Abstract
Biliverdin reductase B (BLVRB) is a newly identified cellular redox regulator that catalyzes the NADPH-dependent reduction of multiple substrates. Through mass spectrometry analysis, we identified high levels of BLVRB in mature red blood cells, highlighting the importance of BLVRB in redox regulation. The BLVRB conformational changes that occur during conezyme/substrate binding and the role of dynamics in BLVRB function, however, remain unknown. Through a combination of NMR, kinetics, and isothermal titration calorimetry studies, we determined that BLVRB binds its coenzyme 500-fold more tightly than its substrate. While the active site of apo BLVRB is highly dynamic on multiple timescales, active site dynamics are largely quenched within holo BLVRB, in which dynamics are redistributed to other regions of the enzyme. We show that a single point mutation of Arg78➔Ala leads to both an increase in active site micro-millisecond motions and an increase in the microscopic rate constants of coenzyme binding. This demonstrates that altering BLVRB active site dynamics can directly cause a change in functional characteristics. Our studies thus address the solution behavior of apo and holo BLVRB and identify a role of enzyme dynamics in coenzyme binding.
Keywords: dynamics, enzyme, network, coenzyme, nuclear magnetic resonance
Introduction
BLVRB is an NADPH-dependent oxidoreductase that acts upon multiple substrates and has emerged as a critical regulator of cellular redox [1, 2]. Two biliverdin reductases (BLVRs) are encoded in humans, BLVRB and BLVRA. These enzymes share virtually no sequence similarity but do share structural similarity typical of the short-chain dehydrogease/reductase superfamily (SDR, see Fig. 1a). SDR members contain a similar N-terminal domain with a typical Rossmann-fold, but differ in the C-terminal domains that define their substrate specificities [4]. The primary role of BLVRs was initially thought to be confined to the second step in heme degradation in which biliverdin is reduced to bilirubin in an isomer-specific manner. Specifically, BLVRB reduces biliverdin-IXβ and BLVRA reduces the more abundant biliverdin-IXα [5]. However, BLVRB was later found to be a general flavin reductase (FR) that reduces numerous substrates, suggesting a more general role in redox regulation [2]. This generalized role is supported by a recent elegant study that discovered BLVRB mRNA levels increase during erythropoieses and suggests that BLVRB is in high abundance in mature red blood cells (RBCs) where it helps regulate cellular redox [6]. An inherent active site loss-of-function mutant was discovered, BLVRBS111L, which rewires the cellular redox state of hematopoietic cells to redirect cell fate to platelet production [6]. BLVRB is therefore the first identified therapeutic target proposed to treat patients with thrombocytopenia, a disease with high morbidity [7, 8], and becomes particularly acute in cancer patients [9, 10]. Elevated expression of BLRVB is also correlated with several cancers [11, 12], potentially protecting cells from cellular stress related to changes in their redox state.
Fig. 1.
Structural and biological comparison of BLVRB and BLVRA. (a) The holo enzyme structures of BLVRB (PDB accession 1HE4, protein in white and NADP+ in red) with both the relatively conserved SDR domain and variable C-terminal domain shown. (b) BLVRB protein levels dominate in RBCs relative to BLVRA, as assessed by two independent methods [3].
Despite the biological importance of BLVRB for regulating cellular redox and its high therapeutic potential, the conformational changes that occur upon coenzyme and substrate binding and the role of dynamics remain unknown. The elucidation of high-resolution structures of holo BLVRB in complex with several substrates have significantly contributed to our molecular understanding of the catalytic mechanism [13]. For example, the BLVRB catalytic mechanism was proposed to involve a two-step catalytic reaction of water-mediated substrate protonation followed by hydride transfer from the coenzyme [14–16], which is analogous to that proposed for dihydrofolate reductase (DHFR) [17, 18]. Interestingly, DHFR and BLVRB may invoke a similar “clamping” of the coenzyme despite their complete lack of structural similarity. For DHFR, solution studies have shown that the Met20 loop exchanges between at least a “closed” and “occluded” conformation, which regulates access to the active site [19, 20]. For BLVRB, Arg78 folds over the coenzyme and may underlie a similar “clamping” mechanism [13]. A lack of structural information for apo BLVRB has precluded comparisons between the apo and holo conformations, presumably due to the inherently flexible nature of the apo BLVRB active site. Thus, experiments that explicitly probe the movements of BLVRB would address the role Arg78.
Studies presented here identify the BLVRB active site as an inherently dynamic region on multiple timescales and offer insight into the role of dynamics in BLVRB function. Specifically, we discovered that the BLVRB active site comprises elevated ps–ns motions and μs–ms motions, but these motions are largely quenched upon coenzyme engagement (in holo BLVRB) with a redistribution of dynamics to other regions. We also show that BLVRB has a semi-ordered binding mechanism with relatively high affinity to the coenzyme and low affinity to a substrate. Finally, by combining relaxation studies that monitor dynamics with kinetic studies that monitor coenzyme binding, we show that mutation of Arg78 increases the rate of μs–ms motions within the active site that is concomitant with an increased rate of coenzyme binding and release. These studies are the first to elucidate the complicated role of BLVRB dynamics in function and point toward a conserved role for active site “clamping” via Arg78.
Results
BLVRB is present within mature RBCs
While BLVRA is present in mature RBCs where it serves a critical role in heme metabolism, the relative concentrations of BLVRB remain unknown. Thus, two proteomic methods were used to address the relative abundance of BLVRB to BLVRA and reveal that BLVRB is 10-fold higher than BLVRA in mature RBCs (Fig. 1b). These results are consistent with recent findings that have reported the expression of BLVRB increases during hematopoietic development, while other heme degradation genes, such as heme oxygenase and BLVRA, remain relatively constant [6]. Considering that RBCs are the most abundant human cell [21], BLVRB is therefore a highly abundant enzyme.
Apo BLVRB binds more tightly to its coenzyme than holo BLVRB binds to the FAD substrate
Previous UV-kinetics studies have indicated that BLVRB engages its coenzyme with a much tighter affinity than its substrates based on Michaelis–Menten kinetic constants (KM) [1], which were suggested to be due to larger conformational rearrangements associated with coenzyme binding. To further assess such changes, we first purified BLVRB, which gives rise to a well-dispersed 15N heteronuclear single quantum coherences (HSQC) spectrum. As the half-life of NADPH is less than 12 h [22] and precludes substrate titrations that would induce catalysis during titration experiments, we used the oxidized NADP+ product to model the holo BLVRB complex. Both NADPH and NADP+ bind with similar affinity (Fig. S1a), lending credence to utilizing the more stable oxidized NADP+. There are chemical shift perturbations (CSPs) due to this single hydride (Fig. S1b,c), which are nearly as large as those previously observed between reduced and oxidized forms bound to DHFR [23]. However, unlike DHFR where full assignments were necessary at lower temperatures to avoid NADPH oxidation [23], the lower CSPs between the BLVRB/NADPH and BLVRP/NADP+ readily allowed all amide resonances to be identified to also map onto the structure (Fig. 1d). As observed for other systems, oxidation is relatively fast and can be observed approximately 12 h after sample preparation (Fig. 1e). Thus, the oxidized NADP+ form of the coenzyme has also been used to model the Michaelis–Menton complex [20] and we will refer NADP+ as the coenzyme unless otherwise stated.
The coenzyme binds BLVRB within the slow NMR timescale, as apo BLVRB resonances (i.e., BLVRB alone) disappear upon titration with the coenzyme and resonances of holo BLVRB appear at new positions. Like the apo BLVRB HSQC, the holo BLVRB HSQC spectrum is also well dispersed but markedly different (Fig. 2a). To address changes induced by a substrate, we then used flavin adenine dinucleotide (FAD) that has previously been shown to be catalyzed by BLVRB [1]. In contrast to coenzyme binding, the FAD substrate binding to holo BLVRB is in fast exchange, and thus, CSPs could readily be followed with titrations that result in HSQC spectra similar to that of the holo BLVRB HSQC spectrum (Fig. 2b). We therefore assigned the backbone resonance of both apo and holo in order to identify the regions affected by binding both coenzyme and substrate, respectively. Over 90% of the backbone resonances of BLVRB were identified in both apo and holo BLVRB, noting that the majority of unassigned residues lie in two short stretches of residues 167–174 and 199–203 that are adjacent to each other within the C-terminal domain.
Fig. 2.
