Abstract
The appearance of people associated with the Lapita culture in the South Pacific ~3,000 years ago1 marked the beginning of the last major human dispersal to unpopulated lands. However, the relationship of these pioneers to the long established Papuans of the New Guinea region is unclear. We report genome-wide ancient DNA data from four individuals from Vanuatu (~3100-2700 years before present) and Tonga (~2700-2300 years before present), and co-analyze them with 778 present-day East Asians and Oceanians. Today, indigenous peoples of the South Pacific harbor a mixture of ancestry from Papuans and a population of East Asian origin that does not exist in unmixed form today, but is a match to the ancient individuals. Most analyses have interpreted the minimum of twenty-five percent Papuan ancestry in the region today as evidence that the first humans to reach Remote Oceania, including Polynesia, were derived from population mixtures near New Guinea, prior to the further expansion into Remote Oceania2–5. However, our finding that the ancient individuals had little to no Papuan ancestry implies later human population movements that spread Papuan ancestry through the South Pacific after the islands’ first peopling.
Pacific islanders today derive from a mixture of two highly divergent ancestral populations3. One arrived in island southeast Asia more than 40,000 years before present (BP), and contributed to the ancestry of both indigenous Australians and Papuans, and hence to other Pacific islanders4. The second ancestral population is more closely related to mainland East Asians4, and is not found in unadmixed form today. The first humans to reach Remote Oceania—a term we use to refer to the region unoccupied prior to ~3,000 BP beyond the main Solomon Islands and in this case excluding Micronesia—were associated with the Lapita culture that spans 3,450-3,250 to 2,700-2,500 BP. These people spread into Remote Oceania using the first boats capable of long-distance sea travel, introduced new domesticated animals and plants, and their successors reached the most isolated islands of the eastern and southern Pacific by 1,000-700 BP6. Several hypotheses have been proposed to explain why present-day indigenous people of Near Oceania (New Guinea, the Bismarck Islands, and the Solomon Islands area) and Remote Oceania have ancestry both from Papuans and from populations of ultimate East Asian origin. In one set of models that has been favored by recent genetic studies3–5,7, the mixture occurred >3,000 BP during the expansion of populations of East Asian origin through the New Guinea region8. In the other set of models, the population of ultimate East Asian origin initially mixed little with Papuans9 and it is later gene exchanges that account for the ubiquitous Papuan ancestry today2,10.
We obtained genome-wide ancient DNA data from three individuals from the Teouma site on Efate island, Vanuatu (Supplementary Information section 1), all directly radiocarbon dated to between 3110-2740 BP overlapping the Lapita period (Extended Data Table 1). We also obtained genome-wide ancient DNA data from an individual from the Talasiu site on Tongatapu island, Tonga, directly radiocarbon dated to 2680-2340, a period spanning the late Lapita and immediately post-Lapita period (Supplementary Information section 2; Extended Data Table 1). In dedicated clean rooms, we prepared powder from petrous bones11, extracted DNA12, and prepared up to four double-stranded libraries from each extract13. We enriched the libraries for 1.24 million targeted single nucleotide polymorphisms (SNPs)14, sequenced the products, and restricted to a single randomly drawn sequence for each SNP. This procedure resulted in 139,461–231,944 SNPs covered at least once in each of the individuals. The low ratio of sequences aligning to Y chromosome targets compared to targets on other chromosomes15 reveals that all four individuals are females (Extended Data Table 1). We obtained three mitochondrial DNA sequences (all from the Vanuatu site) and all were haplogroup B4a1a1a, the classic “Polynesian motif.”16
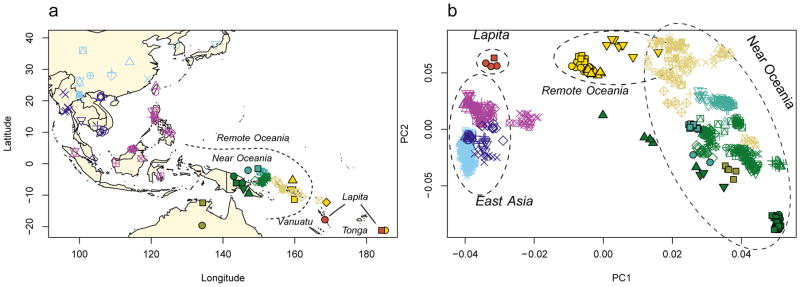
Multiple features of the data suggest authentic and minimally contaminated DNA. First, in all individuals, ~40% of all sites that are cytosines in the human reference sequence appear as thymines in the terminal nucleotide, as expected for genuine ancient DNA (Extended Data Figure 1A). Second, when we carried out Principal Component Analysis (PCA) (Figure 1) of 778 present-day people from 83 East Asian and Oceanian populations genotyped at 621,799 SNPs (Extended Data Table 2), and projected the ancient individuals, we found that all clustered tightly with each other and with data from the same individuals restricting to sequences with cytosine-to-thymine changes at the terminal nucleotide (these sequences are unlikely to be contaminants17,18) (Extended Data Figure 1B). Third, the cluster of ancient individuals does not overlap with present-day populations, indicating that the data are from a population that is not present in unmixed form today (Figure 1). The distinctiveness of the ancient individuals is also highlighted by their high differentiation from all present-day groups (0.05<FST<0.26 between all modern individuals and the ancient Vanuatu individuals) (Extended Data Table 3).
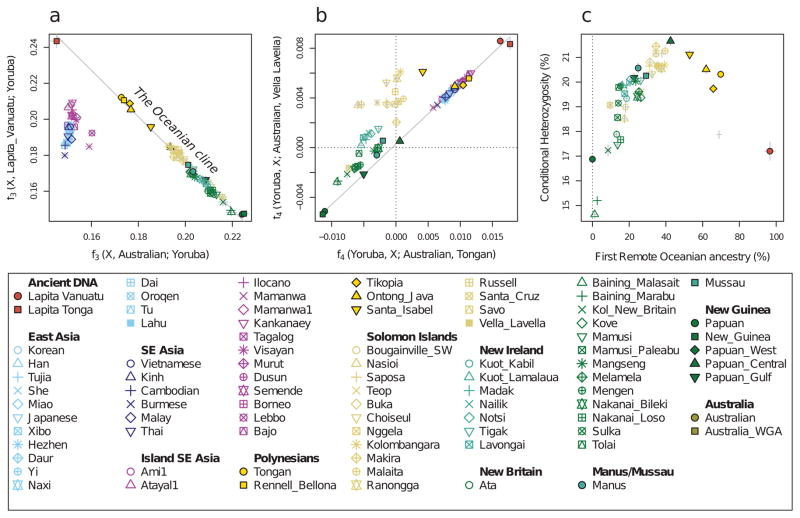
Figure 1. Data from ancient and present-day populations.
a, Locations of 778 present-day individuals genotyped on the Affymetrix Human Origins Array and 4 ancient individuals. b, Ancient individuals projected onto principal component 1 and 2 computed using only present-day samples. Individual population labels are given in Extended Data Figure 2.
The ancient Vanuatu and Tongan individuals are not shifted in the PCA in the direction of Papuan ancestry, in contrast to all present-day Remote Oceanians. In this respect, they are similar to indigenous Taiwanese such as Ami and Atayal as well as to populations from the Philippines such as the Kankanaey that have no detectable Papuan ancestry (Figure 1). To test if the ancient individuals have any evidence of Papuan ancestry, we used qpWave/qpAdm to analyze allele frequency correlation statistics19. The ancient individuals and the Taiwanese Ami are consistent with descending from a common ancestral population to the exclusion of 14 worldwide outgroups (P ≫ 0.05 for the ancient individuals from both Vanuatu and Tonga). We estimate the possible range of Papuan ancestry in the Vanuatu individuals to be 0–11% and in the Tongan individual to be 0–17% (99% confidence intervals truncated at zero), significantly lower than the >25% Papuan ancestry in all present-day Oceanians (Figure 2A). To test the hypothesis that the ancient Remote Oceanian individuals might be from the source population of the non-Papuan ancestry in Oceanians today, we computed the statistic f4(Africa, Test; Australian, Polynesian), which evaluates the degree of allele sharing of a candidate Test population with Polynesians (at sites where Polynesians differ from Australians), and found that it is maximized when Test=Lapita_Vanuatu or Test=Lapita_Tonga (Extended Data Figure 2B), as expected if Lapita were the true source. We conclude that the non-Papuan ancestry that is ubiquitous in Oceania is derived from a population related to the ancient individuals we analyzed, and that this ancestry reached uninhabited islands in Remote Oceania with little or possibly no mixture with Papuans. We call the population of which both the ancient Vanuatu and Tongan individuals were a part the ‘First Remote Oceanians’, and find that the ancestry fraction from this population is the single most important factor shaping genetic variation among Pacific islanders, accounting for most variation in measurements including genetic diversity (Pearson’s R=0.86, P=2×10−12 for 42 non-Polynesian groups) (Extended Data Figure 2) and proportion of archaic Denisovan ancestry (R=−0.96, P < 10−16 for all 56 Oceanian groups) (Figure 2).
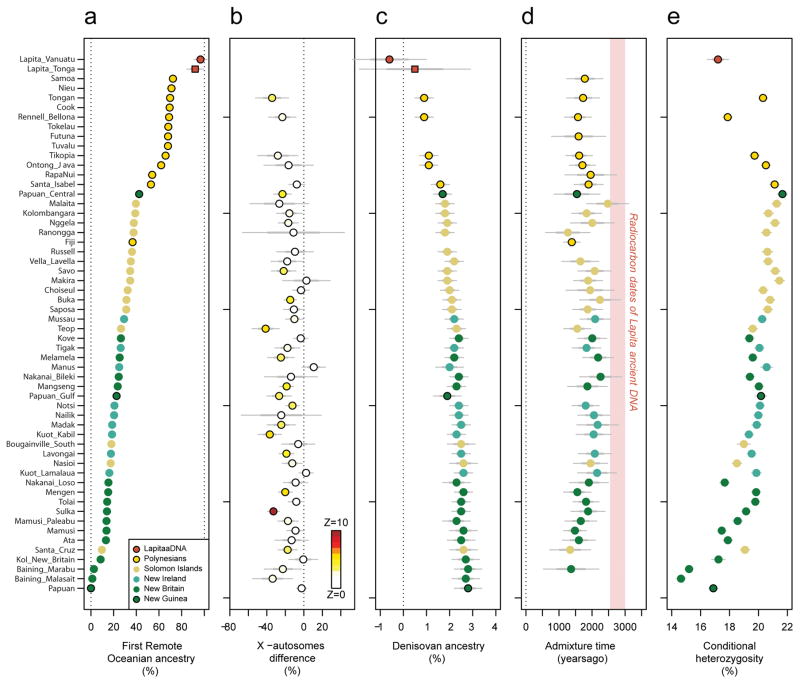
Figure 2. Genetic characteristics of the Oceanian ancestry cline.
