Abstract
Vesicomyid clams in deep-sea chemosynthetic ecosystems harbor sulfur-oxidizing bacteria in their gill epithelial cells. These symbionts, which are vertically transmitted, are species-specific and thought to have cospeciated with their hosts. However, recent studies indicate incongruent phylogenies between some vesicomyid clams and their symbionts, suggesting that symbionts are horizontally transmitted. To more precisely understand the evolution of vesicomyid clams and their symbionts, we compared the evolution of vesicomyid clams and their symbionts through phylogenetic analyses using multi-gene data sets. Many clades in the phylogenetic trees of 13 host species (Abyssogena mariana, Ab. phaseoliformis, Akebiconcha kawamurai, Calyptogena fausta, C. laubieri, C. magnifica, C. nautilei, C. pacifica, Isorropodon fossajaponicum, Phreagena kilmeri, Ph. okutanii, Ph. soyoae, and Pliocardia stearnsii) and their symbionts were well resolved. Six of the 13 host-symbiont pairs (C. fausta, C. magnifica, C. pacifica, Ph. kilmeri, Ph. okutanii, and Ph. soyoae, and their respective symbionts) showed topological congruence. However, the remaining seven pairs (Ak. kawamurai, Ab mariana, Ab. phaseoliformis, C. laubieri, C. nautilei, I. fossajaponicum, and Pl. stearnsii and their corresponding symbionts) showed incongruent topologies, which were supported by the approximately unbiased and Bayes factor tests. Coevolution analyses indicated that six pairs cospeciated, whereas host switching events occurred in the remaining seven pairs. Markedly, multiple host switching events may have occurred in the lineages from the common ancestral symbiont of C. pacifica and C. fausta. Our phylogenetic and coevolution analyses provide additional evidence for host switching during the evolution of vesicomyids.
Keywords: cospeciation, endosymbiosis, host switching, phylogenetic tree, vertical transmission
Introduction
Chemosynthetic ecosystems in hydrothermal vents and seeps are dominated by a wide variety of invertebrates, many of which establish symbiosis with chemosynthetic bacteria. The hosts nutritionally depend on the organic compounds synthesized by the chemosynthetic symbionts, as shown by stable isotope analyses (Le Pennec etal. 1995). Therefore, acquisition of symbionts is key for the host’s survival (Bright and Bulgheresi 2010). The symbionts are transmitted from one generation to the next through either vertical or horizontal transmissions (Dubilier etal. 2008; Bright and Bulgheresi 2010). In vertical transmission, the symbionts are directly transferred from the parent to the offspring via the host’s egg or sperm, and most of the symbionts are maternally inherited. In horizontal transmission, the symbiont is acquired from the environment during the host’s development. Horizontal transmission of the symbiont is independent of the host’s reproduction. The phylogenetic trees of hosts and horizontally transmitted symbionts are incongruent in their topology (Won etal. 2008; Bright and Bulgheresi 2010), whereas those of hosts and vertically transmitted symbionts are congruent (Peek etal. 1998; Bright and Bulgheresi 2010). The latter suggests cospeciation of hosts and their symbionts (Peek etal. 1998).
The deep-sea clams belonging to the family Vesicomyidae are endemic to chemosynthetic ecosystems and are dominant members of these ecosystems. Over 100 extant vesicomyid species have been identified worldwide (Krylova and Shaling 2010). Traditionally, Vesicomyidae has been classified into two genera, Calyptogena and Vesicomya. However, species of Calyptogena (sensu lato) have recently been classified into as many as 15 distinct genera (Krylova and Sahling 2010; Decker etal., 2012). Vesicomyid clams harbor intracellular sulfur-oxidizing bacterial symbionts in the gill epithelial cells; these symbionts are species-specific and are generally believed to be transmitted to the offspring vertically via eggs (Endow and Ohta 1990; Cary and Giovannoni 1993; Ikuta etal. 2016a). However, Stewart etal. (2008) reported that the positions of several host species and their symbionts in their respective phylogenetic trees were incongruent, suggesting that horizontal transmission may have occurred between host-symbiont pairs in the course of evolution. In addition, two phylogenetically distant vesicomyid species, Christineconcha regab and Laubiericoncha chuni, which cooccur in the same aggregations, each harbor small populations of the other species’ symbionts in the gills, along with much larger populations of their original symbiont species (Decker etal. 2013). These findings suggest that at least a few of the symbionts in vesicomyid clams are occasionally transmitted horizontally, and that not all of the vesicomyid clams and their symbionts have coevolved.
In previously reported phylogenetic trees of both the vesicomyid clams and their symbionts, many nodes were poorly supported, owing to the small gene data sets used for the phylogenetic analyses (Peek etal. 1998). Thus, larger gene data sets are required to reveal the more detailed phylogenetic relationships of the clams and their symbionts. To date, complete genomes of the symbionts from two vesicomyid clams, Phreagena okutanii and Calyptogena magnifica, have been sequenced (Kuwahara etal. 2007; Newton etal. 2007). In addition, eight gene sequences [16S rRNA, 23S rRNA, UTP-glucose-1-phosphate uridylyltransferase (galU), Chaperonin EL subunit (groEL), Chaperonin ES subunit (groES), transcription-repair coupling factor (mfd), excinuclease ABC subunit A (uvrA), and DNA helicase II/ATP-dependent (uvrD paralog)] have been determined for several other vesicomyid clam symbiont species (Kuwahara etal. 2011; Shimamura etal. 2017). With regard to hosts, the mitochondrial genomes of five vesicomyid clams were sequenced recently (Liu etal. 2016; Ozawa etal. 2017). Mitochondrial genomes are frequently used to resolve phylogenetic relationships within a metazoan group, because they have relatively high mutation rates and generally contain the 13 protein-coding genes, lacking introns, within one compact genome, providing easily accessible gene sequences.
To understand the evolution of vesicomyid clams and their symbionts, the evolutionary events in 13 vesicomyid clams (Abyssogena mariana, Ab. phaseoliformis, Akebiconcha kawamurai, Calyptogena fausta, C. laubieri, C. magnifica, C. nautilei, C. pacifica, Isorropodon fossajaponicum, Phreagena kilmeri, Ph. okutanii, Ph. soyoae, and Pliocardia stearnsii) and their symbionts were analyzed by comparing the phylogenetic trees of concatenated large sequence data sets composed of 11 mitochondrial genes and eight symbiont genes. Our findings strongly suggest that host switching of symbionts has occurred, based on the topological incongruence between the phylogenetic trees of vesicomyid clams and their corresponding symbionts. On the basis of our current knowledge of the intracellular symbiosis between vesicomyid clams and their symbionts, we discuss the possible mechanism of host switching in vesicomyid clams.
Materials and Methods
Sample Collection and DNA Extraction
Ten vesicomyid clam species (Ab. mariana, Ak. kawamurai, C. fausta, C. laubieri, C. nautilei, C. pacifica, I. fossajaponicum Ph. kilmeri, Ph. soyoae, and Pl. stearnsii) were collected from seeps, using a Deep Submergence Vehicle (DSV) or Remotely Operated Vehicle (ROV), and immediately stored at −80 °C in a freezer until analysis (table 1). For DNA extraction, after the clam samples were thawed, the mantle and/or gill were dissected and cut into small pieces on ice. Total DNA was isolated from ∼25 mg of each tissue using a DNeasy Tissue Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions, and isolated DNA was stored at −20 °C. All of these experiments were conducted in accordance with the Guidelines for Proper Conduct of Animal Experiments (Science Council of Japan).
