Abstract
Cancer is a major public health problem as one of the leading causes of death worldwide. Deciphering the molecular regulation mechanisms of tumor progression can make way for tumor diagnosis and therapy. Paternally expressed gene 10 (PEG10), located on human chromosome 7q21.3, has turned out to be an oncogene implicated in the proliferation, apoptosis and metastasis of tumors. PEG10 has been found to be positively expressed in a variety of cancers with seemingly complex expression regulation mechanisms. In this review, we focus on the most vital factors influencing PEG10 expression and recapitulate some of the currently known and potential mechanisms of PEG10 affecting tumor progression, as understanding the molecular regulatory mechanisms of tumor progression can provide potential PEG10 related diagnosis and biomarker specific targeted therapies.
Keywords: PEG10, Regulatory mechanisms, Proliferation, Apoptosis, Metastasis, Targeted therapies
Background
Paternally expressed gene 10 (PEG10), shows 61.4% homology with murine myelin expression factor 3 (MyEF-3), which encodes a distinctive protein functioning as a transcriptional factor during brain development. Conservation of CCHC-type zinc finger motif suggests that PEG10 may also function as a transcriptional factor [1–3]. PEG10 gene is highly conserved in eutherian mammals which indicates its essential functions, and its protein shows high similarity to retroviral gag-pol proteins [4, 5]. Researches on mice suggest that peg10 plays an important role in placenta formation and adipocyte differentiation, and its knockout can cause embryonic lethality [5, 6]. PEG10 is strongly expressed in placenta, ovary and testis as well as somatic adult organs including brain, kidney and lung [1, 7–9]. However, dysregulated expression of PEG10 has been closely associated with the cell proliferation, apoptosis and development of malignancies. However, the comprehensive mechanism of the regulation of PEG10 expression is still in its infancy.
This review will be an update on the current state of PEG10 in cancer related researches. We try to gain insights into the factors mostly influencing PEG10 expression and the profound mechanisms of PEG10 in tumor progression and potential PEG10 related therapeutic targets.
The structure of PEG10 gene
PEG10 is derived from the Ty3/Gypsy retrotransposon family which is located on human chromosome 7q21.3 in a head-to-head orientation with another paternally expressed gene SGCE. There is an 800 bp CpG (cytosine-phosphate-guanine) island between SGCE and PEG10. Li et al. [10] found epigenetic silencing of PEG10 by promoter methylation led to the low expression of PEG10 mRNA, but the activation of PEG10 was not necessarily associated with hypomethylation. Suzuki et al. [11] analyzed the CpG island methylation status and found that the methylation started about 60 bp downstream from the transcription start site of PEG10, suggesting that it is the methylation of downstream regulatory elements rather the promoter methylation that inhibits maternal transcription.
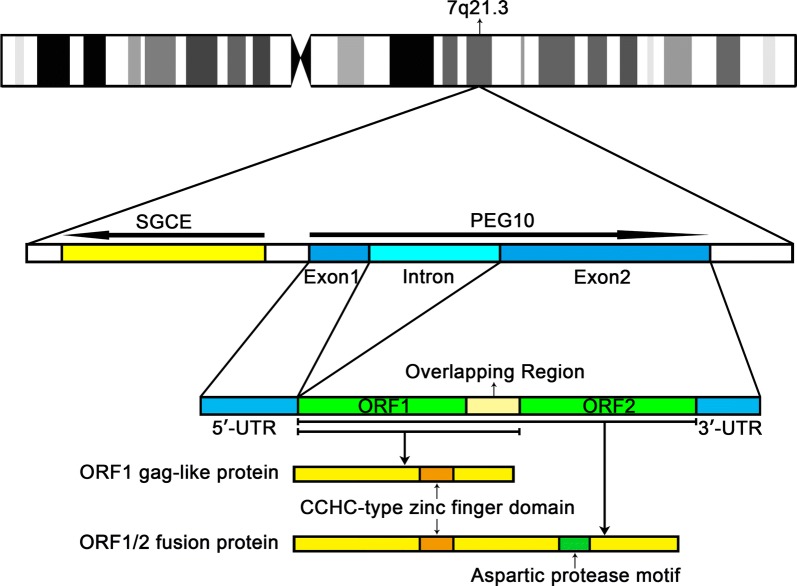
PEG10 gene consists of two exons, separated by a 6.8 kb intron, and its major transcript is 6639 bp (NM_001172438.2) [12]. The exon 1 of PEG10 contains the 5′-untranslated region (UTR) and exon 2 contains two overlapping open reading frames (ORFs) and a 4 kb 3′-UTR sequence [13]. The ORF1 codes for the gag-like PEG10-RF1 protein with a coiled-coil domain in N-terminal and a zinc finger domain in C-terminal, while the pol-like PEG10-RF2 protein is synthesized by a programmed-1 frameshift translation. During the programmed − 1 frameshifting, the ORF1 and the ORF2 translated a gag-pol-like fusion protein named PEG10-RF1/2 from one mRNA (Fig. 1) [4, 14]. Evidence suggests that the translation of PEG10 is also initiated at CUG codon except for the conventional AUG codon [12]. This new finding will add novel cognition to the PEG10’s − 1 frameshifting translation mechanism.
Fig. 1.
