Abstract
Bacterial autotransporters are proteins that use a C-terminal porin-like domain to facilitate the transport of an upstream “passenger domain” across the outer membrane. Although autotransporters are translocated across the inner membrane (IM) via the Sec pathway, some of them contain exceptionally long signal peptides distinguished by a unique N-terminal sequence motif. In this study, we used the Escherichia coli O157:H7 autotransporter EspP as a model protein to investigate the function of the unusual signal peptides. We found that removal of the N-terminal motif or replacement of the EspP signal peptide did not affect translocation of the protein across the IM. Remarkably, modification of the signal peptide caused EspP to misfold in the periplasm and blocked transport of the passenger domain across the outer membrane. Further analysis suggested that the EspP signal peptide transits slowly through the Sec machinery. Based on these results, we propose that the unusual signal peptides not only function as targeting signals, but also prevent misfolding of the passenger domain in the periplasm by transiently tethering it to the IM.
Keywords: Escherichia coli, microbial pathogenesis, protein folding, protein translocation
The autotransporter or “type V” secretion pathway represents a unique mechanism of delivering proteins to the cell surface in Gram-negative bacteria. In this pathway, all of the information required to transport a polypeptide across the outer membrane (OM) is contained within the polypeptide itself (reviewed in ref. 1). Autotransporters are comprised of two domains, a large N-terminal “passenger” domain that frequently exceeds 100 kDa and a C-terminal ≈30-kDa “β domain.” Passenger domains encode a diverse array of effectors that are often virulence factors including adhesins, toxins, proteases, and lipases. The β domains are less well characterized, but they form barrel-like structures typical of OM proteins (2). Although the sequence of the β domain is highly variable, a consensus motif has been identified (3). Upon translocation across the inner membrane (IM), the β domain anchors the AT into the OM and facilitates transport of the passenger domain into the extracellular space by an unknown mechanism. The observation that tightly folded globular domains fused to autotransporters remained trapped in the periplasm (4, 5) strongly suggests that passenger domains are translocated across the OM in a loosely folded conformation. The passenger domains of some autotransporters are cleaved from the OM either by cellular proteases or by an autoproteolytic mechanism. It is generally assumed that the excision of passenger domains from the “pro” form of autotransporters occurs after their translocation across the OM.
Although recent reports have provided evidence that autotransporters are translocated across the IM via the classical Sec pathway (6, 7), some of them contain unusual signal peptides that exceed 50 aa in length. The C-terminal half closely resembles a typical signal peptide in that it contains canonical N, H, and C regions (8) but otherwise shows high sequence variability. This segment is distinguished only by an uncommonly high net positive charge in the N region. The N-terminal half of the long autotransporter signal peptides, however, contains a unique, highly conserved sequence motif (Fig. 7, which is published as supporting information on the PNAS web site) (9). There is no obvious correlation between the presence of this N-terminal extension and the effector function of the passenger domain, but it is found in the signal peptides of all of the serine protease autotransporters of Escherichia coli and Shigella (SPATEs). Interestingly, closely related N-terminal extensions are also found in the signal peptides of some proteins that are secreted via the so-called two partner secretion (TPS) pathway (10). Proteins secreted by this pathway use dedicated, coordinately expressed porin-like translocators to cross the OM. Despite very limited sequence similarity and an uncertain evolutionary relationship, the TPS exoproteins could be thought of as autotransporter passenger domains that are unlinked from their β domains.
Because signal peptides are highly degenerate, the conservation of the N-terminal extension found on autotransporter signal peptides suggests that it has an important function. It is conceivable that the N-terminal extension is required for either the targeting of autotransporters or their translocation across the IM. Indeed, in mammalian cells, IL-15 and plasminogen activator inhibitor-2 have exceptionally long signal peptides that influence the efficiency of their translocation into the endoplasmic reticulum (ER) (11, 12). Consistent with this hypothesis, a recent study presented evidence that a serine protease autotransporter of Escherichia coli and Shigella (SPATE) called Hbp is targeted to the IM cotranslationally by the signal recognition particle (7) and suggested that the unusual signal peptide might serve to prevent the protein from adopting a translocation-incompetent conformation in the cytoplasm. However, there is also evidence that long signal peptides have functions entirely unrelated to protein targeting. The IL-15 signal peptide appears to anchor the protein in the ER transiently (12). Remarkably, a long signal peptide associated with the Lassa virus glycoprotein was recently shown to act in trans to promote proteolytic processing of the protein after its transport into the ER (13).
In this study, we investigated the function of the unique autotransporter signal peptides in more detail by using EspP, a serine protease autotransporter of Escherichia coli and Shigella (SPATE) produced by E. coli O157:H7 (14, 15), as a model protein. We found that the N-terminal extension of the 55-aa EspP signal peptide was not required for efficient translocation of the protein across the IM. Surprisingly, however, deletion of this segment or replacement of the signal peptide led to mis-folding of EspP in the periplasm and severely impaired late stages of protein biogenesis. The observation that the EspP signal peptide appears to “jam” the Sec machinery provided a framework to rationalize our results. This finding raised the possibility that the signal peptide transiently anchors the passenger domain to the periplasmic side of the IM and thereby prevents it from folding into a conformation that is incompatible with transport across the OM. Our data provide evidence for a previously undescribed signal peptide function and also yield significant insights into the biogenesis of both autotransporters and proteins secreted via the two partner secretion pathway.
