ABSTRACT
Efficient protein secretion is often a valuable alternative to classic cellular expression to obtain homogenous protein samples. Early on, bacterial type I secretion systems (T1SS) were employed to allow heterologous secretion of fusion proteins. However, this approach was not fully exploited, as many proteins could not be secreted at all or only at low levels. Here, we present an engineered microbial secretion system which allows the effective production of proteins up to a molecular mass of 88 kDa. This system is based on the hemolysin A (HlyA) T1SS of the Gram-negative bacterium Escherichia coli, which exports polypeptides when fused to a hemolysin secretion signal. We identified an A/U-rich enhancer region upstream of hlyA required for effective expression and secretion of selected heterologous proteins irrespective of their prokaryotic, viral, or eukaryotic origin. We further demonstrate that the ribosomal protein S1 binds to the hlyA A/U-rich enhancer region and that this region is involved in the high yields of secretion of functional proteins, like maltose-binding protein or human interferon alpha-2.
IMPORTANCE A 5′ untranslated region of the mRNA of substrates of type I secretion systems (T1SS) drastically enhanced the secretion efficiency of the endogenously secreted protein. The identification of ribosomal protein S1 as the interaction partner of this 5′ untranslated region provides a rationale for the enhancement. This strategy furthermore can be transferred to fusion proteins allowing a broader, and eventually a more general, application of this system for secreting heterologous fusion proteins.
KEYWORDS: 5′ untranslated region, hemolysins, mRNA, protein-mRNA interaction, protein secretion
INTRODUCTION
The Gram-negative bacterium Escherichia coli is used as the organism of choice for the production of recombinant proteins (1), and the FDA has approved more than 150 recombinant peptide- or protein-based therapeutics of which one-third are produced in E. coli (2). Polypeptide expression and secretion into the extracellular media represent a valuable alternative to the classical approach of expression within the cytosol (3–5). Especially, product titers can be impressive, reaching up to 30 g/liter (5, 6). However, these titers are usually only obtained for homologous proteins in optimized processes and are uncommon for heterologous protein secretion (6–10).
The secretion strategy described herein is based on the hemolysin A (HlyA) type I secretion system (T1SS) of E. coli (11). HlyA is a member of the repeats-in-toxin (RTX) family that forms pores in the cell membrane of human host cells, resulting in lysis. Early on, HlyA was recognized as an interesting carrier for the secretion of heterologous proteins, including vaccines (12, 13). Despite great efforts, until now, an application as a secretion platform has not been achieved, and the majority of polypeptides are secreted with only low yields of approximately 100 μg/liter, if at all (12, 14–16).
The hly operon of E. coli comprises hlyC, hlyA, hlyB, and hlyD (17) (Fig. 1a). The cytoplasmic protein HlyC is required for a posttranslational modification, i.e., the acylation of HlyA at two lysine residues (Lys564 and Lys690) (18, 19). This modification converts the inactive HlyA into an active toxin but is dispensable for efficient secretion (20). hlyB and hlyD encode an ABC transporter and a periplasmic adapter protein, respectively (11), which form the inner membrane complex of the T1SS. Its interaction with HlyA recruits the multifunctional outer membrane protein TolC, completing the secretion apparatus (21). This complex catalyzes the transport of HlyA from the cytoplasm across the two membranes into the extracellular space, without any detectable periplasmic intermediate(s) (22). The C terminus of hemolysin A contains the secretion sequence (approximately the last 60 amino acids) (18) and upstream six repeats of a nonapeptide sequence that binds Ca2+ in the extracellular space, triggering protein folding (23). Secretion of the hemolysin A type I secretion system is independent of substrate size and operates at a rate of approximately 16 amino acids per transporter per second (24). Furthermore, it was shown that the secretion efficiency and Ca2+ binding affinity of the C-terminal fragment of HlyA (amino acids 807 to 1024; around 24 kDa) called HlyA1 is comparable to full-length HlyA (18). Furthermore, HlyA1 does not possess any of the toxic properties of HlyA, and this signal sequence domain has been extensively used for the secretion of various heterologous proteins (12, 16, 25, 26).
FIG 1.
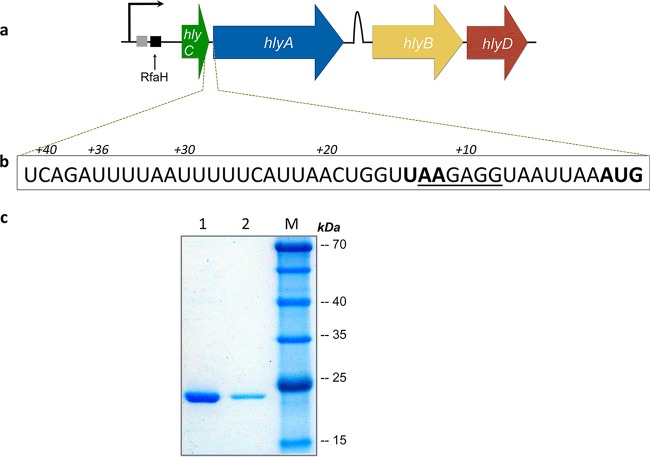
Schematic representation of the hly operon and the effect of the enhancer region on the expression of HlyA1. (a) The hly operon consists of four genes in the order hlyC, hlyA, hlyB, and hlyD (73). Downstream of the promoter (depicted with a black arrow), a JUMPstart (shown in gray box) (28) and an ops sequence (represented by a black box) (29) are present. The stem-loop termination sequence located between the hlyA and hlyB genes is shown as a kink. (b) The AU-rich upstream region of hlyA is highlighted in the box. The start codon of hlyA (AUG) and the stop codon of hlyC (UAA) are shown in bold, while the ribosome-binding site is underlined. (c) SDS-PAGE analysis of nonconcentrated supernatant obtained after expression of hlyA1 gene containing either 159 (lane 1) or 17 (lane 2) nucleotides upstream of the hlyA region and a molecular weight marker (lane M).
In E. coli, mRNA production of the components of the hly operon employ the anti-termination factor RfaH (27) to read through the termination signal that is located between the hlyA and hlyB genes (Fig. 1a). It is proposed that the RNA polymerase stops at the JUMPstart (just upstream of many polysaccharide-associated starts) sequence, which allows binding of RfaH (28). The operon-polarity suppressor (ops) sequence serves the same function (29). Binding of RfaH allows readthrough of a stem-loop termination sequence (30) (Fig. 1a), which is located between the hlyA and hlyB genes, suggesting that mRNA production of the acylase gene hlyC and the structural gene hlyA are uncoupled from the expression of hlyB and hlyC in the absence of RfaH (28). However, the exogenous signal(s) that induce production of HlyA in the stationary phase are currently unknown.
In general, expression of high-abundance proteins, like phage coat proteins (31), ATP synthase proteins (32), translational factors (31), or nucleoid proteins (33), is accompanied by an A/U-rich region upstream of the translation initiation codon and the ribosome-binding site, and this region acts as an enhancer by attracting ribosomal protein S1, also known as RpsA. Except for leaderless mRNAs that lack a 5′ untranslated region (5′ UTR), S1 is required for the translation of almost all mRNAs in E. coli (34), and loss-of-function mutations of RpsA are lethal for E. coli (35); even S1 depletion and overexpression cause cell growth impairment (36). Ribosomal protein S1 is the largest ribosomal protein with high affinity for mRNA (37, 38) and is only weakly associated with the ribosome (39). It associates late during assembly of the 30S ribosomal subunit (40) and serves as a catching arm for mRNA (31). Besides having a critical role in translation, S1 has been implicated in several other cellular processes, such as enhancement of transcription through transcriptional cycling (41), protection of single-stranded RNA from degradation by RNase E (42), stabilization of mRNA (43), rescue of stalled ribosomes (44), and promotion of RNA unwinding by binding 10 nucleotides of RNA in a multistep fashion (45). It interacts with pyrimidine-rich regions in the 5′ UTR of mRNAs, which have been identified in cross-linking experiments (31). Free S1 activates the process of mRNA binding by 30S particles lacking S1 and acts as a translational repressor if added in molar excess over the 30S particles (31). A/U-rich sequences placed upstream of native or foreign genes provide a high efficiency of expression (31, 46, 47), where translation enhancers with a Shine-Dalgarno sequence regulate ribosomal liberation from translation initiation to determine the translation efficiency of the downstream localized coding region (47).
