Abstract
Aldehyde dehydrogenase 1 (ALDH1A1)–positive dopaminergic (DA) neurons at the ventral substantia nigra pars compacta (SNpc) preferentially degenerate in Parkinson’s disease (PD). Their projection pattern and dopamine release properties, however, remains uncharacterized. Here we show that ALDH1A1–positive axons project predominantly to the rostral two–thirds of dorsal striatum. A portion of these axons converge on a small fraction of striosome compartments restricted to the dorsolateral striatum (DLS), where less dopamine release was measured compared to the adjacent matrix enriched with the ALDH1A1–negative axons. Genetic ablation of Aldh1a1 substantially increases the dopamine release in striosomes, but not in matrix. Additionally, the presence of PD-related human α-synuclein A53T mutant or dopamine transporter (DAT) blockers also differentially affects the dopamine output in striosomes and matrix. Together, these results demonstrate distinct dopamine release characteristics of ALDH1A1–positive DA fibers, supporting a regional specific function of ALDH1A1 in regulating dopamine availability/release in striatum.
Introduction
ALDH1A1 belongs to a large family of aldehyde dehydrogenases that oxidize a variety of reactive aldehyde species1. ALDH1A1 is predominantly expressed by a subgroup of DA neurons in the midbrain2. In DA neurons, ALDH1A1 mediates the oxidation of the cytotoxic dopamine intermediate, 3, 4-dihydroxyphenylacetaldehyde (DOPAL), to the less reactive 3, 4-dihydroxyphenylacetic acid (DOPAC)3, 4. ALDH1A1 was also involved in the synthesis of retinoic acid (RA), which plays a crucial role in neuronal patterning, differentiation and survival5. Most recently, ALDH1A1 is reported to mediate the synthesis of the inhibitory neurotransmitter γ-aminobutyric acid (GABA) in DA neurons6. Intriguingly, two subpopulations of DA neurons at SNpc have been segregated along the dorsal to ventral axis for their ability to express ALDH1A17. DA neurons located in the dorsomedial SNpc lack ALHD1A1 expression, whereas DA neurons in the ventrolateral tier express this enzyme. We recently demonstrated that ALDH1A1–positive DA neurons are more susceptible to degeneration in PD8. However, little is known about the connectivity and functionality of this group of neurons. It is unclear whether the axons of these ALDH1A1–expressing DA neurons terminate at distinct locations at the dorsal striatum and exhibit different dopamine release dynamics.
Previous neuron tracing studies suggest that DA neurons at the dorsal tier of SNpc project to the matrix compartment of dorsal striatum, whereas neurons at the ventral tier send their axons to the complementary striosome or patch compartment9. Striosome and matrix compartments are distinguished by expression of different biochemical markers, their embryonic development, and afferent and efferent connectivity10, 11. Striosomes are characterized by enhanced expression of µ opiate receptor (MOR1)12, enkephalin13, and dopamine transporter DAT14. In contrast, matrix is rich in acetylcholinesterase12 and calbindin15. Three different types of axon fibers have been described within the nigrostriatal projections14. Type A fibers arise from the dorsal tier of SNpc and target mainly to the matrix compartment of dorsal striatum. Type B fibers are from the ventral tier DA neurons and terminate predominantly in the striosomes. Considering ALDH1A1–positive DA neurons are mainly distributed at the ventral SNpc, it is reasonable to assume that they project mainly to the striosomes. However, the precise projection pattern remains to be determined. Dopamine release also appears to be different in the striosome and matrix compartments of dorsal striatum16. As we demonstrated recently, dopamine release at striosomes is lower compared to the matrix16, 17. However, more study is required to understand how the transmitter release is differentially regulated.
The goal of the present study aims to determine the projection pattern and dopamine release properties of ALDH1A1–positive DA axons at the dorsal striatum. Since overexpression of PD-related α-synuclein A53T missense mutation substantially impairs dopamine release at the striatum18, we additionally examined whether DA release at ALDH1A1–positive and –negative axon terminals are differentially affected by mutant α-synuclein.
Results
ALDH1A1–positive DA axon fibers project unevenly to the dorsal striatum
Previous neuron tracing studies suggest that DA neurons at the ventral tier of SNpc send their axons to the striosome compartment of dorsal striatum9. However, there is a lack of molecular markers to reliably identify these neurons and their projections. Since ALDH1A1 is predominantly expressed by the ventral SNpc DA neurons in the brain2, 8, 19, 20, we stained sequential coronal sections of 1–month–old wild-type C67BL6 mice to examine the projection pattern of ALDH1A1–positive SNpc axon fibers at the striatum using antibodies against ALDH1A1, the striosomal marker MOR1, and the DA axon terminal marker tyrosine hydroxylase (TH). We found that ALDH1A1–positive DA fibers project unevenly along the rostrocaudal, dorsoventral, and mediolateral axis (Fig. 1a). In the rostral one–third (Bregma: ~2 to 1 mm), ALDH1A1–positive DA fibers mainly terminated at the medial part of nucleus accumbens shell (open arrowheads, top row), and the dorsal half of dorsal striatum. More condensed fibers were observed at the superficial layers of dorsal striatum, whereas no apparent segregation to the striosome or matrix compartments was found. In the middle one–third (Bregma: ~1 to 0 mm), ALDH1A1–positive DA fibers predominantly projected to the most dorsal layers of dorsal striatum. In the dorsomedial striatum (DMS), projections onto striosomes and matrix were not clearly segregated (Fig. 1a,b). By contrast, in the dorsolateral striatum (DLS), ALDH1A1–positive DA fibers largely converged to the most dorsal striosomes (arrowheads) and subcallosal streaks (arrows) (Fig. 1b). Finally, in the caudal one–third of striatum (Bregma: ~0 to –1.2 mm), there were fewer ALDH1A1–positive DA fibers projections, where they were present mainly in the superficial layers of DLS (Fig. 1a). Noticeably, there were a few clusters of ALDH1A1–positive fiber like structures in the thalamus, but they were negative for TH staining (arrows, top row, Fig. 1a). Furthermore, the projections only converge at the DLS striosomes and subcallosal streaks in the middle segment of dorsal striatum. The same projection pattern of ALDH1A1–positive DA fibers in the striatum was also observed in 3-month-old mice (Supplementary Fig. S1). To further specify the origin of ALDH1A1–positive signals in the striatum, we injected Aldh1a1 siRNA plus green florescent protein (GFP)–expressing recombinant adeno-associated viruses6 in the SNpc in one hemisphere of 3-month-old wild type mice (Supplementary Fig. S2a). In line with the previous publication6, this AAV vector effectively inhibited ALDH1A1 expression in the SNpc (Supplementary Fig. S2a). More importantly, the ALDH1A1 staining was substantially suppressed in the ipsilateral dorsal striatum marked by GFP signals (Supplementary Fig. S2b). Taken together, these results demonstrate an uneven projection pattern of ALDH1A1–positive DA fibers in striatum, in which the innervation appears to follow a decreased gradient along the rostrocaudal and dorsoventral axis (Fig. 1c).
Figure 1.