Experimental Procedures
Reagents, Media, and Bacterial Strains. Polyclonal rabbit antisera were generated against the peptides SQMDISNFYIRDYMDFAQNKGC and CEKSAFGKYNVDNAVNANFRYS, which were derived from the N and C termini of proEspP, respectively, after coupling to keyhole limpet hemocyanin. Other rabbit antisera were obtained from the following sources: maltose-binding protein (MBP) was from New England Biolabs, DsbA was from Stressgen, OmpA and β-lactamase were from P.C. Tai (Georgia State University, Atlanta), and OmpF was from Greg Phillips (Iowa State University, Ames). An anti-Ffh antiserum has been described (16). Unless otherwise noted, all experiments were conducted at 37°C in M9 medium containing 0.2% glycerol, and all of the l-amino acids except cysteine and methionine. Selective media contained 100 μg/ml ampicillin or 30 μg/ml chloramphenicol as required. The bacteria used in this study were the E. coli K-12 strains HDB114 (MC4100 ompT::kan ara+), CU164 (MC4100 secY39 zhd33::tet; ref. 17) and HDB58 (MC4100 zhd33::tet), and the E. coli O157:H7 strain EDL933 (American Type Culture Collection).
Plasmid Construction. Details of constructions are provided in full in Supporting Text, which is published as supporting information on the PNAS web site. In brief, espP was amplified by PCR using genomic DNA from EDL933 as a template and cloned into a version of pTRC99A (Amersham Pharmacia) that lacks an NdeI site. A derivative of this plasmid containing unique NdeI and EagI sites flanking the EspP signal peptide (pRLS6) was then constructed. Plasmids encoding espP with the EspP(C), Esp(N) and MBP signal peptides (see Fig. 1A) were designated pRLS7, pRLS8, and pRLS10, respectively, and were generated by cloning NdeI–EagI fragments harboring the appropriate signal peptide into pRLS6. A plasmid encoding an EspP passenger domain cleavage mutant (EspP*) was created by site-directed mutagenesis of pRLS6 and was designated pRLS11. Plasmid pRLS12 was generated by cloning an NdeI–EagI fragment harboring the EspP(C) signal peptide into pRLS11. To construct plasmid pJH62, the β domain of EspP plus the C-terminal 116 aa of the EspP passenger domain were amplified by PCR and cloned into the EagI and HindIII sites of pHL36 (18). NdeI–EagI fragments containing the EspP and EspP(C) SPs were cloned into pJH62 to generate pJH63 and pJH64, respectively. To construct plasmids pJH68, pJH69, and pJH70, wild-type MBP or MBP containing the EspP or EspP(C) signal peptide was first amplified by using pJH28, pJH46, or pJH48 (19, 20) as a template. The amplified DNA was then cloned into a modified version of pBAD33 (21).
Fig. 1.
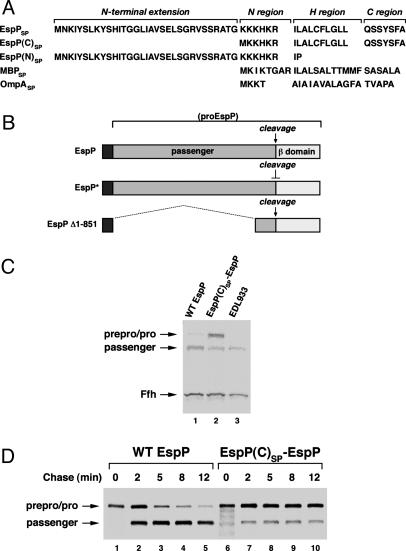
The native EspP signal peptide plays a key role in protein biogenesis. (A) The EspP signal peptide contains the characteristic N, H, and C regions found in typical signal peptides plus a long N-terminal extension. A sequence motif that is conserved in other autotransporter signal peptides is embedded within this extension (see Fig. 7). EspP(C)SP and EspP(N)SP are derivatives that contain only the C-terminal half of the signal peptide or only the N-terminal extension, respectively. (B) ProEspP is cleaved into discrete passenger and β domains at a late step in protein biogenesis. EspP* contains two point mutations that block this cleavage. Most of the passenger domain is deleted in EspP Δ1–851. (C) HDB114 transformed with a plasmid encoding wild-type EspP (pRLS6) or EspP(C)SP-EspP (pRLS7) under the control of the trc promoter and E. coli O157:H7 strain EDL933 were grown in LB. EspP biogenesis was analyzed by Western blot. The blot was probed with antisera raised against an N-terminal peptide of proEspP and Ffh. (D) HDB114 transformed with pRLS6 or pRLS7 were grown in M9 and subjected to pulse–chase labeling after the addition of 100 μM IPTG. EspP-containing polypeptides were immunoprecipitated with the N-terminal antipeptide antiserum.
Analysis of EspP Biogenesis at Steady State. Overnight cultures in LB were washed and diluted into fresh medium at an optical density (OD550) of 0.04. When the cultures reached OD = 0.8, aliquots were trichloroacetic acid (TCA) precipitated and proteins were analyzed by Western blot as described (16).