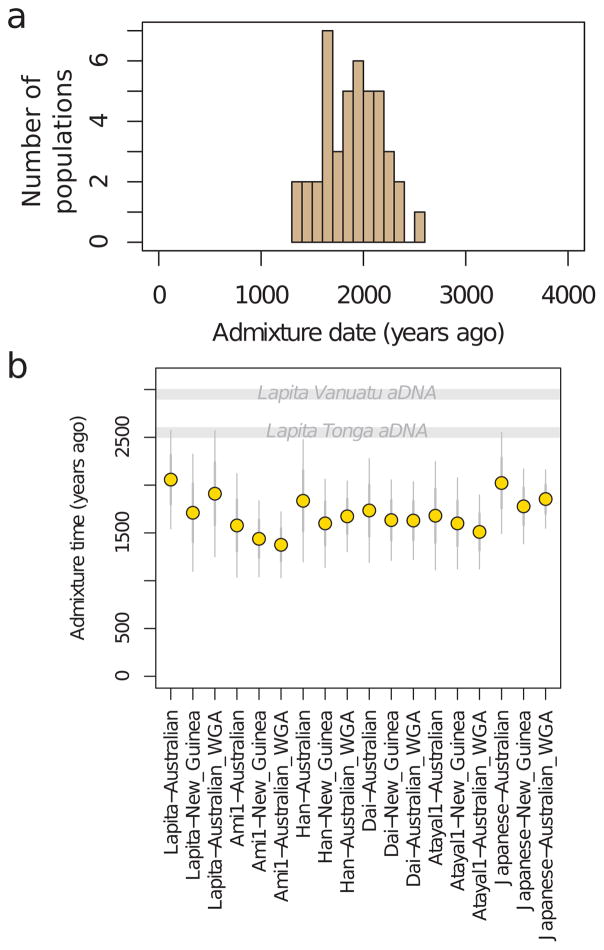
a, Estimated proportion of First Remote Oceanian ancestry. b, Difference between First Remote Oceanian ancestry estimates on chromosome X and the autosomes. c, Denisovan ancestry estimates are inversely related to First Remote Oceanian ancestry estimates. d, Estimated date of admixture in all populations with at least 4 individuals and significant evidence of decay of weighted admixture linkage disequilibrium as measured in ALDER. We used Han and New Guinean Highlanders as surrogates for the ancestral populations. We assumed a generation interval of 28.1 years, and show 95% confidence intervals (thin whiskers) incorporating uncertainty both in the ALDER date and the value of the human generation interval. We show the range of radiocarbon dates for the ancient individuals. e, Conditional heterozygosity (genetic diversity) estimated by drawing two random chromosomes from different individuals at each locus, using only SNPs ascertained in a single Yoruba, and restricting to transversion SNPs to avoid any concerns about inflated heterozygosity due to ancient DNA degradation. Thick and thin error bars in all five panels correspond to 1, and 1.96 standard error of the estimate.
Our evidence that early and geographically diverse Remote Oceanian individuals had little if any Papuan ancestry contradicts models that suggest significant Papuan contributions to Lapita people prior to their dispersal into Remote Oceania3–5. Instead, our results show that the Papuan genetic signature only appeared in many Remote Oceanian populations subsequent to initial settlement. To gain further insight into when the Papuan ancestry may have become ubiquitous in Remote Oceanians, we leveraged the fact that chromosome segments from ancestral populations break up at a known rate due to recombination, and that the inverse of the length of the segments translates to a date of mixture20. We estimate dates of ~50–80 generations ago using ALDER21,, or 1,500–2,300 BP assuming 28.1 years (Methods) per generation22 (Figure 2D; Extended Data Figure 3). We combined the statistical error of the genetic estimate and the uncertainty about the generation interval, and obtained a 95% confidence interval of 1239–1927 BP for a pool of Polynesians all of whom have similar Papuan ancestry proportions (Methods). This finding that Papuan-First Remote Oceanian mixture continued long after the Lapita period implies that the Polynesian ancestral population was not fully formed at that time, although we caution that alternative methods for dating Papuan mixture in Remote Oceanians arrived at older dates4,23–25. However, our ALDER dates are supported by direct ancient DNA evidence, as the Tongan individual at 2680-2340 BP carried little or no Papuan ancestry, providing unambiguous confirmation that the ancestral population of Polynesians was not fully formed by the end of the Lapita period.
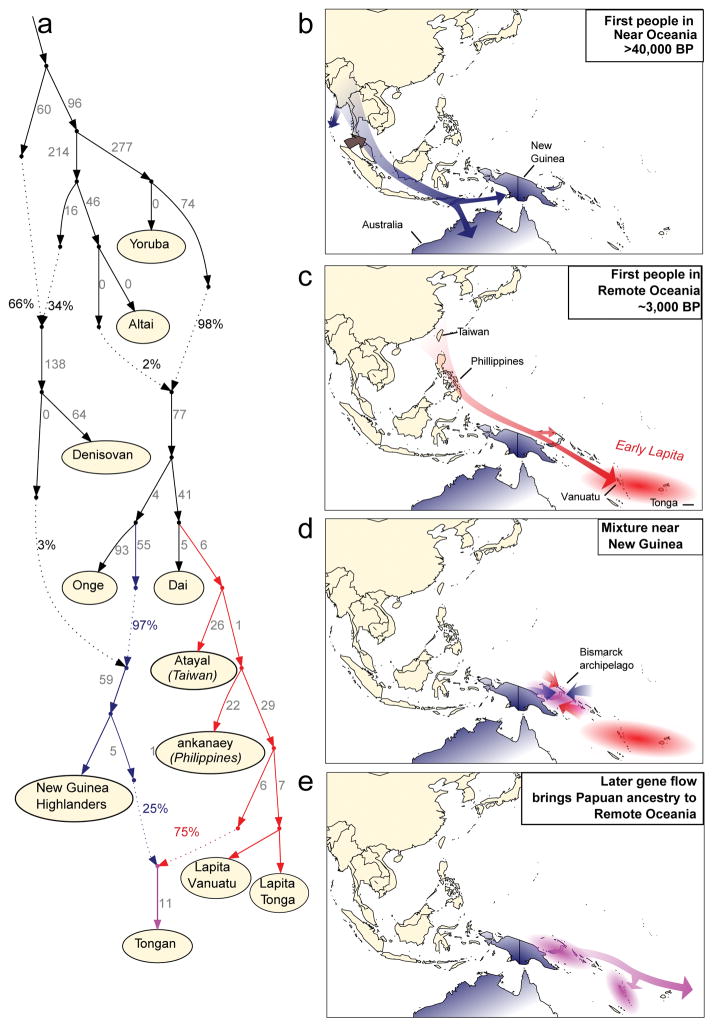
We used qpGraph to explore models of population separation and mixture that might accommodate the ancient DNA data26 (Supplementary Information section 3). We obtain fits using models in which Polynesians today are mixtures of First Remote Oceanians and a Papuan population related to Highland New Guineans (Figure 3A). We also obtained consistent findings using TreeMix27 (Extended Data Figure 4). In Figure 3 we show the best fitting model, which suggests that the ancient individuals from Vanuatu and Tonga descend from an ancestral (presumably Lapita) population that separated earlier from the population that is the primary component in present-day Polynesians. This implies that not just Papuan ancestry but also deeply branching First Remote Oceanian ancestry was introduced to Remote Oceania through later movement of people. Thus, the minimum 25% Papuan ancestry seen in present-day Remote Oceanians is a conservative underestimate of the later population displacement. It is unlikely that there was 100% replacement, however, since we observe weak excess affinity of present-day Tongans to Lapita_Tonga in symmetry tests (Methods). More deeply in time, our modeling indicates that Philippine populations (Kankaney) are the closest outgroup to the First Remote Oceanians, indigenous Taiwanese (Atayal) second closest, and mainland southeast Asians such as Dai most remote, consistent with models of population movement along a route from Taiwan to the Philippines to Near Oceania to Remote Oceania28. We were surprised that we could not fit Australians as outgroups to New Guinean Highlanders and the Papuan ancestry in Polynesians (Extended Data Figure 5). However, we could fit Australians as deriving from a mixture of an ancient Australian lineage and a Papuan lineage from the same group that expanded into Polynesia. This is plausible if there was continuing gene flow between New Guinea and Australia. Another parsimonious model is that the ancestry in present-day Polynesians is not all Papuan, but a Papuan-Australian mix.
Figure 3. A model of population history.
a, A model of population relationships incorporating admixture that fits the allele frequency patterns (all empirical f-statistics within 3 standard errors of expectation). Branch lengths are shown in units of FST × 1000. b, A model of population movements more than 40,000 years ago in which modern humans arrived in the Australia-New Guinea region (blue shading) and mixed with archaic Denisovans (brown shading). c, A model of events prior 3,000 years ago, in which the First Remote Oceanian population formed by spread of a population of ultimate East Asian origin to a region including Vanuatu and Tonga, and experienced little or no mixture with the Papuans they encountered along their journey (red shading). Note that geographic routes are speculative. d, A model of populations of mixed Papuan-First Remote Oceanian ancestry in Near Oceania after 3,000 years ago in a patchwork of islands with different proportions of First Remote Oceanian ancestry (pink shading). e, A model of secondary expansion of admixed populations bringing Papuan ancestry into Remote Oceania, which was still not complete in Tonga by the date of the Talasiu individual at 2680-2340 BP.
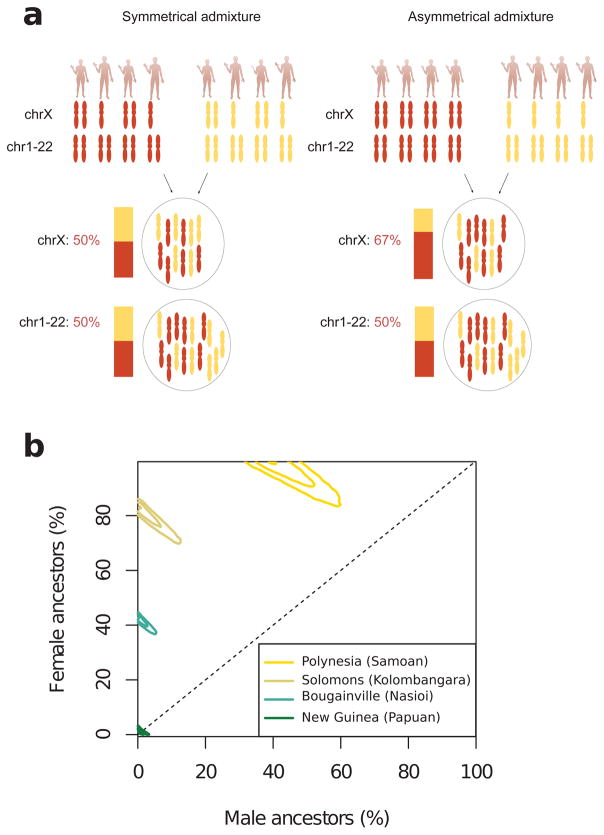
Previous studies of mitochondrial DNA and Y chromosomes suggested that present-day people of the South Pacific harbor more East Asian ancestry from female than from male ancestors3. Our genome-wide analyses confirm a significant excess of First Remote Oceanian ancestry on the X-chromosome compared to the autosomes (Z-scores up to 10) (Figure 2B). Females carry 2/3 of X chromosomes in a population but only 1/2 of the autosomes (Extended Data Figure 6), and we compared the ancestry estimates in these two parts of the genome to obtain the most accurate estimates of sex-biased admixture in diverse Oceanians to date (Extended Data Figure 6; Extended Data Table 4). It has been suggested that matrilocal social structure in the primarily First Remote Oceanian ancestry populations of the region is one likely factor explaining these patterns29,30. However, it is also possible that some of these patterns reflect a scenario in which the later movement of Papuan ancestry into Remote Oceania was largely mediated by males who then mixed with resident females.