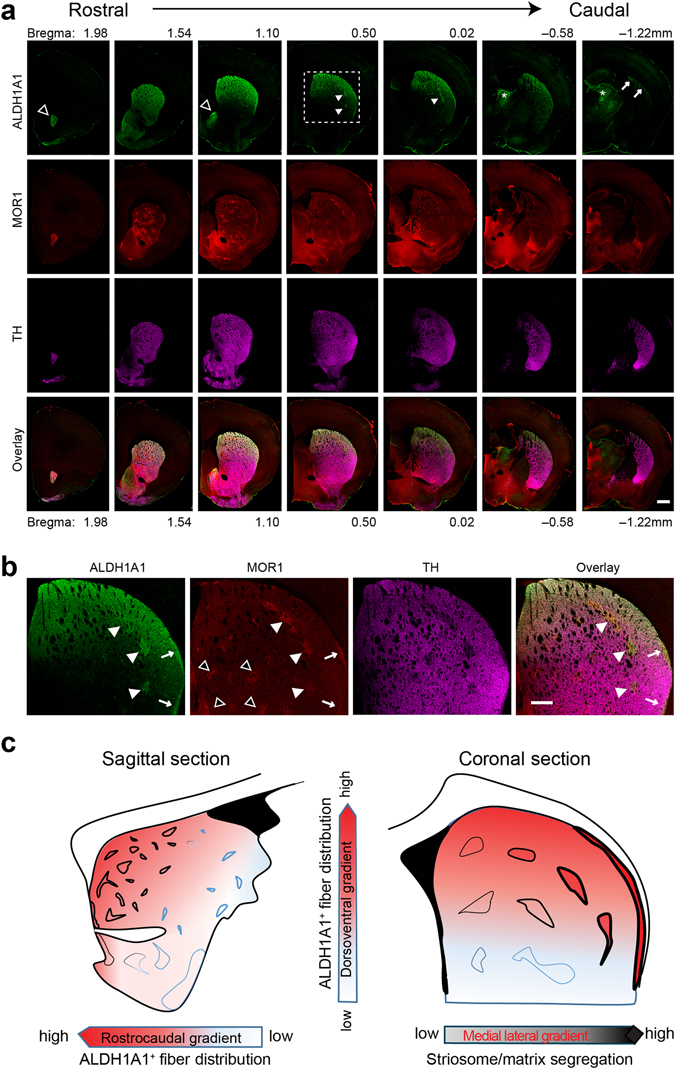
Distinct projection pattern of ALDH1A1–positive DA axons at striatum. (a) Representative images show ALDH1A1 (green), MOR1 (red), and TH (purple) co-staining in sequential striatal coronal sections of 1-month-old mice. Open arrowheads point to nucleus accumbens shell. Solid arrowheads indicate the striosomes. Arrows identify subcallosal streaks. Asterisks mark bundles positive to ALDH1A1 staining, but negative to TH staining. Scale bar: 500 µm. (b) Enlarged images of boxed area (a) show co-staining of ALDH1A1 and MOR1 in strisomes (solid arrowheads) and subcallosal streaks (arrows) at DLS, but not in striosomes (open arrow) at DMS. Scale bar: 100 µm. (c) Schematic diagram outlines the projection pattern of ALDH1A1–positive SNpc DA axon fibers in dorsal striatum.
ALDH1A1–positive fibers release less dopamine at the DLS striosomes
We recently characterized a line of transgenic mice that express GFP mainly in the striatal neurons at striosomes under the transcriptional control of nuclear receptor Nr4a121. Consistent with this early finding, the GFP signals overlapped with ALDH1A1 and MOR1 staining in the DLS striosomes of 3-month-old Nr4a1–GFP transgenic mice (Fig. 2a). Using GFP as a visible striosomal marker using fluorescence microscopy, we measured the dopamine release of ALDH1A1–positive fibers at the striosomes and ALDH1A1–negative fibers in adjacent matrix by fast scan cyclic voltammetry (FSCV) (Fig. 2b). In confirmation of our previous observations16, the magnitude of afferent stimulus-evoked DA release amplitudes was substantially lower in the DLS striosomes than the surrounding matrix areas in Nr4a1-GFP mice with the Aldh1a1 wild-type (Aldh1a1 +/+) background (Fig. 2c,d; ANOVA interaction. Stim. Intensity × Compartment: F(3,30) = 5.363, p < 0.01; Bonferroni’s post hoc test: *p < 0.05, ***p < 0.001). Meanwhile, no obvious alteration of τ was found (for stimulation of 200 µA, Matrix: 0.4773 ± 0.08174 Vs Striosome: 0.4283 ± 0.06379; t(10) = 0.4726 p > 0.05). These data suggestdifferent dopamine release in ALDH1A1–positive fibers that converge to the DLS striosomes.
Figure 2.
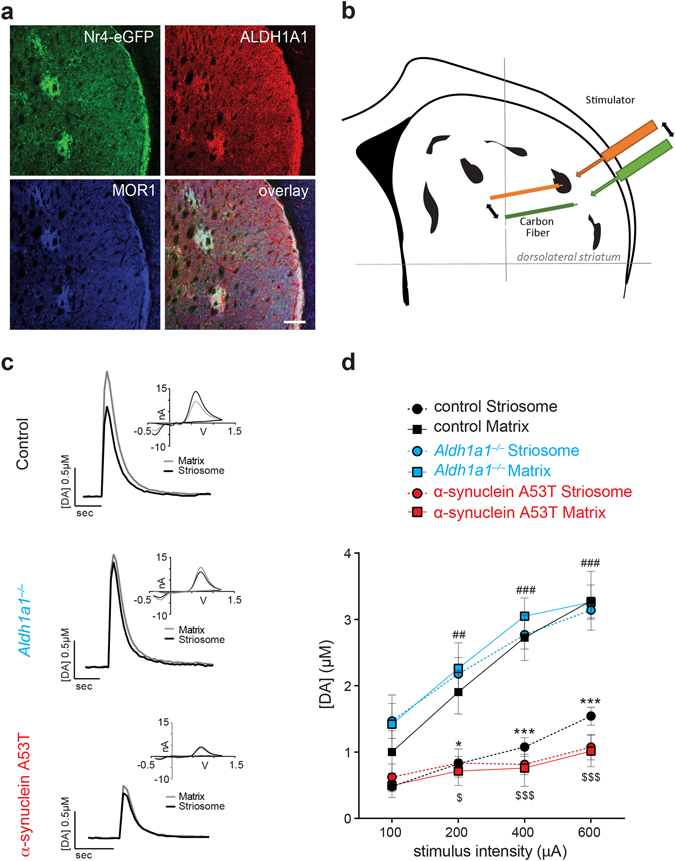
Different dopamine release properties of ALDH1A1–positive and negative fibers at DLS. (a) Representative images show GFP (green), ALDH1A1 (red), and MOR1 (purple) staining at DLS of Nr4a1-GFP mice on control Aldh1a1 +/+ background. Scale bar: 100 µm. (b) Schematic diagram of electrode placement in acute striatal slice for FSCV recording. Distances between electrodes (~100 µm) and from corpus callosum were maintained constant for dopamine release recording in striosome (orange) and matrix (green). (c) Representative traces of electrically evoked dopamine release in dorsolateral striatal compartments of control, Aldh1a1 −/−, and α-synuclein A53T mice, with current and voltage (CV) curve as inset. (d) Input/Output curves of peak dopamine release in dorsolateral striatal compartments of control, Aldh1a1 −/−, and α-synuclein A53T mice (n = 5 animals per genotype, 7 to 10 striosomes-matrix pairs per animal). Data were presented as mean ± SEM. Two-way ANOVA was used for statistical analysis, followed by Bonferroni’s post hoc tests.
ALDH1A1 regulates dopamine release at the DLS striosomes
To evaluate the contribution of ALDH1A1 activity to DLS striosomal DA release, we performed FSCV in Nr4a1-GFP mice crossbred into the Aldh1a1 knockout (Aldh1a1 −/−) background. As shown in the I/O curves, ALDH1A1-deficiency led to increases of DA release peak amplitude selectively in striosomes (Fig. 2c,d; ANOVA Interaction for Genotype × Stim. Intensity × Compartment: F(15,112) = 1.925, p < 0.05; Post hoc test for striosomal area compared to controls: ##p < 0.01, ###p < 0.001). These results demonstrate that ALDH1A1 actively regulates dopamine release in ALDH1A1–positive fibers projecting to the DLS striosomes, but not the surrounding matrix area.
Overexpression of human α–synuclein A53T mutation suppresses DA release mainly in matrix
We previously observed a drastic reduction of dopamine release at the dorsal striatum of α-synuclein A53T transgenic mice18. The transgenic α-synuclein was uniformly expressed throughout TH–positive fibers innervating striosomes and matrix (Supplementary Fig. S3). However, we do not know whether the presence of PD-related mutated α-synuclein affects the dopamine release similarly at the ALDH1A1–positive and negative terminals. Here we crossbred Nr4a1-GFP mice with the α-synuclein A53T transgenic mice to mark the ALDH1A1–positive terminals in the DLS striosomes. Data collected from the α-synuclein A53T mice revealed a substantial reduction of dopamine release in the matrix, while the striosomes were largely spared (Fig. 2c,d; ANOVA interaction Genotype × Stim. Intensity × Compartment: F(9, 60) = 7.763, p < 0.001). These data indicate that ALDH1A1–positive DA fibers are more resistant to α-synuclein overexpressioninduced dopamine release impairments compared to the negative ones.