Kinetic Analysis of EspP Biogenesis and Protein Export. Overnight cultures were washed and diluted into fresh medium at an optical density (OD550) = 0.02. When the cultures reached OD550 = 0.2–0.3, synthesis of EspP- or MBP-containing polypeptides was induced by the addition of isopropylthiogalactoside (IPTG) at a concentration of 100 μM, except where noted, or 0.2% arabinose. Pulse–chase labeling was conducted as described (16) 30 min after the addition of IPTG or 20 min after the addition of arabinose. In experiments that did not involve any further manipulations, aliquots from each culture were immediately subjected to TCA precipitation. In all other experiments, radiolabeled cells were first poured over ice and all further steps were conducted at 0–4°C. To examine protein translocation across the IM, cells were collected by centrifugation at 3,800 × g for 8 min, converted to spheroplasts, and divided in half. One portion was then subjected to proteinase K treatment as described (16), and proteins in all samples were TCA precipitated. A similar protocol was used to examine translocation across the OM, except that 5 mM 2-mercaptoethanol was added to cells 5 min before labeling to prevent the formation of a disulphide bond that interfered with the proteolysis of proEspP. Cells were centrifuged at 3,000 × g for 5 min and resuspended in M9 lacking a carbon source before the addition of protease. In cell fractionation experiments, radiolabeled cells were centrifuged at 2,500 × g for 10 min and converted to spheroplasts as described above. After the addition of 10 mM MgSO4, a periplasmic fraction was obtained by pelleting the spheroplasts at 16,500 × g for 5 min. Spheroplasts were then resuspended in 80 μl of 33 mM Tris·HCl (pH 8.0)/40% sucrose and lysed by the addition of 3.2 ml of H2O. Unbroken spheroplasts were removed by centrifugation at 2,500 × g for 5 min. Subsequently, the lysate was centrifuged at 100,000 × g for 30 min to obtain cytoplasmic and membrane fractions. The membrane pellet was then separated into IM and OM fractions as described (22). Proteins in each fraction were collected by TCA precipitation. In membrane floatation experiments, the 100,000 × g pellets were resuspended in 100 μl of buffer A (3 mM EDTA, pH 7.3/1 mM PMSF/0.25 M sucrose), mixed with 400 μl of buffer A containing 2.3 M sucrose, placed in 11 × 34-mm tubes, and overlaid with 0.5 ml of buffer A containing 1.7 M sucrose. Tubes were filled to the top with buffer A and centrifuged in a Beckman TLS 55 rotor for 4 h at 54,000 rpm. Cell membranes were then collected at the interface between the 1.7 M and 0.25 M sucrose layers. To assess the heat modifiability of EspP, cells were centrifuged at 2,500 × g for 10 min, resuspended in PBS containing 1 mM PMSF, and disrupted by sonication. After the addition of 1% Elugent (Calbiochem), detergent extracts were centrifuged at 20,800 × g for 10 min to remove cell debris.
In most experiments, TCA-precipitated proteins were solubilized and immunoprecipitatations were performed as described (16). In experiments in which the heat modifiability of EspP was examined, native immunoprecipitations were conducted by adding antisera directly to detergent extracts. Proteins were resolved by SDS/PAGE on 8–16% minigels (Novex) except where noted.
Results
The Native EspP Signal Peptide Is Not Required for Translocation Across the IM. Initially we examined the effect of deleting the N-terminal extension of the EspP signal peptide on protein biogenesis by monitoring the conversion of proEspP (Fig. 1B) into discrete passenger and β domain fragments. An E. coli K-12 strain (HDB114) was transformed with a plasmid encoding either wild-type espP or espP containing a signal peptide that lacks the N-terminal extension [“EspP(C)SP-EspP”; Fig. 1 A] under the control of the trc promoter (pRLS6 or pRLS7, respectively). Both signal peptides target EspP and heterologous proteins to the Sec pathway (ref. 20 and unpublished data). To examine the fate of EspP produced at a basal level, cells were grown in LB without IPTG, and portions of each culture were subjected to TCA precipitation. Western blots were then probed with an antipeptide antiserum raised against the N terminus of the passenger domain. This antiserum detects the free passenger domain (≈105 kDa), proEspP (≈135 kDa), and preproEspP (≈140 kDa), the form of the protein that contains a signal peptide. Blots were also probed with an antiserum raised against a constitutively expressed protein (Ffh) to provide an internal standard. Cells containing pRLS6 and pRLS7 synthesized roughly the same amount of EspP as the E. coli O157:H7 strain EDL933 grown under identical conditions (Fig. 1C). Whereas the passenger domain of wild-type EspP was efficiently excised from proEspP, only about a third of EspP(C)SP-EspP was properly processed (Fig. 1C, lanes 1 and 2). The data suggested that truncation of the signal peptide either delays or blocks EspP biogenesis. To distinguish between these two possibilities, cells were grown in M9 and subjected to pulse–chase labeling after the addition of 100 μM IPTG, and EspP was immunoprecipitated from aliquots of each culture with the N-terminal antipeptide antiserum. Only ≈30% of the passenger domain of EspP(C)SP-EspP was cleaved within the first 2 min after synthesis, and no subsequent processing was observed (Fig. 1D, lanes 6–10). These data show that truncation of the EspP signal peptide imposes a complete block on the secretion of a large fraction of the protein.
