse received a US foot shock. To achieve reliable and robust fear learning, we used a relatively long (2 s) and strong foot shock (0.6 mA), which led to conditioned, CS+-evoked freezing levels (70–90%) comparable to those reported previously for similar US parameters in mice41. This is a form of auditory, associative fear conditioning that is amygdala-dependent1,12,14 (Extended Data Fig. 3) and hippocampal-independent4. We analyzed the behavioral performance of all mice tested and retained the data regardless of freezing levels.
Behavior controls
We examined whether microendoscope implantation affected motor behavior by monitoring mouse locomotion during the first two sessions in Chamber A, for mice that had no, unilateral, or bilateral microendoscope implants. We used a standard video camera (AVT, GuppyPro, F125B) and the image acquisition toolbox in MATLAB to acquire movies of the mouse’s behavior at a 20 Hz frame rate. To extract the mouse’s locomotor trajectory we used a custom video-tracking routine written as a plugin for the ImageJ (NIH) image analysis software environment. From these trajectories we calculated the total distance traveled, mean speed and mean acceleration (Extended Data Fig. 3a, b).
We also investigated if microendoscope implantation affected fear learning by comparing conditioned freezing behaviors for the different groups of mice (Extended Data Fig. 3c–f). In addition to the three groups of mice used for locomotor studies, we also included a group of mice that had bilaterally implanted guide tubes through which we administered the GABAA agonist muscimol 10-15 min before the Day 3 conditioning session. These metal guide tubes had the same outer diameter as the implant used for Ca2+ imaging, and we connected them to a 10-μL micro-syringe (Hamilton) via polyethylene (PE 20) tubing. We dissolved muscimol (Sigma-Aldrich) in artificial cerebrospinal fluid (pH 7.4) and infused this solution bilaterally into each BLA through 33-gauge infusion cannulae, each of which extended 0.5 mm beyond their corresponding metal guide tube. 10-15 min before the Day 3 conditioning session, into each BLA we infused a small volume of 0.3 μL that we delivered using a syringe pump (UMP3, World Precision Instruments) at a rate of 0.2 μL/min. The infusion cannulae remained in place for 1 min after the infusion. The final dosage and volume of muscimol delivered was 2.6 nmol and 0.3 μL per BLA, as in prior fear-conditioning studies in mice42.
Ca2+ imaging using the miniature microscope
We first characterized the optical working distance between the glass surface of the microendoscope inside the brain and the cells at the focal plane, by using a combination of empirical measurements and computational ray tracing simulations of the optical pathway. First, we empirically determined the distance between the back focal plane, where the image of the cells was projected outside the microendoscope, and the microendoscope’s external surface protruding from the cranium. To do this, starting with the miniature microscope in a position such that the cells of interest were in focus, we lowered the microscope toward the microendoscope until we focused upon the microendoscope’s external surface. After noting the distance change between these two focal positions, we used the position of the back focal plane in combination with the microendoscope’s optical design to determine computationally the optical working distance between the cells and the surface of the microendoscope inside the brain. For these computations we used optical ray tracing software (Zemax). This yielded values for the working distance within the range 77–181 μm. Histological reconstructions showed that the tip of the microendoscope generally lay in the lateral amygdala (LA). However, because the optical focal plane often spanned ventral parts of LA and dorsal parts of the basal amygdala (BA), we use the joint term basal and lateral amygdala (BLA) throughout the paper.
To mount the base plate of the miniature microscope on the cranium, we attached the microscope to the base plate and lowered the pair toward the implanted microendoscope until we observed green fluorescent cells. We selected a 600 µm × 600 µm field-of-view (FOV) medial from the non-fluorescent axonal fiber tract that separated the BLA and the cortex (Extended Data Fig. 1c). We glued the base plate onto the skull using blue-light curable glue (Flow-it, Pentron). Afterward, we detached the microscope and returned the mouse to its home cage.
Before each Ca2+ imaging session, we briefly head-fixed the mouse using its metal head-bar while allowing it to walk or run in place on a wheel. We then attached the miniature microscope to its base plate and returned the mouse to its home cage for 50–60 min until the start of the imaging session. Each session involved 22–28 min of Ca2+ imaging across a field-of-view of approximately 600 µm × 600 µm, which we always verified matched that seen in any prior sessions in the same animal. After each session we detached the microscope and returned the mouse directly to its home cage for ~22 h.
To acquire fluorescence Ca2+ imaging videos, we used 100-150 μW of illumination intensity at the specimen and took 12 bit images (1000 × 1000 pixels) at a frame rate of 20 Hz. Each pixel covered 0.6 µm × 0.6 µm in tissue. We streamed the video data directly to hard disk (90–100 MB/s).
Two-photon imaging
To check the expression of GCaMP6m in the BLA, we used two-photon imaging to image the BLA in isoflurane-anesthetized mice (1–2% isoflurane in O2). We head-fixed the mice via the implanted metal head bar and positioned the implanted microendoscope under a 20× microscope objective (Olympus, LCPLFLN20xLCD) of an upright two-photon fluorescence microscope (Bruker). We first used wide-field epi-fluorescence imaging to visualize the BLA tissue through the microendoscope. We then switched to two-photon laser-scanning imaging and acquired images of 256 × 256 pixels at a 3 Hz frame rate.
Basic processing of the Ca2+ imaging videos
To account for slowly varying illumination non-uniformities across the field-of-view, we normalized each image frame by dividing it by a spatially low-pass filtered (length constant: 120 µm) version of the frame using ImageJ software (NIH). Next, we used the ImageJ plugin TurboReg43 to correct for lateral motions of the brain by performing a rigid image registration across all frames of the movie. After motion correction, we temporally smoothed and down-sampled each movie from 20 Hz to 5 Hz. We then re-expressed each image frame in units of relative changes in fluorescence, ΔF(t)/F0 = (F(t) – F0)/F0, where F0 is the mean image obtained by averaging the entire movie.
Cell sorting
We identified spatial filters corresponding to individual neurons using an established, automated cell sorting routine based on principal and independent component analyses7,44. As in prior Ca2+ imaging studies using the miniature microsope7,45, after motion correction we identified cells’ spatial filters based on the Ca2+ data acquired over the entire session. For each filter, we then zeroed all pixels with values <50% of that filter’s maximum intensity. To obtain time traces of Ca2+ activity, for each cell we applied its thresholded spatial filter to the ΔF(t)/F0 movie.
As previously described44, the extracted spatial filters generally had sizes, morphologies and activity traces that were characteristic of individual neurons, but there were also some spatial filters that were obviously not neurons and that we discarded (Extended Data Fig. 2b). For the 4–10% of candidates with less common characteristics, we were conservative and accepted only those that were plainly cells by human visual scrutiny. We verified every cell included in the analyses by visual inspection.
Registration of cell identities across imaging sessions
We generated cell maps for each day by projecting thresholded versions of each cell’s spatial filter onto a single image7 (Extended Data Fig. 5a). Taking the map from Day 3 as a reference, we aligned the other cell maps to this one via a scaled image alignment using the TurboReg image registration algorithm43. This corrected slight translations, rotations, or focus-dependent magnification changes between sessions and yielded each cell's location in the reference coordinate system.
We then identified candidate cells across sessions that might be the same neuron seen on multiple occasions. To do this, we applied the observations that our image registration procedure had sub-micron precision, and that the distance between the centroids of neighboring somata was always >6 µm (Extended Data Fig. 5d). We thus enforced that all observed cells deemed to be the same neuron had all pair-wise separations ≤ 6 µm (Extended Data Fig. 5e). The distribution of pair-wise separations between cells assigned the same identity yielded the conservative estimate that 99.7% of these assignments were correct (Extended Data Fig. 5e inset).