DAT blockade differentially affects dopamine release at striatal compartments in Aldh1a1+/+, Aldh1a1−/− and α-synuclein A53T mice
Our previous study demonstrated that DAT blockade differentially potentiates peak DA release in striosome and matrix compartments16. We confirmed this finding in which the DAT blocker, cocaine, enhanced dopamine release more in the striosomes than in the proximal matrix areas in control mice (Fig. 3a,b; left panel: ANOVA interaction Stim. Intensity × Compartment: F(8, 64) = 3.187, p < 0.01; Bonferroni’s post hoc Test *p < 0.05). However, this inter-compartmental difference was not detectable in the Aldh1a1 −/− mice (Fig. 3a,b; left panel: ANOVA interaction Genotype × Stim. Intensity × Compartment: F(24, 128) = 2.880, p < 0.001; Bonferroni’s post hoc Test: striosomes $p < 0.05, $$p < 0.01). A similar lack of inter-compartmental difference was observed in α-synuclein A53T mutant mice, where both matrix and striosomes showed increased DA release to a similar extent after cocaine application (Fig. 3a,b; left panel: ANOVA interaction Genotype × Stim. Intensity × Compartment: F(24, 128) = 2.880, p < 0.001; Bonferroni’s post hoc Test: matrix #p < 0.05, ##p < 0.01, ###p < 0.001; striosomes @p < 0.05). On the other hand, no inter-compartmental difference in τ was observed following DAT blockage in any of the three mouse lines (Fig. 3a,b; right panel). It is worth noting that after DAT blockade the τ was significantly higher in α-synuclein A53T mutant mice compared to Aldh1a1 +/+ mice (Fig. 3a, right panel; ANOVA interaction Genotype × Stim. Intensity × Compartment: F(24, 128) = 2.603, p < 0.001; Bonferroni’s post hoc Test: matrix ###p < 0.001; striosomes @p < 0.05). These results suggest different functional properties of DAT located at Aldh1a1 +/+ and Aldh1a1 −/− axon terminals, suggesting that changes in ALDH1A1 and α–synuclein expression can alter DAT function in axon terminals.
Figure 3.
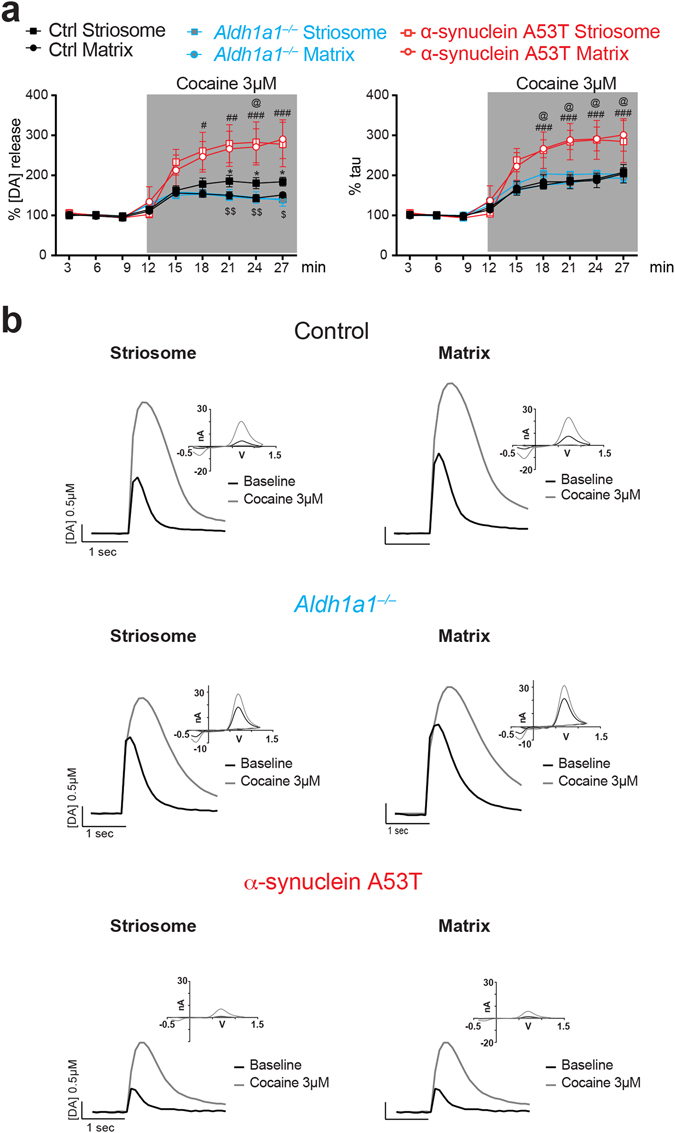
DAT blockade differentially affects dopamine release in ALDH1A1–positive and negative axons at DLS of control and mutant mice. (a) Percentages of electrical evoked dopamine amplitude (left) and uptake time constant (right) normalized to their baseline values (n = 5 animals per genotype, 2 sections per animal). Data were presented as mean ± SEM. Two-way ANOVA was used for statistical analysis, followed by Bonferroni’s post hoc tests. (b) Representative traces of electrically evoked dopamine release in DLS sub-compartments of control, Aldh1a1 −/− and α-synuclein A53T mutant mice, with CV curve as inset.
Dopamine D2 receptors modulate DA release similarly between compartments in Aldh1a1+/+, Aldh1a1−/− and α-synuclein A53T mice
D2 autoreceptors (DRD2s) provide a a presynaptic feedback system that regulates DA neurotransmission in striatum22, 23. We examined the ability of the DRD2 agonist quinpirole to depress dopamine release in slices from the different mouse lines. Application of quinpirole (25 nM) decreased dopamine release amplitude to a similar extent in Aldh1a1 +/+, Aldh1a1 −/− and α-synuclein A53T mice, with little effect on τ in all groups (Fig. 4a,b). These data suggest the presence of ALDH1A1 or mutant α-synuclein do not affect the function of presynaptic DRD2s that alter DA release.
Figure 4.
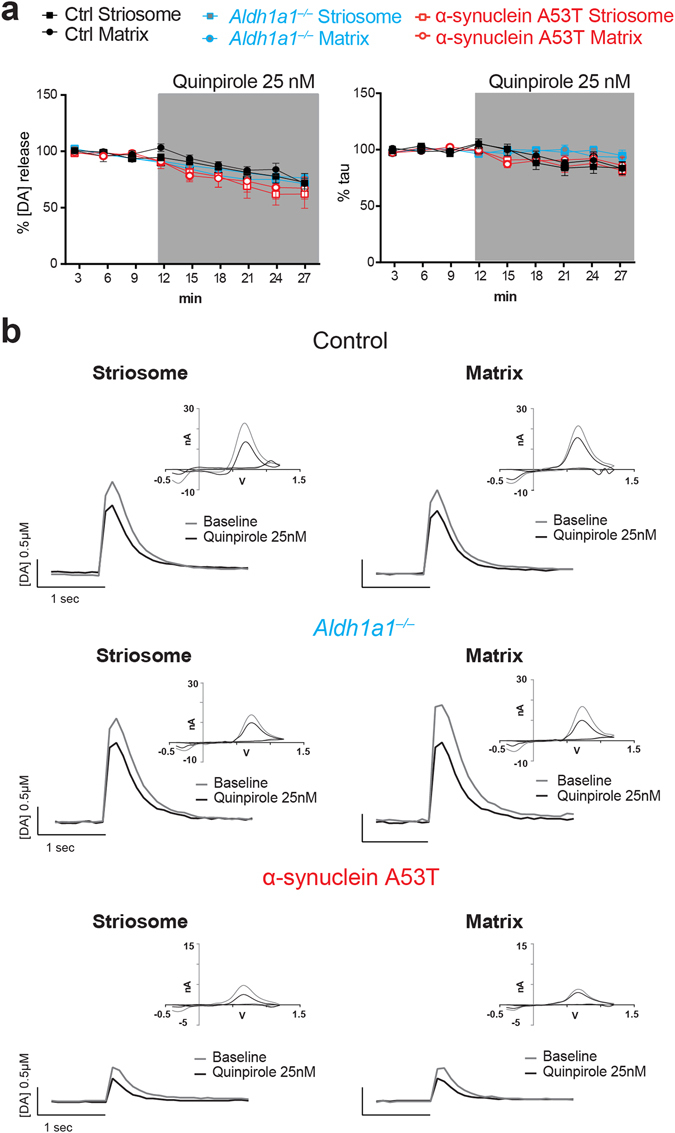
DRD2 autoreceptors modulate dopamine release similarly between ALDH1A1–positive and negative axons at DLS. (a) Percentages of electrical evoked dopamine amplitude (left) and uptake time constant (right) normalized to their baseline values (n = 5 animals per genotype, 2 sections per animal). Data were presented as mean ± SEM. Two-way ANOVA was used for statistical analysis, followed by Bonferroni’s post hoc tests. (b) Representative traces of electrically evoked dopamine release in DLS sub-compartments of control, Aldh1a1 −/− and α-synuclein A53T mutant mice, with CV curve as inset.