To our surprise, further analysis showed that truncation or replacement of the EspP signal peptide did not impair translocation of the protein across the IM. Because we could not resolve the prepro and pro forms of EspP containing short signal peptides by SDS/PAGE, we used a protease protection assay to examine the effect of altering the signal peptide on transport across the IM. HDB114 was transformed with pRLS6, pRLS7, or a plasmid encoding espP with either the MBP signal peptide [“MBPSP-EspP”; Fig. 1 A] or only the N-terminal extension of the native signal peptide [“EspP(N)SP-EspP”; Fig. 1 A]. Cells were subjected to pulse–chase labeling as described above, collected by centrifugation, and converted to spheroplasts. Proteinase K was then added to half of the spheroplast suspension (which included the contents of the periplasm), and EspP-containing polypeptides were immunoprecipitated from treated and untreated samples as well as from the growth medium with the N-terminal antipeptide antiserum. Consistent with the expectation that wild-type EspP is translocated efficiently across the IM, almost all of the ≈135-kDa form of the protein observed at each time point was protease-sensitive (Fig. 2A Top). Interestingly, the pulse-labeled ≈135-kDa form of EspP(C)SP-EspP was also nearly completely protease-sensitive (Fig. 2 A Upper Middle). This result showed that the signal peptide mutant was rapidly translocated across the IM. The observation that the cytoplasmic Ffh protein was protease-resistant (unless spheroplasts were lysed with detergent) confirmed that the spheroplasts remained intact during the experiment (Fig. 2B). MBPSP-EspP was likewise translocated across the IM rapidly despite the fact that replacement of the native signal peptide caused a signficant defect in protein biogenesis (Fig. 2 A Lower Middle). By contrast, EspP(N)SP-EspP was retained in the cytoplasm and was completely resistant to proteolysis (Fig. 2 A Bottom). This observation implies that the N-terminal extension is not an independent targeting signal.
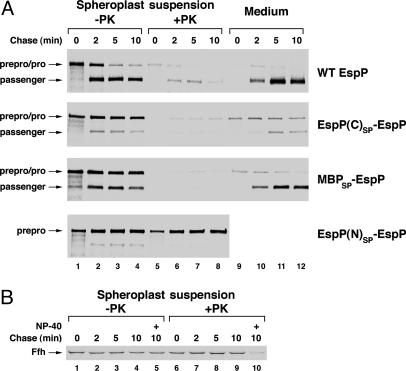
Fig. 2.
Modification of the EspP signal peptide does not inhibit translocation across the IM. (A) HDB114 transformed with pRLS6, pRLS7, or a plasmid encoding MBPSP-EspP or EspP(N)SP-EspP (pRLS10 or pRLS8, respectively) were subjected to pulse–chase labeling. Cells were pelleted and converted to spheroplasts, and proteinase K was added to half of each spheroplast suspension. EspP-containing polypeptides were then immunoprecipitated with an N-terminal antipeptide antiserum. (B) Ffh was immunoprecipitated from the samples used in A Upper Middle. Nonidet P-40 (0.1%) was added to the spheroplast suspension before proteinase K digestion as indicated. -PK, no protease added; +PK, proteinase K added; Medium, protein recovered in culture medium.
Truncation of the Native EspP Signal Peptide Affects Late Steps of Protein Biogenesis. To test the possibility that truncation of the signal peptide impairs integration of EspP into the OM, we next compared the localization of EspP(C)SP-EspP to the wild-type protein. Because we wanted to focus on the trafficking of proEspP and not its subsequent cleavage, we used a noncleavable version of the protein in which the two Asn residues at the junction of the passenger and β domains were mutated to Ser (“EspP*”; Fig. 1B). Cells transformed with a pRLS6 derivative encoding wild-type espP* or espP* with the EspP(C) signal peptide [“EspP(C)SP-EspP*”] (pRLS11 or pRLS12, respectively) were subjected to pulse–chase labeling as described above and fractionated. EspP-containing polypeptides were then immunoprecipitated with an antipeptide antiserum that recognizes the C terminus of the protein. Most of the EspP(C)SP-EspP* localized to the OM within 1 min, although ≈10% of the protein remained in the periplasm even after a 5-min chase (Fig. 3A, lanes 6–10). The observation that most of the EspP(C)SP-EspP* cofractionated with cell membranes when they were isolated by floatation instead of sedimentation (Fig. 3B) confirmed that the protein did not fortuitously pellet with OM vesicles. Overall, the data suggest that truncation of the signal peptide does not seriously impair the intracellular localization of EspP.
Fig. 3.
