Abstract
All living organisms contain a unique class of molecular chaperones called 60 kDa heat shock proteins (HSP60 – also known as GroEL in bacteria). While some organisms contain more than one HSP60 or GroEL isoform, at least one isoform has always proven to be essential. Because of this, we have been investigating targeting HSP60 and GroEL chaperonin systems as an antibiotic strategy. Our initial studies focused on applying this antibiotic strategy for treating African sleeping sickness (caused by Trypanosoma brucei parasites) and drug-resistant bacterial infections (in particular Methicillin-resistant Staphylococcus aureus – MRSA). Intriguingly, during our studies we found that three known antibiotics – suramin, closantel, and rafoxanide – were potent inhibitors of bacterial GroEL and human HSP60 chaperonin systems. These findings prompted us to explore what other approved drugs, natural products, and known bioactive molecules might also inhibit HSP60 and GroEL chaperonin systems. Initial high-throughput screening of 3,680 approved drugs, natural products, and known bioactives identified 161 hit inhibitors of the Escherichia coli GroEL chaperonin system (4.3% hit rate). From a purchased subset of 60 hits, 29 compounds (48%) re-confirmed as selective GroEL inhibitors in our assays, all of which were nearly equipotent against human HSP60. These findings illuminate the notion that targeting chaperonin systems might be a more common occurrence than we previously appreciated. Future studies are needed to determine if the in vivo modes of action of these approved drugs, natural products, and known bioactive molecules are related to GroEL and HSP60 inhibition.
Keywords: GroEL, GroES, HSP60, HSP10, molecular chaperone, chaperonin, proteostasis, small molecule inhibitors, natural products
Graphical Abstract

Molecular chaperones are a class of proteins that cells have developed to help fold polypeptides to their native states, or target them for degradation.1–7 The 60 kDa heat shock proteins (HSP60 – also known as GroEL in bacteria) are a unique class of the molecular chaperone family that function by encapsulating unfolded polypeptides in the central cavity of an HSP60 ring, allowing them to fold while sequestered from the cellular milieu.8–14 HSP60 chaperonins accomplish this in an ATP-dependent fashion with the assistance of co-chaperones called HSP10 (GroES in bacteria), which are “lid” structures that cap off folding-active HSP60 rings (hence, we typically refer to them as HSP60/10 and GroEL/ES chaperonin systems). While some organisms contain more than one HSP60 or GroEL isoform, one has always proven to be essential, at least in the micro-organisms thus far evaluated.15–33 Because of this, we hypothesize that these chaperonin systems are viable antibiotic targets. Since HSP60/10 and GroEL/ES chaperonin systems are highly conserved across all organisms, we envision that this could be a broad-spectrum antibiotic strategy. While human cells also contain an HSP60/10 homolog, located in mitochondria, our accumulating evidence supports that even if compounds can inhibit HSP60/10 in vitro, many of these inhibitors are non-toxic to human cells and in vivo.34–37 No drugs have been developed to specifically inhibit HSP60/10 or GroEL/ES chaperonins, and thus we believe that targeting these molecular machines offers significant promise to address the problem of antibiotic-resistant infectious organisms.
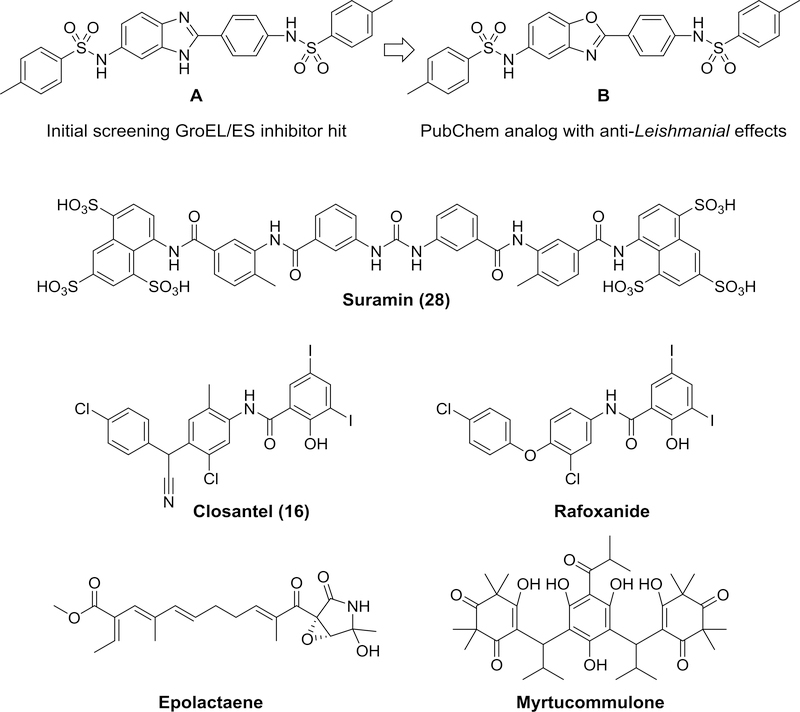
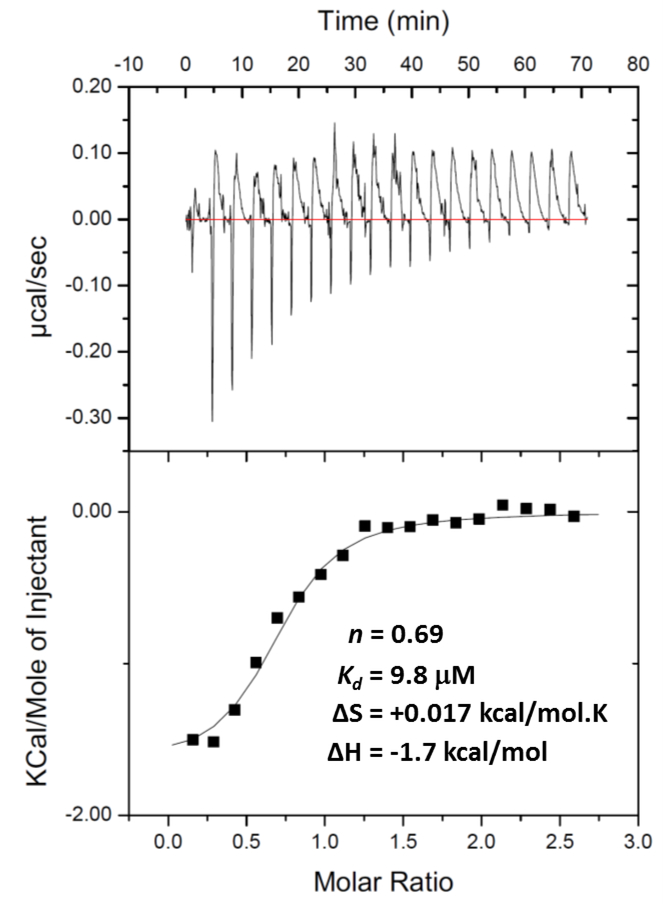
As a first step in our research, we previously conducted a high-throughput screen of ~700,000 small molecules and identified 235 inhibitors of the prototypical GroEL/ES chaperonin system from Escherichia coli.38 One lead hit, the bis-sulfonamido-2-arylbenzimidazole compound A (Figure 1), was isostructural to another molecule identified in the PubChem database (compound B), which was reported to inhibit the growth of Leishmania parasites.39 As an extension, we developed a library of compound B analogs and screened them against Trypanosoma brucei, a related parasite that causes African sleeping sickness.35 Surprisingly, in