Genomic information of human PEG10 gene. PEG10 gene is located on the q21.3 of chromosome 7. The bold arrows show the orientations of PEG10 and SGCE. Exon 1 transcribes the 5’-UTR and exon 2 transcribes two ORFs and the 3’-UTR. ORF1 encodes a gag-like PEG10-RF1 protein, which contains a CCHC-type zinc finger domain. This component protein regulates cell proliferation and apoptosis as a main functional protein. By -1 frameshift translation, the ORF1 and the ORF2 translated PEG10-RF1/2 fusion protein with an aspartic protease motif
The expression levels of PEG10 in cancers
The expression levels of PEG10 in normal and cancer tissues
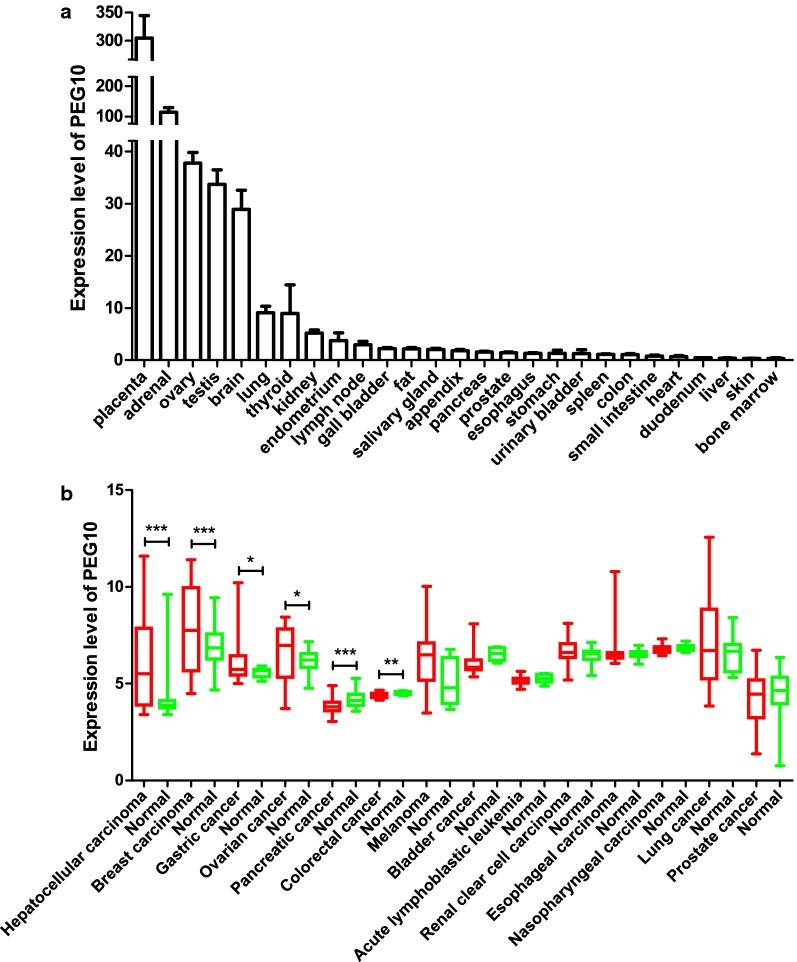
We searched NCBI Gene database to access the expression levels of PEG10 in normal tissues [15]. As shown in Fig. 2a, PEG10 is relatively highly expressed in placenta, adrenal, ovary, testis and brain, but the expression levels are pretty low in other tissues, which are consistent with the previous literature [1, 7–9]. As shown in Table 1, several studies have reported PEG10 is positively expressed in a variety of cancers such as hepatocellular carcinoma (HCC) [9, 16–19], pancreatic carcinoma [20], breast cancer [10], prostate cancer [10], gallbladder carcinoma [21], thyroid cancer [22], oral squamous cell carcinoma [23], colon cancer [24], enchondromas [25] and B-cell chronic lymphocytic leukemia (B-CLL) [26]. Worth noting was that the amplification of PEG10 gene copy numbers detected in HCC also contributed to PEG10 overexpression [17, 27–29].
Fig. 2.
Differential expression pattern of PEG10 in normal tissues and cancers. a) The expression levels of PEG10 in normal tissues. PEG10 is relatively highly expressed in placenta, adrenal, ovary, testis and brain, but shows pretty low expression levels in other normal tissues. b) The expression levels of PEG10 in cancers. PEG10 is overexpressed in HCC and breast cancer, but downregulated in pancreatic carcinoma and colorectal cancer. *P < 0.05, **P < 0.01, ***P < 0.001 by Student’s t test
Table 1.
The expression of PEG10 in cancer patients and cancer cell lines
| Tumors | Type | Methods | Expression | References |
|---|---|---|---|---|
| HCC | Primary HCC | IHC | 15/16a | [9] |
| HCC tissues and cell lines | cDNA microarray | Significantly higher | [16] | |
| HCC cell lines | QPCR | 18/20b | [17] | |
| Primary HCC | QPCR | Significantly higher | ||
| HCC tissues | IHC | 148/218a | [19] | |
| Pancreatic carcinoma | Pancreatic carcinoma tissues | IHC | 85/160b | [20] |
| Breast cancer | Ductal carcinoma | IHC | 6/11a | [10] |
| Invasive ductal carcinomas | IHC | 7/22a | ||
| Breast cancer tissue (n = 161) | IHC | 36%a | ||
| Prostate cancer | Prostate cancer tissue (n = 30) | IHC | 37%a | [10] |
| Gallbladder carcinoma | Gallbladder adenocarcinoma tissues | IHC | 52/108a | [21] |
| Thyroid cancer | 24 thyroid cancer tissues and 14 normal thyroid tissues | QPCR | Significantly higher (P = 0.034) | [22] |
| OSCC | OSCC tissues | QPCR | 83/118b | [23] |
| Colon cancer | Colon cancer tissues | QPCR and WB | 9/20a | [24] |
| Enchondromas | Enchondromas tissues | IHC and QPCR | Strongly positive expression | [25] |
| B-CLL | B-CLL PBMNCs | QPCR | 40/42b | [26] |
IHC immunohistochemistry, QPCR quantitative polymerase chain reaction, OSCC oral squamous cell carcinoma, WB western blot, PBMNC peripheral blood mononuclear cell
aProportion of PEG10-positive tumor tissues/cells
bProportion of tumor tissues/cells occurred PEG10 upregulation compared to normal tissues/cells
To further confirm the expression levels of PEG10 in cancers, we used Gene Expression Omnibus (GEO) datasets to analyze [30]. As shown in Table 2 and Fig. 2b, we observed that PEG10 was overexpressed in several cancers especially HCC and breast cancer. However, PEG10 was shown to be downregulated in pancreatic carcinoma and colorectal cancer, which may be contradictory with the previous literatures. These contradictory results may be due to the data we analyzed were at mRNA level, which may not represent the protein level exactly. In addition, although PEG10 was not shown to be overexpressed in some tumors with regard to the result we analyzed, it may be highly expressed in some specific tumor subtypes since the results we showed are overall. For example, Akamatsu et al. [31] reported that PEG10 was upregulated in neuroendocrine tumors such as neuroendocrine prostate cancer and lung cancer but no significant upregulation in other subtypes.
Table 2.
Basic information of the 14 GEO datasets
| Cancer type | Accession number | Number of samples (tumor/normal) | P-value | References |
|---|---|---|---|---|
| Upregulated | ||||
| Hepatocellular carcinoma | GSE14520 | 233/233 | < 0.0001 | [57, 58] |
| Breast carcinoma | GSE10780 | 42/143 | 0.0002 | [59] |
| Gastric cancer | GSE13861 | 71/19 | 0.0388 | [60] |
| Ovarian cancer | GSE14407 | 12/12 | 0.0438 | [61] |
| Downregulated | ||||
| Pancreatic cancer | GSE28735 | 45/45 | 0.0003 | [62, 63] |
| Colorectal cancer | GSE32323 | 17/17 | 0.0011 | [64] |
| Insignificant | ||||
| Melanoma | GSE3189 | 45/7 | 0.0848 | [65] |
| Bladder cancer | GSE3167 | 41/9 | 0.1243 | [66] |
| Acute lymphoblastic leukemia | GSE26713 | 117/7 | 0.2168 | [67] |
| Renal clear cell carcinoma | GSE36895 | 23/23 | 0.2445 | [68] |
| Esophageal carcinoma | GSE23400 | 53/53 | 0.2526 | [69, 70] |
| Nasopharyngeal carcinoma | GSE12452 | 31/10 | 0.2710 | [71–73] |
| Lung cancer | GSE30219 | 293/14 | 0.2938 | [74] |
| Prostate cancer | GSE6919 | 58/58 | 0.7478 | [75, 76] |
P-values were acquired through t-test for the comparison of PEG10 expression between cancer and normal
PEG10 overexpression is correlated with poor clinicopathological characteristics
Many studies have shown that the expression of PEG10 is closely related to the prognosis of clinicopathological characteristics. Ge et al. [32] conducted meta-analysis to systematically evaluate the correlations between PEG10 and the clinicopathological characteristics in patients with solid tumors. They found that PEG10 overexpression was associated with the higher risk of solid tumors incidence, lower degree of differentiation, increased lymph node metastasis and advanced TNM stage. Moreover, a high level of PEG10 expression was closely correlated to poor overall survival (OS) and it could be used as an independent prognostic biomarker for patients with solid tumors. Furthermore, Bang et al. [19] indicated that PEG10 protein could be a potential biomarker for predicting early recurrence and recurrence-free survival (RFS) in HCC patients after curative resection, even in those with normal serum α-fetoprotein levels.