Nicotinic acetylcholine receptors (nAChRs) have no regionally specific effect on dopamine dynamics in striosomes and matrix, but have reduced effects on striatal dopamine release in Aldh1a−/− and α-synuclein A53T mice
Striatal nAChRs influence many aspects of striatal function and related behavior, in part through a strong interaction with the DA system24. ACh stimulates dopamine release through nACh receptors (nAChRs) on DA axons, mainly via β2-subunit-containing receptors25, 26. We found that DHβE (1 μM), an antagonist ofpresynaptic β2-subunit-containing nAChRs on DA terminals, had the same inhibitory effect on dopamine release in both striatal compartments, but produced a smaller magnitude effect in Aldh1a1 −/−- and A53T mutants (Fig. 5a, left panel; Fig. 5b). Both compartments of Aldh1a1 −/− mice showed lower DHβE inhibition compared with control littermates (ANOVA interaction Compartment × Time: F(24, 128) = 8.71; Bonferroni’s post hoc test: Striosomes: $$$p < 0.001; Matrix: &&p < 0.01, &&&p < 0.001). In A53T mutant mice, the effect of presynaptic nAChR blockade was also proportionally lower than in control (Fig. 5a, left panel; ANOVA interaction Time × Compartment: F(24, 128) = 2.061, p < 0.01; Bonferroni’s post hoc test: Control vs α-synuclein A53T: Matrix: #p < 0.05; Striosomes: @p < 0.05). Meanwhile, the DHβE-induced reduction in τ was significantly reduced in controls in both Aldh1a1 −/− (Fig. 5a, right panel; ANOVA interaction Time × Compartment: F (24, 128) = 2.301, p < 0.01; Bonferroni’s post hoc test: Striosomes $p < 0.05, $$p < 0.01, $$$p < 0.001; Matrix &p < 0.05) and A53T mutant mice (Fig. 5a, right panel; ANOVA interaction Time × Compartment: F (24, 128) = 1.636, p < 0.05. Bonferroni’s post hoc test: Matrix: #p < 0.05). These results suggest that dopamine release in the DLS compartments of Aldh1a1 −/−- and A53T mice is less affected by the inhibition of β2-nAChR.
Figure 5.
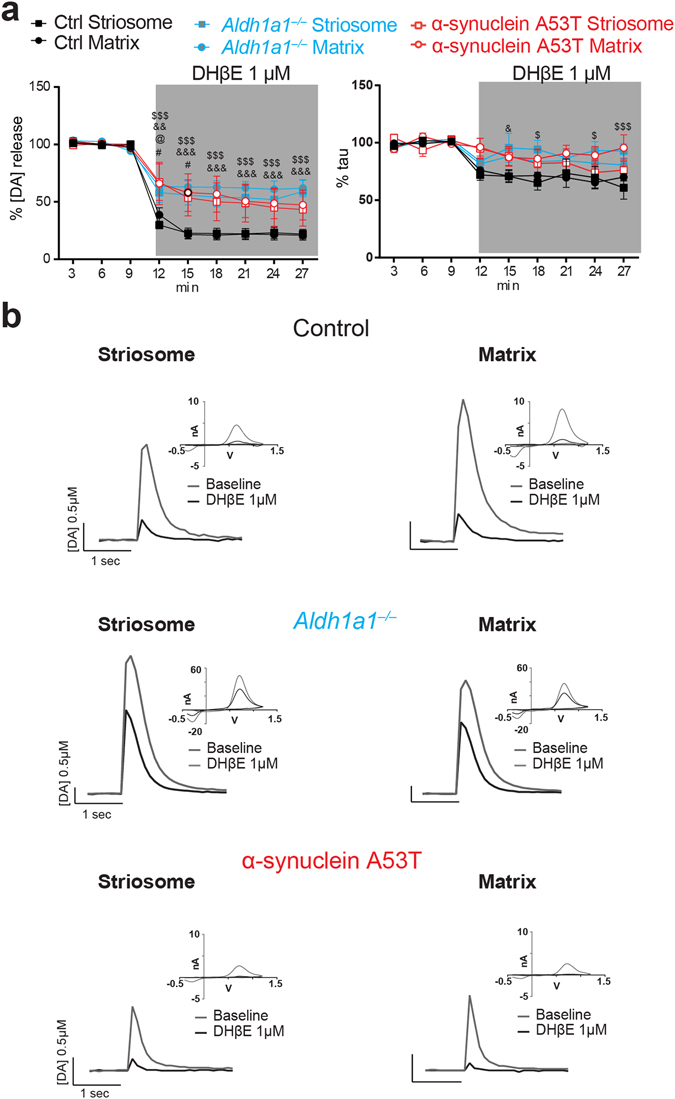
Nicotinic acetylcholine blockade does not have differential effects on DA release from ALDH1A1–positive and negative axons, but is less effective in both Aldh1a1 −/− and α-synuclein A53T mutant mice. (a) Percentages of electrical evoked dopamine amplitude (left) and uptake time constant (right) normalized to their baseline values (n = 5 animals per genotype, 2 sections per animal). Data were presented as mean ± SEM. Two-way ANOVA was used for statistical analysis, followed by Bonferroni’s post hoc tests. (b) Representative traces of electrically evoked dopamine release in DLS sub-compartments of control, Aldh1a1 −/− and α-synuclein A53T mutant mice, with CV curve as inset.
ALDH1A1–positive DA axon fibers show differential expression of TH, DAT, and VMAT2 in striosomes and matrix
Given that TH, DAT, vesicular monoamine transporter 2 (VMAT2), and ALDH1A1 are key proteins participating in synthesis, transport, and catabolism of dopamine27–29, we examined their expression to explore the underlying molecular mechanisms of differential dopamine release dynamics of ALDH1A1–positive fibers in the DLS. Using the same NR4a1-GFP reporter to outline the striosomes, we co-stained the middle one–third of striatal coronal sections of 1–month–old Aldh1a1 +/+, Aldh1a1 −/−, and α-synuclein A53T mice with antibodies against ALDH1A1, TH, DAT, and VMAT2 (Fig. 6a). The intensities of ALDH1A1, DAT, and VMAT2–positive fluorescence signals were substantially higher in the striosomes compared to the adjacent matrix compartments (Fig. 6a,b). A similar weaker TH staining was also observed in the DLS striosomes of 3-month-old mice (Supplementary Fig. S1). By contrast, the intensity of TH staining was weaker in the striosomes (Fig. 6a,b). The absence of Aldh1a1 did not alter the overall distribution of TH, DAT and VMAT2 between striosomes and matrix (Fig. 6a,b). On the other hand, expression of the transgenic α-synuclein A53T mutant protein increased the expression of TH but suppressed the expression of ALDH1A1 (ANOVA Genotype × Protein Interaction: F (3, 73) = 5.576; p < 0.01; Bonferroni’s post hoc test: *p < 0.05) but not significantly for DAT and VMAT2 (Fig. 6a,b). The differential expression of ALDH1A1, TH, at the ALDH1A1–positive fibers may contribute to the distinct dopamine release dynamics in the DLS striosomes.
