Abstract
This study aimed to determine if there is an association between dysbiosis and nonalcoholic fatty liver disease (NAFLD) independent of obesity and insulin resistance (IR). This is a prospective cross-sectional study assessing the intestinal microbiome (IM) of 39 adults with biopsy-proven NAFLD (15 simple steatosis [SS]; 24 nonalcoholic steatohepatitis [NASH]) and 28 healthy controls (HC). IM composition (llumina MiSeq Platform) in NAFLD patients compared to HC were identified by two statistical methods (Metastats, Wilcoxon). Selected taxa was validated using quantitative PCR (qPCR). Metabolites in feces and serum were also analyzed. In NAFLD, 8 operational taxonomic units, 6 genera, 6 families and 2 phyla (Bacteroidetes, Firmicutes) were less abundant and; 1 genus (Lactobacillus) and 1 family (Lactobacillaceae) were more abundant compared to HC. Lower abundance in both NASH and SS patients compared to HC were confirmed by qPCR for Ruminococcus, Faecalibacterium prausnitzii and Coprococcus. No difference was found between NASH and SS. This lower abundance in NAFLD (NASH+SS) was independent of BMI and IR. NAFLD patients had higher concentrations of fecal propionate and isobutyric acid and serum 2-hydroxybutyrate and L-lactic acid. These findings suggest a potential role for a specific IM community and functional profile in the pathogenesis of NAFLD.
Introduction
Nonalcoholic fatty liver disease (NAFLD) is the most prevalent chronic liver disease in the Western world1 and it is closely associated with obesity, insulin resistance (IR) and diabetes, dyslipidemia, and coronary artery disease2, all of which are manifestations of the metabolic syndrome.
The pathogenesis of NAFLD is complex. Research suggests that factors such as genetics3, lipid peroxidation4, IR associated with obesity5, and diet6 may contribute. In addition, emerging research suggests a role for the intestinal microbiome (IM) where dysbiosis and bacterial metabolism and products7 may influence NAFLD pathogenesis through effects on nutrient digestion and absorption, appetite regulation, host gene expression, and immune function8.
Recent human studies have shown associations between IM composition and NAFLD9–16, but whether these associations were due to the presence of NAFLD itself or other factors associated with NAFLD, such as increased body mass index (BMI) and IR, was not clear. Most studies did not even consider these factors10,12–14. In addition, some of these studies did not perform liver biopsies in either the patient group10,12,16 or controls10–14,16, or did not assess other factors that may affect IM such as nutritional intake, environment, or physical activity12–16. In a previous study9 using qPCR, we found an inverse association between the presence of NASH and percentage of Bacteroidetes in the stool, and this was independent of dietary intake and BMI. However, the method only allowed for the assessment of a very limited number of taxa. Also, bacterial products were not analyzed and these products may play a role in the relationship between NAFLD and IM17.
The goal of the present study was to determine if there is an association between dysbiosis and NAFLD independently of obesity and IR, by using advanced sequencing technology to characterize IM, in patients with biopsy-proven NAFLD and healthy living liver donors as healthy controls (HC). To ensure robustness of our conclusions to statistical assumptions, the differences in taxa abundances were assessed by two different methods. We additionally quantified select microbial taxa found to be differentially abundant in next generation sequencing data using qPCR, and examined their relationship with NAFLD, BMI and IR. In addition, we measured serum and fecal metabolites using nuclear magnetic resonance spectrometry.
Methods
Subjects
In this prospective cross-sectional study, adult participants were recruited at the University Health Network (UHN), Toronto, Ontario, Canada. For NAFLD, subjects with persistently elevated liver enzymes were assessed by hepatologists at UHN and NAFLD was confirmed using standard medical practice to rule out other liver conditions. Patients who accepted to have a liver biopsy were then referred for the study. Inclusion criteria were: age >18 years and biopsy-confirmed NAFLD. Exclusion criteria for patients were: liver disease other than NAFLD, anticipated need for liver transplantation within a year or complications of end-stage liver disease such as variceal bleeding or ascites; concurrent medical illnesses; and contraindications for liver biopsy.
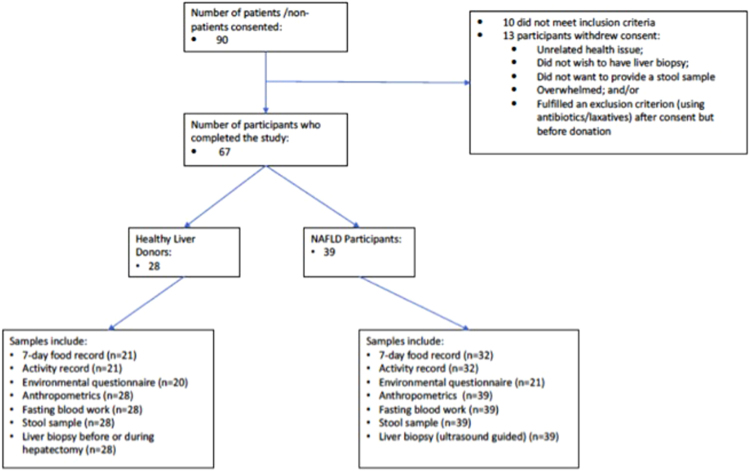
Healthy liver donors from the living donor transplant program served as healthy controls (HC) and as per program protocol a healthy liver was confirmed either by biopsy, computed tomography, and/or medical resonance imaging before partial hepatectomy. For HC the exclusion criteria were: any medical reason excluding them from the live liver donation as per program protocol. Other exclusion criteria for all subjects were: use of medications known to cause or exacerbate steatohepatitis or antibiotics, pre- or probiotics in the preceding 6 months; consumption of more than 20 g of alcohol/day; use of vitamin E or fish oil supplements; chronic gastrointestinal diseases, previous gastrointestinal surgery modifying the anatomy; pregnancy or lactating state. Figure 1 provides an overview of subject recruitment and study completion.
Figure 1.
Subject recruitment and sample collection flow chart.
This study was registered with ClinicalTrials.gov (NCT02148471). Additional trial outcomes are reported in previous publications18,19. A manuscript reporting on immune response is pending submission.
Ethics
The study protocol was approved by the UHN and University of Toronto Research Ethics Boards and conformed to the ethical guidelines of the 1975 Declaration of Helsinki. All participants gave their informed written consent. All methods were performed in accordance with the relevant guidelines and regulations of UHN and the University of Toronto.
Measurements
Patients provided one stool sample and one fasting blood sample, underwent anthropometric measurements (height, weight, BMI) and completed a 7-day food record, a 7-day activity log and an environmental questionnaire before the liver biopsy. Healthy adults undergoing assessment for living liver donation were approached during their first screening appointment. Upon consent, subjects completed the same tests. Histology was obtained during a pre-donation biopsy (performed to verify the healthy state of the liver) or during hepatectomy. The study participants’ demographics, smoking, alcohol consumption history, medications and supplement use were reviewed and collected.
The food record included all food and beverages consumed over seven days using the 2D Food Portion Visual Chart (Nutrition Consulting Enterprises, Framingham, MA) to estimate portion sizes. Macro- and micronutrient intakes were calculated using Food Processor Diet and Nutrition Analysis Software (Version 7, ESHA Research, Salem, OR).
Physical activity logs were recorded for seven days concurrent with the food records. Participants were asked to list any activity, duration, and intensity level (mild, moderate, strenuous, and very strenuous). Daily physical activity units were calculated: 1 unit = 30 minutes mild, 20 minutes moderate, 10 minutes strenuous, or 5 minutes very strenuous activity20.