Factors regulating the expression levels of PEG10
The expression level of PEG10 is regulated by many factors. Transcription factors like E2F, c-MYC and androgen receptor (AR) have been reported to participate in PEG10 expression regulation. Wang et al. [18] reported that both E2F-1 and -4 could directly bind to the PEG10 promoter, and upregulate its transcription in HCC, which was confirmed by chromatin immunoprecipitation (ChIP) and dual luciferase report assay. E2F-1’s direct upregulation of PEG10 expression via binding to the PEG10 promoter was also verified in prostate and pancreatic cancer [18, 20, 31]. Besides, in lung cancer cells, GSK3β/USP11/E2F-1/PEG10 pathway was shown to play an imperative role in PEG10 overexpression [33]. Li et al. [10] reported that c-MYC knockdown in Panc1 cells resulted in a subsequent PEG10 downregulation, and ChIP assays validated that PEG10 was a direct downstream target of c-MYC. In prostate cancer, AR was confirmed to bind to the PEG10 promoter region, thus repressing the transcription of PEG10. Treating prostate cancer cells with synthetic androgen R1881 resulted in an increased AR occupancy at the PEG10 promoter, while decreased when treated with AR antagonist Enzalutamide [31].
Transforming growth factor β (TGF-β) signaling pathway plays a biphasic role in cancer progression [34]. In HCC, PEG10 was increased after treating HepG2 cells with TGF-β1 [44]. However, the mutual inhibition effect of PEG10 and TGF-β signaling was found in chondrosarcoma and enchondroma [25]. Shinohara et al. proposed that TGF-β might inhibit PEG10 expression through the downregulation of c-MYC [10, 25, 35]. Besides, Wang et al. suggested that TGF-β might also inhibit the expression of PEG10 by keeping Rb dephosphorylated to inhibit the release of E2F [18, 36].
Several miRNAs have also been proven to regulate the expression level of PEG10. In HCC, miR-122 repressed the translation level of PEG10 via directly binding to sites 2310 and 2403 in PEG10 3′-UTR [37, 38]. Additionally, miR-491 has also been confirmed to negatively regulate the expression of PEG10 directly in colorectal cancer [24].
Functions of PEG10 and the mechanisms thereof
PEG10 promotes tumor proliferation
Uncontrolled proliferation is an important factor in tumor progression. Numerous researches have reported that PEG10 plays a significant role in promoting the cancer proliferation. The proliferation ability of cancer cells was improved after overexpressing PEG10 in HCC, while decreasing endogenous expression of PEG10 showed prominent growth retardation [9]. PEG10’s role in proliferation was also confirmed in colorectal cancer cell line HCT-116, as Curcumin was able to diminish the proliferation effect by upregulating the expression of miR-491 [24]. In addition, our data revealed that the proliferation ability of Raji cells and A549 cells was decreased after being transfected with PEG10 siRNA [39, 40], and that PEG10 promoted breast cancer cell proliferation after being overexpressed [41]. The function of PEG10 was also certified in gastric cancer, where knockdown of PEG10 in MKN7 cells reduced anchorage-independent colony formation [42]. Additionally, it has also been reported that GSK3β increases the interaction of E2F1 with USP11, which results in the deubiquitination and stabilization of E2F1, which in turn activates PEG10 expression to promote proliferation in A549 [33]. Furthermore, c-MYC protein binds to E-box sequences in the first PEG10 intron and activates its transcription, which further promotes the proliferation of several tumor cells [10, 33]. In vivo experiments showed that volume and weight of the tumors obtained from xenograft tumorigenicity assays were both lower and the Ki-67 score reduced significantly after PEG10 knockdown [20, 31]. The underlying mechanism of PEG10 promoting the proliferation may be due to its cell-cycling promoting effect. In pancreatic and neuroendocrine prostate cancers, the levels of p21, p27 (cell-cycle-dependent kinase inhibitors) and Cyclin E1 (which accumulates at G0/G1 to S phase and reduced smoothly from S to G2/M phase) were upregulated after PEG10 knockdown, which indicate PEG10 drives cell cycle progression from G0/G1 to S phase [20, 31].
PEG10 inhibits the apoptosis of cancer cells
Apoptosis is a physiological process of programmed cell death, which is indispensable in cell development and homeostasis [43]. Dysfunction in apoptosis pathways is a typical characteristic of cancer cells. As an oncogene, PEG10 also has been shown to play an anti-apoptosis role in cancer progression. In Raji cells and HCT-116 cells, PEG10 was found to inhibit the apoptosis of cancer cells [24, 39], while in HepG2 cells PEG10 was shown to increase Bcl-2 expression and decrease Bax expression, and diminish apoptosis induced by doxorubicin [44]. PEG10 overexpression decreases cell death mediated by SIAH1 in HCC, while SIAH1 also reduces the amount of PEG10 protein, thus inducing growth arrest and apoptosis in hepatoma cells [9, 45]. In addition, mRNA interference of PEG10 in human hepatocyte L02 cells resulted in elevated expression levels of anti-apoptosis protein BCL-xL [18]. Similarly, our early studies have also demonstrated that in B-cell acute lymphoblastic leukemia (B-ALL) and B-cell chronic lymphocytic leukemia (B-CLL), CXCL13 and CCL19 together upregulate PEG10 expression in CD23+CD5+ or CD19+CD34+ B cells and then hamper the activation of caspase-3 and caspase-8 to gain the apoptosis resistance induced by tumor necrosis factor-α (TNF-α) [46, 47].