BLVRB binding to both coenzyme and FAD substrate. (a) HSQC spectrum of apo BLVRB (black) superimposed with holo BLVRB (red). (b) HSQC spectrum of holo BLVRB (red) superimposed with addition of 3.2 mM FAD (green). (c) Binding isotherm of apo BLVRB to the reduced NADP+ coenzyme monitored through ITC at 20°C. (d) Binding isotherms of holo BLVRB to FAD monitored through NMR titration for amides of Thr15 (red), Lys178 (green), Leu74 (purple), Ser88 (black), Arg174 (blue), and an example of the CSPs of Thr15. Binding isotherms were globally fit to extract the KD. (e) CSPs between apo BLVRB and holo BLVRB with 10 mM NADP+. (f) CSPs between holo BLVRB and holo BLVRB titrated with 3.2 mM FAD. (g) CSPs between apo and holo BLVRB versus the distance of each residue's amide to the coenzyme and mapped onto the X-ray crystal structure (purple) for CSPs >0.2ppm (PDB accession 1HE2). (e) CSPs between holo BLVRB and holo BLVRB titrated with 3.2 mM FAD substrate versus the distance of each residue's amide to the substrate and mapped onto the X-ray crystal structure (purple) for CSPs >0.2ppm (PDB accession 1HE4). Five proline residues comprise the FAD binding substrate pocket that includes 118, 122, 123, 151, and 152, and thus, there are no closer distances to amide than 6.5 Å. All NMR data were collected at 900 MHz at 20 °C.
The marked difference in coenzyme and substrate affinity is shown by full titration experiments to extract their dissociation constants (KD) from their respective binding isotherms. Due to the different timescales of binding, isothermal titration calorimetry (ITC) was used to determine a binding isotherm for both the reduced and oxidized coenzyme and NMR was used to determine a binding isotherm for the substrate. For the coenzyme, a KD of 0.3 ± 0.2 μM was calculated for BLVRB/NADPH and a KD of 0.4 ± 0.2 μM was calculated for BLVRB/NADP+ association (Figs. 2c and S1a). For the FAD substrate, a KD of 257 ± 22 μM was calculated for the holo BLVRB/FAD association (Fig. 2d). Thus, BLVRB binds its coenzyme some 500-fold tighter than its substrate.
We also addressed whether FAD can directly bind apo BLVRB (in the absence of coenzyme) and observed that many resonances perturbed in holo BLVRB upon titration of FAD are not perturbed within apo BLVRB (Fig. S2a compared to Fig. 2b). Instead, the FAD substrate induces only a subset of perturbations to apo BLVRB. These perturbations were used to calculate a KD for the apo BLVRB/FAD interaction of 590 ± 80 μM (Fig. S2b). Thus, apo BLVRB engages the FAD substrate 2-fold weaker than holo BLVRB.
Larger chemical shifts are induced upon apo BLVRB binding to the coenzyme than those induced upon holo BLVRB binding to the FAD substrate
Chemical shifts were compared between apo BLVRB and saturating concentrations of coenzyme (Fig. 2e) and between holo BLVRB and saturating concentrations of the FAD substrate (Fig. 2f). The coenzyme induced CSPs that are approximately an order of magnitude larger than those induced by the FAD substrate. A quantitative comparison of CSPs induced by either the coenzyme or substrate and the distance between each amide to the coenzyme or substrate reveals the long-range effects of binding on the chemical shifts (Fig. 2g, h). For example, the coenzyme induces CSPs over 1 ppm more than 15 Å away from the binding site (Fig. 2g). Several loops that include residues 9–14 and 72–87 that directly interact with the coenzyme in the X-ray crystal structure and have CSPs induced by coenzyme binding, also have measurable CSPs induced by substrate binding. Although there is no X-ray crystal structure of holo BLVRB bound to FAD, this FAD substrate induces CSPs to the C-terminal region of holo BLVRB that are consistent with the X-ray crystal structure of a similar ternary complex bound to flavin mononucleodite [13]. Compared to the CSPs induced by the FAD substrate to holo BLVRB, CSPs induced by the FAD substrate binding to apo BLVRB are generally lower (Fig. S2c, d).
Coenzyme binding induces changes to CA resonances within the BLVRB active site
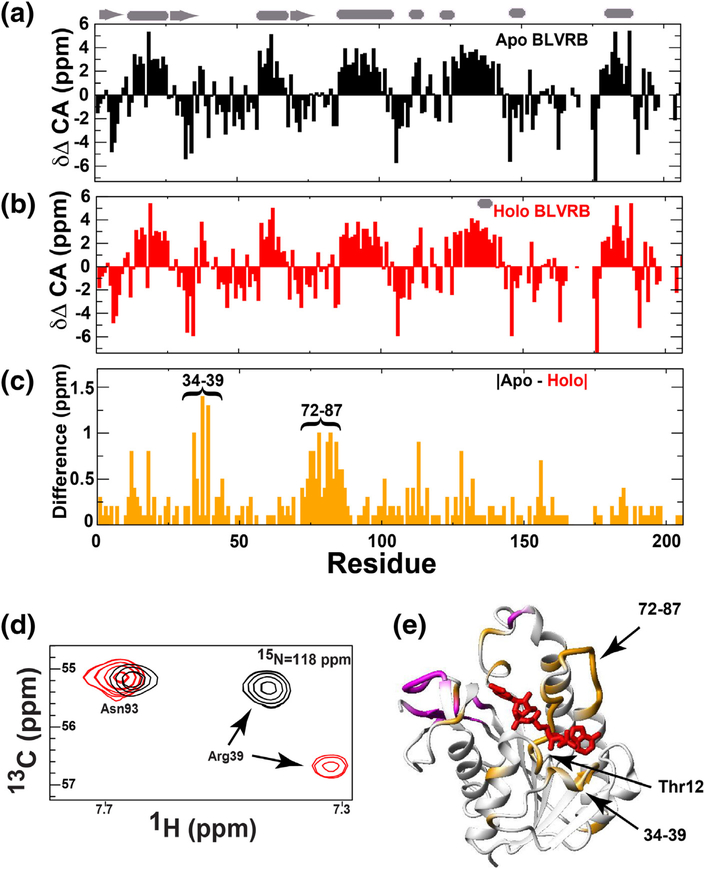
In addition to the high sensitivity of NMR in monitoring environmental changes through CSPs described above, NMR is also sensitive to subtle changes in secondary structure monitored through CA resonances and used here as a proxy for potential structural rearrangements that occur between apo and holo BLVRB (Fig. 3). Resonance assignments obtained for both apo and holo BLVRB were used to calculate the CA chemical shift differences relative to a random coil (δΔ CA), which are the most sensitive indicators of secondary structure (Fig. 3a, b), with positive and negative differences indicative of α-helical and β-strand secondary structure, respectively [24]. While these δΔ CA values are largely in agreement with the secondary structure found within the BLVRB ternary complexes solved by X-ray crystallography [13], our solution assignments also allow us to calculate the associated changes of δΔ CA between apo and holo BLVRB (Fig. 3c). Such changes are exemplified by the resonance shifts of residues such as Arg39 CA (Fig. 3d) and pinpoint several regions adjacent to the BLVRB active site that likely undergo secondary structural changes upon coenzyme engagement, including residues 34–39 and residues 72–87 (Fig. 3e). The majority of these changes to δΔ CA values are reflective of an increase in secondary structure propensity that is consistent with an increase in structural integrity upon coenzyme binding. From the holo BLVRB X-ray crystal structures, there are several arginine residues within these regions that properly align the coenzyme [13]. For example, Arg35 and Arg39 form stabilizing interactions for the adenosine moiety of the coenzyme, while Arg35 and Arg78 form an electrostatic interaction with the phosphate group of the coenzyme. In addition, Arg78 straddles over the coenzyme and forms a hydrogen bond with the carbonyl backbone of Thr12. Thus, these two regions of 34–39 and 72–87 exhibit large changes to their CA resonances upon coenzyme binding.
Fig. 3.
BLVRB secondary structure propensities and changes upon coenzyme binding. (a) CA chemical shift differences between apo BLVRB and CA resonances within a random coil (δΔ CA). (b) CA chemical shift differences between holo BLVRB and a random coil. (c) The absolute difference in CA chemical shift differences between apo BLVRB and a random coil and between holo BLVRB and a random coil. CA chemical shift propensities were calculated by subtracting random coil CA resonance reported at the BMRB (http://www.bmrb.wisc.edu/). (d) An example of the 3D-HNCA spectra for both apo and holo BLVR showing no change for Asn93 CA resonances and large changes for Arg39 CA resonances. (e) These CA chemical shift differences >0.4ppm in (d) are mapped onto the X-ray crystal structure of BLVRB (orange) with unassigned resonances also shown (pink). For secondary structure, positive values indicate α-helical secondary structure, and negative values indicate β-strand secondary structure. Cartoon representation of secondary structure from the X-ray crystal structure is shown at the top (PDB accession 1HE4).