HlyA was recognized previously as an interesting carrier for the secretion of heterologous proteins (12), and its signal sequence domain (HlyA1) has been extensively used for the secretion of various heterologous proteins (12, 16, 25, 26). In principle, the fusion of the C terminus of a heterologous protein to HlyA1 would lead to its secretion in the presence of the inner membrane components HlyB and HlyD. However, the huge biotechnological relevance of this system has been limited due to the fact that not all of the proteins were efficiently secreted (16). Thus, we were interested in exploiting and optimizing the HlyA T1SS for the production of heterologous proteins. Here, we report a hemolysin-specific A/U-rich enhancer region, which we identified upstream of the hlyA start codon. The presence of this enhancer elevated the secretion levels. This is highlighted by our results that in combination with our engineered two-plasmid system, the HlyA-based secretion system including the enhancer sequence is now converted into a platform for the secretion of heterologous proteins in E. coli.
RESULTS
The upstream region of hlyA affects HlyA1 production.
For cloning and secretion studies, the C-terminal part of HlyA (HlyA1) derived from plasmid pSU2726 (48, 49) expressing full-length HlyA was used. While subcloning the hlyA gene into another vector, it was observed that the secretion of HlyA was comparably low. Sequence analysis of the parental vector revealed the presence of an additional sequence of 159 nucleotides (nt) derived from the hlyC gene (cloning artifact resulting from restriction sites) and noticed that this natural upstream region of hlyA (Fig. 1b) had a strong influence on the level of secretion.
With a 159-nucleotide (nt) 5′ sequence containing the 3′ end of the hlyC gene, a much greater amount of HlyA1 was secreted than with a shorter fragment of only 17 nt (Fig. 1c), indicating that this upstream sequence enhanced the expression and/or secretion by an unknown mechanism.
Known hemolysin regulatory proteins are not responsible for enhanced expression.
The hly operon is regulated by several mechanisms (30, 50–54). To test whether one of the known regulatory proteins is involved in the observed increased levels of HlyA1 secretion, we obtained E. coli mutant strains from the Coli Genetic Stock Center (CGSC) lacking individual factors previously shown to affect HlyA secretion and transformed the respective transporter plasmids together with either the hlyA1 expression plasmid containing the truncated hlyA upstream region of 17 nt or the plasmid with the long upstream region of 159 nt (Table 1). E. coli K-12 BW25113 (wild type) was used as a reference strain, which behaved like the expression strain E. coli BL21(DE3). Mutants lacking typical hemolysin regulators, such as the stationary-phase sigma factor RpoS (53) (JW5437-1), nucleoid-associated protein H-NS (53, 55) (JW1225-2), expression-modulating protein Hha (55) (JW0449-5), anti-terminator factor RfaH (28) (JW3818-1), and the small RNA DsrA (53, 56) (NM6003) did not show any significant differences in the levels of secretion compared to the reference strain. Mutants lacking proteins involved in posttranslational regulation or RNA stability, such as the chaperone Hfq (57–59) (JW41-30-1), RNase E (60) (N3431), or RNase R (JW5741-1), were also analyzed, and again, no significant differences in expression levels were observed (data not shown).
TABLE 1.
Sequences of hlyA enhancer regions used in this studya

Sequences of wild-type, truncated, and mutated hlyA enhancer regions are shown. The RBS is underlined, part of the original enhancer sequence up to 37 nt upstream is depicted in bold.
Identification of an A/U-rich enhancer region upstream of hlyA.
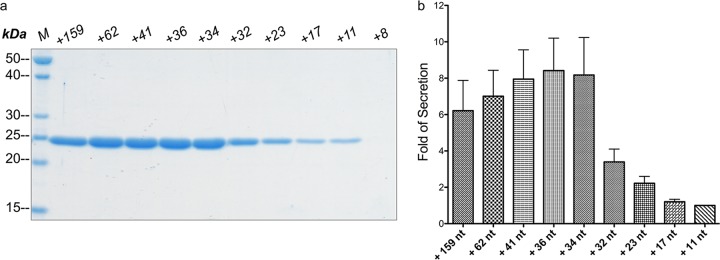
Next, the 5′ upstream region of hlyA was analyzed in more detail by conducting a comprehensive mutational analysis (Fig. 2 and Table 1). Secretion experiments with HlyA1 combined with 5′ truncations of various lengths uncovered that an upstream region of at least 34 nt is responsible and required for optimal high-level production. Strikingly, this region is rich in the nucleotides adenine and uracil (Fig. 1b). As expected, the strain without a ribosome-binding site (termed +8) did not express detectable levels of HlyA1. However, elongation of the 5′ sequence to 11 nucleotides or 17 nt showed at least detectable levels of secretion (Fig. 2a). Conversely, elongation of the 5′ sequence to at least 34 nucleotides increased secretion significantly to approximately 8-fold, but further elongation did not improve secretion (Fig. 2).
FIG 2.
The hemolysin A enhancer region. (a) SDS-PAGE of nonconcentrated supernatant from E coli strains carrying hlyA1 with 5′ enhancer region of different lengths. (b) Quantification (fold change; analyzed with ImageJ) of the amount of HlyA1 secreted in the supernatant deduced from a representative gel based on the lowest detectable expression level (+11). Error bars represent the standard deviation of the results from at least three independent biological replicates.
Mutations within the enhancer sequence and the above-mentioned secretion experiments revealed that the uracils at positions +33 to +36 are important for the production of HlyA1 (see Fig. S1 in the supplemental material). Additionally, the substitution of adenine with thymine in the sequence between nucleotides +29 and +35 (AAAUUAA) and vice versa did not significantly impair production, indicating that no specific binding motif exists in this region. Taken together, this extended mutational analysis provides compelling evidence for the importance of the A/U-rich region for efficient expression and subsequent secretion of HlyA1.
Ribosomal protein S1 binds to the hlyA enhancer region.
As demonstrated above, an A/U-rich region upstream of the hlyA gene is involved in a strong enhancing effect on the production of HlyA1. A/U-rich sequences upstream of the translation initiation codon are known binding sites for the ribosomal protein S1 (31, 46, 47). Consequently, we investigated the potential binding of S1 to the 5′ upstream sequence of hlyA by incubating bead-immobilized RNA containing the enhancer region with E. coli cell extract. Subsequently, bound proteins were eluted from the beads and analyzed via SDS-PAGE. Here, only one prominent protein band was detected (Fig. 3, lane 1) and identified as RpsA-ribosomal protein S1 by liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis, proving binding of S1 to the A/U-rich region. Other proteins possibly binding to this region could not be detected.
FIG 3.
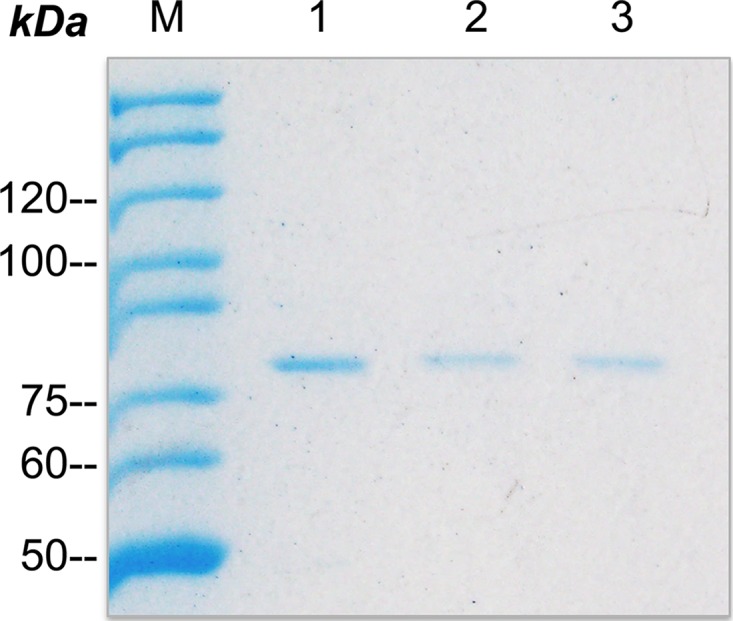
Pulldown assay with RNA. SDS-PAGE of eluted proteins of the hlyA-enhancer region immobilized to magnetic beads and the dedicated sequences of the low-expression strains +17 (lane 2) and 33u>g-34u>g (lane 3).
Furthermore, beads containing the RNA sequence encoding the complete enhancer region resulted in a detectable band of the S1 protein, whereas the RNA sequences derived from the strains displaying reduced secretion efficiencies (+17, Fig. 3, lane 2; and 33u>g-34u>g, Fig. 3, lane 3) resulted in protein bands of reduced intensity. This clearly implies that binding of ribosomal protein S1 to the A/U-rich enhancer region may result in the observed high-level expression and subsequent secretion of proteins. Moreover, the sequence changes result in reduced binding of S1 with the final consequence of decreased secretion levels.