Table 1.
Vesicomyid Clam Species Used in the Present Study
| Species | Date of Collection | Depth (m) | Latitude | Longitude | Location | Collection Method | Dive No | No of Samples |
|---|---|---|---|---|---|---|---|---|
| Abyssogena mariana | 9/9/2013 | 5633 | 11.65 N | 143.04 E | Mariana Trench | DSV Shinkai 6500 | YK13-08 6K #1362 | 1 |
| Isorropodon fossajaponicum | 5/31/2006 | 6181 | 40.10 N | 144.18 E | Japan trench | DSV Shinkai 6500 | YK06-05 6K #950 | 1 |
| Akebiconcha kawamurai | 6/13/2005 | 608 | 34.08 N | 137.79 E | Daini Tenryu Knoll, Nankai Trough | DSV Shinkai 6500 | YK05-08 6K#881 | 1 |
| Calyptogena fausta | 6/10/1996 | 1490 | 34.92 N | 138.65 E | Suruga Bay | DSV Shinkai 2000 | NT96-09 2K#869 | 1 |
| Calyptogena laubieri | 8/23/1997 | 3761 | 33.65 N | 137.91 E | Nankai Trough | ROV Kaiko | KR97-05 Kaiko#45 | 1 |
| Calyptogena nautilei | 6/18/2005 | 3257 | 32.58 N | 134.69 E | Off Muroto, Nankai Trough | DSV Shinkai 6500 | YK05-08 6K#884 | 1 |
| Calyptogena pacifica | 3/29/1996 | 659–683.5 | 36.77 N | 122.05 W | Monterey Bay | ROV Ventana | dive #1037 | 1 |
| Phreagena kilmeri | 3/25/1996 | 900 | 36.73 N | 122.04W | Monterey Bay | ROV Ventana | dive #1032 | 1 |
| Phreagena soyoae | 1/14/2010 | 1171 | 35.00 N | 139.23 E | Off Hatsushima, Sagami Bay | ROV Hyper-Dolphin | NT10-01 HPD#1074 | 1 |
| Pliocardia stearnsii | 3/29/1996 | 659–683.5 | 36.77 N | 122.05 W | Monterey Bay | ROV Ventana | dive #1037 | 1 |
Polymerase Chain Reaction (PCR) Amplification and Sequencing
The mitochondrial genomes in five vesicomyid clams (Ab. mariana, Ab. phaseoliformis, C. magnifica, I. fossajaponicum, and Ph. okutanii) used in the present study were recently sequenced (Liu etal. 2016; Ozawa etal. 2017). We obtained the sequences of 11 mitochondrial genes [ATP synthase F0 subunit 6 (atp6), ATP synthase F0 subunit 8 (atp8), cytochrome b (cob), cytochrome c oxidase subunit I (cox1), cytochrome c oxidase subunit II (cox2), cytochrome c oxidase subunit III (cox3), NADH dehydrogenase subunit 1 (nad1), NADH dehydrogenase subunit 3 (nad3), NADH dehydrogenase subunit 4 (nad4), NADH dehydrogenase subunit 5 (nad5), and large subunit ribosomal RNA gene (rrnL)] from the remaining eight vesicomyid clams (Ak. kawamurai, C. fausta, C. laubieri, C. nautilei, C. pacifica, Ph. kilmeri, Ph. soyoae, and Pl. stearnsii). To obtain the partial mitochondrial genome sequences, including those of 11 mitochondrial genes, three or four PCR primer sets per vesicomyid species were designed based on the consensus mitochondrial genome sequences (table 2). With regard to the symbionts, we used seven sets of PCR primers for eight symbiont gene sequences [16S rRNA (Lane 1991), 23S rRNA (Kuwahara etal. 2011), and galU, groEL, groES, mfd, uvrA, and uvrD paralog (Shimamura etal. 2017)] from the symbiont of Ab. mariana and one primer set for the galU gene sequence from the symbiont of I. fossajaponicum (table 2). The mitochondrial or symbiont gene fragments were amplified in 25-µL PCR reactions that contained ∼100 ng DNA template, 1× LA-Taq buffer (Mg2+ plus; TaKaRa Bio, Inc., Shiga, Japan), 0.25 mM each deoxynucleotide (dNTP) solution, 0.2 µM each primer, and 0.625 U LA-Taq polymerase (TaKaRa Bio, Inc.). The PCR reaction mixtures were incubated under the conditions described in table 2 after the initial denaturation step (96 °C for 2 min), using a TaKara TP600 Thermal cycler (TaKaRa Bio, Inc.). To check the size of the PCR-amplified fragments, 2.0-µL aliquots of the PCR reaction mixtures were electrophoresed on 0.6–1.0% agarose gels and stained with 0.06% ethidium bromide. The unincorporated primers and dNTPs in the PCR reaction mixtures were removed as described previously (Werle etal. 1994). The amplified DNA fragments were then directly sequenced using a Big Dye Terminator v3.1 Cycle Sequencing Kit (Life Technologies, Carlsbad, CA, USA) and either an ABI PRIZM 3130xl or 3730xl Genetic Analyzer (Life Technologies), according to the manufacturer’s instructions.
Table 2.