Figure 6.
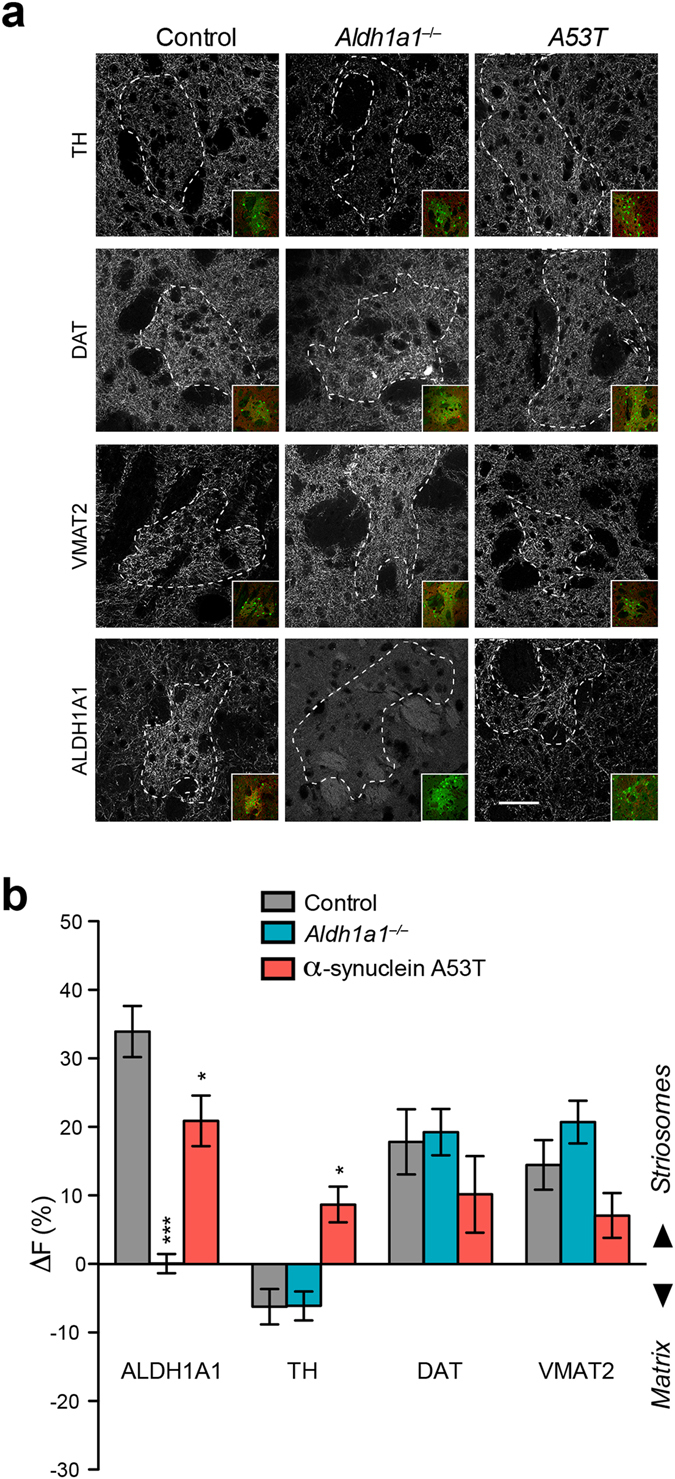
Distribution of TH, DAT, VMAT2, and ALDH1A1 proteins in the striosomes and matrix of DLS. (a) Representative images show TH, DAT, VMAT2, and ALDH1A1 staining in the DLS of 1-month-old Nr4a-GFP mice in control, Aldh1a1 −/− and α-synuclein A53T mutant background. Dashed lines outline the striosomes. Inserts show co-staining with GFP to mark the strisomes. Scale bar: 50 µm. (b) Bar graph depicts the relative fluorescence intensity (ΔF) of ALDH1A1, TH, DAT, and VMAT2 staining in the striosome and matrix compartments (n = 5 animals per genotype, x matrix-striosome pairs per animal). Data were presented as mean ± SEM. One-way ANOVA was used for statistical analysis for each protein, followed by Bonferroni’s post hoc tests where Aldh1a1 −/− and α-synuclein A53T mutant were compared to Controls (*p < 0.05, ***p < 0.001).
Discussion
It has been well documented that DA neurons located at the ventrolateral tier of SNpc preferentially degenerate in the postmortem brains of PD patients30, 31. Recently, we demonstrated that those DA neurons selectively express ALDH1A1, a multi-functional aldehyde dehydrogenase8. Given that ALDH1A1 is predominantly expressed by the midbrain DA neurons2, 8, 19, 20, the appearance of ALDH1A1 staining in the dorsal striatum most likely reflects the trafficking of ALDH1A1 proteins to the axon terminals. Indeed, ALDH1A1 signals were co-localized with the DA neuron marker TH in the striatum, but not with the striosomal neuron marker MOR1. ALDH1A1 staining reveals a more complex projection pattern of ventral SNpc DA neurons compared to early neuron tracing studies9. While the cell bodies of ALDH1A1–positive DA neurons mostly reside at the ventral tier of SNpc, the axon fibers more heavily innervate the most dorsal portions of dorsal striatum. Furthermore, ALDH1A1–positive DA axon fibers only show a clear convergence in the striosomes in the DLS. The more homogenous projections to DMS likely result from the presence of overlapping DA innervation from ALDH1A1–positive DA neurons in the VTA20, likely the same DA neurons involved in a precise form of reward prediction32. DLS-projecting SNpc DA neurons have been implicated in motor control and show similar responses to both rewarding and aversive stimuli33. DMS-projecting neurons, on the other hand, respond differently to rewarding and aversive inputs33, 34. Future studies will need to further distinguish subpopulations of ALDH1A1–positive SNpc DA neurons based on their distinct molecular composition, connectivity and functionality.
Degeneration of DLS–projecting ALDH1A1–positive SNpc neurons is strongly implicated in the pathogenesis of PD2, 30, 31, 35. As an initial step to characterize the functional properties of these neurons, we systematically measured their dopamine release and reuptake parameters under different genetic and pharmacological manipulations. Normally, dopamine release amplitudes are lower in striosomes than matrix areas in control mice16, 17, which is correlated with a lower level of TH expression in the axons terminating in the striosomes of human36 and mouse brains, These observations uggest that TH expression level is a factor contributing to reduced dopamine release in DLS–projecting ALDH1A1–positive SNpc neurons. However, genetic ablation of Aldh1a1 substantially increased DA release selectively in the DLS striosomes to a level that matches dopamine release to the adjacent matrix, arguing that ALDH1A1 plays a more important role in determining the inter-compartmental differences in dopamine release. This finding is also consistent with an early report that shows a higher than normal extracellular striatal dopamine level in Aldh1a1 −/− mice as measured by in vivo microdialysis27. Since ALDH1A1 inhibition does not affect the overall expression of TH, DAT and VMAT2 proteins27 or their relative distributions in DA fibers ending in striosomes and matrix, ALDH1A1 likely limits the dopamine release through increasing metabolism of cytosolic dopamine37. Therefore, a lack of ALDH1A1 may promote cytosolic dopamine buildup, resulting in more vesicular loading and release upon stimulation. Noticeably, the expression of ALDH1A1 is substantially suppressed in α-synuclein A53T transgenic mice and PD brains8. The reduced ALDH1A1 expression, together with a relative increase of TH expression, likely serves as a compensatory mechanism to enhance dopamine release. Paradoxically, the loss of ALDH1A1 also leads to higher levels of highly reactive DOPAL intermediates, which are detrimental to the survival of DA neurons2, 8, 38. Understanding how ALDH1A1–positive fibers react to PD-related genetic and environmental insults could be informative to develop new therapeutic approaches in the treatment of PD27.
