Abstract
Background
Maternal transcripts are accumulated in the oocyte during oogenesis to provide for protein synthesis from oocyte maturation through early embryonic development, when nuclear transcription is silenced. The maternal mRNAs have short poly(A) tails after undergoing post-transcriptional processing necessary for stabilizing them for storage. The transcripts undergo cytoplasmic polyadenylation when they are to be translated. Transcriptome analyses comparing total mRNA and elongated poly(A) mRNA content among eggs of different quality can provide insight into molecular mechanisms affecting egg developmental competence in rainbow trout. The present study used RNA-seq to compare transcriptomes of unfertilized eggs of rainbow trout females yielding different eyeing rates, following rRNA removal and poly(A) retention for construction of the libraries.
Results
The percentage of embryos to reach the 32-cell stage at 24 h post fertilization was significantly correlated to family eyeing rate, indicating that inviable embryos were developmentally compromised before zygotic genome activation. RNA sequencing identified 2 differentially expressed transcripts (DETs) from total mRNA sequencing comparing females with low-quality (< 5% eyeing), medium-quality (30–50% eyeing), and high-quality (> 80% eyeing) eggs. In contrast, RNA sequencing from poly(A) captured transcripts identified 945 DETs between low- and high-quality eggs, 1012 between low- and medium-quality eggs, and only 2 between medium- and high-quality eggs. The transcripts of mitochondrial genes were enriched with polyadenylated transcript sequencing and they were significantly reduced in low-quality eggs. Similarly, mitochondrial DNA was reduced in low-quality eggs compared with medium- and high-quality eggs. The functional gene analysis classified the 945 DETs between low- and high-quality eggs into 31 functional modules, many of which were related to ribosomal and mitochondrial functions. Other modules involved transcription, translation, cell division, apoptosis, and immune responses.
Conclusions
Our results indicate that differences in egg quality may be derived from differences in maternal nuclear transcript activation and cytoplasmic polyadenylation before ovulation, as opposed to accumulation and storage of maternal nuclear transcripts during oogenesis. Transcriptome comparisons suggest low-quality eggs suffered from impaired oxidative phosphorylation and translation. The DETs identified in this study provide insight into developmental competence in rainbow trout eggs.
Electronic supplementary material
The online version of this article (10.1186/s12864-019-5690-5) contains supplementary material, which is available to authorized users.
Keywords: Rainbow trout, Egg quality, Polyadenylation, mRNA, Mitochondria
Background
Reliable production of high-quality eggs is essential for meeting production cycle demands for seed stock. Fertility is high in the rainbow trout industry when fish are maintained under optimal conditions. Nevertheless, quality of the eggs or ova can be affected by many intrinsic and extrinsic factors including the genetics, age and diet of brood fish [1–6]; pre-spawning exposure to stressors and photo-thermal cycles [7–10]; and postovulatory aging of the eggs [11–13]. Understanding mechanisms by which egg quality becomes compromised in response to suboptimal genetics, management, nutrition, and environmental conditions is critical to optimizing hatchery productivity.
The oocyte becomes transcriptionally inactive following oocyte growth and remains transcriptionally silent or greatly repressed until zygotic genome activation (ZGA) which usually takes place around the time of the mid-blastula transition (MBT) in most vertebrates. Therefore, the oocyte serves as a reservoir of biomolecules including proteins, lipids and RNAs deposited into the egg during oogenesis, for utilization from oocyte maturation through early embryonic development [14, 15]. Levels of certain proteins and lipids have been linked to egg viability in many fish species including rainbow trout [16]. A relationship between the maternal transcriptome and developmental competence has also been supported in a variety of fishes using an assortment of molecular approaches, although few investigations involved next-generation sequencing [17, 18]. In rainbow trout, genes linked to decreased egg quality caused by postovulatory aging were identified using quantitative reverse transcription PCR [19–21] and transcripts associated with decreased egg quality in response to the use of photoperiod to shorten the time to spawning or hormone-induced ovulation were identified by microarray analyses [22, 23]. One microarray study compared the transcriptome of un-manipulated female rainbow trout that exhibited either 100% viability through eyeing or less than 64%, but confirmed only one differentially expressed transcript (DET) [24]. Less than 200 DETs were identified among the studies with few overlapping transcripts between them. Although deep sequencing approaches have been used to identify some miRNAs and mitochondrial genome encoded small RNAs related to egg deterioration due to postovulatory aging [25, 26], global mRNA analysis techniques based on deep-sequencing technologies have not been applied to investigate a possible connection between mRNA content in unfertilized eggs and egg quality in fish.
Stored maternal transcripts generally have shortened polyadenylic acid (poly(A)) tails and are masked to inhibit both translation and degradation [27–30]. In the oocyte, new transcripts intended for sequestration are polyadenylated in the nucleus and then translocated to the cytoplasm where they are subsequently partially deadenylated for storage [30–32]. The stored maternal transcripts require cytoplasmic polyadenylation to allow translation. The cytoplasmic polyadenylation or deadenylation of stored transcripts is a critical control mechanism for translation during oocyte maturation and early embryonic development [28, 33, 34]. When comparing eggs of different quality, differences in expressed transcripts based on total mRNA content may indicate differences in accumulation or degradation of the transcripts throughout oogenesis and maturation, whereas differences in mRNAs with longer poly(A) tails may indicate differences and changes in the translational activity of the transcripts [35–38]. To further understand the relationship between the maternal transcriptome and egg quality, we used RNA-seq to compare mRNA transcriptomes of ovulated eggs from 20 individual females that produced eggs of disparate quality as determined by eyeing rate. We compared transcriptomes sequenced from libraries prepared following rRNA removal or by oligo(dT) capture of polyadenylated RNA. Oligo(dT) capture methodologies are not efficient at capturing transcripts with short poly(A) tails [35, 39, 40], and therefore, mRNA transcriptomes sequenced from libraries prepared by oligo(dT) capture of polyadenylated RNA should be enriched in activated transcripts with elongated poly(A) tails.
Results and discussion
Eyeing rate and early embryo viability
Viability was assessed at ~ 250 accumulated thermal units (ATUs) post fertilization, which we refer to as eyeing in the present manuscript. The 250 ATU mark is actually after retinal pigmentation but is often used by industry because mortality is generally very low after retinal pigmentation [41]; it is well after embryos are resistant to mechanical shock [42, 43], and is several days before hatching which allows time for the eggs to be sorted to remove dead and subviable eggs before shipment to production facilities. Eyeing rates for the families in the selective breeding program was typical of rainbow trout aquaculture operations [5], exhibiting a mean eyeing rate of 79.3% (Fig. 1). Only 16 of the 192 families evaluated had less than 50% eyeing and were classified as subfertile. Furthermore, only six families exhibited an eyeing rate of less than 10%, whereas eyeing rates were greater than 30% for all others. Sperm used to fertilize each of the subfertile families also yielded families with eyeing rates over 78%, substantiating the eggs and not the sperm as the cause of the subfertility. Visual inspection of eggs collected before fertilization did not show obvious signs of the eggs being compromised in ways that would allow for their being discarded by hatchery personnel.
Fig. 1.
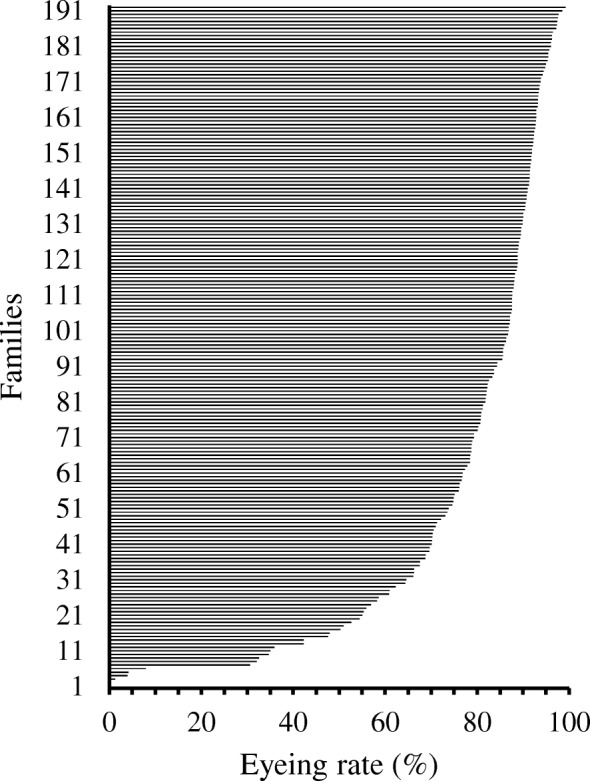
Eyeing rates of the 192 surveyed rainbow trout families in the selective breeding program
Embryo cleavage was assessed for the 20 selected families at about 24 h post fertilization (Table 1). On average about 80% of the embryos in the high-quality families reached at least the 32-cell stage at this time. The percentage of embryos reaching the 32-cell stage was similar to the eyeing rates for most families, suggesting that most of the embryos that would not survive had died or had very delayed development by the 32-cell stage. The percentage of embryos reaching the 32-cell stage was significantly correlated to eyeing rate (R = 0.85; P < 0.001). Few embryos failed to reach the 8-cell stage by 24 h post fertilization. Together these data support most of the non-viable embryos were fertilized but failed developmentally before the 32-cell stage, which should be before the major wave of ZGA. Although the timing of the major wave of ZGA has not been characterized in rainbow trout, in general, more cleavage divisions are completed before ZGA in animals that develop more slowly [44]. The ZGA for most fish species investigated, all of which develop more rapidly than rainbow trout, have been shown to begin during the MBT at about cell cycle 10 or ~ 1000 cells [45–49]. An exception is the medaka (Oryzias latipes) in which the ZGA begins at about the 64-cell stage; before the MBT [50]. A previous study on the same rainbow trout broodstock population as in the present study reported most embryo mortality in subfertile families took place by the second cleavage interval [51]; earlier than in our study and well before ZGA. Although the timing of embryonic mortality cannot specify a cause of the mortality, most of the embryonic mortality reported for this population of rainbow trout has taken place while the embryos were dependent upon maternal transcripts and before many of the other forms of stored biomolecules would be either required or exhausted by the embryos, leaving aberrations in levels or activation of maternal transcripts as possible contributors to subfertility.
Table 1.