Primers and PCR Conditions for the Amplification of Vesicomyid Mitochondrial and Symbiont’s Gene Regions
| Species | Host or Symbiont | Forward Primer | Sequence | Reverse Primer | Sequence | PCR Amplification Condition (30 cycles) | Genomic Region |
|---|---|---|---|---|---|---|---|
| Abyssogena mariana | Symbiont | 27Fa | AGAGTTTGATCCTGGCTCAG | 1492Rb | GGTTACCTTGTTACGACTT | 96 °C 20 s—56 °C 10 s—72 °C 2 min | 16S rRNA |
| Symbiont | 23S_Fc | GGGAACTGAAACATCTAAGTACC | 23S_Rc | CCCGCTTAGATGCTTTCAG | 96 °C 20 s—56 °C 10 s—72 °C 2 min | 23S rRNA | |
| Symbiont | UvrA_Fd | GCRCGTARTGCTGCRCCAGTGTC | UvrA_Rd | RCTTGATGGAGGYCCWATTATTGCC | 96 °C 20 s—56 °C 10 s—72 °C 5 min | UvrA | |
| Symbiont | UvrDpara_Fd | AARGYTTCCATGTCAACCCGTAGTG | UvrDpara_Rd | AAGCATTTGATGCTGGMTGTGATGC | 96 °C 20 s—56 °C 10 s—72 °C 5 min | UvrD paralog | |
| Symbiont | mfd_Fd | ACCAACAACCTCACAACCATCAGC | mfd_Rd | CGATATTATCACCATCGCTACTAGC | 96 °C 20 s—56 °C 15 s—72 °C 5 min | mfd | |
| Symbiont | galU_Fd | GTTAAGGCTGTATCCCTATATGCAC | galU_Rd | GCTTGTGGYACTAGTTATAAYGCAG | 96 °C 20 s—56 °C 10 s—72 °C 5 min | galU | |
| Symbiont | GroELS_Fc | YGCACCAGAACAATAYTTAACCAATG | GroELS_Rc | WAYCCTGCAATTGYYTCATCAACAC | 96 °C 20 s—56 °C 10 s—72 °C 4 min | groES-groEL | |
| Isorropodon fossajaponicum | Symbiont | Ifos_galU_F | TTAAATAATACCTTGTTGTTTAGC | galU_Rd | GCTTGTGGYACTAGTTATAAYGCAG | 96 °C 20 s—56 °C 10 s—72 °C 5 min | galU |
| Akebiconcha kawamurai | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | AkaMit_rrnL_F | TTACAGGGCTGTTAATGGCTGCTC | nad1_R2 | AATCAAATGGCGCTCGATTAGTCTC | 96 °C 20 s—60 °C 10 s—68 °C 5 min | rrnL - nad1 | |
| Host | AkaMit_nad1_F | ATTGGCAGAGACTAATCGAGCTC | LNCR_R2 | TGATGTATTGCTACATTTGTCGCGC | 96 °C 20 s—60 °C 10 s—68 °C 2 min | nad1 - LNCR | |
| Calyptogena fausta | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | rrnL_F | CTACCATAGGGATAACAGCGTTATC | tRNA_Q_R | AGAGAACTTAATCTCATCTCTGACTC | 96 °C 20 s—60 °C 10 s—68 °C 6 min | rrnL - tRNA Gln | |
| Host | CfaMit_nad4_F | AGCCTGTGTTTAACTTGAGAGAACG | nad1_R | ATACTCAACCCTATACCCCGCTAC | 96 °C 20 s—60 °C 10 s—68 °C 2 min | nad4 - nad1 | |
| Host | CfaMit_nad1_F | TATCTGGGTGAGTTTGTGATTCTCG | LNCR_R2 | TGATGTATTGCTACATTTGTCGCGC | 96 °C 20 s—60 °C 10 s—68 °C 2 min | nad1 - LNCR | |
| Calyptogena laubieri | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | ClaMit_rrnL_F | AGATTATGTGGACTATGGTGCCTGC | nad1_R | ATACTCAACCCTATACCCCGCTAC | 96 °C 20 s—60 °C 10 s—68 °C 6 min | rrnL - nad1 | |
| Host | ClaMit_nad1_F | ATGGCTTGTGGCCACATTGGCAG | LNCR_R2 | TGATGTATTGCTACATTTGTCGCGC | 96 °C 20 s—60 °C 10 s—68 °C 2 min | nad1 - LNCR | |
| Calyptogena nautilei | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | CnaMit_rrnL_F | GTTAGTAGGGTTGATATTGGCCATC | nad1_R2 | AATCAAATGGCGCTCGATTAGTCTC | 96 °C 20 s—60 °C 10 s—68 °C 6 min | rrnL - nad1 | |
| Host | CnaMit_atp6_F | TATGCGGGCTGTAGCTCAGAGAG | LNCR_R | AGGCAGRCCTCACGGAGCCACGC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | atp6 - LNCR | |
| Calyptogena pacifica | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | CpaMit_rrnL_F | AGAGTGCTTGGTGGGTGTTCTATC | nad1_R2 | AATCAAATGGCGCTCGATTAGTCTC | 96 °C 20 s—60 °C 10 s—68 °C 6 min | rrnL - nad1 | |
| Host | CpaMit_atp6_F | GTATGATTCAATTGCGGCAAGGGC | LNCR_R | AGGCAGRCCTCACGGAGCCACGC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | atp6 - LNCR | |
| Phreagena kilmeri | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | rrnL_F | CTACCATAGGGATAACAGCGTTATC | nad1_R2 | AATCAAATGGCGCTCGATTAGTCTC | 96 °C 20 s—60 °C 10 s—68 °C 6 min | rrnL - nad1 | |
| Host | PkiMit_atp6_F | TGAAGGTTGCTTAGCTTCTTTGCAG | LNCR_R | AGGCAGRCCTCACGGAGCCACGC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | atp6 - LNCR | |
| Phreagena soyoae | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | PsoMit_rrnL_F | TTGCCTAAGGCTCATGTGGAGGC | nad1_R2 | AATCAAATGGCGCTCGATTAGTCTC | 96 °C 20 s—60 °C 10 s—68 °C 6 min | rrnL - nad1 | |
| Host | AkaMit_nad1_F | ATTGGCAGAGACTAATCGAGCTC | LNCR_R | AGGCAGRCCTCACGGAGCCACGC | 96 °C 20 s—60 °C 10 s—68 °C 2 min | nad1 - LNCR | |
| Pliocardia stearnsii | Host | rrnS_F | GGAATTCAGGTACACGTATAGGATC | rrnL_R | TTAATCCAACATCGAGGTCGCAAAC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnS - rrnL |
| Host | PstMit_rrnL_F | AAGAGGTTAGGCCTGCCCGGTG | nad1_R2 | AATCAAATGGCGCTCGATTAGTCTC | 96 °C 20 s—60 °C 10 s—68 °C 8 min | rrnL - nad1 | |
| Host | PstMit_atp6_F | GGCATGTGAACTGCTAGTGTAGTG | LNCR_R | AGGCAGRCCTCACGGAGCCACGC | 96 °C 20 s—60 °C 10 s—68 °C 5 min | atp6 - LNCR |
Assembly and Analysis of DNA Sequences
The nucleotide sequences of the PCR-amplified fragments were assembled using Sequencher Ver 4.10.1 (Gene Codes Co., MI, USA). The open-reading frames of the protein-coding genes were then determined using Sequencher Ver 4.10.1 and identified using BLASTP against the NCBI nonredundant protein sequence database. The rrnL gene was identified using BLASTN against the NCBI nonredundant nucleotide sequence database. The sequences of the host mitochondrial genes and symbiont genes were submitted to the DNA Data Bank of Japan.
Phylogenetic Analyses
Phylogenetic analyses of the 13 vesicomyid clam species (Ab. mariana, Ab. phaseoliformis, Ak. kawamurai, C. fausta, C. laubieri, C. magnifica, C. nautilei, C. pacifica, I. fossajaponicum, Ph. kilmeri, Ph. okutanii, Ph. soyoae, and Pl. stearnsii) were conducted using the concatenated nucleotide sequences of 11 mitochondrial genes (atp6, atp8, cob, cox1, cox2, cox3, nad1, nad3, nad4, nad5, and rrnL). The accession numbers for mitochondrial gene sequences used in this study are provided in supplementary data S1, Supplementary Material online. The analyses of the corresponding symbiont species were conducted using the concatenated nucleotide sequences of eight bacterial genes (16S rRNA, 23S rRNA, galU, groEL, groES, mfd, uvrA, and uvrD paralog), of which accession numbers are provided in supplementary data S2, Supplementary Material online. A set of five Venerid species (Meretrix lusoria, M. meretrix, M. petechialis, Paphia euglypta, and Venerupis philippinarum) were included as an outgroup for the phylogenetic analysis of the host species (supplementary data S1, Supplementary Material online). The symbiont of the vent mussel Bathymodiolus septemdierum, which is closely related to the vesicomyid symbionts (Ikuta etal. 2016b), was used as an outgroup for the phylogenetic analysis of the symbiont species (supplementary data S2, Supplementary Material online).