Coenzyme binding induces changes to BLVRB dynamics
To further identify the regions that are responsible for coenzyme engagement, we compared the dynamic regions of apo and holo BLVRB. NMR relaxation techniques provide a broad range of methodologies that can probe the conformational landscape of proteins on a variety of timescales, which depend on the unique characteristics of each system. For apo BLVRB, both locally elevated ps–ns motions and slower μs–ms motions have been identified here that are specifically affected by coenzyme binding upon formation of the holo enzyme. The fact that the coenzyme induces such changes to most of these motions is significant, as it suggests that their dynamics are coupled to coenzyme binding.
15N R1 relaxation rates were used as a proxy for ps–ns motions (higher rates indicative of more disorder, lower rates indicative of more order) and clearly reveal a quenching of active site flexibility upon coenzyme binding (Fig. 4a). Specifically, two stretches of elevated R1 relaxation rates within apo BLVRB are observed. These include residues 76–82 within the active site loop and residues 36–38 that are immediately adjacent to the active site (Fig. 4b). Interestingly, while R1 relaxation rates are diminished within the active site upon coenzyme binding, R1 relaxation rates increase around the periphery of holo BLVRB (Fig. 4b, yellow versus cyan, respectively). There are only several residues that are relatively unperturbed by coenzyme binding, such as Ser42 and Gln206 that exhibit elevated R1 relaxation rates in both apo and holo BLVRB. Examples of fitted data used to calculate R1 relaxation rates are also shown (Fig. S3a)
Fig. 4.
BLVRB dynamics are altered upon coenzyme binding. (a) 15N R1 relaxation rates for both apo BLVRB (black) and holo BLVRB (red). Inset: blowup of 15N R1 relaxation rates for the region of 72–86. (b) Residues with 15N R1 relaxation rates greater than one SDEV above the mean for apo BLVRB (left, yellow) and holo BLVRB (right, cyan) are mapped onto the X-ray crystal structure (PDB accession 1HE2). (c) R2 relaxation rates for both apo BLVRB (black) and holo BLVRB (red) extracted for each residue from 15N amide R2-CPMG dispersions at the lowest CPMG field of 50 Hz. (d) Residues exhibiting R2-CPMG dispersions greater than 1 Hz at a static field of 900 MHz are mapped onto the holo BLVRB structure for apo BLVRB (yellow) and for holo BLVRB (cyan). (e–i) R2-CPMG dispersions for residues within apo BLVRB (black) and holo BLVRB (red) individually fit (fitted line). All atoms from residues exhibiting elevated R1 relaxation rates and R2-CPMG dispersions are shown based on their respective 15N relaxation data.
R2-Carr-Purcell-Meiboom-Gill (R2-CPMG) dispersion was used to identify regions within both apo and holo BLVRB exhibiting chemical exchange on the μs–ms timescale and also reveals a change the dynamics between the two forms (Fig. 4c–l). For example, while over 30 residues exhibit chemical exchange in apo BLVRB, the chemical exchange contribution (Rex) was quenched for all of these residues in holo BLVRB (Fig. 4c and d, yellow). In fact, the majority of residues that exhibit chemical exchange in apo BLVRB exhibit no detectable dispersion within holo BLVRB (Fig. 4c, e–j, l). Only two of these residues still exhibit detectable R2-CPMG dispersion in holo BLVRB, Ser111 (Fig. 4j) and Thr164 (Fig. 4k). Conversely, only three residues exhibit exchange in holo BLVRB, which include Asp36, Ser37, and Arg39 (Fig. 4c and d, cyan), but exhibit no detectable exchange in apo BLVRB (Fig. 4f). Chemical exchange was on the fast NMR timescale for those residues that could be accurately fit to the Carver–Richards equations (see Materials and Methods) and the associated rates of chemical exchange (kex) were extrapolated by collecting R2-CPMG dispersion at two fields for apo BLVRB (Fig. S3b). However, while kex is well determined for these residues, the extracted parameters that include the chemical shift difference between sampled conformers and their relative populations cannot be accurately deconvoluted under such an exchange regime.
Despite our findings that show all residues exhibiting R2-CPMG dispersions are impacted by coenzyme binding (i.e., either quenched or altered), the localized and complicated nature of enzyme dynamics is immediately apparent. For example, while some exchange rates could not be accurately extracted from R2-CPMG dispersions, there is a wide range of extracted exchange rates from those that could (Table S1), which suggests that many motions are at least partially localized. For example, R2-CPMG dispersion is linear for Thr164 and cannot be suppressed by the highest imparted refocusing field (νcpmg) within either apo or holo BLVRB (Fig. 4k). In contrast, several residues that are within close proximity to each other and within the active site of apo BLVRB exhibit broadly similar rates of chemical exchange, which include Thr77 (Fig. 4g), Cys109 (Fig. 4i), and Ser111 (Fig. 4j) and could be globally fit to a single dynamic presented below in the context of a comparison to a BLVRB mutant. However, Thr77 exemplifies the complicated nature of dynamics, as its amide is involved in at least two dynamic exchange processes in apo BLVRB. Specifically, although Thr77 exhibits an exchange rate similar to both Cys109 and Ser111, its neighboring residues of Arg78 and Asn79 do not exhibit R2-CPMG dispersion (Fig. S4a–c). As exchange could not be suppressed for residues 77–79 in apo BLVRB even at the highest imparted R2-CPMG frequency (Fig. S4d), this means that there is at least another dynamic exchange process for these residues that occurs on a much faster μs timescale on top of the exchange process identified for Thr77 through R2-CPMG dispersion. While exchange appears to persist at multiple temperatures for residues 77–79 as observed by their severe line broadening (Fig. S4e–g), these exchange processes are quenched upon coenzyme binding in holo BLVRB (Fig. 4c).
Mutation of Arg78 alters coenzyme binding
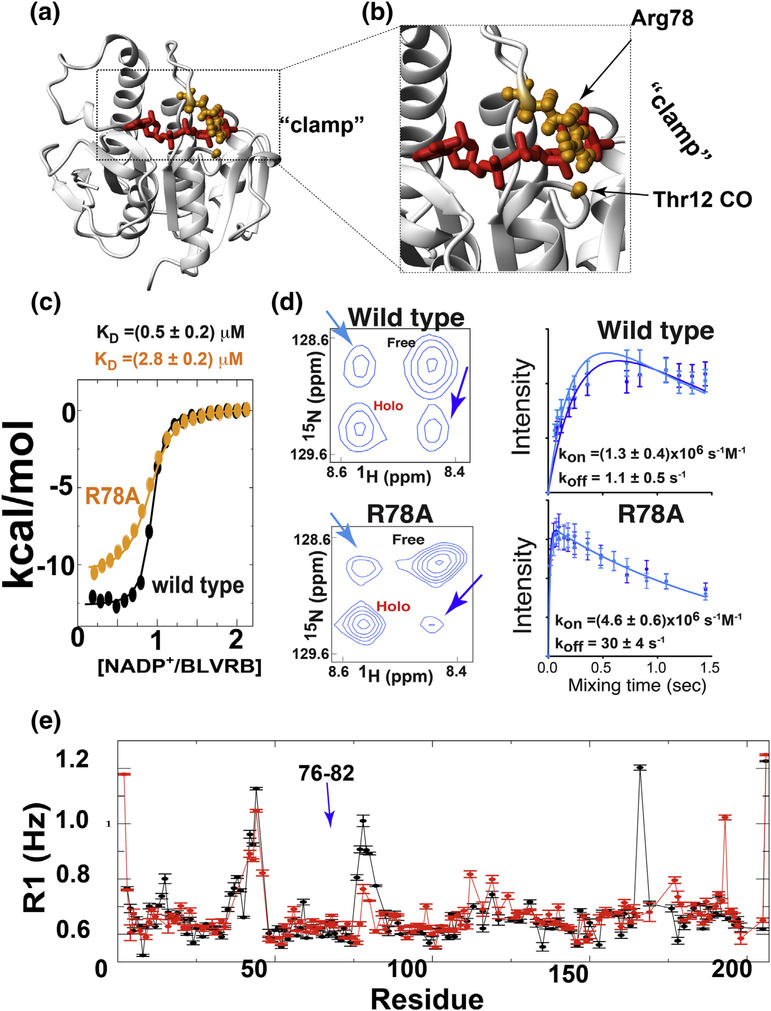
A mechanistic role of Arg78 that was previously implicated from holo BLVRB X-ray crystal structures [13] and is now supported by NMR solution studies prompted further investigation to specifically address its importance. Specifically, X-ray crystal structures of holo BLVRB clearly show that Arg78 straddles the coenzyme to form a hydrogen bond with the Thr12 carbonyl backbone, suggesting a “clamping” role (Fig. 5a, b), which is consistent with NMR solution studies that reveal that dynamics on multiple timescales are quenched upon binding. Therefore, to experimentally probe the role for Arg78 in “clamping” of the coenzyme, we engineered R78A (Arg78➔Ala) for a comparative analysis with the wild type BLVRB. The R78A mutant gave rise to a well-resolved HSQC similar to the wild type enzyme (Fig. S5a). Both the thermodynamics and the kinetics of coenzyme binding of BLVRB R78A were compared to that of wild type BLVRB using ITC and NMR studies, respectively (Fig. 5c, d).