Our study has documented that many of the first humans in Remote Oceania had little, if any, Papuan ancestry, a stark contrast to the situation today. While our findings cannot rule out the possibility that multiple groups—some of which carried substantial amounts of Papuan ancestry—settled Remote Oceania early on, the lack of such ancestry both in Vanuatu and Tonga can be more parsimoniously explained by later population movements bringing the Papuan ancestry. The scenario emerging from ancient DNA analysis is thus radically different from that suggested by previous genetic studies, which have generally posited that the first people in Remote Oceania and Polynesia2–5 had substantial Papuan ancestry. Our finding of major post-Lapita movements of Papuan ancestry into Remote Oceania also cannot be related to the later arrival of Papuan ancestry that has been suggested for Fiji, which is estimated to have occurred at least a millennium later at 500 BP4 or 1,100 BP24 (Figure 2). Systematic study of ancient DNA from throughout Remove Oceania should make it possible to provide a detailed chronicle of the population movements and sex-biased population mixtures that shaped the ancestry of present-day Oceanians.
Online Content Methods, along with any additional Extended Data display items and Source Data, are available in the online version of the paper; references unique to these sections appear only in the online paper.
METHODS
Ancient DNA sampling, extraction, library preparation, enrichment and sequencing
The Vanuatu skeletal samples B30A, B10B, B17 were analyzed with permission from the Vanuatu National Museum and the excavators of the Teouma site. The Tonga skeletal sample SK10 was analyzed with permission from the excavators of the Talasiu site.
All preparation of skeletal samples, DNA extraction, and library preparation was carried out in dedicated ancient DNA laboratories at University College Dublin, Ireland (sample preparation of the three Lapita_Vanuatu individuals), at Harvard Medical School in Boston, USA (DNA extraction and library preparation of the three Lapita_Vanuatu individuals), and at the Max Planck Institute for the Science of Human History in Jena, Germany (sample preparation, DNA extraction and library preparation of the the Lapita_Tonga individual). Each of these facilities is spatially separated from other molecular biology laboratories, and measures are taken to protect ancient individuals from contamination including HEPA filtered air, head-to-toe suits, face masks with visors, multiple layers of gloves, bleaching of all surfaces, ultraviolet light (UVC) decontamination of (non-sensitive) consumables and chemicals, and UVC decontamination of the facility when researchers are not in the room31. The final step of the library preparation (amplification) was performed outside the ancient DNA lab.
We prepared powder from the cochlea of petrous bones, extracted DNA12, and prepared libraries with standard protocols (ref. 13 for the Lapita_Vanuatu individuals and ref 32 for the Lapita_Tonga individual). For the three Lapita_Vanuatu individuals, the first library was prepared in the presence of uracil DNA glycosylase (UDG) to cut out errors due to ancient DNA damage, whereas the remaining three libraries as well as the Lapita_Tonga library were prepared without UDG as this preserves more DNA for any given sample. We performed in-solution enrichment using previously reported protocols13,14,33,34 for a targeted set of 1,237,207 SNPs that comprises two previously separately reported sets of 394,577 SNPs34 (390k capture), and 842,630 SNPs14. We sequenced the product on an Illumina NextSeq500 instrument for 2×75cycles. Following demultiplexing, and, for the Lapita_Vanuatu samples removal of both oligonucleotide barcodes that were used to identify the libraries and trailing adapter sequences, we merged the forward and reverse reads of each read pair requiring a 15 base pair overlap (allowing one mismatch). We then aligned merged sequences to the human genome hg19 using BWA 0.6.135. We removed sequences aligned to identical outer coordinates, choosing the highest quality sequence for each duplication cluster. We merged the data from the four libraries for each Lapita_Vanuatu individual.
Genomic analysis
We determined sex by comparing the number of X and Y chromosome alignments15. We estimated damage patterns using PMDtools v0.5918, separating damage patterns observed inside and outside a CpG context. Since all four individuals were female, we could not estimate contamination using X-chromosome data. We investigated whether there was evidence of excess relatedness between any pair of individuals among the Lapita_Vanuatu individuals, but found that the pairwise mismatch rate using panel 5 of the Affymetrix Human Origins array (see below) was 19.8%±0.4% for I1368/I1369, 19.7%±0.6% for I1368/I1370, and 20.5%±0.4% for I1369/I1370. This suggests no atypical pair of individuals and a similar within-population mismatch rate as some present-day Polynesian populations (Figure 2).
Genotyping of present-day humans
We genotyped 356 individuals from 38 southeast Asian and Oceanian populations on the Affymetrix Human Origins array (Extended Data Table 2). The individuals all contributed DNA samples voluntarily and provided informed consent consistent with studies of human genetic variation and history. Ethical approval of the component studies was provided by the Singapore Health IRB, the Research Ethics Committee at the Facultés de Médecine de Toulouse, the Brunei Medical and Health Research Ethics Committee, the University of Cambridge Biology Research Ethics Committee, the Government of Papua New Guinea Medical Research Advisory Committee, and the Temple University IRB. The collection of genome-wide variation data on de-identified samples was approved by the Harvard Human Research Protection Program (Protocol 11681), re-reviewed on 12 July 2016.
We restricted analysis to samples that had >95% genotyping completeness and that were not visual outliers in PCA with respect to the main cluster of samples in the group. We merged with previously reported Affymetrix Human Origins SNP array data26,36–39. We also co-analyzed our data with samples genotyped on the Affymetrix 6.0 platform where we removed three previously published39 Rapa Nui individuals (5s5j, XB3B, and 3p3p), and two previously published40 Samoan individuals (PLY_07 and PLY_11), that all appeared to have recent European ancestry based on clustering analyses. We finally compared our data to high-coverage genomes from an archaic Neanderthal and an archaic Denisovan, both from Denisova Cave in the Altai Mountains of Siberia41–43.
Population genetic analysis
When overlapping with the Affymetrix Human Origins SNP array data set of present-day human populations, we have between 74,000–126,000 SNPs covered at least once for each of the four individuals (Extended Data Table 1). This is more than the minimum coverage required for high resolution analysis using allele frequency correlation statistics, which is 10,000 SNPs per individual according to Supplementary Information section 6.2 of ref. 44, a study that had the same median coverage (0.19x) as ours (the range in the present study is 0.14–0.26x). For all analyses, we called genotypes by randomly sampling a single non-duplicate sequence read at each position45. This procedure is standard for analysis of low-coverage ancient DNA data and is also often used for higher-coverage data to minimize reference genome biases that can be introduced when determining diploid genotypes14,17,34,36,41,44–50. For the qpAdm, qpWave and qpGraph analyses we excluded transition SNPs to avoid biases from postmortem damage (see below).
We performed PCA using smartpca51, with the option inbreed: YES in order to sample a single genotype from each individual randomly to match the pseudo-haploid nature of the ancient DNA genotypes from the ancient individuals52. We computed f3-, f4-, D-statistics, and FST as in ref. 26, randomly sampling a single haploid chromosome for each individual, using popstats38. We estimated the date of admixture using ALDER21. We tested the consistency of a matrix of f4-statistics with one or more sources of ancestry with respect to a set of outgroups (New_Guinea, Denisova, Sardinian, English, Yakut, Chukchi, Mala, Japanese, Ju_hoan_North, Mixe, Onge, Yoruba, and Mbuti) using qpWave19.
For the ancient individuals and all present-day populations genotyped on the Human Origins array, we used qpAdm34, which estimates ancestry proportions from two or more proxy source populations assuming that the proxies are more closely related to the real source populations than they are to a set of outgroups (qpAdm also provides a formal statistical test for whether this is the case, which passes in the context that we use it here). We estimated First Remote Oceanian and Papuan ancestry using Denisova, Sardinian, English, Yakut, Chukchi, Mala, Japanese, Ju_hoan_North, Mixe, Onge, Yoruba, and Mbuti as outgroups and New_Guinea and Ami as proxies for the source populations. For the ancient individuals, we excluded all transition SNPs to avoid possible biases due to postmortem damage, resulting in 35,194 transversion SNPs for Lapita_Vanuatu and 22,030 for Lapita_Tonga. For estimating ancestry proportions in the Affymetrix 6.0 Polynesian data, we used whole-genome sequences from the same populations as outgroups53. We estimated Denisovan ancestry using the Denisovan genome and Japanese as the two sources, and chimpanzee, Ju_hoan_North, Mbuti, Yoruba, Dinka and the Altai Neanderthal genome as outgroups.
We computed conditional heterozygosity using panel 5 of the Affymetrix Human Origins array, which contains SNPs ascertained as heterozygous in a single West African Yoruba individual. This provides an unbiased estimate of relative heterozygosity since the Yoruba individual is approximately symmetrically related to all Oceanians (Denisovan ancestry violates this assumption but should not change the ranking of populations). We estimated heterozygosity as the average pairwise mismatch rate when sampling 2 chromosomes from 2 different individuals using popstats38, restricting to transversion SNPs for all populations.
For authentication, we used PMDtools18 to extract sequences with clear evidence of postmortem damage patterns (PMD score of at least 3), disregarding individual bases with phred-scaled base quality < 30. We randomly sampled new haploid genotypes from the resulting set of sequences and projected the ancient individuals on the principal components inferred from the present-day populations as above. After this filtering, we retained 68,450 SNPs for I1368; 98,722 SNPs for I1369; 83,024 SNPs for I1370; and 117,023 SNPs for CP30. Ninety-nine percent confidence intervals for qpAdm estimates of Papuan ancestry (see above) using the PMD score-restricted data was 0–21% for Lapita_Vanuatu and 0–24% for Lapita_Tonga, consistent with the confidence intervals obtained from the full data.