Assessment of early embryo development in 20 selected families. The percentage of embryos reaching each cell stage by ~ 24 h post fertilization, and eyeing rate, are indicated
| Egg quality group | Family ID # | Embryos collected at ~ 24 h post fertilization | Eyeing rate (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Total embryos | Embryos with ≥2 cells (%) | Embryos with ≥4 cells (%) | Embryos with ≥8 cells (%) | Embryos with ≥16 cells (%) | Embryos with ≥32 cells (%) | ||||
| Subfertile | Low quality | 48 | 26 | 62 | 62 | 58 | 42 | 12 | 0.0 |
| 63 | 42 | 81 | 81 | 74 | 21 | 0 | 1.3 | ||
| 129 | 31 | 97 | 97 | 90 | 42 | 0 | 0.0 | ||
| 136 | 53 | 98 | 98 | 92 | 81 | 9 | 4.2 | ||
| Medium quality | 13 | 30 | 93 | 93 | 93 | 93 | 63 | 30.6 | |
| 60 | 61 | 82 | 82 | 75 | 41 | 3 | 42.2 | ||
| 71 | 54 | 91 | 91 | 91 | 69 | 13 | 47.5 | ||
| 106 | 45 | 93 | 93 | 91 | 91 | 40 | 34.7 | ||
| 108 | 43 | 88 | 88 | 88 | 53 | 19 | 35.9 | ||
| 114 | 49 | 100 | 100 | 100 | 94 | 65 | 42.2 | ||
| Fertile | High quality | 19 | 32 | 94 | 94 | 94 | 94 | 63 | 87.6 |
| 47 | 25 | 100 | 100 | 100 | 100 | 88 | 83.6 | ||
| 58 | 39 | 92 | 92 | 92 | 92 | 59 | 80.6 | ||
| 59 | 52 | 98 | 98 | 98 | 96 | 83 | 97.7 | ||
| 87 | 50 | 98 | 98 | 98 | 98 | 84 | 89.1 | ||
| 97 | 54 | 94 | 94 | 94 | 94 | 81 | 87.4 | ||
| 99 | 51 | 100 | 100 | 100 | 100 | 100 | 92.6 | ||
| 102 | 59 | 100 | 100 | 100 | 100 | 95 | 92.1 | ||
| 103 | 34 | 100 | 100 | 100 | 100 | 100 | 97.5 | ||
| 119 | 47 | 98 | 98 | 98 | 96 | 87 | 88.8 | ||
Mapping of sequencing reads following rRNA removal and poly(a) retention
RNA-seq analysis of the eggs of 20 females generated 31 to 58, and 43 to 68 million reads from the libraries constructed with the Illumina® TruSeq® Stranded Total RNA Library Prep Kit with Ribo-Zero Gold (rRNA removal) and TruSeq® Stranded mRNA Sample Prep Kit (poly(A) retention) respectively (Table 2). Similar percentages of the reads were mapped to rRNA gene sequences using the two approaches, with an average of 5.4% derived from the libraries constructed by rRNA removal and 4.7% derived from the libraries constructed by poly(A) retention kits respectively. Whereas the percentage of reads mapped to the nuclear transcriptome was half as great with poly(A) retention compared with rRNA removal, the percentage of reads mapped to mitochondrial RNA was more than 10-fold greater for poly(A) retention compared with rRNA removal. This reversal in trends is consistent with a higher proportion of the mitochondrial mRNA transcripts being polyadenylated than the proportion of the nuclear mRNA transcripts that are polyadenylated or efficiently captured by the poly(A) retention procedures.
Table 2.
Overview of RNA-seq read alignments
| Egg quality group | Female ID# | rRNA removal | Poly(A) retention | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Total reads | Mapped to transcriptome (%) | Mapped to mitochondrial RNA (%) | Mapped to rRNA (%) | Total reads | Mapped to transcriptome (%) | Mapped to mitochondrial RNA (%) | Mapped to rRNA (%) | ||
| Low quality | 48 | 37,157,250 | 43.6 | 3.1 | 8.3 | 61,999,239 | 30.3 | 32.7 | 2.4 |
| 63 | 39,461,498 | 48.5 | 2.4 | 5.4 | 64,750,933 | 39.3 | 22.3 | 2.0 | |
| 129 | 58,281,853 | 54.8 | 2.4 | 3.7 | 59,599,504 | 39.1 | 18.4 | 3.3 | |
| 136 | 36,865,244 | 47.3 | 1.6 | 5.7 | 43,578,497 | 34.8 | 20.8 | 3.0 | |
| Mean | 42,941,461 | 48.6 | 2.4 | 5.8 | 57,482,043 | 35.9 | 23.6 | 2.7 | |
| Medium quality | 13 | 37,890,847 | 44.9 | 3.0 | 7.6 | 64,816,319 | 20.9 | 53.9 | 5.7 |
| 60 | 36,566,538 | 49.0 | 2.9 | 4.4 | 60,757,076 | 13.8 | 62.5 | 4.2 | |
| 71 | 33,120,194 | 51.2 | 5.0 | 4.1 | 55,561,085 | 12.9 | 61.2 | 12.7 | |
| 106 | 37,362,260 | 49.0 | 2.7 | 5.1 | 62,699,350 | 24.7 | 45.5 | 3.0 | |
| 108 | 53,796,039 | 52.9 | 6.8 | 5.5 | 61,835,604 | 24.5 | 45.6 | 2.6 | |
| 114 | 32,683,461 | 41.3 | 6.5 | 7.7 | 59,163,406 | 11.7 | 64.7 | 2.4 | |
| Mean | 38,569,890 | 48.1 | 4.5 | 5.7 | 60,805,473 | 18.1 | 55.6 | 5.1 | |
| High quality | 19 | 37,448,462 | 46.1 | 3.3 | 7.7 | 66,890,967 | 32.3 | 35.5 | 2.8 |
| 47 | 37,299,964 | 48.1 | 3.0 | 4.6 | 61,202,732 | 34.6 | 27.9 | 2.2 | |
| 58 | 36,637,099 | 43.5 | 2.1 | 6.2 | 61,065,293 | 37.6 | 26.6 | 1.8 | |
| 59 | 31,040,335 | 50.2 | 4.9 | 5.2 | 52,972,699 | 10.5 | 64.2 | 15.0 | |
| 87 | 38,856,780 | 49.7 | 2.8 | 5.0 | 55,700,910 | 18.8 | 53.3 | 5.9 | |
| 97 | 38,254,940 | 51.0 | 2.5 | 4.1 | 61,667,813 | 20.8 | 48.1 | 7.4 | |
| 99 | 55,069,888 | 52.7 | 4.8 | 3.2 | 68,472,008 | 25.4 | 46.3 | 1.7 | |
| 102 | 32,044,365 | 49.0 | 5.8 | 5.4 | 57,053,445 | 10.6 | 65.2 | 9.1 | |
| 103 | 39,749,239 | 48.1 | 1.7 | 4.1 | 59,550,700 | 23.8 | 43.2 | 4.3 | |
| 119 | 51,515,745 | 51.4 | 8.1 | 5.6 | 62,330,132 | 15.9 | 61.7 | 2.7 | |
| Mean | 39,791,682 | 49.0 | 3.9 | 5.1 | 60,690,670 | 23.0 | 47.2 | 5.3 | |
| All | Mean | 40,055,100 | 48.6 | 3.8 | 5.4 | 60,083,386 | 24.1 | 45.0 | 4.7 |
Mitochondrial mRNA transcripts are polyadenylated with a tail of approximately 50 nucleotides in vertebrates as part of transcript processing and therefore most would be expected to be captured by the poly(A) retention [52, 53]. Similarly, nuclear mRNA transcripts are also polyadenylated as part of processing. In most cells the majority of cytosolic nuclear transcripts are polyadenylated with a poly(A) tail greater than 80 nucleotides [29, 54]. However, stored maternal nuclear transcripts possess a short poly(A) tail around 15–40 nucleotides that are elongated to over 80 nucleotides through cytoplasmic polyadenylation during activation [27, 29, 36, 40, 55]. In general, oligo(dT) capture approaches are not very efficient at capturing mRNAs with shorter poly(A) tails [35, 39, 40]. As far as we are aware, the capture efficiency of the Illumina® TruSeq® Stranded mRNA Sample Preparation Kit for short poly(A) tails such as those in stored maternal mRNA, has not been characterized. It is therefore likely that data collected following poly(A) retention represent primarily activated mRNAs with longer poly(A) tails and data following rRNA removal represent primarily the more abundant stored maternal transcripts with short poly(A) tails, which is consistent with a lower percentage of the transcript reads aligning to the nuclear transcriptome following poly(A) retention compared with rRNA removal.
Eggs of vertebrates possess large stockpiles of mitochondria that are active in providing energy during maturation and stockpiling ATP for energy to drive early embryonic events including cleavage [56–59]. Furthermore, there is extensive mitochondrial DNA (mtDNA) replication during maturation that then ceases until after ZGA. High proportions of mitochondrial transcripts as observed in our study have been reported in early cod and halibut embryos [48, 60]. The percentage of reads that were aligned as mitochondrial transcripts was reduced in the low-quality eggs following poly(A) retention as well as rRNA removal. The percentage of reads aligned to mitochondrial RNAs in the low-quality eggs following poly(A) retention was 23.6% compared with 47.2% in the high-quality eggs, and 2.4% compared with 3.9% respectively following rRNA removal (Table 2). A reduction in mitochondria and mtDNA has been identified as a cause of reduced fertility in several mammalian species and serves as a marker of oocyte quality in mammals [56, 57]. Furthermore, augmentation of mitochondrial number or mtDNA in maturing oocytes has been recognized as a method to improve oocyte quality in mammals including humans [56, 57, 61, 62]. We therefore used real-time quantitative PCR to measure abundance of mtDNA in our samples by measuring two mitochondrial genes, mt-atp6 and mt-cyb. Abundance of both genes were reduced 1.6 log2 fold change (log2FC) in the low-quality eggs (Fig. 2) suggesting a reduction in mitochondria or mtDNA may contribute to differences in the percentage of reads aligned to the mitochondrial transcriptome among egg quality groups.
Fig. 2.
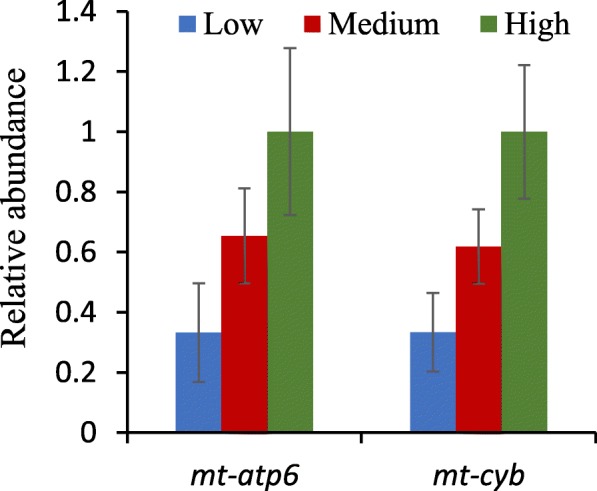
Relative abundance of mitochondrial DNA. Real-time quantitative PCR measurement of mt-atp6 and mt-cyb genes normalized to 18S, in low-, medium-, and high-quality eggs. (mean ± SEM)
Gene expression profiling revealed greater differences in polyadenylated mRNA than total mRNA abundance with fertility
We detected 44,330 and 39,133 transcripts with at least three normalized reads expressed in the libraries constructed by rRNA removal and poly(A) retention kits respectively. About 89% of the transcripts following rRNA removal and 77% of the transcripts following poly(A) retention were shared among the three treatment groups (Fig. 3) and many of those that were not shared had low transcript numbers. No DETs were identified comparing the 10 subfertile and 10 fertile or high-quality females from either the rRNA removal or poly(A) retention libraries by DESeq2 with the criteria of a false discovery rate (FDR) < 0.05. We therefore divided the subfertile group into a low- and medium-quality group (Table 1).