Fig. 5.
Functional and dynamic impact of BLVRB R78A mutation. (a) Arg78 “clamps” over the coenzyme (red) to form a hydrogen bond with the backbone carbonyl of Thr12. Atoms are shown as balls (orange) in holo BLVRB (PDB accession 1HE2). (b) Blow-up of the active site and the hydrogen bond between Arg78 and Thr12. (c) ITC titration for BLVRB binding to NADP+ with wild type (black) and R78A (orange) with an extracted KD of 0.5 ± 0.2 μM and 2.8 ± 0.2 μM, respectively. (d) ZZ-exchange buildup curves for exchange peaks of Val52 with 500 μM 15N-labeled wild type BLVRB or the R78A mutant in the presence of 250 μM NADP+. Left: Spectrum of each is shown for a 1.08-s delay, with arrows indicating the exchange peaks. Right: Build-up curves shown for exchange peaks with extracted rates derived from fits to all four intensities of kon=1.3 ± 0.4 (s μM)−1/koff = 1.1 ± 0.5 s−1 and kon = 4.6 ± 0.6 (s μM)−1/koff = 30 ± 4 s−1. (e) R1 relaxation rates of the R78A mutant for both apo (black) and holo (red) forms. We note that in the holo R78A mutant, that residues 36–38 are absent as 36–37 are poorly fit and Ser38 is overlapped.
ITC reveals nearly a 6-fold diminishment in affinity of NADP+ binding to BLVRB R78A relative to the wild type enzyme (Fig. 5c). This is consistent with previous kinetics studies that also indicated a weaker coenzyme affinity [15]. The difference in coenzyme binding affinity would not be expected to significantly affect substrate turnover, which is collected at concentrations significantly higher than the KD of either the wild type or R78A mutant. Indeed, UV-kinetics data indicate that substrate turnover is within uncertainty for both wild type BLVRB and the R78A mutant (Fig. S5b).
ZZ-exchange spectroscopy was used to determine the kinetic impact of perturbing Arg78 to the on-rate (kon) and the off-rate (koff) of the BLVRB/coenzyme complex using 500 μM 15N-labeled BLVRB (wild type or the R78A mutant) and 250 μM coenzyme (Fig. 5d). This allows both the apo (i.e., free) and holo (i.e., bound) resonances to be observed simultaneously, thereby giving rise to exchange peaks between the apo and holo forms. Spectral analysis reveals the apo and holo BLVRB resonances could be resolved along with the exchange peaks for only a subset of residues. However, the amide of Val52 was resolved in both wild type BLVRB and the R78A mutant (Fig. 5d, left). From these data, kon and koff were extracted by simultaneously fitting all four resonances as a function of time (two auto peaks and two exchange peaks), as recently described [25, 26]. These data clearly indicate that exchange is substantially faster in the BLVRB R78A mutant relative to the wild type enzyme (Fig. 5d, right). For example, the large increase in these microscopic rate constants is immediately apparent in the shifted maximum intensities of the exchange peaks for BLVRB R78A relative to the wild type enzyme. Quantitatively, the R78A mutation induces a 3-fold increase in kon and a 27-fold increase in koff, indicating that the large increase in koff underlies the weakening of the KD. Interestingly, despite the critical role that Arg78 plays in coenzyme binding, the mobility of residues 76–82 is still restricted upon coenzyme binding similar to that found in wild type BLVRB, as shown by a comparative analysis of R1 relaxation rates between apo and holo BLVRB R78A (Fig. 5e). Thus, while mutation of Arg78 does induce weaker coenzyme binding to apo BLVRB with a more dramatic change to the koff of the coenzyme, the active site loop has become relatively restricted in the holo form of the mutant as it is in the holo form of the wild type enzyme.
Arg78 is coupled to global dynamics on the slow timescale
A comparative analysis of R2-CPMG relaxation data for wild type BLVRB and the R78A mutant reveals that the dynamics of the BLVRB active site are coupled to distal motions within the enzyme on the μs–ms timescale (Fig. 6). Specifically, R2-CPMG relaxation dispersion was used to monitor changes to μs–ms motions induced by the R78A mutation (Table S1), revealing that the exchange contribution (Rex) increases for most residues when compared to the wild type enzyme (Fig. 6a). These changes can be mapped along one side of the enzyme (Fig. 6b) and comprise the same region that exhibits R2-CPMG dispersion within the wild type enzyme (Fig. 4d), suggesting that these residues form a dynamically coupled network. Interestingly, some residues that exhibit no detectable exchange in the wild type enzyme have exchange within the R78A mutant, such as Lys105 that is 24 Å away from the mutation site of Arg78 and also includes Val107 and Ala108 within the same β-strand (Fig. 6b, bottom). While R2-CPMG dispersions for many residues could not be accurately fit for either the R78A mutant or the wild type enzyme, a subset of active site residues could be fit to estimate local exchange rates (kex), such as Ser111 and Leu125. Specifically, mutation-induced increases in exchange rates from the wild type enzyme to the R78A mutant are observed in Ser111, which increases its exchange rate from 1200 ± 300 s−1 to 1900 ± 200 s−1, and Leu125, which increases from 2200 ± 540 s−1 to 4800 ± 1300 s−1(Table S1), indicating that at least these local rates of motions are increased in the mutant.
Fig. 6.
Dynamics within BLVRB are distally coupled to the active site. (a) R2-CPMG dispersions for both wild type BLVRB (black) and R78A (orange) are shown for Asp80, Lys105, Val107, Ala108, Cys109, Ser111, Leu125, and Tyr156. (b) Changes between wild type BLVRB and the R78A mutant R2-CPMG dispersions mapped onto the structure of holo BLVRB (orange). Two orientations are shown along with the largest distance between coupled residues (red arrows), and all atoms are depicted for those residues in which the full 15N R2-CPMG dispersions are shown.
Several active site residues could be globally fit for comparisons between the wild type and R78A active site dynamics. However, caution should be taken in over-interpreting global fits without any a priori reason other than their close spatial proximity to each other as has been previously reported for other systems [27–29]. For example, the R78A mutation induces more complicated R2-CPMG dispersion profiles for two residues relative to wild type BLVRB, Thr77 and Thr85, which are near the potential “hinge” region of the larger Arg78 loop and near each other (Fig. S6a). Nevertheless, several residues within the BLVRB active site exhibit similar exchange rates within the wild type enzyme (Cys109, Ser111, and Thr77) and the R78A mutant (Ala108, Cys109, Ser111, and Asp80), which were extracted from R2-CPMG dispersions (Table S1). These residues could be global fit within each enzyme and once again suggests an increase in active site motions upon mutating Arg78 (Fig. S6b). Regardless of whether these active site residues are globally fit or independently fit, several residues such as Ser111 and Leu125 are faster within the BLVRB R78A mutant than the wild type enzyme. Thus, faster motions within the active site induced by this single point mutation are coupled to an increase in kon and koff rates monitored through ZZ-exchange (Fig. 5d), thereby identifying a causal link between dynamic control of the BLVRB active site and coenzyme binding.
Discussion
BLVRB affinities to its coenzyme and substrate and potential conformational rearrangements
The NMR solution studies performed here facilitate comparisons of the conformational changes that ensue upon coenzyme and substrate binding to BLVRB. Of particular relevance are the large-scale CSPs induced upon coenzyme binding that are in sharp contrast to the more localized CSPs induced upon substrate binding. For example, some of the largest CSPs induced by the coenzyme include residues 72–87 that are immediately within the BLVRB active site and the adjacent region of residues 34–39. This is consistent with a recent study that has shown that Arg35 is at least partially responsible for the selectivity of NADPH over NADH, as its side chain forms a direct interaction with the phosphate of the coenzyme [15]. Although in the absence of an apo BLVRB structure, it is difficult to deduce whether CSPs due to coenzyme binding are localized to side chain rearrangements or larger secondary structure rearrangements, it is worth noting that coenzyme binding to BLVRB induces higher CSPs than those previously reported for coenzyme binding to DHFR [23]. Thus, while there is no straightforward relationship between CSPs and structural changes, it is clear that the chemical environment changes for BLVRB upon coenzyme binding include regions distal to the active site.