Identification of neuronal sub-classes
We identified functional sub-classes of neurons by comparing the stimulus-evoked fluorescence Ca2+ signals of individual cells to their baseline fluorescence levels, using 1 s time bins. To compute each cell’s baseline activity level, we averaged its fluorescence signal over the complete imaging session excluding all stimulus presentations. For the analyses of neural responses to a CS– or CS+ (always in the form of 25 tone pulses, 200 ms in duration, delivered at 1 Hz), we defined the stimulus response period as the 25-s-period that began at the onset of the first tone pulse and extended 800 ms beyond the offset of the 25th pulse (i.e., up to the start of the US). To analyze neural responses to a shock US, we defined the stimulus response period as the 2 s period of eyelid or foot shock delivery. To analyze the neural responses to sucrose water, we defined the stimulus response period as the 1 s interval starting from the onset of stimulus delivery. After computing each cell’s stimulus-evoked fluorescence responses in 1 s time bins, we compared the set of all such responses to the cell’s baseline activity level using the Wilcoxon rank-sum test. All cells with stimulus-evoked responses that were significantly different from baseline activity (significance criterion: P ≤ 0.01) were classified as CS- or US-responsive.
We also verified that the definition of baseline activity had little effect on the sets of cells identified as having stimulus-evoked responses, by comparing the results obtained using two different definitions. In one case, we determined each cell’s level of baseline activity by finding its average activity across the full imaging session, excluding stimulus presentations. Alternatively, we used the 20-s-period immediately prior to each stimulus presentation to assess the magnitude of the stimulus-evoked response. Using all 3655 cells for this validation analysis, we found that 3524 cells (96%) were categorized identically under the two definitions of baseline activity, indicating that the choice of definition had little effect on our subsequent analysis results.
To identify neurons that significantly increased or decreased their CS-evoked activity during the five paired CS-US presentations on Day 3, we compared their CS-evoked Ca2+ signals for CS presentations early in the session (presentations 1 and 2) versus those late in the session (presentations 3-5) (Wilcoxon rank-sum test, using a significance threshold of P ≤ 0.05). To identify cells with significantly increased or decreased their CS-evoked activity after conditioning, we compared CS-evoked Ca2+ signals from the days before (Days 1, 2) and after (Days 4–6) the training session on Day 3 (Wilcoxon rank-sum test, using a significance threshold of P ≤ 0.05).
Population vector analyses
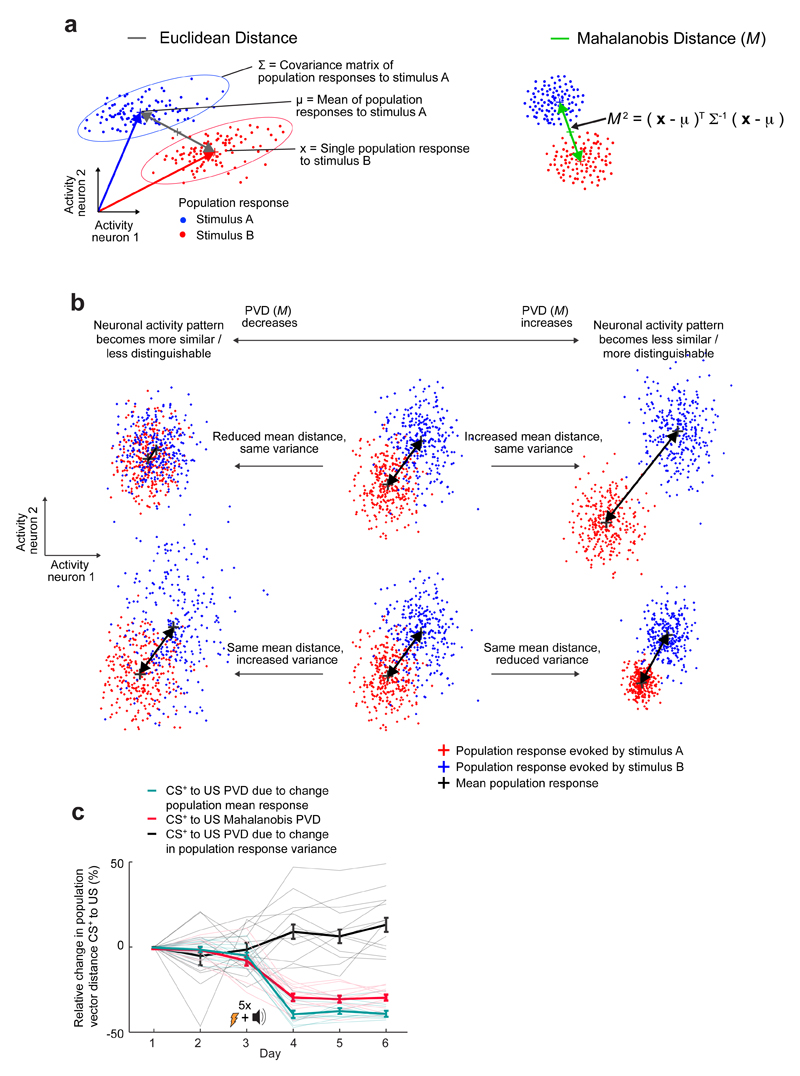
We analyzed our data with MATLAB (Mathworks) using the imaging and machine learning toolboxes. For population vector analysis, decoder training and testing we used neuronal Ca2+ signals expressed as relative fluorescence changes (ΔF/F), down-sampled the traces to 1 s time bins, and organized the data to contain equal numbers of time points for baseline, CS+, CS– or US presentations. We chose 1 s bins, because this choice yielded superior decoding performance compared to the use of either shorter or longer time bins. To quantify the similarity of two sets of neuronal ensemble response patterns, we calculated the Mahalanobis distances between the two sets of population activity vectors21. To do this, we created a group of N-dimensional (N = number of neurons) activity vectors, x, for each stimulus type (baseline, CS–, CS+ or US) and calculated the population vector distances (PVD) between the two groups (Extended Data Fig. 10). For example, the Mahalanobis PVD (M) between sets of CS- and US-evoked ensemble activity patterns is:
where x and μ are individual and mean population vectors for CS and US ensemble responses, respectively, and xT and μT are their transposes. Σ is the covariance matrix for the set of ensemble responses. The Mahalanobis distance takes into account the differences in the means of the two sets of ensemble responses as well as their co-variances (Extended Data Fig. 9).
To track the CS-US PVDs across the Day 3 training session, we down-sampled all neural activity traces to one-second time bins. This resulted in 25 time bins for each twenty-five-second CS presentation and two time bins for each two-second US presentation. Next, we constructed the mean CS+, CS– and US population vectors by averaging the evoked neural responses over all five presentations of each stimulus and all the time bins associated with each stimulus presentation. To calculate the change in the CS+ population vector expected under a cellular, Hebbian model of associative potentiation, we restricted the changes to the CS+ population vector to those cells that were US-responsive and used the empirically determined mean stimulus-evoked responses of these cells to calculate the vector entries.
Decoding ensemble neural activity
We constructed all binary (Fig. 3b; Extended Data Fig. 8) and three-way (Fig. 3a; Extended Data Fig. 7) Fisher linear decoders21 in MATLAB. To construct the three-way decoders, we used an established approach based on multiclass Fisher linear discriminant analysis that maximizes the ratio of the mean variances between the different classes to that within the individual classes21. We used the set of neural ensemble Ca2+ response traces (ΔF/F) from each mouse and trained decoders to discriminate the Ca2+ activity patterns that occurred during baseline epochs, CS+ or CS– presentations. Before training we down-sampled the data into 1 s time bins. We determined decoder performance values as the mean rate of correct predictions over a 10-fold cross-validation. For cross-validation, we split each dataset into ten equally sized blocks and randomly assigned each time bin to one of the ten blocks; we used nine of the blocks for decoder training and one for testing. To evaluate the statistical significance of decoding performance, we trained control decoders on temporally shuffled datasets, and compared the mean, cross-validated performance values to those of the real decoders.
When making comparisons across decoders involving unequal numbers of cells (Fig. 3a), we confirmed all results via a control analysis that used statistical re-sampling methods46 to construct decoders based on equal numbers of cells; this yielded decoding results virtually indistinguishable from those shown in Fig. 3a. As a further check, we also verified that the small performance difference between decoders based on all cells and those based only on CS-responsive neurons was not simply due to the smaller number of cells used for the latter decoders, as opposed to the information content of their activity traces. For this purpose, we constructed control decoders (Fig. 3a; dashed green curve) based on the same number of cells as used for the decoders of CS-responsive cells, but with the cells randomly chosen. The accuracy difference between these control decoders and that of the decoders of CS-responsive neurons was dramatic, as the control decoders performed at levels very close to chance and no better than decoders based on temporally shuffled neural activity traces (Fig. 3a; dashed gray curve).