The predicted amino acid sequences of the protein-coding genes from each mitochondrion and symbiont were separately aligned using MAFFT v7.164 b (Katoh and Standley 2013), using the default parameters. On the basis of amino acid alignment, the nucleotide sequences were aligned using the Transalign program in EMBOSS package (Rice etal. 2000). The nucleotide sequences of rRNA genes from both the host mitochondria and symbionts were also individually aligned using MAFFT v7.164 b, with the default parameters. Gaps and insert sequences in the alignments of nucleotide sequences were removed prior to phylogenetic analyses with trimAl 1.2 rev59 (Capella-Gutiérrez etal. 2009). The appropriate evolutionary model for the nucleotide sequence alignment of each gene was selected based on Akaike Information Criteria (AIC) and the Bayesian Information Criterion (BIC) for the maximum likelihood (ML) and Bayesian analyses, respectively, using Kakusan4 software (Tanabe 2011).
ML and Bayesian trees were reconstructed from the alignment of each gene, using an appropriate model for each analysis (supplementary data S3, Supplementary Material online). The ML analyses of the host clams and symbionts were conducted using RAxML version 8.2.10 (Stamatakis 2014), with 40 distinct randomized maximum parsimony starting trees (supplementary data S3, Supplementary Material online), and the ML bootstrap support (BS) was obtained by 100 resamplings. The Bayesian inference was conducted using MrBayes 3.2.6, with the appropriate evolutionary model for each gene (supplementary data S3, Supplementary Material online; Ronquist etal. 2012), and the Bayesian trees for the individual mitochondrial and symbiont genes were generated from random starting trees over one million generations, with one cold and three heated Markov chains and sampling every 100 generations. The first 250,000 generations were excluded as burn-in, and the remaining trees were summarized to obtain Bayesian posterior probability (PP). The phylogenetic trees of respective genes in host and symbiont are shown in supplementary data S4 and S5, Supplementary Material online. The resulting individual gene trees showed no instances in which the taxa were placed in conflicting positions with a zero-branch length or with extremely long branches. Hence, the concatenated alignments of the host mitochondrial genes (10,496 nucleotide sites/18 taxa) and the symbiont genes (15,847 nucleotide sites/14 taxa) were subjected to ML and Bayesian analyses, as described above (supplementary data S3, Supplementary Material online). The alignment data sets are available from Figshare (accession html: https://figshare.com/s/c28afd62efc648d87d5b).
Tree Topology Comparison
The significance of topological difference between the host and symbiont trees was assessed using nonparametric tests. An approximately unbiased (AU) test (Shimodaira 2002) and Bayes factor test were implemented in CONSEL (Shimodaira and Hasegawa 2001) and MrBayes 3.2.6. (Ronquist etal. 2012), respectively. Both the constrained and unconstrained topologies of the host and symbiont trees (table 3) were estimated using RAxML with the GTR + G model. The likelihood of the best constrained tree was compared with the likelihoods of the other trees, thereby providing probability values (P values) of AU for each tree. Tree topologies with P values of <0.05 were rejected. The marginal likelihoods were estimated by stepping-stone sampling for the hypotheses (Rambaut and Drummond 2009) (table 3), then Bayes factors were calculated by taking the differences between the marginal likelihood values. A log difference of Bayes factors in the range of 3–5 is typically considered to be strong evidence in favor of a model, and a log difference above 5 is considered to be very strong evidence (Kass and Raftery 1995).
Table 3.
Topology Tests of the Phylogenies of 13 Vesicomyid Host Clams and Their Symbionts
| Constraint Topology Patterns | AUa (p) | Log Likelihoods | BF01b | |
|---|---|---|---|---|
| (A) Host | Best host topology | 1.0000 | −79310.243 | – |
| Constraint topology including Ak. kawamurai and C. laubieri on symbiont tree | 0.0050 | −79338.346 | 12.205 | |
| Constraint topology including Ak. kawamurai, C. laubieri, Ph. kilmeri, Ph. okutanii and Ph. soyoae on symbiont tree | 0.0030 | −79337.531 | 11.851 | |
| Constraint topology including C. fausta, C. pacifica and Pl. stearnsii on symbiont tree | 0.0030 | −79365.117 | 23.832 | |
| Constraint topology including C. fausta, C. nautilei, C. pacifica and Pl. stearnsii on symbiont tree | 0.0020 | −79377.612 | 29.258 | |
| Constraint topology including Ak. kawamurai, C. fausta, C. laubieri, C. nautilei, C. magnifica, C. pacifica, Ph. kilmeri, Ph. okutanii, Ph. soyoae and Pl. stearnsii on symbiont tree | 0.0001 | −79404.52 | 40.944 | |
| Constraint topology including Ab. mariana and I. fossajaponicum on symbiont tree | 0.0000 | −79624.583 | 136.516 | |
| Constraint topology including Ab. mariana, Ab. phaseoliformis and I. fossajaponicum on symbiont tree | 0.0000 | −79624.675 | 136.556 | |
| Best symbiont topology | 0.0002 | −79715.059 | 175.809 | |
| (B) Symbiont | Best symbiont topology | 1.0000 | −61438.662 | – |
| Constraint topology including Ak. kawamurai, Ph. kilmeri, Ph. okutanii and Ph. soyoae on host tree | 0.0160 | −61448.226 | 9.564 | |
| Constraint topology including Ak. kawamurai, C. laubieri, Ph. kilmeri, Ph. okutanii and Ph. soyoae on host tree | 0.0190 | −61448.765 | 10.104 | |
| Constraint topology including Ab. mariana and Ab. phaseoliformis on host tree | 0.0000 | −61837.220 | 398.559 | |
| Constraint topology including Ab. mariana, Ab. phaseoliformis, C. fausta and C. pacifica on host tree | 0.0000 | −63718.042 | 2279.381 | |
| Constraint topology including Ab. mariana, Ab. phaseoliformis, C. fausta, C. pacifica and I. fossajaponicum on host tree | 0.0000 | −64030.448 | 2591.786 | |
| Constraint topology including Ak. kawamurai, Ab. mariana, Ab. phaseoliformis, C. fausta, C. laubieri, C. magnifica, C. pacifica, I. fossajaponicum, Ph. kilmeri, Ph. okutanii and Ph. soyoae on host tree | 0.0000 | −64040.829 | 2602.167 | |
| Constraint topology including Ak. kawamurai, Ab. mariana, Ab. phaseoliformis, C. fausta, C. laubieri, C. magnifica, C. pacifica, I. fossajaponicum, Ph. kilmeri, Ph. okutanii, Ph. soyoae and Pl. stearnsii on host tree | 0.0000 | −64041.791 | 2603.129 | |
| Best host topology | 0.0000 | −64041.88 | 2603.218 | |
The AU test was performed in CONSEL. BF test was estimated by MrBayes.
Approximately unbiased test.
Bayes factor test.