Solution studies performed here also allow us to directly quantify coenzyme and substrate binding affinities for the first time as well as the order of binding through CSPs using the apo and holo enzyme. For example, thermodynamically, the relatively high binding affinity of the coenzyme compared to the FAD substrate indicates that the coenzyme would normally be bound first. NMR perturbation analysis also reveals that the holo enzyme associates with the substrate approximately 2-fold tighter than the apo enzyme. Collectively, these data suggest that BLVRB adopts a semi-ordered binding mechanism that is consistent with the structure of the ternary complexes. For example, the coenzyme is relatively buried within BLVRB where the nicotinamide ring facilitates a stacking interaction with the substrate isoalloxazine ring that defines flavins [9]. Thus, the previously proposed ordered binding mechanism of BLVRB determined through UV-kinetics studies is consistent with our findings here based on NMR studies [1].
BLVRB exhibits both similarities and clear distinctions from other reductases in regard to coenzyme and substrate binding. For example, reductases that include DHFR, BLVRA, and BLVRB engage their coenzyme with similar micromolar binding affinities [30, 31]. In fact, the quantitative analysis here of BLVRB binding to NADP+ extracted from ZZ-exchange spectroscopy has revealed that the microscopic rate constants are nearly identical to that previously determined for DHFR [32]. Both DHFR and BLVRB coenzyme on-rates are substantially slower than diffusion-limited on-rates, which could be due to occlusion of the active site discussed further below. In contrast to coenzyme binding, BLVRA binding to its biliverdin-IXα substrate and DHFR binding to its dihydrofolate (DHF) substrate are both in the low micromolar range [30, 31], whereas BLVRB binding to the FAD substrate is over 2 orders of magnitude weaker. In general, a weaker substrate binding of BLVRB could represent an evolutionary tradeoff with substrate promiscuity for BLVRB, whereas both BLVRA and DHFR are selective for their substrates. BLVRB's substrate promiscuity may also underlie its role in regulating cellular redox and hematopoietic cell fate [6], which is consistent with its relatively high abundance in mature RBCs shown here using two methods of MS analysis.
BLVRB is an inherently dynamic enzyme on multiple timescales
In addition to the conformational changes imparted by coenzyme binding, BLVRB is a dynamic enzyme in both its apo and holo forms. NMR relaxation studies performed here reveal that the inherent dynamics of BLVRB occur on multiple timescales. Moreover, the fact that monitored R1 relaxation rates and R2-CPMG dispersions change throughout the enzyme upon coenzyme binding means that these motions are coupled to the active site and potentially coupled to function. For example, the apo BLVRB active site exhibits elevated R1 relaxation rates, indicating local disorder, and exhibits R2-CPMG dispersion representative of μs–ms motions, which are both largely quenched upon coenzyme binding. Coenzyme binding induces a redistribution of motions, which can be seen by plotting the changes to R1 relaxation rates. These changes are associated with a decrease in ps–ns motions within the enzyme active site but an increased in ps–ns motions to the periphery of BLVRB (Fig. 7a), potentially suggesting a conservation of global ps–ns dynamics. Slow timescale dynamics are also redistributed, but to a lesser extent. For example, while most residues exhibit a quenching in their slow timescale dynamics, residues within the loop of 36–39 exhibit an increase (Fig. 7b). It should also be noted that R2CPMG dispersions suggest that intermediate motions on the μs timescale are also largely quenched, as R2 relaxation rates that remain elevated for the highest imparted CPMG field within apo BLVRB are diminished in holo BLVRB (meaning, elevated R20 values in apo BLVRB shown in Fig. S4d such as residues 77–79 are suppressed in holo BLVRB shown in Fig. 4c). Such changes indicate that BLVRB is inherently dynamic on a wide range of timescales and that these dynamics are all specifically affected by coenzyme binding. As the currently known flavin substrates bind relatively weakly, assessing the dynamics in the substrate-bound state may be complicated by their relatively fast on-/off-rates [33]. However, current efforts to identify the broad range of substrates catalyzed by BLVRB are underway and may help identify tighter binding substrates to facilitate such dynamic comparisons to ternary complexes. Nonetheless, our studies suggest that the inherent dynamics of BLVRB are a form of conformational selection that allows for coenzyme binding and that once the holo BLVRB complex is formed, the creation of a relatively rigid binding surface allows binding of multiple substrates.
Fig. 7.
The BLVRB active site is inherently dynamic. (a) Monitored changes to disorder within the ns-ps timescale. Entire residues are colored if their 15N R1 relaxation rates are diminished (yellow) or increased (cyan) greater than 0.5 SDEV (0.029 s−1) above the average change (0.032 s−1) for all residues, and the active site is denoted by an arrow (blue) to Arg78. (b) All three loops surrounding the BLVRB active site of residues 10–15, 76–82, and 109–114 exhibit μs–ms motions quenched upon coenzyme binding (yellow) as monitored through R2-CPMG dispersion. Only residues Asp36, Ser37, and Arg39 exhibit an increase in exchange upon coenzyme binding (cyan). Only the backbone atoms are colored. (c) The BLVRB loop region of 75–82 (orange) that comprises Arg78 is dynamic on multiple timescales and becomes ordered upon coenzyme binding. For wild type BLVRB, an exchange rate of this loop region estimated from a global fit of several adjacent residues (Fig. S6b) is shown and the microscopic exchange rates between apo and holo BLVRB are shown (from ZZ-exchange spectroscopy, see Fig. 5d). Two structures of apo BLVRB were modeled using harmonic constraints using Chemistry at Harvard Macromolecular Mechanics (CHARMM, version 36b1) for all backbone residues except for residues within the Arg78 loop. These include residues 75–84 that comprise elevated R1 relaxation rates for 76–82. Although R1 relaxation rates for Leu74, Pro83, and Thr84 were not determined due to overlap and the absence of a proline amide, these residues were not constrained as they reside within this loop region.
While the instability of NADPH currently precludes relaxation measurements, we have identified measurable differences between holo BLVRB bound of NADPH and NADP+ due to the single hydride using HSQCs that may have a dynamic link (Fig. S1). Specifically, many of the same amides that exhibit large CSPs between the two bound coenzyme forms are also the same amides that are inherently dynamic within apo BLVRB. These include residues distant to the active site such as Thr164 and Ile176, which at the very least means that residues that are inherently dynamics in apo BLVRB are also in direct communication with the hydride in holo BLVRB. Whether the single hydride difference results in shifts in populations that induce some chemical shift changes in the holo forms or whether these are mechanical structural changes will await further studies. For example, elegant combinations of other enzymes to recycle NADPH may be used to increase the lifetime of NMR samples and facilitate both relaxation and structural studies [34].
Although progress has been made in understanding the role of active site fluctuations in enzyme function [33, 35–40], recent studies that include our own have explicitly shown that dynamic couplings over long distances are a form of allosteric communication tied to active site motions and function [41–43]. Coupled dynamic networks have been shown to exist in multiple proteins regardless of whether their rates of exchange are identical, which suggests a more complicated coupling than direct mechanically coupled networks. In fact, the increase in exchange (Rex) observed within the β-strand of residues 105–108 within the R78A mutant could collectively suggest an increase in a sampled minor conformation induced by the R78A mutation (Fig. 6a). What this explicit conformation physically encompasses will likely await further structural studies that probe apo BLVRB. The identification of coupled dynamic networks within BLVRB that are altered by both coenzyme binding and the R78A active site mutation now facilitates studies to address how these distal dynamic segments regulate active site dynamics and, in-turn, how this may modulate function. This is the basis of our recently proposed methodology referred to as Relaxation And Single Site Multiple Mutations (RASSMM) [41], which relies first on identifying potentially coupled residues to an enzyme's active site as shown here for R78A induced dynamic changes that are distant to the BLVRB active site (Fig. 6). A series of mutations are then constructed along these identified networks whereby both the dynamics and function are compared to deduce the role of dynamic communication in enzyme function without directly perturbing the active site. Future studies using this approach will allow us to determine the role of motions distal from the BLVRB active site, such as the dynamics of Thr164 that resides within the C-terminal region of BLVRB.
A link between active site dynamics and coenzyme binding identified via BLVRB Arg78
BLVRB Arg78 plays a role in both active site dynamics and function, as assessed by NMR relaxation experiments and coenzyme binding experiments, respectively. A mechanistic role was initially postulated based on the holo BLVRB X-ray crystal structures where Arg78 “clamps” down over the coenzyme and forms multiple contacts with both the coenzyme and backbone carbonyl of BLVRB Thr12 [13]. Regardless of whether motions are localized to individual residues or represent a more concerted dynamic, they are nonetheless faster within the R78A mutant than within wild type BLVRB. In addition to this increase in the exchange rates induced by the R78A mutation in apo BLVRB, the coenzyme binding kinetics are also faster as determined by ZZ-exchange spectroscopy (Fig. 5d). Thus, faster dynamics imparted by this active site mutation are associated with a concomitant increase in microscopic rate constants of coenzyme binding, suggesting that there is a link between inherent motions and coenzyme binding.