Liver biopsies were taken percutaneously (needle biopsy) for NAFLD patients and intra-operatively (wedge biopsy) for HC. In some HC a pre-donation needle biopsy was used. A portion of each biopsy was fixed in 10% formalin, embedded in paraffin wax, sectioned, and stained with hematoxylin and eosin for morphologic evaluation and Prussian Blue stain to rule out iron loading. Steatosis, inflammation, and fibrosis were assessed using the Brunt system21. Disease severity was additionally evaluated using the NAFLD Activity Score (NAS)22. The pathologist was blinded to the study.
Blood was drawn in a fasting state. Measures of glucose metabolism included plasma glucose, insulin, hemoglobin A1c (HbA1c), and homeostasis model assessment estimated IR (HOMA-IR)23. Liver enzymes and lipid profile (total cholesterol, low density lipoprotein (LDL), high density lipoprotein (HDL) and triglycerides (TG)) were measured at the UHN Laboratory Medicine Program using the Architect c8000 system (Abbott Laboratories, Abbott Park, Illinois). LDL was calculated from total cholesterol – HDL. Fasting plasma glucose and plasma insulin were measured by the enzymatic hexokinase method and radioimmunoassay, respectively24.
Each patient and HC collected one stool sample, which was frozen immediately in their home freezer (−20 °C). Within 24 hours, they brought the frozen sample in an insulated bag with cooling elements to UHN, where it was stored at −80 degree centigrade. Subsequently, stool samples were transferred into a sterile masticator bag, thawed and homogenized using a masticator blender (IUL, S.A., Barcelona, Spain). The samples were aliquoted, immediately placed on dry ice, and then transferred to −80 °C for storage until DNA extraction and metabolite analysis.
For IM analysis, bacterial DNA was extracted using the E.Z.N.A.TM Stool DNA Isolation Kit (Omega Bio-Tek, Doraville, GA, USA) as previously described9. Fecal DNA samples were stored at −80 °C until being amplified by PCR using the Earth Microbiome V4 primer set25, with the addition of combinatorial in-line barcodes so that all the samples could be sequenced in the same sequencing run26. The amplified DNA was then sequenced on the Illumina MiSeq platform with paired end 220 nucleotide reads, producing 25 million reads in total. Reads were overlapped with PANDAseq assembler27, clustered into Operational Taxonomic Units (OTUs) using UCLUST28, and annotated with the SILVA database29 using mothur30, producing a table of counts per OTU per sample. 7.5 million of the reads were successfully overlapped and annotated into 532 OTUs. A generalized workflow for processing 16S rRNA gene sequencing reads is available at https://github.com/ggloor/miseq_bin. Two sequencing runs were conducted with the results combined into one data matrix which was used for the analysis. The resulting data matrix was filtered at 1% OTU relative abundance (OTUs had to be at least 1% abundant in at least 1 sample) and the remaining OTUs (161) were included into the analysis. The lowest read count per sample after OTU filtering was 15,790, and the highest read count per sample was 210,867. The sequencing and pre-processing data for analysis was performed at the Microbiota Profiling Service, Department of Biochemistry, University of Western Ontario, London, Ontario, Canada.
To further evaluate the sequencing data, the abundance of selected taxa was estimated in fecal samples using qPCR. Primers and probe sequences used in this study are listed in Supplementary Table 2. These included: total bacteria, Bacteroidetes, Alistipes, Coprococcus, Ruminococcus, Lactobacillus and F. prausnitzii. The runs were performed in triplicates using 50 ng of fecal DNA in the 7900HT thermocycler (Applied Biosystems, Foster City, CA) as previously described9. The number of cells for each bacterial group in fecal samples was calculated from standard curves and expressed as colony forming unit per gram (CFU/g) of wet feces, normalized to total counts (relative abundance).
Serum metabolites, including ethanol, were measured at the Metabolomic Innovation Centre (University of Alberta, Edmonton, AB, Canada) using nuclear magnetic resonance (NMR) spectrometry on a 500 MHz Inova (Varian Inc., Palo Alto, CA) spectrometer (Varian Inc., Palo Alto, CA). For stools, NMR spectroscopy was also used from the same center to identify eight metabolites of interest (acetic acid, butyric acid, formic acid, isobutyric acid, isovalerate, L-lactic acid, propionate, succinate). For the measurement, fecal samples were prepared by mixing 20 mg of frozen fecal material with 1 mL of saline phosphate buffer in deuterium oxide, followed by centrifugation (18,000 × g, 1 min). Fecal supernatants were filtered through 0.2 μm membrane filters31.
Statistical Analysis
Descriptive summaries of all measurements were calculated: means and standard deviations (SD) for normal variables, medians and first and third quartiles for skewed variables, and proportions for categorical variables. Plots were used as appropriate to examine the data. Bivariate relationships were assessed using Pearson’s or Spearman’s correlation coefficients, Chi-square and Fisher exact tests where appropriate. Comparisons across diagnosis groups were performed using one-way ANOVA followed by the t-test (normally distributed data), data were log-transformed to normalize distribution if necessary, or using the Kruskal-Wallis test followed by Wilcoxon rank-sums test (not normally distributed data). All tests were two-sided and performed at the 5% significance (alpha) level. Bonferroni’s correction method was used to account for multiple comparisons between diagnostic groups.
For next generation sequencing, differences in overall IM characteristics of diagnostic groups were assessed in QIIME32. Principal coordinate analysis based on weighted UniFrac distance matrix33 was performed to visualize and examine differences in overall microbial community compositions among the three groups. For differential abundance analysis read counts were quantile normalized34. In order to ensure robustness of our conclusions to statistical assumptions we applied 3 different analysis methods and made inferences if the results of all analyses were consistent. First, zero-inflated Gaussian models to perform 3-group comparisons35. Then pairwise differences between groups were assessed by two non-parametric methods: Metastats35,36 and Wilcoxon rank sum test (details in Supplemental Materials). Correction for multiple comparisons was done using Benjamini-Hochberg false discovery rate37, and q-value < 0.05 was considered statistically significant. In some cases, following a recent publication15 we were less stringent in controlling for multiple comparisons, and considered higher q-value thresholds (but less than 0.1).
For qPCR data, due to non-normality and heteroscedasticity of distribution for most groups of bacteria, we used non-parametric methods: Kruskal-Wallis and Wilcoxon rank sum tests for between group comparisons, and Spearman correlation coefficients and partial Spearman correlation coefficients to examine relationship between relative abundances and other variables (diagnosis (treated as ordinal variable), BMI, IR, metabolites).
Analysis was performed using tools SAS 9.4, R 3.2.5 and QIIME software.
Data Availability
The datasets generated during and analysed during the current study are not publicly available due to privacy restrictions but some blinded data may be made available from the corresponding author on reasonable request.
Results
Patient characteristics
The study included 15 patients with biopsy-proven simple steatosis (SS), 24 with nonalcoholic steatohepatitis (NASH) and 28 healthy liver donors from the living donor transplant program as HC. Patient demography and clinical characteristics are presented in Table 1. The HC group was younger compared to both SS and NASH with significantly lower median BMI compared to NASH. There were no significant differences in gender distribution. As expected, liver transaminases were progressively increased from HC to SS to NASH. Four NASH and 1 SS patients who were taking anti-diabetic drugs (none were on insulin therapy) as well as 3 patients with HOMA-IR >15 (possibly a consequence of non-fasting before the blood test), were excluded from analyses related to diabetes parameters (HbA1c, HOMA-IR). SS and NASH patients had significantly higher HbA1C compared to HC, and NASH had higher HOMA-IR compared to both SS and HC.
Table 1.