Fig. 3.
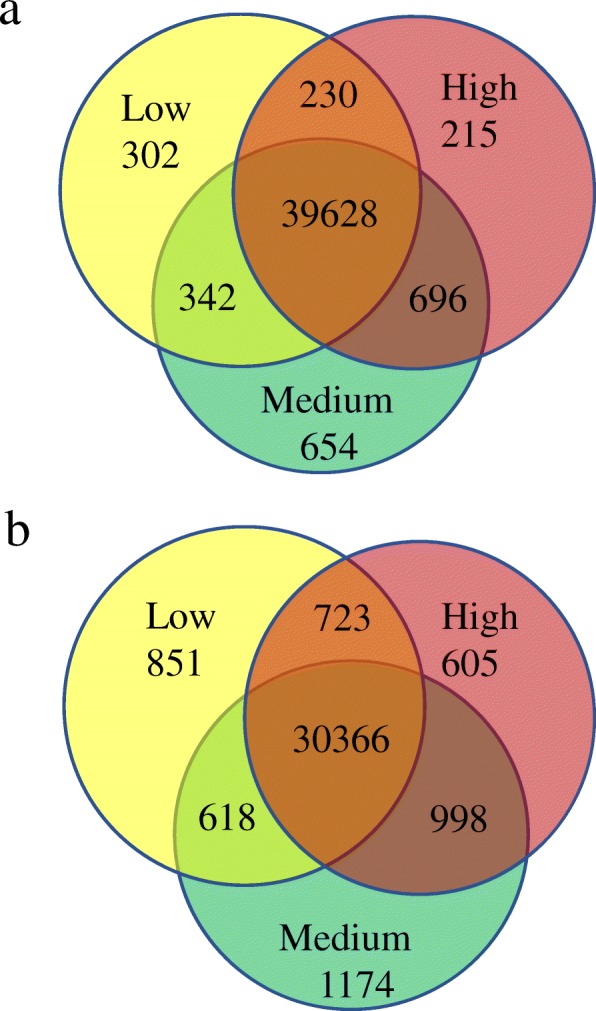
Gene transcripts detected in low-, medium-, and high-quality eggs. a Libraries constructed by rRNA removal. b Libraries constructed by poly(A) retention. Genes expressed with average normalized reads greater than 1 were counted
Only two DETs were identified among the three treatment groups; low-, medium-, and high-quality eggs, from our dataset sequenced from the libraries constructed with an rRNA removal kit by DESeq2 with the criteria of log2FC ≥ 1, FDR ≤ 0.05, and a total number of normalized reads in a given comparison being ≥500 (Additional file 1: Table S1). The transcript agfg1 (Arf-GAP domain and FG repeats-containing protein 1) was enriched in low-quality eggs compared with high-quality eggs (1.81 log2FC, FDR = 0.0172) and ahnak (neuroblast differentiation-associated protein ahnak-like) was enriched in low-quality eggs compared with medium-quality eggs (2.01 log2FC, FDR = 0.0026). AGFG1 has been shown to affect the accumulation of a small set of non-polyadenylated cellular mRNAs in mammalian cells [63], and mRNA of Drongo (Drosophila neural GTS1-like), the homolog of AGFG1 in the fruit fly, associates with Me31B (maternal expression of 31B) which is required for the translational repression of maternal mRNAs in the oocyte [64]. AHNAK is considered to be involved in calcium flux regulation and has been proposed to interact with s100 proteins to regulate cellular Ca2+ homeostasis [65]. Notably, an s100 protein, s100a1 (S100 calcium binding protein A1), is among the most reduced polyadenylated transcripts in low-quality eggs. Most important, the lack of DETs following rRNA removal supports the concept that low fertility was not a result of differences in the accumulation or degradation of transcripts throughout oogenesis or maturation.
A total of 1339 transcripts were differentially expressed among the egg quality groups when the libraries were constructed with a poly(A) retention kit, using the same criteria as with rRNA removal (Additional file 1: Table S1). There were only 2 DETs comparing medium- and high-quality eggs, esr2b (estrogen receptor beta 2) and pltp (phospholipid transfer protein), both of which were higher in medium quality eggs compared with both low- and high-quality eggs. There were 945 DETs comparing low- and high-quality eggs, and 1012 comparing low- and medium-quality eggs. There were 619 DETs shared between the low- versus high-quality egg comparison and the low- versus medium-quality comparison, and both DETs from the medium- versus high-quality comparison were also differentially expressed between the low- and medium-quality eggs. The shared DETs between the low- versus high- quality comparison and the low- versus medium-quality comparisons were consistent in direction of change for any given transcript. Taken together, there was little difference in the transcriptomes of medium-quality eggs with between 30 and 50% eyeing rates, and the high-quality eggs with greater than 80% eyeing rates; therefore, we focused our further analyses and discussions on the low- versus high-quality comparison. The 945 DETs included 732 unique gene descriptions, with many of the more differentially expressed transcripts sharing gene descriptions with multiple transcripts or locus tags.
Among those 945 DETs, 724 were decreased and 221 were increased in low-quality eggs (Additional file 1: Table S1). There were many nuclear genes associated with ribosome production and function among the most significantly reduced DETs. Over 100 of the 945 DETs were for 40s or 60s ribosomal proteins. In general, the functions of the transcripts were diverse even among the most differentially expressed transcripts (Fig. 4). The transcripts in Fig. 4 were associated with 62 different GO terms for Biological process and 30 for Molecular function, even with 14 of these transcripts having no associated GO terms. Among the 221 enriched transcripts in the low-quality eggs, the gene transcript with the largest increase, 2.07 log2FC, is tob1 (protein tob1-like) (Additional file 1: Table S1). This transcript, however, was highly variable among the low-quality groups (FDR = 0.0010). Whereas the high-quality eggs ranged from 46 to 280 normalized reads, one of the low-quality groups with no survival at eyeing and another with 1.3%, had over 1300 normalized reads each, whereas the other two had below 80 reads. TOB1 is characterized as an anti-proliferative protein whose activity is mediated through interactions with the Caf1a/Caf1b deadenylases leading to target mRNA deadenylation and decay [66–68]. Considerable differences in expression among the groups of low-quality eggs, including a transcript involved with mRNA deadenylation and decay, might suggest disparate causes leading to the reduction in quality or differences in the progression of viability among the low-quality families.
Fig. 4.

Heat map of 50 differentially expressed transcripts. The top 10 up-regulated and 40 down-regulated genes in low-quality eggs were selected based on false discovery rate (FDR) value. The red bar indicates females with low-quality eggs and the green bar indicates females with high-quality eggs. * Indicates the gene has not been officially named. The gene name abbreviations are listed in Additional file 1: Table S7
Because egg quality is a major issue in the aquaculture industry [69], many microarray and quantitative reverse transcription PCR studies have revealed sets of genes associated with egg quality [17]. In rainbow trout, eggs with reduced egg quality in response to postovulatory aging had lower expression levels of tubb (tubulin b) and npm2 (nucleoplasmin-2) [20], and eggs with reduced quality in response to photoperiod treatment to accelerate the time of spawning had increased expression of pyc (pyruvate carboxylase) [22]. Consistent with those findings, our data show transcript levels of tubb and npm2 are decreased, and pyc is increased, in low-quality eggs. On the other hand, we observed decreased expression of krt8 (keratin 8), krt18 (keratin 18), rpl24 (60s ribosomal protein l24), and apoc1 (apolipoprotein c1) with reduced egg quality, whereas krt8 and krt18 have been reported to increase with reduced egg quality due to postovulatory aging [20], and rpl24 and apoc1 to increase with reduced egg quality in response to hormone implantation or photoperiod manipulation to accelerate the time of spawning [22]. The three studies used poly(A) retention and therefore likely measured changes in activated transcripts. In the present study total transcript abundance based on rRNA removal libraries did not significantly differ between high- and low-quality eggs for any of the seven genes mentioned, with the mean normalized reads differing by less than 10% for rpl24, pyc, npm2, and tubb, and 68, 80 and 146% for krt8, krt18 and apoc1 respectively. Moreover, the directions of the trends were similar with poly(A) retention and rRNA removal for each of these transcripts. The differences in the direction of the expression of specific genes among studies may be due to the treatments behind the reductions in egg quality or criteria for assessing developmental competence. Alternatively, differences may reflect where in the activation and deactivation cycle of the transcripts the eggs were sampled, and not just a measure of how many transcripts are activated overall. Further study of the kinetics of cytoplasmic adenylation and deadenylation in rainbow trout may help with interpretations of transcript levels and their associations with egg quality.
Even with poly(A) retention, only two transcripts were differentially expressed between medium- and high-quality eggs (esr2b and pltp) despite the medium-quality eggs ranging from 31 to 48% eyeing, and also representing only the lower 8% of clutches in terms of eyeing rate. The lack of DETs between medium- and high-quality eggs may suggest transcript variance did not contribute to the reduced egg quality, or transcript disparities were not detected due to the nature of the study or data analyses. Transcript signatures associated with less drastic differences in fertility have been identified in rainbow trout eggs in response to specific treatments [20, 22, 23]. Since the cause or causes of the reduced egg quality in the present study is undefined, there is likely a host of maladies among the egg clutches resulting in reduced fertility that do not share the same molecular pathways, making transcriptome signatures of any one difficult to discern. More robust analyses such as use of artificial neural networks and supervised machine learning which has been able to identify molecular signatures composed of many minor changes in transcript levels in oocytes of striped bass that associated with fertility [70] may be helpful. Regardless, the expressions of many genes have been linked to egg quality in the present study including genes not previously associated with egg quality, providing new insight into how an egg can become compromised.