A simplified model based on both relaxation experiments and binding experiments may be drawn that includes the role of Arg78 (Fig. 7c). Specifically, the disordered loop comprising Arg78 may undergo an inherent conformational dynamic in apo BLVRB that either leaves the active site partially accessible (Fig. 7c, top right) and partially closed (Fig. 7c, bottom right). The rate of exchange between these two forms can be estimated from a global fit of the R2-CPMG dispersion (Fig. S6b). Conversely, upon substrate engagement, the active site within holo BLVRB is relatively ordered and does not exhibit this extensive dynamic exchange. Whether Arg78 motions are involved in the chemical step or solely coenzyme binding will require additional studies that probe hydride transfer [32]. Regardless, motions within the Arg78 loop identified in apo BLVRB appear to be similar to that of the Met20 loop of DHFR that closes over its substrate [19, 44]. Studies here have therefore characterized the inherent dynamics of the active site and particularly the role of Arg78 that contributes to coenzyme binding.
Finally, the role of Arg78 is highlighted by its evolutionary dynamics. Normally, residues with key roles in function, such as the substrate-binding role of Arg78, lead to extreme evolutionary conservation. However, in BLVRB, Arg78 is not conserved, but is replaced by the radically different glycine amino acid in lemurs, for example (Fig. 8). How can Arg78 be so important in human BLVRB, but seemingly not conserved? This conundrum is apparent upon inspection of the three-dimensional structure, as the arginine has not disappeared but has been swapped into the adjacent loop at active site residue 14. Most BLVRB enzymes have an arginine at either residue 14 or residue 78 (human numbering), although a subset maintain an arginine at both sites (Fig. S7). In primates, the two substitutions were made on the same branch separating the common ancestor of the Simiiformes (apes, new and old-world monkeys) with the common ancestor of the Haplorhini (i.e., after the split from the common ancestor with the tarsier). This suggests possible adaptive coevolution at these two sites and, from our mechanistic studies, plausibly leads to the idea that adjusting the location of this arginine serves to fine-tune coenzyme binding in BLVRB enzymes.
Fig. 8.
Evolutionary swapping of human Arg78 within the BLVRB active site. The phylogenetic tree graphic is derived from the tree shown in Figure S7, but focused on the Primates. Multiple BLVRBs from the Macaques, New World Monkeys, and Lemurs are shown as filled triangles on the tree. As in SFX, the two focal amino acids, at positions 14 and 78 (numbered relative to human positions), are shown along with the surrounding 3 amino acids on either side in the Ensembl amino acid alignment. The sequence examples shown illustrate the double substitution at sites 14 and 78 from the common ancestor of haplorrhynes to the common ancestor of new and old world monkeys (Simiiformes), along with the general conservation of the surrounding sites. The human BLVRB structure is shown with positions 14 and 78 in spacefill to illustrate their relationship to the rest of the structure. The lemur sequence shown is from the mouse lemur, Microcebus murinus.
Materials and Methods
Proteomics analysis via GeLC–MS and high-pH reversed phase fractionation coupled to nano-UHPLC–MS/MS
Whole blood was collected from healthy donor volunteers after signing of informed consent in agreement with the Declaration of Helsinki. Upon centrifugation for 10 min at 4 C at 2500g, RBC pellets were collected and further leukocyte filter (log4 white blood cell depletion, as per transfusion service standards) prior to further washing three times in PBS and centrifugation to remove any residual non-RBC contaminant. Cells were thus lysed through hyposmotic stress with cytosol and membranes separated via ultracentrifugation and washes in PBS for three cycles at 15,000g. Proteins were thus processed via GeLC (1DSDS-PAGE followed by digestion of the single lane excised into 12 bands, tryptic digestion, and nano-UHPLC–MS/MS). In parallel, a two-dimensional chromatography method was used. Proteins were separated by high-pH reversed phase fractionation into 64 fractions, 32 for RBC cytosol and membranes each, prior to tryptic digestion. Peptide fractions were then analyzed by nano-UHPLC–MS/MS analyses (nanoEasy LC II coupled to QExactive HF; Thermo Fisher, Bremen, Germany) for peptide identification. Analytical details of these workflows have been described extensively [3, 45, 46].
Cloning, protein expression, and purification
Human BLVRB (UniProt P30043) codon optimized for bacterial expression with an N-terminal 6xHis-tag and thrombin cleavage site was cloned into pET21 for expression in BL21/DE3 cells. The BLVRB R78A mutation was engineered through site-directed mutagenesis using the forward primer, catcgtgctgctg ggcaccgccaatgacctcagtcccacg, and its reverse complement. Unlabeled proteins were grown in Luria broth (LB) while labeled proteins were grown in M9 minimal media (6 g/l Na2HPO4, 3 g/l KH2PO4, 0.5 g/l NaCl, 1 g/l NH4Cl, 2 g/l glucose, 2 ml of 1 M MgSO4, 100 ml of 1 M NaCl CaCl2, 10 mg/l thiamine). 15N,13C-labeled proteins for backbone assignments were initially grown in M9 media with both 15N ammonium chloride and 13C glucose (protonated), yet production in 100% D2O to afford 2H,15N,13C-labeling facilitated further assignments of residues initially too line broadened. Proteins for R2-CPMG dispersion were grown with 15N ammonium chloride in D2O as previously described [29]. Although the level of deuteration was not quantified, R2 rates were significantly lower than non-deuterated counterparts (data not shown). Transformed BL21/DE3 cells supplemented with 100 mg/ml ampicillin were grown until an OD (600nm) of approximately 0.6 and induced for 3–4 h prior to harvesting. Cell pellets were homogenized via sonication in “denaturation buffer“ [5 M guanidine, 50 mM Na2HPO4 (pH 7.5), 500 mM NaCl, 10 mM imidazole], applied to Ni-affinity resin column (Sigma) and eluted with “denaturation buffer” supplemented with 400 mM imidazole. Elutions comprising BLVRB were acidified to pH 2–3 for application to a 3-ml Resource 15RPC (GE Healthcare Life Sciences) in order to remove residual coenzymes from bacterial growths. This reverse phase purification was found to be absolutely necessary, as even dialysis under denaturing conditions was not able to remove residual impurities that led to binding within 1%–3% of holo BLVRB as assessed through HSQC spectra with the appearance of coenzyme bound resonances (data not shown), which were also previously found in different purification protocols [15]. Denatured protein was then subjected to a series of dialysis steps that we have previously described [29]. Briefly, elutions were first diluted with “refold buffer” [100 mM Tris (pH 7.5), 100 mM NaCl, 1 mM DTT] supplemented with 5 M guanidine-HCl for dialyses for at least 36 h against “refold buffer” supplemented with 1 M arginine. All buffers were at a final pH of 7.5. Dialysis buffer was changed at least two times using “refold buffer” alone, concentrated, the 6xHis tag removed at room temperature overnight, and then applied to a Superdex-75 (GE Healthcare Life Sciences) equilibrated in “NMR buffer” [50 mM Bis–Tris (pH 6.5), 50 mM NaCl, 1 mM DTT]. BLVRB solutions were then further concentrated using Amicon centrifuge filters (3 kDa MWCO) as needed and stored at −80 °C until further use.
ITC and UV-kinetic assays
For ITC, a MicroCal VP-ITC was used with well samples containing 100 μM BLVRB (and BLVRB R78A) with either NADPH or NADP+ titrant at 1 mM and at 20 °C. NADPH was prepared immediately prior to ITC in order to avoid significant oxidation. All buffers were identical to that described above for “NMR buffer.” ITC experiments were performed in duplicate and processed using Origin software provided with the MicroCal VP-ITC. For UV-kinetic assays, a Bioteck Synergy 2 multi-mode detection plate reader was used with 100 μl total volumes to observe the diminishment in NADPH at 340 nm as previously described [1]. NADPH concentrations were kept constant at 300 μM, BLVRB (and BLVRB R78A) was kept constant at 10 μM, and the FAD substrate was varied as indicated. Each velocity was determined from an average of at least three kinetic runs with initial velocities calculated for the first 20–30 s and fit to the Michaelis–Menton equation using GraphPad Prism version 4.0 (GraphPad Software Inc., San Diego, CA).