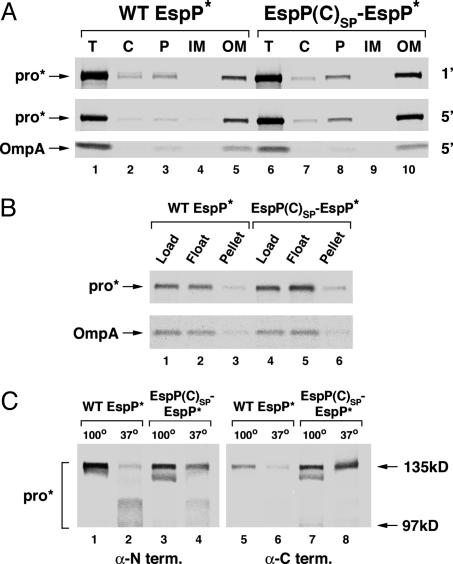
Truncation of the signal peptide causes incomplete folding of EspP in the OM. HDB114 were transformed with a plasmid encoding wild-type EspP* (pRLS11) or EspP(C)SP-EspP* (pRLS12). (A) Cells were pulse-labeled and subjected to a 1-min (1′) or 5-min (5′) chase. Cell extracts were fractionated, and preproEspP* and proEspP* were immunoprecipitated by using a C-terminal antipeptide antiserum. OmpA was also immunoprecipitated as a control. Lanes 1 and 6, total lysate (T); lanes 2 and 7, cytoplasm (C); lanes 3 and 8, periplasm (P); lanes 4 and 9, inner membranes (IM); lanes 5 and 10, outer membranes (OM). (B) Cells were pulse-labeled and subjected to a 5-min chase, and a high-speed pellet was obtained from spheroplast lysates. This pellet was resuspended, and membranes were isolated by floatation on a discontinuous sucrose gradient. Immunoprecipitations were performed as in A. Lanes 1 and 4, original high-speed pellet (Load); lanes 2 and 5, membrane fraction (Float); lanes 3 and 6, sucrose gradient pellet (Pellet). (C) Cells were pulse-labeled, subjected to a 5-min chase, and lysed in the presence of detergent. ProEspP* was isolated by native immunoprecipitation using N-and C-terminal antipeptide antisera and resolved by SDS/PAGE on 6% gels after heating to 100°C (lanes 1, 3, 5, and 7) or 37°C (lanes 2, 4, 6, and 8).
Although the trafficking of EspP containing the EspP(C) signal peptide seemed relatively normal, further analysis revealed that the protein does not fold into a native structure in the OM. Previous studies have shown that, upon integration into the OM, the β-barrel domains of many bacterial OM proteins adopt a tightly folded conformation that remains stable in SDS even after the protein is heated to moderate temperatures. The compact folding of the β-barrel causes the proteins to migrate rapidly on SDS/PAGE (23). To determine whether the β domains of wild-type EspP* and EspP(C)SP-EspP* undergo the same type of conformational change as other OM proteins, cells transformed with pRLS11 or pRLS12 were pulse-labeled as described above and subjected to a 5-min chase. Cell lysates were then prepared in the presence of a nonionic detergent, and proEspP* was isolated by native immunoprecipitation. When wild-type proEspP* was immunoprecipitated with the N-terminal antiserum and then boiled, the protein was completely denatured and migrated at ≈135 kDa (Fig. 3C, lane 1). Interestingly, most of the protein migrated as a diffuse band of ≈100–115 kDa if it was heated to only 37°C before SDS/PAGE (Fig. 3C, lane 2). The simplest explanation of this result is that the β domain folds compactly, but conformational variation in the passenger domain causes the protein to migrate heterogeneously. Curiously, very little wild-type proEspP* was immunoprecipitated with the C-terminal antiserum (Fig. 3C, lanes 5 and 6). The data suggest that the extreme C terminus is inaccessible when the protein folds into a native conformation. By contrast, most of the signal peptide mutant migrated at ≈135 kDa after it was heated to only 37°C, and it was recognized effectively by both antisera (Fig. 3C, lanes 3–4 and 7–8). Both the gel mobility and antibody accessibility data strongly suggest that truncation of the EspP signal peptide impairs the folding of the protein after translocation across the IM and prevents the β domain from adopting a compact conformation in the OM.
Consistent with the observation that the native EspP signal peptide is required for complete folding of the β domain, we found that truncation of the signal peptide strongly inhibits the translocation of the passenger domain across the OM. We monitored the sensitivity of the EspP* passenger domain to proteolysis in intact cells to assess the effectiveness of translocation. HDB114 transformed with pRLS11 or pRLS12 were radiolabeled as described above and divided in half. Proteinase K was then added to one of the two aliquots, and proEspP* was immunoprecipitated with the C-terminal antiserum. Most of the wild-type EspP* (but not the periplasmic protein DsbA) was accessible to the exogenous protease after a 2-min chase (Fig. 4A, lanes 1–6). By contrast, almost all of the EspP(C)SP-EspP* was resistant to proteolysis even after a 5-min chase (Fig. 4A, lanes 7–12). In addition, the synthesis of DsbA was elevated in cells that produced the signal peptide mutant, most likely as a result of periplasmic stress. These results strongly suggest that, whereas the passenger domain of wild-type EspP is efficiently translocated across the OM, the passenger domain of most of the signal peptide mutant remains trapped in the periplasm. To obtain evidence that the impairment of passenger domain cleavage observed in Figs. 1 and 2 was a direct consequence of the passenger domain translocation defect, we repeated the above experiment using cells that produced the cleavable version of EspP containing the Esp(C) signal peptide. EspP-containing polypeptides were immunoprecipitated with the N-terminal antiserum. As expected, most of the free passenger domain was degraded by proteinase K, but all of the proEspP was resistant to digestion (Fig. 4B).