Characteristics of patients included in the study.
| HC (n = 28)* n (%) |
SS (n = 15)* n (%) |
NASH (n = 24)* n (%) |
p-value** | |
|---|---|---|---|---|
| Gender, male | 15 (54%) | 9 (60%) | 11 (46%) | 0.7 |
| Ethnicity, Caucasian (n = 41) | 16 (80%) | 4 (44%) | 7 (58%) | 0.15 |
| Median (min, max) | Median (min, max) | Median (min, max) | ||
| Age, yrs | 36.5A,B (21, 58) | 48B (33, 61) | 46.5A (29, 68) | 0.0003 |
| BMI, kg/m2 | 26.6A (19.5, 35.3) | 27.4 (23.5, 44.2) | 32.1A (24.17, 49.53) | <0.0001 |
| AST, U/L | 19.5B,C (12, 29) | 26A,C (16, 53) | 45A,B (18, 114) | <0.0001 |
| ALT, U/L | 17.5B,C (7, 41) | 45A,C (14, 116) | 70A,B (22, 168) | <0.0001 |
| HbA1c*** (n = 62) | 0.05A,B (0.04, 0.07) | 0.06B (0.05, 0.09) | 0.06A (0.05, 0.07) | 0.0003 |
| HOMA-IR*** (n = 48) | 1.16A (0.53, 7.59) | 1.12B (0.54, 6.83) | 3.27A,B (1.21, 14.54) | 0.0006 |
| NAFLD Activity Score (n = 53) | 0 | 1 (1, 4) | 4.5 (3, 8) | <0.0001 |
*For some variables numbers were lower due to missing values/exclusions.
**Kruskal Wallis test or Chi-square test as appropriate.
***Patients on diabetes drugs excluded.
Same subscript letters denotes significant differences.
HC: healthy controls, SS: simple steatosis, NASH: nonalcoholic steatohepatitis, AST: aspartate transaminase, ALT: alanine transaminase, HbA1c: Hemoglobin A1c, HOMA-IR: homeostasis model assessment estimated insulin resistance.
For the liver histology, NAFLD Activity Score for SS patients ranged from 1 to 4 (median 1), and for NASH patients from 3 to 8 (median 4.5). Details on liver histology (Supplementary Table 3), level of activity (Supplementary Table 4), environmental questionnaire (Supplementary Table 5), and diet (Supplementary Tables 6 and 7) are reported in the supplementary section. Overall, there were no significant differences in level of activity and diet composition. There was a higher proportion of HC born in Canada but ethnicity was similar between groups for those who filled out the questionnaire.
Intestinal Microbiome
Next generation sequencing data
161 OTUs passed a 1% filter and were retained for analysis. The filtered OTUs were aggregated (after normalization) into higher taxonomic levels: 8 phyla, 25 families and 44 genera.
The analysis of overall community structure and composition, including Simpson diversity metric and principal coordinate analysis did not reveal differences between the groups. Simpson diversity was not significantly different between the 3 groups (p-value 0.5) (Supplementary Figure 1). There was no clear separation between groups in weighted UniFrac principal coordinate analysis plot (Supplementary Figure 2).
To investigate differences between diagnostic groups in differential abundance of individual taxa, we first fit ANOVA ZIG model comparing 3 groups on OTU and genus levels, followed by pairwise comparisons between each diagnostic category (Supplementary Data). Size and direction of differences for HC vs SS and HC vs NASH comparisons were similar in both OTU and genus levels, and differentially abundant groups were similar in both of these comparisons. At the same time, the differences between SS and NASH groups were never significant and size of differences was small, suggesting similar microbiome characteristics for NASH and SS in our study. Based on this and taking into account low sample size in SS group, we combined NASH and SS into one NAFLD group for further analysis.
Based on the results of Metastats and Wilcoxon tests we identified 7 OTUs, 10 genera (for 4 of those q-value in Wilcoxon test did not reach 0.05 threshold, but was <0.1), and 5 families as less abundant in NAFLD than in HC. One OTU, 1 genus (Lactobacillus), and 1 family (Lactobacillaceae) were more abundant in NAFLD. Two dominant phyla, Bacteroidetes and Firmicutes were less abundant in NAFLD (Table 2). Differentially abundant genera are further summarized in Fig. 2.
Table 2.
Taxa identified as differentially abundant in patients with NAFLD (SS or NASH) compared to HC by Metastats and Wilcoxon rank sum test.
| Taxa | Metastats | Wilcoxon test | Quantile normalized counts | ||||||
|---|---|---|---|---|---|---|---|---|---|
| median (min, max) | |||||||||
| log fold change1 | p-value min2 | p-value max2 | q-value min2 | q-value max2 | p-value | q-value | HC | NAFLD | |
| n = 28 | n = 39 | ||||||||
| OTU level | |||||||||
| Firmicutes;Clostridia;Clostridiales; Lachnospiraceae;Anaerostipes;11 | −0.761 | 0.0002 | 0.0007 | 0.008 | 0.028 | 0.0003 | 0.017 | 873.8 | 513.2 |
| (412.1, 3191.3) | (181.8, 2543.3) | ||||||||
| Firmicutes;Clostridia;Clostridiales; Lachnospiraceae;Incertae_Sedis; 13 | −0.681 | 0.002 | 0.003 | 0.039 | 0.06 | 0.001 | 0.021 | 636.6 | 399.4 |
| (142.5, 3652.8) | (183.2, 2090) | ||||||||
| Firmicutes;Clostridia;Clostridiales; Ruminococcaceae;Faecalibacterium; 2 | −0.751 | 0.002 | 0.002 | 0.039 | 0.046 | 0.001 | 0.029 | 1670.9 | 946.2 |
| (308.6, 8565.5) | (375.5, 5075.3) | ||||||||
| Firmicutes;Clostridia;Clostridiales; Lachnospiraceae;unclassified; 20 | −0.577 | 0.001 | 0.002 | 0.039 | 0.046 | 0.001 | 0.025 | 434.3 | 277.3 |
| (133.9, 3712.2) | (159.3, 831.9) | ||||||||
| Bacteroidetes;Bacteroidia;Bacteroidales; Bacteroidaceae;Bacteroides; 23 | −0.989 | 0 | 0.0001 | 0 | 0.016 | 0.0004 | 0.017 | 492.4 | 194.4 |
| (128.7, 1768.7) | (95.9, 882.5) | ||||||||
| Firmicutes;Clostridia;Clostridiales; Lachnospiraceae;Dorea; 31 | −0.77 | 0.0002 | 0.0003 | 0.008 | 0.016 | 0.0004 | 0.017 | 267.6 | 140.5 |
| (86.6, 1280.1) | (64.3, 503.7) | ||||||||