Coevolution Analyses
To estimate the most probable coevolution scenario between the host clams and their symbionts, two reconciliation methods, the Jane 4.0 (Conow etal. 2010) and CoRe-PA (Merkle etal. 2010) were used. These topology-based programs assign costs to four coevolution events (cospeciation, duplication, sorting or loss, host switching) that link the two trees. In addition, Jane 4.0 also assigns a cost to failure-to-diverge events (a host speciates while the symbiont does not speciate and remains in both new host species). CoRe-PA calculates the optimal cost values, whereas Jane 4.0 requires the cost values to be inputted. Both methods seek to minimize the total costs of all events that occur. After symbiont phylogeny was mapped onto host phylogeny, several coevolutionary phylogenetic patterns exhibiting minimal total cost were retrieved. Our criteria for selecting a result in coevolutionary phylogenetic patterns took into account only cospeciation and host switching events, as sorting, duplication, and failure-to-diverge events of the symbiont are not expected in a comparison of phylogenetic trees. For the Jane 4.0 analysis, the default cost setting was used, and mapping was performed with 100 generations and a population size of 100. CoRe-PA was performed with automatic estimation of the optimal cost setting and computed reconstructions of 100,000 random data sets. The event costs obtained by CoRe-PA were 0.0999 for cospeciation, 0.3000 for sorting, 0.2999 for duplication, and 0.3000 for host switching.
Results
Phylogeny of Host Vesicomyid Clams
Most clades in the phylogenetic tree reconstructed from the concatenated gene data sets (10,469 nucleotide sites of 11 mitochondrial genes from 18 taxa) could be resolved with high statistical support (fig. 1A). Six species (Ph. soyoae, Ph. kilmeri, Ph. okutanii, Ak. kawamurai, C. laubieri, and C. magnifica) formed a monophyletic clade with ML BS of 58% and Bayesian PP of 0.9959 (fig. 1A; clade a). Two monophyletic clades that included either C. pacifica and C. fausta (fig. 1A; clade b), or Ab. phaseoliformis and Ab. mariana (fig. 1A; clade c) were highly supported with ML BS of 98–100% and Bayesian PP of 1.0 (fig. 1A), and they formed a monophyletic clade (fig. 1A; clade d). Isorropodon fossajaponicum was branched with clade d with ML BS of only 45% and Bayesian of PP of 0.9848 (fig. 1A; clade e). Calyptogena nautilei was the earliest branching lineage within the vesicomyid clam radiation, with ML BS of 100% and PP of 1.0, and the next most basal taxon was Pl. stearnsii with ML BS of 80% and PP of 1.0.
Fig. 1.
—Comparison of Maximum Likelihood (ML) phylogenies for 13 species of vesicomyid clams and their symbionts. (A) ML tree of vesicomyid clams based on the concatenated data sets of nucleotide sequences of 11 mitochondrial genes (atp6, atp8, cox1, cox2, cox3, cob, nad1, nad3, nad4, nad5, and rrnL). Venerid species (Meretrix lusoria, M. meretrix, M. petechialis, Paphia euglypta, and Venerupis philippinarum) were used as outgroups. (B) ML phylogenetic tree of 13 vesicomyid symbionts based on concatenated data sets of the nucleotide sequences of eight genes (16S rRNA, 23S rRNA, galU, groEL, groES, mfd, uvrA, and uvrD paralog). The symbiont of Bathymodiolus septemdierum was used as an outgroup. Dotted lines between vesicomyid clam species and their symbionts’ trees indicate symbiotic pairs. Numbers at the bipartitions indicate ML bootstrap supports of < 100% (left) and Bayesian posterior probabilities of < 1.0 (right). Unlabeled nodes are fully supported (100% bootstrap support and posterior probability at 1.0). Small letters in the squares of some nodes indicate the monophyletic clades described in the text. The orange branches are those exhibiting incongruence between the topologies of the host clams and symbionts, as indicated by an approximately unbiased (AU) test (P < 0.05) and Bayes factor test (>5). Host and symbiont trees were not drawn to the same rate scale, and scale bars indicate the rate of internal nodes except for outgroups.
Phylogeny of Vesicomyid Symbionts
The phylogenetic tree of the 13 vesicomyid symbionts based on the concatenated gene data set (15,847 nucleotide sites of eight symbiont genes from 14 taxa) is shown in figure 1B. The phylogenetic lineages were well resolved at most nodes, showing high statistical support with ML BS of 96–100% and Bayesian PP of 0.9997–1.0 (fig. 1B). The symbionts of Ph. soyoae, Ph. kilmeri, Ph. okutanii, Ak. kawamurai, C. laubieri, and C. magnifica (fig. 1B; clade a); those of C. pacifica, C. fausta, Pl. stearnsii, and C. nautilei (fig. 1B; clade b); and those of Ab. mariana, I. fossajaponicum, and Ab. phaseoliformis (fig. 1B; clade c), respectively, formed monophyletic clades supported with ML BS of 71–100% and Bayesian PP of 0.9681–1.0 (fig. 1B). The affiliation of the symbiont of Pl. stearnsii with the symbionts of C. pacifica and C. fausta was low (ML BS of 75% and a Bayesian PP of 0.8440) (fig. 1B; clade d).
Comparison of the Host and Symbiont Phylogenetic Trees
A comparison of the host and symbiont phylogenetic trees showed that the phylogenetic lineages of six hosts (Ph. soyoae, Ph. kilmeri, Ph. okutanii. C. magnifica, C. pacifica, and C. fausta) and of their corresponding symbionts were congruent (fig. 1; black branches). However, those of seven other hosts (Ak. kawamurai, C. laubieri, Ab. phaseoliformis, Ab. mariana, I. fossajaponicum, Pl. stearnsii, and C. nautilei) and their symbionts were not congruent (fig. 1; orange branches). The results of the AU and Bayes factors tests of both constrained and unconstrained topologies in host and symbiont rejected the phylogenetically congruent positions of the seven host-symbiont pairs (P < 0.05; BF > 5; table 3 and fig. 1; orange branches).
Coevolution analyses between host clams and their symbionts were performed with Jane 4.0 and CoRe-PA. The Jane 4.0 analysis estimated three coevolution patterns, of which two were selected by our criteria described in the Materials and Methods section (fig. 2A and B). CoRe-PA yielded six coevolution patterns, only one of which was selected by our criteria; it exhibited the same coevolution pattern as one of the two patterns generated by Jane 4.0 (fig. 2A).