Increasing evidence suggests that α-synuclein mutations directly interfere with the synaptic vesicle release machinery and inhibit neurotransmission39–41. Compared to the other α-synuclein mouse models, this line of α-synuclein transgenic mice develop much more robust pathological abnormalities18. At one month of age, these α-synuclein transgenic mice show >15% loss of SNc DA neurons, and >80% reduction of dopamine release18. More α-synuclein aggregates were accumulated in the ALDH1A1-negative DA neurons compared to the positive ones8, which may contribute to the preferential loss of ALDH1A1-negative DA neurons in these mice. Although transgenic α-synuclein is expressed comparably by both ALDH1A1–positive and negative SNpc DA neurons8 and fibers, the reduction of dopamine release was more severe in the ALDH1A1-negative axons terminating in the matrix. The preferential degeneration of ALDH1A1–negative SNpc DA neurons and axon fibers8 likely contributes to the overall reduction of dopamine release in the matrix. The presence of excessive α-synuclein also substantially increased the effect of cocaine on the DA peak amplitude. The greater relative efficacy of cocaine to affect dopamine dynamics in the α–synuclein transgenic mice may be attributable to a compensatory increase in basal DAT activity following DA terminal loss in these mice18. In addition, under our experimental conditions we did not observe any difference in DRD2 autoreceptor-mediated inhibition of dopamine release in either striatal compartment by the DRD2. Therefore, the observed DAT adaptation may be independent of any indirect effect from DRD2 autoreceptor activity.
Striatal ACh influences many aspects of striatal function, and related behaviors, through a strong interaction with the DA system24. ACh stimulates DA release through nAChRs on DA axons, mainly via β2-subunit-containing receptors25, 26. As mentioned previously16, we did not observe a robust difference of presynaptic nAChR blockade between the DA terminals ended in striosomes and matrix, suggesting that this cholinergic effect has no preferential effects within a specific striatal compartment under current experimental conditions. However, dopamine release in the DLS of Aldh1a1 −/− and α-synuclein A53T mutant mice was less affected by antagonism of β2-nAChRs. A floor effect may prohibit our ability to determine the efficacy of nAChR inhibition on DA release in α-synuclein transgenic mice, as well as any intercompartmental contrast. On the other hand, augmented extracellular dopamine in Aldh1a1 −/− striata very likely hampers the ability of the local ACh system to enhance DA release, although more detailed investigation is needed to understand why Aldh1a1 −/− DA neurons are less responsive to nAChR blockade.
In summary, our findings reveal a distinct projection pattern of an ALDH1A1–positive subpopulation of DA neurons to the dorsal striatum. We further characterized the dopamine release properties of a fraction of these axons that converge on the DLS striosomes, supporting an important role of ALDH1A1 in regulating dopamine availability and release.
Methods
Animal subjects
Pitx3 +/IRES2–tTA knock–in mice and tetO–A53T α-synuclein transgenic mice were created as described previously18, 42. Aldh1a knockout (Aldh1a1 −/−) mice43 were obtained from the Jackson Laboratory (Bar Harbor, ME), and backcrossed with C57BL6J mice for more than five generations. Nr4a1–EGFP139Gsat (Nr4a1–GFP) transgenic mice21 were obtained from GENSAT and backcrossed with C57BL6J mice for more than five generations. The Pitx3 +/IRES2–tTA mice were bred with Nr4a1–GFP mice for the generation of Nr4a1–GFP::Pitx3 +/IRES2–tTA bigenic mice (referred as control mice), which then bred with tetO–A53T transgenic mice to generate Nr4a1–GFP::Pitx3 +/IRES2–tTA::tetO–A53T triple-transgenic mice (referred as α-synuclein A53T mice). Nr4a1–GFP mice were also crossbred with Aldh1a1 −/− mice to generate Nr4a1–GFP::Aldh1a1 −/− bigenic mice (referred as Aldh1a1 −/− mice). All mice were housed in a 12-hour-light/dark cycle and fed regular diet ad libitum. The Animal Care and Use Committee (ACUC) of the National Institute on Aging, NIH approves all the mouse-related experimental procedures performed in this study. We carried out all the mouse experiments following the approved guidelines.
Genotyping
Genomic DNA was prepared from tail biopsy using DirectPCR Lysis Reagent (Viagen Biotech Inc.) and subjected to PCR amplification using specific sets of PCR primers for each genotype, including Pitx3 +/IRES2–tTA knock–in mice (Pitx3-F: GACTGGCTTGCCCTCGTCCCA and Pitx3-R: GTGCACCGAGGCCCCAGATCA), tetO–A53T transgenic mice (PrpEx2-F: TACTGCTCCATTTTGCGTGA and SNCA-R: TCCAGAATTCCTTCCTGTGG), Aldh1a1 −/− mice (ALDH1A1muF: CTATCGCCTTCTTGACGAGTTCTT and ALDH1A1muR: CCTTGTACATCTTAACGGTGCACA), Aldh1a1 wild-type (Aldh1a1 +/+) mice (ALDH1A1wtF: TAAAGACCTGGATAAGGCCATCA and ALDH1A1wtR: ACGGTGCACAAAATAAACATCTG), and Nr4a1–GFP mice were genotyped at birth by fluorescence of the thymus, tails and ears using a hand held LED light source.
Stereotaxic viral injection
Stereotaxic AAV injections were conducted on 3–month–old C57BL6/J mice. Before surgery, mice were deeply anesthetized by intraperitoneal injection of ketamine (100 mg/kg)/xylazine (10 mg/kg) solution. A total volume of 500 nl virus solution was injected unilaterally into Substantia Nigra pars compacta (SNpc, coordinates used, AP: −1.5 mm, ML: ±0.9 mm from bregma, DV: −4.1 mm from exposed dura mater). Virus solution was injected at an infusion rate of 100 nl/min and withdrawn 10 min after the end of injection. Following virus injection, the scalp was sutured and mice were returned to their home cages. Virus-injected mice were used for experiment at least 4 weeks after virus infusion.
Immunohistochemistry and light microscopy
Mice were anesthetized with ketamine and then transcardially perfused with a 4% formaldehyde/PBS solution as described previously44. The brains were then extracted and post fixed in 4% formaldehyde overnight, then submerged in 30% sucrose for 24 hours at 4 °C for later sectioning. Series of 40 μm sections were collected using a cryostat (Leica Biosystems, Richmond IL). The sections were blocked in a 10% normal donkey serum, 1% bovine serum albumin, 0.3% Triton X-100, PBS solution for 1 hour at room temperature on an orbital shaker at low speed. The sections were then incubated in a primary antibody/PBS solution over two nights as follows: goat anti-ALDH1A1 (1:1,000; R&D system), or rabbit anti-ALDH1A1 (1:1,000; Sigma-Aldrich), or rabbit anti-MOR1 (1:3,000; Immunostar), or mouse anti-TH (1:1,000; Immunostar), or rat anti-DAT (1:1,000; Millipore), or rabbit anti-VMAT2 antibody (1:50,000, obtained from Dr. Gary W. Miller) or mouse anti-human α-synuclein antibody (1:1,000; sc211, Santa Cruz). Sections were then washed three times in PBS before being incubated in secondary antibody solutions with Alexa 488- or Alexa Fluor 546–, or Alexa Fluor 633-conjugated secondary antibody (1:500, Invitrogen) at room temperature for 1 hour. Following three washes in PBS, sections were mounted onto subbed slides, and coverslipped with mounting media (ProLong® Gold Antifade Mountant, Life technology). Stained sections were imaged using a laser scanning confocal microscope (LSM 780; Zeiss). The paired images in the figures were collected at the same gain and offset settings. After collection, processing was applied uniformly to all paired images. The images were either presented as a single optic layer from individual fields or displayed as maximum intensity projections to represent confocal stacks.