| Firmicutes;Clostridia;Clostridiales; Lachnospiraceae;Blautia; 6 | −0.685 | 0 | 0.0002 | 0 | 0.016 | 0.0004 | 0.017 | 1182.7 | 657.1 |
| (398.9, 3296.7) | (338.4, 1959) | ||||||||
| Firmicutes;Bacilli;Lactobacillales; Lactobacillaceae;Lactobacillus; 84 | 1.258 | 0.007 | 0.008 | 0.076 | 0.078 | 0.001 | 0.019 | 9.8 | 23.8 |
| (2.1, 740) | (1.6, 701.6) | ||||||||
| Genus level | |||||||||
| Alistipes | −1.307 | 0.0003 | 0.001 | 0.004 | 0.005 | 0.001 | 0.005 | 126.8 | 38.9 |
| (21.8, 1779.6) | (18.5, 696) | ||||||||
| Anaerostipes | −0.761 | 0.001 | 0.001 | 0.005 | 0.009 | 0.0003 | 0.005 | 873.8 | 513.2 |
| (412.1, 3191.3) | (181.8, 2543.3) | ||||||||
| Bacteroides | −0.874 | 0.0004 | 0.001 | 0.004 | 0.006 | 0.0004 | 0.005 | 2798.4 | 1117.2 |
| (654.2, 6083.6) | (425.9, 7385.7) | ||||||||
| Blautia | −0.48 | 0.002 | 0.003 | 0.014 | 0.021 | 0.003 | 0.016 | 4205.6 | 2823.6 |
| (2040, 8236) | (1154, 10964) | ||||||||
| Coprococcus | −0.778 | 0.005 | 0.006 | 0.027 | 0.03 | 0.014 | 0.06 | 850.5 | 413.6 |
| (174.6, 2877.3) | (151.1, 1848.1) | ||||||||
| Dorea | −0.596 | 0.004 | 0.006 | 0.027 | 0.031 | 0.01 | 0.048 | 1414.4 | 890 |
| (574.1, 6919.3) | (425.7, 4320.7) | ||||||||
| Faecalibacterium | −0.599 | 0.006 | 0.006 | 0.027 | 0.031 | 0.003 | 0.017 | 2837.1 | 1663.1 |
| (552.3, 8886.2) | (699.4, 8785.9) | ||||||||
| Lactobacillus | 1.232 | 0.013 | 0.016 | 0.046 | 0.05 | 0.001 | 0.005 | 20.7 | 54.6 |
| (2.9, 5067.1) | (9.2, 720.9) | ||||||||
| Parabacteroides | −1.045 | 0.012 | 0.012 | 0.046 | 0.047 | 0.017 | 0.068 | 76.6 | 24.5 |
| (7, 438.9) | (4.7, 423.6) | ||||||||
| Roseburia | −0.426 | 0.015 | 0.016 | 0.047 | 0.05 | 0.021 | 0.071 | 600 | 432.9 |
| (266.7, 1926.5) | (218.7, 1412.6) | ||||||||
| Ruminococcus | −0.7 | 0.008 | 0.011 | 0.035 | 0.047 | 0.019 | 0.07 | 1846.1 | 692.3 |
| (323.5, 4628) | (347.5, 3160.2) | ||||||||
| Family level | |||||||||
| Bacteroidaceae | −0.874 | 0.0004 | 0.001 | 0.003 | 0.004 | 0.0004 | 0.003 | 2798.4 | 1117.2 |
| (654.2, 6083.6) | (425.9, 7385.7) | ||||||||
| Lachnospiraceae | −0.563 | 0.0001 | 0.0002 | 0.001 | 0.003 | 0.0001 | 0.002 | 16822.1 | 12328 |
| (9212, 34155) | (6477, 26139) | ||||||||
| Lactobacillaceae | 1.232 | 0.012 | 0.013 | 0.043 | 0.046 | 0.001 | 0.004 | 20.7 | 54.6 |
| (2.9, 5067.1) | (9.2, 720.9) | ||||||||
| Porphyromonadaceae | −1.413 | 0.001 | 0.001 | 0.003 | 0.004 | 0.001 | 0.005 | 163.8 | 37.3 |
| (9.3, 7343.7) | (10.1, 463.1) | ||||||||
| Rikenellaceae | −1.307 | 0.0001 | 0.001 | 0.001 | 0.004 | 0.001 | 0.004 | 126.8 | 38.9 |
| (21.8, 1779.6) | (18.5, 696) | ||||||||
| Ruminococcaceae | −0.774 | 0 | 0.0001 | 0 | 0.001 | 0 | 0.001 | 8298.6 | 4881.6 |
| (3328, 17756) | (2139, 14877) | ||||||||
| Phylum level | |||||||||
| Bacteroidetes | −0.763 | 0.003 | 0.003 | 0.012 | 0.013 | 0.003 | 0.011 | 3744 | 1497.7 |
| (787.4, 12620.1) | (613.1, 10942.2) | ||||||||
| Firmicutes | −0.622 | 0 | 0 | 0 | 0 | 0 | 0.0003 | 30975.9 | 21209.9 |
| (19486, 57571) | (10571, 46704) | ||||||||
1Negative parameter denotes that taxon is more abundant in HC, positive – that it is more abundant in NAFLD.
2Estimated in 3 rounds of permutations, 10,000 instances each.
HC: healthy controls, SS: simple steatosis, NAFLD: nonalcoholic fatty liver disease, NASH: nonalcoholic steatohepatitis.
Figure 2.
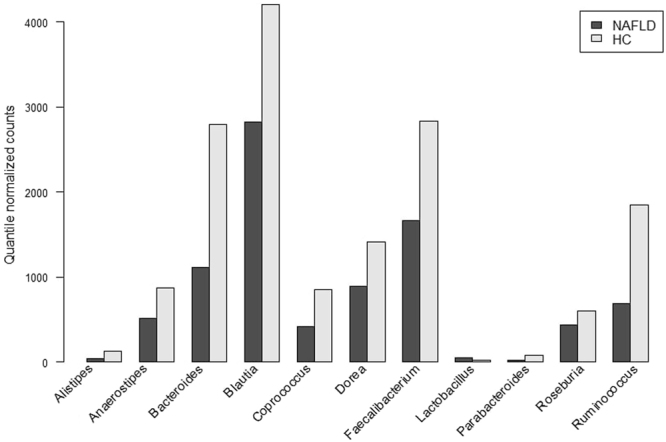
Medians of differentially abundant genera based on 16S sequencing data.
qPCR Data
Based on 16S data analysis results and similar taxa previously reported11,12,14,15, the following taxa were selected to be quantified by qPCR: Bacteroidetes which included Parabacteroides and Bacteroides genera found on sequencing as well as F. prausnitzii, Alistipes, Coprococcus, Ruminococcus and Lactobacillus. The relative abundance of the targeted bacterial groups is presented in Table 3.
Table 3.
Relative abundances of bacterial taxa quantified by quantitative PCR.
| HC | SS | NASH* | p-value** | ||||
|---|---|---|---|---|---|---|---|
| n=28 | n=15 | n=24 | |||||
| Median | (min, max) | Median | (min, max) | Median | (min, max) | ||
| Bacteroidetes | 2.4 | (0.1, 30.5) | 3.5 | (0.1, 10.8) | 0.78 | (0, 13.3) | 0.19 |
| Alistipes | 1 | (0, 5.1) | 0.2 | (0, 4.6) | 0.6 | (0, 5.9) | 0.14 |
| Coprococcus | 0.4A, B | (0, 15.4) | 0.0A | (0, 2.6) | 0.0B | (0, 1.2) | 0.005 |
| Lactobacillus | 0 | (0, 0.4) | 0 | (0, 0.6) | 0 | (0, 0.3) | 0.7 |
| Ruminococcus | 13.2A, B | (7.9, 27.4) | 2.6A | (0.3, 16.4) | 2.0B | (0, 9.6) | <0.0001 |
| Faecalibacterium prausnitzii | 11.7A, B | (0.4, 22.1) | 2.2A | (0, 10.2) | 1.7B | (0, 12.4) | <0.0001 |
Abundances were calculated as ratio of colony forming units (CFU) to total bacteria CFU in the sample. Same subscript denotes significant difference (after Bonferroni correction).
HC: Healthy controls, SS: simple steatosis, NASH: nonalcoholic steatohepatitis.
*For 1 NASH patient there was not enough DNA for Coprococcus, Alistipes and F. prausnitzii, so the sample size was reduced to n = 23.
**Kruskal Wallis test. Same subscript letters denote significant differences.
The following groups were less abundant in NASH and SS patients compared to HC: Ruminococcus, F. prausnitzii and Coprococcus (Table 3). Even though the median abundance for all taxa except Coprococcus were lower in NASH than in SS, the differences between the two patient groups did not reach statistical significance. There were no statistically significant differences between diagnostic groups for Bacteroidetes, Alistipes and Lactobacillus.