PEG10 promotes metastasis of cancer cells
Tumor metastasis is an important factor in promoting poor prognosis in cancer patients and is mainly characterized by the migration, invasion and epithelial–mesenchymal transition (EMT) of cancer cells. It has been shown that PEG10 is overexpressed in metastatic prostate cancer and rectal adenocarcinoma versus primary and benign tumors, thus indicating that PEG10 may be involved in cancer metastasis [31, 48]. Our studies revealed that suppressing the expression of PEG10 in human Raji cells and A549 cells resulted in the reduction of migration and invasion capabilities of cells, and the decrease of matrix metalloproteinases (MMPs) like MMP-2 and -9 [39, 40]. In addition, we also observed that overexpression of PEG10 promoted the migration and invasion of breast cancer cell line MDA-MB-231 cells, in which the expression levels of MMP-1, -2 and -9 were increased while that of TIMP-1 and -2 decreased [41]. Besides, PEG10 also turned out to promote pancreatic cancer cells migration and invasion through ERK/MMP7 pathway [20]. In HCC, overexpression of PEG10 in HepG2 cells decreased the expression levels of epithelial marker protein E-cadherin and increased the expression levels of mesenchymal marker protein vimentin. PEG10 may be involved in the activation of canonical TGF-β pathway to promote the EMT of cancers. Although PEG10 could inhibit TGF-β receptors to block TGF-β pathway and may diminish the inhibition of proliferation induced by TGF-β signaling, the invasion and EMT induced by TGF-β could be removed after PEG10 knockdown [44]. In prostate cancer cells, after TGF-β treatment, PEG10 knockdown decreased Smad2 and Smad3 phosphorylation, SBE-4 (which contains four copies of Smad binding elements) luciferase reporter activity and decreased the expression of mesenchymal transcription factor Snail1 and Zeb1 (which are the direct mediators of the TGF-β pathway) [31]. A series of clinical investigations revealed that HCC [19], gallbladder cancer [21, 49], lung cancer [40], oral squamous cell carcinoma [23], pancreatic cancer [20] and gastric cancer [42] are more vulnerable to metastasis or invasion with PEG10 overexpression. A diagram of the expression regulation factors of PEG10 and the underlying mechanisms of the oncogenic role of PEG10 in cancer progression is shown in Fig. 3.
Fig. 3.
Schematic representation of PEG10 expression regulation mechanisms and its role in tumor progression. miRNAs indicate miR-122 and miR-491; Caspases indicate caspase 3 and caspase 8; MMPs indicate MMP-1, -2 and -9; TIMPs indicate TIMP-1 and -2. The solid arrows indicate that these signaling pathways have been confirmed; the dashed arrows show that these signaling pathways need to be further validated
Therapy
Immunotherapy of HCC
Recent years have witnessed the rapid development of carcinoma immunotherapy. Dendritic cells (DCs) are powerful professional antigen presenting cells which were discovered in 1973 and have been used in anti-tumor immunotherapy since the 1980s. In 2010, the first therapeutic tumor DCs vaccine sipuleucel-T for the treatment of metastatic prostate cancer was approved by the US Food and Drug Administration (FDA) based on its confirmed safe and non-toxic side effects [50]. After being transfected with recombinant adenovirus (Ad) vectors encoding tumor associated antigen (TAA), DCs are able to process the TAA to peptides and then bind them to MHC class I molecules for recognition by CD8+ T cells. Besides, DCs express costimulators to provide the signals needed for differentiation of CD8+ T cells into anti-tumor specific cytotoxic T lymphocytes (CTLs) which are able to recognize and kill tumor cells without a requirement for costimulation [50–53].
Numerous evidence demonstrated that PEG10 was overexpressed in HCC and contributed to the oncogenesis, thus it might be a TAA of HCC. Peng et al. [54] transfected DCs with PEG10 recombinant Ad and found the DCs could specifically elicit CTLs to secrete interferon-γ (IFN-γ) and lyse HepG2. This Ad-PEG10 transduced DCs could induce an anti-tumor immune response against PEG10 positive HCC with HLA-A2 restricted both in vitro and in vivo. The findings indicate that DCs transfected with Ad-PEG10 might be an ideal target for HCC immunotherapy.
Others
In recent years, there have been growing interest in RNA-targeted therapies [55]. As described above, PEG10 may be a target for intervention in cancer. Using antisense oligonucleotides or siRNA to interfering PEG10 mRNA for treatment can be taken into consideration. Besides, to further explore the mechanisms of PEG10 promoting tumor progression and find it is which domain of PEG10 protein that actually works, may pave the path for the design of the small molecular inhibitors which may also be an effective strategy for PEG10 inactivation. In addition, Kempinska et al. [56] demonstrated that the menin–MLL1 complex binds to PEG10 gene directly and catalyzes H3K4me3 to upregulate the expression of PEG10 through the epigenetic mechanism. The menin–MLL inhibitor MI-503 showed an anti-tumor effect of HCC, and the underlying mechanism may be owing to the indirect downregulation of PEG10. Therefore, exploring the agents that can inhibit the function of PEG10 indirectly may also be a potential approach for tumor therapy.
Conclusions
PEG10 is an imprinting gene that plays a key role in tumor proliferation, apoptosis and metastasis. The studies of PEG10 in tumors mainly focused on HCC, but in recent years extensive studies have shown that PEG10 also contributes to the progression of many types of cancer. The expression level of PEG10 is regulated by several factors, but their effects may vary in different tumors. Since PEG10 is highly expressed in tumors, it may serve as a TAA which can be utilized in tumor immunotherapy or a potential target for new anti-cancer regimens and cancer diagnosis. Nevertheless, further studies are needed to gain insights into the molecular mechanisms of the role PEG10 played in cancer.
Authors’ contributions
TX and SP performed the selection of literature and drafted the manuscript. HZ, ZL, ZY and YY collected the related references. JX, QY, MW and WY prepared the figures. KMT and MJ revised the language. QZ and JX carried out the design of the review. All authors contributed to this manuscript. All authors read and approved the final manuscript.
Acknowledgements
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The datasets analyzed during the current study are available in the NCBI Gene database (https://www.ncbi.nlm.nih.gov/gene/) and GEO database (https://www.ncbi.nlm.nih.gov/geo/).
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81770180).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- PEG10
paternally expressed gene 10
- MyEF-3
murine myelin expression factor 3
- CpG
cytosine-phosphate-guanine
- UTR
untranslated region
- ORF
open reading frame
- HCC
hepatocellular carcinoma
- B-CLL
B-cell chronic lymphocytic leukemia
- GEO
Gene Expression Omnibus
- OS
overall survival
- RFS
recurrence-free survival
- AR
androgen receptor
- ChIP
chromatin immunoprecipitation
- GSK3β
glycogen synthase kinase-3β
- USP11
ubiquitin-specific protease 11
- TGF-β
transforming growth factor β
- B-ALL
B-cell acute lymphoblastic leukemia
- TNF-α
tumor necrosis factor-α
- EMT
epithelial–mesenchymal transition
- MMPs
matrix metalloproteinases
- TIMPs
tissue inhibitor of metalloproteinases
- DCs
dendritic cells
- Ad
adenovirus
- TAA
tumor associated antigen
- CTLs
cytotoxic T lymphocytes
- IFN-γ
interferon-γ
- MLL1
mixed lineage leukemia 1
- H3K4me3
trimethylation of H3K4
Contributor Information
Tian Xie, Email: 2016302160014@whu.edu.cn.