Effect of the enhancer region on mRNA levels.
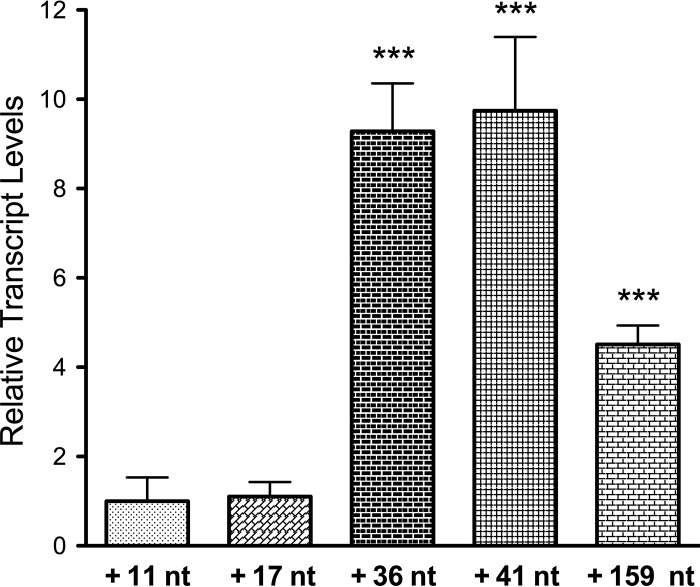
To determine whether the secretion level of HlyA1 based on the 5′ enhancer region is correlated with the mRNA levels, a quantitative PCR analysis of strains with 5′ truncations of different lengths was performed to test the expression profile of the hlyA1 gene. The E. coli housekeeping gene hcaT was used as the reference gene. The expression level of hlyA1 in strains containing the highly truncated 5′ enhancer regions (+11 nt and +17 nt) was significantly lower than the strains having +36 or +41 nt enhancer regions (Fig. 4). Furthermore, an increase of around 10-fold in terms of relative transcript levels of hlyA1 was observed in these strains (+36 nt and +41 nt), which had previously demonstrated increased secretion levels of HlyA1 (Fig. 2). Even for the longest extension (159 nt), an approximately 5-fold increase in mRNA level was detected.
FIG 4.
Transcriptional regulation of hlyA1 gene with 5′ enhancer region of different lengths. For each construct, three biological and technical replicates each were assayed by quantitative PCR (qPCR) using gene-specific primers. mRNA levels are expressed as the mean ± standard deviation (SD) of transcript levels relative (RLT) to the hcaT transcript and the lowest-expressing strain (11 nt). A two-tailed unpaired t test comparing the RTL values of the lowest-expressing strain (11 nt) and others was used to determine the statistical relevance. ***, P < 0.001. Error bars represent the standard deviations of the results from at least three independent biological replicates.
Overall, these data suggest that the observed increase in mRNA levels results in an enhanced secretion efficiency (Fig. 2 and 4). This is consistent with the many functions of ribosomal protein S1 in translation, such as stabilizing mRNA and enhancing transcript release from RNA polymerase, which enables the core enzyme for continuous reinitiation of transcription (41). However, our data cannot pinpoint the enhanced secretion efficiency to a single function of ribosomal protein S1. On the other hand, expression from the strong T7 promoter with the A/U-rich enhancer region derived from the pET-system (Novagen) did not improve or even reduced the amount of secreted protein (Fig. S2), implying that enhanced transcript levels and efficient translation cannot be the only reasons for the enhanced production of extracellular protein. Therefore, we also compared the stability and secondary structure of the enhancer region and the corresponding A/U-rich region of pET promoters at the RNAstructure server (https://rna.urmc.rochester.edu/RNAstructureWeb/) or the ViennaRNA Web Services (http://rna.tbi.univie.ac.at). Both analyses revealed that the A/U-rich region of the pET promoters was more stable and possessed high free energy. Additionally, highly efficient expression may lead to the degradation of heterologous proteins or the formation of inclusion bodies. We assume that usage of the weaker lac promoter and the lower copy number of the plasmids might result in an optimal number and assembly of transporters in the membrane, which in combination with the native hlyA enhancer result in an efficient finely tuned transcription, translation, and secretion of fusion proteins.
Fusion of passengers to the hlyA1 carrier protein.
The importance of the 5′ untranslated region and the interaction of the enhancer sequence with ribosomal protein S1 suggested the possibility that this mechanism is of more general nature. Thus, the role of the enhancer sequence for the expression and secretion of heterologous proteins was investigated.
Various proteins of interest were chosen, covering a molecular mass range from 19 to 88 kDa, which are known to provoke particular problems during production: (i) those which were not secreted by T1SS (maltose-binding protein [MBP] [61]), (ii) those which require intramolecular disulfide bridges for functionality (interferon alpha-2 [IFN-α2] [62]), and (iii) those with hydrolytic activities, which are frequently toxic for the expressing host cells (LipA and CalB).
Employing our expression system containing the hlyA enhancer region of 159 nt, all proteins analyzed were expressed and secreted with extracellular yields in the milligrams per liter range. After purification by size-exclusion chromatography (SEC) from cell-free supernatants, highly pure IFN-α2-HlyA1 and MBP-HlyA1 were obtained with a yield of 6 mg/liter each, while the yields of LipA-HlyA1 and CalB-HlyA1 were in the range of 3 mg/liter.
An inverse correlation between folding kinetics of passengers and secretion efficiencies has been demonstrated previously (25, 61). However, our engineered and improved system also catalyzes the transport of fast-folding proteins, such as wild-type MBP.
Furthermore, fusion proteins of HlyA1 with interferon alpha-2 (IFN-α2) and lipase A (LipA) with the 159-nt and 34-nt upstream regions of hlyA were constructed. Consistent with the results obtained for HlyA1, the 34-nt region resulted in increased levels of protein secretion (Fig. 5).
FIG 5.
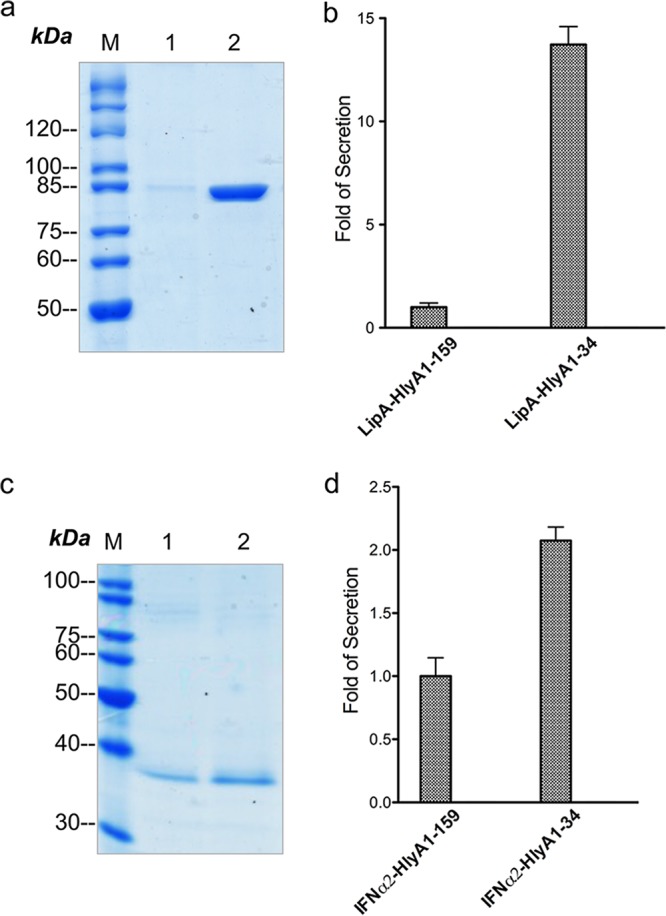
Secretion of fusion proteins with various lengths of the enhancer region of hlyA. SDS-PAGE of nonconcentrated supernatant samples from E. coli strains expressing hlyA1 with 5′ enhancer regions of 159 (lane 1) and 34 (lane 2) nucleotides with LipA (a) and IFN-α2 (c). Quantification (fold change; analyzed with ImageJ) of the amounts of LipA-HlyA1 (b) and IFN-α2-HlyA1 (d) secreted in the supernatant deduced from a representative gel based on the strains with 159 nt. Error bars represent the standard deviations of the results from at least three independent biological replicates.