that study we found that suramin (28), the first-line treatment for T. brucei infections in humans, was capable of inhibiting Escherichia coli GroEL/ES (which we typically use as a surrogate for compound evaluation) as well as human HSP60/10. To further support that this interaction was real, in the present study, we analyzed the suramin-GroEL binding properties using Isothermal Titration Calorimetry (see the Supporting Information for a detailed protocol for this experiment). An isotherm for a representative suramin-GroEL binding analysis is presented in Figure 2, with the thermodynamic parameters, binding affinities, and binding stoichiometries averaged from triplicate analyses presented in Table 1. We found that suramin had a Kd of 21 μM for binding to E. coli GroEL, which corresponds reasonably well with the IC50 values for suramin inhibition in our assays that monitor GroEL/ES-mediated refolding of dMDH and dRho. While this analysis shows the suramin-GroEL interaction is indeed real, what remains to be seen is what contribution suramin’s binding to the three HSP60s in T. brucei make to its anti-trypanosomal effects. Future studies will need to explore this, but are beyond the scope of the present study.
Figure 1.
Structures of compounds previously found to inhibit E. coli GroEL/ES and/or human HSP60/10 chaperonin systems.
Figure 2.
Representative analysis of the binding of suramin (28) to E. coli GroEL measured by Isothermal Titration Calorimetry (ITC). The top panel shows a representative binding isotherm obtained by titrating suramin (2 mM) into a solution of GroEL (150 μM monomer concentration) in the ITC cell. The lower panel shows the integrated data (solid squares) fit to a single-site binding model (solid line). The molar ratio refers to the binding stoichiometry of suramin to monomeric GroEL. Average results for the various binding parameters (Kd, n, ΔH, ΔS, and ΔG) obtained from triplicate analyses are presented in Table 1.
Table 1.
IC50 values for compounds tested in our GroEL/ES-mediated substrate enzyme refolding assays, the native enzymatic reporter counter screens (vide infra), and ITC analytical results for suramin (28) binding to E. coli GroEL. Binding parameter results are averaged from three replicate analyses. Binding is predominantly entropically driven, with a moderate enthalpic contribution to affinity. Particularly interesting is that suramin binds with a stoichiometry of ~9 molecules per GroEL tetradecamer and, since suramin does not inhibit GroEL ATPase activity, it is likely binding to unknown sites outside of the ATP pockets.
| Suramin (28) | |
|---|---|
| Assay IC50 (μM) | |
| GroEL/ES-dMDH Refolding | 8.1 |
| GroEL/ES-dRho Refolding | 16 |
| Native MDH reporter activity | >63 |
| Native Rho reporter activity | >100 |
| Stoichiometry n | |
| Molecule : GroELmonomer | 0.67 ± 0.05 |
| Molecule : GroELoligomer | 9.4 ± 0.6 |
| Thermodynamic parameters | |
| Log(K d /μM) | 1.31 ± 0.30 |
| K d (μM) | 21 |
| ΔH (kcal/mol) | −2.07 ± 0.36 |
| ΔS (kcal/mol·K) | 0.014 ± 0.002 |
| TΔS (kcal/mol) | 4.15 ± 0.72 |
| ΔG (kcal/mol) | −6.22 ± 0.41 |
In three additional follow up studies to our high-throughput screen, we explored the antibacterial properties of a subset of 22 of our hit GroEL inhibitors, plus additional compound B analogs and a series of analogs based on a bisarylamide hit-to-lead scaffold.34, 36, 37 During these studies, we discovered that two known anthelmintics used in veterinary medicine, closantel and rafoxanide, were also potent GroEL/ES and HSP60/10 inhibitors.37, 40 In addition, other groups have identified a handful of natural products, such as epolactaene and myrtucommulone, that were able to target the human HSP60/10 chaperonin system.41–44 Taken together, these accumulating findings prompted us to consider the possibility that targeting chaperonin systems with small molecule inhibitors may be more common than we previously thought. To shed further light on this possibility, we designed the present study to identify what other approved drugs, natural products, or known bioactive molecules might also inhibit HSP60/10 and/or GroEL/ES chaperonin systems.