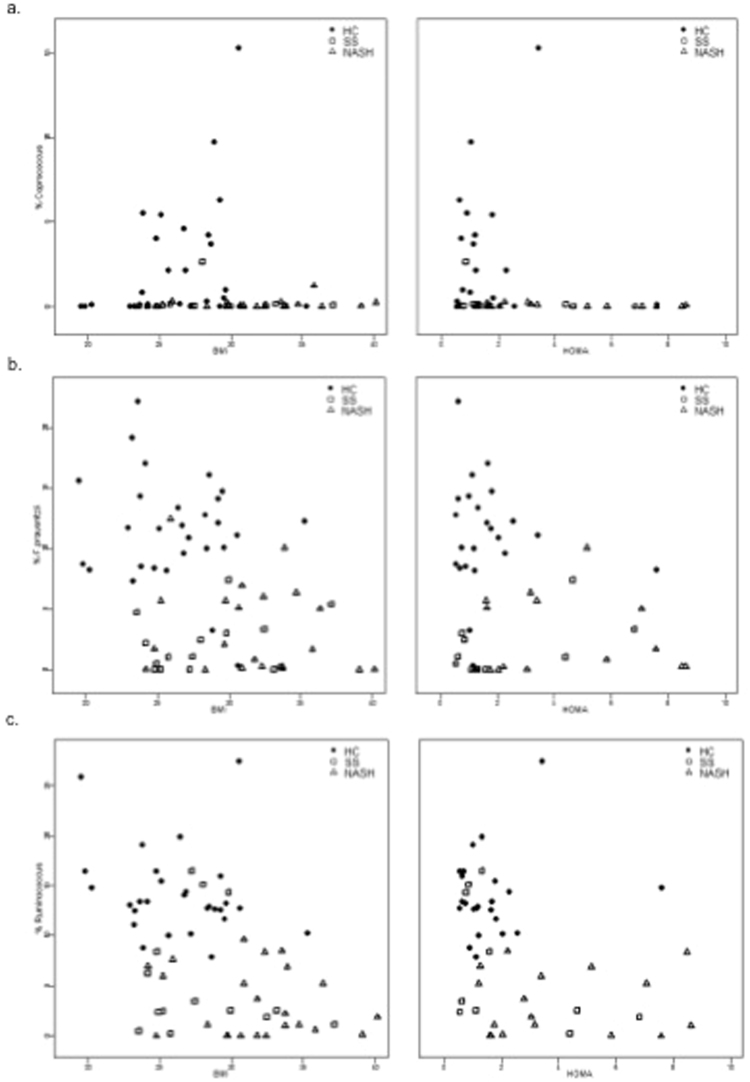
To investigate whether the association between taxa and NAFLD was due to confounding factors like BMI and IR, we calculated two sets of partial Spearman correlations: (1) between these factors and bacterial abundances, controlling for diagnosis and; (2) between bacterial abundances and diagnosis controlling for BMI and HOMA-IR (Table 4). For the taxa that differed between HC and NAFLD, partial correlations between diagnosis status and abundance were highly significant after controlling for BMI or IR. After controlling for diagnosis, there was no evidence of relationship between BMI and bacterial abundances, and for HOMA-IR only weak evidence of a positive relationship with F. prausnitzii (Fig. 3). Therefore, our analysis suggests that NALFD is associated with reduced abundance of Ruminococcus, Coprococcus and F. prausnitzii independent of BMI and IR.
Table 4.
Partial Spearman correlation coefficients (p-value); diagnosis was treated as ordinal variable (1-healthy control, 2-simple steatosis, 3-nonalcoholic steatohepatitis).
| BMI (controlling for diagnosis) | HOMA-IR (controlling for diagnosis) | diagnosis (controlling for BMI) | diagnosis (controlling for HOMA IR) | |
|---|---|---|---|---|
| %Coprococcus | 0.18 (0.1) | −0.11 (0.5) | −0.40 (0.0009) | −0.40 (0.006) |
| %Faecalibacterium prausnitzii | 0.02 (0.9) | 0.28 (0.057) | −0.61 (<0.0001) | −0.68 (<0.0001) |
| %Ruminococcus | −0.09 (0.5) | −0.04 (0.8) | −0.69 (<0.0001) | −0.70 (<0.0001) |
HOMA-IR: homeostasis model assessment estimated insulin resistance.
Figure 3.
Scatterplots illustrating relationship between relative abundance of relevant bacterial taxa by qPCR, diagnosis and body mass index (left panel), and homeostasis model assessment estimated insulin resistance (HOMA-IR, right panel) (a) Coprococcus; (b) F. prauznitzii (c) Ruminococcus.
We then tested whether there was a relationship between liver histology (% steatosis, lobular inflammation, ballooning, fibrosis, NAFLD activity score, as seen in Supplementary Table 3) and relative abundances of quantified taxa within the NAFLD group (NASH+SS, n = 38). The results were negative except for a weak negative relationship between Ruminococcus and lobular inflammation (ρ = −0.33, p-value 0.04) (Supplementary Table 8). Considering that these p-values were not controlled for multiple comparisons, this result can be due to Type 1 error.
Bacterial Products
NAFLD patients had significantly higher concentrations of fecal propionate and isobutyric acid as well as serum 2-hydroxybutyrate and L-lactic acid (Table 5). The ratio of the short-chain fatty acids (SCFA) acetate:propionate:butyrate in our HC and NAFLD patients was approximately 62:23:15 and 61.5:22.5:16, respectively, which is similar to commonly quoted ‘normal’ ratios of 60:20:20 and 60:25:1538. Serum ethanol was not different between groups. The abundance of Ruminococcus, F. prausnitzii and Coprococcus was not correlated with the concentration of the measured bacterial products (Supplementary Table 9).
Table 5.
Bacteria-related fecal and serum metabolites.
| Fecal Metabolite (μM) | Healthy Controls | NAFLD Patients | p-value | ||
|---|---|---|---|---|---|
| n | Median (Q1, Q3) | n | Median (Q1, Q3) | ||
| Butyric Acid | 28 | 1366 (993, 2072) | 38 | 1753 (1130, 2266) | 0.2 |
| Propionate | 28 | 2052 (1598, 2419) | 38 | 2485 (1985, 3649) | 0.019 |
| Acetic Acid | 28 | 5665 (4025, 7061) | 38 | 6781 (4619, 8384) | 0.088 |
| Formic Acid | 28 | 60 (50, 73) | 38 | 56 (42, 68) | 0.3 |
| Total SCFA | 28 | 9122 (6747, 11757) | 38 | 11366 (8097, 13320) | 0.05 |
| (sum of above) | |||||
| Isobutyric Acid | 28 | 249 (199, 285) | 38 | 293 (239, 372) | 0.017 |
| Isovalerate | 28 | 181 (115, 223) | 38 | 172 (136, 208) | 0.9 |
| Succinate | 28 | 121 (67, 175) | 38 | 94 (58, 187) | 0.5 |
| L-Lactic Acid | 28 | 37 (29, 47) | 38 | 41 (30, 58) | 0.3 |
| Serum Metabolite (μM) | Healthy Controls | NAFLD Patients | p-value | ||
| n | Median (Q1, Q3) | n | Median (Q1, Q3) | ||
| Acetic Acid | 26 | 48 (3767) | 33 | 37 (33, 59) | 0.09 |
| Formic Acid | 26 | 55 (47, 63) | 33 | 58 (51, 61) | 0.4 |
| 2-Hydroxybutyrate | 26 | 48 (39, 66) | 33 | 69 (48, 78) | 0.0096 |
| Isobutyric Acid | 26 | 0 (0, 0) | 33 | 0 (0, 0) | 0.2 |
| L-Lactic Acid | 26 | 1370 (1069, 1807) | 33 | 1958 (1523, 2288) | 0.0056 |
| Ethanol | 26 | 23.5 (0, 50) | 33 | 0 (0, 41) | 0.8 |
| Endotoxin (EU/mL) | 24 | 6.0 (5.4, 6.4) | 38 | 6.1 (5.3, 6.5) | 0.7 |
NAFLD: nonalcoholic fatty liver disease = patients with simple steatosis and those with nonalcoholic steatohepatitis combined, SCFA: short-chain fatty acids.
