Abstract
Acinetobacter baumannii causes severe, fulminant, community-acquired pneumonia (CAP) in tropical and subtropical regions. We compared the population structure, virulence and antimicrobial resistance determinants of northern Australian community-onset A. baumannii strains with local and global strains. We performed whole-genome sequencing on 55 clinical and five throat colonization A. baumannii isolates collected in northern Australia between 1994 and 2016. Clinical isolates included CAP (n=41), healthcare-associated pneumonia (n=7) and nosocomial bloodstream (n=7) isolates. We also included 93 publicly available international A. baumannii genome sequences in the analyses. Patients with A. baumannii CAP were almost all critically unwell; 82 % required intensive care unit admission and 18 % died during their inpatient stay. Whole-genome phylogenetic analysis demonstrated that community-onset strains were not phylogenetically distinct from nosocomial strains. Some non-multidrug-resistant local strains were closely related to multidrug-resistant strains from geographically distant locations. Pasteur sequence type (ST)10 was the dominant ST and accounted for 31/60 (52 %) northern Australian strains; the remainder belonged to a diverse range of STs. The most recent common ancestor for ST10 was estimated to have occurred in 1738 (95 % highest posterior density, 1626–1826), with evidence of multiple introduction events between Australia and Southeast Asia between then and the present day. Virulence genes associated with biofilm formation and the type 6 secretion system (T6SS) were absent in many strains, and were not associated with in-hospital mortality. All strains were susceptible to gentamicin and meropenem; none carried an AbaR resistance island. Our results suggest that international dissemination of A. baumannii is occurring in the community on a contemporary timescale. Genes associated with biofilm formation and the T6SS may not be required for survival in community niches. The relative contributions of host and bacterial factors to the clinical severity of community-onset A. baumannii infection require further investigation.
Keywords: Acinetobacter baumannii, community-acquired pneumonia, whole-genome sequencing, epidemiology, virulence, antimicrobial resistance
Data Summary
Raw sequence data for northern and central Australian A. baumannii strains sequenced as part of this study are available in the Short Read Archive in Bioproject PRJNA478282 (https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA478282) and accession numbers are listed in Table S1, available in the online version of this article. Accession numbers for global A. baumannii strains used in the analyses are listed in Table S2. GenBank accession numbers for annotated K loci are included in Table S3.
Impact Statement.
Although best known for causing multidrug-resistant infection in hospitals, Acinetobacter baumannii is also an important cause of severe community-acquired pneumonia in tropical regions. This condition usually affects individuals with hazardous alcohol consumption, and often leads to death. In this study, we used whole-genome sequencing to investigate the genetic relationship between northern Australian community-onset A. baumannii strains and a collection of global strains. We also investigated the relationship between putative virulence genes and clinical outcomes. We found that some community-onset non-multidrug-resistant study strains were closely related to multidrug-resistant A. baumannii strains from geographically distant international locations, suggesting that global spread of A. baumannii is occurring in the community at the present day. We found variation in the profile of virulence determinants, although these were not associated with increased disease severity. We hypothesize that host factors play an important role in the susceptibility to and severity of community-onset A. baumannii infection.
Introduction
Acinetobacter baumannii is a ubiquitous Gram-negative bacterium which is a frequent cause of multidrug-resistant nosocomial infection across the globe. The ability to survive for prolonged periods in the hospital environment and to readily acquire antimicrobial resistance determinants via horizontal gene transfer have contributed to the propensity of this organism to cause hospital outbreaks involving multiple facilities [1]. Three major international global clones have emerged as the dominant lineages associated with hospital outbreaks [2].
In contrast to nosocomial-acquired A. baumannii , little is known about community-acquired disease, which predominantly occurs in tropical and subtropical climates with a seasonal peak during the rainy season [3]. Unlike nosocomial infection, community-onset disease is not associated with multidrug resistance and usually affects individuals with risk factors such as hazardous alcohol consumption and diabetes [3, 4]. The clinical presentation is typically with fulminant lobar pneumonia with septic shock, with in-hospital mortality as high as 64 % [5].
The relative contributions of host and bacterial factors to the severity of community-onset A. baumannii pneumonia are uncertain. Virulence determinants in A. baumannii are incompletely understood, although biofilm formation, iron acquisition, lipo-oligosaccharide, capsular polysaccharide, outer membrane proteins and protein secretion systems are thought to be important [6]. Furthermore, alcohol intoxication may contribute to A. baumannii pathogenesis via impairment of phagocytosis and by influencing bacterial gene expression [5, 7–9].
To further our understanding of the genomic epidemiology and pathogenesis of community-onset A. baumannii , we used whole-genome sequencing to characterize the largest series of community-onset A. baumannii clinical isolates to date. We used comparative genomics to determine the phylogenetic relationship between these strains and a large collection of publicly available global A. baumannii strains, and to explore virulence determinants and genes associated with antimicrobial resistance.
Methods
Isolates and clinical definitions
Sixty A. baumannii strains isolated between 1994 and 2016 were sequenced, including 55 isolates from patients with clinical infection, and five throat colonization isolates from community subjects without clinical evidence of infection (Table S1) [3, 10]. All available clinical A. baumannii isolates were sequenced. All apart from three clinical isolates were collected from patients at Royal Darwin Hospital, which is the referral centre for the tropical north (‘Top End’) of the Northern Territory, Australia. The Top End has an area of roughly 245 000 km2 and is sparsely populated. Darwin is the only city and has a population of approximately 122 000 people; the remaining population lives in towns or remote communities separated by vast geographical distances. Three isolates were from Alice Springs Hospital, in the desert region of Central Australia. The five colonization isolates had been collected from patients presenting from the community to the Royal Darwin Hospital Emergency Department with non-respiratory presentations and a history of excess alcohol consumption [10].