Shan Pan, Email: panshan@whu.edu.cn.
Hang Zheng, Email: wangxqzh@163.com.
Zilv Luo, Email: 2015302180093@whu.edu.cn.
Kingsley M. Tembo, Email: kintembo@yahoo.co.uk
Muhammad Jamal, Email: jamalbiotech@yahoo.com.
Zhongyang Yu, Email: 2015302180105@whu.edu.cn.
Yao Yu, Email: 2014302180057@whu.edu.cn.
Jing Xia, Email: 2014302180066@whu.edu.cn.
Qian Yin, Email: yinqian@whu.edu.cn.
Meng Wang, Email: wangmengsy@whu.edu.cn.
Wen Yuan, Email: 2015203010044@whu.edu.cn.
Qiuping Zhang, Email: qpzhang@whu.edu.cn.
Jie Xiong, Email: jiexiong@whu.edu.cn.
References
- 1.Ono R, Kobayashi S, Wagatsuma H, Aisaka K, Kohda T, Kaneko-Ishino T, Ishino F. A retrotransposon-derived gene, PEG10, is a novel imprinted gene located on human chromosome 7q21. Genomics. 2001;73:232–237. doi: 10.1006/geno.2001.6494. [DOI] [PubMed] [Google Scholar]
- 2.Youngson NA, Kocialkowski S, Peel N, Ferguson-Smith AC. A small family of sushi-class retrotransposon-derived genes in mammals and their relation to genomic imprinting. J Mol Evol. 2005;61:481–490. doi: 10.1007/s00239-004-0332-0. [DOI] [PubMed] [Google Scholar]
- 3.Steplewski A, Krynska B, Tretiakova A, Haas S, Khalili K, Amini S. MyEF-3, a developmentally controlled brain-derived nuclear protein which specifically interacts with myelin basic protein proximal regulatory sequences. Biochem Biophys Res Commun. 1998;243:295–301. doi: 10.1006/bbrc.1997.7821. [DOI] [PubMed] [Google Scholar]
- 4.Clark MB, Janicke M, Gottesbuhren U, Kleffmann T, Legge M, Poole ES, Tate WP. Mammalian gene PEG10 expresses two reading frames by high efficiency − 1 frameshifting in embryonic-associated tissues. J Biol Chem. 2007;282:37359–37369. doi: 10.1074/jbc.M705676200. [DOI] [PubMed] [Google Scholar]
- 5.Ono R, Nakamura K, Inoue K, Naruse M, Usami T, Wakisaka-Saito N, Hino T, Suzuki-Migishima R, Ogonuki N, Miki H, et al. Deletion of Peg10, an imprinted gene acquired from a retrotransposon, causes early embryonic lethality. Nat Genet. 2006;38:101–106. doi: 10.1038/ng1699. [DOI] [PubMed] [Google Scholar]
- 6.Hishida T, Naito K, Osada S, Nishizuka M, Imagawa M. peg10, an imprinted gene, plays a crucial role in adipocyte differentiation. FEBS Lett. 2007;581:4272–4278. doi: 10.1016/j.febslet.2007.07.074. [DOI] [PubMed] [Google Scholar]
- 7.Ono R, Shiura H, Aburatani H, Kohda T, Kaneko-Ishino T, Ishino F. Identification of a large novel imprinted gene cluster on mouse proximal chromosome 6. Genome Res. 2003;13:1696–1705. doi: 10.1101/gr.906803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Smallwood A, Papageorghiou A, Nicolaides K, Alley MK, Jim A, Nargund G, Ojha K, Campbell S, Banerjee S. Temporal regulation of the expression of syncytin (HERV-W), maternally imprinted PEG10, and SGCE in human placenta. Biol Reprod. 2003;69:286–293. doi: 10.1095/biolreprod.102.013078. [DOI] [PubMed] [Google Scholar]
- 9.Okabe H, Satoh S, Furukawa Y, Kato T, Hasegawa S, Nakajima Y, Yamaoka Y, Nakamura Y. Involvement of PEG10 in human hepatocellular carcinogenesis through interaction with SIAH1. Cancer Res. 2003;63:3043–3048. [PubMed] [Google Scholar]
- 10.Li CM, Margolin AA, Salas M, Memeo L, Mansukhani M, Hibshoosh H, Szabolcs M, Klinakis A, Tycko B. PEG10 is a c-MYC target gene in cancer cells. Cancer Res. 2006;66:665–672. doi: 10.1158/0008-5472.CAN-05-1553. [DOI] [PubMed] [Google Scholar]
- 11.Suzuki S, Ono R, Narita T, Pask AJ, Shaw G, Wang C, Kohda T, Alsop AE, Marshall Graves JA, Kohara Y, et al. Retrotransposon silencing by DNA methylation can drive mammalian genomic imprinting. PLoS Genet. 2007;3:e55. doi: 10.1371/journal.pgen.0030055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lux H, Flammann H, Hafner M, Lux A. Genetic and molecular analyses of PEG10 reveal new aspects of genomic organization, transcription and translation. PLoS ONE. 2010;5:e8686. doi: 10.1371/journal.pone.0008686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Volff J, Korting C, Schartl M. Ty3/Gypsy retrotransposon fossils in mammalian genomes: did they evolve into new cellular functions? Mol Biol Evol. 2001;18:266–270. doi: 10.1093/oxfordjournals.molbev.a003801. [DOI] [PubMed] [Google Scholar]
- 14.Shigemoto K, Brennan J, Walls E, Watson CJ, Stott D, Rigby PWJ, Reith AD. Identification and characterisation of a developmentally regulated mammalian gene that utilises-1 programmed ribosomal frameshifting. Nucleic Acids Res. 2001;29:4079–4088. doi: 10.1093/nar/29.19.4079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.NCBI Gene database. https://www.ncbi.nlm.nih.gov/gene/23089/?report=expression. Accessed 10 Dec 2017.