In comparison to the strain carrying a 159-nt enhancer region, the amount of LipA-HlyA1 secreted with the 34-nt region was approximately 13 times higher (Fig. 5a and b). In the case of IFN-α2, the difference was less pronounced but still detectable, with the 34-nt region resulting in an increase in secretion (Fig. 5c and d).
Functional studies on secreted proteins and purification steps.
Besides yield and stability, the bioactivity of secreted proteins is crucial; thus, the above-mentioned four proteins were examined with respect to functionality.
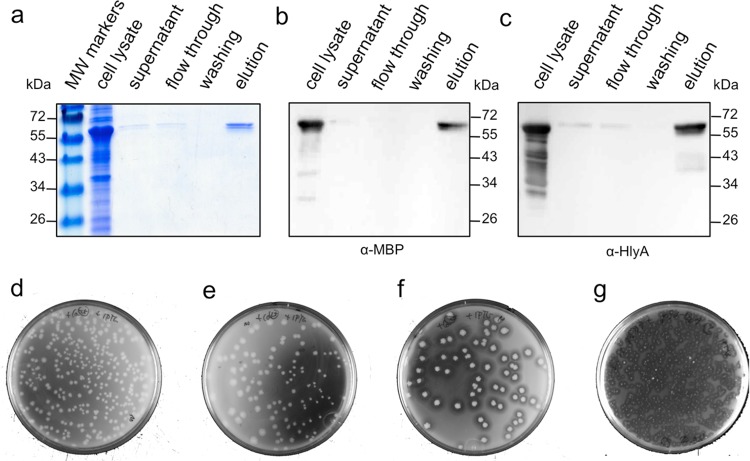
First, the binding of secreted MBP to amylose was examined. MBP-HlyA1 was bound to amylose and was eluted by maltose, clearly indicating the bioactivity of MBP-HlyA1 (Fig. 6a to c). Second, the lipolytic activity of secreted CalB was analyzed using a tributyrin-glycerol-based agar plate test (Fig. 6d to g). Cells lacking the transport complex (HlyB-HlyD) (Fig. 6d) or cells secreting HlyA1 alone (Fig. 6e) did not form halos, whereas secreted CalB-HlyA1 formed halos, demonstrating its lipolytic activity (Fig. 6f). The lipolytic activity of secreted LipA-HlyA1 was also demonstrated in a similar manner (Fig. 6g).
FIG 6.
Purification of secreted protein. (a) SDS-PAGE analysis of E. coli cells expressing and secreting MBP-HlyA1 fusion protein, its subsequent purification on amylose resin. MW, molecular weight. (b and c) Corresponding Western blot analysis of (a) using antibodies against MBP (b) and HlyA1 (c), respectively. Please note the additional bands in the case of the anti-HlyA antibody in the cell lysate. Since these bands are not present in the final elution, we believe that these correspond to C-terminal degradation products that would not be secreted due to the absence of the secretion sequence. (d to g) Analyses of lipolytic activity of secreted fusion proteins CalB and LipA. Tributyrin-glycerol-based agar plates are shown representing cells expressing calB alone (d), cells expressing only hlyA1, hlyB, and hlyD (e), cells expressing calB-hlyA1, hlyB, and hlyD (f), and cells expressing lipA-hlyA1, hlyB, and hlyD (g). Halo formation is only found around cell colonies expressing both the lipases and hlyB and hlyD (f and g).
Secreted IFN-α2-HlyA1 was analyzed for its antiviral activity on human fibroblasts. A luciferase-based reporter gene assay was used to quantify the specific activity (Fig. 7). Secreted HlyA1 alone was inert, whereas secreted IFN-α2-HlyA1 had a specific activity of 2.9 ± 0.9 × 106 U/mg. Factor Xa-catalyzed separation of HlyA1 increased the specific activity of IFN-α2 to 7.0 ± 0.9 × 108 U/mg, a value comparable to that of a commercial reference IFN-α2 (2.0 × 108 U/mg; PLB Assay Science).
FIG 7.
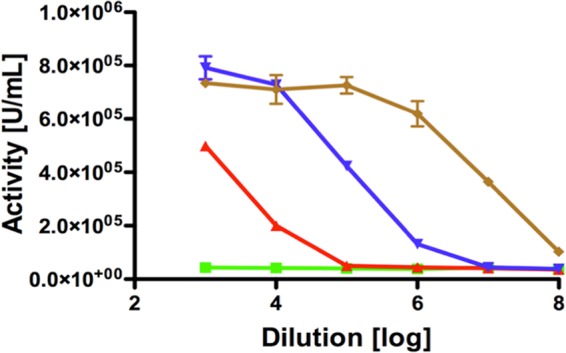
Luciferase reporter gene assay with secreted IFN-α2. A reference IFN-α2 (undiluted 1.0 × 105 U/ml or 2.0 × 108 U/mg; red curve) stimulated the expression of the luciferase gene in contrast to HlyA1 (23 μM, green curve). The specific activity of secreted IFN-α2-HlyA1 is shown in blue (23 μM), while the specific activity of IFN-α2 after HlyA1 separation (23 μM) is depicted in brown. Error bars represent the standard deviations of the results from at least three independent biological replicates.
DISCUSSION
In general, the secretion of proteins has distinct advantages over cellular production of the biomolecules. In the past, various hosts have been optimized for certain proteins resulting in impressive secretion efficiencies. The organisms employed include, for example, E. coli, Bacillus subtilis, Aspergillus species, or Pichia pastoris (3, 4, 63, 64). Despite the intensive efforts in optimizations, no general platform for protein secretion has been identified so far.
Our efforts increased the efficiency of the HlyA T1SS approximately 250-fold in terms of HlyA (>100 mg/liter compared to 400 μg/liter supernatant [49]) and about 25-fold in terms of C-terminal HlyA fusion constructs (180 mg of HlyA1/optical density [OD] unit compared to 7 mg of HlyA1/OD unit [61]; Fig. 1c). The increased efficiency relied mainly on the presence of the 5′ enhancer sequence upstream of the start codon of the fusion gene (Fig. 1b and 2) and its interaction with ribosomal protein S1 (Fig. 3). Furthermore, the mRNA transcript levels were also proportional to the secretion efficiency (Fig. 4). Studies in unrelated type III secretion system of Yersinia enterocolitica also identified a nontranslated RNA element upstream of the transport substrate that increased secretion levels, which stimulates the secretion of heterologous polypeptides (65). However, in contrast to the HlyA T1SS developed for E. coli, the synthesis, secretion, assembly, and function of the type III systems is far more complex, with over 50 gene products involved (66).
In addition to an optimized secretion of the natural substrate and its C-terminal fragment, heterologous proteins were also efficiently secreted (Fig. 5 to 7). Obviously, the secretion process is initiated more efficiently with the optimized system than with the HlyA T1SS system lacking the enhancer sequence (61).
The four secreted proteins tested here, LipA, CalB, IFN-α2, and MBP, are all biologically active (Fig. 6 and 7). Notably, the specific activity of IFN-α2 increased after separation from HlyA1 and outperformed a commercially available reference of IFN-α2 (PLB Assay Science) (Fig. 7). Considering that the biological activity of IFN-α2 critically relies on at least one disulfide bridge (67), the secreted IFN-α2 must have acquired this posttranslational modification (Fig. S2). Since it is unlikely within the reducing environment of E. coli (5), this capacity represents an obvious additional advantage of the HlyA T1SS over intracellular production strategies.
In summary, the enhancer sequence and its interaction with ribosomal protein S1 appear to be the keys to efficient protein secretion. The yields of the passenger fusions investigated in this study were in the range of milligrams of protein per liter of cell culture and offer a valuable avenue for the production of heterologous proteins. Previously, only slow-folding proteins were successfully secreted (61), and now, this bottleneck might be circumvented by use of the identified enhancer sequence.
MATERIALS AND METHODS
Construction of plasmids and dual-plasmid system for secretion.
Bicistronic hlyB and hlyD were amplified with the oligonucleotides BD_for and BD_rev (Table 2) and cloned into vector pET401. Subsequently, hlyB and hlyD were cleaved with NcoI/SacI and shuttled into pK184-BspHI to create plasmid p3S_HlyB-HlyD, having kanamycin resistance and being inducible with isopropyl β-d-1-thiogalactopyranoside (IPTG) under the lac promoter.
TABLE 2.