Discussion
We demonstrated, in a well-characterized adult population, that NAFLD was associated with reduced abundance of several bacterial taxa (Ruminococcus, Coprococcus and F. prausnitzii) independent of BMI and IR. Additionally, selected bacterial products related to fermentation (SCFA) were higher in NAFLD patients compared to HC. Our results are consistent with previous studies, comparing NAFLD and HC11–14 and investigating the association between IM and NAFLD severity15. In all of these studies, comparisons were cross-sectional, except for Wong et al. who also compared IM of NASH patients before and after probiotic intervention. All reported evidence of dysbiosis in NAFLD patients when compared to HC. Only one study14 used RT-PCR to validate the results of the next-generation sequencing analyses but only with one genus (Lactobacillus). A pediatric study11 is also the only one that controlled for confounding factors when assessing the association between IM and NAFLD.
The patient groups in this study differed as expected based on the disease state. SS and NASH patients were older and NASH had a significantly higher BMI than HC. Liver enzymes and insulin resistance measures were also higher in the NAFLD groups. All of these findings were expected given the typical NAFLD population39. In regard to age, previous studies have identified changes in IM over the lifespan, however, most changes occur in early childhood and elderly adults >70 years old40–42. Research into IM changes in the elderly has often been confounded by dietary changes, medication use, and declining overall health. Therefore, a cut-off age for IM change is undeterminable43. Our total population, with a maximum age of 68 years is therefore unlikely to have experienced a confounding effect of age on IM. In regard to BMI, the median BMI of the HC was in the overweight range 26.6 (19.5, 35.3) [median (min, max)]. However, the groups were characterized by liver histology, and HC were confirmed to have a normal liver. This is a strength in our study as, to our knowledge, no other studies published on NAFLD had a HC group with confirmed normal liver biopsies. Another strength is the HC group with a median BMI in the overweight range. This BMI range in the HC group is reflective of the current Canadian population44. The smaller difference in BMI between NAFLD and HC reduces potential confounding effects on IM when comparing the groups.
The lower abundance of Coprococcus or Faecalibacterium in NAFLD was consistent with other studies11,13,14. Ruminococcus was previously reported only by Zhu et al., but the family Ruminococcaceae was consistently reported as less abundant in NAFLD12,14. In our study, liver biopsy was used to diagnose NAFLD while some of these previous studies12,14 relied mostly on imaging. Also, their healthy controls had no liver biopsy, were much leaner compared to NAFLD12–14 and differences in BMI were not considered in the analysis. This could have influenced the results.
Our analyses did not show any association between IM and NAFLD severity or correlation between specific taxa and histological parameters. This is contrary to the findings by Boursier et al.15, who reported higher abundance of Bacteroides in NASH compared to SS patients and a positive association between Ruminococcus and fibrosis severity, independent of metabolic factors (assessed only by the presence or absence of diabetes). The diverging results can have multiple explanations. First, there can be differences in disease severity in study samples: our patients had milder disease (NAFLD activity score median 1 for SS and 4.5 for NASH) and only 18% had severe fibrosis or cirrhosis (stage 3 or 4). Secondly, differences in patient populations, environmental factors and dietary habits in Canada vs Europe, could have influenced associations between IM and disease severity. Finally, in our study, more stringent criteria to select differentially abundant species by controlling for multiple comparisons were applied compared to Boursier’s study, which did not correct for increased chances of type I error when testing the hypothesis of differential abundance for multiple taxa.
Although the sequencing data showed that the Bacteroidetes phylum and one OTU within the phylum were lower in NAFLD versus HC, this was not confirmed by qPCR. Using qPCR, we previously showed9 a lower percentage of Bacteroidetes in NASH versus HC and SS. However, it is possible that since the qPCR primers targeted all Bacteroidetes, we may have missed sub-phylum differences if the differences were less pronounced in this study compared with results from Mouzaki et al.9. Others also reported conflicting results for this taxon. Boursier et al. found higher proportion of Bacteroidetes in NASH compared to SS15, Zhu et al. also found a higher proportion in pediatric NASH patients and those with obesity compared to lean controls11, whereas others12,14 did not mention it among differentially abundant taxa in NAFLD.
We found lower abundance of F. prausnitzii, Coprococcus and Ruminococcus in NAFLD patients. F. prausnitzii is an anti-inflammatory commensal bacterium and its abundance is reduced in inflammatory bowel disease45 and metabolic syndrome46. Coprococcus was also lower in NAFLD patients, similar to the report by Zhu et al.11. A lower abundance of Coprococcus has also been observed in other inflammatory conditions47. Therefore, it is conceivable that lower abundance can promote chronic inflammation that may contribute to NAFLD pathogenesis. Ruminococcus belong to the family Ruminococcaceae which are also fermenting anaerobes that lead to the production of SCFA48. Both Zhu et al.11 and Raman et al.12 found a lower abundance of Ruminococcaceae in NAFLD patients, however, not in this particular genus.
We expected to see differences in SCFA between groups. However, in feces, only propionate and isobutyric acid were higher in NAFLD compared to HC, whereas concentrations of butyrate, acetate, formate, and total SCFA were not different. There were no correlations between SCFA and bacterial abundances. The ratios of acetate:propionate:butyrate were similar to the normal values in both groups38. However, the amount and type of products can vary depending on species. For example, Ruminococcus genus contains species and strains which can be metabolically versatile49–51 which means the amount and type of fermentation products vary according to species. Some can use different substrates like mucus or cellulose, resulting in production of acetate, ethanol, succinate, lactate and formate, but very little butyrate as end products of glucose metabolism51. We also found higher serum 2-hydroxybutyrate and L-lactic acid. However, it is difficult to compare our results to previous studies as human data on SCFA in NAFLD are scarce. Wong et al. found that NASH patients had a lower prevalence of butyrate producing Faecalibacterium (similar to our study), but they did not measure butyrate levels13. Raman et al.12, comparing obese NAFLD diagnosed on imaging versus lean HC subjects, measured fecal volatile compounds and also showed higher levels of SCFA with elevated fecal propionic, butyric and acetic acids in NAFLD. Similarly increased volatile compounds were observed in NAFLD patients16 and higher total fecal SCFA, particularly propionate, was also found in overweight and obese adults compared to lean controls52. Higher SCFA from bacterial fermentation can lead to increased energy absorption of up to 150 kcal per day53. It is conceivable that the higher concentration of fecal propionate and isobutyric acid found in our NAFLD group reflect this phenomenon. Mechanisms other than IM could have also contributed to the higher level of SCFA in NAFLD versus HC, such as slow transit time38 reported in obesity54 which may have increased fermentation, or differences in absorption rate55. 2-hydroxybutyrate (alpha-hydroxybutyrate) is a metabolite formed during amino acid catabolism and glutathione anabolism56. Although this compound has been mentioned as an intermediary in the bacterial metabolism of amino acids, it is not clear how much is actually produced and absorbed in the human colon and absorbed into the blood57. Oxidative stress in the liver that can be triggered by IR may have contributed to higher 2-hydroxybutyrate level by increasing the production of hepatic glutathione56,58.