Clinical isolates were from patients with community-acquired pneumonia (CAP; n=41), community-onset of pneumonia but recent healthcare contact (HCAP; n=7) or nosocomial bloodstream infection (BSI; n=7). Pneumonia was defined as clinical and radiological evidence of pneumonia [3]. For CAP and HCAP, A. baumannii was isolated within 48 h of hospital admission, and in nosocomial BSI ≥48 h after hospital admission [3, 11]. Only one isolate per episode of infection was included. Bloodstream isolates were the preferred sample type if available, and were considered significant if there was pure growth of A. baumannii in blood cultures and/or there was concurrent growth of A. baumannii in blood cultures and sputum. In the absence of a bloodstream isolate, sputum and pleural fluid isolates were considered significant if there was pneumonia and no alternative pathogen was identified during the episode of infection.
Antimicrobial susceptibility testing, DNA extraction and whole genome sequencing
Antimicrobial susceptibility testing was performed on the Vitek 2 (bioMérieux) on all isolates except for two throat colonization isolates. Multidrug resistance was defined as resistance to two or more of the following drug classes: anti-pseudomonal cephalosporins, anti-pseudomonal carbapenems, fluoroquinolones and aminoglycosides [1]. Genomic DNA was extracted from the isolates using the DNeasy Blood and Tissue kit (Qiagen) according to the manufacturer’s instructions. Whole genome sequencing was performed at the Australian Genome Research Facility using Next-Era XT libraries and 100 bp paired-end reads on an Illumina HiSeq2500 sequencer. The raw sequence data are available in the Short Read Archive in Bioproject PRJNA478282.
Bioinformatic analysis
Genomes were assembled using a reference-assisted approach with MGAP (v0.0.1) [12]. The Pasteur multilocus sequence type (MLST) of the study isolates was determined using the whole genome assemblies and https://pubmlst.org/abaumannii/. Genomes were annotated with Prokka 1.10 [13]. An additional 93 A. baumannii strains were obtained from public databases, selected to represent diverse geographical locations and sequence types (STs) (Table S2). Multiple sequence alignment and variant calling was undertaken using the SPANDx pipeline (v3.2.1) [14]. A. baumannii strain ATCC 17978 was used as the reference genome [15]. Phylogenetic analyses were performed using PAUP* [16] for maximum-parsimony and IQ-TREE [17] for maximum-likelihood analyses. For maximum-likelihood analyses, model selection was undertaken using the Bayesian information criterion and 1000 bootstrap replicates were performed to select a consensus tree. Adjustment of maximum-likelihood phylogenies for recombination was done using ClonalFrameML [18]. Temporal analysis was performed with BEAST [19] using the approach outlined by Holt et al. [20]. Phylogenetic trees were visualized using Interactive Tree of Life [21]. We determined the presence or absence of the A. baumannii virulence genes from the Virulence Factor Database [22], the adeR/S two-component regulatory system [23] and the type 6 secretion system (T6SS) cluster [24] in our collection using SRST2 [25] and Large Scale Blast Score Ratio (LS-BSR) [26]. Genes associated with antimicrobial resistance were identified using Abricate [27] and the NCBI Bacterial Antimicrobial Resistance Reference Gene Database version 1 [28].
Results
A. baumannii CAP usually requires intensive care unit admission and has a high mortality rate
The study included 55 clinical A. baumannii isolates; 41 were from patients with CAP, seven with HCAP and seven with nosocomial BSI including six with bacteraemic hospital-acquired pneumonia and one with catheter-associated bloodstream infection. The clinical isolates included one isolate per episode of infection, and included 51 bloodstream isolates. Three sputum isolates and a pleural fluid isolate from four patients with severe CAP requiring intensive care unit (ICU) admission were also included; these four patients had negative blood cultures and no alternative pathogens were identified. Clinical details for a subset of CAP and HCAP episodes have been previously reported [3]; the demographic and clinical features of all 55 patients with clinical A. baumannii infection included in this study are summarized in Table 1. All but three patients were Indigenous Australians. The number of cases of HCAP and nosocomial BSI in this study was small, but in all groups ≥50 % required ICU admission, and mortality was high and ranged from 14 to 29 %.
Table 1. Demographics and severity indicators for CAP, HCAP and nosocomial BSI episodes.
| CAP (n=41) | HCAP (n=7) | Nosocomial BSI (n=7) | P-value | |
|---|---|---|---|---|
| Median age (years), (range) | 45.6 (29.8–68.8) | 42.1 (31.1–48.5) | 51.5 (41.2–76.1) | 0.11 |
| Male sex | 18/39 (46 %) | 4/7 (57 %) | 2/7 (29 %) | 0.55 |
| Indigenous | 33/36 (92 %) | 7/7 (100 %) | 6/6 (100 %) | 1.0 |
| Hazardous alcohol consumption | 25/32 (78 %) | 6/6 (100 %) | 2/6 (33 %) | 0.03 |
| ICU admission | 28/34 (82 %) | 5/7 (71 %) | 3/6 (50 %) | 0.14 |
| Inpatient death | 7/40 (18 %) | 1/7 (14 %) | 2/7 (29 %) | 0.84 |
Clinical A. baumannii isolates comprise diverse and novel STs, but ST10 is the most common ST among CAP, HCAP and nosocomial strains
In silico MLST by the Pasteur scheme revealed that ST10 was the most common ST in this study, and accounted for 31/60 (52 %) isolates including 24/41 (59 %) CAP isolates, 4/7 (57 %) HCAP isolates and 3/7 (43 %) nosocomial BSI isolates (Table S1). Patient outcomes were similar in the ST10 and non-ST10-infected groups. ICU admission was required in 21/28 (75 %) ST10-infected patients versus 15/19 (79 %) infected with other STs, and in-hospital death occurred in 5/31 (16 %) and 5/23 (22 %), respectively (Table 2).