To test whether the Lapita_Vanuatu and the Lapita_Tonga form a clade, we used qpWave to test a model of Dai, Ami, Kankanaey and a fourth population were consistent with being outgroups to the two ancient sample groups (we used Dai, Ami and Kankanaey as these span present-day Mainland East Asia, Taiwan, and the Philippines, and lack Papuan ancestry to the limits of our resolution). The analysis used the ~12,000 SNPs that remained after excluding transition SNPs and SNPs missing in one of the two ancient sample groups. We found that the model was consistent with the data for all tested Oceanian and Asian populations shown in Figure 1, but that the lowest P-value was observed for present-day Tongans (P = 0.09). We also found that f4(Ami, Tongan; Lapita_Vanuatu, Lapita_Tonga) = 0.006, Z=3.2, when using all SNPs. This documents an affinity between present-day Tongans and Lapita_Tonga, consistent with the hypothesis that the ancient population of Tonga with little Papuan ancestry may have contributed some of the ancestry of present-day Tongans.
Admixture date estimation
To estimate the date of historical admixture between First Remote Oceanians and Papuans, we used ALDER21,25 on the full Human Origins array data, with New Guinean Highlanders and Han Chinese as the two sources. We use Han Chinese for this analysis due to their substantial sample size compared to populations more closely related to the ancestral First Remote Oceanian population such as the ancient individuals we analyzed, indigenous Taiwanese, and indigenous Philippine groups. ALDER estimates are robust even when using an imperfect surrogate for the ancestral populations26. We estimate an admixture date for a pool of Polynesian populations by combining data from Tongans, Tikopia, Russell and Bellona, all genotyped on the Affymetrix Human Origins SNP array.
ALDER and other methods based on admixture linkage disequilibrium estimate dates in units of generations, which need to be converted to years. For this purpose we require an estimate of the generation interval—the average age of a parent at the time their gametes were formed—weighted by the fraction of recombination events that occur in each sex (62.3% of all autosomal crossovers are estimated to occur in females based on Table 1 of ref. 54). Using estimates from the anthropological literature, this quantity is 27.8 years for hunter-gathering societies, 28.6 years for developed nation states, and 29.6 years for less developed nation states22. These numbers are in the range of the point estimate we use of 28.1 years based on breakdown of admixture linkage disequilibrium in radiocarbon-dated ancient genomes55 (Methods). To account for the substantial variability in generation intervals across human societies, we use the sample standard error of 2.15 years measured across eleven diverse hunter-gatherer groups based on Table 4 of ref. 22. The date estimates in Figure 2 and Extended Data Figure 4 thus use a generation interval of 28.1 years, and combine the standard error from ALDER (a) with the uncertainty in generation time, that is, , where A is the ALDER point estimate in number of generations.
We do not subtract 66 years from the dates produced by ALDER to obtain BP dates (conventionally the date before 1950 CE, 66 years ago), because what ALDER is estimating is a number that is close to the BP date. To see this, note that ALDER estimates the date between when chromosomes of the two ancestries began crossing over (one generation after mixing began), and the date of the last cross-over (when the germ cells that mixed to produce the present-day samples in our study were formed, likely one or two generations prior to 2016 CE). Accounting for these corrections means that ALDER is estimating a date of mixture that is likely to be within a generation of the true BP date.
Fitting models of population history
We used qpGraph26,56 to assess the fit of admixture graph models to allele frequency correlation patterns as measured by f2, f3-, and f4-statistics. We started with a skeleton phylogenetic tree consisting of Yoruba, New_Guinea, Dai, Atayal, Kankanaey and Lapita_Vanuatu. We added Tongan, Mamanwa (a Philippine Negrito group), Nasioi and Kolombangara, respectively, to all possible edges in the tree, and retained only the graph solutions that provided no individual f4 statistics with |Z| > 3 between empirical and predicted statistics. For the extended version of the admixture graph, we also added Australians to all possible edges of the graph that included these populations. Finally, we modeled the previously documented admixture history relating Denisovans and the Altai Neanderthal genome to the outgroup chimpanzee and the anatomically modern human populations, to which we added the Andamanese Onge and the Lapita_Tonga. The final graph visualized in Figure 3 used 10,893 SNPs after restricting to transversion SNPs to avoid complications due to ancient DNA damage and also SNPs with coverage in all groups. For more information on the admixture graph inference procedure, see Supplementary Information section 3.
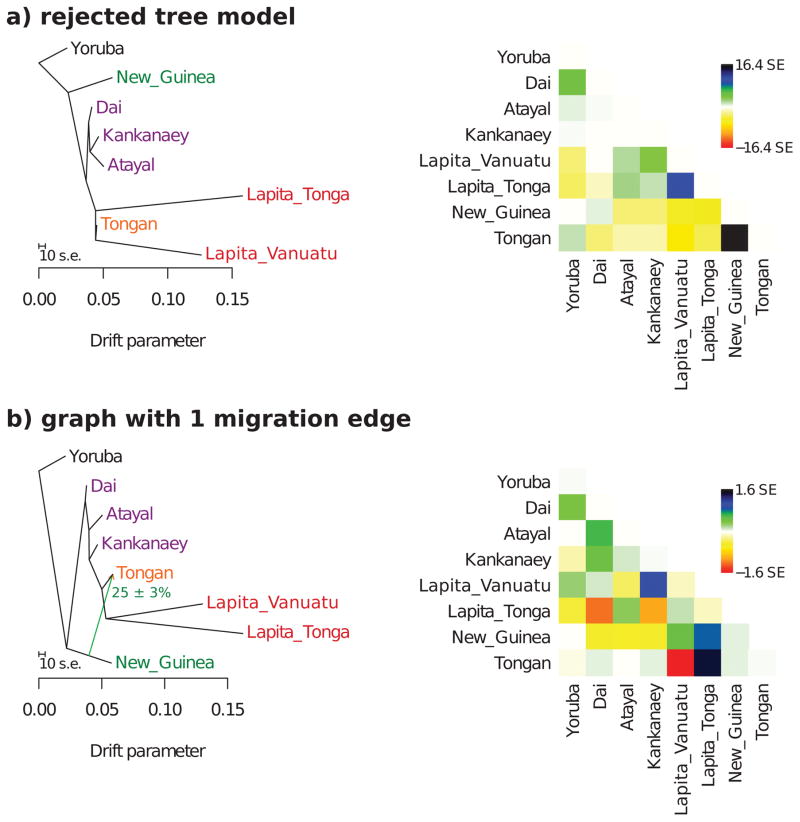
As an alternative inference method, we used Treemix v1.1227 to test models for Yoruba, Dai, Atayal, Kankanaey, Tongan, New Guinean Highlanders, Lapita_Vanuatu and Lapita_Tonga. The total number of SNPs after excluding transitions, SNPs with minor allele count of less than 4 in the selected data, and loci where one population had missing data, was 10,119, which we divided into 337 blocks of 30 SNPs each to estimate the covariance matrix. We first fitted a maximum likelihood tree of all populations, but found that several of the fitted allele frequency covariances deviated from those empirically observed by up to 16.4 standard errors. We then used the automated heuristic optimization in Treemix to infer a graph model with 1 admixture event using the same populations, and found that the optimal fit was for a model with an admixture event in the history of Tongans, where one portion of their ancestry diverged prior to the split of the ancestors of the ancient Vanuatu and Tonga individuals, and the other (25% ± 3%) derived from the New Guinean lineage. This maximum deviation between empirical and model covariances observed for the graph with 1 admixture edge was 1.6, indicating good fit, consistent with our investigation of models using qpGraph.
Female and male ancestral contributions
To estimate the proportion of female ancestors (F) and male ancestors (M) for a given population, we used two different methods both based on the point estimates of ancestry for the X-chromosome and autosomes. Both used the same underlying model that the observed admixture proportion estimates Ĥauto and ĤX for the autosomes and X-chromosome, respectively, depend on M and F such that:
| (1) |
| (2) |
The first approach obtains unbounded point estimates of M and F by rearranging equations:
| (3) |
| (4) |
Similarly, we obtained standard errors for M and F using the weighted block jackknife standard errors for Ĥauto and ĤX, SEauto and SEX, as
| (5) |
| (6) |
As an alternative to estimating M and F, we took an approximate Bayesian approach by performing 1 million simulations where M and F were sampled from a uniform prior distribution (0, 1). We then simulated ancestry estimates specifying normal distributions with means and standard errors matching the empirical values (Equations 1–2). We used the abc R package57 to run a rejection algorithm retaining the 1% of all simulation replicates with the closest Euclidean distances to the empirical Ĥauto and ĤX, and performed local linear regression on log-transformed summary statistics to obtain a posterior distribution. The results of the two methods are qualitatively similar. In Extended Data Figure 6, we plot the posterior intervals of these distributions for selected populations.
Sample size
No statistical methods were used to predetermine sample size.
Extended Data
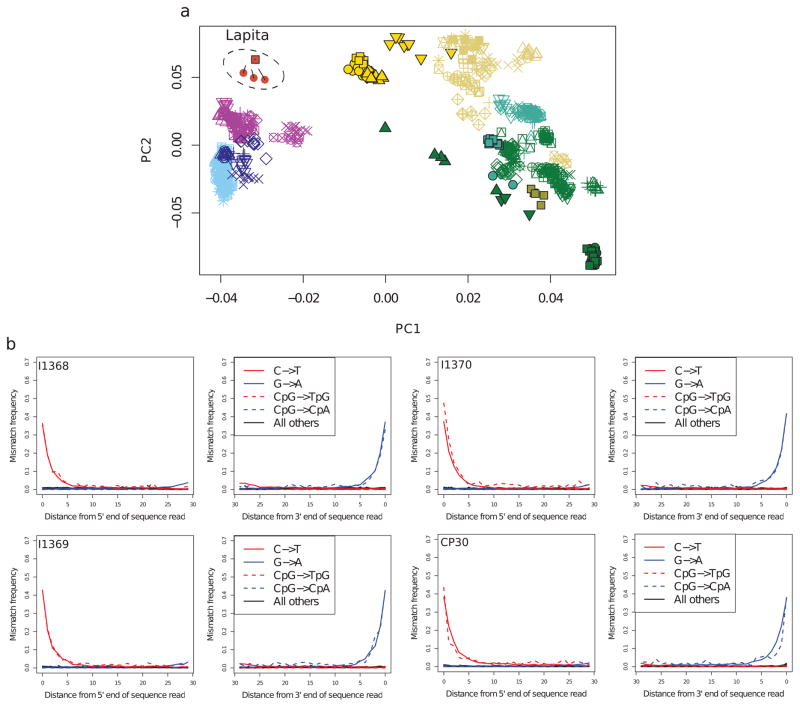
Extended Data Figure 1. Ancient DNA authenticity.
a, Principal component analysis performed as for Figure 1, but with the four ancient individuals represented only by sequences that show clear evidence of postmortem damage (PMD score of at least 3) to remove contaminating sequences that might be present17,18. The numbers of SNPs remaining after restricting to damaged sequences is 68,450 SNPs for I1368; 98,722 SNPs for I1369; 83,024 SNPs for I1370; and 117,023 SNPs for CP30. The lines indicate the projection of the samples when no damage-restriction is performed. The large number of SNPs retained, and the fact that the ancient individuals cluster tightly and have the same qualitative positioning in the plot as Figure 1, indicates that contamination is not contributing to our findings. We also find that estimates of Papuan ancestry using PMD score restricted data is consistent with those obtained using the full data (Methods). b, Postmortem damage patterns for genome-wide in-solution enrichment data from four ancient individuals.