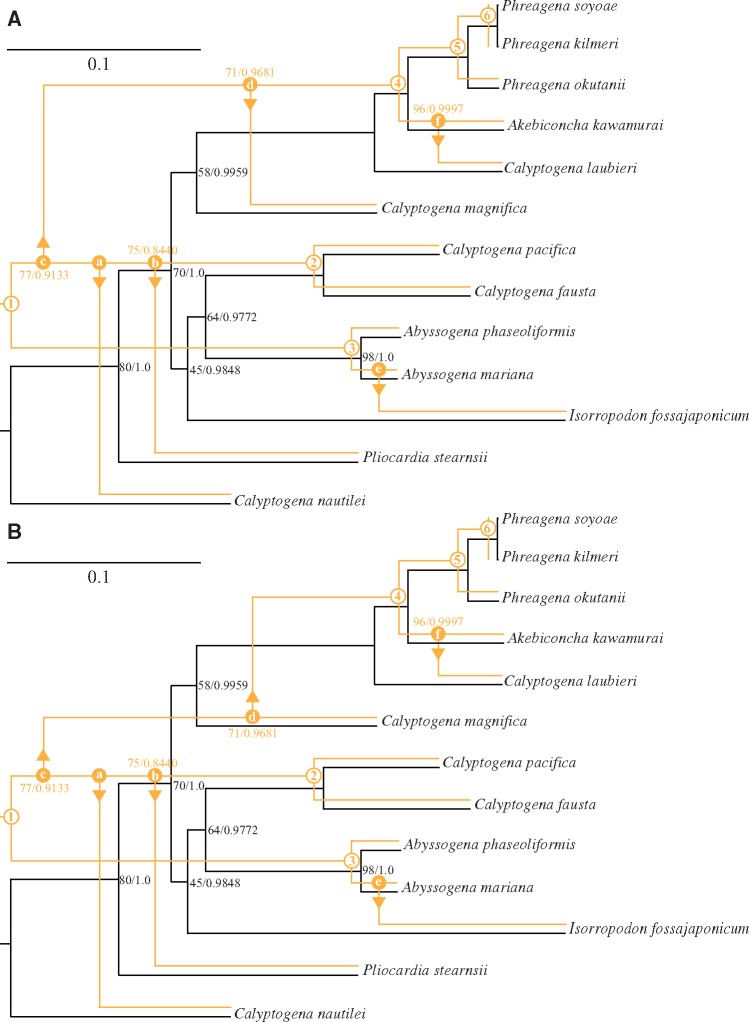
Fig. 2.
—Coevolution scenarios between the vesicomyid clams and their symbionts. The coevolution analysis was performed using two reconciliation methods, the Jane 4.0 and CoRe-PA. Two preferred coevolutionary patterns exhibiting minimal total cost were selected, (A) Jane 4.0 and CoRe-PA, and (B) Jane 4.0. The black and orange topologies indicate the topologies of the hosts and symbionts, respectively, and the black and orange numbers at the bipartitions indicate the ML bootstrap supports of < 100% (left) and Bayesian posterior probabilities of < 1.0 (right) on host and symbiont trees, respectively. Unlabeled nodes are fully supported (100% bootstrap support and posterior probability of 1.0). The numbers in the orange open circles and small letters in the orange closed circles indicate cospeciation and host switching events, respectively.
The two selected coevolution patterns were essentially similar (fig. 2A and B) and contained six cospeciation events and six host switching events (fig. 2A and B). The six cospeciation events (fig. 2A and B; open circles No. 1–6) and four host switching events (fig. 2A and B; closed circles a, b, e, and f) in the two selected coevolution patterns were identical. Among the six cospeciation events, three (fig. 2A and B; open circles No. 2, 5, and 6) were also supported by phylogenetic trees (fig. 1); however, the cospeciation of Ab. phaseoliformis (fig. 2A and B; open circle No. 3) was not supported by the phylogenetic trees (fig. 1). Among the four host switching events, two were suggested in the lineage of the common ancestral symbiont of C. pacifica and C. fausta. Our analyses indicated that the common ancestral symbiont of these two clam species shifted to two other clam species as hosts: C. nautilei or its ancestor (fig. 2A and B; closed circle a), and P. stearnsii or its ancestor (fig. 2A and B; closed circle b). The remaining two host switching events were suggested to have occurred in the lineage of the ancestral symbionts of Ak. kawamurai and Ab. mariana. The ancestral symbionts of these two clam species shifted to other clam species as hosts, C. laubieri and I. fossajaponicum, respectively (fig. 2A and B; closed circles e and f).
The host switching events associated with the common ancestor of the symbionts of C. pacifica and C. fausta were suggested to be different between two coevolution patterns. Based on figure 2A, the ancestral symbiont of these clam species shifted to the common ancestor of the lineage containing Ph. soyoae, Ph. kilmeri, Ph. okutanii, Ak. kawamurai, and C. laubieri (fig. 2A; closed circle c). After this event, an additional host switching event, to C. magnifica, was estimated to have occurred in this lineage (fig. 2A; closed circle d). On the other hand, figure 2B suggests that the common ancestor of symbionts of C. pacifica and C. fausta first shifted to the lineage of C. magnifica (fig. 2B; closed circle c) and then secondarily to the ancestral lineage of Ph. soyoae, Ph. kilmeri, Ph. okutanii, Ak. kawamurai, and C. laubieri (fig. 2B; closed circle d).
Discussion
Occasional Host Switching of the Symbionts of Vesicomyid Clams
The phylogenetic positions of the clams selected in the present study are widely distributed in a previously published phylogenetic tree that was based on sequences of the mitochondrial cox1 gene (Decker etal. 2012). In the present study, we used large gene data sets (over 10 kbp) to enhance the resolution of the phylogenetic trees of vesicomyid clams and their symbionts, an improvement over previously reported phylogenetic trees (Stewart etal. 2008; fig. 1). The phylogenetic positions of six vesicomyid hosts (Ph. soyoae, Ph. kilmeri, Ph. okutanii. C. magnifica, C. pacifica, and C. fausta) and their symbionts were congruent (fig. 1). Cospeciation of these host-symbiont pairs was also well supported by coevolution analyses using Jane 4.0 and CoRe-PA, except in C. magnifica (fig. 2). However, the tree topologies of seven other host taxa (Ak. kawamurai, C. laubieri, Ab. phaseoliformis, Ab. mariana, I. fossajaponicum, Pl. stearnsii, and C. nautilei) and their symbionts were incongruent, and such differences were statistically supported by AU and Bayes factor tests (table 3 and fig. 1). Our coevolution analyses using Jane 4.0 and CoRe-PA estimated that multiple host switching events have occurred. Remarkably, the common ancestral lineage of the symbionts of C. fausta and C. pacifica was repeatedly associated with such events (fig. 2). Possible host switching may also have occurred between the host lineages of I. fossajaponicum and Ab. mariana and between those of Ak. kawamurai and C. laubieri. Indeed, recent studies have reported the possibility of horizontal transfers of symbionts in vesicomyid clams, based on the incongruence between host and symbiont phylogenies and detection of symbionts from other host species along with its own symbiont in the gill using in situ hybridization (Stewart etal. 2008; Decker etal. 2013). Therefore, it seems likely that vesicomyid symbionts are predominantly transmitted vertically but also are transmitted horizontally on occasion. The present study also suggests this hypothesis.