Fig. 4.
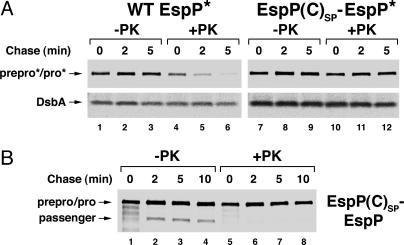
Truncation of the signal peptide inhibits translocation of the EspP passenger domain across the OM. HDB114 transformed with pRLS11 or pRLS12 (A) or pRLS7 (B) were subjected to pulse–chase labeling. Cells were pelleted, and proteinase K was added to half of each sample. EspP-containing polypeptides were immunoprecipitated by using C-terminal (A) or N-terminal (B) antipeptide antisera. DsbA was immunoprecipitated from the samples in A as a control. -PK, no protease added; +PK, proteinase K added.
We next wished to determine whether the protein folding and translocation defects that we observed are a consequence of the large size or unique structural features of the EspP passenger domain. To this end, we constructed a deletion mutant that contains only the last 116 aa of the passenger domain (“EspP Δ1–851”; Fig. 1B). We then examined the effect of replacing the EspP signal peptide with either the EspP(C) or OmpA signal peptide on protein biogenesis. Cells transformed with a plasmid containing a derivative of EspP Δ1–851 under the control of the trc promoter were subjected to pulse–chase labeling after the addition of 10 μM IPTG. At this concentration of inducer EspP Δ1–851 was synthesized at a higher level than full-length EspP in the presence of 100 μM IPTG (data not shown). EspP-containing polypeptides were immunoprecipitated with the C-terminal antiserum, and secretion was assessed by monitoring the accumulation of the cleaved β domain. When we used this assay, we found that alteration of the signal peptide had no significant effect on either the rate or efficiency of secretion (Fig. 5). These observations imply that the long signal peptide is required specifically to prevent the full length passenger domain from adopting a conformation that is incompatible with its translocation across the OM.
Fig. 5.
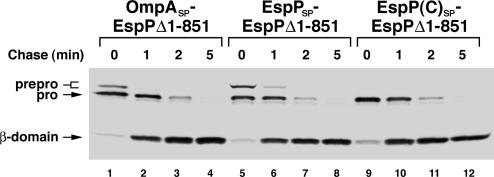
Alteration of the signal peptide does not affect the biogenesis of an EspP deletion mutant. HDB114 transformed with a plasmid encoding EspP Δ1–851 with an OmpA, EspP or EspP(C) signal peptide (pJH61, pJH62, or pJH63, respectively) were subjected to pulse–chase labeling. EspP-containing polypeptides were immunoprecipitated with a C-terminal antipeptide antiserum.
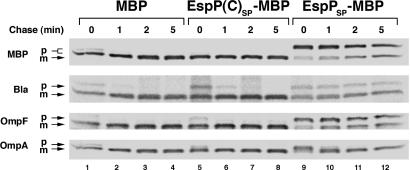
The EspP Signal Peptide Appears to Dissociate Slowly from the Sec Machinery. To explain the apparently paradoxical observation that a targeting signal affects protein folding in the periplasm, we conjectured that the EspP signal peptide optimizes events that occur during or shortly after translocation but before its removal by leader peptidase. Support for this hypothesis emerged from experiments that were designed to test an alternative explanation of the results. To determine whether the signal peptide acts in trans after its cleavage by leader peptidase, we examined the effect of overproducing a heterologous protein (MBP) that harbors the native EspP signal peptide on the biogenesis of EspP(C)SP-EspP. Overproduction of the MBP derivative (“EspPSP-MBP”) did not suppress the biogenesis defect associated with truncation of the EspP signal peptide (data not shown) but instead nonspecifically impaired protein translocation across the IM. This phenomenon is illustrated in an experiment in which 0.2% arabinose was added to HDB114 transformed with a plasmid that encodes wild-type MBP, MBP containing the EspP(C) signal peptide (“EspP(C)SP-MBP”) or EspPSP-MBP under the control of the araBAD promoter. Overproduction of EspPSP-MBP (but not the other versions of MBP) inhibited its own translocation across the IM as well as the translocation of several other proteins (Fig. 6). It is unlikely that the transport defect resulted from competition for a limiting targeting factor because the proteins examined are targeted by different pathways. One simple explanation of the results is that the EspP signal peptide impeded the access of presecretory proteins to the Sec complex by transiting through the translocation machinery (and presumably becoming accessible to leader peptidase) relatively slowly. By implying that the EspP signal peptide acts as a temporary membrane anchor, this interpretation provides a plausible framework for explaining the role of the signal peptide in late stages of protein biogenesis. Consistent with the idea that the EspP signal peptide transiently “jams” the Sec complex, we found that even moderate overproduction of EspPSP-MBP in a strain harboring the secY39(Cs) allele was lethal (Fig. 8, which is published as supporting information on the PNAS web site). The synthetic lethal effect suggests that EspPSP-MBP exacerbates the translocation defect that results from the secY mutation.