Extended Data Figure 2. f-statistics document the Oceanian ancestry cline.
a, Shared genetic drift with Lapita_Vanuatu is negatively related to shared drift with Australians. Except for Lapita_Tonga, populations from Taiwan, the Philippines and Polynesia share the most genetic drift with the Lapita_Vanuatu, who are not shown in the plot because they are used as reference in the computation. The trend line was fitted without the East Asian populations in the off-cline cluster. b, The Lapita_Vanuatu and Lapita_Tonga maximize statistics of the form f4(Yoruba, Test; Australian, Oceanian), suggesting that they are the most closely related to the East Asian ancestry in Oceanians of any sampled population. The trend line was fitted using populations >0.005 on the x-axis, together with the two populations with the lowest values on the x-axis (Papuan and New_Guinea). c, biplot of First Remote Oceanian ancestry proportions against conditional heterozygosity. Populations with intermediate admixture proportion show the greatest genetic diversity. Thick and thin error bars in all panels are 1, and 1.96 standard errors of the estimate, respectively.
Extended Data Figure 3. Admixture date estimates.
a, histogram of the point estimate dates in Figure 2d. b, Admixture date estimates for Tongans using different pairs of source populations (“Lapita” in this figure refers to Lapita_Vanuatu). Error bars show 1 (thick whiskers) and 1.96 (thin whiskers) standard errors, respectively. (“WGA” refers to whole-genome amplified DNA.)
Extended Data Figure 4. Admixture graph inferred using Treemix.
a, A simple tree-like model without admixture fits the data poorly, as can be seen from the matrix of residuals between empirical and modeled allele frequency covariance on the right. b, The optimal placement of a single 25% admixture event is from the lineage related to New Guinean Highlanders into the lineage leading to Tongans. Tongans derive the other portion of their ancestry from the lineage leading to the two ancient groups of individuals. This graph has no significant deviations between empirical and modeled allele frequency covariances.
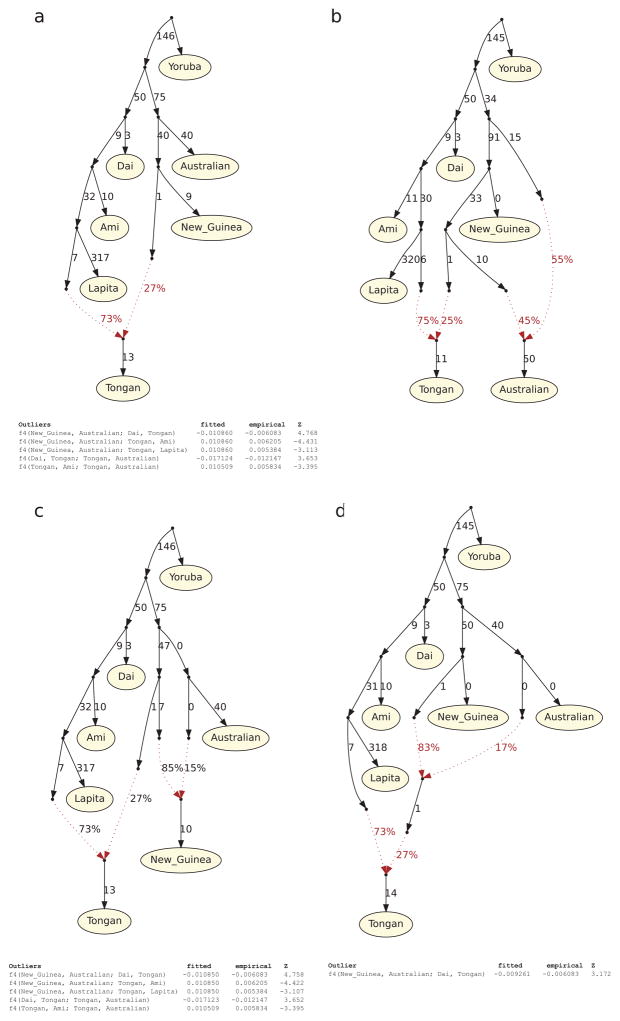
Extended Data Figure 5. Admixture graphs modeling the population history of Australians.
Outlier f4-statistics are shown (|Z| > 3). a, A model with a single admixture edge positing that Australians are an outgroup to the Papuan ancestry in Tongans does not fit the data (5 outlier statistics). b, An alternative model with 2 admixture edges where the Papuan ancestry in Tongans also contributed to Australians fits the data (no outliers). c, A model with 2 admixture edges where New Guinean Highlanders are admixed from an Australian source after the divergence of the Papuan source in Tongans does not fit the data (5 outliers). d, A model with 2 admixture edges where the Papuan ancestry in Tongans is intermediate between the New Guinean Highlander lineage and the Australian lineage. Branch lengths are in units of FST × 1000. Lapita in this figure refers only to Lapita_Vanuatu, which is the only group for which we have multiple individuals (needed to compute FST).
Extended Data Figure 6. First Remote Oceanian ancestry today comes primarily from females.
a, Illustration of the rationale for using the X-chromosome to study asymmetrical admixture between males and females. The example on the left illustrates admixture with equal proportion of males and females in both the red and the yellow ancestral population. The example on the right illustrates an extreme case of asymmetrical admixture where the red ancestral population only contributes females and the yellow ancestral population only contributes males to the admixed generation, demonstrating the disproportional contribution of X chromosomes by females to the admixed population. b, Female and male ancestral contributions based on an admixture model fitted to estimated ancestry proportions on the autosomes and X-chromosome. We show the 95%, 70%, and 5% highest posterior intervals for four selected populations from Polynesia (Samoans), the Solomon Islands (Kolombangara), Bougainville (Nasioi), and mainland New Guinea (Papuans).
Extended Data Table 1.
In-solution DNA enrichment and sequencing of ancient individuals.All dates are calibrated using OxCal v4.2.458 with a mixture of the Marine13 and Intcal13 curves59 as determined by linear interpolation between dietary terrestrial/marine δ13C isotopic endpoints (−21‰/−12‰) with an uncertainty of ±10% on the per-cent marine carbon result following previous recommendations60. Two of the dates have been previously reported (for I1368/B30A and I1370/B17)61, and in this study we add two new dates: for I1369/B10B from Tonga (on the same petrous bone used for ancient DNA analysis) and on CP30/SK10 from Tonga (on a fibula). Measured 13C and 15N values for I1369/B10B are −14.5 and 13.7‰ respectively, and for SK10 −16.44‰ and 10.48‰. As justified in ref. 61, we also applied a location-specific reservoir correction (ΔR) of 40±44 14C years to the marine curve to adjust for regional oceanic variation in 14C around Vanuatu, and 11±83 14C years for Tongatapu62.
| Sample information | Coverage on chromosomes 1–22 | Sex determination | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Location | ID1 | ID2 | Bone for aDNA | Bone for dating | 14C Date: Calibrated 95.4% Conf. Int. (Uncalibrated date, Lab number) | Mean depth | All SNPs | SNPs over-lapping array | Y SNPs | X SNPs | Sex |
| Vanuatu | I1368 | B30A | Petrous | Skull | 2990-2740 BP (2983±32 BP, Wk-22657) | 0.26 | 139,461 | 74,631 | 321 | 18,231 | F |
| Vanuatu | I1369 | B10B | Petrous | Petrous | 3000-2750 BP (3045±30 BP, Poz-81126) | 0.14 | 199,500 | 107,523 | 341 | 24,255 | F |
| Vanuatu | I1370 | B17 | Petrous | Skull | 3110-2780 BP (3083±26 BP, Wk-21026) | 0.21 | 167,311 | 90,402 | 231 | 19,303 | F |
| Tonga | CP30 | SK10 | Petrous | Fibula | 2680-2340 BP (2594±20 BP, Wk-41883) | 0.16 | 231,994 | 125,908 | 75 | 25,943 | F |
Extended Data Table 2.
356 individuals newly genotyped on the Human Origins Array.