Mitochondrial RNA expression
The rainbow trout mitochondrial genome encodes 13 polypeptides, 22 tRNAs, two rRNAs and has a noncoding D-loop region [71]. Transcripts for the 13 mitochondrial protein coding genes and D-loop region were found to be significantly reduced in the low-quality eggs by RNA-seq with poly(A) retention libraries (Table 3). In addition, mt-tn (tRNA-Asn) was also decreased in low-quality eggs but few transcripts were detected for this or all other tRNA genes. There were also abundant reads for the rRNAs; mt-rnr1 (mitochondrion 12S) and mt-rnr2 (mitochondrion 16S), and despite the trends being similar for the rRNAs and mRNAs, the DESeq2 analysis did not detect a significant reduction in mt-rnr1 or mt-rnr2 in low-quality eggs (FDR > 0.1; Table 3). Although there are some reports of mitochondrial rRNAs being polyadenylated [72, 73], mitochondrial rRNAs are generally not polyadenylated or contain no more than 10 nucleotides of the tail in most vertebrate cells [52, 53, 74]. It is worth noting that whereas the rRNAs comprised about half of the mitochondrial RNA reads following rRNA removal (Additional file 1: Table S2), they comprised less than 5% of the mitochondrial RNA reads following poly(A) retention, supporting the idea that few of the rRNA transcripts were polyadenylated or possessed long poly(A) tails. We do not know if the antibodies used for rRNA removal recognize mt-rnr1 or mt-rnr2 to any extent. Mitochondrial rRNA transcripts are polyadenylated as part of the degradation process [72] and therefore the differences in polyadenylated rRNA transcripts among groups may represent differences in transcripts undergoing degradation.
Table 3.
DESeq2 statistics for mitochondrial ribosomal RNA and differentially expressed mRNA transcripts
| Gene | Base mean | Log2 fold change | P-value | FDR |
|---|---|---|---|---|
| dlp | 119,812 | −1.73 | 3.86E-05 | 1.94E-03 |
| mt-atp6 | 1,338,170 | −1.63 | 6.52E-05 | 2.89E-03 |
| mt-atp8 | 113,992 | −1.77 | 5.49E-05 | 2.58E-03 |
| mt-col | 8,076,707 | −1.94 | 2.65E-05 | 1.46E-03 |
| mt-coll | 7,324,055 | −1.88 | 2.09E-05 | 1.23E-03 |
| mt-colll | 4,570,645 | −1.81 | 2.31E-05 | 1.31E-03 |
| mt-cyb | 8,530,201 | −2.10 | 4.01E-06 | 3.51E-04 |
| mt-nd1 | 914,918 | −1.70 | 5.69E-05 | 2.63E-03 |
| mt-nd2 | 418,048 | −1.70 | 9.08E-05 | 3.76E-03 |
| mt-nd3 | 265,015 | −1.35 | 9.77E-04 | 2.05E-02 |
| mt-nd4 | 994,427 | −1.67 | 9.67E-05 | 3.96E-03 |
| mt-nd4l | 86,839 | −1.73 | 8.50E-05 | 3.55E-03 |
| mt-nd5 | 702,120 | −1.55 | 3.57E-04 | 1.01E-02 |
| mt-nd6 | 238,040 | −1.64 | 1.14E-04 | 4.43E-03 |
| my-tn | 136 | −1.80 | 5.90E-04 | 1.43E-02 |
| mt-rnr1 | 327,067 | −1.06 | 4.83E-02 | 2.43E-01 |
| mt-rnr2 | 1,289,619 | −1.30 | 1.47E-02 | 1.20E-01 |
The changes in expression of the 13 mitochondrial protein genes between high-quality and low-quality eggs are very consistent, ranging between − 1.34 and − 2.10 log2FC as calculated by DESeq2 (Table 3). This consistency may be due to coordinated regulation of the transcripts as they are all needed for oxidative phosphorylation [52, 57], or to a reduction in mitochondria in the low-quality eggs. However, DESeq2 uses shrinkage estimators [75] that resulted in reduced log2FC estimates for many DETs compared with calculating log2FC based only on normalized read values. If log2FC for the 13 protein mRNAs are calculated based simply on the normalized reads the average is about − 2.21 log2FC for the low-quality eggs compared with the high-quality eggs (Additional file 1: Table S2). This magnitude of change is greater than the − 1.6 log2FC difference in mitochondrial DNA (Fig. 2), supporting an additional reduction in expression beyond just a reduction in mitochondrial number although the measurements were conducted using different assays. It is also worth noting that using the same calculation approach, mt-rnr1 and mt-rnr2 are also about a 2.3 log2FC, suggesting similar circumstances behind the reduced levels of the rRNAs and the mRNAs in low-quality eggs.
Interestingly DESeq2 analysis of data from rRNA removal libraries revealed no significant differences in mitochondrial gene expression among egg quality groups (FDR > 0.999). Nevertheless, the numerical means for the protein and D-loop mRNAs, and rRNAs, were lower in the low-quality eggs compared to the high-quality eggs. Again, it is worth noting that whereas there was an average − 0.17 log2FC between high- and low-quality eggs for these transcripts with DESeq2 (Additional file 1: Table S1), the log2FC calculated based simply on the normalized reads averaged − 0.78 (Additional file 1: Table S2). This is in line with the differences in the proportion of aligned mitochondrial transcripts among treatments with rRNA removal libraries (Table 2). Differences in percentage of reads aligning with mitochondrial genes among groups are not consistent with a − 1.6 log2FC difference in mitochondrial DNA or mitochondrial numbers between low- and high-quality eggs. Among possible explanations for differences in mtDNA being greater than differences in total transcript reads for mitochondrial genes may lie in differences in the timing of transcription and mtDNA replication. During maturation there is an explosion in mtDNA replication, increasing 1000-fold in some species [59]. It is not known when the transcripts being measured in ovulated eggs are transcribed or processed in relation to this increase in mtDNA replication. Perhaps a lower content of mtDNA in low-quality eggs has not yet been reflected in transcription rates. Alternatively, there could be a compensatory increase in mitochondrial transcription in those oocytes with reduced mtDNA. The greater reduction in polyadenylated mitochondrial transcripts than mtDNA in low-quality eggs may reflect a combination of less mtDNA and transcript processing within mitochondria.
The requirement for oxidative phosphorylation by mitochondria to provide ATP in the early embryo varies among species [56, 76]. In the zebrafish, the maternal ATP pool is insufficient to execute the ubiquitin proteasomal pathway required for protein degradation required to advance beyond the 32-cell stage [76]. Although the 32-cell stage is well before ZGA in this species [45, 47], mitochondrial transcription has been shown to be active prior to mtDNA replication in zebrafish embryos [58]. Mitochondria in zebrafish embryos are active and free fatty acids serve as substrate for oxidative phosphorylation to supply the required ATP [76]. Less is known about how rainbow trout meet energy demands in the early embryo but oxidative metabolism is present even in the unfertilized eggs, and continues through early development [77]. Although the timing of the mortality in zebrafish embryos deficient in the ability to produce ATP is similar to that of the present study, egg ATP levels were found to not correlate with fertility in rainbow trout eggs [10, 77]. The associations among mitochondrial mtDNA abundance, mitochondrial transcript levels, ATP levels, and egg quality in rainbow trout are unresolved. Mitochondria serve many functions in addition to ATP production that are essential to embryo survival such as sequestration and release of intracellular calcium [78, 79]. Furthermore, deficiencies in mitochondrial activity and mtDNA number have been shown to have separate although overlapping impacts on egg quality [56, 80].
Functional classification of differentially expressed genes
Gene ontology (GO) analysis of the 945 DETs from the comparison of polyadenylated transcript enriched libraries for low- versus high-quality eggs revealed one or more associated GO terms for 811 of the transcripts. The five most common GO terms in Biological process are ribosome biogenesis, translation, metabolic process, oxidation-reduction process, and DNA templated regulation of transcription (Fig. 5). Clustering of DETs based on associated GO terms, using the Database for Annotation, Visualization and Integrated Discovery (DAVID) gene functional classification algorithms under kappa value of 0.3, resulted in 31 functional modules [81, 82] (Table 4). The enrichment scores of the modules ranged from 1.52 to 13.40 with the number of genes in each module ranging from 4 to 164, which included 547 of the DETs in total.
Fig. 5.
The 10 most represented gene ontology terms in biological process. The analysis included the 945 differentially expressed transcripts comparing low- and high-quality eggs. The number of significantly enriched gene ontology terms is shown in parenthesis, (P < 0.05)
Table 4.