Table 2. Outcomes of A. baumannii infection in the presence and absence of putative virulence attributes.
Fisher's exact test P-values were >0.05 for all comparisons
| Attribute (ST or virulence factor) present (+) or absent (–) | ICU admission | Died | |
|---|---|---|---|
| ST10 | + | 21/28 (75 %) | 5/31 (16 %) |
| − | 15/19 (79 %) | 5/23 (22 %) | |
| hemO | + | 23/30 (77 %) | 8/37 (22 %) |
| − | 13/17 (76 %) | 2/17 (12 %) | |
| csuA-E | + | 14/17 (82 %) | 4/20 (20 %) |
| − | 22/30 (73 %) | 6/34 (18 %) | |
| abaI/R | + | 9/11 (82 %) | 5/15 (33 %) |
| − | 27/36 (75 %) | 5/39 (13 %) | |
| adeR/S | + | 15/19 (79 %) | 4/22 (18 %) |
| − | 21/28 (75 %) | 6/32 (19 %) | |
| T6SS | + | 18/24 (75 %) | 7/30 (23 %) |
| − | 18/23 (78 %) | 3/24 (13 %) | |
The remaining isolates were a diverse range of STs; these included ST216 (n=5) and ST267 (n=2), with the remainder being singletons including nine novel STs (Table S1). Only one isolate belonged to one of the major international global clones (GC1); this was a throat colonization isolate that was ST1. Both community and nosocomial strains were diverse, with 15 different STs among the 41 CAP isolates and five different STs among the seven nosocomial isolates.
Whole-genome phylogenetic analysis shows that Australian community-onset and nosocomial strains are not unique, and are interspersed with global isolates
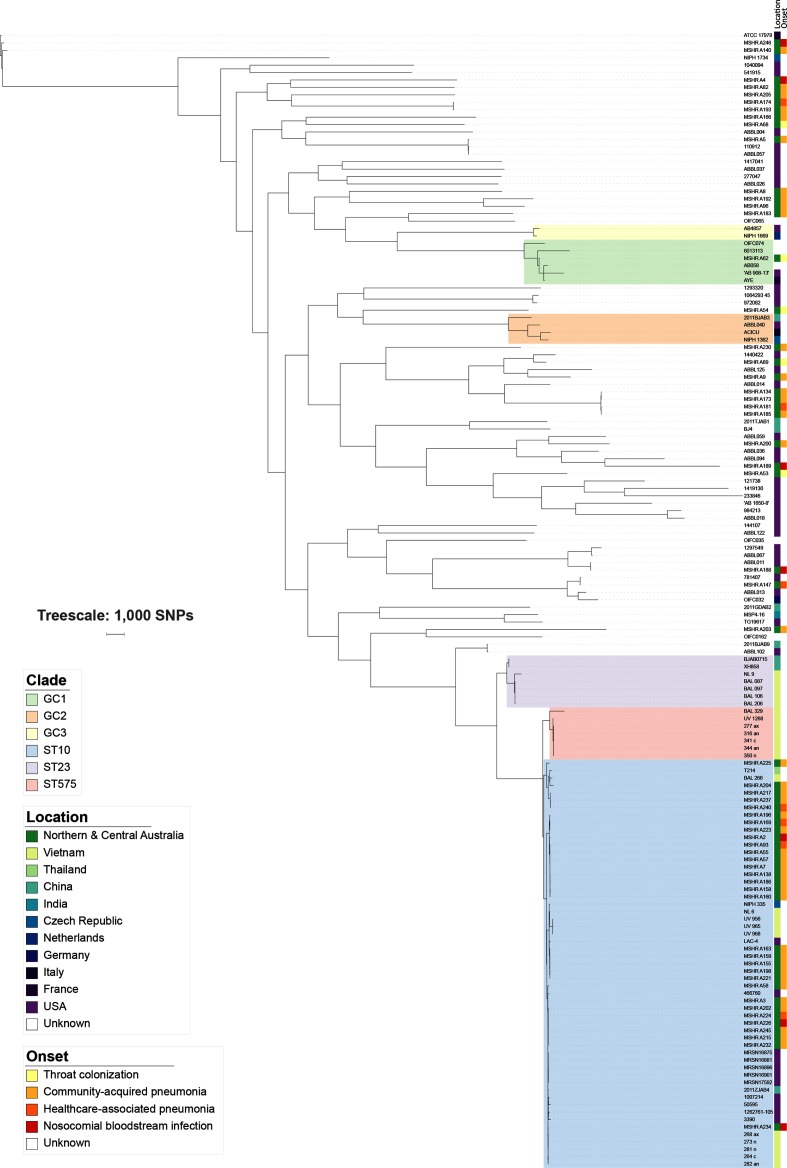
We undertook whole genome phylogenetic analysis of the 60 study isolates and an additional 93 international A. baumannii isolates selected for geographical and genetic diversity. The core alignment revealed 129 530 SNPs, which were used to inform a maximum-parsimony phylogeny (consistency index 0.2391, homoplasy index 0.7609; Fig. 1). Maximum-likelihood (Fig. S1) and recombination-adjusted (Fig. S2) analyses were also performed. In each of these analyses, the CAP, HCAP, nosocomial and colonization isolates were diverse and did not form distinct populations; some CAP strains were closely related to nosocomial BSI strains. Furthermore, the study isolates were spread among many different clades, and were interspersed with international isolates from diverse geographical locations throughout the phylogeny. There was no discernible phylogeographical signal.