| Population | N | Country of origin | Land mass | Language | Lat. | Long. | Co-authors for samples | Protocol Numbers for informed consent | Data distribution |
|---|---|---|---|---|---|---|---|---|---|
| Ata | 8 | Papua New Guinea | New Britain | Papuan | −5.7 | 150.9 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Baining_Malasait | 5 | Papua New Guinea | New Britain | Papuan | −4.47 | 151.9 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Baining_Marabu | 10 | Papua New Guinea | New Britain | Papuan | −4.63 | 152.3 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Bajo | 10 | Indonesia | Sulawesi | Austronesian | −3.97 | 122.59 | M.P.C, P.K., F.-X.R. | 4.13.2013 approval AMIS-UPS Ethics Committee | Signed Letter |
| Buka | 8 | Papua New Guinea | Bougainville | Austronesian | −5.42 | 154.67 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Burmese | 10 | Myanmar | Asia | Sino-Tibetan | 16.41 | 95.89 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
| Dusun | 10 | Brunei | Borneo | Austronesian | 4.71 | 114.67 | T.K., S.A. | MHREC/EDU/2012/3(1) and HBREC.2011.01 | Fully public |
| Ilocano | 2 | Philippines | Luzon | Austronesian | 14.6 | 120.98 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
| Kankanaey | 10 | Philippines | Luzon | Austronesian | 17.07 | 121.03 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
| Kol_New_Britain | 2 | Papua New Guinea | New Britain | Papuan | −5.38 | 151.63 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Kove | 18 | Papua New Guinea | New Britain | Austronesian | −5.47 | 148.95 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Kuot_Kabil | 9 | Papua New Guinea | New Ireland | Papuan | −3.07 | 151.7 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Kuot_Lamalaua | 8 | Papua New Guinea | New Ireland | Papuan | −3 | 151.5 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Lavongai | 15 | Papua New Guinea | New | Austronesian | −2.57 | 150.43 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Lebbo | 8 | Indonesia | Borneo | Austronesian | 1.66 | 117.16 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Madak | 10 | Papua New Guinea | New Ireland | Austronesian | −3.1 | 151.7 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Malay | 9 | Singapore | Asia | Austronesian | 1.35 | 103.82 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
| Mamusi | 20 | Papua New Guinea | New Britain | Austronesian | −6 | 151 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Mamusi_Paleabu | 6 | Papua New Guinea | New Britain | Austronesian | −5.95 | 150.9 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Mangseng | 6 | Papua New Guinea | New Britain | Austronesian | −5.93 | 150.7 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Manus | 2 | Papua New Guinea | Manus | Austronesian | −2.08 | 147 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Melamela | 10 | Papua New Guinea | New Britain | Austronesian | −5 | 151.25 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Mengen | 10 | Papua New Guinea | New Britain | Austronesian | −5.1 | 151.4 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Murut | 10 | Brunei | Borneo | Austronesian | 4.62 | 115.14 | T.K., J.T.S.W. | MHREC/EDU/2012/3(1) and HBREC.2011.01 | Fully public |
| Mussau | 10 | Papua New Guinea | St. Matthias | Austronesian | −1.58 | 149.73 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Nakanai_Bileki | 10 | Papua New Guinea | New Britain | Austronesian | −5.75 | 150.8 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Nakanai_Loso | 7 | Papua New Guinea | New Britain | Austronesian | −5.48 | 150.8 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Nailik | 9 | Papua New Guinea | New Ireland | Austronesian | −2.98 | 151.52 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Notsi | 9 | Papua New Guinea | New Ireland | Austronesian | −3.05 | 151.65 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Saposa | 10 | Papua New Guinea | Bougainville | Austronesian | −5.58 | 154.67 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Southwest_Bougainville | 2 | Papua New Guinea | Bougainville | Papuan | −6.6 | 155.5 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Sulka | 20 | Papua New Guinea | New Britain | Papuan | −4.5 | 152.3 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Tagalog | 5 | Philippines | Luzon | Austronesian | 14.6 | 120.98 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
| Teop | 10 | Papua New Guinea | Bougainville | Austronesian | −5.85 | 155.18 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Tigak | 10 | Papua New Guinea | New Ireland | Austronesian | −2.57 | 150.83 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Tolai | 24 | Papua New Guinea | New Britain | Papuan | −4.31 | 152.14 | F.R.F., J.S.F., G.K., D.A.M. | MRAC.1998.2000.2010 and 99-226.4320 | Signed Letter |
| Vietnamese | 10 | Vietnam | Asia | Austroasiatic | 10.82 | 106.64 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
| Visayan | 4 | Philippines | Mindanao | Austronesian | 9.76 | 125.51 | T.K., J.T.S.W. | 2010/820/A and HBREC.2011.01 | Fully public |
Extended Data Table 3.
f-statistics for populations on the Oceanian cline. Standard error (SE) is shown for FST between each Test population and Lapita_Vanuatu. The Z-score is given for the statistic f3(Lapita_Vanuatu, Australian; Test), where Z < 3 provides significant evidence that the Test is admixed between sources related to Lapita_Vanuatu and Australians.
| FST(Lapita_Vanuatu, Test) | f3(Lapita_Vanuatu, Australian; Test) | |||
|---|---|---|---|---|
|
| ||||
| Test population | Estimate | SE | Estimate | Z-score |
| Baining Malasait | 0.263 | 0.005 | 0.143 | 38.4 |
| Baining_Marabu | 0.249 | 0.004 | 0.115 | 40.1 |
| Papuan | 0.225 | 0.004 | 0.066 | 33.3 |
| Kol_New_Britain | 0.216 | 0.006 | 0.086 | 16.7 |
| Mamusi | 0.204 | 0.004 | 0.059 | 26.5 |
| Ata | 0.197 | 0.004 | 0.050 | 21.6 |
| Nakanai_Loso | 0.194 | 0.004 | 0.055 | 21.9 |
| Mamusi_Paleabu | 0.187 | 0.004 | 0.040 | 17.8 |
| Santa_Cruz | 0.185 | 0.004 | 0.021 | 10.7 |
| Nasioi | 0.178 | 0.004 | 0.028 | 14.7 |
| Bougainville_South | 0.176 | 0.006 | 0.034 | 8.1 |
| Sulka | 0.174 | 0.004 | 0.022 | 13.1 |
| Mengen | 0.168 | 0.004 | 0.017 | 9.1 |
| Tolai | 0.163 | 0.004 | 0.009 | 5.6 |
| Kuot_Kabil | 0.162 | 0.004 | 0.014 | 7.7 |
| Lavongai | 0.160 | 0.004 | 0.009 | 5.2 |
| Kuot_Lamalaua | 0.160 | 0.004 | 0.008 | 4.3 |
| Nakanai_Bileki | 0.153 | 0.004 | 0.013 | 7.0 |
| Melamela | 0.152 | 0.004 | 0.010 | 5.1 |
| Madak | 0.151 | 0.004 | 0.003 | 1.8 |
| Papuan_Gulf | 0.151 | 0.005 | 0.002 | 0.7 |
| Kove | 0.150 | 0.004 | 0.013 | 7.4 |
| Mangseng | 0.145 | 0.004 | 0.003 | 1.7 |
| Nailik | 0.145 | 0.004 | 0.000 | −0.1 |
| Teop | 0.144 | 0.004 | 0.005 | 2.8 |
| Notsi | 0.144 | 0.004 | −0.002 | −1.4 |
| Manus | 0.141 | 0.006 | −0.005 | −1.4 |
| Tigak | 0.141 | 0.004 | 0.002 | 1.2 |
| Mussau | 0.132 | 0.004 | −0.005 | −3.3 |
| Choiseul | 0.123 | 0.004 | −0.007 | −3.7 |
| Saposa | 0.122 | 0.004 | −0.011 | −6.5 |
| Buka | 0.118 | 0.004 | −0.016 | −9.9 |
| Vella_Lavella | 0.112 | 0.004 | −0.016 | −8.6 |
| Ranongga | 0.110 | 0.004 | −0.015 | −8.2 |
| Savo | 0.109 | 0.004 | −0.020 | −12.1 |
| Russell | 0.108 | 0.005 | −0.017 | −6.8 |
| Kolombangara | 0.108 | 0.004 | −0.015 | −8.0 |
| RenBel | 0.106 | 0.004 | 0.035 | 14.0 |
| Gela | 0.103 | 0.004 | −0.024 | −14.3 |
| Makira | 0.101 | 0.004 | −0.024 | −16.1 |
| Malaita | 0.096 | 0.004 | −0.025 | −15.5 |
| Papuan_Central | 0.094 | 0.004 | −0.031 | −18.2 |
| Bajo | 0.082 | 0.004 | 0.022 | 12.8 |
| Isabel | 0.079 | 0.004 | −0.024 | −15.9 |
| Tikopia | 0.077 | 0.004 | −0.003 | −1.3 |
| Ontong_Java | 0.069 | 0.004 | −0.018 | −10.4 |
| Tongan | 0.053 | 0.004 | −0.018 | −9.9 |
Extended Data Table 4.
Ancestry estimates for populations on the Oceanian cline.“Auto.” gives the estimate on the autosomes (chromosomes 1–22). “Diff” gives the difference between the autosome and X chromosome estimates.