Summary of the plasmids and oligonucleotides used in this studya
| Plasmid | Oligonucleotide | Sequence | Template | Cloning method |
|---|---|---|---|---|
| p3S_HlyB-HlyD | BD_for | GCTATCCATGGCGAATTCTGATTCTTGTCATAAAATTGATTATGGG | hlyBD | See Materials and Methods |
| BD_rev | CGTTGTCTAGATTCTTAACGCTCATGTAAACTTTCTGTTACAGACTC | |||
| pSU_HlyA1+159 | pSU-HlyA1_for | TTAAGAGGTAATTAAATGGGAAATTCTCTTGCAAAAAATG | pSU2726 | In-fusion |
| pSU-HlyA1_rev | ATTTAAATTAATATATTATGCTGATGTGGTCAG | |||
| pSU_HlyA1+62 | Gap62_for | ATAACTGAAGTAAAAAGAAAGTCA | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+41 | Gap41_for | TCAGATTTTAATTTTTCATTAACTGG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+36 | Gap36_for | TTTTAATTTTTCATTAACTGGTTAAG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+34 | Gap34_for | TTAATTTTTCATTAACTGGTTAAGAG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+32 | Gap32_for | AATTTTTCATTAACTGGTTAAGAG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+23 | Gap23_for | TTAACTGGTTAAGAGGTAATTAAATG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+17 | Gap17_for | GGTTAAGAGGTAATTAAATGGG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+11 | Gap13_for | GAGGTAATTAAATGGGAAATTCTC | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1+8 | Gap8_for | GTAATTAAATGGGAAATTCTCTTG | pSU__HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_HlyA1_Δ14-23 | Gap13_for | AAGAGGTAATTAAATGGGAAATTC | pSU__HlyA1+159 | Ligation |
| GapΔ14-23_rev | TGAAAAATTAAAATCTGACTTTCT | |||
| pSU_HlyA1_Δ24-32 | Gap23_for | TTAACTGGTTAAGAGGTAATTAAATG | pSU__HlyA1+159 | Ligation |
| Δ24-32_rev | AAAATCTGACTTTCTTTTTACTTCA | |||
| pSU_HlyA1_Δ33-41 | Gap32_for | AATTTTTCATTAACTGGTTAAGAG | pSU__HlyA1+159 | Ligation |
| Δ33-41_rev | CTTTCTTTTTACTTCAGTTATTAACTC | |||
| pSU_HlyA1_Δ36-41 | Gap35_for | TTTAATTTTTCATTAACTGGTTAAG | pSU__HlyA1+159 | Ligation |
| Δ33-41_rev | CTTTCTTTTTACTTCAGTTATTAACTC | |||
| pSU_HlyA1_ 33u>g | Gap17_for | GGTTAAGAGGTAATTAAATGGG | pSU__HlyA1+41 | Ligation |
| Gap_u33g_rev | AGTTAATGAAAAATTCAAATCTGAGC | |||
| pSU_HlyA1_ 34u>g | Gap17_for | GGTTAAGAGGTAATTAAATGGG | pSU__HlyA1+41 | Ligation |
| Gap_u34g_rev | AGTTAATGAAAAATTACAATCTGAGC | |||
| pSU_HlyA1_33u>g-34u>g | Gap17_for | GGTTAAGAGGTAATTAAATGGG | pSU__HlyA1+41 | Ligation |
| Gap_u33-34g_rev | AGTTAATGAAAAATTCCAATCTGAGCT | |||
| pSU_HlyA1_AAATTAA | Gap23_for | TTAACTGGTTAAGAGGTAATTAAATG | pSU__HlyA1+41 | Ligation |
| AAATTAA_rev | TGAAATTAATTTATCTGACTTTCTTTTTAC | |||
| pSU_HlyA1+all AU | all AU_for | ATTAAATGGGAAATTCTCTTGC | pSU__HlyA1+41 | Ligation |
| all AU_rev | TACCTCTTAATTAATTAATTAAAAATTAAAATCTG | |||
| pSU_LipA-HlyA1+159 | LipA_for | AGAGGTAATTAAATGGGCATCTTTAGCTATAAGG | lipA | In-fusion |
| LipA_rev | TGCAAGAGAATTTCCGGCCAACACCACCTG | |||
| pSU_LipA-HlyA1+34 | Gap34_for | TTAATTTTTCATTAACTGGTTAAGAG | pSU_LipA-HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG | |||
| pSU_MBP-HlyA1+159 | MBP_for | AGAGGTAATTAAATGAAAATCGAAGAAGGTAAAC | mbp | In-fusion |
| MBP_rev | TGCAAGAGAATTTCCACGGCCATCAATCTTGGTGATACGAGTCTG | |||
| pSU_CalB-HlyA1+159 | CalB_for | AGAGGTAATTAAATGCTACCTTCCGGTTCG | calB | In-fusion |
| CalB_rev | TGCAAGAGAATTTCCACGGCCATCAATGGGGGTGACGATGCCG | |||
| pSU_IFN-α2-HlyA1+159 | IFN-α2_for | AGAGGTAATTAAATGTGTGATCTGCCGCAGACTC | IFN-α2 | In-fusion |
| IFN-α2_rev | GTGACGGCCATCAATTTCCTTACTTCTTAAACTTTCTTGCAAGTTT | |||
| pSU_IFN-α2-HlyA1+34 | Gap34_for | TTAATTTTTCATTAACTGGTTAAGAG | pSU_IFN-α2-HlyA1+159 | Ligation |
| Gap_rev | GCTCGAATTCGTAATCATG |
The oligonucleotides and templates used for the generation of the plasmids are listed. Either an in-fusion or ligation method was used (refer to Materials and Methods). Plasmid pSU2726 served as parental vector except for the p3S_HlyB-HlyD plasmid.
Plasmid pSU2726 (containing ampicillin resistance with a lac promoter) (49) encoding full-length HlyA was used as the template for all plasmids encoding truncations of HlyA (including HlyA1, amino acid residues 807 to 1024) and the fusion proteins. Either an in-fusion or ligation method was used for their production, according to the manufacturer's protocols (New England BioLabs, USA, and TaKaRa, USA). All plasmids and oligonucleotides used in this study are summarized in Table 2.
For secretion via the Hly system, a dual plasmid-based system was constructed as previously described (61). In brief, a two-plasmid approach was used, where one plasmid, p3S_HlyB-HlyD, encodes HlyB and HlyD, comprising the inner membrane complex, while the other plasmid, pSU_HlyA1-X (X refers to the additional nucleotides present upstream of the start codon), encodes the C-terminal fragment of HlyA containing the secretion signal. Since both plasmids are under the control of lac promoter, the expression of all the genes was simultaneously inducible by IPTG. In case of the secretion of heterologous proteins, the second plasmid contained the secretion signal fused to its C terminus.
Expression and secretion experiments.
E. coli BL21(DE3) competent cells were cotransformed with p3S_HlyB-HlyD and pSU_HlyA1-X (X refers to the additional nucleotides present upstream of the start codon) (Tables 2 and S2) bearing the transport substrate and grown on 2YT (10 g yeast extract, 16 g tryptone/peptone, 5 g sodium in 1 liter) agar plates supplemented with 100 μg · ml−1 ampicillin and 30 μg · ml−1 kanamycin. Overnight cultures using single colonies (bearing the double plasmids) were used to inoculate 25 ml of 2YT medium supplemented with both antibiotics at an optical density at 600 nm (OD600) of 0.1. The cultures were grown at 37°C and 180 rpm. The expression of HlyA1, HlyB, and HlyD was induced at an OD600 of 0.4 to 0.6 using 1 mM IPTG; additionally, CaCl2 was also added to the medium at a final concentration of 5 mM. Cells were expressed for 3 to 6 h. The secreted protein samples were quantified after SDS-PAGE analysis and colloidal Coomassie staining (68) using the ImageJ (Image Processing and Analysis in Java) software (24, 69) with bovine serum albumin (BSA) standard.
RNA pulldown assay.
3′-Biotin TEG-labeled RNA containing either the enhancer sequence (5′AAAGUCAGAUUUUAAUUUUUCAUUAACUGGUUAAGA) or its mutated version (33u>g-34u>g, 5′ AAAGUCAGAUUGGAAUUUUUCAUUAACUGGUUAAGA) or the corresponding sequence from the low-expression strain +17 (5′-CAGCUAUGACAUGAUUACGAAUUCGAGCGGUUAAGA) was purchased from Eurofins Genomics. The RNA-protein pulldown assay was performed using the Thermo Scientific Pierce magnetic RNA-protein pulldown kit, according to the instructions of the manufacturer. Biotin elution buffer was used for the harvest of bound proteins.