Fig. 6.
The EspP signal peptide appears to “jam” the Sec machinery. HDB114 transformed with a plasmid encoding MBP, EspP(C)SP-MBP or EspPSP-MBP (pJH68, pJH69, or pJH70, respectively) and the cloning vector pHDB3 (ampR; ref. 16) were subjected to pulse–chase labeling after the addition of 0.2% arabinose to induce synthesis of the MBP derivative, and the indicated presecretory protein was immunoprecipitated. p, precursor; m, mature.
Discussion
In this study we discovered that an exceptionally long signal peptide associated with a bacterial autotransporter plays an unprecedented role in protein biogenesis. We found that the C-terminal half of the EspP signal peptide (which closely resembles a classical signal peptide) as well as the MBP signal peptide promote effective translocation of EspP across the IM. Curiously, the EspP(C) signal peptide acts as a cotranslational targeting signal whereas both the MBP and native EspP signal peptides route proteins into posttranslational targeting pathways (ref. 20 and unpublished data). Because there is no clear correlation between effective translocation of EspP across the IM and the mechanism of targeting, it seems unlikely that the unusual autotransporter signal peptides have been conserved to prevent aggregation in the cytoplasm. Consistent with this view, we found that alteration of the EspP signal peptide inhibited proper folding of the β domain and effective translocation of the passenger domain across the OM. The results imply that the unusual EspP signal peptide plays an important role in steps in protein biogenesis that occur after it performs its targeting function. Although our experiments do not show whether altered folding of the β domain causes the passenger domain translocation defect or vice versa, they do indicate that the EspP signal peptide offsets a tendency of the passenger domain to fold nonproductively in the periplasm and to consequently become trapped. In addition, experiments using a truncated version of EspP (EspP Δ1–851) indicate that the potential for misfolding in the periplasm is due to unique structural features of the full-length passenger domain.
How might the EspP signal peptide influence the fate of the passenger domain after it has been transported across the IM? To explain this apparent paradox, we propose that the signal peptide acts as a transient membrane anchor that prevents the passenger domain from misfolding in the periplasm. Our model is based largely on results which suggest that the EspP signal peptide remains in proximity to the Sec complex longer than a typical signal peptide. Most notably, we found that overproduction of a heterologous protein containing the EspP signal peptide caused a nonspecific inhibition of protein translocation across the IM that is highly reminiscent of the “jamming” phenomenon observed in previous studies in which the Sec complex appeared to be occluded (24, 25). During a prolonged association with the translocation channel, the EspP signal peptide would presumably remain inaccessible to leader peptidase. Because the EspP signal peptide does not appear to be cleaved especially slowly in pulse–chase studies (unpublished data); however, any delay in its release from the Sec complex is probably brief. Nevertheless, proteins fold on a millisecond time scale, and even a very transient tethering of the N terminus of EspP to the IM during and possibly after translocation might prevent the formation of inappropriate intramolecular or inter-molecular contacts or prevent the passenger domain from folding compactly. Indeed attachment of MBP to the IM by replacing its signal peptide with a transmembrane segment prevents the protein from attaining a native folded state in the periplasm (19). Attachment of the EspP passenger domain to the IM might preserve its translocation-competence simply by facilitating the binding of periplasmic chaperones before misfolding can occur. Consistent with this possibility, we found that the magnitude of the secretion defect associated with truncation of the EspP signal peptide is influenced by both growth conditions and the level of EspP synthesis (unpublished data). The data suggest that the native EspP signal peptide offsets a potential imbalance in the ratio of newly synthesized autotransporter molecules to a limiting periplasmic chaperone.
Although there is growing evidence that at least some signal peptides contain significant functional information, signal peptides that act in cis to promote effective protein biogenesis at a posttranslocation step have not been previously reported. Nevertheless, there is precedent in the literature for many of the features that our model attributes to the EspP signal peptide. A few very long (or very basic) signal peptides have been described in mammalian cells that are cleaved slowly and thereby act as transient membrane anchors. By retaining proteins in the ER, these signal peptides do not influence protein folding, but instead appear to control the amount of protein that reaches its physiological site of action (12, 26). In addition, an unusual signal peptide sequence motif has been shown to affect the efficiency of precursor processing in Staphylococcus aureus (27). Finally, a recent study showed that variation in signal peptide sequence alters the ratio of three distinct topological forms of the prion protein (28). In this study, significant differences in the ability of individual signal peptides to gate open the translocation machinery appeared to dramatically affect the biogenesis of the protein. These results are particularly intriguing because they challenge the long-standing assumption that all signal peptides interact with the Sec machinery in a stereotyped fashion.
Supplementary Material
Acknowledgments
We thank Manu Hegde for helpful discussions and for critical reading of the manuscript.
Author contributions: R.L.S. and H.D.B. designed research; R.L.S., J.H.P., and H.D.B. performed research; K.M.S. provided new reagents/analytic tools; R.L.S., J.H.P., and H.D.B. analyzed data; and H.D.B. wrote the paper.
This paper was submitted directly (Track II) to the PNAS office.