| Test | First Remote Oceanian ancestry estimate | Anc. contrib. (method of moments) | Two source model | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Auto. | SE | chrX | SE | Diff. | SE | Z | Male | SE | Female | SE | P-value | |
| Tongan | 69.8% | 1.0% | 104.1% | 8.9% | −34.3% | 9.0% | −3.8 | −33.1% | 27.0% | 172.7% | 22.0% | 0.22 |
| Rennel & Bellona | 68.9% | 1.2% | 92.1% | 7.3% | −23.2% | 7.4% | −3.1 | −0.7% | 22.4% | 138.5% | 32.8% | 0.89 |
| Tikopia | 65.8% | 1.0% | 93.9% | 10.9% | −28.1% | 10.9% | −2.6 | −18.5% | 32.9% | 150.1% | 39.9% | 0.58 |
| Ontong_Java | 61.9% | 0.9% | 78.4% | 13.3% | −16.5% | 13.3% | −1.2 | 12.4% | 40.1% | 111.4% | 12.7% | 0.83 |
| Santa Isabel | 52.9% | 0.9% | 60.5% | 4.2% | −7.6% | 4.3% | −1.8 | 30.1% | 13.1% | 75.7% | 14.2% | 0.87 |
| Papuan_Central | 42.5% | 0.9% | 65.6% | 4.7% | −23.1% | 4.8% | −4.8 | −26.8% | 14.6% | 111.8% | 47.7% | 0.14 |
| Malaita | 39.6% | 0.9% | 66.1% | 15.9% | −26.5% | 15.9% | −1.7 | −39.9% | 47.8% | 119.1% | 20.8% | 0.25 |
| Kolombangara | 39.1% | 1.0% | 54.7% | 6.9% | −15.6% | 7.0% | −2.2 | −7.7% | 21.1% | 85.9% | 16.3% | 0.87 |
| Nggela | 37.8% | 0.9% | 54.6% | 5.4% | −16.8% | 5.5% | −3.1 | −12.6% | 16.6% | 88.2% | 82.5% | 0.59 |
| Ranongga | 37.6% | 1.0% | 48.8% | 27.5% | −11.2% | 27.5% | −0.4 | 4.0% | 82.6% | 71.2% | 29.8% | 0.15 |
| Russell | 36.2% | 1.2% | 45.8% | 9.9% | −9.6% | 10.0% | −1.0 | 7.4% | 30.1% | 65.0% | 26.2% | 0.19 |
| Vella_Lavella | 35.2% | 1.0% | 53.2% | 8.7% | −18.0% | 8.8% | −2.1 | −18.8% | 26.4% | 89.2% | 19.9% | 0.10 |
| Savo | 34.6% | 0.9% | 56.4% | 6.6% | −21.8% | 6.7% | −3.3 | −30.8% | 20.1% | 100.0% | 38.4% | 0.18 |
| Makira | 34.6% | 0.9% | 31.8% | 12.8% | 2.8% | 12.8% | 0.2 | 43.0% | 38.6% | 26.2% | 13.4% | 0.08 |
| Choiseul | 32.4% | 1.0% | 35.5% | 4.4% | −3.1% | 4.5% | −0.7 | 23.1% | 13.8% | 41.7% | 9.8% | 0.20 |
| Buka | 31.3% | 0.9% | 46.0% | 3.2% | −14.7% | 3.3% | −4.4 | −12.8% | 10.3% | 75.4% | 16.9% | 0.52 |
| Saposa | 31.1% | 0.9% | 41.8% | 5.6% | −10.7% | 5.7% | −1.9 | −1.0% | 17.2% | 63.2% | 14.2% | 0.25 |
| Mussau | 29.2% | 0.9% | 39.4% | 4.7% | −10.2% | 4.8% | −2.1 | −1.4% | 14.6% | 59.8% | 22.0% | 0.54 |
| Teop | 26.5% | 0.9% | 67.8% | 7.3% | −41.3% | 7.4% | −5.6 | −97.4% | 22.2% | 150.4% | 11.5% | 0.86 |
| Kove | 26.4% | 0.9% | 29.8% | 3.8% | −3.4% | 3.9% | −0.9 | 16.2% | 12.0% | 36.6% | 20.2% | 0.01 |
| Tigak | 26.2% | 0.9% | 43.9% | 6.7% | −17.7% | 6.8% | −2.6 | −26.9% | 20.4% | 79.3% | 21.4% | 0.87 |
| Melamela | 25.3% | 0.9% | 50.1% | 7.1% | −24.8% | 7.2% | −3.5 | −49.1% | 21.6% | 99.7% | 18.8% | 0.94 |
| Manus | 24.9% | 1.2% | 14.2% | 6.2% | 10.7% | 6.3% | 1.7 | 57.0% | 19.2% | −7.2% | 42.9% | 0.87 |
| Nakanai_Bileki | 24.5% | 0.9% | 38.4% | 14.3% | −13.9% | 14.3% | −1.0 | −17.2% | 43.1% | 66.2% | 11.2% | 0.28 |
| Mangseng | 23.6% | 0.9% | 42.2% | 3.7% | −18.6% | 3.8% | −4.9 | −32.2% | 11.7% | 79.4% | 19.9% | 0.34 |
| Papuan_Gulf | 22.6% | 1.1% | 49.3% | 6.6% | −26.7% | 6.7% | −4.0 | −57.5% | 20.3% | 102.7% | 10.9% | 0.07 |
| Notsi | 20.7% | 0.8% | 33.0% | 3.6% | −12.3% | 3.7% | −3.3 | −16.2% | 11.3% | 57.6% | 64.8% | 0.13 |
| Nailik | 20.4% | 0.8% | 44.8% | 21.6% | −24.4% | 21.6% | −1.1 | −52.8% | 64.9% | 93.6% | 22.9% | 0.08 |
| Madak | 18.7% | 0.8% | 43.1% | 7.6% | −24.4% | 7.6% | −3.2 | −54.5% | 23.0% | 91.9% | 18.7% | 0.11 |
| Kuot_Kabil | 18.6% | 0.9% | 55.4% | 6.2% | −36.8% | 6.3% | −5.9 | −91.8% | 18.9% | 129.0% | 26.5% | 0.04 |
| Bougainville_Sout | 18.0% | 1.4% | 24.1% | 8.8% | −6.1% | 8.9% | −0.7 | −0.3% | 27.0% | 36.3% | 11.8% | 0.26 |
| Lavongai | 17.6% | 0.8% | 36.4% | 3.9% | −18.8% | 4.0% | −4.7 | −38.8% | 12.1% | 74.0% | 16.0% | 0.07 |
| Nasioi | 17.4% | 1.0% | 29.9% | 5.3% | −12.5% | 5.4% | −2.3 | −20.1% | 16.4% | 54.9% | 11.5% | 0.40 |
| Kuot_Lamalaua | 16.2% | 0.8% | 13.8% | 3.8% | 2.4% | 3.9% | 0.6 | 23.4% | 11.8% | 9.0% | 19.6% | 0.07 |
| Nakanai_Loso | 15.4% | 1.1% | 24.3% | 6.5% | −8.9% | 6.6% | −1.4 | −11.3% | 20.0% | 42.1% | 11.2% | 0.37 |
| Mengen | 15.2% | 0.8% | 35.3% | 3.7% | −20.1% | 3.8% | −5.3 | −45.1% | 11.6% | 75.5% | 13.0% | 0.41 |
| Tolai | 14.3% | 0.8% | 22.5% | 4.3% | −8.2% | 4.4% | −1.9 | −10.3% | 13.3% | 38.9% | 9.4% | 0.01 |
| Sulka | 14.1% | 0.8% | 47.0% | 3.1% | −32.9% | 3.2% | - | −84.6% | 9.8% | 112.8% | 16.3% | 0.48 |
| Mamusi_Paleabu | 13.7% | 1.0% | 30.9% | 5.4% | −17.2% | 5.5% | −3.1 | −37.9% | 16.7% | 65.3% | 13.9% | 0.21 |
| Mamusi | 13.7% | 0.9% | 22.7% | 4.6% | −9.0% | 4.7% | −1.9 | −13.3% | 14.3% | 40.7% | 27.1% | 0.35 |
| Ata | 13.2% | 1.0% | 26.4% | 9.0% | −13.2% | 9.1% | −1.5 | −26.4% | 27.3% | 52.8% | 15.4% | 0.24 |
| Santa_Cruz | 9.6% | 0.9% | 27.1% | 5.1% | −17.5% | 5.2% | −3.4 | −42.9% | 15.7% | 62.1% | 23.3% | 0.17 |
| Kol_New_Britain | 8.5% | 1.4% | 9.1% | 7.7% | −0.6% | 7.8% | −0.1 | 6.7% | 23.8% | 10.3% | 30.1% | 0.66 |
| Baining_Marabu | 2.6% | 1.0% | 25.3% | 10.0% | −22.7% | 10.0% | −2.3 | −65.5% | 30.3% | 70.7% | 32.5% | 0.24 |
| Baining_Malasait | 1.2% | 1.2% | 34.9% | 10.8% | −33.7% | 10.9% | −3.1 | −99.9% | 32.8% | 102.3% | 8.2% | 0.12 |
| Papuan | 0.0% | 0.5% | 2.4% | 2.7% | −2.4% | 2.7% | −0.9 | −7.2% | 8.3% | 7.2% | 0.0% | 0.58 |
Supplementary Material
Acknowledgments
We thank the 356 volunteers who donated samples for genome-wide analysis. We thank Mark Stoneking for co-funding genotyping of the Bismarck samples. We thank Murray Brilliant, Heather Norton, and Laura Scheinfeldt, for help in the preparation of the Bismarck samples and establishment of a repository for them at the Marshfield Foundation. We thank Alexander Kim, Irina Pugach, and Mark Stoneking for critical comments, and Iain Mathieson for critiques and advice on estimating sex-specific ancestral contributions. The Fig. 1a and the Fig. 3b–e maps were plotted in R using the world() map of the ‘fields’ and ‘maps’ packages (using public domain data from the CIA World Data Bank II). P.S. was supported by the Wenner-Gren foundation and the Swedish Research Council (VR grant 2014-453). The Teouma research by M.S. and S.B. was supported by the Australian Research Council (Discovery Grants DP0880789 and DP110101415), the National Geographic Society, and by the Australia-Pacific Science Foundation. F.V. was supported by CNRS-UMR 7041. M.N. was supported by an Irish Research Council grant (GOIPD/2013/1). D.F. was supported by an Irish Research Council grant (GOIPG/2013/36). Q.F. was funded by the National Natural Science Foundation of China (L1524016), the Chinese Academy of Sciences Discipline Development Strategy Project (2015-DX-C-03) and the Bureau of International Cooperation of the Chinese Academy of Sciences. T.K. was supported by ERC starting grant FP7-261213. C.P. and J.K. were supported by the Baden Wuerttemberg Foundation. J.K was supported by the DFG grant KR 4015/1-1 and the Max Planck Society. R.P. was supported by ERC starting grant ADNABIOARC (263441). D.R. was supported by NIH grant GM100233, by NSF HOMINID BCS-1032255, and is a Howard Hughes Medical Institute investigator.
Footnotes
The authors declare no competing financial interests. Readers are welcome to comment on the online version of the paper
Supplementary Information is available in the online version of the paper.
Author contributions J.K., R.P. and D.R. supervised the study. M.S., F.V., S.B., G.A.C., and C.R. assembled archaeological material and information. P.S., C.P., Q.F., M.L., S.M., N.R. and D.R. analyzed genetic data. C.P., K.Si., F.P., D.F., E.H., N.R, and K.St. performed laboratory work. S.A., M.P.C., F.R.F., J.S.F., T.K., G.K., P.K., D.A.M., F-X.R., and T.S.W. assembled the sample collection from present-day populations. P.S. and D.R. wrote the manuscript with major input from C.P., M.S., F.V., G.A.C., M.P.C., J.S.F, J.K., R.P. and additional input from all other co-authors.