Construction of the CS+ rescue decoder
We constructed and validated the rescued time–lapse CS+ decoder in five main steps (Extended Data Fig. 10a,b). Step 1: We recorded CS+ ensemble activity before conditioning (Day 1 and 2). Step 2: We recorded neuronal population activity during conditioning with five CS-US paired presentations (Day 3) and identified individual neurons that altered their CS+-evoked responses [Wilcoxon rank-sum test, comparing CS+-evoked responses between the early (CS-US pairings 1 and 2) and late phase (CS-US pairings 3-5) of conditioning, significance threshold P < 0.15]. Step 3: We simulated the full, consolidated CS+ ensemble transformation by gradually extrapolating changes of individual neuron responses and adding them to their CS+ responses before conditioning (Extended Data Fig. 10c). Step 4: We trained a new rescue decoder and evaluated its performance for different extrapolation magnitudes. Step 5: To validate the simulated transformation of ensemble coding, we compared the average performance of the rescue decoder to the average performance of the stable CS– time-lapse decoder.
Relating neural population vectors to freezing behavior
To examine how ensemble neural activity related to each mouse’s overall level of conditioned freezing (Fig. 5a), we first calculated for each individual CS+ (or CS–) presentation the PVD to the mean US population vector, and then normalized the resulting CS–US PVD by the value of the CS–US PVD computed for the mouse’s first CS+ (or CS–) presentation. We plotted these normalized CS–US PVD values as a function of the percentage of time during each 25-s CS presentation that the mouse spent freezing (Fig. 5a).
To examine whether BLA ensemble neural activity differed between the moments within individual CS+ presentations when a mouse was freezing versus not freezing (Fig. 5b), we divided each 25-s CS+ presentation into 1 s time bins. For each CS+ presentation we then found the ratio of the CS+–US PVDs, as computed for the 1-s time bins when the mouse was freezing versus those when the mouse was not freezing. We plotted this ratio as a function of the proportion of time during the 25-s CS+ that the mouse spent freezing (Fig. 5b).
Next, we examined how the change in each mouse’s CS+–US PVD during learning related to the change in its freezing behavior (Fig. 5c). For each mouse we computed the percentage change in the CS+–US PVD occurring between the last six CS+ presentations before learning (Days 1 and 2) and the first six CS+ presentations after learning (Day 4). We plotted the resulting values versus the changes in freezing behavior across the same time periods.
We performed a similar analysis to examine how the change in each mouse’s CS+–US PVD during extinction training related to the consolidated change in its freezing behavior (Fig. 5d). We compared the first six CS+ presentations from the first day of extinction training (Day 4) to the first six CS+ presentations on the last day of extinction learning (Day 6). For each mouse we computed the percentage differences in CS+–US PVDs across these two groups of CS+ presentations, and we compared the resulting values to the ratio of the mouse’s freezing levels during these two sets of CS+ presentations.
Histological verification of cell identity
Four weeks after injection of the GCaMP6m viral construct or two weeks after the imaging experiments. we transcardially perfused mice with phosphate-buffered saline (PBS) followed by ice-cold 4% paraformaldehyde (PFA). Next, we extracted mouse brains and kept them for post-fixation in PFA for 24-48 h. We then cut 100-μm-thick coronal brain slices using a Vibratome (VT1000s, Leica) and stored all slices in PBS.
To validate the implant positions of the microendoscopes relative to the BLA we mounted all coronal brain slices on microscopy slides and acquired large field-of-view fluorescence images using a standard fluorescence macroscope (Z16, Leica). We then overlaid all images with a validated reference image47, marked the position of the endoscope tip for every mouse (Extended Data Fig. 1b), and determined the ventral depth of the implant with respect to bregma, using the coordinate system of the reference image.
To stain inhibitory or excitatory neurons, we followed standard immunostaining procedures. In brief, we incubated brain slices with the primary antibodies, rabbit anti-GAD65 (1:500, catalogue no. AB1511, EMD Millipore) or anti-Neurogranin (1:10000, catalogue no. 07-425, EMD Millipore) at 4° C overnight followed by a second overnight incubation at 4 °C with secondary anti-rabbit Alexa 647 antibodies (1:1,000, both Invitrogen).
Data analyses and statistical tests
We conducted all analyses using custom routines written in MATLAB (Mathworks) and ImageJ (NIH) software. Throughout the paper we used two-tailed, non-parametric statistical tests to avoid assumptions of normal distributions and equal variance across groups. All signed-rank tests were Wilcoxon signed-rank tests. All rank-sum tests were Wilcoxon rank-sum tests. For analyses of variance (ANOVA), we used the Friedman and Kruskal-Wallis tests, respectively, for ANOVAs with and without repeated measures. Supplementary Table 1 summarizes the results from these ANOVA analyses. The sizes of our mice samples were chosen to approximately match those of previous work, as there was no pre-specified effect size. Investigators were not blind to an animal’s experimental cohort.
Code and Data availability
The algorithm used for image registration is available on its author’s website43. The algorithm used for cell sorting is available as published supplementary material44. Other software code and the data that support the findings of this study are available from the corresponding author upon reasonable request.
Extended Data
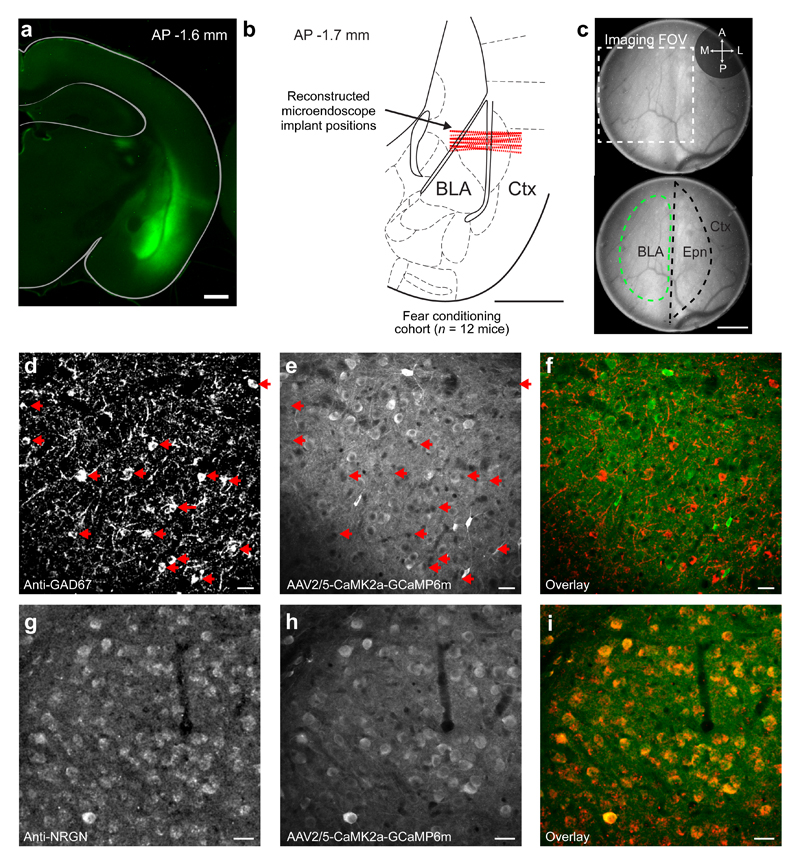
Extended Data Fig. 1. Mouse preparation for Ca2+ imaging in excitatory BLA neurons.
(a) Coronal slice of a mouse brain showing expression in the BLA of the GCaMP6m Ca2+ indicator, five weeks after injection of the AAV2/5-CaMK2a-GCaMP6m virus. Scale bar: 1 mm.