Fig. 1.
Maximum-parsimony phylogenetic analysis including Australian study and global A. baumannii isolates.
Australian non-multidrug-resistant ST10 strains are closely related to international multidrug-resistant ST10 strains
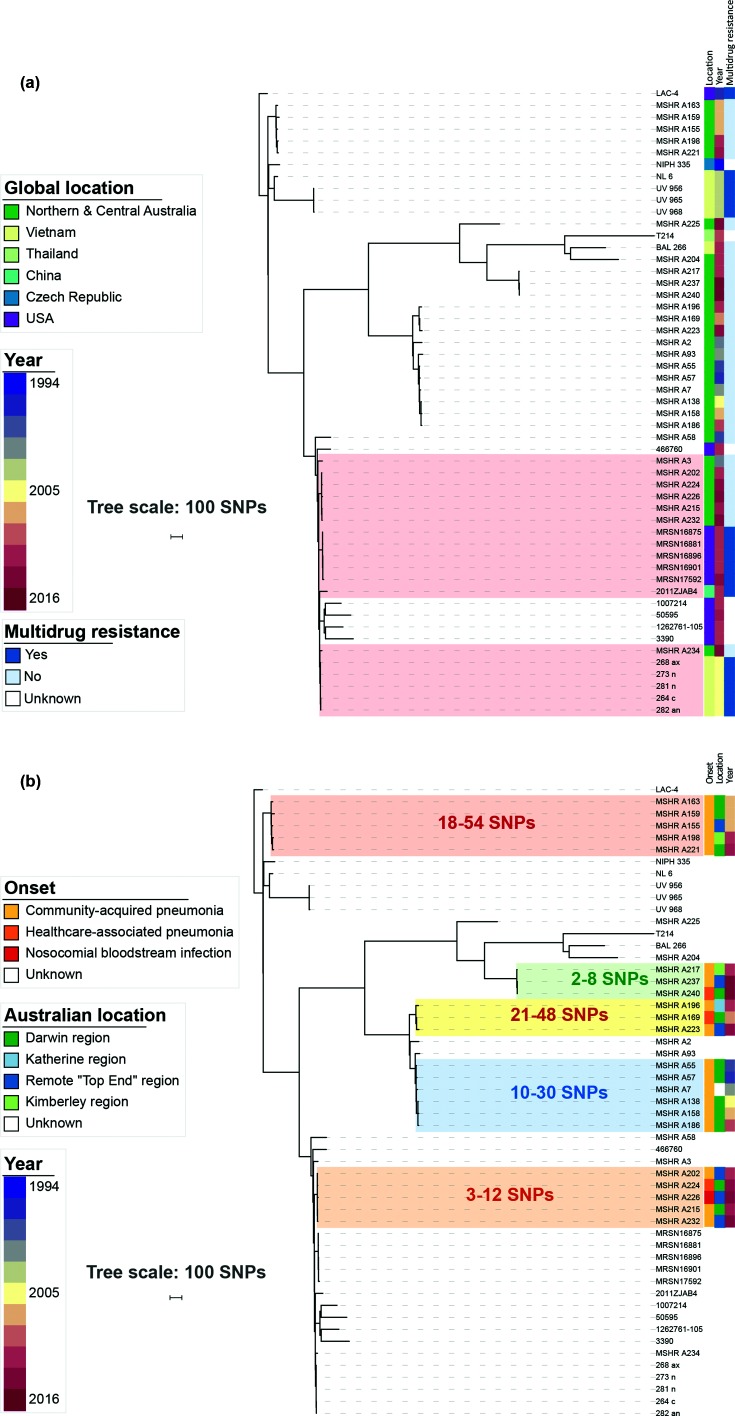
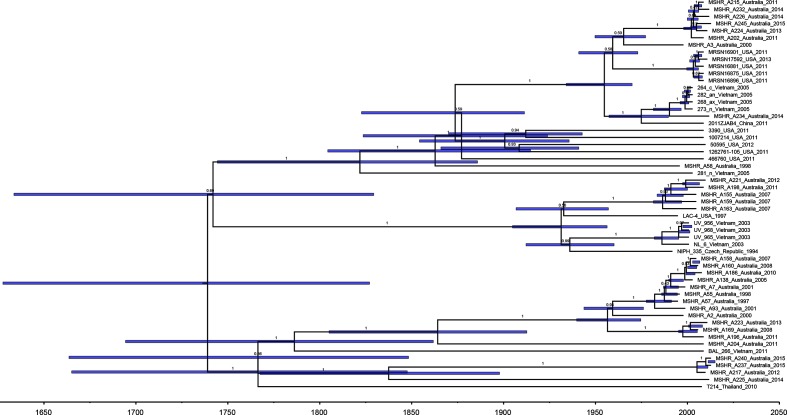
ST10 strains formed a single clade including several smaller clusters of closely related isolates. To examine this further, local and international ST10 sequences were mapped against the ST10 LAC-4 genome [29] resulting in a maximum-parsimony phylogeny based on 6560 core SNPs (Fig. 2). Maximum-likelihood and recombination-adjusted ST10 phylogenies are presented in Figs S3 and S4. A dated ST10 phylogeny is shown in Fig. 3. The most recent common ancestor (MRCA) for ST10 was estimated to date back to 1738 (95 % highest posterior density, HPD, 1626–1826). There were three deeply branching clades, each including strains from northern Australia and Southeast Asia, suggesting multiple introduction events between then and the present day.