RNA purification.
The same protocol of secretion experiments (see above) was performed using the selected HlyA1 plasmids containing enhancer regions of varied lengths. However, after induction with IPTG and the addition of CaCl2, the cells were allowed to grow only until an OD600 of 1.0 and were collected by centrifugation. Total RNA was isolated and purified using an RNeasy minikit (Qiagen, Germany).
Quantitative real-time PCR.
Three micrograms of purified RNA was treated with DNase I (Turbo DNA-free kit; Invitrogen) and subsequently with RNA Clean & Concentrator (Zymo Research), according to the manufacturer's protocol. cDNA was synthesized using 1 μg of RNA (Maxima first-strand cDNA synthesis kit for quantitative real-time PCR [RT-qPCR] with dsDNase; Thermo Scientific). Transcript levels were measured during RT-qPCR using my-Budget 5× EvaGreen QPCR-mix II (Bio-Budget) with an Mx3000P (Stratagene) cycler using primers hlyA1 Fwd (CCGGATAATCACACCTGATTCCC), hlyA1 Rev (CCCTGACTTCCATAGGCTAATGC), hcaT Fwd (GCTGCTCGGCTTTCTCATCC), and hcaT Rev (CCAACCACGCTGACCAACC). hlyA1 transcript levels were normalized to hcaT transcript levels to determine their relative transcript levels (RTL). Mean standard deviations of ratios were calculated considering the standard deviations of the threshold cycle (CT) values of both reference and target genes by using the Gauss equation for error propagation (70). RTL values were further compared to the lowest-expressing strain (11 nt).
Purification of secreted fusion proteins.
Cells bearing the double plasmids (p3S_HlyB-HlyD and pSU_HlyA1-X with the passenger protein) were grown for 6 h after induction with 1 mM IPTG. A volume of 45 ml of cell-free supernatant was concentrated to 2 ml by ultrafiltration (10,000 molecular weight cutoff [MWCO]) and applied to an SEC column equilibrated in 10 mM Tris-HCl (pH 7.3) and 120 mM NaCl.
Functionality of secreted MBP.
Cell-free supernatant containing MBP-HlyA1 was loaded onto an amylose resin (NEB). The column was washed with 20 mM HEPES (pH 7.4), 150 mM NaCl, and MBP-HlyA1 and was eluted with 10 mM maltose.
Functionality of secreted lipases.
Cells were either transformed with pSU_LipA-HlyA1 or pSU_CalB-HlyA1 or together with p3S_HlyB-HlyD and grown on LB-agar plates supplemented with 1% glycerol tributyrate, 1 mM IPTG, and the appropriate antibiotics for either 18 h (pSU_LipA-HlyA1-159) or 36 h (pSU_CalB-HlyA1-159) at 37°C.
Separation of the passenger protein from HlyA1.
Fifty micrograms of secreted fusion proteins was digested with 1 μg of factor Xa (NEB). IFN-α2 was separated from HlyA1 by anion-exchange chromatography and concentrated by ultrafiltration to a concentration of 0.5 mg · ml−1.
Luciferase reporter gene assay of secreted IFN-α2.
The interferon reporter cell line has been previously described (71). Briefly, human retinal pigment epithelial cells have been cotransfected with a plasmid containing interferon-stimulated response element (ISRE) promoter/enhancer elements driving a luciferase reporter gene (pISRE; Stratagene) together with a plasmid conferring Geneticin resistance, as described previously (72). Confluent cells were incubated with the indicated dilutions of a commercial reference (rec-IFN-α2 and PBL), HlyA1, secreted and purified IFN-α2-HlyA1 or IFN-α2 for 6 h at 37°C and 5% CO2. After incubation, cells were lysed, and the luciferase activity was quantified using the luciferase reporter gene assay (Roche, Germany).
Supplementary Material
ACKNOWLEDGMENTS
We thank the Heinrich Heine University Düsseldorf and the Ministry of Innovation, Science and Research of the German Federal State North Rhine-Westphalia (NRW) (scholarship from CLIB Graduate Cluster Industrial Biotechnology to C.S.) and the Bioeconomy Science Center for funding (project BioBreak to L.S.). M.T. received funding from the Deutsche Forschungsgemeinschaft (through GRK1949 TP.13 and TRR60 project A7N).
We also thank Prashant Desai, Christoph Bürth, Lasse van Wijlick, and Joachim Ernst (Institute of Molecular Mycology, Heinrich Heine University) for real-time PCR machine usage and support, and Diana Kleinschrodt and Laura Kuschmierz for valuable support.
Footnotes
Supplemental material for this article may be found at https://doi.org/10.1128/AEM.01163-17.
REFERENCES
- 1.Rosano GL, Ceccarelli EA. 2014. Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol 5:172. doi: 10.3389/fmicb.2014.00172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Huang CJ, Lin H, Yang X. 2012. Industrial production of recombinant therapeutics in Escherichia coli and its recent advancements. J Ind Microbiol Biotechnol 39:383–399. doi: 10.1007/s10295-011-1082-9. [DOI] [PubMed] [Google Scholar]
- 3.Harwood CR, Cranenburgh R. 2007. Bacillus protein secretion: an unfolding story. Trends Microbiol 16:73–79. doi: 10.1016/j.tim.2007.12.001. [DOI] [PubMed] [Google Scholar]
- 4.Ni Y, Chen R. 2009. Extracellular recombinant protein production from Escherichia coli. Biotechnol Lett 31:1661–1670. doi: 10.1007/s10529-009-0077-3. [DOI] [PubMed] [Google Scholar]
- 5.Choi J, Lee S. 2004. Secretory and extracellular production of recombinant proteins using Escherichia coli. Appl Microbiol Biotechnol 64:625–635. doi: 10.1007/s00253-004-1559-9. [DOI] [PubMed] [Google Scholar]
- 6.Gouka RJ, Punt PJ, van den Hondel CA. 1997. Efficient production of secreted proteins by Aspergillus: progress, limitations and prospects. Appl Microbiol Biotechnol 47:1–11. doi: 10.1007/s002530050880. [DOI] [PubMed] [Google Scholar]
- 7.Kakeshita H, Kageyama Y, Ozaki K, Nakamura K, Ara K. 2012. Improvement of heterologous protein secretion by Bacillus subtilis. In Petre PM. (ed), Advances in applied biotechnology InTech, Rijeka, Croatia. http://www.intechopen.com/books/advances-in-applied-biotechnology/improvement-of-heterologous-protein-secretion-by-bacillus-subtilis.