Possible Mechanism of Horizontal Transfer Causing Host Switching
Stewart etal. (2008) proposed three mechanisms (i.e., host hybridization, environmental acquisition, and horizontal transmission) that could facilitate the horizontal acquisition (nonmaternal transmission) of symbionts that could trigger host switching and result in incongruent tree topologies of vesicomyid clams and their symbionts. The host-hybridization hypothesis states that both the mitochondria and symbionts could be paternally transferred via sperm during hybridization between distinct species. In this scenario, hybrid hosts with different symbionts and mitochondrial phylotypes would occur by the displacement or loss of the maternal symbiont or mitochondrion, causing changes in the phylogenetic positions of the resulting symbiont and mitochondrial types. Several bivalve species of Heterodonta, to which vesicomyid clams belong, have a sperm-transmitted mitochondrial genome, along with a standard maternally transmitted one. This mechanism is called doubly uniparental inheritance of mitochondria (Skibinski etal. 1994; Zouros etal. 1994). Bivalve species with doubly uniparental inheritance have two distinct sex-specific mitochondrial DNA types (female-type and male-type) that potentially lead to misunderstanding regarding their evolution (Doucet-Beaupré etal. 2010). The paternal transmission of symbionts has been observed in Rickettsia, a facultative symbiont of insects (Watanabe etal. 2014). The symbiont 16S rRNA sequence was not amplified from the testis of Ph. okutanii, suggesting that vesicomyid sperm does not harbor the symbiont (Ikuta etal. 2016a). Paternal symbiont or mitochondrial transfer has not been reported in vesicomyids (Stewart etal. 2008). These data suggest that neither the transmission of mitochondria nor that of symbionts is likely to occur paternally. Thus, the horizontal transfer of mitochondria and symbionts is unlikely to have occurred through hybridization.
The second hypothesis, the environmental acquisition hypothesis, states that symbionts are directly acquired from the environment. However, the genomes of symbionts from Ph. okutanii and C. magnifica (Kuwahara etal. 2007; Newton etal. 2007) lack genes associated with flagella for motility in the environment and genes for the related type IV secretory system for infection into host cells (Kuwahara etal. 2007). Moreover, vesicomyid symbionts have not been found in bacterial mats or sediments in their hosts’ habitats (Moyer etal. 1995; Polz and Cavanaugh 1995). Therefore, this hypothesis is also unlikely to explain the observed patterns.
The horizontal transfer hypothesis states that symbionts are transmitted between distinct hosts without hybridization. Horizontal transmission could occur through the direct contact between eggs and eggs, eggs and host, or released symbionts and hosts. Symbionts that have no infection system are unlikely to be transferred into other host tissues via direct contact. Moreover, horizontal transmission of symbionts in the gill epithelial cells of host clam is also unlikely, because of their intracellular localization. Recently, Ikuta etal. (2016a) reported that in Ph. okutanii, ∼400 symbiont cells are located at the external surface of eggs. The symbiont cells are localized at the egg surface in the vitelline coat, which may prevent the symbionts from making direct contact with other eggs or hosts, reducing the frequency of host switching and making such horizontal transmission occasional and historical (Ikuta etal. 2016a). In host switching by direct contact events, horizontally transmitted symbionts must enter the maternal germ cells (such as eggs) of the new host to transfer the symbiont to the next generation. The process by which the symbiont enters both the maternal germ cells and gill epithelial cells may occur during the developmental stage of the host. Thus, among the three hypotheses proposed by Stewart etal. (2008), the most plausible mechanism for the horizontal transfer of symbionts in vesicomyid clams is that the symbionts on the egg surface are transferred by direct contact of eggs of different clam species before the developmental process begins.
The Process of Host Switching
Horizontal transfer of the symbiont from the original host to a new host is considered to be the first step of a host switching event. After the horizontal transfer of a symbiont occurs, the original symbiont may be eliminated by evolutionary change or selection through a possible bottleneck of vertical transmission via egg, and then the horizontally transmitted symbiont speciates in its new host. Recently, Decker etal. (2013) reported that each of two clam species possesses a small population of the symbiont from the other respective species, along with its own symbiont, when the hosts cooccur within the same colony. Cooccurrence of multiple symbionts in a single individual may represent an initial process of host switching via horizontal transfer by direct contact. Moreover, the physical proximity of distinct vesicomyid clam species likely facilitates horizontal symbiont transfer (Decker etal. 2013). Two vesicomyid species, Ph. okutanii and Ph. soyoae, that coexist in a seep colony in Sagami Bay (Kojima and Ohta, 1997) exhibit synchronized spawning triggered by an increase in water temperature (Fujikura etal. 2007). Although there is currently no evidence of host switching between Ph. okutanii and Ph. soyoae, it is possible that the extracellular localization of symbionts, such as on the surface of eggs, could facilitate the horizontal transmission of symbionts between cooccurring species. Of the 13 vesicomyids in the present study, C. pacifica, Ph. kilmeri, and Pl. stearnsii are known to cooccur in colonies in Monterey Bay (Barry etal. 2007), and data from the present study suggest that host switching between C. pacifica and Pl. stearnsii occurred. Other vesicomyids that harbor possible host-switching symbionts are currently isolated geographically; however, it is possible that these clam species lived in close proximity in the past.
Conclusion
Previous studies have proposed that the intracellular symbionts of vesicomyid clams are vertically (maternally) transmitted to offspring and that the hosts and symbionts cospeciated (Endow and Ohta 1990; Cary and Giovannoni 1993; Peek etal. 1998; Ikuta etal. 2016a). However, several recent studies have suggested that the vesicomyid and symbiont phylogenies are decoupled and that at least some symbionts are horizontally transmitted (Stewart etal. 2008; Decker etal. 2013). In the present study, to more closely investigate the coevolution of these vesicomyid clams and their symbionts, the phylogenetic trees were reconstructed using large gene data sets from both the host clam species and their symbionts. Our results suggest that multiple host switching events are highly likely to have occurred during the early evolution of vesicomyids, and such host switching events were most likely mediated by horizontal egg-to-egg transmission. Our data provide strong support for the hypothesis that vesicomyid symbionts are predominantly transmitted vertically but are occasionally transmitted horizontally. These findings contribute to the understanding of the evolutionary process of intracellular symbiosis by which symbionts are generally considered to be vertically transmitted.
Supplementary Material
Supplementary data are available at Genome Biology and Evolution online.
Supplementary Material
Acknowledgments
We are grateful to the captains and crews of the research vessels Yokosuka, Natsushima, and Kairei, and to the operation teams of the DSV Shinkai 6500, the ROV Hyper-Dolphin, and the ROV Kaiko, which were used to collect deep-sea bivalves for their assistance during the JAMSTEC Yokosuka, Natsushima, and Kairei cruises. We would also like to thank Dr. T. Okutani for helpful discussion about bivalve classification. This work was supported by a grant from the Japanese Society for the Promotion of Science, awarded to T.Y. (JSPS KAKENHI Grant Number JP24570252).