The aligned sequences are available through the European Nucleotide Archive under accession number PRJEB14728. The newly reported SNP genotyping data for the subset of individuals who provided informed consent consistent with fully public distribution are available at (http://genetics.med.harvard.edu/reichlab/Reich_Lab/Datasets.html). To access data for the remaining samples, researchers should send a signed letter to D.R. containing the following text: “(a) I will not distribute the data outside my collaboration; (b) I will not post the data publicly; (c) I will make no attempt to connect the genetic data to personal identifiers for the samples; (d) I will use the data only for studies of population history; (e) I will not use the data for any selection studies; (f) I will not use the data for medical or disease-related analyses; (g) I will not use the data for commercial purposes.” Extended Data Table 2 specifies which samples are consistent with which type of data distribution
References
- 1.Sheppard PJ, Chiu S, Walter R. Re-dating Lapita Movement into Remote Oceania. Journal of Pacific Archaeology. 2015;6:26–36. [Google Scholar]
- 2.Kayser M, et al. Genome-wide Analysis Indicates More Asian than Melanesian Ancestry of Polynesians. The American Journal of Human Genetics. 2008;82:194–198. doi: 10.1016/j.ajhg.2007.09.010. http://dx.doi.org/10.1016/j.ajhg.2007.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kayser M. The human genetic history of Oceania: near and remote views of dispersal. Current Biology. 2010;20:R194–R201. doi: 10.1016/j.cub.2009.12.004. [DOI] [PubMed] [Google Scholar]
- 4.Wollstein A, et al. Demographic history of Oceania inferred from genome-wide data. Current Biology. 2010;20:1983–1992. doi: 10.1016/j.cub.2010.10.040. [DOI] [PubMed] [Google Scholar]
- 5.Matisoo-Smith E. Ancient DNA and the human settlement of the Pacific: A review. Journal of Human Evolution. 2015;79:93–104. doi: 10.1016/j.jhevol.2014.10.017. http://dx.doi.org/10.1016/j.jhevol.2014.10.017. [DOI] [PubMed] [Google Scholar]
- 6.Bellwood PS. First farmers: the origins of agricultural societies. Blackwell Publishing; 2005. [Google Scholar]
- 7.Duggan AT, et al. Maternal history of Oceania from complete mtDNA genomes: contrasting ancient diversity with recent homogenization due to the Austronesian expansion. The American Journal of Human Genetics. 2014;94:721–733. doi: 10.1016/j.ajhg.2014.03.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kayser M, et al. Melanesian origin of Polynesian Y chromosomes. Current Biology. 2000;10:1237–1246. doi: 10.1016/s0960-9822(00)00734-x. http://dx.doi.org/10.1016/S0960-9822(00)00734-X. [DOI] [PubMed] [Google Scholar]
- 9.Blust R. Remote Melanesia: one history or two? An addendum to Donohue and Denham. Oceanic Linguistics. 2008;47:445–459. [Google Scholar]
- 10.Friedlaender JS, et al. The Genetic Structure of Pacific Islanders. PLoS Genet. 2008;4:e19. doi: 10.1371/journal.pgen.0040019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pinhasi R, et al. Optimal Ancient DNA Yields from the Inner Ear Part of the Human Petrous Bone. PLoS ONE. 2015;10:e0129102. doi: 10.1371/journal.pone.0129102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dabney J, et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proceedings of the National Academy of Sciences. 2013;110:15758–15763. doi: 10.1073/pnas.1314445110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rohland N, Harney E, Mallick S, Nordenfelt S, Reich D. Partial uracil–DNA–glycosylase treatment for screening of ancient DNA. Philosophical Transactions of the Royal Society of London B: Biological Sciences. 2015;370 doi: 10.1098/rstb.2013.0624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Fu Q, et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature. 2015 doi: 10.1038/nature14558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Skoglund P, Storå J, Götherström A, Jakobsson M. Accurate sex identification of ancient human remains using DNA shotgun sequencing. Journal of Archaeological Science. 2013;40:4477–4482. [Google Scholar]
- 16.Melton T, et al. Polynesian genetic affinities with Southeast Asian populations as identified by mtDNA analysis. American journal of human genetics. 1995;57:403. [PMC free article] [PubMed] [Google Scholar]
- 17.Skoglund P, et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science. 2012;336:466–469. doi: 10.1126/science.1216304. [DOI] [PubMed] [Google Scholar]
- 18.Skoglund P, et al. Separating endogenous ancient DNA from modern day contamination in a Siberian Neandertal. Proceedings of the National Academy of Sciences. 2014 doi: 10.1073/pnas.1318934111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Reich D, et al. Reconstructing Native American population history. Nature. 2012;488:370– 374. doi: 10.1038/nature11258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Moorjani P, et al. The history of African gene flow into Southern Europeans, Levantines, and Jews. PLoS genetics. 2011;7:e1001373. doi: 10.1371/journal.pgen.1001373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Loh P-R, et al. Inference of admixture parameters in human populations using weighted linkage disequilibrium. 2012 [Google Scholar]
- 22.Fenner JN. Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. American Journal of Physical Anthropology. 2005;128:415–423. doi: 10.1002/ajpa.20188. [DOI] [PubMed] [Google Scholar]
- 23.Xu S, Pugach I, Stoneking M, Kayser M, Jin L. Genetic dating indicates that the Asian–Papuan admixture through Eastern Indonesia corresponds to the Austronesian expansion. Proceedings of the National Academy of Sciences. 2012;109:4574–4579. doi: 10.1073/pnas.1118892109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Pugach I, Matveyev R, Wollstein A, Kayser M, Stoneking M. Dating the age of admixture via wavelet transform analysis of genome-wide data. Genome Biol. 2011;12:R19. doi: 10.1186/gb-2011-12-2-r19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lipson M, et al. Reconstructing Austronesian population history in Island Southeast Asia. Nature communications. 2014:5. doi: 10.1038/ncomms5689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Patterson N, et al. Ancient admixture in human history. Genetics. 2012;192:1065–1093. doi: 10.1534/genetics.112.145037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pickrell JK, Pritchard JK. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS genetics. 2012;8:e1002967. doi: 10.1371/journal.pgen.1002967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bellwood P. Holocene Population History in the Pacific Region as a Model for Worldwide Food Producer Dispersals. Current Anthropology. 2011;52:S363–S378. doi: 10.1086/658181. [DOI] [Google Scholar]
- 29.Jordan FM, Gray RD, Greenhill SJ, Mace R. Matrilocal residence is ancestral in Austronesian societies. Proceedings of the Royal Society of London B: Biological Sciences. 2009 doi: 10.1098/rspb.2009.0088. rspb. 2009.0088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Stephen Lansing J, et al. An ongoing Austronesian expansion in Island Southeast Asia. Journal of Anthropological Archaeology. 2011;30:262–272. http://dx.doi.org/10.1016/j.jaa.2011.06.004. [Google Scholar]
- 31.Knapp M, Clarke AC, Horsburgh KA, Matisoo-Smith EA. Setting the stage – Building and working in an ancient DNA laboratory. Annals of Anatomy - Anatomischer Anzeiger. 2012;194:3–6. doi: 10.1016/j.aanat.2011.03.008. http://dx.doi.org/10.1016/j.aanat.2011.03.008. [DOI] [PubMed] [Google Scholar]
- 32.Meyer M, Kircher M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harbor Protocols. 2010;2010 doi: 10.1101/pdb.prot5448. pdb. prot5448. [DOI] [PubMed] [Google Scholar]
- 33.Fu Q, et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proceedings of the National Academy of Sciences. 2013;110:2223–2227. doi: 10.1073/pnas.1221359110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Haak W, et al. Massive migration from the steppe is a source for Indo-European languages in Europe. 2015 doi: 10.1038/nature14317. arXiv preprint arXiv:1502.02783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lazaridis I, et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature. 2014;513:409–413. doi: 10.1038/nature13673. http://www.nature.com/nature/journal/v513/n7518/abs/nature13673.html - supplementary-information. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Qin P, Stoneking M. Denisovan Ancestry in East Eurasian and Native American Populations. Molecular Biology and Evolution. 2015 doi: 10.1093/molbev/msv141. [DOI] [PubMed] [Google Scholar]
- 38.Skoglund P, et al. Genetic evidence for two founding populations of the Americas. Nature. 2015 doi: 10.1038/nature14895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Moreno-Mayar JV, et al. Genome-wide Ancestry Patterns in Rapanui Suggest Pre-European Admixture with Native Americans. Current Biology. 2014;24:2518–2525. doi: 10.1016/j.cub.2014.09.057. http://dx.doi.org/10.1016/j.cub.2014.09.057. [DOI] [PubMed] [Google Scholar]
- 40.Reich D, et al. Denisova Admixture and the First Modern Human Dispersals into Southeast Asia and Oceania. The American Journal of Human Genetics. 2011;89:516–528. doi: 10.1016/j.ajhg.2011.09.005. http://dx.doi.org/10.1016/j.ajhg.2011.09.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Reich D, et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature. 2010;468:1053–1060. doi: 10.1038/nature09710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Meyer M, et al. A High-Coverage Genome Sequence from an Archaic Denisovan Individual. Science. 2012;338:222–226. doi: 10.1126/science.1224344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Prufer K, et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature. 2014;505:43–49. doi: 10.1038/nature12886. http://www.nature.com/nature/journal/v505/n7481/abs/nature12886.html - supplementary-information. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Allentoft ME, et al. Population genomics of Bronze Age Eurasia. Nature. 2015;522:167–172. doi: 10.1038/nature14507. http://www.nature.com/nature/journal/v522/n7555/abs/nature14507.html - supplementary-information. [DOI] [PubMed] [Google Scholar]
- 45.Green RE, et al. A Draft Sequence of the Neandertal Genome. Science. 2010;328:710–722. doi: 10.1126/science.1188021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Raghavan M, et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature. 2014;505:87–91. doi: 10.1038/nature12736. http://www.nature.com/nature/journal/v505/n7481/abs/nature12736.html - supplementary-information. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Raghavan M, et al. The genetic prehistory of the New World Arctic. Science. 2014:345. doi: 10.1126/science.1255832. [DOI] [PubMed] [Google Scholar]
- 48.Rasmussen M, et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature. 2014;506:225–229. doi: 10.1038/nature13025. http://www.nature.com/nature/journal/v506/n7487/abs/nature13025.html - supplementary-information. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rasmussen M, et al. The ancestry and affiliations of Kennewick Man. Nature. 2015;523:455–458. doi: 10.1038/nature14625. http://www.nature.com/nature/journal/v523/n7561/abs/nature14625.html - supplementary-information. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Skoglund P, et al. Genomic Diversity and Admixture Differs for Stone-Age Scandinavian Foragers and Farmers. Science. 2014;344:747–750. doi: 10.1126/science.1253448. [DOI] [PubMed] [Google Scholar]
- 51.Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS genetics. 2006;2:e190. doi: 10.1371/journal.pgen.0020190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Skoglund P, Jakobsson M. Archaic human ancestry in East Asia. Proceedings of the National Academy of Sciences. 2011 doi: 10.1073/pnas.1108181108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Sudmant PH, et al. Global diversity, population stratification, and selection of human copy-number variation. Science. 2015:349. doi: 10.1126/science.aab3761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kong A, et al. A high-resolution recombination map of the human genome. Nat Genet. 2002;31:241–247. doi: 10.1038/ng917. http://www.nature.com/ng/journal/v31/n3/suppinfo/ng917_S1.html. [DOI] [PubMed] [Google Scholar]
- 55.Moorjani P, et al. A genetic method for dating ancient genomes provides a direct estimate of human generation interval in the last 45,000 years. Proceedings of the National Academy of Sciences. 2016;113:5652–5657. doi: 10.1073/pnas.1514696113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Reich D, Thangaraj K, Patterson N, Price AL, Singh L. Reconstructing Indian population history. Nature. 2009;461:489–494. doi: 10.1038/nature08365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Csillery K, Blum MGB, Gaggiotti O, Francois O. Approximate Bayesian Computation (ABC) in practice. Trends in Ecology and Evolution. 2010;25:410– 418. doi: 10.1016/j.tree.2010.04.001. [DOI] [PubMed] [Google Scholar]
- 58.Bronk Ramsey C. OxCal Program v4.2.4. Radiocarbon Accelerator Unit, University of Oxford; 2016. [Google Scholar]
- 59.Reimer PJ, et al. IntCal13 and Marine13 Radiocarbon Age Calibration Curves 0–50,000 Years cal BP. 2013. [Google Scholar]
- 60.Ambrose SH. Food and nutrition in history and anthropology (USA) 1993. Isotopic analysis of paleodiets: methodological and interpretive considerations. [Google Scholar]
- 61.Petchey F, Spriggs M, Bedford S, Valentin F, Buckley H. Radiocarbon dating of burials from the Teouma Lapita cemetery, Efate, Vanuatu. Journal of Archaeological Science. 2014;50:227–242. http://dx.doi.org/10.1016/j.jas.2014.07.002. [Google Scholar]
- 62.Petchey F, Anderson A, Zondervan A, Ulm S, Hogg A. New marineΔ R values for the South Pacific subtropical gyre region. Radiocarbon. 2008;50:373–397. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.