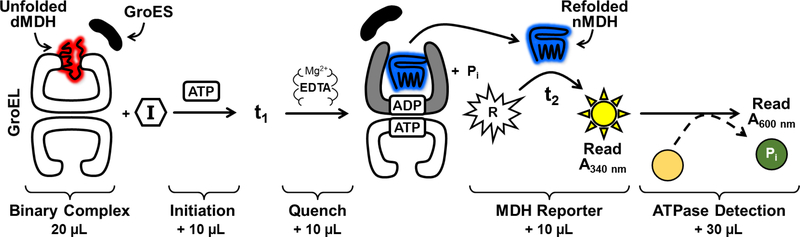
In this study, we screened against the Library of Pharmaceutically Active Compounds (LOPAC) and the MicroSource Spectrum libraries, which together contain 3,680 approved drugs, natural products, and known bioactive molecules. For the primary high-throughput screen, we developed a new protocol that combined our traditional GroEL/ES-dMDH refolding and chaperonin-mediated ATPase assays into one multiplexed assay. A schematic representation of this multiplexed assay is presented in Figure 3, with a detailed description of the protocol provided in the Supporting Information. Using this assay, we screened the 3,680 compound library and identified 219 compounds that inhibited >50% of the reporter enzymatic reaction, and thus putatively the refolding of the MDH reporter enzyme by the GroEL/ES chaperonin system. We found that none of the hits reduced ATPase activity, which was perhaps not surprising since in our previous high-throughput screen of ~700,000 compounds, only a handful of hits were able to inhibit ATP hydrolysis by GroEL.38 We next conducted a secondary high-throughput screen of the entire 3,680 compounds using our traditional GroEL/ES-dRho refolding assay, which employs rhodanese (Rho) as the reporter enzyme to be refolded by the chaperonin system.34, 36, 37 A detailed description of the protocol for this assay is presented in the Supporting Information. Since MDH and Rho have orthogonal enzymatic reactions (i.e. MDH catalyzes the oxidation of NADH in the presence of mesoxilic acid, while Rho catalyzes the conversion of cyanide to thiocyanate – two very different enzymatic reactions), we have typically found that these two refolding assays are effective at removing false-positive compounds that inhibit the refolded reporter enzymes. Percent inhibition results for these two assays are presented in Table S2 in the Supporting Information. While suramin was within the 3,680 compound library, we found that it did not meet the 50% inhibition cutoff in both of these assays, likely because the compound concentrations in the screens were near its IC50 values for inhibiting GroEL/ES refolding functions (~10–15 μM). While this suggests that if we tested the libraries at higher concentrations, or relaxed our inhibition cutoffs, we may find even more hits, we refrained from doing so as we had already obtained a high hit rate of 4.3% for compounds exhibiting >50% in both of the screens.
Figure 3. Protocol for the primary multiplexed high-throughput screening assay.
For primary screening of the LOPAC and MicroSource Spectrum libraries, we employed a new assay where we combined our individual GroEL/ES-dMDH refolding and GroEL/ES-dMDH ATPase assays into one multiplexed format. In this assay, a solution containing GroES and a binary complex of denatured malate dehydrogenase (dMDH) bound to GroEL was dispensed into the wells of a 384-well microplate. Compounds from the LOPAC and MicroSource Spectrum libraries (single concentrations) were then pin-transferred into the wells. The chaperonin-mediated refolding cycle was initiated by addition of ATP, the plates were incubated at 37°C for ~30 minutes (t1 – until ~90% of the dMDH would have been refolded in the absence of inhibitors), and EDTA was then added to quench the refolding cycle. The substrates for the refolded, native MDH (nMDH) were added (sodium mesoxalate and NADH) and the enzymatic reporter reaction was monitored over time by reading well absorbance at 340 nm (t2 – until the DMSO control wells had reached ~90% conversion of NADH to NAD+). In this coupled assay, the extent of chaperonin inhibition is proportional to the amount of enzymatic activity, and thus refolded MDH, present. In the same plate, we then added the malachite green phosphate reporter reagents to evaluate chaperonin-mediated hydrolysis of ATP.
Upon examination of the 161 hits, we found a wide range of molecular structures, from as small as the single aryl mesalamine (23) to large, macrocyclic natural products like thiostrepton (24) and ivermectin (31). While singleton hits were identified, several hits were found that fell into distinct scaffold clusters, including analogs of suramin (1, 22, 28), chalcones (35, 43, 44), ivermectin (31–33), agaric acid (7, 8, 11, 20, 25, 27, 39, 42), and porphyrins (5, 19, 36). To further verify that hits were not false-positives owing to potential abnormalities in the library parent stocks or compounds that inhibit both the native MDH and Rho reporter enzymes, we selected a set of 60 hits to purchase purified powders of for confirmatory screening in our wider panel of established chaperonin-mediated biochemical assays. The structures of these hits are presented in Figure S1 in the Supporting Information, which were selected to maintain structural diversity while still probing some of the scaffold clusters. These 60 purchased compounds were re-screened in dose-response format in both of our traditional GroEL/ES-dMDH and GroEL/ES-dRho refolding assays, counter-screened in assays that monitor for false-positive inhibition of the refolded native MDH or Rho enzymatic reactions, and evaluated for inhibition of GroEL-mediated ATPase activity. Please refer to the Supporting Information for detailed protocols of these assays, and Table 2 for a compilation of IC50 values obtained in these assays. Consistent with results from the primary high-throughput screen, none of the compounds showed appreciable inhibition of GroEL-mediated ATPase activity. However, from the GroEL/ES-mediated refolding assays and native MDH and Rho reporter counter-screens, we were able to classify the 60 hits into four distinct inhibitor categories in order of decreasing selectivity for the GroEL/ES chaperonin system (Table 2): 12 compounds (20%) had >10x selectivity in both of the refolding assays (blue classification); 17 compounds (28%) exhibited >10x selectivity in one of the refolding assays, but not the other (green classification); 19 compounds (32%) exhibited between 1–10x selectivity in both of the refolding assays (yellow classification); and 12 compounds (20%) generally did not re-confirm as GroEL/ES inhibitors (orange classification).
Table 2.