- 16.Tsou AP, Chuang YC, Su JY, Yang CW, Liao YL, Liu WK, Chiu JH, Chou CK. Overexpression of a novel imprinted gene, PEG10, in human hepatocellular carcinoma and in regenerating mouse livers. J Biomed Sci. 2003;10:625–635. doi: 10.1159/000073528. [DOI] [PubMed] [Google Scholar]
- 17.Ip WK, Lai PB, Wong NL, Sy SM, Beheshti B, Squire JA, Wong N. Identification of PEG10 as a progression related biomarker for hepatocellular carcinoma. Cancer Lett. 2007;250:284–291. doi: 10.1016/j.canlet.2006.10.012. [DOI] [PubMed] [Google Scholar]
- 18.Wang C, Xiao Y, Hu Z, Chen Y, Liu N, Hu G. PEG10 directly regulated by E2Fs might have a role in the development of hepatocellular carcinoma. FEBS Lett. 2008;582:2793–2798. doi: 10.1016/j.febslet.2008.07.009. [DOI] [PubMed] [Google Scholar]
- 19.Bang H, Ha SY, Hwang SH, Park CK. Expression of PEG10 is associated with poor survival and tumor recurrence in hepatocellular carcinoma. Cancer Res Treat. 2015;47:844–852. doi: 10.4143/crt.2014.124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Peng YP, Zhu Y, Yin LD, Zhang JJ, Wei JS, Liu X, Liu XC, Gao WT, Jiang KR, Miao Y. PEG10 overexpression induced by E2F-1 promotes cell proliferation, migration, and invasion in pancreatic cancer. J Exp Clin Cancer Res. 2017;36:30. doi: 10.1186/s13046-017-0500-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Liu DC, Yang ZL, Jiang S. Identification of PEG10 and TSG101 as carcinogenesis, progression, and poor-prognosis related biomarkers for gallbladder adenocarcinoma. Pathol Oncol Res. 2011;17:859–866. doi: 10.1007/s12253-011-9394-7. [DOI] [PubMed] [Google Scholar]
- 22.Boot A, Oosting J, de Miranda NF, Zhang Y, Corver WE, van de Water B, Morreau H, van Wezel T. Imprinted survival genes preclude loss of heterozygosity of chromosome 7 in cancer cells. J Pathol. 2016;240:72–83. doi: 10.1002/path.4756. [DOI] [PubMed] [Google Scholar]
- 23.Singh SS, Kumar R, Kushwaha VS, Bhatt M, Singh A, Mishra A, Ram H, Parmar D, Gupta R. Expression of radioresistant gene PEG10 in OSCC patients and its prognostic significance. Asian Pac J Cancer Prev. 2017;18:1513–1518. doi: 10.22034/APJCP.2017.18.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Li B, Shi C, Li B, Zhao JM, Wang L. The effects of Curcumin on HCT-116 cells proliferation and apoptosis via the miR-491/PEG10 pathway. J Cell Biochem. 2018;119:3091–3098. doi: 10.1002/jcb.26449. [DOI] [PubMed] [Google Scholar]
- 25.Shinohara N, Maeda S, Yahiro Y, Sakuma D, Matsuyama K, Imamura K, Kawamura I, Setoguchi T, Ishidou Y, Nagano S, Komiya S. TGF-beta signalling and PEG10 are mutually exclusive and inhibitory in chondrosarcoma cells. Sci Rep. 2017;7:13494. doi: 10.1038/s41598-017-13994-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kainz B, Shehata M, Bilban M, Kienle D, Heintel D, Kromer-Holzinger E, Le T, Krober A, Heller G, Schwarzinger I, et al. Overexpression of the paternally expressed gene 10 (PEG10) from the imprinted locus on chromosome 7q21 in high-risk B-cell chronic lymphocytic leukemia. Int J Cancer. 2007;121:1984–1993. doi: 10.1002/ijc.22929. [DOI] [PubMed] [Google Scholar]
- 27.Tsuji K, Yasui K, Gen Y, Endo M, Dohi O, Zen K, Mitsuyoshi H, Minami M, Itoh Y, Taniwaki M, et al. PEG10 is a probable target for the amplification at 7q21 detected in hepatocellular carcinoma. Cancer Genet Cytogenet. 2010;198:118–125. doi: 10.1016/j.cancergencyto.2010.01.004. [DOI] [PubMed] [Google Scholar]
- 28.Huang J, Sheng HH, Shen T, Hu YJ, Xiao HS, Zhang Q, Zhang QH, Han ZG. Correlation between genomic DNA copy number alterations and transcriptional expression in hepatitis B virus-associated hepatocellular carcinoma. FEBS Lett. 2006;580:3571–3581. doi: 10.1016/j.febslet.2006.05.032. [DOI] [PubMed] [Google Scholar]
- 29.Dong H, Zhang H, Liang J, Yan H, Chen Y, Shen Y, Kong Y, Wang S, Zhao G, Jin W. Digital karyotyping reveals probable target genes at 7q21.3 locus in hepatocellular carcinoma. BMC Med Genomics. 2011;4:60. doi: 10.1186/1755-8794-4-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.NCBI GEO database. https://www.ncbi.nlm.nih.gov/geo/. Accessed 8 Dec 2017.
- 31.Akamatsu S, Wyatt AW, Lin D, Lysakowski S, Zhang F, Kim S, Tse C, Wang K, Mo F, Haegert A, et al. The placental gene PEG10 promotes progression of neuroendocrine prostate cancer. Cell Rep. 2015;12:922–936. doi: 10.1016/j.celrep.2015.07.012. [DOI] [PubMed] [Google Scholar]
- 32.Ge H, Yan Y, Wu D, Huang Y, Tian F. Prognostic value of PEG10 in Asian solid tumors: a meta-analysis. Clin Chim Acta. 2018;483:197–203. doi: 10.1016/j.cca.2018.04.041. [DOI] [PubMed] [Google Scholar]
- 33.Wang D, Zhao J, Li S, Wei J, Nan L, Mallampalli RK, Weathington NM, Ma H, Zhao Y. Phosphorylated E2F1 is stabilized by nuclear USP11 to drive Peg10 gene expression and activate lung epithelial cells. J Mol Cell Biol. 2018;10:60–73. doi: 10.1093/jmcb/mjx034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Massague J. TGFbeta in Cancer. Cell. 2008;134:215–230. doi: 10.1016/j.cell.2008.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yagi K, Furuhashi M, Aoki H, Goto D, Kuwano H, Sugamura K, Miyazono K, Kato M. c-myc is a downstream target of the Smad pathway. J Biol Chem. 2002;277:854–861. doi: 10.1074/jbc.M104170200. [DOI] [PubMed] [Google Scholar]
- 36.Li JM, Hu PPC, Shen X, Yu Y, Wang XF. E2F4-RB and E2F4-p107 complexes suppress gene expression by transforming growth factor beta through E2F binding sites. Proc Natl Acad Sci USA. 1997;94:4948–4953. doi: 10.1073/pnas.94.10.4948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Shyu YC, Lee TL, Lu MJ, Chen JR, Chien RN, Chen HY, Lin JF, Tsou AP, Chen YH, Hsieh CW, Huang TS. miR-122-mediated translational repression of PEG10 and its suppression in human hepatocellular carcinoma. J Transl Med. 2016;14:200. doi: 10.1186/s12967-016-0956-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Li J, Huang L, Xiao X, Chen Y, Wang X, Zhou Z, Zhang C, Zhang Y. Photoclickable MicroRNA for the intracellular target identification of microRNAs. J Am Chem Soc. 2016;138:15943–15949. doi: 10.1021/jacs.6b08521. [DOI] [PubMed] [Google Scholar]