NMR sample preparation, spectroscopy, and data analysis
All apo BLVRB samples were prepared in 50 mM Bis–Tris (pH 6.5), 50 mM NaCl, and 1 mM DTT at 500 μM with 5% D2O and holo BLVRB samples contained 10 mM NADPH or NADP+. NADPH was again prepared immediately prior to sample preparation in order to avoid oxidation, which was monitored through chemical shifts reverting back to that with the reduced NADP+ form (Fig. S1). Backbone assignments using a 2H,15N,13C-labeled enzyme were initially collected on a Varian 600 spectrometer equipped with a cryo-probe at 20 °C for both apo and holo BLVRB using standard BioPack sequences that included an HNCA, HN(co)CA, HNCACB, and a CBCA(co)NH. As the CA resonances are readily obtained from both the HNCA and HN(co)CA, both the HNCACB and CBCA(co)NH were optimized for CB coupling, specifically by setting the delay for CC coupling to 6 ms. In addition, an HNCA and HN(co) CA were collected at 10 °C for apo BLVRB on a Bruker 800 at the High Magnetic Field Laboratory along with temperature titrations, which helped facilitate the assignment of several residues such as Arg78 and Asn79 (Fig. S4). Specifically, while the amides of these residues are still line broadened at 10 °C, their respective CA correlations were more readily identified at the lower temperature. For the R78A mutation, most amide resonances were not significantly perturbed. However, amide assignments were confirmed via a 3D 15N-NOESY–HSQC by comparison to the wild type 3D 15N-NOESY–HSQC, whereby both spectra were collected on a Varian 900 MHz spectrometer. Spectra were processed using NMRPipe [47] and all data were analyzed using CCPNmr software [48]. All 3D assignment spectra collected on Varian 600- and 900-MHz spectrometers utilized non-uniform sampling using the method of Hyberts et al. [49] reconstructed to 51 and 96 increments in the nitrogen and carbon dimensions, respectively. All relaxation experiments were conducted on the Rocky Mountain Varian 900 spectrometer equipped with a cryo-probe at 20 °C with R1 relaxation experiments utilizing a standard BioPack sequence, and R2-CPMG dispersions were collected with TROSY selection as previously described [29, 41]. R2-CPMG dispersions for apo BLVRB were also collected on a Varian 600 MHz spectrometer at the High Magnetic Field Laboratory at 20 °C. As this 600-MHz spectrometer is not equipped with a cryo-probe, full decay curves at each applied CPMG refocusing field were collected to increase signal-to-noise. Specifically, as opposed to the standard intensity-based method to extract each R2 relaxation rate at each imparted field strength from a single 2D spectrum, R2 relaxation rates were determined by simultaneously arraying the number of imparted CPMG refocusing pulses together with the mixing time and directly determine each R2 relaxation rate. R1 relaxation rates were calculated in CCPNmr and R2-CPMG dispersions were fit to the full Carver–Richards equation using CPMG_fit as previously described [29]. Briefly, CPMG_fit performs a least squares analytical fit to the dispersion data using the generalized Carver–Richards equations, and exchange rates are only reported for those fits that have errors less than the fitted rates themselves (Table S1). Uncertainties for R2-CPMG dispersions were estimated from residues exhibiting no exchange (i.e., flat dispersion curves) and were 0.5–1.0 Hz. All R1 relaxation rates were collected on 15N-labeled proteins, R2-CPMG dispersions were collected on 2H,15N-labeled proteins, and assignments were collected on both 13C,15N-labeled and 2H,13C,15N-labeled proteins. ZZ-exchange was collected on samples containing 500 μM 15N-labeled enzyme and 250 μM NADP+, and all four peaks were fit to the full set of relaxation equations using GraphPad Prism version 4.0 (GraphPad Software Inc.), as we have previously described [50]. The only modifications to the ZZ exchange equations from our previous studies are that now the rates of exchange reflect the coenzyme off-rate (koff) and the coenzyme on-rate (kon), which have recently been described [25, 26]. Initial values for the exchange rates of koff and kon were set to reflect the KD values that we measured from ITC (KD=koff/kon).
NMR assignments
The NMR assignments for apo BLVRB and holo BLVRB have been deposited in the BioMagResBank, accession numbers 27462 and 27463, respectively.
Supplementary Material
Acknowledgments
E.Z.E. was supported by NIH application number 1RO1GM107262. B.G.M. is supported by NIH application number GM115308. A.D.S. is supported by the 2017 Webb-Waring Early Career Investigator award via the Boettcher Foundation. The National High Magnetic Field Laboratory is supported by the National Science Foundation through NSF/DMR-1644779 and the State of Florida.
Abbreviations used:
- BLVRB
biliverdin reductase B
- BLVRA
biliverdin reductase A
- FR
flavin reductase
- CSP
Chemical Shift Perturbation
- RBC
red blood cell
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jmb.2018.06.015.
References
- [1].Cunningham O, Gore MG, Mantle TJ, Initial-rate kinetics of the flavin reductase reaction catalysed by human biliverdin-IX beta reductase (BVR-B), Biochem. J 345 (2000) 393–399. [PMC free article] [PubMed] [Google Scholar]
- [2].Shalloe F, Elliott G, Ennis O, Mantle TJ, Evidence that biliverdin-IX beta reductase and flavin reductase are identical, Biochem. J 316 (1996) 385–387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Reisz JA, Chessler KM, Dzieciatkowska M, D'Alessandro A, Hansen CH, Blood and plasma proteomics: targeted quantitation and posttranslational redox modifications, Methods Mol. Biol 1619 (2017) 353–371. [DOI] [PubMed] [Google Scholar]
- [4].Kallberg Y, Oppermann U, Persson B, Classification of the short-chain dehydrogenase/reductase superfamily using hidden Markov models, FEBS J. 277 (2010) 2375–2386. [DOI] [PubMed] [Google Scholar]
- [5].Yamaguchi T, Komoda Y, Nakajima H, Biliverdin-IX-alpha reductase and biliverdin-IX-beta reductase from human liver— purification and characterization, J. Biol. Chem 269 (1994) 24343–24348. [PubMed] [Google Scholar]
- [6].Wu S, Li ZD, Gnatenko DV, Zhang BB, Zhao L, Malone LE, Markova N, Mantle TJ, Nesbitt NM, Bahou WF, BLVRB redox mutation defines heme degradation in a metabolic pathway of enhanced thrombopoiesis in humans, Blood 128 (2016) 699–709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [7].Bolton-Maggs PHB, Chalmers EA, Collins PW, Harrison P, Kitchen S, Liesner RJ, Minford A, Mumford AD, Parapia LA, Perry DJ, Watson SP, Wilde JT, Williams MD, A review of inherited platelet disorders with guidelines for their management on behalf of the UKHCDO, Br. J. Haematol 135 (2006) 603–633. [DOI] [PubMed] [Google Scholar]
- [8].Kistangari G, McCrae KR, Immune thrombocytopenia, Hematol. Oncol. Clin. North Am 27 (2013) 495 (+). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Piatek C, O'Connell CL, Liebman HA, Treating venous thromboembolism in patients with cancer, Expert. Rev. Hematol 5 (2012) 201–209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Liebman HA, Thrombocytopenia in cancer patients, Thromb. Res 133 (2014) S63–S69. [DOI] [PubMed] [Google Scholar]
- [11].Zhou P, Kalakonda N, Comenzo RL, Changes in gene expression profiles of multiple myeloma cells induced by arsenic trioxide (ATO): possible mechanisms to explain ATO resistance in vivo, Br. J. Haematol 128 (2005) 636–644. [DOI] [PubMed] [Google Scholar]
- [12].Zhang JT, Wang KJ, Zhang JZ, Liu SS, Dai LP, Zhang JY, Using proteomic approach to identify tumor-associated proteins as biomarkers in human esophageal squamous cell carcinoma, J. Proteome Res 10 (2011) 2863–2872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Pereira PJB, Macedo-Ribeiro S, Parraga A, Perez-Luque R, Cunningham O, Darcy K, Mantle TJ, Coll M, Structure of human biliverdin IX beta reductase, an early fetal bilirubin IX beta producing enzyme, Nat. Struct. Biol 8 (2001) 215–220. [DOI] [PubMed] [Google Scholar]
- [14].Fu G, Liu H, Doerksen RJ, Molecular modeling to provide insight into the substrate binding and catalytic mechanism of human biliverdin-IX alpha reductase, J. Phys. Chem. B 116 (2012) 9580–9594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Chu W-T, Nesbitt NM, Gnatenko DV, Li Z, Zhang B, Seeliger MA, Browne S, Mantle TJ, Bahou WF, Wang J, Enzymatic activity and thermodynamic stability of biliverdin IX beta reductase are maintained by an active site serine, Chem Eur J 23 (2017) 1891–1900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Smith LJ, Browne S, Mulholland AJ, Mantle TJ, Computational and experimental studies on the catalytic mechanism of biliverdin-IX beta reductase, Biochem. J 411 (2008) 475–484. [DOI] [PubMed] [Google Scholar]