(b) Schematic of a coronal mouse brain section shown with the reconstructed positions (dashed red lines) of the microendoscope implants in the BLA, for the 12 mice subject to the experimental protocol of Fig. 1c. The focal planes for in vivo Ca2+ imaging were 77–181 µm below the indicated implant positions, as determined through computational modeling of the microendoscope’s optical pathway done using the empirically determined value of the back focal length. Hence, the optical focal plane often spanned ventral parts of lateral amygdala (LA) and dorsal parts of the basal amygdala (BA), motivating our use the joint term basal and lateral amygdala (BLA) throughout the paper. AP: anterior posterior. Ctx: cortex. Scale bar: 1 mm. The mouse brain section in this figure has been reproduced with permission from47.
(c) Top, Wide-field fluorescence image of BLA tissue acquired through an implanted microendoscope, six weeks after injection of the AAV2/5-CaMK2a-GCaMP6m virus. The outer fiber tract enclosing the BLA does not express GCaMP6m and appears as a vertical dark stripe in the center of the field-of-view. The dashed box shows the position of the camera’s field-of-view, which we positioned over the BLA by using the fiber tract as a reference marker. Bottom, The same image as in the top panel, but with the boundaries of the BLA and endopiriform nucleus (Epn) marked in green and black dashed lines, respectively. Ctx: piriform cortex. Scale bar: 0.2 mm.
(d–f) Coronal section of a mouse brain showing, d, inhibitory neurons in the BLA immuno-labeled with a monoclonal anti-GAD67 antibody; e, neurons expressing GCaMP6m under the control of the CaMK2a promoter; and f, the overlay of the images in d and e. Red arrows in d and e mark GAD67-positive interneurons that are not expressing GCaMP6m. Scale bars: 20 μm.
(g–i) Coronal brain section showing, g, excitatory neurons in the BLA immuno-labeled using a polyclonal anti-Neurogranin (NRGN) antibody48; h, neurons expressing GCaMP6m; and i, an overlay of the images in d and e, showing that the set of NRGN-positive excitatory neurons (labeled red) strongly overlap with the set of cells expressing GCaMP6m (labeled green). Scale bars: 20 μm.
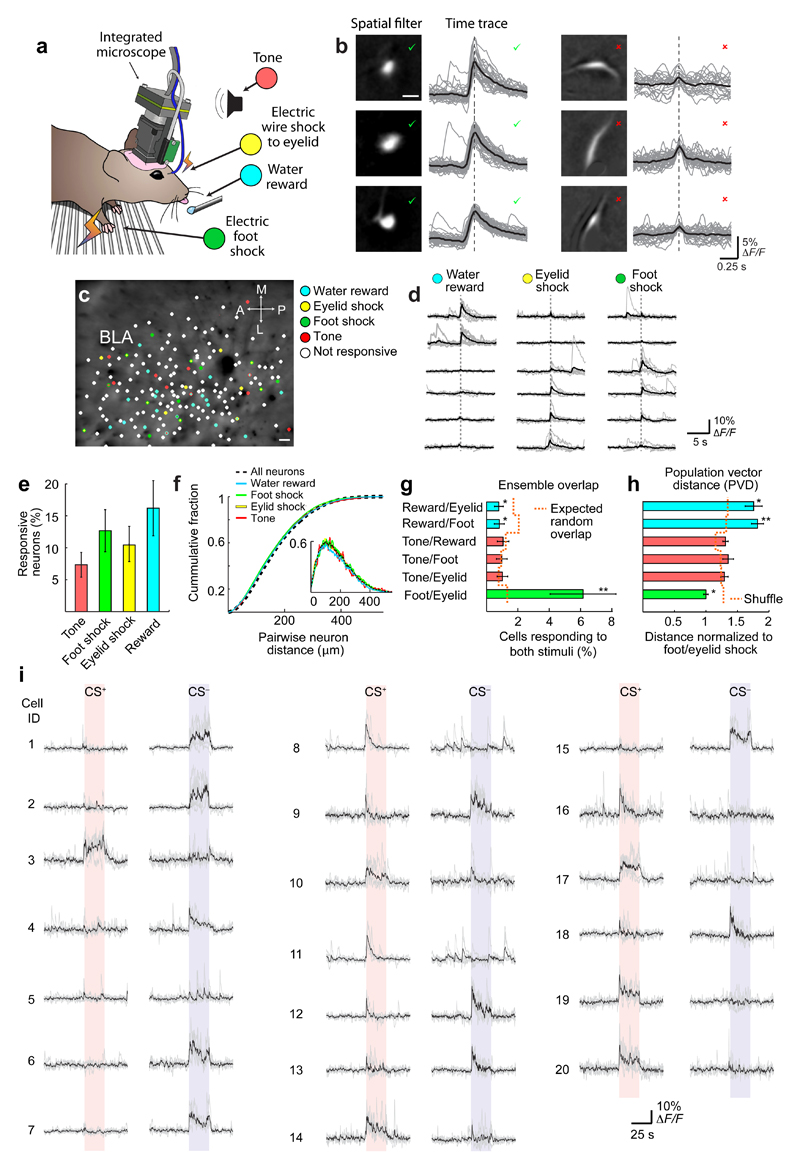
Extended Data Fig. 2. Stimuli of neutral, positive, and negative valence activate sparse, largely distinct, spatially intermingled subsets of neurons in the BLA.
(a) A miniature fluorescence microscope enabled large-scale neural Ca2+ imaging in the BLA of awake behaving mice as we presented stimuli of different valences to the animals.
(b) Candidate cells identified using an automated cell sorting routine7,44 were easily segregated into those (left column) with sizes, morphologies and Ca2+ activity traces (gray traces, individual activity transients; black traces, mean waveforms) characteristic of individual neurons, and those that were obviously not neurons (right column). For the 4–10% of candidates with less common characteristics, we were conservative and accepted only those that were plainly cells by human visual scrutiny. We verified every cell included in the analyses by visual inspection.
(c) An example cell map in the BLA, as determined from a Ca2+ imaging dataset 28 min in duration. Colors indicate the subsets of BLA neurons that responded to rewards (light blue), electric foot shocks (green) or eyelid shocks (yellow), or neutral tones (red). Scale bar: 20 μm. (Significance threshold: P ≤ 0.01, rank-sum test, comparing evoked Ca2+ signals to baseline levels).
(d) Ca2+ responses of six example neurons in the same mouse following the delivery of individual water rewards (left), eyelid shocks (middle) or foot-shocks (right). Gray traces show the Ca2+ responses from eight individual trials. Black traces show the mean responses.
(e) Mean ± s.e.m percentages of cells (n = 1251 neurons in total from 8 mice) with significant Ca2+ responses to the four different stimuli (Threshold for a significant response: P ≤ 0.01, comparing evoked versus baseline Ca2+ levels for n = 8 presentations of the stimulus; Wilcoxon rank-sum test).
(f) Cumulative probability distributions, each determined as a mean over 8 mice (1251 total cells), of the centroid separations between all pairs of cells in each mouse (dashed black curve), and between pairs of cells that both had significant responses to one of the four different stimuli (colored curves). Inset: The corresponding probability densities.
(g) Mean ± s.e.m. percentages of all neurons (n = 8 mice; 1251 cells in total) that had significant responses to each of the two stimuli in each pair listed on the vertical axis. Dashed orange line indicates the expected levels of overlap due to random chance. * denotes P < 0.05 and ** denotes P < 0.01, comparing the actual percentages versus those determined from datasets in which we randomly shuffled the cells’ identities (1000 random shuffles; Wilcoxon signed-rank test).
(h) Mean ± s.e.m. Mahalanobis population vector distances (PVD) between the ensemble neural representations of the two stimuli of each pair listed on the vertical axis. All PVD values are normalized to the PVD between the neural representations of eyelid-shock and foot-shock. Pairs of stimuli with smaller PVD values have ensemble neural representations of greater similarity than pairs with larger PVD values. Dashed orange line indicates the PVDs between ensembles in which we randomly shuffled the cells’ identities (1000 random shuffles). * denotes P < 0.05 and ** denotes P < 0.01, comparing the actual PVD values versus those determined for the shuffled datasets (Wilcoxon signed-rank test). Data are based on the same 1251 cells as in e–g.