- 39.Xiong J, Qin J, Zheng Y, Peng X, Luo Y, Meng X. PEG10 promotes the migration of human Burkitt’s lymphoma cells by up-regulating the expression of matrix metalloproteinase-2 and -9. Clin Invest Med. 2012;35:E117–E125. doi: 10.25011/cim.v35i3.16587. [DOI] [PubMed] [Google Scholar]
- 40.Deng X, Hu Y, Ding Q, Han R, Guo Q, Qin J, Li J, Xiao R, Tian S, Hu W, et al. PEG10 plays a crucial role in human lung cancer proliferation, progression, prognosis and metastasis. Oncol Rep. 2014;32:2159–2167. doi: 10.3892/or.2014.3469. [DOI] [PubMed] [Google Scholar]
- 41.Li X, Xiao R, Tembo K, Hao L, Xiong M, Pan S, Yang X, Yuan W, Xiong J, Zhang Q. PEG10 promotes human breast cancer cell proliferation, migration and invasion. Int J Oncol. 2016;48:1933–1942. doi: 10.3892/ijo.2016.3406. [DOI] [PubMed] [Google Scholar]
- 42.Ishii S, Yamashita K, Harada H, Ushiku H, Tanaka T, Nishizawa N, Yokoi K, Washio M, Ema A, Mieno H, et al. The H19-PEG10/IGF2BP3 axis promotes gastric cancer progression in patients with high lymph node ratios. Oncotarget. 2017;8:74567–74581. doi: 10.18632/oncotarget.20209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Khan KH, Blanco-Codesido M, Molife LR. Cancer therapeutics: targeting the apoptotic pathway. Crit Rev Oncol Hematol. 2014;90:200–219. doi: 10.1016/j.critrevonc.2013.12.012. [DOI] [PubMed] [Google Scholar]
- 44.Zhang M, Sui C, Dai B, Shen W, Lu J, Yang J. PEG10 is imperative for TGF-beta1-induced epithelial–mesenchymal transition in hepatocellular carcinoma. Oncol Rep. 2017;37:510–518. doi: 10.3892/or.2016.5282. [DOI] [PubMed] [Google Scholar]
- 45.Yoshibayashi H, Okabe H, Satoh S, Hida K, Kawashima K, Hamasu S, Nomura A, Hasegawa S, Ikai I, Sakai Y. SIAH1 causes growth arrest and apoptosis in hepatoma cells through beta-catenin degradation-dependent and -independent mechanisms. Oncol Rep. 2007;17:549–556. [PubMed] [Google Scholar]
- 46.Hu C, Xiong J, Zhang L, Huang B, Zhang Q, Li Q, Yang M, Wu Y, Wu Q, Shen Q, et al. PEG10 activation by co-stimulation of CXCR5 and CCR7 essentially contributes to resistance to apoptosis in CD19+CD34+ B cells from patients with B cell lineage acute and chronic lymphocytic leukemia. Cell Mol Immunol. 2004;1:280–294. [PubMed] [Google Scholar]
- 47.Chunsong H, Yuling H, Li W, Jie X, Gang Z, Qiuping Z, Qingping G, Kejian Z, Li Q, Chang AE, et al. CXC chemokine ligand 13 and CC chemokine ligand 19 cooperatively render resistance to apoptosis in B cell lineage acute and chronic lymphocytic leukemia CD23+CD5+ B cells. J Immunol. 2006;177:6713–6722. doi: 10.4049/jimmunol.177.10.6713. [DOI] [PubMed] [Google Scholar]
- 48.Hua Y, Ma X, Liu X, Yuan X, Qin H, Zhang X. Identification of the potential biomarkers for the metastasis of rectal adenocarcinoma. APMIS. 2017;125:93–100. doi: 10.1111/apm.12633. [DOI] [PubMed] [Google Scholar]
- 49.Liu Z, Yang Z, Liu D, Li D, Zou Q, Yuan Y, Li J, Liang L, Chen M, Chen S. TSG101 and PEG10 are prognostic markers in squamous cell/adenosquamous carcinomas and adenocarcinoma of the gallbladder. Oncol Lett. 2014;7:1128–1138. doi: 10.3892/ol.2014.1886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Palucka K, Banchereau J. Cancer immunotherapy via dendritic cells. Nat Rev Cancer. 2012;12:265–277. doi: 10.1038/nrc3258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Xia D, Moyana T, Xiang J. Combinational adenovirus-mediated gene therapy and dendritic cell vaccine in combating well-established tumors. Cell Res. 2006;16:241–259. doi: 10.1038/sj.cr.7310032. [DOI] [PubMed] [Google Scholar]
- 52.Melief CJ. Cancer immunotherapy by dendritic cells. Immunity. 2008;29:372–383. doi: 10.1016/j.immuni.2008.08.004. [DOI] [PubMed] [Google Scholar]
- 53.Gardner A, Ruffell B. Dendritic cells and cancer immunity. Trends Immunol. 2016;37:855–865. doi: 10.1016/j.it.2016.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Peng W, Zhao G, Ma Y, Yu H, Wang X. Dendritic cells transfected with PEG10 recombinant adenovirus elicit anti-tumor immune response in vitro and in vivo. Vaccine. 2011;29:3501–3506. doi: 10.1016/j.vaccine.2011.02.027. [DOI] [PubMed] [Google Scholar]
- 55.Crooke ST, Witztum JL, Bennett CF, Baker BF. RNA-targeted therapeutics. Cell Metab. 2018;27:714–739. doi: 10.1016/j.cmet.2018.03.004. [DOI] [PubMed] [Google Scholar]
- 56.Kempinska K, Malik B, Borkin D, Klossowski S, Shukla S, Miao H, Wang J, Cierpicki T, Grembecka J. Pharmacologic inhibition of the Menin–MLL interaction leads to transcriptional repression of PEG10 and blocks hepatocellular carcinoma. Mol Cancer Ther. 2018;17:26–38. doi: 10.1158/1535-7163.MCT-17-0580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Roessler S, Jia HL, Budhu A, Forgues M, Ye QH, Lee JS, Thorgeirsson SS, Sun Z, Tang ZY, Qin LX, Wang XW. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 2010;70:10202–10212. doi: 10.1158/0008-5472.CAN-10-2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Roessler S, Long EL, Budhu A, Chen Y, Zhao X, Ji J, Walker R, Jia HL, Ye QH, Qin LX, et al. Integrative genomic identification of genes on 8p associated with hepatocellular carcinoma progression and patient survival. Gastroenterology. 2012;142(957–966):e912. doi: 10.1053/j.gastro.2011.12.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Chen DT, Nasir A, Culhane A, Venkataramu C, Fulp W, Rubio R, Wang T, Agrawal D, McCarthy SM, Gruidl M, et al. Proliferative genes dominate malignancy-risk gene signature in histologically-normal breast tissue. Breast Cancer Res Treat. 