Literature Cited
- Barry JP, Whaling PJ, Kochevar RK.. 2007. Growth, production, and mortality of the chemosynthetic vesicomyid bivalve, Calyptogena kilmeri from cold seeps off central California. Mar Ecol. 28(1):169–182. [Google Scholar]
- Bright M, Bulgheresi S.. 2010. A complex journey: transmission of microbial symbionts. Nat Rev Microbiol. 8(3):218–230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T.. 2009. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25(15):1972–1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cary SC, Giovannoni SJ.. 1993. Transovarial inheritance of endosymbiotic bacteria in clams inhabiting deep-sea hydrothermal vents and cold seeps. Proc Natl Acad Sci USA. 90(12):5695–5699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conow C, Fielder D, Ovadia Y, Libeskind-Hadas R.. 2010. Jane: a new tool for the cophylogeny reconstruction problem. Algorithms Mol Biol. 5:16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Decker C, Olu K, Arnaud-Haond S, Duperron S.. 2013. Physical proximity may promote lateral acquisition of bacterial symbionts in vesicomyid clams. PLoS One 8(7):e64830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Decker C, Olu K, Cunha RL, Arnaud-Haond S.. 2012. Phylogeny and diversification patterns among vesicomyid bivalves. PLoS One 7(4):e33359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doucet-Beaupré H, et al. 2010. Mitochondrial phylogenomics of the Bivalvia (Mollusca): searching for the origin and mitogenomic correlates of doubly uniparental inheritance of mtDNA. BMC Evol Biol. 10:50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubilier N, Bergin C, Lott C.. 2008. Symbiotic diversity in marine animals: the art of harnessing chemosynthesis. Nat Rev Microbiol. 6:725–740. [DOI] [PubMed] [Google Scholar]
- Endow K, Ohta S.. 1990. Occurrence of bacteria in the primary oocytes of vesicomyid clam Calyptogena soyoae. Mar Ecol Prog Ser. 64:309–311. [Google Scholar]
- Fujikura K, et al. 2007. Long-term in situ monitoring of spawning behavior and fecundity in Calyptogena spp. Mar Ecol Prog Ser. 333:185–193. [Google Scholar]
- Ikuta T, et al. 2016a. Surfing the vegetal pole in a small population: extracellular vertical transmission of an ‘intracellular’ deep-sea clam symbiont. R Soc Open Sci. 3(5):160130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikuta T, et al. 2016b. Heterogeneous composition of key metabolic gene clusters in a vent mussel symbiont population. ISME J. 10(4):990–1001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kass RE, Raftery AE.. 1995. Bayes factors. J Am Stat Assoc. 90(430): 773–795. [Google Scholar]
- Katoh K, Standley DM.. 2013. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 30(4):772–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kojima S, Ohta S.. 1997. Calyptogena okutanii n. sp., a sibling species of Calyptogena soyoae Okutani, 1957 (Bivalvia: Vesicomyidae). Venus 56:189–195. [Google Scholar]
- Krylova EM, Sahling H.. 2010. Vesicomyidae (Bivalvia): current taxonomy and distribution. PLoS One 5(4):e9957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuwahara H, et al. 2007. Reduced genome of the thioautotrophic intracellular symbiont in a deep-sea clam, Calyptogena okutanii. Curr Biol. 17(10):881–886. [DOI] [PubMed] [Google Scholar]
- Kuwahara H, et al. 2011. Loss of genes for DNA recombination and repair in the reductive genome evolution of thioautotrophic symbionts of Calyptogena clams. BMC Evol Biol. 11:285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lane DJ. 1991. 16S/23S rRNA sequencing In:Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. . New York: John Wiley and Sons; p. 115–175. [Google Scholar]
- Le Pennec M, Beninger PG, Herry A.. 1995. Feeding and digestive adaptations of bivalve molluscs to sulphide-rich habitats. Comp Biochem Physiol A Physiol. 111(2):183–189. [Google Scholar]
- Liu H, Cai S, Zhang H, Vrijenhoek RC.. 2016. Complete mitochondrial genome of hydrothermal vent clam Calyptogena magnifica. Mitochondrial DNA A 27(6):4333–4335. [DOI] [PubMed] [Google Scholar]
- Merkle D, Middendorf M, Wieseke N.. 2010. A parameter-adaptive dynamic programming approach for inferring cophylogenies. BMC Bioinformatics 11(Suppl 1):S60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moyer CL, Dobbs FC, Karl DM.. 1995. Phylogenetic diversity of the bacterial community from a microbial mat at an active, hydrothermal vent system, Loihi Seamount, Hawaii. Appl Environ Microbiol. 61(4):1555–1562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newton IL, et al. 2007. The Calyptogena magnifica chemoautotrophic symbiont genome. Science 315(5814):998–1000. [DOI] [PubMed] [Google Scholar]
- Ozawa G, et al. 2017. Updated mitochondrial phylogeny of Pteriomorph and Heterodont Bivalvia, including deep-sea chemosymbiotic Bathymodiolus mussels, vesicomyid clams and the thyasirid clam Conchocele cf. bisecta. Mar Genomics 31:43–52. [DOI] [PubMed] [Google Scholar]
- Peek AS, Feldman RA, Lutz RA, Vrijenhoek RC.. 1998. Cospeciation of chemoautotrophic bacteria and deep sea clams. Proc Natl Acad Sci USA. 95(17):9962–9966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polz MF, Cavanaugh CM.. 1995. Dominance of one bacterial phylotype at a Mid-Atlantic Ridge hydrothermal vent site. Proc Natl Acad Sci USA. 92(16):7232–7236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rambaut A, Drummond AJ.. 2009. Tracer v1.5, URL. [Accessed June 30, 2017]. Available from: http://tree.bio.ed.ac.uk/software/tracer/.
- Rice P, Longden I, Bleasby A.. 2000. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 16(6):276–277. [DOI] [PubMed] [Google Scholar]
- Ronquist F, et al. 2012. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 61(3):539–542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimamura S, et al. 2017. Loss of genes related to nucleotide excision repair (NER) and implications for reductive genome evolution in symbionts of deep-sea vesicomyid clams. PLoS One 12(2): e0171274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimodaira H. 2002. An approximately unbiased test of phylogenetic tree selection. Syst Biol. 51(3):492–508. [DOI] [PubMed] [Google Scholar]
- Shimodaira H, Hasegawa M.. 2001. CONSEL: for assessing the confidence of phylogenetic tree selection. Bioinformatics 17(12):1246–1247. [DOI] [PubMed] [Google Scholar]
- Skibinski DOF, Gallagher C, Beynon CM.. 1994. Mitochondrial DNA inheritance. Nature 368(6474):817–818. [DOI] [PubMed] [Google Scholar]
- Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart FJ, Young CR, Cavanaugh CM.. 2008. Lateral symbiont acquisition in a maternally transmitted chemosynthetic clam endosymbiosis. Mol Biol Evol. 25(4):673–687. [DOI] [PubMed] [Google Scholar]
- Tanabe AS. 2011. Kakusan4 and Aminosan: two programs for comparing nonpartitioned, proportional and separate models for combined molecular phylogenetic analyses of multilocus sequence data. Mol Ecol Resour. 11(5):914–921. [DOI] [PubMed] [Google Scholar]
- Watanabe K, et al. 2014. Intrasperm vertical symbiont transmission. Proc Natl Acad Sci USA. 111(20):7433–7437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Werle E, Schneider C, Renner M, Völker M, Fiehn W.. 1994. Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res. 22(20):4354–4355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Won YJ, Jones WJ, Vrijenhoek RC.. 2008. Absence of cospeciation between deep-sea mytilids and their thiotrophic endosymbionts. J Shellfish Res. 27:129–138. [Google Scholar]
- Zouros E, Ball AO, Saavedra C, Freeman KR.. 1994. Mitochondrial DNA inheritance. Nature 368(6474):818. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.