- 8.Rojas Contreras JA, Pedraza-Reyes M, Ordonez LG, Estrada NU, Barba de la Rosa AP, De Leon-Rodriguez A. 2010. Replicative and integrative plasmids for production of human interferon gamma in Bacillus subtilis. Plasmid 64:170–176. doi: 10.1016/j.plasmid.2010.07.003. [DOI] [PubMed] [Google Scholar]
- 9.Zhang XZ, Cui ZL, Hong Q, Li SP. 2005. High-level expression and secretion of methyl parathion hydrolase in Bacillus subtilis WB800. Appl Environ Microbiol 71:4101–4103. doi: 10.1128/AEM.71.7.4101-4103.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Mergulhao F, Summers DK, Monteiro GA. 2005. Recombinant protein secretion in Escherichia coli. Biotechnol Adv 23:177–202. doi: 10.1016/j.biotechadv.2004.11.003. [DOI] [PubMed] [Google Scholar]
- 11.Holland IB, Schmitt L, Young J. 2005. Type 1 protein secretion in bacteria, the ABC-transporter dependent pathway (review). Mol Membr Biol 22:29–39. doi: 10.1080/09687860500042013. [DOI] [PubMed] [Google Scholar]
- 12.Holland IB, Kenny B, Steipe B, Plückthun A. 1990. Secretion of heterologous proteins in Escherichia coli. Methods Enzymol 182:132–143. doi: 10.1016/0076-6879(90)82013-R. [DOI] [PubMed] [Google Scholar]
- 13.Spreng S, Dietrich G, Goebel W, Gentschev I. 1999. The Escherichia coli haemolysin secretion apparatus: a potential universal antigen delivery system in Gram-negative bacterial vaccine carriers. Mol Microbiol 31:1596–1598. doi: 10.1046/j.1365-2958.1999.01290.x. [DOI] [PubMed] [Google Scholar]
- 14.Mackman N, Baker K, Gray L, Haigh R, Nicaud JM, Holland IB. 1987. Release of a chimeric protein into the medium from Escherichia coli using the C-terminal secretion signal of haemolysin. EMBO J 6:2835–2841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fernández LA, Sola I, Enjuanes L, de Lorenzo V. 2000. Specific secretion of active single-chain Fv antibodies into the supernatants of Escherichia coli cultures by use of the hemolysin system. Appl Environ Microbiol 66:5024–5029. doi: 10.1128/AEM.66.11.5024-5029.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Blight MA, Holland IB. 1994. Heterologous protein secretion and the versatile Escherichia coli haemolysin translocator. Trends Biotechnol 12:450–455. doi: 10.1016/0167-7799(94)90020-5. [DOI] [PubMed] [Google Scholar]
- 17.Nicaud JM, Mackman N, Gray L, Holland IB. 1985. Characterisation of HlyC and mechanism of activation and secretion of haemolysin from E. coli 2001. FEBS Lett 187:339–344. doi: 10.1016/0014-5793(85)81272-2. [DOI] [PubMed] [Google Scholar]
- 18.Nicaud JM, Mackman N, Gray L, Holland IB. 1986. The C-terminal, 23 kDa peptide of E. coli haemolysin 2001 contains all the information necessary for its secretion by the haemolysin (Hly) export machinery. FEBS Lett 204:331–335. doi: 10.1016/0014-5793(86)80838-9. [DOI] [PubMed] [Google Scholar]
- 19.Stanley P, Packman LC, Koronakis V, Hughes C. 1994. Fatty acylation of two internal lysine residues required for the toxic activity of Escherichia coli hemolysin. Science 266:1992. doi: 10.1126/science.7801126. [DOI] [PubMed] [Google Scholar]
- 20.Hardie KR, Issartel JP, Koronakis E, Hughes C, Koronakis V. 1991. In vitro activation of Escherichia coli prohaemolysin to the mature membrane-targeted toxin requires HlyC and a low molecular-weight cytosolic polypeptide. Mol Microbiol 5:1669–1679. doi: 10.1111/j.1365-2958.1991.tb01914.x. [DOI] [PubMed] [Google Scholar]
- 21.Wandersman C, Delepelaire P. 1990. TolC, an Escherichia coli outer membrane protein required for hemolysin secretion. Proc Natl Acad Sci U S A 87:4776–4780. doi: 10.1073/pnas.87.12.4776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Thanabalu T, Koronakis E, Hughes C, Koronakis V. 1998. Substrate-induced assembly of a contiguous channel for protein export from E.coli: reversible bridging of an inner-membrane translocase to an outer membrane exit pore. EMBO J 17:6487–6496. doi: 10.1093/emboj/17.22.6487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Sánchez-Magraner L, Viguera AR, García-Pacios M, Garcillán MP, Arrondo J-LR, de la Cruz F, Goñi FM, Ostolaza H. 2007. The calcium-binding C-terminal domain of Escherichia coli α-hemolysin is a major determinant in the surface-active properties of the protein. J Biol Chem 282:11827–11835. doi: 10.1074/jbc.M700547200. [DOI] [PubMed] [Google Scholar]
- 24.Lenders MH, Beer T, Smits SH, Schmitt L. 2016. In vivo quantification of the secretion rates of the hemolysin A type I secretion system. Sci Rep 6:33275. doi: 10.1038/srep33275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Schwarz CK, Landsberg CD, Lenders MH, Smits SH, Schmitt L. 2012. Using an E. coli type 1 secretion system to secrete the mammalian, intracellular protein IFABP in its active form. J Biotechnol 159:155–161. doi: 10.1016/j.jbiotec.2012.02.005. [DOI] [PubMed] [Google Scholar]
- 26.Schwarz CK, Lenders MH, Smits SH, Schmitt L. 2012. Secretion of slow-folding proteins by a type 1 secretion system. Bioengineered 3:289–292. doi: 10.4161/bioe.20712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lindberg AA, Hellerqvist CG. 1980. Rough mutants of Salmonella Typhimurium: immunochemical and structural analysis of lipopolysaccharides from rfaH mutants. J Gen Microbiol 116:25–32. [DOI] [PubMed] [Google Scholar]
- 28.Leeds JA, Welch RA. 1997. Enhancing transcription through the Escherichia coli hemolysin operon, hlyCABD: RfaH and upstream JUMPStart DNA sequences function together via a postinitiation mechanism. J Bacteriol 179:3519–3527. doi: 10.1128/jb.179.11.3519-3527.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bailey MJ, Hughes C, Koronakis V. 1997. RfaH and the ops element, components of a novel system controlling bacterial transcription elongation. Mol Microbiol 26:845–851. doi: 10.1046/j.1365-2958.1997.6432014.x. [DOI] [PubMed] [Google Scholar]
- 30.Artsimovitch I, Landick R. 2002. The transcriptional regulator RfaH stimulates RNA chain synthesis after recruitment to elongation complexes by the exposed nontemplate DNA strand. Cell 109:193–203. doi: 10.1016/S0092-8674(02)00724-9. [DOI] [PubMed] [Google Scholar]
- 31.Boni IV, Isaeva DM, Musychenko ML, Tzareva NV. 1991. Ribosome-messenger recognition: mRNA target sites for ribosomal protein S1. Nucleic Acids Res 19:155–162. doi: 10.1093/nar/19.1.155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.McCarthy JE, Schairer HU, Sebald W. 1985. Translational initiation frequency of atp genes from Escherichia coli: identification of an intercistronic sequence that enhances translation. EMBO J 4:519–526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nafissi M, Chau J, Xu J, Johnson RC. 2012. Robust translation of the nucleoid protein Fis requires a remote upstream AU element and is enhanced by RNA secondary structure. J Bacteriol 194:2458–2469. doi: 10.1128/JB.00053-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sørensen MA, Fricke J, Pedersen S. 1998. Ribosomal protein S1 is required for translation of most, if not all, natural mRNAs in Escherichia coli in vivo. J Mol Biol 280:561–569. doi: 10.1006/jmbi.1998.1909. [DOI] [PubMed] [Google Scholar]
- 35.Kitakawa M, Isono K. 1982. An amber mutation in the gene rpsA for ribosomal protein S1 in Escherichia coli. Mol Gen Genet 185:445–447. doi: 10.1007/BF00334137. [DOI] [PubMed] [Google Scholar]
- 36.Briani F, Curti S, Rossi F, Carzaniga T, Mauri P, Deho G. 2008. Polynucleotide phosphorylase hinders mRNA degradation upon ribosomal protein S1 overexpression in Escherichia coli. RNA 14:2417–2429. doi: 10.1261/rna.1123908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Draper DE, von Hippel PH. 1978. Nucleic acid binding properties of Escherichia coli ribosomal protein S1. I. Structure and interactions of binding site I. J Mol Biol 122:321–338. [DOI] [PubMed] [Google Scholar]
- 38.Bycroft M, Hubbard TJ, Proctor M, Freund SM, Murzin AG. 1997. The solution structure of the S1 RNA binding domain: a member of an ancient nucleic acid-binding fold. Cell 88:235–242. doi: 10.1016/S0092-8674(00)81844-9. [DOI] [PubMed] [Google Scholar]
- 39.Subramanian AR, van Duin J. 1977. Exchange of individual ribosomal proteins between ribosomes as studied by heavy isotope-transfer experiments. Mol Gen Genet 158:1–9. doi: 10.1007/BF00455113. [DOI] [PubMed] [Google Scholar]