(i) Twenty sets of fluorescence Ca2+ traces, showing evoked responses to presentations of the CS+ and CS– from 20 example neurons prior to fear conditioning. Light gray traces show the cells’ individual responses to each of five stimulus presentations; black traces are average responses. Traces were down-sampled to 5 Hz to aid visualization.
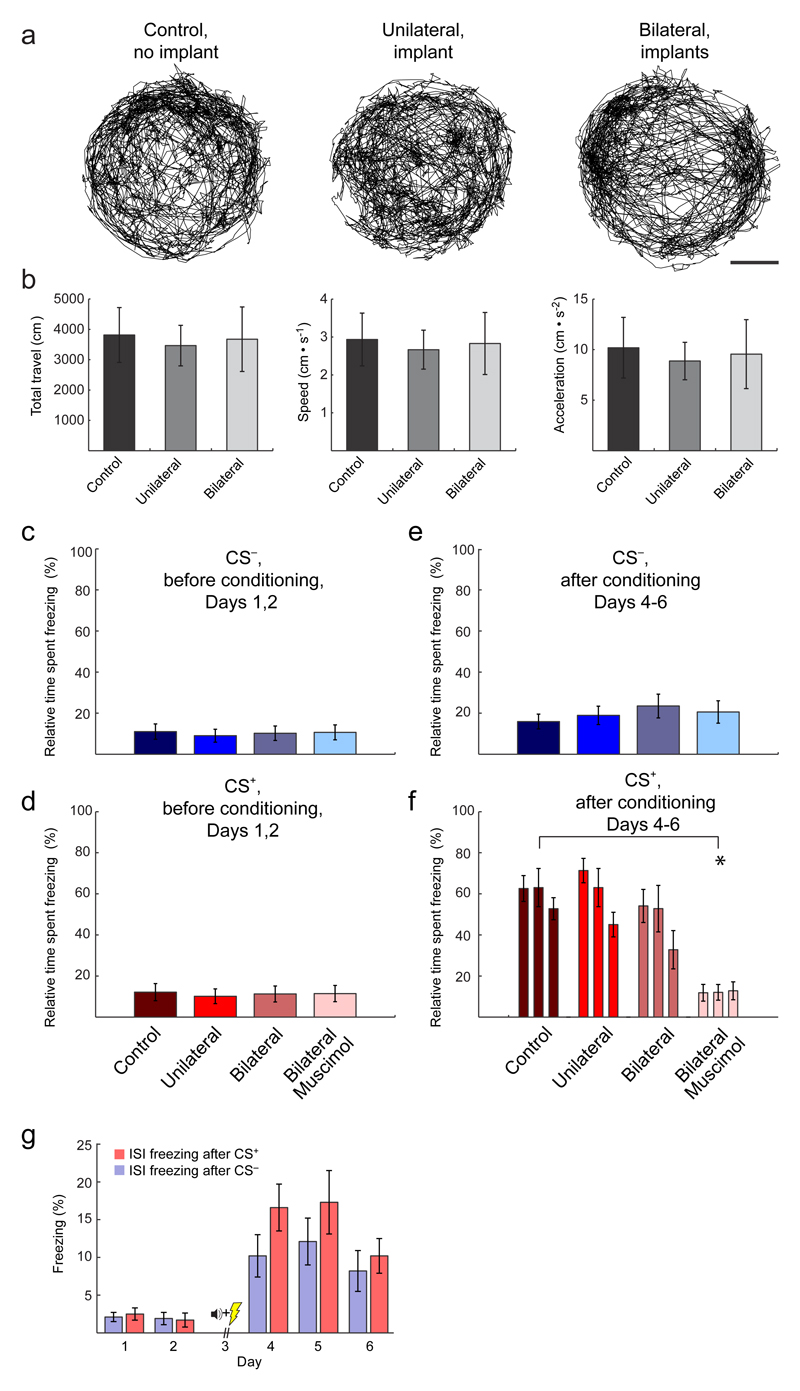
Extended Data Fig. 3. Unilateral implantation of a microendoscope implantation does not alter conditioned freezing; bilateral implantation minimally alters conditioned freezing without affecting locomotion.
(a) Traces of locomotor activity across an entire (22 min) habituation session (Day 1), for one example mouse in each of the three different experimental groups indicated. Scale bar: 5 cm.
(b) Mean ± s.e.m. values of the total distance traveled (left), locomotor speed (middle) and acceleration (right) for the three different groups of mice during the Day 1 habituation session. There were no significant differences between the three experimental groups [no-implant control (12 mice); unilateral implant (12 mice); bilateral implant (10 mice)] in any of the three movement-related parameters (One-way Kruskal-Wallis Test; degrees of freedom: dfgroup = 2, dferr = 31, dftotal = 33 for 3 groups and 34 total mice; χ2 = 10–12; P ≥ 0.05 for all three parameters).
(c, d) Mean ± s.e.m. percentages of time mice spent freezing before conditioning (Days 1, 2) in response to 5 presentations of the CS–, c, and 5 presentations of the CS+, d, in control mice with no implant (12 mice), mice with a unilateral implant (12 mice), mice with a bilateral implant (10 mice), and mice that had a bilateral implant and received a muscimol injection into the BLA before the Day 3 conditioning session (8 mice). There were no significant differences in freezing time between any of the groups (One-way Kruskal-Wallis Test; degrees of freedom: dfgroup = 3, dferr = 42, dftotal = 45 for 4 groups and 42 total mice; χ2 = 10.2 and 11.8 for CS+ and CS–, P ≥ 0.05 for both CS+ and CS–).
(e, f) Mean ± s.e.m. percentages of time mice spent freezing after conditioning (Days 4–6) in response to 4 presentations of the CS–, e, and during 3 sets each comprising 4 presentations of the CS+, f, in the same 42 mice as in panels c, d. * denotes P = 0.005 (Wilcoxon signed-rank test; bilateral muscimol group vs. control; significance threshold = 0.02 after Dunn-Šidák correction for 3 comparisons). These data are consistent with past work showing the necessity of BLA for auditory fear conditioning49 and further demonstrate that the BLA we are imaging are functional and necessary for the behavior.
g) Mean ± s.e.m. percentages of time mice (n = 12) spent freezing during the 20–180 s inter-stimulus intervals following either a CS+ or CS– presentation. CS+ and CS– freezing values are averages over the numbers of stimulus presentations shown in Fig. 1c. After fear conditioning, CS–-evoked freezing levels were above those during the inter-stimulus intervals, indicating the CS– did not serve as a learned safety signal.
Extended Data Fig. 4. Ca2+ transient responses of individual BLA neurons to CS presentations closely resemble expectations based on electrical recordings of these responses.
To check whether fluorescence Ca2+ imaging in the BLA captured similar forms of neural activity as prior extracellular electrical recordings in this brain area, we compared individual neurons’ responses to CS presentations, as observed using the two recording modalities in two different sets of mice presented the same CS stimuli. Across the two datasets, there was close agreement between the shapes of the empirically determined Ca2+ transient waveforms and the expected waveforms based on the electrically recorded CS-evoked spiking responses.
(a) We took a recording of CS-evoked spiking activity from an individual BLA cell (left), convolved the spike train with a decaying exponential function (700 ms time constant) to account for the kinetics of the GCaMP6m indicator (middle), and subtracted the baseline fluorescence level to yield a predicted CS-evoked Ca2+ fluorescence signal (ΔF/F) whose waveform shape closely matched the actual CS-evoked Ca2+ fluorescence signal of a BLA cell that we had monitored using the miniature microscope (right).
(b) Six additional examples of individual BLA neurons’ CS-evoked spiking responses, as monitored via extracellular electrical recordings (black traces). From these spike trains, we used the approach of panel a to predict the Ca2+ fluorescence signals that these cells would produce (red traces) and compared these predictions to the actual CS-evoked Ca2+ fluorescence signals of another six BLA cells that we had studied by Ca2+ imaging and that had similar responses (blue traces).