Abbreviations: OM, outer membrane; IM, inner membrane; ER, endoplasmic reticulum; MBP, maltose-binding protein; TCA, trichloroacetic acid; IPTG, isopropylthiogalactoside.
References
- 1.Desvaux, M., Parham, N. J. & Henderson, I. R. (2004) Res. Microbiol. 155, 53-60. [DOI] [PubMed] [Google Scholar]
- 2.Oomen, C. J., van Ulsen, P., Van Gelder, P., Feijen, M., Tommassen, J. & Gros, P. (2004) EMBO J. 23, 1257-1266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Loveless, B. J. & Saier, M. H., Jr. (1997) Mol. Memb. Biol. 14, 113-123. [DOI] [PubMed] [Google Scholar]
- 4.Klauser, T., Pohlner, J. & Meyer, T. F. (1990) EMBO J. 9, 1991-1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Veiga, E., de Lorenzo, V. & Fernandez, L. A. (1999) Mol. Microbiol. 33, 1232-1243. [DOI] [PubMed] [Google Scholar]
- 6.Brandon, L. D., Goehring, N., Janakiraman, A., Yan, A. W., Wu, T., Beckwith, J. & Goldberg, M. B. (2003) Mol. Microbiol. 50, 45-60. [DOI] [PubMed] [Google Scholar]
- 7.Sijbrandi, R., Urbanus, M. L., ten Hagen-Jongman, C. M., Bernstein, H. D., Oudega, B., Otto, B. & Luirink, J. (2003) J. Biol. Chem. 278, 4654-4659. [DOI] [PubMed] [Google Scholar]
- 8.von Heijne, G. (1985) J. Mol. Biol. 184, 99-105. [DOI] [PubMed] [Google Scholar]
- 9.Henderson, I. R., Navarro-Garcia, F. & Nataro, J. P. (1998) Trends Microbiol. 6, 370-378. [DOI] [PubMed] [Google Scholar]
- 10.Jacob-Dubuisson, F., Locht, C. & Antoine, R. (2001) Mol. Microbiol. 40, 306-313. [DOI] [PubMed] [Google Scholar]
- 11.Belin, D., Bost, S., Vassalli, J.-D. & Strub, K. (1996) EMBO J. 15, 469-478. [PMC free article] [PubMed] [Google Scholar]
- 12.Kurys, G., Tagaya, Y., Bamford, R., Hanover, J. A. & Waldmann, T. A. (2000) J. Biol. Chem. 275, 30653-30659. [DOI] [PubMed] [Google Scholar]
- 13.Eichler, R., Lenz, O., Strecker, T., Eickmann, M., Klenk, H.-D. & Garten, W. (2003) EMBO Rep. 4, 1084-1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Brunder, W., Schmidt, H. & Karch, H. (1997) Mol. Microbiol. 24, 767-778. [DOI] [PubMed] [Google Scholar]
- 15.Djafari, S., Ebel, F., Deibel, C., Krämer, S., Hudel, M. & Chakraborty, T. (1997) Mol. Microbiol. 25, 771-784. [DOI] [PubMed] [Google Scholar]
- 16.Ulbrandt, N. D., Newitt, J. A. & Bernstein, H. D. (1997) Cell 88, 187-196. [DOI] [PubMed] [Google Scholar]
- 17.Baba, T., Jacq, A., Brickman, E., Beckwith, J., Taura, T., Ueguchi, C., Akiyama, Y. & Ito, K. (1990) J. Bacteriol. 172, 7005-7010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lee, H. C. & Bernstein, H. D. (2002) J. Biol. Chem. 277, 43527-43535. [DOI] [PubMed] [Google Scholar]
- 19.Lee, H. C. & Bernstein, H. D. (2001) Proc. Natl. Acad. Sci. USA 98, 3471-3476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Peterson, J. H., Woolhead, C. A. & Bernstein, H. D. (2003) J. Biol. Chem. 278, 46155-46162. [DOI] [PubMed] [Google Scholar]
- 21.Guzman, L.-M., Belin, D., Carson, M. J. & Beckwith, J. (1995) J. Bacteriol. 177, 4121-4130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Qi, H.-Y. & Bernstein, H. D. (1999) J. Biol. Chem. 274, 8993-8997. [DOI] [PubMed] [Google Scholar]
- 23.Freudl, R., Schwarz, H., Stierhof, Y.-D., Gamon, K., Hindennach, I. & Henning, U. (1986) J. Biol. Chem. 261, 11355-11361. [PubMed] [Google Scholar]
- 24.Snyder, W. B. & Silhavy, T. J. (1995) J. Bacteriol. 177, 953-963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Samuelson, J. C., Jiang, F., Yi, L., Chen, M., de Gier, J. W., Kuhn, A. & Dalbey, R. E. (2001) J. Biol. Chem. 276, 34847-34852. [DOI] [PubMed] [Google Scholar]
- 26.Li, Y., Luo, L., Thomas, D. Y. & Kang, C. Y. (1994) Virology 204, 266-278. [DOI] [PubMed] [Google Scholar]
- 27.Bae, T. & Schneeewind, O. (2003) J. Bacteriol. 185, 2910-2919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kim, S. J., Mitra, D., Salerno, J. R. & Hegde, R. S. (2002) Dev. Cell 2, 207-217. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.