Gene cluster analysis of differentially expressed genes
| Cluster | Number of genes | Cluster enrichment | Maximum number of GOs in biological process | Maximum number of GOs in molecular function | Major involvement | ||
|---|---|---|---|---|---|---|---|
| GO description | P-value | GO description | P-value | ||||
| Cluster 1 | 164 | 8.43 | ribosome biogenesis | 1.01E-130 | structural constituent of ribosome | 4.07E-159 | Ribosome biogenesis |
| Cluster 2 | 74 | 5.82 | transport | 5.81E-03 | transporter activity | 2.56E-02 | Mitochondrion related transporter |
| Cluster 3 | 55 | 2.09 | autophagosome assembly | 9.55E-04 | protein binding | 1.09E-05 | Protein transport and modification |
| Cluster 4 | 48 | 2.44 | proteolysis | 9.53E-02 | protein binding | 1.09E-05 | Protein transport |
| Cluster 5 | 32 | 3.22 | metabolic process | 4.74E-02 | GTP binding | 1.29E-04 | Translation |
| Cluster 6 | 30 | 2.39 | phosphorylation | 4.84E-02 | ATP binding | 9.37E-03 | Phosphorylation |
| Cluster 7 | 29 | 2.68 | regulation of transcription, DNA-templated | 7.14E-03 | zinc ion binding | 4.92E-02 | Transcription |
| Cluster 8 | 22 | 13.4 | rRNA processing | 3.43E-07 | nucleic acid binding | 2.32E-02 | Ribosome processing |
| Cluster 9 | 22 | 10.73 | metabolic process | 4.74E-02 | metal ion binding | 5.60E-02 | Transcription |
| Cluster 10 | 20 | 10.7 | oxygen transport | 8.43E-20 | iron ion binding | 5.84E-07 | Oxygen transport |
| Cluster 11 | 17 | 2.35 | metabolic process | 4.74E-02 | zinc ion binding | 4.92E-02 | Zinc ion binding |
| Cluster 12 | 13 | 5.26 | hydrogen ion transmembrane transport | 6.72E-10 | NADH dehydrogenase (ubiquinone) activity | 5.13E-14 | NADH dehydrogenase (ubiquinone) activity |
| Cluster 13 | 13 | 10.9 | cellular iron ion homeostasis | 1.69E-12 | ferric iron binding | 1.69E-16 | Iron ion homeostasis and oxidation reduction |
| Cluster 14 | 12 | 2.96 | ATP synthesis coupled proton transport | 1.42E-11 | proton-transporting ATP synthase activity | 4.16E-09 | ATP generation |
| Cluster 15 | 11 | 1.74 | G-protein coupled receptor signaling pathway | 1.22E-02 | structural molecule activity | 1.40E-02 | structural molecule activity |
| Cluster 16 | 10 | 1.56 | regulation of transcription from RNA polymerase II promoter | 1.35E-01 | nucleic acid binding | 2.32E-02 | Transcription |
| Cluster 17 | 10 | 5.65 | oxidation-reduction process | 2.83E-04 | glutathione peroxidase activity | 1.93E-06 | Protect from oxidative damage |
| Cluster 18 | 9 | 1.7 | signal transduction | 7.68E-04 | transposase activity | 8.42E-07 | Immune response, |
| Cluster 19 | 9 | 5.28 | positive regulation of RNA polymerase II transcriptional preinitiation | 4.86E-05 | TBP-class protein binding | 3.31E-04 | Cell division |
| Cluster 20 | 8 | 1.86 | regulation of cell proliferation | 7.20E-02 | calcium ion binding | 6.47E-04 | Regulate cell proliferation |
| Cluster 21 | 8 | 1.67 | midbrain-hindbrain boundary structural organization | 4.06E-04 | transcription factor activity, sequence-specific DNA binding | 1.32E-02 | Transcription regulation |
| Cluster 22 | 7 | 4.46 | multicellular organism development | 2.96E-02 | nucleic acid binding | 2.32E-02 | Cell cycle and division |
| Cluster 23 | 7 | 3.39 | proton transport | 1.63E-05 | NADH dehydrogenase (ubiquinone) activity | 5.13E-14 | Mitochondrial electron transport |
| Cluster 24 | 6 | 1.76 | interstrand cross-link repair | 3.82E-03 | metal ion binding | 5.60E-02 | Transcription regulation |
| Cluster 25 | 6 | 1.92 | transmembrane transport | 1.35E-02 | ATP binding | 9.37E-03 | Regulate apoptosis |
| Cluster 26 | 6 | 5.01 | heart contraction | 1.13E-02 | metal ion binding | 5.60E-02 | Transcription regulation |
| Cluster 27 | 6 | 1.52 | microtubule-based movement | 1.03E-01 | metal ion binding | 5.60E-02 | Transcription regulation |
| Cluster 28 | 6 | 1.97 | proteolysis | 9.53E-02 | zinc ion binding | 4.92E-02 | Regulate cell cycle and apoptosis |
| Cluster 29 | 5 | 1.7 | metabolic process | 4.74E-02 | metal ion binding | 5.60E-02 | ATP binding |
| Cluster 30 | 4 | 2.96 | regulation of transcription, DNA-templated | 7.14E-03 | DNA binding | 5.37E-04 | Transcription regulation |
| Cluster 31 | 4 | 10.23 | neurotrophin TRK receptor signaling pathway | 2.92E-07 | structural constituent of ribosome | 4.07E-159 | Transcription regulation |
The largest cluster included 164 transcripts involved in ribosome biogenesis of which only 7 were increased in low-quality eggs. As mentioned, over 100 DETs are for 40s and 60s ribosomal proteins. Moreover, the categories of the GO terms in gene clusters 2, 3, 4, 5, 7, 8, 9, and 31, are also associated with ribosome function. Clusters 2, 12, 14, 17, and 23 are involved in mitochondrial function. Together, many of the clusters are associated with translation and the production of energy by mitochondria to drive early cell division. The other gene clusters are mainly involved in regulation of transcription, cell division, apoptosis, and immune responses. Transcripts increased in low-quality eggs were distributed among 22 clusters.
There were 398 DETs that were not included in the clusters. This includes the 134 DETs without associated GO terms such as n1/n2 protein (histone-binding protein n1 n2-like), senp7 (sentrin-specific protease 7-like) and parp8 (kisutch poly [ADP-ribose] polymerase 8-like); and 264 that were orphan genes such as mettl3 (n6-adenosine-methyltransferase 70 kda subunit), bmp10 (bone morphogenetic protein 10-like), and haus3 (haus augmin-like complex subunit 3-like). The listed transcripts are among the top DETs based on FDR values and therefore may also serve as important indicators of egg quality (see Fig. 4).
Conclusions
The present study identifies differences in the transcriptome among ovulated eggs of different quality for which most of the mortality occurred between fertilization and the 32-cell stage, which is before ZGA. The identification of only two DETs by RNA-seq of libraries constructed by rRNA removal kits, compared with 1339 DETs derived from libraries following poly(A) retention kits, supports transcriptome differences with egg quality arose from differences in cytoplasmic polyadenylation or deadenylation of stored maternal transcripts as opposed to being the result of differences in the accumulation of maternal transcripts during oogenesis. Furthermore, few DETs were identified between medium- and high-quality eggs. Nine clusters of DETs which encompassed 375 DETs or about 40% of all DETs identified between high- and low-quality eggs, were associated with ribosome biogenesis and processing. The multitude of ribosome related DETs in low-quality eggs suggests inadequate ribosome production required for maternal mRNA translation, which could lead to a cascade of developmental dysfunction. This reduction in ribosomal gene expression was true of mitochondrial transcripts as well as nuclear transcripts. Moreover, mtDNA abundance was reduced by 1.6 log2FC in low-quality eggs compared with high-quality eggs further supporting the ability of the egg to provide energy following fertilization was compromised. In addition to genes associated with ribosome and mitochondrion biogenesis and function, GO analysis indicates levels of transcripts involved in the regulation of transcription, translation, cell division, apoptosis, and immune responses were altered in the low-quality eggs. Many of these genes have not previously been reported to contribute to egg quality in rainbow trout. The present study provides insights into how dysfunction of the egg transcriptome can affect developmental competence in fish eggs.
Methods
Sample collection
Eggs were collected from rainbow trout that were part of the selective breeding program at Troutlodge Inc. Sumner, WA, USA. Eggs from individual two-year-old broodstock rainbow trout were stripped into plastic bags. About 90 unfertilized eggs from each female were collected and immediately frozen in liquid nitrogen, and another 50 eggs were collected and placed into modified Davidson’s fixative [83] for examination to eliminate samples with overripe eggs or other abnormalities. The remaining eggs were fertilized with sperm harvested from neomales. The semen derived from each sire was used to fertilize eggs from two to three females. The fertilized eggs were incubated as individual families as part of the Troutlodge Inc. selective breeding program which evaluates eyeing rate at about 250 ATUs calculated as the sum of mean daily water temperature in degrees Celsius, which we refer to as eyeing in the present manuscript. The eggs were incubated at 10 °C for about the first 24 h post fertilization, after which time a sample of about 25–60 embryos were collected from each family and fixed in Stockard’s solution [84] to evaluate early embryonic survival and viability at about the 32-cell stage by enumerating the embryos reaching each stage of cell cleavage (Table 1). The frozen samples were kept in a − 80 °C freezer at Troutlodge Inc. until they were shipped on dry ice to the National Center for Cool and Cold Water Aquaculture (NCCCWA) after which they were again placed at − 80 °C until RNA isolation. The fixed samples were shipped at ambient temperature to NCCCWA for evaluation.
Selection of rainbow trout females for RNA-seq analysis of eggs
Selection of egg samples for RNA-seq analysis was based primarily on eyeing rate. A range of 130–218 individuals from each family were examined for survival and viability at eyeing. Dead and subviable eggs included those that were unfertilized, had precipitated yolk in response to shocking the eggs, or were considered to have poorly developed eyes. We considered an eyeing rate of 50% as the demarcation between fertile and subfertile families. Only 16 of the 192 families generated had eyeing rates that were less than 50% (Fig. 1). In addition, the sperm from the sires used for each of these matings also yielded at least one family with an eyeing rate greater than 78% confirming the sperm used for fertilization was not the cause of the poor eyeing rates. Ten of the subfertile families (0–47.5% eyeing) and 10 fertile or high-quality families with eyeing rates greater than 80% (80.6–97.7% eyeing) that shared sires with the ten subfertile families, were selected for RNA-seq analyses. Since there were no families with eyeing rates between 10 and 30%, we subsequently further divided the subfertile families as low- (0–4.2%) and medium- (30.6–47.5%) quality families or females. Visual examination of the fixed eggs from these 20 females revealed no obvious signs of poor egg quality before fertilization.
Assessment of early embryo development
All embryos collected at about 24 h post fertilization from each of the females selected for RNA-seq analysis were examined to determine viability. The fixed embryos were immersed in 0.5% methylene blue overnight. The cell number of each embryo was counted or confirmed to be greater than 32, using a stereo microscope (Nikon SMZ660). Those embryos with less than 32 cells were considered subviable.
RNA isolation and sequencing
RNAs were isolated from frozen eggs which were homogenized in Tri Reagent (Sigma, St. Louis, MO) with a Qiagen Retsch MM300 TissueLyser Shaker Mixer Grinder Agitator Mill (Retsch Inc., Haan, Germany). Total RNA was isolated following the manufacturer’s protocol with the modification of using Phase Lock Gel (5 PRIME, Inc., Gaithersburg, MD, USA) and Phase Separation Reagent (Molecular Research Center, Cincinnati, OH, USA) to separate the aqueous phase from the organic phase. The isolated RNAs were further purified by lithium chloride precipitation and treated with DNase. The RNA integrity was evaluated by gel electrophoresis (Additional file 2: Figure S1), a NanoDrop ND-1000 (Thermo SCIENTIFIC, Wilmington, DE, USA) with λ260/280 great than 1.98, and a 2100 Bioanalyzer (Santa Clara, CA, USA) with RNA Integrity Number between 7.2 and 9.1 (Additional file 1: Table S3). DNaseI treated RNA was used to construct libraries with the Illumina® TruSeq® Stranded Total RNA Library Prep Kit with Ribo-Zero Gold (rRNA removal) and TruSeq® Stranded mRNA Sample Prep Kit (poly(A) retention). The libraries were sequenced by HiSeq2500 with 100 nt paired-end reads. Raw reads were deposited in NCBI Sequence Read Archive database (SRA accession: SRP108797). We sequenced the libraries for rRNA removal in two batches. The first batch of four samples (99, 119, 108, and 129) had less than 4% of the reads aligned to rRNA whereas the second batch of 16 samples contained 14.4 to 55.7% rRNAs indicating the rRNAs were not effectively removed (Additional file 1: Table S4). Hence, the 16 samples were re-sequenced using a newly purchased kit which resulted in the effective removal of rRNAs to below an average of 6% of the reads. The data from the three rRNA removal sequencing runs were included in subsequent transcriptome analyses.