Image analysis
For the quantitative assessment of various marker proteins in striatum, the dorsal lateral striatum (DLS) patches were chosen randomly imaged by a 40× oil immersion lens from three mice per group. A single optical layer was taken using identical settings and exported to ImageJ (NIH) for imaging analyses. Images were converted to an 8-bit color scale (fluorescence intensity from 0 to 255) using ImageJ. Areas of interest, striosomes and adjacent matrix, were first selected by Polygon or Freehand selection tools and then subjected to measurement by mean optical intensities. The mean intensity for the background area was subtracted from the selected area to determine the net mean intensity.
Fast-Scan Cyclic Voltammetry
FSCV was used to investigate dopamine release evoked by electrical stimulation in matrix and striosomal compartments localized in the dorsolateral quadrant of the striatum. Five mice of 10–14 weeks old from each of the three experimental groups (control, Aldh1a1 −/− and A53T) were considered in this analysis. Since no significant difference was observed between Nr4a1–GFP and Nr4a1–GFP::Pitx3 +/IRES2–tTA mice, the latter were considered as a valid control for all group comparisons. Coronal brain slices (250 μm) containing dorsal stratum from mice were prepared as described previously45. Slices were kept in oxygenated modified Kreb’s buffer as follows (in mM): NaCl 126, KCl 2.5, NaH2PO4 1.2, CaCl2 2.4, MgCl2 1.2, NaHCO3 25, glucose 11, HEPES 20, L-ascorbic acid 0.4; at room temperature until required. Recordings were made at 32 °C in a chamber perfused at a rate of 1.5 ml/min. Cylindrical carbon-fiber microelectrodes (75–125 μm exposed fiber) were prepared with T650 fibers (6 μm diameter, Goodfellow) and inserted into a glass pipette. The carbon-fiber electrode was held at −0.4 V, and the potential was ramped to +1.2 V and back at 400 V/s every 100 ms. Dopamine (DA) release was evoked by a rectangular, electrical pulse stimulation (if not specified otherwise: 200 μA, 1.2 ms, monophasic) generated by a DS3 Constant Current Stimulator (Digitimer, Hertfordshire, UK). Delivery of an electrical stimulation was applied every 3 min by a concentrical bipolar electrode placed 100 μm from the recording electrode and always from matrix (Nr4a1–GFP–negative) area. Data collection was done using DEMON software46. Ten cyclic voltammograms of charging currents were recorded as background before stimulation, and the average was subtracted from data collected during and after stimulation. Matrix/striosome ratio of DA peak amplitude responses were obtained from six to ten striosome-matrix paired recordings (pseudo-randomly performed from individual striosomes and proximal matrix area) across the rostral-caudal extent of the dorsolateral striatum, and averaged for each single mouse. Input/output function (I/O) curves were constructed by plotting peak amplitude of DA release as a function of stimulus currentover a range of stimulus intensities. The time constant (τ) of the evoked DA response was used as an index of DA uptake. Two of the most representative striosome/matrix area pairs were chosen for each mouse, and selected randomly for pharmacological manipulation experiments.
Drugs
Quinpirole, sulpiride, dihydro-β-erythrodine (DhβE) and cocaine hydrochloride were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Statistics
GraphPad Prism 6 (GraphPad Software, La Jolla, CA) was used for all statistical tests. Unpaired t-tests and two-ways ANOVA were used for the experiments examining dopamine release between groups and/or compartments, followed by Bonferroni’s post hoc test in case of significant interaction between factors, and reported in the figures.
Electronic supplementary material
Acknowledgements
We want to thank Yolanda Mateo for her excellent suggestions and technical assistance, and Dr. Lu Chen of Stanford University for providing the Aldh1a1 siRNA AAV vectors. This work was supported by the intramural research programs of National Institute on Aging (HC) and the National Institutes of Health, National Institute on Alcohol Abuse and Alcoholism, Division of Intramural Clinical and Biological Research (MID and DML).
Author Contributions
C.S., H.C. and D.L. designed the experiments and wrote the manuscript. C.S. and H.C. prepared the figures. J.W. contributed to Figures 1 and S1. C.S. contributed to Figures 2–5. W.Z. contributed to Figure 6. J.P. contributed to Figure S2. X.C. contributed to Figure S3. A.S., and M.D. provided technical support.
Competing Interests
The authors declare that they have no competing interests.
Footnotes
Carmelo Sgobio, Junbing Wu and Wang Zheng contributed equally to this work.
Electronic supplementary material
Supplementary information accompanies this paper at doi:10.1038/s41598-017-05598-1
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Vasiliou V, Pappa A, Petersen DR. Role of aldehyde dehydrogenases in endogenous and xenobiotic metabolism. Chem Biol Interact. 2000;129:1–19. doi: 10.1016/S0009-2797(00)00211-8. [DOI] [PubMed] [Google Scholar]
- 2.Cai H, Liu G, Sun L, Ding J. Aldehyde Dehydrogenase 1 making molecular inroads into the differential vulnerability of nigrostriatal dopaminergic neuron subtypes in Parkinson’s disease. Translational neurodegeneration. 2014;3:27. doi: 10.1186/2047-9158-3-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Marchitti SA, Deitrich RA, Vasiliou V. Neurotoxicity and metabolism of the catecholamine-derived 3,4-dihydroxyphenylacetaldehyde and 3,4-dihydroxyphenylglycolaldehyde: the role of aldehyde dehydrogenase. Pharmacol Rev. 2007;59:125–150. doi: 10.1124/pr.59.2.1. [DOI] [PubMed] [Google Scholar]
- 4.Rees JN, Florang VR, Eckert LL, Doorn JA. Protein reactivity of 3,4-dihydroxyphenylacetaldehyde, a toxic dopamine metabolite, is dependent on both the aldehyde and the catechol. Chem Res Toxicol. 2009;22:1256–1263. doi: 10.1021/tx9000557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Duester G. Families of retinoid dehydrogenases regulating vitamin A function: production of visual pigment and retinoic acid. European journal of biochemistry/FEBS. 2000;267:4315–4324. doi: 10.1046/j.1432-1327.2000.01497.x. [DOI] [PubMed] [Google Scholar]
- 6.Kim JI, et al. Aldehyde dehydrogenase 1a1 mediates a GABA synthesis pathway in midbrain dopaminergic neurons. Science. 2015;350:102–106. doi: 10.1126/science.aac4690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.McCaffery P, Drager UC. High levels of a retinoic acid-generating dehydrogenase in the meso-telencephalic dopamine system. Proc Natl Acad Sci USA. 1994;91:7772–7776. doi: 10.1073/pnas.91.16.7772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu G, et al. Aldehyde dehydrogenase 1 defines and protects a nigrostriatal dopaminergic neuron subpopulation. J Clin Invest. 2014;124:3032–3046. doi: 10.1172/JCI72176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bentivoglio M, Morelli M. The organization and circuits of mesencephalic dopaminergic neurons and the distribution of dopamine receptors in the brain. Hand Chem N. 2005;21:1–107. [Google Scholar]
- 10.Crittenden JR, Graybiel AM. Basal Ganglia disorders associated with imbalances in the striatal striosome and matrix compartments. Frontiers in neuroanatomy. 2011;5:59. doi: 10.3389/fnana.2011.00059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Smith JB, et al. Genetic-Based Dissection Unveils the Inputs and Outputs of Striatal Patch and Matrix Compartments. Neuron. 2016;91:1069–1084. doi: 10.1016/j.neuron.2016.07.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Herkenham M, Pert CB. Mosaic distribution of opiate receptors, parafascicular projections and acetylcholinesterase in rat striatum. Nature. 1981;291:415–418. doi: 10.1038/291415a0. [DOI] [PubMed] [Google Scholar]
- 13.Graybiel AM, Ragsdale CW, Jr., Yoneoka ES, Elde RP. An immunohistochemical study of enkephalins and other neuropeptides in the striatum of the cat with evidence that the opiate peptides are arranged to form mosaic patterns in register with the striosomal compartments visible by acetylcholinesterase staining. Neuroscience. 1981;6:377–397. doi: 10.1016/0306-4522(81)90131-7. [DOI] [PubMed] [Google Scholar]