Compilation of IC50 values from the respective biochemical assays. Compounds are ordered from most selective in the refolding assays at the top, to lower selectivity going down the table. Compounds colored blue exhibit >10x selectivity in both of the GroEL/ES-mediated refolding assays compared to the native MDH and Rho enzymatic reporter counter screens; compounds colored green exhibit >10x selectivity in one of the refolding assays, but not the other; compounds colored yellow exhibit between 1–10x selectivity in both of the refolding assays; and compounds colored orange generally did not re-confirm as GroEL/ES inhibitors.
| Biochemical Assay IC50 (0μM) | |||||||
|---|---|---|---|---|---|---|---|
| Compound | Native Rho | Native MDH | GroEL/ES-mediated refolding of |
HSP60/10-dMDH | GroEL | ||
| # | Name | Reporter | Reporter | dRho | dMDH | Refolding | ATPase |
| 1 | Evans blue | 8.7 | 19 | 0.087 | 0.039 | 0.17 | 161 |
| 2 | (±)-Gossypol-acetic acid | 11 | 48 | 0.43 | 0.66 | 1.2 | >250 |
| 3 | Adapalene | >100 | >63 | 5.1 | 2.1 | 2.3 | >250 |
| 4 | Curcumin | 56 | >63 | 2.9 | 3.1 | 8.3 | >250 |
| 5 | Protoporphyrin IX | >100 | >63 | 3.0 | 4.4 | 4.9 | >250 |
| 6 | Ethacrynic acid | >100 | >63 | 4.7 | 5.2 | 8.5 | >250 |
| 7 | Agaric acid | >100 | >63 | 8.9 | 3.5 | 4.0 | >250 |
| 8 | Cetylpyridinium chloride | >100 | >63 | 8.9 | 4.2 | 6.0 | 152 |
| 9 | Epigallocatechin gallate (EGCG) | 7.3 | 16 | 0.58 | 0.36 | 2.3 | >250 |
| 10 | Hexachlorophene | 51 | 18 | 4.7 | 1.4 | 1.3 | 153 |
| 11 | Hexadecyltrimethylammonium bromide | >100 | >63 | 12 | 6.5 | 7.4 | 210 |
| 12 | Hematein | 12 | 46 | 1.2 | 1.8 | 3.6 | >250 |
| Spearman Correlation Coefficient & p value | 0.746 (p = 0.0071) | ||||||
| 0.853 (p = 0.0008) | |||||||
| 13 | Aurintricarboxylic acid | >100 | 3.0 | 0.57 | 0.90 | 3.2 | >250 |
| 14 | Erythrosin B | 9.2 | 33 | 1.2 | 0.50 | 2.9 | 222 |
| 15 | β-laphachone | 0.12 | >63 | 0.59 | 1.3 | 2.1 | >250 |
| 16 | Closantel | >100 | 6.2 | 2.0 | 2.2 | 2.8 | >250 |
| 17 | Tannic acid | 39 | 4.6 | 1.0 | 0.87 | 3.8 | 137 |
| 18 | Gossypetin | 0.33 | >63 | 0.39 | 2.1 | 7.5 | >250 |
| 19 | Chlorophyllin sodium copper salt | 11 | >63 | 1.2 | 2.3 | 4.4 | >250 |
| 20 | L-Ascorbyl palmitate | 53 | >63 | 8.5 | 3.5 | 3.1 | 177 |
| 21 | Plumbagin | 0.060 | >63 | 0.29 | 3.8 | 6.3 | >250 |
| 22 | Pontamine sky blue | 8.5 | >63 | 2.0 | 3.9 | 6.8 | >250 |
| 23 | Mesalamine | 0.28 | >63 | 0.49 | 6.0 | 12 | >250 |
| 24 | Thiostrepton | >100 | >63 | 16 | 6.8 | 13 | >250 |
| 25 | Retinoic acid | 91 | >63 | 21 | 6.7 | 5.3 | >250 |
| 26 | Morin hydrate | 57 | 47 | 5.2 | 8.5 | 11 | >250 |
| 27 | Arachidonic acid | >100 | >63 | 18 | 7.3 | 4.5 | >250 |
| 28 | Suramin | >100 | >63 | 16 | 8.1 | 7.2 | >250 |
| 29 | Bithionol | >100 | 24 | 13 | 4.3 | 2.8 | >250 |
| Spearman Correlation Coefficient & p value | 0.556 (p = 0.0223) | ||||||
| 0.595 (p = 0.0132) | |||||||
| 30 | Menadione | 1.0 | >63 | 2.5 | 9.0 | 15 | >250 |
| 31 | Ivermectin | >100 | >63 | 23 | 9.5 | 16 | >250 |
| 32 | Abamectin (Avermectin b1a) | >100 | >63 | 32 | 9.8 | 15 | >250 |
| 33 | Eprinomectin | >100 | >63 | 38 | 11 | 17 | >250 |
| 34 | Crystal violet | 16 | >63 | 15 | 11 | 5.8 | >250 |
| 35 | 4’-Methoxychalcone | >100 | >63 | 30 | 13 | 16 | >250 |
| 36 | Cyanocobalamin (vtamin B12) | 0.66 | >63 | 0.12 | 16 | 12 | >250 |
| 37 | Purpurin | 2.3 | >63 | 4.0 | 15 | 23 | >250 |
| 38 | Lithocholic acid | >100 | >63 | 59 | 16 | 10 | >250 |
| 39 | Benzethonium chloride | >100 | >63 | 26 | 24 | 23 | >250 |
| 40 | Benzbromarone | >100 | >63 | 49 | 18 | 9.1 | >250 |
| 41 | Cholecalciferol (vtamin D3) | >100 | >63 | 31 | 22 | 11 | >250 |
| 42 | 4-Nonylphenol | 72 | 47 | 29 | 12 | 9.5 | >250 |
| 43 | 4-Hydroxychalcone | >100 | >63 | 40 | 21 | 22 | >250 |
| 44 | 4’-Hydroxychalcone | >100 | >63 | 56 | 21 | 20 | >250 |
| 45 | Dichlorophen | >100 | >63 | 66 | 24 | 17 | >250 |
| 46 | Calciferol (Vtamin D2) | 94 | >63 | 41 | 28 | 45 | >250 |
| 47 | Phenoxybenzamine HCl | >100 | >63 | 129 | 40 | 78 | >250 |
| 48 | Paclitaxel | >100 | >63 | 114 | 56 | >100 | >250 |
| Spearman Correlation Coefficient & p value | 0.601 (p = 0.0064) | ||||||
| 0.626 (p = 0.0042) | |||||||
| 49 | Primiquine | 2.9 | >63 | 6.2 | 75 | >100 | 103 |
| 50 | Riboflavin-5-phosphate sodium salt | 0.38 | >63 | 0.77 | 85 | 80 | >250 |
| 51 | Ethidium bromide | 0.83 | >63 | 2.4 | >100 | >100 | >250 |
| 52 | Methylene blue trihydrate | 6.1 | >63 | 3.3 | >100 | >100 | >250 |
| 53 | Riboflavin | 8.8 | >63 | 18 | >100 | >100 | >250 |
| 54 | Harmalol | 9.7 | >63 | 16 | >100 | >100 | >250 |
| 55 | (−)-Erythromycin | >100 | >63 | 178 | >100 | >100 | >250 |
| 56 | Candesartan | >100 | >63 | >250 | >100 | >100 | >250 |
| 57 | Albuterol | >100 | >63 | >250 | >100 | >100 | >250 |
| 58 | Nalbuphine | >100 | >63 | >250 | >100 | >100 | >250 |
| 59 | SB 204070 | >100 | >63 | >250 | >100 | >100 | >250 |
| 60 | Fluvastatin sodium | >100 | >63 | >250 | >100 | >100 | >250 |
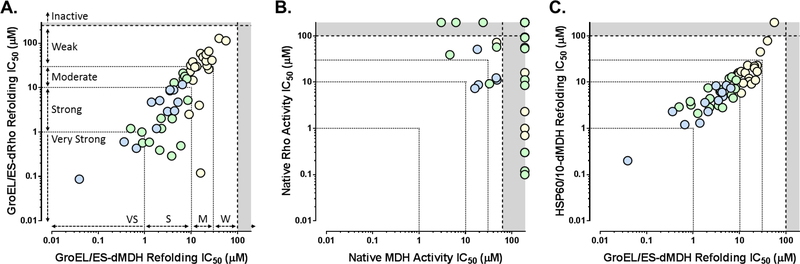
Upon further analysis of the biochemical results, for the highly selective blue series molecules, we see a strong correlation between IC50 values obtained from the GroEL/ES-dMDH and -dRho refolding assays (Figure 4A), with a Spearman correlation coefficient of 0.746 (p = 0.0071). While the green and yellow series are less selective, each still have moderate correlations between their IC50 values obtained in the two refolding assays, with Spearman correlation coefficients of 0.556 (p = 0.0223) and 0.601 (p = 0.0064), respectively. Few of these molecules, however, inhibited in both the native MDH and Rho reporter counter-screens (Figure 4B), suggesting they are likely still on target for inhibiting the GroEL/ES chaperonin system.
Figure 4.