Fig. 2.
Maximum-parsimony phylogenetic analysis of ST10 A. baumannii strains. (a) International and Australian isolates separated by <100 SNPs are highlighted. (b) Local Australian genomic clusters are highlighted, and the range of SNP distances between isolates in each cluster is listed.
Fig. 3.
Bayesian phylogenetic analysis of ST10 A. baumannii strains. Blue bars indicate 95% HPD. Branch labels indicate posterior probabilities. Timescale indicated in years.
There were several instances where our non-multidrug-resistant study strains were closely related to multidrug-resistant strains from geographically distant locations including Vietnam, the USA and China (Figs 2a and 3). A 2014 nosocomial BSI isolate from our study was separated from a cluster of colonization isolates collected from patients entering an ICU in Vietnam in 2005 [30] by 22 core SNPs, and was separated by 83 core SNPs from a 2011 carbapenem-resistant bloodstream isolate from Zhejiang, China (2011ZJAB4) [31]. Six community and nosocomial isolates from our study collected between 2011 and 2015 were separated from a clade of 2011 multidrug-resistant outbreak strains from Maryland, USA [32] (labelled ‘MRSN’ in the tree), by 59 core SNPs. The patients infected with these strains had no known epidemiological links to these international locations, and some of the patients were from very remote locations in the Top End. The MRCA for this clade was estimated to have occurred in 1957 (95 % HPD 1936–1972) (Fig. 3), the approximate time that both Australia and the USA were participating in the Vietnam war.
Also within the ST10 tree were five distinct genomic clusters of closely related study strains (Fig. 2b). Each of these clusters comprised community-onset strains; the clusters spanned 4–14 years with 4/5 clusters involving multiple northern Australian regions separated by hundreds of kilometres. Despite distances of as little as two SNPs separating some strains, the local genomic clusters each spanned a long time period and/or broad geographical space, and there were no clear epidemiological links between patients. The MRCAs for each of these clusters were estimated to have occurred between 7 and 22 years prior to the earliest strain in each cluster.
Putative virulence genes are absent in many northern Australian strains
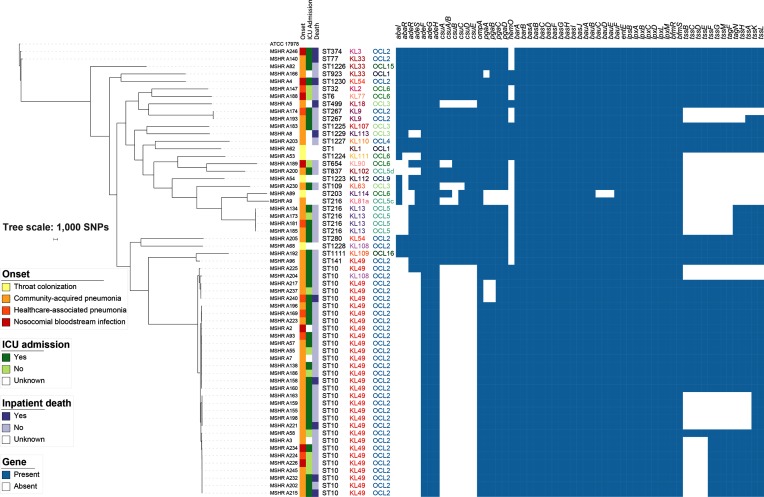
Interrogation of the study and international genomes for the presence of putative virulence genes revealed an absence of several virulence genes. The presence or absence of virulence genes varied according to lineage but not disease onset (Fig. 4). Associated with biofilm formation, the chaperone-usher type 1 pili system is encoded by the csuAB-E gene cluster [33]; 36/60 (60 %) study isolates did not have a complete csuAB-E cluster, and the locus was entirely absent in all ST10 strains. The abaI and abaR autoinducer synthase genes [34] are associated with quorum sensing and biofilm formation; one or both of these genes were absent in 43/60 (72 %) study isolates, and both were absent in all ST10 strains. The two-component regulatory system adeRS regulates expression of adeABC and is associated with increased biofilm formation [23]; one or both of these genes were absent in 35/60 (58 %) study strains and both were absent in all ST10 strains except one. T6SS genes were also absent in some isolates; the locus was either partly or completely absent in 28/60 (47 %) study isolates including 15/31 (48 %) ST10 strains. A haem utilization cluster including the hemO gene was previously found to be present in hypervirulent ST10 strains [29] and variably present in other A. baumannii genomes. The hemO gene was found in 39/60 (65 %) strains in our study, including all ST10 strains.
Fig. 4.
Virulence genes, and onset and outcomes of A. baumannii infection.
The proportion of patients infected with strains with and without these virulence factors requiring ICU admission and who died during their hospital admission is presented in Table 2. None of these virulence factors demonstrated a statistically significant association with ICU admission or inpatient death (Fisher's exact test P>0.05 for all comparisons), although the number of cases was small.