2010;119:335–346. doi: 10.1007/s10549-009-0344-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Cho JY, Lim JY, Cheong JH, Park YY, Yoon SL, Kim SM, Kim SB, Kim H, Hong SW, Park YN, et al. Gene expression signature-based prognostic risk score in gastric cancer. Clin Cancer Res. 2011;17:1850–1857. doi: 10.1158/1078-0432.CCR-10-2180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Bowen NJ, Walker LD, Matyunina LV, Logani S, Totten KA, Benigno BB, McDonald JF. Gene expression profiling supports the hypothesis that human ovarian surface epithelia are multipotent and capable of serving as ovarian cancer initiating cells. BMC Med Genomics. 2009;2:71. doi: 10.1186/1755-8794-2-71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhang G, Schetter A, He P, Funamizu N, Gaedcke J, Ghadimi BM, Ried T, Hassan R, Yfantis HG, Lee DH, et al. DPEP1 inhibits tumor cell invasiveness, enhances chemosensitivity and predicts clinical outcome in pancreatic ductal adenocarcinoma. PLoS ONE. 2012;7:e31507. doi: 10.1371/journal.pone.0031507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zhang G, He P, Tan H, Budhu A, Gaedcke J, Ghadimi BM, Ried T, Yfantis HG, Lee DH, Maitra A, et al. Integration of metabolomics and transcriptomics revealed a fatty acid network exerting growth inhibitory effects in human pancreatic cancer. Clin Cancer Res. 2013;19:4983–4993. doi: 10.1158/1078-0432.CCR-13-0209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Khamas A, Ishikawa T, Shimokawa K, Mogushi K, Iida S, Ishiguro M, Mizushima H, Tanaka H, Uetake H, Sugihara K. Screening for epigenetically masked genes in colorectal cancer using 5-Aza-2′-deoxycytidine, microarray and gene expression profile. Cancer Genomics Proteomics. 2012;9:67–75. [PubMed] [Google Scholar]
- 65.Talantov D, Mazumder A, Yu JX, Briggs T, Jiang Y, Backus J, Atkins D, Wang Y. Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res. 2005;11:7234–7242. doi: 10.1158/1078-0432.CCR-05-0683. [DOI] [PubMed] [Google Scholar]
- 66.Dyrskjot L, Kruhoffer M, Thykjaer T, Marcussen N, Jensen JL, Moller K, Orntoft TF. Gene expression in the urinary bladder: a common carcinoma in situ gene expression signature exists disregarding histopathological classification. Cancer Res. 2004;64:4040–4048. doi: 10.1158/0008-5472.CAN-03-3620. [DOI] [PubMed] [Google Scholar]
- 67.Homminga I, Pieters R, Langerak AW, de Rooi JJ, Stubbs A, Verstegen M, Vuerhard M, Buijs-Gladdines J, Kooi C, Klous P, et al. Integrated transcript and genome analyses reveal NKX2-1 and MEF2C as potential oncogenes in T cell acute lymphoblastic leukemia. Cancer Cell. 2011;19:484–497. doi: 10.1016/j.ccr.2011.02.008. [DOI] [PubMed] [Google Scholar]
- 68.Pena-Llopis S, Vega-Rubin-de-Celis S, Liao A, Leng N, Pavia-Jimenez A, Wang S, Yamasaki T, Zhrebker L, Sivanand S, Spence P, et al. BAP1 loss defines a new class of renal cell carcinoma. Nat Genet. 2012;44:751–759. doi: 10.1038/ng.2323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Su H, Hu N, Yang HH, Wang C, Takikita M, Wang QH, Giffen C, Clifford R, Hewitt SM, Shou JZ, et al. Global gene expression profiling and validation in esophageal squamous cell carcinoma and its association with clinical phenotypes. Clin Cancer Res. 2011;17:2955–2966. doi: 10.1158/1078-0432.CCR-10-2724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Li WQ, Hu N, Burton VH, Yang HH, Su H, Conway CM, Wang L, Wang C, Ding T, Xu Y, et al. PLCE1 mRNA and protein expression and survival of patients with esophageal squamous cell carcinoma and gastric adenocarcinoma. Cancer Epidemiol Biomarkers Prev. 2014;23:1579–1588. doi: 10.1158/1055-9965.EPI-13-1329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Dodd LE, Sengupta S, Chen IH, den Boon JA, Cheng YJ, Westra W, Newton MA, Mittl BF, McShane L, Chen CJ, et al. Genes involved in DNA repair and nitrosamine metabolism and those located on chromosome 14q32 are dysregulated in nasopharyngeal carcinoma. Cancer Epidemiol Biomarkers Prev. 2006;15:2216–2225. doi: 10.1158/1055-9965.EPI-06-0455. [DOI] [PubMed] [Google Scholar]
- 72.Sengupta S, den Boon JA, Chen IH, Newton MA, Dahl DB, Chen M, Cheng YJ, Westra WH, Chen CJ, Hildesheim A, et al. Genome-wide expression profiling reveals EBV-associated inhibition of MHC class I expression in nasopharyngeal carcinoma. Cancer Res. 2006;66:7999–8006. doi: 10.1158/0008-5472.CAN-05-4399. [DOI] [PubMed] [Google Scholar]
- 73.Hsu WL, Tse KP, Liang S, Chien YC, Su WH, Yu KJ, Cheng YJ, Tsang NM, Hsu MM, Chang KP, et al. Evaluation of human leukocyte antigen-A (HLA-A), other non-HLA markers on chromosome 6p21 and risk of nasopharyngeal carcinoma. PLoS ONE. 2012;7:e42767. doi: 10.1371/journal.pone.0042767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Rousseaux S, Debernardi A, Jacquiau B, Vitte AL, Vesin A, Nagy-Mignotte H, Moro-Sibilot D, Brichon PY, Lantuejoul S, Hainaut P, et al. Ectopic activation of germline and placental genes identifies aggressive metastasis-prone lung cancers. Sci Transl Med. 2013;5:186ra166. doi: 10.1126/scitranslmed.3005723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Chandran UR, Ma C, Dhir R, Bisceglia M, Lyons-Weiler M, Liang W, Michalopoulos G, Becich M, Monzon FA. Gene expression profiles of prostate cancer reveal involvement of multiple molecular pathways in the metastatic process. BMC Cancer. 2007;7:64. doi: 10.1186/1471-2407-7-64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Yu YP, Landsittel D, Jing L, Nelson J, Ren B, Liu L, McDonald C, Thomas R, Dhir R, Finkelstein S, et al. Gene expression alterations in prostate cancer predicting tumor aggression and preceding development of malignancy. J Clin Oncol. 2004;22:2790–2799. doi: 10.1200/JCO.2004.05.158. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets analyzed during the current study are available in the NCBI Gene database (https://www.ncbi.nlm.nih.gov/gene/) and GEO database (https://www.ncbi.nlm.nih.gov/geo/).