Correlation plots of IC50 values for compounds tested in the respective biochemical assays. Each data point represents results for individual compounds tested in the respective assays, with color coding of points corresponding to the selectivity classifications of compound results presented in Table 2. Compounds inhibited nearly equipotently in both the GroEL/ES-dMDH and GroEL/ES-dRho refolding assays (panel A, Spearman correlation coefficients presented in Table 2), with few that inhibited both the native MDH and Rho reporter counter-screens (panel B), supporting on-target effects against the chaperonin-mediated refolding cycle. Compounds inhibited the human HSP60/10 and E. coli GroEL/ES chaperonin systems nearly equipotently, suggesting binding sites may be highly conserved between the two (panel C, Spearman correlation coefficients presented in Table 2). As indicated in panel A, For the purposes of categorizing inhibitor potencies in the various biochemical assays, we consider compounds with IC50 values plotted in the grey zones to be inactive (i.e. greater than the maximum concentrations tested), >30 μM to be weak inhibitors, 10–30 μM moderate inhibitors, 1–10 μM potent inhibitors, and <1 μM very potent and acting near stoichiometrically since the concentration of GroEL tetradecamer is 50 nM during the refolding cycle (i.e. 700 nM GroEL monomeric subunits).
As an extension on potential selectivity concerns, because human HSP60 shares high homology to bacterial E. coli GroEL (~48% sequence identity), we counter-screened compounds 1–60 in an analogous HSP60/10-dMDH refolding assay. As seen in Figure 4C, there is a high correlation for compounds inhibiting both E. coli GroEL/ES and human HSP60/10: Spearman correlation coefficients are 0.853 (p = 0.0008) for the blue series, 0.595 (p = 0.0132) for the green series, and 0.626 (p = 0.0042) for the yellow series compounds. These high correlations were not surprising based on results we have reported for other inhibitors, where we have only found a few scaffolds that can selectively inhibit E. coli GroEL/ES.34–37 While this raises the possibility of potential HSP60-dependent cytotoxicity to human cells, our ongoing studies support the notion that even though compounds can inhibit human HSP60/10 in vitro, this does not necessarily translate to cytotoxicity in vivo and would not preclude an inhibitor series from further development as an antibiotic candidate. For example, suramin inhibits human HSP60/10 in vitro, yet has been safely used for over 100 years as a first-line therapeutic for treating African sleeping sickness. A lack of cytotoxicity in vivo despite inhibition of HSP60/10 in vitro could be explained by the fact that compounds would need to be cell permeable and penetrate the highly impermeable mitochondrial membrane to even interact with HSP60/10 in the mitochondrial matrix. For chaperonin inhibitors that do affect the viability of human cells, whether they are natural products, synthetically derived, or even approved drugs or not, it will be important for future studies to determine whether or not their cytotoxicities are HSP60-dependent. As a first-pass indicator of general cellular toxicity, we employ Alamar Blue-based cell viability assays with a small panel of human cell lines from different tissues.
To further inform on the possibility that some of these compounds may exhibit promiscuous and pan-assay interference effects, we searched the PubChem database to see how many assays each of these compounds has been screened in, and the number of assays in which they have been classified as active hits.39 We have compiled these results in Table S1 in the Supporting Information. While some hits have only been reported tested in a handful of assays, most have been tested in hundreds, or even thousands, of assays. To more easily assimilate the data, in Figure S2A and S2B, we binned the aggregate results of the percentage assays that each molecule was reported active hits in, compared to their total number of reported assays. For example, 38.3% of compounds 1–60, and 37.4% of all hit compounds, were reported to be active in 0–10% of the assays (i.e. least promiscuous) they were tested in, as reported in the PubChem Database. The nearly identical distribution of the two series may indicate the compounds purchased for confirmatory evaluation (1–60) are a suitable subset that is representative of the complete set of 161 GroEL/ES inhibitor hits identified from the initial screens. We further compared the PubChem % Actives results for compounds 1–60 to their IC50 values from the GroEL/ES refolding assays to see if there was any evidence of more potent GroEL/ES inhibitors being inherently selective or promiscuous; however, we did not observe any correlation (Figure S2C). We exercise a word of caution in over-interpreting these results, though, as we have not further dissected the actual assays that each compound was tested in – for example, it would actually be beneficial if compounds were tested and reported active in a high proportion of infectious organism proliferation assays. With some compounds being tested in hundreds to thousands of assays, such a detailed analysis would be best suited for future studies of individual compounds. As an overview, we have compiled a brief listing of reported bioactivities for each of the 161 hits in Table S1 in the Supporting Information. While the bioactivities presented are far from exhaustive, they begin to paint a picture that targeting chaperonin systems could play a significant role in a number of indications, including inflammation, autoimmunity, cancer, and, in the context of the present study, antibiotic applications.