Data analysis
To classify reads as belonging to the nuclear transcriptome, the mitochondrial transcriptome, or as rRNA, after the adaptor was trimmed by bcl2fastq v2.17.1.14, the reads passing FastQC evaluation (http://www.bioinformatics.babraham.ac.uk/projects/fastqc), were respectively aligned to the nucleotide sequences corresponding to all CDS features annotated on the rainbow trout genome assembly (GCA_900005705.1) [85] with 72 additional genes selected from gene bank (Additional file 1: Table S5), the mitochondrial genome which includes 38 genes and the D-loop region [25, 86] and rRNA genes [87] by using Bowtie2 tool under default settings [88]. To avoid redundancy between the transcripts assigned to the mitochondrial and nuclear transcriptomes, we used the mitochondrial transcripts as queries to blast the rainbow trout [85] transcriptome data mentioned above. The blast results showed that GSONMT00007417001 was aligned with mitochondrial mt-rnr2, mt-tl1, and mt-nd1, and GSONMT00007419001 aligned with mt-nd4l. Therefore, GSONMT00007417001 and GSONMT00007419001 were removed from the nuclear transcriptome reference list. In addition, although mt-rnr1 and mt-rnr2 transcripts were included in the mitochondrial transcriptome, some of the rRNAs in the rRNA reference list have high similarity to these genes and therefore some of the reads aligned under rRNA may have been mt-rnr1 and mt-rnr2. The raw reads aligned to nuclear transcriptome and mitochondrial mRNAs were merged and used as input to DESeq2 to identify DETs among groups at a FDR < 0.05, log2FC greater than 1, and total number of normalized reads within comparisons being greater than 500.
Gene ontology analysis of the identified DETs was conducted using Blast2GO PRO platform (BioBam Bioinformatics S.L., Spain) [89]. An R script written according to the algorithms proposed by the Database for Annotation, Visualization and Integrated Discovery (DAVID) gene functional classification was used to cluster the identified DETs into functional gene modules based on the results of the GO analyses [81, 82]. Module enrichment scores were generated by calculating the geometric mean of the P-values which were derived from hypergeometric test of the input gene sets, followed by negative log transformation of the geometric mean [90]. False discovery rate was calculated using the Benjamini-Hochberg procedure. Spearman correlation coefficient for eyeing rate and early embryo viability was estimated using R (R × 64 3.3.0).
Mitochondrial gene quantification by real-time quantitative PCR
The insoluble materials leftover following homogenization in Tri Reagent during RNA isolation was mixed with 180 μl of 1 × TE buffer and shaken at speed setting 30 for 2 min with a Qiagen TissueLyser and then incubated for 10 min at room temperature. The mixture was centrifuged at 12,000 rpm for 10 min, and the supernatant was transferred to a tube containing Phase Lock Gel. The tube was shaken and centrifuged as above. The supernatant was collected, and DNA was isolated using a Quick-DNA™ Universal Kit (Zymo Research, Irvine, CA, USA). The DNA isolate was digested with RNaseA and the DNA was purified using a ZR-Duet™ DNA/RNA MiniPrep kit (Zymo Research, Irvine, CA, USA). The DNA from about 90 eggs was eluted in 20 μl of elution buffer and then diluted by adding 20 μl of water. The relative quantity of the mt-atp6 and mt-cyb genes was measured on an ABI 7900HT sequence detection system (Applied Biosystems, Foster City, CA). Each reaction consisted of 1.5 μL of diluted DNA, 3 μl of each primer (5 μM) and 1× SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA). The thermal cycling profile was 50 °C for 2 min, 95 °C for 10 min and 40 cycles of 95 °C for 30 s, 60 °C for 20 s, and 72 °C for 30 s. A final dissociation step was performed to assess the specificity of the reaction. Relative quantification of the mitochondrial DNA was estimated by the standard curve method with three technical replications, and mean differences of the mitochondrial DNA were reported as relative change using the value for the high-quality eggs as a calibrator. An 18 s rRNA gene was used as the reference control [91]. Primer sequences are shown in Additional file 1: Table S6.
Additional files
Table S1. Differentially expressed transcripts among low-, medium- and high-quality eggs identified by DESeq2. Table S2. Mean and log2FC of normalized mitochondrial DET reads. Table S3. Quality analyses for RNAs treated with DNase and submitted for RNA sequencing. Table S4. Overview of RNA-seq read alignments from the libraries constructed with defective rRNA removal kit. Table S5. Additional gene transcripts used as reference. Table S6. Primers used in real-time quantitative PCR. Table S7. Gene name abbreviations. (XLSX 506 kb)
Figure S1. Electrophoresis of egg RNAs isolated from different families selected for RNA sequencing. The families labeled in red are from the low-quality group; the families labeled in green are from the medium quality group; the families labeled in black are from the high-quality group. About 400 ng/sample of RNA was loaded to each well. Family 10 was not used for RNA sequencing. (PPTX 122 kb)
Acknowledgements
We thank Dr. Guangtu Gao for assistance in providing ribosomal reference sequences and help in data analysis and discussion; and Jill Birkett for assistance with embryo assessments. The use of trade, firm, or corporation names in this publication is for the information and convenience of the reader. Such use does not constitute an official endorsement or approval by the USDA or the ARS of any product or service to the exclusion of others that may be suitable. USDA is an equal opportunity provider and employer.
Funding
This work was supported by USDA-ARS CRIS Project 8082–31000-012. The funding bodies had no role in the design of the experiment, interpretation of the data, or writing of the manuscript.
Availability of data and materials
The sequences used in this study were deposited into NCBI Sequence Read Archive under accession number SPR108797 (https://www.ncbi.nlm.nih.gov/sra?term=SRP108797).
Abbreviations
- ATUs
Accumulated thermal units
- DAVID
Database for Annotation, Visualization and Integrated Discovery
- DET
Differentially expressed transcript
- FDR
False discovery rate
- GO
Gene ontology
- Log2FC
Log2 fold change
- MBT
Mid-blastula transition
- mtDNA
Mitochondrial DNA
- NCCCWA
National Center for Cool and Cold Water Aquaculture. Gene name abbreviations are presented in Additional file 1: Table S7
- Poly(A)
Polyadenylic acid
- ZGA
Zygotic genome activation
Authors’ contributions
G. W., H. M., K. M., and D. D. designed the study and collected the samples. A. G. H. consulted on design of the approach to the RNA-seq and evaluation of the data; and constructed the libraries and conducted the sequencing. H. M. analyzed the data. H. M. and G. W. drafted the manuscript and all authors contributed to the final version. All authors have read and approved the manuscript.
Ethics approval
All animal experiments were conducted under a protocol approved by the USDA/ARS National Center for Cool and Cold Water Aquaculture Institutional Animal Care and Use Committee (protocol #50).
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Hao Ma, Email: Hao.Ma@ars.usda.gov.
Kyle Martin, Email: Kyle.Martin@hendrix-genetics.com.
Doug Dixon, II, Email: Doug.Dixon@hendrix-genetics.com.
Alvaro G. Hernandez, Email: aghernan@illinois.edu
Gregory M. Weber, Phone: 1-304-724-8340, Email: greg.weber@ars.usda.gov
References
- 1.Brooks S, Tyler CR, Sumpter JP. Egg quality in fish: what makes a good egg? Rev Fish Biol Fish. 1997;7(4):387–416. [Google Scholar]
- 2.Vehvilainen H, Kause A, Koskinen H, Paananen T. Genetic architecture of rainbow trout survival from egg to adult. Genet Res. 2010;92(1):1–11. doi: 10.1017/S0016672310000017. [DOI] [PubMed] [Google Scholar]
- 3.Su GS, Liljedahl LE, Gall GAE. Genetic and environmental variation of female reproductive traits in rainbow trout (Oncorhynchus mykiss) Aquaculture. 1997;154(2):115–124. [Google Scholar]
- 4.Blom JH, Dabrowski K. Reproductive success of female rainbow-trout (Oncorhynchus-Mykiss) in response to graded dietary Ascorbyl monophosphate levels. Biol Reprod. 1995;52(5):1073–1080. doi: 10.1095/biolreprod52.5.1073. [DOI] [PubMed] [Google Scholar]
- 5.Bromage NRaC, P.R.T. (ed.): Egg production in rainbow trout. Croom Helm., London; 1988.