Northern Australian A. baumannii strains have diverse capsule K loci and lipo-oligosaccharide OC loci
In A. baumannii , the polysaccharide capsule which forms a protective layer on the cell surface is considered a major virulence determinant [35, 36]. The enzymes required for capsule biosynthesis are encoded at the K locus, but there is extensive variation at this site leading to differences in the composition and structure of the capsule [37]. Consistent with the diversity of the isolates in this collection, the study isolates had 22 different capsule types (Fig. 4, Table S1) including eight novel capsule types designated KL108–114; annotated sequences for these have been uploaded to GenBank, and accession numbers are listed in Table S3. All ST10 strains with the exception of one had the KL49 capsule locus associated with the production of 8-epilegionaminic acid. The remaining ST10 isolate carried KL108, which carries genes for legionaminic acid synthesis. The isolate carrying KL108 was closely related to the remaining local ST10 isolates, suggesting the replacement event had occurred recently and locally. Four isolates belonging to ST216 had the KL13 locus, which is associated with the production of acinetaminic acid, a recently described non-2-ulosonic acid that had not previously been identified in a natural biological source [38]. The outer core of lipo-oligosaccharide also varies significantly due variation in gene sets at the OC locus responsible for directing synthesis [39], and several different OC loci were also seen in the collection, including four that had not been detected previously (OCL13–16) (Fig. 4, Table S1).
Community-onset strains are not multidrug-resistant and do not harbour an AbaR resistance island
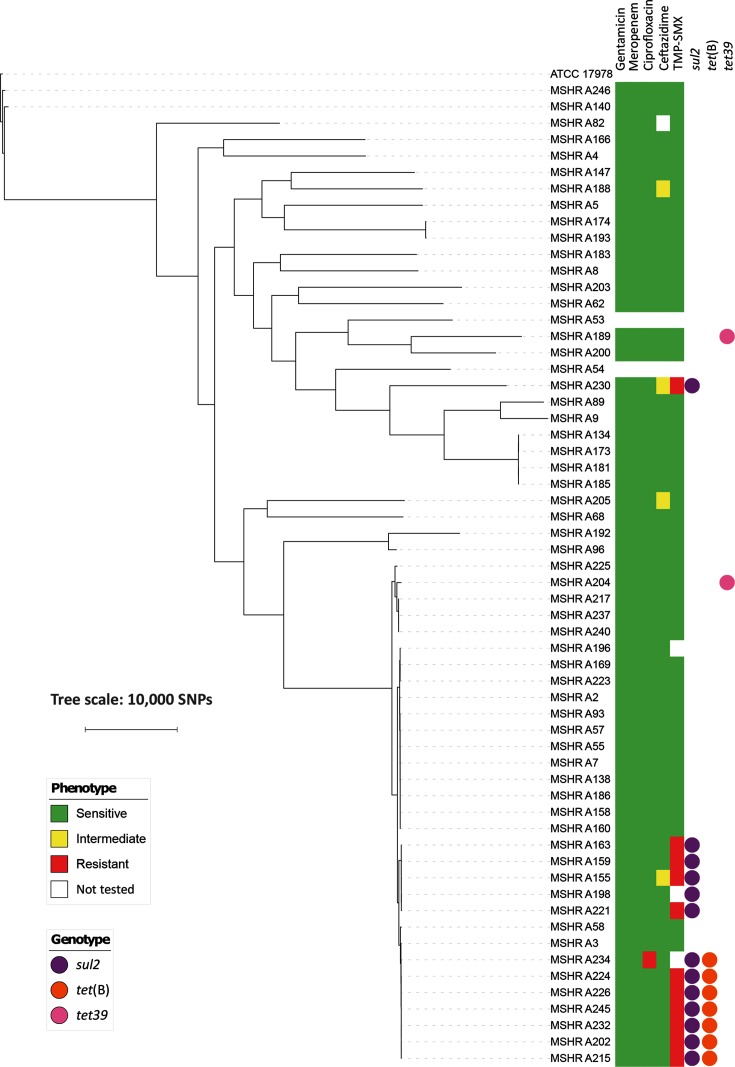
Antimicrobial susceptibility results were available for all isolates apart from two throat colonization isolates (Fig. 5). All isolates tested were susceptible to gentamicin and meropenem. One isolate was resistant to ciprofloxacin, four had intermediate susceptibility to ceftazidime, and 13 were resistant to trimethoprim-sulfamethoxazole (TMP-SMX). The comM gene is the usual site of insertion for AbaR resistance islands in GC1 and GC2 A. baumannii isolates, and was intact in 51/60 isolates. The nine strains with genes inserted within comM did not have genes associated with antimicrobial resistance at this site; instead, a wide variety of genes associated with metabolism, transcriptional regulation and heavy metal transport were identified.
Fig. 5.
Phenotypic antimicrobial susceptibilities and antimicrobial resistance genes.
The one ciprofloxacin-resistant isolate (MSHR_A234) was nosocomial, and had an S83L substitution in gyrA and an S80L substitution in parC. This combination of mutations is common in A. baumannii , is associated with high-level resistance to quinolones [40, 41] and was the probable cause of ciprofloxacin resistance in this strain. Isolates with phenotypic resistance to TMP-SMX all had the sul2 gene, known to be associated with resistance to sulfamethoxazole in Gram-negative bacteria [42]. In total, 10/11 of these isolates were ST10 strains. Seven of the TMP-SMX-resistant isolates also possessed tet(B) and an additional two isolates had the tet39 gene; these genes are associated with resistance to tetracyclines [43]. Phenotypic testing was not undertaken for this class of antimicrobial because it is not used for treatment of A. baumannii infection at our institution, and the clinical significance of the tet genes is uncertain.
Discussion
In this study, we used comparative genomics to characterize the largest reported collection of community-onset A. baumannii isolates. We found that northern Australian community-onset isolates were diverse, were not phylogenetically distinct from international isolates and in some cases were closely related to multidrug-resistant nosocomial strains from geographically distant locations [44]. We found no evidence of a difference in the presence of putative virulence genes in community-onset isolates. These findings suggest that the typical presentation of severe, rapidly progressing community-acquired Acinetobacter pneumonia may relate more to known host risk factors than to genetic variants in the infecting pathogen. There may be important as yet undiscovered virulence determinants among these strains, and a genome-wide association study using a larger and more diverse dataset could be used to uncover these. Another possible explanation is that acquisition of genes associated with antimicrobial resistance may have an attenuating effect on the virulence of nosocomial Acinetobacter strains, leading to a more indolent presentation.