Through our own ongoing studies, we continue to accumulate evidence that targeting the GroEL/ES or HSP60/10 chaperonin systems of infectious organisms is a promising strategy for antibiotic development.34–37 We have identified chaperonin system inhibitors that are cytocidal to T. brucei parasites and Gram-positive bacteria, even towards S. aureus bacteria that reside in already established biofilms. As discussed above, despite having the ability to inhibit human HSP60/10 in vitro, many of our most potent GroEL/ES inhibitors exhibit low-to-no cytotoxicity to human cells in culture. Intriguingly, the present study screening the LOPAC and MicroSource Spectrum libraries helps to further substantiate these findings as 30% of the 161 GroEL/ES hit inhibitors are reported to have antimicrobial effects against a variety of pathogens, ranging from in vitro observations to practical use in household surface disinfectants and approved broad-spectrum antibiotics used clinically. Examples of these in the identified GroEL/ES and HSP60/10 inhibitor hits include suramin, ivermectin, retinoic acid, adapalene, cetylpyridinium chloride, abamectin, eprinomectin, benzethonium chloride, dichlorophen, curcumin, hexachlorophene, and hexadecyltrimethylammonium bromide. For some of these compounds, like suramin, definitive mechanisms of action have yet to be elucidated.45–51 Given the results presented herein, it is possible that some of these compounds may owe their antimicrobial effects to inhibition of the GroEL/ES and HSP60/10 chaperonin systems of the infecting pathogens.
In summary, in the present study, we screened a library of 3,680 known drugs, natural products, and bioactive compounds for their ability to inhibit the prototypical GroEL/ES chaperonin system from E. coli. The premise for this study was rooted in accumulating evidence, from us and others, indicating that it might not be uncommon for known drugs and natural products to target GroEL/ES and HSP60/10 chaperonin systems. From our two high-throughput screens, we obtained an unusually high hit rate of 4.3%, from which we purchased purified powders of a subset of 60 hits for confirmatory evaluation in a panel of our chaperonin-mediated biochemical assays. Of this 60 hit subset, 29 compounds (49%) reconfirmed as GroEL/ES inhibitors with >10x selectivity in at least one of our chaperonin-mediated refolding assays over our native MDH or Rho enzymatic reporter counter-screens, supporting that hits were on-target. With such a high hit and re-confirmation rate, this study suggests that targeting chaperonin systems might be a more common occurrence than we previously appreciated. These findings further incentivize broader screening of additional known drugs, clinical candidates, and natural product libraries and extracts to identify more compounds that may be functioning against chaperonin systems. Intriguingly, we found that 30% of the hits were reported to have antibiotic properties, thus further supporting the viability of a chaperonin-targeting antibiotic strategy; however, the other 70% have other reported bioactivities, including effects on inflammation, autoimmunity, and cancer, to name a few. While this may not be surprising as we found that most compounds were nearly equipotent inhibitors of both the human HSP60/10 and E. coli GroEL/ES chaperonin systems, a new question emerges: how much, if any, can the bioactivities of these known drugs and natural products be attributed to targeting GroEL/ES and HSP60/10 chaperonin systems?
Supplementary Material
Acknowledgments:
Research reported in this publication was supported by the National Institute of General Medical Sciences (NIGMS) of the National Institutes of Health (NIH) under Award Number R01GM120350. QQH and YP additionally acknowledge support by NIH grants 5R01GM111639 and 5R01GM115844. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. This work was also supported in part by startup funds from the IU School of Medicine (SMJ) and the University of Arizona (EC). The human HSP60 expression plasmid (lacking the 26 amino acid N-terminal mitochondrial signal peptide) was generously donated by Dr. Abdussalam Azem from Tel Aviv University, Faculty of Life Sciences, Department of Biochemistry, Israel.