- 6.Palace VP, Werner J. Vitamins a and E in the maternal diet influence egg quality and early life stage development in fish: a review. Sci Mar. 2006;70:41–57. [Google Scholar]
- 7.Contreras-Sanchez WM, Schreck CB, Fitzpatrick MS, Pereira CB. Effects of stress on the reproductive performance of rainbow trout (Oncorhynchus mykiss) Biol Reprod. 1998;58(2):439–447. doi: 10.1095/biolreprod58.2.439. [DOI] [PubMed] [Google Scholar]
- 8.Campbell PM, Pottinger TG, Sumpter JP. Stress reduces the quality of gametes produced by rainbow-trout. Biol Reprod. 1992;47(6):1140–1150. doi: 10.1095/biolreprod47.6.1140. [DOI] [PubMed] [Google Scholar]
- 9.Bonnet E, Fostier A, Bobe J. Characterization of rainbow trout egg quality: a case study using four different breeding protocols, with emphasis on the incidence of embryonic malformations. Theriogenology. 2007;67(4):786–794. doi: 10.1016/j.theriogenology.2006.10.008. [DOI] [PubMed] [Google Scholar]
- 10.Aegerter S, Jalabert B. Effects of post-ovulatory oocyte ageing and temperature on egg quality and on the occurrence of triploid fry in rainbow trout, Oncorhynchus mykiss. Aquaculture. 2004;231(1–4):59–71. [Google Scholar]
- 11.Lahnsteiner F. Morphological, physiological and biochemical parameters characterizing the over-ripening of rainbow trout eggs. Fish Physiol Biochem. 2000;23(2):107–118. [Google Scholar]
- 12.Springate JRC, Bromage NR, Elliott JAK, Hudson DL. The timing of ovulation and stripping and their effects on the rates of fertilization and survival to eying, hatch and swim-up in the rainbow-trout (Salmo-Gairdneri R) Aquaculture. 1984;43(1–3):313–322. [Google Scholar]
- 13.Craik JCA, Harvey SM. Egg quality in rainbow-trout - the relation between egg viability, selected aspects of egg composition, and time of stripping. Aquaculture. 1984;40(2):115–134. [Google Scholar]
- 14.Tadros W, Lipshitz HD. The maternal-to-zygotic transition: a play in two acts. Development. 2009;136(18):3033–3042. doi: 10.1242/dev.033183. [DOI] [PubMed] [Google Scholar]
- 15.Lyman-Gingerich J, Pelegri F. Maternal factors in fish oogenesis and embryonic development. In: Babin PJ, Cerda J, Labadie K, editors. The fish oocyte: from basic studies to biotechnological applications. Dordrecht, The Netherlands: Springer; 2007. pp. 141–174. [Google Scholar]
- 16.Lubzens E, Bobe J, Young G, Sullivan CV. Maternal investment in fish oocytes and eggs: the molecular cargo and its contributions to fertility and early development. Aquaculture. 2017;472(1):37. [Google Scholar]
- 17.Sullivan CV, Chapman RW, Reading BJ, Anderson PE. Transcriptomics of mRNA and egg quality in farmed fish: some recent developments and future directions. Gen Comp Endocrinol. 2015;221:23–30. doi: 10.1016/j.ygcen.2015.02.012. [DOI] [PubMed] [Google Scholar]
- 18.Klangnurak W, Fukuyo T, Rezanujjaman MD, Seki M, Sugano S, Suzuki Y, Tokumoto T. Candidate gene identification of ovulation-inducing genes by RNA sequencing with an in vivo assay in zebrafish. PLoS One. 2018;13(5):e0196544. doi: 10.1371/journal.pone.0196544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Aegerter S, Jalabert B, Bobe J. mRNA stockpile and egg quality in rainbow trout (Oncorhynchus mykiss) Fish Physiol Biochem. 2003;28(1–4):317–318. [Google Scholar]
- 20.Aegerter S, Jalabert B, Bobe J. Large scale real-time PCR analysis of mRNA abundance in rainbow trout eggs in relationship with egg quality and post-ovulatory ageing. Mol Reprod Dev. 2005;72(3):377–385. doi: 10.1002/mrd.20361. [DOI] [PubMed] [Google Scholar]
- 21.Aegerter S, Jalabert B, Bobe J. Messenger RNA stockpile of cyclin B, insulin-like growth factor I, insulin-like growth factor II, insulin-like growth factor receptor Ib, and p53 in the rainbow trout oocyte in relation with developmental competence. Mol Reprod Dev. 2004;67(2):127–135. doi: 10.1002/mrd.10384. [DOI] [PubMed] [Google Scholar]
- 22.Bonnet E, Fostier A, Bobe J. Microarray-based analysis of fish egg quality after natural or controlled ovulation. BMC Genomics. 2007;8. [DOI] [PMC free article] [PubMed]
- 23.Bonnet E, Montfort J, Esquerre D, Hugot K, Fostier A, Bobe J. Effect of photoperiod manipulation on rainbow trout (Oncorhynchus mykiss) egg quality: a genomic study. Aquaculture. 2007;268(1–4):13–22. [Google Scholar]
- 24.Nagler JJ, Cavileer TD, Stoddard JW, Parsons JE. Maternal mRNA differences in unfertilized rainbow trout (Oncorhynchus mykiss) eggs from batches exhibiting variable embryonic survival. Cybium. 2008;32(2):233. [Google Scholar]
- 25.Ma H, Weber GM, Wei HR, Yao JB. Identification of mitochondrial genome-encoded small RNAs related to egg deterioration caused by postovulatory aging in rainbow trout. Mar Biotechnol. 2016;18(5):584–597. doi: 10.1007/s10126-016-9719-3. [DOI] [PubMed] [Google Scholar]
- 26.Ma H, Weber GM, Hostuttler MA, Wei H, Wang L, Yao J. MicroRNA expression profiles from eggs of different qualities associated with post-ovulatory ageing in rainbow trout (Oncorhynchus mykiss) BMC Genomics. 2015;16:201. doi: 10.1186/s12864-015-1400-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bachvarova RF. A maternal tail of poly(a) - the long and the short of it. Cell. 1992;69(6):895–897. doi: 10.1016/0092-8674(92)90606-d. [DOI] [PubMed] [Google Scholar]
- 28.Pique M, Lopez JM, Foissac S, Guigo R, Mendez R. A combinatorial code for CPE-mediated translational control. Cell. 2008;132(3):434–448. doi: 10.1016/j.cell.2007.12.038. [DOI] [PubMed] [Google Scholar]
- 29.Villalba A, Coll O, Gebauer F. Cytoplasmic polyadenylation and translational control. Curr Opin Genet Dev. 2011;21(4):452–457. doi: 10.1016/j.gde.2011.04.006. [DOI] [PubMed] [Google Scholar]
- 30.Cui J, Sartain CV, Pleiss JA, Wolfner MF. Cytoplasmic polyadenylation is a major mRNA regulator during oogenesis and egg activation in Drosophila. Dev Biol. 2013;383(1):121–131. doi: 10.1016/j.ydbio.2013.08.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Slater DW, Slater I, Gillespie D. Post-fertilization synthesis of Polyadenylic acid in sea-urchin embryos. Nature. 1972;240(5380):333. doi: 10.1038/240333a0. [DOI] [PubMed] [Google Scholar]
- 32.Rosenthal ET, Tansey TR, Ruderman JV. Sequence-specific Adenylations and Deadenylations accompany changes in the translation of maternal messenger-Rna after fertilization of Spisula oocytes. J Mol Biol. 1983;166(3):309–327. doi: 10.1016/s0022-2836(83)80087-4. [DOI] [PubMed] [Google Scholar]
- 33.Paris J, Philippe M. Poly(a) metabolism and polysomal recruitment of maternal mRNAs during early Xenopus development. Dev Biol. 1990;140(1):221–224. doi: 10.1016/0012-1606(90)90070-y. [DOI] [PubMed] [Google Scholar]
- 34.Brevini-Gandolfi TAL, Favetta LA, Mauri L, Luciano AM, Cillo F, Gandolfi F. Changes in poly(a) tail length of maternal transcripts during in vitro maturation of bovine oocytes and their relation with developmental competence. Mol Reprod Dev. 1999;52(4):427–433. doi: 10.1002/(SICI)1098-2795(199904)52:4<427::AID-MRD12>3.0.CO;2-G. [DOI] [PubMed] [Google Scholar]
- 35.Blower MD, Jambhekar A, Schwarz DS, Toombs JA. Combining different mRNA capture methods to analyze the transcriptome: analysis of the Xenopus laevis transcriptome. PLoS One. 2013;8(10). [DOI] [PMC free article] [PubMed]
- 36.Gohin M, Fournier E, Dufort I, Sirard MA. Discovery, identification and sequence analysis of RNAs selected for very short or long poly a tail in immature bovine oocytes. Mol Hum Reprod. 2014;20(2):127–138. doi: 10.1093/molehr/gat080. [DOI] [PubMed] [Google Scholar]
- 37.Subtelny AO, Eichhorn SW, Chen GR, Sive H, Bartel DP. Poly(a)-tail profiling reveals an embryonic switch in translational control. Nature. 2014;508(7494):66. doi: 10.1038/nature13007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Reyes JM, Chitwood JL, Ross PJ. RNA-Seq profiling of single bovine oocyte transcript abundance and its modulation by cytoplasmic polyadenylation. Mol Reprod Dev. 2015;82(2):103–114. doi: 10.1002/mrd.22445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Meijer HA, Bushell M, Hill K, Gant TW, Willis AE, Jones P, de Moor CH. A novel method for poly(a) fractionation reveals a large population of mRNAs with a short poly(a) tail in mammalian cells. Nucleic Acids Res. 2007;35(19). [DOI] [PMC free article] [PubMed]
- 40.Cabada MO, Darnbrough C, Ford PJ, Turner PC. Differential accumulation of two size classes of poly(a) associated with messenger RNA during oogenesis in Xenopus laevis. Dev Biol. 1977;57(2):427–439. doi: 10.1016/0012-1606(77)90227-5. [DOI] [PubMed] [Google Scholar]
- 41.Nagler JJ, Parsons JE, Cloud JG. Single pair mating indicates maternal effects on embryo survival in rainbow trout, Oncorhynchus mykiss. Aquaculture. 2000;184(1–2):177–183. [Google Scholar]
- 42.Jensen JOT. New mechanical shock sensitivity units in support of criteria for protection of salmonid eggs from blasting or seismic disturbance. Can Tech Rep Fish Aquat Sci. 2003(2452):27.
- 43.Jensen JOT, Alderdice DF. Comparison of mechanical shock sensitivity of eggs of five Pacific salmon (Oncorhynchus) species and steelhead trout (Salmo gairdneri) Aquaculture. 1989;78:19. [Google Scholar]
- 44.Marlow FL. Maternal Control of Development in Vertebrates. CA: My Mother Made Me Do It! San Rafael; 2010. [PubMed] [Google Scholar]
- 45.Kane DA, Kimmel CB. The zebrafish Midblastula transition. Development. 1993;119(2):447–456. doi: 10.1242/dev.119.2.447. [DOI] [PubMed] [Google Scholar]
- 46.Zamir E, Kam Z, Yarden A. Transcription-dependent induction of G1 phase during the zebra fish midblastula transition. Mol Cell Biol. 1997;17(2):529–536. doi: 10.1128/mcb.17.2.529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mathavan S, Lee SGP, Mak A, Miller LD, Murthy KRK, Govindarajan KR, Tong Y, Wu YL, Lam SH, Yang H, et al. Transcriptome analysis of zebrafish embryogenesis using microarrays. PLoS Genet. 2005;1(2):260–276. doi: 10.1371/journal.pgen.0010029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kleppe L, Edvardsen RB, Kuhl H, Malde K, Furmanek T, Drivenes O, Reinhardt R, Taranger GL, Wargelius A. Maternal 3'UTRs: from egg to onset of zygotic transcription in Atlantic cod. BMC Genomics. 2012;13:443. doi: 10.1186/1471-2164-13-443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hall TE, Smith P, Johnston IA. Stages of embryonic development in the Atlantic cod Gadus morhua. J Morphol. 2004;259(3):255–270. doi: 10.1002/jmor.10222. [DOI] [PubMed] [Google Scholar]
- 50.Kraeussling M, Wagner TU, Schartl M. Highly asynchronous and asymmetric cleavage divisions accompany early transcriptional activity in pre-blastula Medaka embryos. PLoS One. 2011;6(7). [DOI] [PMC free article] [PubMed]