Extended Data Fig. 5. Precise spatial registration of the Ca2+ imaging datasets from different behavioral sessions allows unambiguous tracking of individual cells across multiple days.
Using the spatial filters provided for each neuron by the automated cell sorting algorithm7,44, we made maps of all active cells detected in the BLA on each day of the study. We then used standard methods of image alignment43 to register these maps across the different days. Approximately 50% of all neurons observed across the entire experiment were detected as active on individual days.
(a) Example maps of active BLA cells from three mice on the first (left), third (middle), and last (right) day of the six-day experimental protocol (Fig. 1c). Circles indicate cells that were active in only one of the three recordings (gray), on two of the three days (blue), or on all three days (red).
Scale bar: 30 μm. The maximum number of active cells seen in one session was 192.
(b) Thresholded spatial filters from three example cells registered across the six-day experimental protocol. Green asterisks indicate each cell’s centroid position on Day 1. Blue asterisks mark each cell’s centroid positions on subsequent days. Scale bar: 10 μm.
(c) Five examples of neighboring cells detected via their activity patterns on different days of the experiment. In each case, the two individual cells are clearly discernible. Scale bar: 10 μm.
(d) Cumulative histogram of the distances between the centroids of all pairs of cells detected within the same imaging session, plotted with a logarithmic scale on the y-axis. Inset: A magnified view of the portion of the graph enclosed in the dashed box. No pairs of cells were separated by <6 µm.
(e) Cumulative histogram of the distances between the centroids of all pairs of active cells registered as being the same neuron seen in different sessions. Inset: Magnified view of the plot for y-axis values >97%. Because the worst-case alignment error of the image registration algorithm was 1.5 µm, as determined by a bootstrap analysis7, and since all pairs of anatomically distinct cells were separated by ≥ 6 µm (panel d), cell pairs separated by < 4.5 µm were virtually guaranteed to be the same neuron seen on two different occasions. This yielded the worst-case estimate that >99.7% of all cell pairs registered as being the same cells were correctly assigned the same identity. This estimate is conservative in that the image registration errors were usually <1 µm.
(f) A plot of the mean ± s.e.m probability that an active neuron detected in one imaging session will also be active in a subsequent session, for all 3655 neurons in the study (black points) and for CS–-responsive neurons (gray points). Inset: Mean ± s.e.m. probability that a cell detected on any day in the study was present in each of the imaging sessions, for all 3655 neurons in the study (black trace), the CS+-responsive neurons (red), and the CS–-responsive neurons (blue). These probabilities were constant throughout the study and statistically indistinguishable between the three groups of cells examined for all days and all mice [49 ± 2% (s.e.m.) of all cells were active each day; Two-way Friedman Test; degrees of freedom: dfdays = 5, dfgroup = 2, dfinteraction = 10, dferr = 198, dftotal = 215 for 6 days, 3 groups of cells and 12 mice; χ2 = 1.6–7.5; P > 0.05 for all three P-values].
(g) The total number of neurons detected in each mouse was stable across all days of the study (152 ± 14 cells per day; mean ± s.e.m.; n = 12 mice; One-way Friedman Test; degrees of freedom: dfdays = 5, dferr = 55, dftotal = 71 for 6 days and 12 mice; χ2 = 5.9; P = 0.31).
(h) Mean ± s.e.m. percentage of all 3655 cells in the study that were detected in one to six sessions.
Extended Data Fig. 6. Conditioning induces bi-directional changes in CS-evoked responses.
Contrary to the predictions of the cellular, Hebbian model of fear learning, conditioning induced substantial bi-directional changes in the CS+-evoked responses of cells that responded to the US and of cells that did not respond to the US. Notably (panel a), a preponderance of cells that responded to both the CS+ and US before training had decreased CS+-evoked responses after training. Further (panel b), many cells with potentiated CS+-evoked responses after training were not US-responsive. Ensemble level analyses showed that cells with up- and down-regulated responses made equally important contributions to the learning-induced changes in ensemble neural coding (Fig. 3e).
(a) Mean ± s.e.m percentages of CS–-responsive cells that were also US-responsive (blue) and of CS+-responsive cells that were also US-responsive. The latter data are further divided into those cells that increased their CS+-evoked responses after training (maroon), those that underwent no significant changes in their CS+-evoked responses (pink), and those that decreased their CS+-evoked responses after training (red).
(b) Mean ± s.e.m percentages of CS–-responsive cells that were not US-responsive (blue) and of CS+-responsive cells that were not US-responsive. The latter data are further divided into those cells that increased their CS+-evoked responses after training (maroon), those that underwent no significant changes in their CS+-evoked responses (pink), and those that decreased their CS+-evoked responses after training (red).
All data are from the same 12 mice.
Extended Data Fig. 7. BLA ensembles provide sufficient information to decode the CS, and the decoding accuracy improves with successive tone presentations in a series of tones.
(a) Left: A three-way decoder has three possible outputs (CS+, CS– and baseline) and hence different categories of possible errors. When a decoder makes a Type A error, it outputs the wrong CS (i.e. CS+ instead of CS–). When a decoder makes a Type B error, it fails to distinguish a CS presentation from baseline activity. Right: When we used all neurons’ activity traces to train the decoders, they determined the correct answer on 97 ± 1% (mean ± s.e.m.) of all trials from a testing set comprising equal numbers of samples of each type. The success rate was 90 ± 2% when we trained the decoders using only those cells with statistically significant responses to at least one of the CS types.
(b) For trials that were incorrectly decoded, the pie charts show the proportions of the two types of errors, for decoders trained on the activity traces of all neurons (left), and for those trained using only neurons with statistically significant responses to at least one of the two CS types (right).
(c) Type A errors (mean ± s.e.m.) declined sharply during the first 5 of the 25 CS tone pulses (black curve), both before (left panel), and after (right panel) conditioning. After conditioning, as the 25 tone pulses proceeded the mice increasingly distinguished between the CS– (blue curve) and the CS+ (red curve), as seen by the differences in evoked freezing behavior.
(d) Schematic showing how we extracted the principal components (PCs) of the BLA ensemble responses to CS tone presentations. Dashed box encloses two PCs, used in panel e for illustration.
(e) Plots of the first two PCs, determined as in d for four example mice, illustrate that the ensemble responses to the CS+ (red points) and the CS– (blue points) were generally distinguishable. Black stars mark the first out of 25 tone pulses for each CS presentation and illustrate that the initial tones in the series were generally the hardest to categorize correctly.
Extended Data Fig. 8. Fisher linear discriminant analysis of BLA population activity.
(a) A notional schematic showing the Mahalanobis distance and the discrimination boundary of a Fisher linear discriminant analysis (FLDA) decoder (black dotted line), which discriminates the multi-dimensional, neural ensemble responses to CS presentations from the activity patterns during baseline conditions. For simplicity, the schematic shows a hypothetical case in which the ensemble consisted of only two neurons, but the basic principles readily apply to larger ensembles. For a given set of training data, the Fisher decoder provides the a posteriori probability that a representative data sample will be correctly categorized.
(b) Example histogram from one mouse showing BLA ensemble responses (Day 1) to CS presentations, normalized and projected onto the dimension of maximal discriminability. The dashed vertical line marks the classification boundary of the Fisher linear decoder, dividing those ensemble responses classified as baseline from those classified as representing a CS. The separation between the two peaks in the histogram is an empirical estimate of the Mahalanobis distance, which is a multi-dimensional generalization of the discriminability index, d’, used in statistics21.
(c) Mean decoding performance as a function of the number of cells used for training the decoder (open circles) and corresponding parametric fits to a sigmoid function. When the training and testing data came from the same day (black curve), performance asymptotically approached near perfect decoding when more than ~100 cells were used. When the training and testing data came from different days (red curve), our datasets were not large enough to approach the asymptote. However, the sigmoidal fit suggests that the asymptotic performance of time-lapse decoders would be ~90% in cases with more than ~120 cells. Shading indicates s.e.m.