Supporting Information: Supporting information associated with this article can be found in the online version, which includes tabulations of initial % inhibition results from the multiplexed GroEL/ES-dMDH refolding/ATPase and GroEL/ES-dRho high-throughput screens; PubChem assay results and bioactivities of the 161 inhibitor hits; structures of the purchased subset of hits 1–60; log(IC50) results with standard deviations for follow-up testing of purchased compounds; analysis of inhibitor assay bioactivity results reported in the PubChem database; and experimental protocols for biophysical and biochemical experiments.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Hartl FU; Bracher A; Hayer-Hartl M Molecular chaperones in protein folding and proteostasis. Nature 2011, 475, 324–332. [DOI] [PubMed] [Google Scholar]
- 2.Stefani M; Dobson CM Protein aggregation and aggregate toxicity: new insights into protein folding, misfolding diseases and biological evolution. J. Mol. Med. (Berl.) 2003, 81, 678–699. [DOI] [PubMed] [Google Scholar]
- 3.Maisonneuve E; Ezraty B; Dukan S Protein aggregates: An aging factor involved in cell death. J. Bacter 2008, 190, 6070–6075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Carmichael J; Chatellier J; Woolfson A; Milstein C; Fersht AR; Rubinsztein DC Bacterial and yeast chaperones reduce both aggregate formation and cell death in mammalian cell models of Huntington’s disease. Proc. Natl. Acad. Sci. U.S.A 2000, 97, 9701–9705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bao YP; Cook LJ; O’Donovan D; Uyama E; Rubinsztein DC Mammalian, yeast, bacterial, and chemical chaperones reduce aggregate formation and death in a cell model of oculopharyngeal muscular dystrophy. J. Biol. Chem 2002, 277, 12263–12269. [DOI] [PubMed] [Google Scholar]
- 6.Wong P; Houry WA Chaperone networks in bacteria: analysis of protein homeostasis in minimal cells. J. Struct. Biol 2004, 146, 79–89. [DOI] [PubMed] [Google Scholar]
- 7.Mogk A; Huber D; Bukau B Integrating protein homeostasis strategies in prokaryotes. Cold Spring Harb. Perspect. Biol 2011, 3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Braig K; Otwinowski Z; Hegde R; Boisvert DC; Joachimiak A; Horwich AL; Sigler PB The crystal structure of the bacterial chaperonin GroEL at 2.8 A. Nature 1994, 371, 578–586. [DOI] [PubMed] [Google Scholar]
- 9.Sigler PB; Xu Z; Rye HS; Burston SG; Fenton WA; Horwich AL Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem 1998, 67, 581–608. [DOI] [PubMed] [Google Scholar]
- 10.Horwich AL; Farr GW; Fenton WA GroEL-GroES-mediated protein folding. Chem. Rev 2006, 106, 1917–1930. [DOI] [PubMed] [Google Scholar]
- 11.Fenton WA; Kashi Y; Furtak K; Horwich AL Residues in chaperonin GroEL required for polypeptide binding and release. Nature 1994, 371, 614–619. [DOI] [PubMed] [Google Scholar]
- 12.Fenton WA; Horwich AL GroEL-mediated protein folding. Protein Sci. 1997, 6, 743–760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Horwich AL; Fenton WA; Chapman E; Farr GW Two families of chaperonin: physiology and mechanism. Annu. Rev. Cell. Dev. Biol 2007, 23, 115–145. [DOI] [PubMed] [Google Scholar]
- 14.Saibil HR; Fenton WA; Clare DK; Horwich AL Structure and allostery of the chaperonin GroEL. J. Mol. Biol 2013, 425, 1476–1487. [DOI] [PubMed] [Google Scholar]
- 15.Chapman E; Farr GW; Usaite R; Furtak K; Fenton WA; Chaudhuri TK; Hondorp ER; Matthews RG; Wolf SG; Yates JR; Pypaert M; Horwich AL Global aggregation of newly translated proteins in an Escherichia coli strain deficient of the chaperonin GroEL. Proc. Natl. Acad. Sci. U.S.A 2006, 103, 15800–15805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fayet O; Ziegelhoffer T; Georgopoulos C The groES and groEL heat shock gene products of Escherichia coli are essential for bacterial growth at all temperatures. J. Bacteriol 1989, 171, 1379–1385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Neckers L; Tatu U Molecular Chaperones in Pathogen Virulence: Emerging New Targets for Therapy. Cell Host & Microbe 2008, 4, 519–527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Alsford S; Turner DJ; Obado SO; Sanchez-Flores A; Glover L; Berriman M; Hertz-Fowler C; Horn D High-throughput phenotyping using parallel sequencing of RNA interference targets in the African trypanosome. Genome Research 2011, 21, 915–924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ojha A; Anand M; Bhatt A; Kremer L; Jacobs WR Jr.; Hatfull GF GroEL1: a dedicated chaperone involved in mycolic acid biosynthesis during biofilm formation in mycobacteria. Cell 2005, 123, 861–873. [DOI] [PubMed] [Google Scholar]
- 20.Hu YM; Henderson B; Lund PA; Tormay P; Ahmed MT; Gurcha SS; Besra GS; Coates ARM A Mycobacterium tuberculosis mutant lacking the groEL homologue cpn60.1 is viable but fails to induce an inflammatory response in animal models of infection. Infection and Immunity 2008, 76, 1535–1546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhang XB; Cui JA; Nilsson D; Gunasekera K; Chanfon A; Song XF; Wang HN; Xu Y; Ochsenreiter T The Trypanosoma brucei MitoCarta and its regulation and splicing pattern during development. Nucleic Acids Research 2010, 38, 7378–7387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kong TH; Coates ARM; Butcher PD; Hickman CJ; Shinnick TM Mycobacterium-Tuberculosis Expresses 2 Chaperonin-60 Homologs. Proceedings of the National Academy of Sciences of the United States of America 1993, 90, 2608–2612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lund PA Microbial molecular chaperones. Advances in Microbial Physiology, Vol 44 2001, 44, 93–140. [DOI] [PubMed] [Google Scholar]
- 24.Kumar CMS; Mande SC; Mahajan G Multiple chaperonins in bacteria-novel functions and non-canonical behaviors. Cell Stress & Chaperones 2015, 20, 555–574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rao T; Lund PA Differential expression of the multiple chaperonins of Mycobacterium smegmatis. Fems Microbiology Letters 2010, 310, 24–31. [DOI] [PubMed] [Google Scholar]
- 26.Folgueira C; Requena JM A postgenomic view of the heat shock proteins in kinetoplastids. Fems Microbiology Reviews 2007, 31, 359–377. [DOI] [PubMed] [Google Scholar]