The strengths of our study were the precise characterization of HC and NAFLD patients through liver biopsy, and the documentation of dietary intake, physical activity, and environmental factors. We applied robust statistical methodology appropriate for analysis of this type of data, including normalization methods and statistical testing. Particularly, we used 2 types of statistical tests for sequencing data to ensure robustness of conclusions. We also applied multiple comparisons corrections whenever appropriate to avoid identifying false positive taxa and confirmed the results with qPCR. The limitation of our study is the relatively low sample size that does not allow for more multivariate analyses and may have prevented us from showing differences between IM of SS and NASH. However, the sample size was similar to other studies11,13 that used liver biopsy to characterize NAFLD. The expanding study of the IM discovered numerous environmental factors that may influence the IM. Vitamin D deficiency, for instance, has recently been linked to dysbiosis and NAFLD59. In this study, vitamin D status was not examined, but the patient population was from a relatively small geographic region, and vitamin D intake did not differ between groups. Also, we cannot confirm causality owing to the observational design.
In summary, NAFLD had lower abundance of Ruminococcus, F. prausnitzii and Coprococcus independent of BMI and IR, and higher concentrations of select fecal and serum metabolites, which may suggest a specific IM community and functional profile in these patients. Future metagenomic research would allow for better characterization of this functional profile.
Electronic supplementary material
Acknowledgements
We thank the Microbiota Profiling Service, Department of Biochemistry, University of Western Ontario, London, ON, Canada, for sequencing and pre-processing the data and the Metabolomic Innovation Centre, University of Alberta, AB, Canada, for the measurements of fecal and serum metabolites. Some of the equipment used in this study was supported by the 3D (Diet, Digestive Tract and Disease) Centre funded by the Canadian Foundation for Innovation and Ontario Research Fund, project number 19442 and 30961. This study was funded through operating grants from the Canadian Institutes of Health Research (Grants NMD-86922, MOP-89705) and the American College of Gastroenterology. EMC holds the Lawson Family Chair in Microbiome Nutrition Research at the University of Toronto.
Author Contributions
J.P.A., B.M.A. designed research, develop research plan and provided study oversight; B.M.A., H.E.D. conducted research; E.M.C., A.Ta. and S.E.F. provided essential materials necessary for research and A.Te. developed database; H.E.D.S., A.Te., W.L. analyzed data and performed statistical analysis; H.E.D.S., A.Te., E.M.C., J.P.A. wrote the paper; J.P.A. had primary responsibility for final content. All authors read and approved the final manuscript.
Competing Interests
The authors declare that they have no competing interests.
Footnotes
Electronic supplementary material
Supplementary information accompanies this paper at 10.1038/s41598-018-19753-9.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Younossi, Z. M. et al. Changes in the prevalence of the most common causes of chronic liver diseases in the United States from 1988 to 2008. Clin Gastroenterol Hepatol9, 524–530.e521; quiz e560, 10.1016/j.cgh.2011.03.020 (2011). [DOI] [PubMed]
- 2.Vanni E, et al. From the metabolic syndrome to NAFLD or vice versa? Dig Liver Dis. 2010;42:320–330. doi: 10.1016/j.dld.2010.01.016. [DOI] [PubMed] [Google Scholar]
- 3.Rotman Y, Koh C, Zmuda JM, Kleiner DE, Liang TJ. The association of genetic variability in patatin-like phospholipase domain-containing protein 3 (PNPLA3) with histological severity of nonalcoholic fatty liver disease. Hepatology. 2010;52:894–903. doi: 10.1002/hep.23759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Farrell GC, Larter CZ. Nonalcoholic fatty liver disease: from steatosis to cirrhosis. Hepatology. 2006;43:S99–S112. doi: 10.1002/hep.20973. [DOI] [PubMed] [Google Scholar]
- 5.Alam S, Mustafa G, Alam M, Ahmad N. Insulin resistance in development and progression of nonalcoholic fatty liver disease. World journal of gastrointestinal pathophysiology. 2016;7:211–217. doi: 10.4291/wjgp.v7.i2.211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Marchesini G, Petta S, Dalle Grave R. Diet, weight loss, and liver health in nonalcoholic fatty liver disease: Pathophysiology, evidence, and practice. Hepatology. 2016;63:2032–2043. doi: 10.1002/hep.28392. [DOI] [PubMed] [Google Scholar]
- 7.Wieland A, Frank DN, Harnke B, Bambha K. Systematic review: microbial dysbiosis and nonalcoholic fatty liver disease. Aliment Pharmacol Ther. 2015;42:1051–1063. doi: 10.1111/apt.13376. [DOI] [PubMed] [Google Scholar]
- 8.Chassaing B, Etienne-Mesmin L, Gewirtz AT. Microbiota-liver axis in hepatic disease. Hepatology. 2014;59:328–339. doi: 10.1002/hep.26494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mouzaki M, et al. Intestinal microbiota in patients with nonalcoholic fatty liver disease. Hepatology. 2013;58:120–127. doi: 10.1002/hep.26319. [DOI] [PubMed] [Google Scholar]
- 10.Spencer, M. D. et al. Association between composition of the human gastrointestinal microbiome and development of fatty liver with choline deficiency. Gastroenterology140, 976–986, 10.1053/j.gastro.2010.11.049 (2011). [DOI] [PMC free article] [PubMed]
- 11.Zhu L, et al. Characterization of gut microbiomes in nonalcoholic steatohepatitis (NASH) patients: a connection between endogenous alcohol and NASH. Hepatology. 2013;57:601–609. doi: 10.1002/hep.26093. [DOI] [PubMed] [Google Scholar]
- 12.Raman M, et al. Fecal microbiome and volatile organic compound metabolome in obese humans with nonalcoholic fatty liver disease. Clin Gastroenterol Hepatol. 2013;11(868–875):e861–863. doi: 10.1016/j.cgh.2013.02.015. [DOI] [PubMed] [Google Scholar]
- 13.Wong VW, et al. Molecular characterization of the fecal microbiota in patients with nonalcoholic steatohepatitis–a longitudinal study. PLoS One. 2013;8:e62885. doi: 10.1371/journal.pone.0062885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jiang W, et al. Dysbiosis gut microbiota associated with inflammation and impaired mucosal immune function in intestine of humans with non-alcoholic fatty liver disease. Scientific reports. 2015;5:8096. doi: 10.1038/srep08096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Boursier J, et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology. 2016;63:764–775. doi: 10.1002/hep.28356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Del Chierico F, et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology. 2017;65:451–464. doi: 10.1002/hep.28572. [DOI] [PubMed] [Google Scholar]
- 17.Leung C, Rivera L, Furness JB, Angus PW. The role of the gut microbiota in NAFLD. Nat Rev Gastroenterol Hepatol. 2016;13:412–425. doi: 10.1038/nrgastro.2016.85. [DOI] [PubMed] [Google Scholar]
- 18.Arendt BM, et al. Altered hepatic gene expression in nonalcoholic fatty liver disease is associated with lower hepatic n-3 and n-6 polyunsaturated fatty acids. Hepatology. 2015;61:1565–1578. doi: 10.1002/hep.27695. [DOI] [PubMed] [Google Scholar]
- 19.Da Silva HE, et al. A cross-sectional study assessing dietary intake and physical activity in Canadian patients with nonalcoholic fatty liver disease vs healthy controls. Journal of the Academy of Nutrition and Dietetics. 2014;114:1181–1194. doi: 10.1016/j.jand.2014.01.009. [DOI] [PubMed] [Google Scholar]