- 51.Stoddard JW, Parsons JE, Nagler JJ. Early onset of embryonic mortality in sub-fertile families of rainbow trout (Oncorhynchus mykiss) Reprod Fertil Dev. 2005;17(8):785–790. doi: 10.1071/rd05087. [DOI] [PubMed] [Google Scholar]
- 52.Fernandez-Silva P, Enriquez JA, Montoya J. Replication and transcription of mammalian mitochondrial DNA. Exp Physiol. 2003;88(1):41–56. doi: 10.1113/eph8802514. [DOI] [PubMed] [Google Scholar]
- 53.Temperley RJ, Wydro M, Lightowlers RN, Chrzanowska-Lightowlers ZM. Human mitochondrial mRNAs-like members of all families, similar but different. Bba-Bioenergetics. 2010;1797(6–7):1081–1085. doi: 10.1016/j.bbabio.2010.02.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Curanovic D, Cohen M, Singh I, Slagle CE, Leslie CS, Jaffrey SR. Global profiling of stimulus-induced polyadenylation in cells using a poly(a) trap. Nat Chem Biol. 2013;9(11):671. doi: 10.1038/nchembio.1334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Richter JD. Cytoplasmic polyadenylation in development and beyond. Microbiol Mol Biol R. 1999;63(2):446–44+. doi: 10.1128/mmbr.63.2.446-456.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wai T, Ao A, Zhang XY, Cyr D, Dufort D, Shoubridge EA. The role of mitochondrial DNA copy number in mammalian fertility. Biol Reprod. 2010;83(1):52–62. doi: 10.1095/biolreprod.109.080887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Chappel S. The role of mitochondria from mature oocyte to viable blastocyst. Obstet Gynecol Int. 2013;2013:183024. doi: 10.1155/2013/183024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Artuso L, Romano A, Verri T, Domenichini A, Argenton F, Santorelli FM, Petruzzella V. Mitochondrial DNA metabolism in early development of zebrafish (Danio rerio) Bba-Bioenergetics. 2012;1817(7):1002–1011. doi: 10.1016/j.bbabio.2012.03.019. [DOI] [PubMed] [Google Scholar]
- 59.St John J. The control of mtDNA replication during differentiation and development. Bba-Gen Subjects. 2014;1840(4):1345–1354. doi: 10.1016/j.bbagen.2013.10.036. [DOI] [PubMed] [Google Scholar]
- 60.Bai J, Solberg C, Fernandes JMO, Johnston IA. Profiling of maternal and developmental-stage specific mRNA transcripts in Atlantic halibut Hippoglossus hippoglossus. Gene. 2007;386:202–210. doi: 10.1016/j.gene.2006.09.012. [DOI] [PubMed] [Google Scholar]
- 61.Schatten H, Sun QY, Prather R. The impact of mitochondrial function/dysfunction on IVF and new treatment possibilities for infertility. Reprod Biol Endocrin. 2014;12:1–11. doi: 10.1186/1477-7827-12-111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Shoubridge EA, Wai T. Mitochondrial DNA and the mammalian oocyte. Mitochondrion Germline Early Dev. 2007;77:87–111. doi: 10.1016/S0070-2153(06)77004-1. [DOI] [PubMed] [Google Scholar]
- 63.Sanchez-Velar N, Udofia EB, Yu Z, Zapp ML. hRIP, a cellular cofactor for rev function, promotes release of HIV RNAs from the perinuclear region. Genes Dev. 2004;18(1):23–34. doi: 10.1101/gad.1149704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Catrina IE, Bayer LV, Yanez G, McLaughlin JM, Malaczek K, Bagaeva E, Marras SAE, Bratu DP. The temporally controlled expression of Drongo, the fruit fly homolog of AGFG1, is achieved in female germline cells via P-bodies and its localization requires functional Rab11. RNA Biol. 2016;13(11):1117–1132. doi: 10.1080/15476286.2016.1218592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Gentil BJ, Delphin C, Mbele GO, Deloulme JC, Ferro M, Garin J, Baudier J. The giant protein AHNAK is a specific target for the calcium- and zinc-binding S100B protein - potential implications for Ca2+ homeostasis regulation by S100B. J Biol Chem. 2001;276(26):23253–23261. doi: 10.1074/jbc.M010655200. [DOI] [PubMed] [Google Scholar]
- 66.Doidge R, Mittal S, Aslam A, Winkler GS. The anti-proliferative activity of BTG/TOB proteins is mediated via the Caf1a (CNOT7) and Caf1b (CNOT8) Deadenylase subunits of the Ccr4-not complex. PLoS One. 2012;7(12). [DOI] [PMC free article] [PubMed]
- 67.Matsuda S, KawamuraTsuzuku J, Ohsugi M, Yoshida M, Emi M, Nakamura Y, Onda M, Yoshida Y, Nishiyama A, Yamamoto T. Tob, a novel protein that interacts with p185(erbB2), is associated with antiproliferative activity. Oncogene. 1996;12(4):705–713. [PubMed] [Google Scholar]
- 68.Mauxion F, Chen CYA, Seraphin B, Shyu AB. BTG/TOB factors impact deadenylases. Trends Biochem Sci. 2009;34(12):640–647. doi: 10.1016/j.tibs.2009.07.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Migaud H, Bell G, Cabrita E, McAndrew B, Davie A, Bobe J, Herraez MP, Carrillo M. Gamete quality and broodstock management in temperate fish. Rev Aquacult. 2013;5:S194–S223. [Google Scholar]
- 70.Chapman RW, Reading BJ, Sullivan CV. Ovary transcriptome profiling via artificial intelligence reveals a transcriptomic fingerprint predicting egg quality in striped bass, Morone saxatilis. PLoS One. 2014;9(5). [DOI] [PMC free article] [PubMed]
- 71.Zardoya R, GarridoPertierra A, Bautista JM. The complete nucleotide sequence of the mitochondrial DNA genome of the rainbow trout, Oncorhynchus mykiss. J Mol Evol. 1995;41(6):942–951. doi: 10.1007/BF00173174. [DOI] [PubMed] [Google Scholar]
- 72.Slomovic S, Laufer D, Geiger D, Schuster G. Polyadenylation and degradation of human mitochondrial RNA: the prokaryotic past leaves its mark. Mol Cell Biol. 2005;25(15):6427–6435. doi: 10.1128/MCB.25.15.6427-6435.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Baserga SJ, Linnenbach AJ, Malcolm S, Ghosh P, Malcolm ADB, Takeshita K, Forget BG, Benz EJ. Polyadenylation of a human mitochondrial ribosomal-Rna transcript detected by molecular-cloning. Gene. 1985;35(3):305–312. doi: 10.1016/0378-1119(85)90009-5. [DOI] [PubMed] [Google Scholar]
- 74.Rorbach J, Minczuk M. The post-transcriptional life of mammalian mitochondrial RNA. Biochem J. 2012;444:357–373. doi: 10.1042/BJ20112208. [DOI] [PubMed] [Google Scholar]
- 75.Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12). [DOI] [PMC free article] [PubMed]
- 76.Dutta A, Sinha DK. Zebrafish lipid droplets regulate embryonic ATP homeostasis to power early development. Open Biol. 2017;7(7). [DOI] [PMC free article] [PubMed]
- 77.Wendling NC, Bencic DC, Nagler JJ, Cloud JG, Ingermann RL. Adenosine triphosphate levels in steelhead (Oncorhynchus mykiss) eggs: an examination of turnover, localization and role. Comp Biochem Phys A. 2004;137(4):739–748. doi: 10.1016/j.cbpb.2004.02.009. [DOI] [PubMed] [Google Scholar]
- 78.Cagnone GLM, Tsai TS, Makanji Y, Matthews P, Gould J, Bonkowski MS, Elgass KD, Wong ASA, Wu LE, McKenzie M, et al. Restoration of normal embryogenesis by mitochondrial supplementation in pig oocytes exhibiting mitochondrial DNA deficiency. Sci Rep-Uk. 2016;6. [DOI] [PMC free article] [PubMed]
- 79.Dumollard R, Duchen M, Sardet C. Calcium signals and mitochondria at fertilisation. Semin Cell Dev Biol. 2006;17(2):314–323. doi: 10.1016/j.semcdb.2006.02.009. [DOI] [PubMed] [Google Scholar]
- 80.Ge HS, Tollner TL, Hu Z, Dai MM, Li XH, Guan HQ, Shan D, Zhang XJ, Lv JQ, Huang CJ, et al. The importance of mitochondrial metabolic activity and mitochondrial DNA replication during oocyte maturation in vitro on oocyte quality and subsequent embryo developmental competence. Mol Reprod Dev. 2012;79(6):392–401. doi: 10.1002/mrd.22042. [DOI] [PubMed] [Google Scholar]
- 81.Huang DW, Sherman BT, Tan Q, Collins JR, Alvord WG, Roayaei J, Stephens R, Baseler MW, Lane HC, Lempicki RA. The DAVID gene functional classification tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 2007;8(9). [DOI] [PMC free article] [PubMed]
- 82.Ma H, Gao G, Weber GM. Use of DAVID algorithms for clustering custom annotated gene lists in a non-model organism, rainbow trout. BMC Res Notes. 2018;11(63):1–6. doi: 10.1186/s13104-018-3154-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Hershberger WK, Hostuttler MA. Variation in time to first cleavage in rainbow trout Oncorhynchus mykiss embryos: a major factor in induction of tetraploids. J World Aquacult Soc. 2005;36(1):96–102. [Google Scholar]
- 84.Velsen FPJ. Embryonic development in eggs of sockeye salmon, Oncorhynchus nerka. Can Spec Publ Fish Aquat Sci. 1980;49:1–19. [Google Scholar]
- 85.Berthelot C, Brunet F, Chalopin D, Juanchich A, Bernard M, Noel B, Bento P, Da Silva C, Labadie K, Alberti A, et al. The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates. Nat Commun. 2014;5. [DOI] [PMC free article] [PubMed]
- 86.Kodama S, Yamada H, Annab L, Barrett JC. Elevated expression of mitochondrial cytochrome-B and Nadh dehydrogenase Subunit-4/4l genes in senescent human-cells. Exp Cell Res. 1995;219(1):82–86. doi: 10.1006/excr.1995.1207. [DOI] [PubMed] [Google Scholar]
- 87.Abernathy J, Overturf K. Comparison of ribosomal RNA removal methods for transcriptome sequencing workflows in teleost fish. Anim Biotechnol. 2016;27(1):60–65. doi: 10.1080/10495398.2015.1086365. [DOI] [PubMed] [Google Scholar]
- 88.Langmead B, Salzberg SL. Fast gapped-read alignment with bowtie 2. Nat Methods. 2012;9(4):357–U354. doi: 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Conesa A, Gotz S. Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics. 2008;2008:619832. doi: 10.1155/2008/619832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi: 10.1038/nprot.2008.211. [DOI] [PubMed] [Google Scholar]
- 91.Zhang HS, Maguire D, Swarts S, Sun WM, Yang SM, Wang W, Liu CM, Zhang M, Zhang D, Zhang L, et al. Replication of murine mitochondrial DNA following irradiation. Adv Exp Med Biol. 2009;645:43–48. doi: 10.1007/978-0-387-85998-9_7. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. Differentially expressed transcripts among low-, medium- and high-quality eggs identified by DESeq2. Table S2. Mean and log2FC of normalized mitochondrial DET reads. Table S3. Quality analyses for RNAs treated with DNase and submitted for RNA sequencing. Table S4. Overview of RNA-seq read alignments from the libraries constructed with defective rRNA removal kit. Table S5. Additional gene transcripts used as reference. Table S6. Primers used in real-time quantitative PCR. Table S7. Gene name abbreviations. (XLSX 506 kb)
Figure S1. Electrophoresis of egg RNAs isolated from different families selected for RNA sequencing. The families labeled in red are from the low-quality group; the families labeled in green are from the medium quality group; the families labeled in black are from the high-quality group. About 400 ng/sample of RNA was loaded to each well. Family 10 was not used for RNA sequencing. (PPTX 122 kb)
Data Availability Statement
The sequences used in this study were deposited into NCBI Sequence Read Archive under accession number SPR108797 (https://www.ncbi.nlm.nih.gov/sra?term=SRP108797).