- 40.Sykes MT, Williamson JR. 2009. A complex assembly landscape for the 30S ribosomal subunit. Annu Rev Biophys 38:197–215. doi: 10.1146/annurev.biophys.050708.133615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sukhodolets MV, Garges S, Adhya S. 2006. Ribosomal protein S1 promotes transcriptional cycling. RNA 12:1505–1513. doi: 10.1261/rna.2321606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kaberdin VR, Blasi U. 2006. Translation initiation and the fate of bacterial mRNAs. FEMS Microbiol Rev 30:967–979. doi: 10.1111/j.1574-6976.2006.00043.x. [DOI] [PubMed] [Google Scholar]
- 43.Komarova AV, Tchufistova LS, Dreyfus M, Boni IV. 2005. AU-rich sequences within 5′ untranslated leaders enhance translation and stabilize mRNA in Escherichia coli. J Bacteriol 187:1344–1349. doi: 10.1128/JB.187.4.1344-1349.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Saguy M, Gillet R, Skorski P, Hermann-Le Denmat S, Felden B. 2007. Ribosomal protein S1 influences trans-translation in vitro and in vivo. Nucleic Acids Res 35:2368–2376. doi: 10.1093/nar/gkm100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Qu X, Lancaster L, Noller HF, Bustamante C, Tinoco I Jr. 2012. Ribosomal protein S1 unwinds double-stranded RNA in multiple steps. Proc Natl Acad Sci U S A 109:14458–14463. doi: 10.1073/pnas.1208950109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Olins PO, Devine CS, Rangwala SH, Kavka KS. 1988. The T7 phage gene 10 leader RNA, a ribosome-binding site that dramatically enhances the expression of foreign genes in Escherichia coli. Gene 73:227–235. doi: 10.1016/0378-1119(88)90329-0. [DOI] [PubMed] [Google Scholar]
- 47.Takahashi S, Furusawa H, Ueda T, Okahata Y. 2013. Translation enhancer improves the ribosome liberation from translation initiation. J Am Chem Soc 135:13096–13106. doi: 10.1021/ja405967h. [DOI] [PubMed] [Google Scholar]
- 48.Jumpertz T, Chervaux C, Racher K, Zouhair M, Blight MA, Holland IB, Schmitt L. 2010. Mutations affecting the extreme C terminus of Escherichia coli haemolysin A reduce haemolytic activity by altering the folding of the toxin. Microbiology 156:2495–2505. doi: 10.1099/mic.0.038562-0. [DOI] [PubMed] [Google Scholar]
- 49.Soloaga A, Ostolaza H, Goni FM, de la Cruz F. 1996. Purification of Escherichia coli pro-haemolysin, and a comparison with the properties of mature alpha-haemolysin. Eur J Biochem 238:418–422. doi: 10.1111/j.1432-1033.1996.0418z.x. [DOI] [PubMed] [Google Scholar]
- 50.Thomas S, Holland IB, Schmitt L. 2014. The type 1 secretion pathway—the hemolysin system and beyond. Biochim Biophys Acta 1843:1629–1641. doi: 10.1016/j.bbamcr.2013.09.017. [DOI] [PubMed] [Google Scholar]
- 51.Paytubi S, Dietrich M, Queiroz MH, Juarez A. 2013. Role of plasmid- and chromosomally encoded Hha proteins in modulation of gene expression in E. coli O157:H7. Plasmid 70:52–60. doi: 10.1016/j.plasmid.2013.01.006. [DOI] [PubMed] [Google Scholar]
- 52.Jubete Y, Zabala JC, Juarez A, de la Cruz F. 1995. hlyM, a transcriptional silencer downstream of the promoter in the hly operon of Escherichia coli. J Bacteriol 177:242–246. doi: 10.1128/jb.177.1.242-246.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Li H, Granat A, Stewart V, Gillespie JR. 2008. RpoS, H-NS, and DsrA influence EHEC hemolysin operon (ehxCABD) transcription in Escherichia coli O157:H7 strain EDL933. FEMS Microbiol Lett 285:257–262. doi: 10.1111/j.1574-6968.2008.01240.x. [DOI] [PubMed] [Google Scholar]
- 54.Dong T, Schellhorn HE. 2010. Role of RpoS in virulence of pathogens. Infect Immun 78:887–897. doi: 10.1128/IAI.00882-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Madrid C, Nieto JM, Paytubi S, Falconi M, Gualerzi CO, Juarez A. 2002. Temperature- and H-NS-dependent regulation of a plasmid-encoded virulence operon expressing Escherichia coli hemolysin. J Bacteriol 184:5058–5066. doi: 10.1128/JB.184.18.5058-5066.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Fröhlich KS, Vogel J. 2009. Activation of gene expression by small RNA. Curr Opin Microbiol 12:674–682. doi: 10.1016/j.mib.2009.09.009. [DOI] [PubMed] [Google Scholar]
- 57.Chao Y, Vogel J. 2010. The role of Hfq in bacterial pathogens. Curr Opin Microbiol 13:24–33. doi: 10.1016/j.mib.2010.01.001. [DOI] [PubMed] [Google Scholar]
- 58.Wang MC, Chien HF, Tsai YL, Liu MC, Liaw SJ. 2014. The RNA chaperone Hfq is involved in stress tolerance and virulence in uropathogenic Proteus mirabilis. PLoS One 9:e85626. doi: 10.1371/journal.pone.0085626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Nakano M, Takahashi A, Su Z, Harada N, Mawatari K, Nakaya Y. 2008. Hfq regulates the expression of the thermostable direct hemolysin gene in Vibrio parahaemolyticus. BMC Microbiol 8:155. doi: 10.1186/1471-2180-8-155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Moll I, Afonyushkin T, Vytvytska O, Kaberdin VR, Blasi U. 2003. Coincident Hfq binding and RNase E cleavage sites on mRNA and small regulatory RNAs. RNA 9:1308–1314. doi: 10.1261/rna.5850703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Bakkes PJ, Jenewein S, Smits SH, Holland IB, Schmitt L. 2010. The rate of folding dictates substrate secretion by the Escherichia coli hemolysin type 1 secretion system. J Biol Chem 285:40573–40580. doi: 10.1074/jbc.M110.173658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bekisz J, Schmeisser H, Hernandez J, Goldman ND, Zoon KC. 2004. Human interferons alpha, beta and omega. Growth Factors 22:243–251. doi: 10.1080/08977190400000833. [DOI] [PubMed] [Google Scholar]
- 63.Fleissner A, Dersch P. 2010. Expression and export: recombinant protein production systems for Aspergillus. Appl Microbiol Biotechnol 87:1255–1270. doi: 10.1007/s00253-010-2672-6. [DOI] [PubMed] [Google Scholar]
- 64.Laroche Y, Storme V, De Meutter J, Messens J, Lauwereys M. 1994. High-level secretion and very efficient isotopic labeling of tick anticoagulant peptide (TAP) expressed in the methylotrophic yeast, Pichia pastoris. Nat Biotechnol 12:1119–1124. doi: 10.1038/nbt1194-1119. [DOI] [PubMed] [Google Scholar]
- 65.Anderson DM, Schneewind O. 1999. Yersinia enterocolitica type III secretion: an mRNA signal that couples translation and secretion of YopQ. Mol Microbiol 31:1139–1148. doi: 10.1046/j.1365-2958.1999.01254.x. [DOI] [PubMed] [Google Scholar]
- 66.Majander K, Anton L, Antikainen J, Lang H, Brummer M, Korhonen TK, Westerlund-Wikstrom B. 2005. Extracellular secretion of polypeptides using a modified Escherichia coli flagellar secretion apparatus. Nat Biotechnol 23:475–481. doi: 10.1038/nbt1077. [DOI] [PubMed] [Google Scholar]
- 67.Morehead H, Johnston PD, Wetzel R. 1984. Roles of the 29-138 disulfide bond of subtype A of human alpha interferon in its antiviral activity and conformational stability. Biochemistry 23:2500–2507. doi: 10.1021/bi00306a028. [DOI] [PubMed] [Google Scholar]
- 68.Dyballa N, Metzger S. 2009. Fast and sensitive colloidal Coomassie G-250 staining for proteins in polyacrylamide gels. J Vis Exp doi: 10.3791/1431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Schneider CA, Rasband WS, Eliceiri KW. 2012. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Muller PY, Janovjak H, Miserez AR, Dobbie Z. 2002. Short technical report processing of gene expression data generated by quantitative real-time RT-PCR. Biotechniques 32:1372–1379. [PubMed] [Google Scholar]
- 71.Lavender KJ, Gibbert K, Peterson KE, Van Dis E, Francois S, Woods T, Messer RJ, Gawanbacht A, Muller JA, Munch J, Phillips K, Race B, Harper MS, Guo K, Lee EJ, Trilling M, Hengel H, Piehler J, Verheyen J, Wilson CC, Santiago ML, Hasenkrug KJ, Dittmer U. 2016. Interferon alpha subtype-specific suppression of HIV-1 infection in vivo. J Virol 90:6001–6013. doi: 10.1128/JVI.00451-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Zimmermann A, Trilling M, Wagner M, Wilborn M, Bubic I, Jonjic S, Koszinowski U, Hengel H. 2005. A cytomegaloviral protein reveals a dual role for STAT2 in IFN-γ signaling and antiviral responses. J Exp Med 201:1543–1553. doi: 10.1084/jem.20041401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Felmlee T, Pellett S, Welch RA. 1985. Nucleotide sequence of an Escherichia coli chromosomal hemolysin. J Bacteriol 163:94–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.