- 14.Gerfen CR, Herkenham M, Thibault J. The neostriatal mosaic: II. Patch- and matrix-directed mesostriatal dopaminergic and non-dopaminergic systems. J Neurosci. 1987;7:3915–3934. doi: 10.1523/JNEUROSCI.07-12-03915.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gerfen CR, Baimbridge KG, Miller JJ. The neostriatal mosaic: compartmental distribution of calcium-binding protein and parvalbumin in the basal ganglia of the rat and monkey. Proc Natl Acad Sci USA. 1985;82:8780–8784. doi: 10.1073/pnas.82.24.8780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Salinas AG, Davis MI, Lovinger DM, Mateo Y. Dopamine dynamics and cocaine sensitivity differ between striosome and matrix compartments of the striatum. Neuropharmacology. 2016;108:275–283. doi: 10.1016/j.neuropharm.2016.03.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brimblecombe KR, Cragg SJ. Substance P Weights Striatal Dopamine Transmission Differently within the Striosome-Matrix Axis. The Journal of neuroscience: the official journal of the Society for Neuroscience. 2015;35:9017–9023. doi: 10.1523/JNEUROSCI.0870-15.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lin X, et al. Conditional expression of Parkinson’s disease-related mutant alpha-synuclein in the midbrain dopaminergic neurons causes progressive neurodegeneration and degradation of transcription factor nuclear receptor related 1. J Neurosci. 2012;32:9248–9264. doi: 10.1523/JNEUROSCI.1731-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.McCaffery P, Drager UC. High levels of a retinoic acid-generating dehydrogenase in the meso-telencephalic dopamine system. Proceedings of the National Academy of Sciences of the United States of America. 1994;91:7772–7776. doi: 10.1073/pnas.91.16.7772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Poulin JF, et al. Defining midbrain dopaminergic neuron diversity by single-cell gene expression profiling. Cell reports. 2014;9:930–943. doi: 10.1016/j.celrep.2014.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Davis MI, Puhl HL., 3rd Nr4a1-eGFP is a marker of striosome-matrix architecture, development and activity in the extended striatum. PLoS One. 2011;6:e16619. doi: 10.1371/journal.pone.0016619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Rice ME, Cragg SJ. Nicotine amplifies reward-related dopamine signals in striatum. Nature neuroscience. 2004;7:583–584. doi: 10.1038/nn1244. [DOI] [PubMed] [Google Scholar]
- 23.De Mei C, Ramos M, Iitaka C, Borrelli E. Getting specialized: presynaptic and postsynaptic dopamine D2 receptors. Current opinion in pharmacology. 2009;9:53–58. doi: 10.1016/j.coph.2008.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cragg SJ. Meaningful silences: how dopamine listens to the ACh pause. Trends Neurosci. 2006;29:125–131. doi: 10.1016/j.tins.2006.01.003. [DOI] [PubMed] [Google Scholar]
- 25.Rapier C, Lunt GG, Wonnacott S. Stereoselective nicotine-induced release of dopamine from striatal synaptosomes: concentration dependence and repetitive stimulation. J Neurochem. 1988;50:1123–1130. doi: 10.1111/j.1471-4159.1988.tb10582.x. [DOI] [PubMed] [Google Scholar]
- 26.Exley R, Cragg SJ. Presynaptic nicotinic receptors: a dynamic and diverse cholinergic filter of striatal dopamine neurotransmission. Br J Pharmacol. 2008;153(Suppl 1):S283–297. doi: 10.1038/sj.bjp.0707510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Anderson DW, Schray RC, Duester G, Schneider JS. Functional significance of aldehyde dehydrogenase ALDH1A1 to the nigrostriatal dopamine system. Brain Res. 2011;1408:81–87. doi: 10.1016/j.brainres.2011.06.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Vaughan RA, Foster JD. Mechanisms of dopamine transporter regulation in normal and disease states. Trends in pharmacological sciences. 2013;34:489–496. doi: 10.1016/j.tips.2013.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Daubner SC, Le T, Wang S. Tyrosine hydroxylase and regulation of dopamine synthesis. Arch Biochem Biophys. 2011;508:1–12. doi: 10.1016/j.abb.2010.12.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Fearnley JM, Lees AJ. Ageing and Parkinson’s disease: substantia nigra regional selectivity. Brain: a journal of neurology. 1991;114(Pt 5):2283–2301. doi: 10.1093/brain/114.5.2283. [DOI] [PubMed] [Google Scholar]
- 31.Kordower JH, et al. Disease duration and the integrity of the nigrostriatal system in Parkinson’s disease. Brain. 2013;136:2419–2431. doi: 10.1093/brain/awt192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Parker NF, et al. Reward and choice encoding in terminals of midbrain dopamine neurons depends on striatal target. Nat Neurosci. 2016;19:845–854. doi: 10.1038/nn.4287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lerner TN, et al. Intact-Brain Analyses Reveal Distinct Information Carried by SNc Dopamine Subcircuits. Cell. 2015;162:635–647. doi: 10.1016/j.cell.2015.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bromberg-Martin ES, Matsumoto M, Hikosaka O. Dopamine in motivational control: rewarding, aversive, and alerting. Neuron. 2010;68:815–834. doi: 10.1016/j.neuron.2010.11.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kish SJ, Shannak K, Hornykiewicz O. Uneven pattern of dopamine loss in the striatum of patients with idiopathic Parkinson’s disease. Pathophysiologic and clinical implications. N Engl J Med. 1988;318:876–880. doi: 10.1056/NEJM198804073181402. [DOI] [PubMed] [Google Scholar]
- 36.Morigaki R, Goto S. Putaminal Mosaic Visualized by Tyrosine Hydroxylase Immunohistochemistry in the Human Neostriatum. Frontiers in neuroanatomy. 2016;10:34. doi: 10.3389/fnana.2016.00034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Casida JE, et al. Benomyl, aldehyde dehydrogenase, DOPAL, and the catecholaldehyde hypothesis for the pathogenesis of Parkinson’s disease. Chem Res Toxicol. 2014;27:1359–1361. doi: 10.1021/tx5002223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Burke WJ, et al. Aggregation of alpha-synuclein by DOPAL, the monoamine oxidase metabolite of dopamine. Acta neuropathologica. 2008;115:193–203. doi: 10.1007/s00401-007-0303-9. [DOI] [PubMed] [Google Scholar]
- 39.Hunn BH, Cragg SJ, Bolam JP, Spillantini MG, Wade-Martins R. Impaired intracellular trafficking defines early Parkinson’s disease. Trends Neurosci. 2015;38:178–188. doi: 10.1016/j.tins.2014.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Xu J, et al. alpha-Synuclein Mutation Inhibits Endocytosis at Mammalian Central Nerve Terminals. J Neurosci. 2016;36:4408–4414. doi: 10.1523/JNEUROSCI.3627-15.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sulzer D, Surmeier DJ. Neuronal vulnerability, pathogenesis, and Parkinson’s disease. Mov Disord. 2013;28:41–50. doi: 10.1002/mds.25095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lin X, et al. Leucine-rich repeat kinase 2 regulates the progression of neuropathology induced by Parkinson’s-disease-related mutant alpha-synuclein. Neuron. 2009;64:807–827. doi: 10.1016/j.neuron.2009.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Fan X, et al. Targeted disruption of Aldh1a1 (Raldh1) provides evidence for a complex mechanism of retinoic acid synthesis in the developing retina. Mol Cell Biol. 2003;23:4637–4648. doi: 10.1128/MCB.23.13.4637-4648.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Cai H, et al. Loss of ALS2 function is insufficient to trigger motor neuron degeneration in knock-out mice but predisposes neurons to oxidative stress. J Neurosci. 2005;25:7567–7574. doi: 10.1523/JNEUROSCI.1645-05.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sgobio C, et al. Optogenetic measurement of presynaptic calcium transients using conditional genetically encoded calcium indicator expression in dopaminergic neurons. PLoS One. 2014;9:e111749. doi: 10.1371/journal.pone.0111749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yorgason JT, Espana RA, Jones SR. Demon voltammetry and analysis software: analysis of cocaine-induced alterations in dopamine signaling using multiple kinetic measures. J Neurosci Methods. 2011;202:158–164. doi: 10.1016/j.jneumeth.2011.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