- 27.Alsford S; Turner DJ; Obado SO; Sanchez-Flores A; Glover L; Berriman M; Hertz-Fowler C; Horn D High-throughput phenotyping using parallel sequencing of RNA interference targets in the African trypanosome. Genome Res 2011, 21, 915–924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang X; Cui J; Nilsson D; Gunasekera K; Chanfon A; Song X; Wang H; Xu Y; Ochsenreiter T The Trypanosoma brucei MitoCarta and its regulation and splicing pattern during development. Nucleic Acids Res 2010, 38, 7378–7387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bringaud F; Peyruchaud S; Baltz D; Giroud C; Simpson L; Baltz T Molecular characterization of the mitochondrial heat shock protein 60 gene from Trypanosoma brucei. Mol Biochem Parasitol 1995, 74, 119–123. [DOI] [PubMed] [Google Scholar]
- 30.Radwanska M; Magez S; Michel A; Stijlemans B; Geuskens M; Pays E Comparative analysis of antibody responses against HSP60, invariant surface glycoprotein 70, and variant surface glycoprotein reveals a complex antigen-specific pattern of immunoglobulin isotype switching during infection by Trypanosoma brucei. Infect Immun 2000, 68, 848–860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Radwanska M; Magez S; Dumont N; Pays A; Nolan D; Pays E Antibodies raised against the flagellar pocket fraction of Trypanosoma brucei preferentially recognize HSP60 in cDNA expression library. Parasite Immunology 2000, 22, 639–650. [DOI] [PubMed] [Google Scholar]
- 32.Colaco CA; MacDougall A Mycobacterial chaperonins: the tail wags the dog. FEMS Microbiol Lett 2014, 350, 20–24. [DOI] [PubMed] [Google Scholar]
- 33.Brocchieri L; Karlin S Conservation among HSP60 sequences in relation to structure, function, and evolution. Protein Science 2000, 9, 476–486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Abdeen S; Salim N; Mammadova N; Summers CM; Frankson R; Ambrose AJ; Anderson GG; Schultz PG; Horwich AL; Chapman E; Johnson SM GroEL/ES inhibitors as potential antibiotics. Bioorg. Med. Chem. Lett 2016, 26, 3127–3134. [DOI] [PubMed] [Google Scholar]
- 35.Abdeen S; Salim N; Mammadova N; Summers CM; Goldsmith-Pestana K; McMahon-Pratt D; Schultz PG; Horwich AL; Chapman E; Johnson SM Targeting the HSP60/10 chaperonin systems of Trypanosoma brucei as a strategy for treating African sleeping sickness. Bioorg. Med. Chem. Lett 2016, 26, 5247–5253. [DOI] [PubMed] [Google Scholar]
- 36.Abdeen S; Kunkle T; Salim N; Ray AM; Mammadova N; Summers C; Stevens M; Ambrose AJ; Park Y; Schultz PG; Horwich AL; Hoang QQ; Chapman E; Johnson SM Sulfonamido-2-arylbenzoxazole GroEL/ES Inhibitors as Potent Antibacterials against Methicillin-Resistant Staphylococcus aureus (MRSA). J Med Chem 2018, 61, 7345–7357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kunkle T; Abdeen S; Salim N; Ray AM; Stevens M; Ambrose AJ; Victorino J; Park Y; Hoang QQ; Chapman E; Johnson SM Hydroxybiphenylamide GroEL/ES Inhibitors Are Potent Antibacterials against Planktonic and Biofilm Forms of Staphylococcus aureus. J Med Chem 2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Johnson SM; Sharif O; Mak PA; Wang HT; Engels IH; Brinker A; Schultz PG; Horwich AL; Chapman E A biochemical screen for GroEL/GroES inhibitors. Bioorg. Med. Chem. Lett 2014, 24, 786–789. [DOI] [PubMed] [Google Scholar]
- 39.Kim S; Thiessen PA; Bolton EE; Chen J; Fu G; Gindulyte A; Han L; He J; He S; Shoemaker BA; Wang J; Yu B; Zhang J; Bryant SH PubChem Substance and Compound databases. Nucleic Acids Res 2016, 44, D1202–1213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Stromberg BE; Schlotthauer JC; Conboy GA The efficacy of closantel against Fascioloides magna in sheep. J. Parasitol 1984, 70, 446–447. [PubMed] [Google Scholar]
- 41.Meng Q; Li BX; Xiao X Toward Developing Chemical Modulators of Hsp60 as Potential Therapeutics. Front Mol Biosci 2018, 5, 35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Nagumo Y; Kakeya H; Yamaguchi J; Uno T; Shoji M; Hayashi Y; Osada H Structure-activity relationships of epolactaene derivatives: structural requirements for inhibition of Hsp60 chaperone activity. Bioorg Med Chem Lett 2004, 14, 4425–4429. [DOI] [PubMed] [Google Scholar]
- 43.Nagumo Y; Kakeya H; Shoji M; Hayashi Y; Dohmae N; Osada H Epolactaene binds human Hsp60 Cys442 resulting in the inhibition of chaperone activity. Biochem J 2005, 387, 835–840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wiechmann K; Muller H; Konig S; Wielsch N; Svatos A; Jauch J; Werz O Mitochondrial Chaperonin HSP60 Is the Apoptosis-Related Target for Myrtucommulone. Cell Chemical Biology 2017, 24, 614-+. [DOI] [PubMed] [Google Scholar]
- 45.Barrett MP; Boykin DW; Brun R; Tidwell RR Human African trypanosomiasis: pharmacological re-engagement with a neglected disease. Br J Pharmacol 2007, 152, 1155–1171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zhang YL; Keng YF; Zhao Y; Wu L; Zhang ZY Suramin is an active site-directed, reversible, and tight-binding inhibitor of protein-tyrosine phosphatases. Journal of Biological Chemistry 1998, 273, 12281–12287. [DOI] [PubMed] [Google Scholar]
- 47.Hanau S; Rippa M; Bertelli M; Dallocchio F; Barrett MP 6-Phosphogluconate dehydrogenase from Trypanosoma brucei. Kinetic analysis and inhibition by trypanocidal drugs. Eur J Biochem 1996, 240, 592–599. [DOI] [PubMed] [Google Scholar]
- 48.Fairlamb AH; Bowman IB Uptake of the trypanocidal drug suramin by bloodstream forms of Trypanosoma brucei and its effect on respiration and growth rate in vivo. Mol Biochem Parasitol 1980, 1, 315–333. [DOI] [PubMed] [Google Scholar]
- 49.Perie J; Riviere-Alric I; Blonski C; Gefflaut T; Lauth de Viguerie N; Trinquier M; Willson M; Opperdoes FR; Callens M Inhibition of the glycolytic enzymes in the trypanosome: an approach in the development of new leads in the therapy of parasitic diseases. Pharmacol Ther 1993, 60, 347–365. [DOI] [PubMed] [Google Scholar]
- 50.Willson M; Callens M; Kuntz DA; Perie J; Opperdoes FR Synthesis and activity of inhibitors highly specific for the glycolytic enzymes from Trypanosoma brucei. Mol Biochem Parasitol 1993, 59, 201–210. [DOI] [PubMed] [Google Scholar]
- 51.Wang CC Molecular mechanisms and therapeutic approaches to the treatment of African trypanosomiasis. Annu Rev Pharmacol Toxicol 1995, 35, 93–127. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.