The lack of any discernible geographical signal and the close evolutionary relationships between strains from geographically distant locations suggests that international dissemination of A. baumannii is occurring on a contemporary timescale. Temporal analysis revealed that the MRCA for ST10 was estimated to have occurred in approximately 1738, with multiple introduction events between northern Australia and Asia between then and the present day. This timescale is longer than for the emergent, multidrug-resistant GC1 lineage, which was estimated to have an MRCA as recently as 1960 [20]. Outside the healthcare setting, Acinetobacter species are generally considered to be environmental micro-organisms and A. baumannii has previously been isolated from soil and water in tropical and temperate climates [45–48]. A. baumannii also has community niches outside the natural environment, and colonizes humans, their lice, domestic and farm animals, and a wide variety of foods including fruit and vegetables, cheese, meat and milk [10, 45–47]. Each of these is a potential vessel for dissemination, and may contribute to spread of A. baumannii in the community, conceivably over large geographical distances.
Additionally, A. baumannii infection was described as a complication of trauma during the Vietnam war [49], more recently in the Middle East [50] and as a result of the 2006 conflict in East Timor [51], and has also occurred following natural disasters including the 2004 Asian tsunami [52]. Whether infecting strains are acquired from the individual’s microbiome, the local environment or from healthcare facilities is uncertain. Our temporal analyses suggested that the MRCA for a clade of closely related ST10 strains from northern Australia, Vietnam and USA was estimated to have occurred in approximately 1957 and we hypothesize that ST10 A. baumannii dissemination may have occurred as a result of repatriation of trauma victims during the Vietnam war.
ST10 was the most common ST seen among community and nosocomial isolates in this study. Given that the same clone dominated in both settings, we hypothesize that patients with nosocomial infection may have already been colonized at the time of hospital admission prior to developing infection with these strains. This is analogous to infection with methicillin-resistant Staphylococcus aureus , in which genomic sequencing has demonstrated intermingling of community and nosocomial strains [53]. It has previously been suggested on the basis of PFGE of community colonization and hospital A. baumannii strains in New York that the community is not a reservoir for hospital A. baumannii strains [5, 48], although our findings suggest this may not be the case in our region. This suggestion is also challenged by findings of a study using the Oxford MLST scheme in Taiwan, which found that half of the dominant nosocomial STs also caused community-onset disease [54].
Internationally, ST10 strains have been associated with colonization and nosocomial infection in an ICU in Vietnam [30], and an outbreak of extensively drug-resistant infection in the USA [32], suggesting that this ST exists in diverse geographical locations and has the capacity to colonize, cause disease and to acquire antimicrobial resistance determinants. Furthermore, it has previously been suggested that ST10 strains are more virulent than others; evidence to support this has included the high mortality rate associated with an outbreak in the USA among ‘relatively immunocompetent’ individuals [32], and demonstration of hypervirulence of ST10 strains in mouse models of infection [32, 55]. The majority of patients included in our study had severe pneumonia requiring ICU admission; however, within our cohort, ST10 strains were not associated with increased ICU admission or inpatient death.
The remaining community and nosocomial isolates in our study belonged to a wide range of diverse STs. This is consistent with the extensive ST diversity seen among both community and nosocomial strains in Taiwan, with unique STs found in 78 and 31 %, respectively [54]. There was no dominant community ST in Taiwan, but 17 % of community-onset isolates were Oxford ST447, an ST found in 27/31 (87 %) of our Pasteur ST10 strains including community-onset and nosocomial strains. Extensive ST diversity has also been observed in community A. baumannii strains outside the human clinical setting. A survey of soil, water, food, and domestic and farm animals in Lebanon identified 42 A. baumannii isolates belonging to 36 STs, including 24 novel STs [47]. These included two STs observed in our study, including an ST10 strain isolated from cow faeces and an ST216 strain isolated from cheese. In Reunion Island, 12 isolates from dogs and cats belonged to eight STs, and included one ST observed in our study; this was an ST203 strain isolated from a cat [46]. These observations highlight that the A. baumannii STs that cause severe disease may have diverse niches in wide-ranging geographical locations.
We observed that genes associated with biofilm formation and the T6SS varied according to lineage, potentially reflecting adaptation to different ecological and environmental conditions. The absence of T6SS genes and csuAB-E in clinical ST10 strains has been previously described [32, 56] and we observed the absence of these genes in addition to the absence of abaI/R and adeR/S in ST10 strains in our collection, despite the severity of clinical disease. The latter three loci are associated with biofilm formation and may be more important for persistence in the hospital environment than in other niches. This may explain why, although capable of causing nosocomial outbreaks [30, 32], ST10 is not a common cause of nosocomial A. baumannii infection globally. T6SS is thought to have a role in host colonization and killing of competing bacteria, and in Acinetobacter is hypothesized to have a role in the release and uptake of the DNA belonging to bacterial prey, potentially contributing to acquisition of antimicrobial resistance genes [57]. This may also provide a greater advantage for nosocomial compared to community strains.