(d) Mean ± s.e.m. decoding performance of time-lapse CS– decoders, as a function of the elapsed time between the day on which the training dataset was acquired and the day on which the testing dataset was acquired (n = 12 mice). Despite cells’ declining re-occurrence probabilities as a function of elapsed time (Extended Data Fig. 5f), decoding performance remained stable for time-lapse intervals of 1–5 days.
Extended Data Fig. 9. The Mahalanobis distance quantifies the discriminability of two sets of ensemble responses and takes into account the mean and covariance of each response set.
(a) Left: Schematic illustration of two sets of ensemble neural responses (blue and red clouds of data points). The Euclidean distance (gray line) between the means of the two distributions does not take into account the degree to which the ensemble neural responses are variable from trial to trial. Right: To characterize the differentiability of the two response sets in a way that takes into account neural variability, we determined the Mahalanobis distance (M) between the two distributions. To do this, we first used the covariance matrix of the ensemble neural responses (∑) to map the data points into a space in which the distributions have unity variance in all directions. We determined M by calculating the distance between the means of the two resulting distributions.
(b) A change in the Mahalanobis PVD can be due to changes in the means of the two sets of ensemble responses, changes in response variability, or both. The schematic illustrates these two different ways in which the sets of ensemble responses can become more or less differentiable. The top row shows a pair of cases in which changes in the mean ensemble responses dominate the change in the PVD. The bottom row shows a pair of cases in which changes in response variability dominate the change in the PVD.
(c) We divided the total change in the CS+–US PVD (red curve) induced by learning into two components, a component due to changes in the mean CS+-evoked response (cyan curve) and a component due to changes in the variability of the CS+-evoked responses (black curve). After conditioning, the CS+-evoked responses became less variable (black curve) but also more similar to the US-evoked ensemble responses (cyan curve). The latter effect substantially outweighed the former, leading to a net ~32% decline (red curve) in the differentiability of the CS+- and US-evoked responses, as quantified by the net decrease in the Mahalanobis distance. Thin lines show the values from each of 12 individual mice. Thick lines show the mean values. Error bars are s.e.m.
Extended Data Fig. 10. Procedure for computational rescue of the CS+ decoders.
Unlike time-lapse CS– decoders, which worked well across all six days of the experiment, time-lapse CS+ decoders did not work well when the training and testing datasets came from a pair of days that spanned across the conditioning session (Fig. 3b). This failure mode for the CS+ decoders arises from the learning-induced changes in the ensemble representation of the CS+ (Fig. 3c,d). However, we found that by extrapolating the changes in the CS+ representation that occur during the training session on Day 3, we could predict the much greater, subsequent changes in the CS+ representation that occur before the next session on Day 4 and thereby rescue the failures of the time-lapse CS+ decoders. This figure schematizes the procedure for the computational rescue.
(a) Schematic illustration of how conditioning-induced changes in CS+-evoked ensemble activity (light and dark red dots) can impair the performance of a time-lapse decoder trained on data from before fear conditioning and tested on data from after conditioning.
(b) Through five main steps, we computationally simulated the changes in the CS+-representation that occurred during post-training consolidation, by extrapolating by a factor, q, the much smaller changes in the CS+-representation that occurred during the Day 3 training session.
(c) To determine the optimal value of q, the extrapolation factor, we simulated the post-training changes in the CS+-representation by computationally adjusting the CS+ population vectors in increments of ΔA, the modest change in coding that occurred on Day 3. Increments of 3–5 times ΔA were optimal, in that they best rescued the capabilities of two-way decoders trained on either of Days 1 or 2 to detect a CS+ presentation when tested on data from after training (Days 4–6), or vice versa. (Each datum shows the mean ± s.e.m decoding performance, averaged across 12 mice and the 12 possible pair-wise combinations per mouse of one pre- and one post-training day). The same analysis of the CS– representation scarcely yielded any change in decoding performance, because the effects of training (ΔA) for the CS– were negligible. Decoders trained on temporally shuffled data (1000 shuffles; gray curve) and those based only on cells with up- (green) or down-regulated (purple) responses to the CS+ after training performed less successfully than decoders based on all cells (brown).
Supplementary Material
Supplementary Information is linked to the online version of the paper at www.nature.com/nature.
Acknowledgements
G. Venkatraman, B. Ahanonu, J. Li , B. Rossi, C. Herry, S. Ciocchi and J. Bacelo provided technical assistance. We appreciate Swiss National Science Foundation (B.F.G.), Ambizione (J.G.), U.S. National Science Foundation (L.J.K.), Stanford University (L.J.K., J.D.M.), Simons Foundation (L.J.K.), and Helen Hay Whitney Foundation (M.C.L.) fellowships. A.L. received support from the Swiss National Science Foundation, Novartis Research Foundation, and an ERC Advanced grant. M.J.S. received support from HHMI and DARPA.
Footnotes
Author Contributions
B.F.G. designed experiments. B.F.G., J.G.P. and J.G. established the Ca2+ imaging protocol and performed experiments. B.F.G., P.E.J, A.L. and M.J.S. designed analyses. B.F.G. and J.D.M. analyzed data. J.A.L., L.J.K., J.D.M., and M.C.L. provided software code and advised on analyses. J.Z.L. constructed virus. F.G. and A.L. provided electrophysiological data. B.F.G. and M.J.S. wrote the paper. J.G., A.L. and all authors edited the paper. A.L. and M.J.S. supervised the research.
Competing financial interests. M.J.S. is a scientific co-founder of Inscopix, Inc., which produces the miniature fluorescence microscope used in this study.
References
- 1.Herry C, et al. Switching on and off fear by distinct neuronal circuits. Nature. 2008;454:600–606. doi: 10.1038/nature07166. [DOI] [PubMed] [Google Scholar]
- 2.Maren S, Quirk GJ. Neuronal signalling of fear memory. Nat Rev Neurosci. 2004;5:844–852. doi: 10.1038/nrn1535. [DOI] [PubMed] [Google Scholar]
- 3.Quirk GJ, Armony JL, LeDoux JE. Fear conditioning enhances different temporal components of tone-evoked spike trains in auditory cortex and lateral amygdala. Neuron. 1997;19:613–624. doi: 10.1016/s0896-6273(00)80375-x. [DOI] [PubMed] [Google Scholar]
- 4.Chowdhury N, Quinn JJ, Fanselow MS. Dorsal hippocampus involvement in trace fear conditioning with long, but not short, trace intervals in mice. Behav Neurosci. 2005;119:1396–1402. doi: 10.1037/0735-7044.119.5.1396. [DOI] [PubMed] [Google Scholar]
- 5.Quirk GJ, Repa C, LeDoux JE. Fear conditioning enhances short-latency auditory responses of lateral amygdala neurons: parallel recordings in the freely behaving rat. Neuron. 1995;15:1029–1039. doi: 10.1016/0896-6273(95)90092-6. [DOI] [PubMed] [Google Scholar]
- 6.Ghosh KK, et al. Miniaturized integration of a fluorescence microscope. Nat Methods. 2011;8:871–878. doi: 10.1038/nmeth.1694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ziv Y, et al. Long-term dynamics of CA1 hippocampal place codes. Nat Neurosci. 2013;16:264–266. doi: 10.1038/nn.3329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen TW, et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature. 2013;499:295–300. doi: 10.1038/nature12354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Garner AR, et al. Generation of a synthetic memory trace. Science. 2012;335:1513–1516. doi: 10.1126/science.1214985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gore F, et al. Neural Representations of Unconditioned Stimuli in Basolateral Amygdala Mediate Innate and Learned Responses. Cell. 2015;162:134–145. doi: 10.1016/j.cell.2015.06.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Namburi P, et al. A circuit mechanism for differentiating positive and negative associations. Nature. 2015;520:675–678. doi: 10.1038/nature14366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Senn V, et al. Long-range connectivity defines behavioral specificity of amygdala neurons. Neuron. 2014;81:428–437. doi: 10.1016/j.neuron.2013.11.006. [DOI] [PubMed] [Google Scholar]