- [17].Khavrutskii IV, Price DJ, Lee J, Brooks CL, Conformational change of the methionine 20 loop of Escherichia coli dihydrofolate reductase modulates pK(a) of the bound dihydrofolate, Protein Sci. 16 (2007) 1087–1100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Wan Q, Bennett BC, Wilson MA, Kovalevsky A, Langan P, Howell EE, Dealwis C, Toward resolving the catalytic mechanism of dihydrofolate reductase using neutron and ultrahigh-resolution X-ray crystallography, Proc. Natl. Acad. Sci. U. S. A 111 (2014) 18225–18230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [19].Venkitakrishnan RP, Zaborowski E, McElheny D, Benkovic SJ, Dyson HJ, Wright PE, Conformational changes in the active site loops of dihydrofolate reductase during the catalytic cycle, Biochemistry 43 (2004) 16046–16055. [DOI] [PubMed] [Google Scholar]
- [20].McElheny D, Schnell JR, Lansing JC, Dyson HJ, Wright PE, Defining the role of active-site loop fluctuations in dihydrofolate reductase catalysis, Proc. Natl. Acad. Sci. U. S. A 102 (2005) 5032–5037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Sender R, Fuchs S, Milo R, Revised estimates for the number of human and bacteria cells in the body, PLoS Biol. 14 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Wu JT, Wu LH, Knight JA, Stability of nadph—effect of various factors on the kinetics of degradation, Clin. Chem 32 (1986) 314–319. [PubMed] [Google Scholar]
- [23].Osborne MJ, Venkitakrishnan RP, Dyson HJ, Wright PE, Diagnostic chemical shift markers for loop conformation and substrate and cofactor binding in dihydrofolate reductase complexes, Protein Sci. 12 (2003) 2230–2238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Hafsa NE, Wishart DS, CSI 2.0: a significantly improved version of the Chemical Shift Index, J. Biomol. NMR 60 (2014) 131–146. [DOI] [PubMed] [Google Scholar]
- [25].Furukawa A, Konuma T, Yanaka S, Sugase K, Quantitative analysis of protein–ligand interactions by NMR, Prog. Nucl. Magn. Reson. Spectrosc 96 (2016) 47–57. [DOI] [PubMed] [Google Scholar]
- [26].Li Y, Altorelli NL, Bahna F, Honig B, Shapiro L, Palmer AG, Mechanism of E-cadherin dimerization probed by NMR relaxation dispersion, Proc. Natl. Acad. Sci. U. S. A 110 (2013) 16462–16467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Csermely P, Palotai R, Nussinov R, Induced fit, conformational selection and independent dynamic segments: an extended view of binding events, Trends Biochem. Sci 35 (2010) 539–546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].McDonald LR, Boyer JA, Lee AL, Segmental motions, not a two-state concerted switch, underlie allostery in CheY, Structure 20 (2012) 1363–1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Schlegel J, Armstrong GS, Redzic JS, Zhang FL, Eisenmesser EZ, Characterizing and controlling the inherent dynamics of cyclophilin-A, Protein Sci. 18 (2009) 811–824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [30].Cunningham O, Dunne A, Sabido P, Lightner D, Mantle TJ, Studies on the specificity of the tetrapyrrole substrate for human biliverdin-IX alpha reductase and biliverdin-IX beta reductase—structure–activity relationships define models for both active sites, J. Biol. Chem 275 (2000) 19009–19017. [DOI] [PubMed] [Google Scholar]
- [31].Miller GP, Benkovic SJ, Strength of an interloop hydrogen bond determines the kinetic pathway in catalysis by Escherichia coli dihydrofolate reductase, Biochemistry 37 (1998) 6336–6342. [DOI] [PubMed] [Google Scholar]
- [32].Farnum MF, Magde D, Howell EE, Hirai JT, Warren MS, Grimsley JK, Kraut J, Analysis of hydride transfer and cofactor fluorescence decay in mutants of dihydrofolate-reductase—possible evidence for participation of enzyme molecular motions in catalysis, Biochemistry 30 (1991) 11567–11579. [DOI] [PubMed] [Google Scholar]
- [33].Holliday M, Armstrong GS, Eisenmesser E, Determination of the full catalytic cycle among multiple cyclophilin family members and limitations on the application of CPMG-RD in reversible catalytic systems, Biochemistry 54 (2015) 5815–5827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [34].Boehr DD, McElheny D, Dyson HJ, Wright PE, The dynamic energy landscape of dihydrofolate reductase catalysis, Science 313 (2006) 1638–1642. [DOI] [PubMed] [Google Scholar]
- [35].Carroll MJ, Fau Mauldin Rv, Gromova AV, Fau Gromova Av, Singleton SF, Fau Singleton Sf, Collins EJ, Fau Collins Ej, Lee AL, Lee AL, Evidence for dynamics in proteins as a mechanism for ligand dissociation, Nat. Chem. Biol 8 (2012) 246–252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Eisenmesser E, Capodagli GC, Armstrong GS, Holliday M, Isern NG, Zhang F, Pegan SD, Inherent dynamics within the Crimean–Congo hemorrhagic fever virus protease are localized to the same region as substrate interactions, Protein Sci. 24 (2015) 651–660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [37].Torbeev VY, Raghuraman H, Hamelberg D, Tonelli M, Westler WM, Perozo E, Kent SBH, Protein conformational dynamics in the mechanism of HIV-1 protease catalysis, Proc. Natl. Acad. Sci. U. S. A 108 (2011) 20982–20987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Doshi U, Holliday MJ, Eisenmesser EZ, Hamelberg D, Dynamical network of residue-residue contacts reveals coupled allosteric effects in recognition, catalysis, and mutation, Proc. Natl. Acad. Sci. U. S. A 113 (2016) 4735–4740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [39].Lauren C McGowan, D. Hamelberg, Conformational plasticity of an enzyme during catalysis: intricate coupling between cyclophilin A dynamics and substrate turnover, Biophys. J 104 (2013) 216–226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Arora K, Brooks CL, Multiple intermediates, diverse conformations, and cooperative conformational changes underlie the catalytic hydride transfer reaction of dihydrofolate reductase, in: Klinman J, HammesSchiffer S (Eds.), Dynamics in Enzyme Catalysis, 337, 2013, pp. 165–187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Holliday M, Camilloni C, Armstrong GS, Vendruscolo C, Eisenmesser EZ, Networks of dynamic allostery regulate enzyme function, Structure 25 (2017) 1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Motlagh HN, Wrabl JO, Li J, Hilser VJ, The ensemble nature of allostery, Nature 508 (2014) 331–339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Goodey NM, Benkovic SJ, Allosteric regulation and catalysis emerge via a common route, Nat. Chem. Biol 4 (2008) 474–482. [DOI] [PubMed] [Google Scholar]
- [44].Li LY, Falzone CJ, Wright PE, Benkovic SJ, Functional-role of a mobile loop of Escherichia-coli dihydrofolate-reductase in transition-state stabilization, Biochemistry 31 (1992) 7826–7833. [DOI] [PubMed] [Google Scholar]
- [45].D'Alessandro A, Dzieciatkowska M, Nemkov T, Hansen KC, Red blood cell proteomics update: is there more to discover? Blood Transfus. 15 (2017) 182–187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [46].Reisz JA, Wither MJ, Dzieciatkowska M, Nemkov T, Issaian A, Yoshida T, Dunham AJ, Hill RC, Hansen KC, D'Alessandro A, Oxidative modifications of glyceraldehyde 3-phosphate dehydrogenase regulate metabolic reprogramming of stored red blood cells, Blood 128 (2016) E32–E42. [DOI] [PubMed] [Google Scholar]
- [47].Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A, Nmrpipe—a multidimensional spectral processing system based on UNIX pipes, J. Biomol. NMR 6 (1995) 277–293. [DOI] [PubMed] [Google Scholar]
- [48].Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas P, Ulrich EL, Markley JL, Ionides J, Laue ED, The CCPN data model for NMR spectroscopy: development of a software pipeline, Proteins: Struct. Funct. Bioinf 59 (2005) 687–696. [DOI] [PubMed] [Google Scholar]
- [49].Hyberts SG, Milbradt AG, Wagner AB, Arthanari H, Wagner G, Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling, J. Biomol. NMR 52 (2012) 315–327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Schlegel J, Redzic JS, Porter C, Yurchenko V, Bukrinsky M, Armstrong GS, Zhang FL, Isern NG, DeGregori J, Hodges R, Eisenmesser EZ, Solution characterization of the extracellular region of CD147 and its interaction with its enzyme ligand cyclophilin-A, J. Mol. Biol 391 (2009) 518–535. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.