- 20.Pan XR, et al. Effects of diet and exercise in preventing NIDDM in people with impaired glucose tolerance. The Da Qing IGT and Diabetes Study. Diabetes Care. 1997;20:537–544. doi: 10.2337/diacare.20.4.537. [DOI] [PubMed] [Google Scholar]
- 21.Brunt EM, Janney CG, Di Bisceglie AM, Neuschwander-Tetri BA, Bacon BR. Nonalcoholic steatohepatitis: a proposal for grading and staging the histological lesions. Am J Gastroenterol. 1999;94:2467–2474. doi: 10.1111/j.1572-0241.1999.01377.x. [DOI] [PubMed] [Google Scholar]
- 22.Kleiner DE, et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology. 2005;41:1313–1321. doi: 10.1002/hep.20701. [DOI] [PubMed] [Google Scholar]
- 23.Matthews DR, et al. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia. 1985;28:412–419. doi: 10.1007/BF00280883. [DOI] [PubMed] [Google Scholar]
- 24.Psychogios N, et al. The human serum metabolome. PLoS One. 2011;6:e16957. doi: 10.1371/journal.pone.0016957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Caporaso JG, et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. Isme j. 2012;6:1621–1624. doi: 10.1038/ismej.2012.8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gloor GB, et al. Microbiome Profiling by Illumina Sequencing of Combinatorial Sequence-Tagged PCR Products. PLoS One. 2010;5:e15406. doi: 10.1371/journal.pone.0015406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD. PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics. 2012;13:31. doi: 10.1186/1471-2105-13-31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–2461. doi: 10.1093/bioinformatics/btq461. [DOI] [PubMed] [Google Scholar]
- 29.Quast C, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–596. doi: 10.1093/nar/gks1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Schloss PD, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009;75:7537–7541. doi: 10.1128/AEM.01541-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Le Gall G, et al. Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011;10:4208–4218. doi: 10.1021/pr2003598. [DOI] [PubMed] [Google Scholar]
- 32.Caporaso JG, et al. QIIME allows analysis of high-throughput community sequencing data. Nature methods. 2010;7:335–336. doi: 10.1038/nmeth.f.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lozupone CA, Knight R. Species divergence and the measurement of microbial diversity. FEMS Microbiol. Rev. 2008;32:557–578. doi: 10.1111/j.1574-6976.2008.00111.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bullard JH, Purdom E, Hansen KD, Dudoit S. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics. 2010;11:94. doi: 10.1186/1471-2105-11-94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Paulson JN, Stine OC, Bravo HC, Pop M. Differential abundance analysis for microbial marker-gene surveys. Nature methods. 2013;10:1200–1202. doi: 10.1038/nmeth.2658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.White JR, Nagarajan N, Pop M. Statistical methods for detecting differentially abundant features in clinical metagenomic samples. PLoS Comput Biol. 2009;5:e1000352. doi: 10.1371/journal.pcbi.1000352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B. 1995;57:289–300. [Google Scholar]
- 38.Macfarlane S, Macfarlane GT. Regulation of short-chain fatty acid production. Proc Nutr Soc. 2003;62:67–72. doi: 10.1079/PNS2002207. [DOI] [PubMed] [Google Scholar]
- 39.LaBrecque DR, et al. World Gastroenterology Organisation global guidelines: Nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. J Clin Gastroenterol. 2014;48:467–473. doi: 10.1097/MCG.0000000000000116. [DOI] [PubMed] [Google Scholar]
- 40.Lakshminarayanan B, Stanton C, O’Toole PW, Ross RP. Compositional dynamics of the human intestinal microbiota with aging: implications for health. J Nutr Health Aging. 2014;18:773–786. doi: 10.1007/s12603-014-0549-6. [DOI] [PubMed] [Google Scholar]
- 41.Yatsunenko T, et al. Human gut microbiome viewed across age and geography. Nature. 2012;486:222–227. doi: 10.1038/nature11053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Odamaki T, et al. Age-related changes in gut microbiota composition from newborn to centenarian: a cross-sectional study. BMC microbiology. 2016;16:90. doi: 10.1186/s12866-016-0708-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Buford TW. (Dis) Trust your gut: the gut microbiome in age-related inflammation, health, and disease. Microbiome. 2017;5:80. doi: 10.1186/s40168-017-0296-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Table 105-0501 and Catalogue no. 82–221-X. (Statistics Canada, 2016).
- 45.Sokol H, et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA. 2008;105:16731–16736. doi: 10.1073/pnas.0804812105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Haro C, et al. The gut microbial community in metabolic syndrome patients is modified by diet. J Nutr Biochem. 2016;27:27–31. doi: 10.1016/j.jnutbio.2015.08.011. [DOI] [PubMed] [Google Scholar]
- 47.Shaw, K. A. et al. Dysbiosis, inflammation, and response to treatment: a longitudinal study of pediatric subjects with newly diagnosed inflammatory bowel disease. 8, 75, 10.1186/s13073-016-0331-y (2016). [DOI] [PMC free article] [PubMed]
- 48.Scheppach W, Weiler F. The butyrate story: old wine in new bottles? Curr Opin Clin Nutr Metab Care. 2004;7:563–567. doi: 10.1097/00075197-200409000-00009. [DOI] [PubMed] [Google Scholar]
- 49.Sartor RB. Key questions to guide a better understanding of host-commensal microbiota interactions in intestinal inflammation. Mucosal immunology. 2011;4:127–132. doi: 10.1038/mi.2010.87. [DOI] [PubMed] [Google Scholar]
- 50.Crost EH, et al. Utilisation of mucin glycans by the human gut symbiont Ruminococcus gnavus is strain-dependent. PLoS One. 2013;8:e76341. doi: 10.1371/journal.pone.0076341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hynonen U, et al. Isolation and whole genome sequencing of a Ruminococcus-like bacterium, associated with irritable bowel syndrome. Anaerobe. 2016;39:60–67. doi: 10.1016/j.anaerobe.2016.03.001. [DOI] [PubMed] [Google Scholar]
- 52.Schwiertz, A. et al. Microbiota and SCFA in lean and overweight healthy subjects. Obesity (Silver Spring)18, 190–195, 10.1038/oby.2009.167. [DOI] [PubMed]
- 53.Jumpertz R, et al. Energy-balance studies reveal associations between gut microbes, caloric load, and nutrient absorption in humans. Am J Clin Nutr. 2011;94:58–65. doi: 10.3945/ajcn.110.010132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Mushref MA, Srinivasan S. Effect of high fat-diet and obesity on gastrointestinal motility. Annals of translational medicine. 2013;1:14. doi: 10.3978/j.issn.2305-5839.2012.11.01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Vogt JA, Wolever TM. Fecal acetate is inversely related to acetate absorption from the human rectum and distal colon. J Nutr. 2003;133:3145–3148. doi: 10.1093/jn/133.10.3145. [DOI] [PubMed] [Google Scholar]
- 56.2-Hydroxybutyric acid.
- 57.Doelle, H. W. Bacterial Metabolism. Second edn, 416–419 (Academic Press, Inc., 1975).
- 58.Ferrannini E, et al. Early metabolic markers of the development of dysglycemia and type 2 diabetes and their physiological significance. Diabetes. 2013;62:1730–1737. doi: 10.2337/db12-0707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Su D, et al. Vitamin D Signaling through Induction of Paneth Cell Defensins Maintains Gut Microbiota and Improves Metabolic Disorders and Hepatic Steatosis in Animal Models. Frontiers in physiology. 2016;7:498. doi: 10.3389/fphys.2016.00498. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated during and analysed during the current study are not publicly available due to privacy restrictions but some blinded data may be made available from the corresponding author on reasonable request.