- 13.Romanski LM, Clugnet MC, Bordi F, LeDoux JE. Somatosensory and auditory convergence in the lateral nucleus of the amygdala. Behav Neurosci. 1993;107:444–450. doi: 10.1037//0735-7044.107.3.444. [DOI] [PubMed] [Google Scholar]
- 14.Wolff SB, et al. Amygdala interneuron subtypes control fear learning through disinhibition. Nature. 2014;509:453–458. doi: 10.1038/nature13258. [DOI] [PubMed] [Google Scholar]
- 15.Blair HT, et al. Unilateral storage of fear memories by the amygdala. J Neurosci. 2005;25:4198–4205. doi: 10.1523/JNEUROSCI.0674-05.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ciocchi S, et al. Encoding of conditioned fear in central amygdala inhibitory circuits. Nature. 2010;468:277–282. doi: 10.1038/nature09559. [DOI] [PubMed] [Google Scholar]
- 17.Erlich JC, Bush DE, Ledoux JE. The role of the lateral amygdala in the retrieval and maintenance of fear-memories formed by repeated probabilistic reinforcement. Front Behav Neurosci. 2012;6:16. doi: 10.3389/fnbeh.2012.00016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kong E, Monje FJ, Hirsch J, Pollak DD. Learning not to fear: neural correlates of learned safety. Neuropsychopharmacology. 2014;39:515–527. doi: 10.1038/npp.2013.191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mankin EA, et al. Neuronal code for extended time in the hippocampus. Proc Natl Acad Sci U S A. 2012;109:19462–19467. doi: 10.1073/pnas.1214107109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Blair HT, Schafe GE, Bauer EP, Rodrigues SM, LeDoux JE. Synaptic plasticity in the lateral amygdala: a cellular hypothesis of fear conditioning. Learn Mem. 2001;8:229–242. doi: 10.1101/lm.30901. [DOI] [PubMed] [Google Scholar]
- 21.Bishop CM. Pattern Recognition and Machine Learning. Vol. 1. Springer; 2007. [Google Scholar]
- 22.Rodrigues SM, Schafe GE, LeDoux JE. Molecular mechanisms underlying emotional learning and memory in the lateral amygdala. Neuron. 2004;44:75–91. doi: 10.1016/j.neuron.2004.09.014. [DOI] [PubMed] [Google Scholar]
- 23.Sotres-Bayon F, Quirk GJ. Prefrontal control of fear: more than just extinction. Current opinion in neurobiology. 2010;20:231–235. doi: 10.1016/j.conb.2010.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Goosens KA, Hobin JA, Maren S. Auditory-evoked spike firing in the lateral amygdala and Pavlovian fear conditioning: mnemonic code or fear bias? Neuron. 2003;40:1013–1022. doi: 10.1016/s0896-6273(03)00728-1. [DOI] [PubMed] [Google Scholar]
- 25.Danielson NB, et al. Sublayer-Specific Coding Dynamics during Spatial Navigation and Learning in Hippocampal Area CA1. Neuron. 2016;91:652–665. doi: 10.1016/j.neuron.2016.06.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rubin A, Geva N, Sheintuch L, Ziv Y. Hippocampal ensemble dynamics timestamp events in long-term memory. eLife. 2015;4 doi: 10.7554/eLife.12247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Han JH, et al. Neuronal competition and selection during memory formation. Science. 2007;316:457–460. doi: 10.1126/science.1139438. [DOI] [PubMed] [Google Scholar]
- 28.Rashid AJ, et al. Competition between engrams influences fear memory formation and recall. Science. 2016;353:383–387. doi: 10.1126/science.aaf0594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rogerson T, et al. Molecular and Cellular Mechanisms for Trapping and Activating Emotional Memories. PLoS One. 2016;11:e0161655. doi: 10.1371/journal.pone.0161655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Paton JJ, Belova MA, Morrison SE, Salzman CD. The primate amygdala represents the positive and negative value of visual stimuli during learning. Nature. 2006;439:865–870. doi: 10.1038/nature04490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Belova MA, Paton JJ, Morrison SE, Salzman CD. Expectation modulates neural responses to pleasant and aversive stimuli in primate amygdala. Neuron. 2007;55:970–984. doi: 10.1016/j.neuron.2007.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Rumpel S, LeDoux J, Zador A, Malinow R. Postsynaptic receptor trafficking underlying a form of associative learning. Science. 2005;308:83–88. doi: 10.1126/science.1103944. [DOI] [PubMed] [Google Scholar]
- 33.Sah P, Westbrook RF, Luthi A. Fear conditioning and long-term potentiation in the amygdala: what really is the connection? Annals of the New York Academy of Sciences. 2008;1129:88–95. doi: 10.1196/annals.1417.020. [DOI] [PubMed] [Google Scholar]
- 34.Fadok JP, Dickerson TM, Palmiter RD. Dopamine is necessary for cue-dependent fear conditioning. J Neurosci. 2009;29:11089–11097. doi: 10.1523/JNEUROSCI.1616-09.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Johansen JP, et al. Hebbian and neuromodulatory mechanisms interact to trigger associative memory formation. Proc Natl Acad Sci U S A. 2014 doi: 10.1073/pnas.1421304111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Gutig R, Sompolinsky H. The tempotron: a neuron that learns spike timing-based decisions. Nat Neurosci. 2006;9:420–428. doi: 10.1038/nn1643. [DOI] [PubMed] [Google Scholar]
- 37.Fremaux N, Gerstner W. Neuromodulated Spike-Timing-Dependent Plasticity, and Theory of Three-Factor Learning Rules. Frontiers in neural circuits. 2015;9:85. doi: 10.3389/fncir.2015.00085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Pawlak V, Wickens JR, Kirkwood A, Kerr JN. Timing is not Everything: Neuromodulation Opens the STDP Gate. Frontiers in synaptic neuroscience. 2010;2:146. doi: 10.3389/fnsyn.2010.00146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hinton G. The ups and downs of Hebb synapses. PsycARTICLES. 2003;44(1):10–13. [Google Scholar]
- 40.Johansen JP, Tarpley JW, LeDoux JE, Blair HT. Neural substrates for expectation-modulated fear learning in the amygdala and periaqueductal gray. Nat Neurosci. 2010;13:979–986. doi: 10.1038/nn.2594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Courtin J, et al. Prefrontal parvalbumin interneurons shape neuronal activity to drive fear expression. Nature. 2014;505:92–96. doi: 10.1038/nature12755. [DOI] [PubMed] [Google Scholar]
- 42.Raybuck JD, Lattal KM. Double dissociation of amygdala and hippocampal contributions to trace and delay fear conditioning. PLoS One. 2011;6:e15982. doi: 10.1371/journal.pone.0015982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Thevenaz P, Ruttimann UE, Unser M. A pyramid approach to subpixel registration based on intensity. IEEE Trans Image Process. 1998;7:27–41. doi: 10.1109/83.650848. [DOI] [PubMed] [Google Scholar]
- 44.Mukamel EA, Nimmerjahn A, Schnitzer MJ. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron. 2009;63:747–760. doi: 10.1016/j.neuron.2009.08.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Betley JN, et al. Neurons for hunger and thirst transmit a negative-valence teaching signal. Nature. 2015;521:180–185. doi: 10.1038/nature14416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Rigotti M, et al. The importance of mixed selectivity in complex cognitive tasks. Nature. 2013;497:585–590. doi: 10.1038/nature12160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.George Paxinos KBJF. The Mouse Brain in Stereotaxic Coordinates. 2001;2 [Google Scholar]
- 48.Singec I, Knoth R, Ditter M, Volk B, Frotscher M. Neurogranin is expressed by principal cells but not interneurons in the rodent and monkey neocortex and hippocampus. The Journal of comparative neurology. 2004;479:30–42. doi: 10.1002/cne.20302. [DOI] [PubMed] [Google Scholar]
- 49.Maren S, Yap SA, Goosens KA. The amygdala is essential for the development of neuronal plasticity in the medial geniculate nucleus during auditory fear conditioning in rats. J Neurosci. 2001;21:RC135. doi: 10.1523/JNEUROSCI.21-06-j0001.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.