The clinical significance of variation at the K locus is uncertain. The only previous study to assess the effect on virulence investigated the KL1 type and found that this was associated with increased survival in human ascites fluid and human serum, and increased virulence in a rat soft tissue model compared to KL1-negative strains [36]. The KL49 locus is associated with the production of 8-epilegionaminic acid, and has previously been found in the hypervirulent ST10 LAC-4 strain [58], the ST10 isolates associated with a fatal outbreak in the USA [32] and almost all ST10 strains in our collection. Recently, Oxford ST457 strains carrying KL49 were found to be associated with greater mortality and were more virulent in a Galleria mellonella infection model than other CC92 strains [59]. Whether the KL49 capsule type may contribute to increased virulence requires further investigation.
The main limitation of this study was the small sample size. We hypothesize that there are important yet-to-be discovered virulence factors that contributed to the severity of illness observed in our study. These could be identified using a genome-wide association study (GWAS) comparing various clinical outcomes, or comparing disease with colonization. Within our cohort of clinical cases, there was little heterogeneity in terms of disease severity, i.e. almost everyone was critically unwell. Additionally, there were only five colonization isolates. This led to inadequate power to perform GWAS using our current dataset. While there are a large number of publicly available A. baumannii genomes, there are few associated data relating to clinical outcomes, and data related to invasive disease versus colonization may be unreliable due to the clinical complexity of making this distinction. We therefore did not proceed with GWAS including publicly available strains as part of the current study.
Here, we have described the population structure, virulence and antimicrobial resistance determinants in a collection of predominantly community-onset A. baumannii strains from northern and central Australia. We found that ST10 was the dominant ST, and was a cause of CAP, HCAP and nosocomial A. baumannii infection in our setting. The remaining isolates were very diverse, and only one isolate belonged to one of the major international global clones. We found no evidence of phylogeographical restriction, and in some cases non-multidrug-resistant community-onset strains were closely related to multidrug-resistant strains from geographically distant locations. We hypothesize that dissemination may have occurred via humans, ectoparasites, food and/or inanimate objects. Virulence genes associated with biofilm formation and the T6SS were absent from many of our isolates, which suggests these attributes may not be required for survival in community niches. Although there was variation in the virulence gene profile of A. baumannii strains in this collection, pathogen factors contributing to the typical presentation of acute, fulminant pneumonia are uncertain. Key host risk factors such as hazardous alcohol intake and diabetes may play a greater role than variations in bacterial virulence genes in the pathogenesis of severe A. baumannii pneumonia, and further work investigating host–pathogen interactions is required.
Data Bibliography
Raw sequence data for northern and central Australian A. baumannii strains sequenced as part of this study are available in the Short Read Archive in Bioproject PRJNA478282 (https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA478282) and accession numbers are listed in Table S1. References and accession numbers for global A. baumannii strains used in the analyses are listed in Table S2.
Supplementary Data
Funding information
This work was supported by the National Health and Medical Research Council of Australia (Postgraduate Scholarship to E.M.M. #1114696, Fellowships to N.M.A. #1135820 and J.S.D. #1160331, and grants including the HOT NORTH initiative); and Advance Queensland (Fellowship to D.S.S. #AQRF13016-17RD2).
Acknowledgements
We would like to thank Associate Professor Rob Baird and the Department of Pathology at Royal Darwin Hospital for providing the A. baumannii isolates, and Dr Erin Price for advice regarding analyses.
Author contributions
The study was conceptualized by J.S.D., N.M.A., B.J.C., D.S.S. and E.M.M. Funding was acquired by J.S.D., N.M.A., B.J.C. and D.S.S. Data curation and investigation were undertaken by E.M.M., K.A.P., J.S.D. and N.M.A. Formal analysis was done by E.M.M., D.S.S., J.J.K. and R.M.H. E.M.M. wrote the original draft, and D.S.S., J.S.D., N.M.A., B.J.C., J.J.K. and R.M.H. revised and edited the manuscript.
Conflicts of interest
The authors declare that there are no conflicts of interest.
Ethical statement
The study was approved by the Human Research Ethics Committee of the Northern Territory Department of Health and Menzies School of Health Research #04/20.
Footnotes
Raw sequence data from this study are available in the Short Read Archive in Bioproject PRJNA478282 (https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA478282).
Abbreviations: BSI, bloodstream infection; CAP, community-acquired pneumonia; GWAS, genome-wide association study; HCAP, healthcare-acquired pneumonia; HPD, highest posterior density; ICU, intensive care unit; MLST, multilocus sequence typing; MRCA, most recent common ancestor; ST, sequence type; TMP-SMX, trimethoprim-sulfamethoxazole.
All supporting data, code and protocols have been provided within the article or through supplementary data files. Three supplementary tables and four supplementary figures are available with the online version of this article.
References
- 1.Peleg AY, Seifert H, Paterson DL. Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev. 2008;21:538–582. doi: 10.1128/CMR.00058-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Diancourt L, Passet V, Nemec A, Dijkshoorn L, Brisse S. The population structure of Acinetobacter baumannii: expanding multiresistant clones from an ancestral susceptible genetic pool. PLoS One. 2010;5:e10034. doi: 10.1371/journal.pone.0010034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Davis JS, McMillan M, Swaminathan A, Kelly JA, Piera KE, et al. A 16-year prospective study of community-onset bacteremic Acinetobacter pneumonia: low mortality with appropriate initial empirical antibiotic protocols. Chest. 2014;146:1038–1045. doi: 10.1378/chest.13-3065. [DOI] [PubMed] [Google Scholar]
- 4.Anstey NM, Currie BJ, Withnall KM. Community-acquired Acinetobacter pneumonia in the Northern Territory of Australia. Clin Infect Dis. 1992;14:83–91. doi: 10.1093/clinids/14.1.83. [DOI] [PubMed] [Google Scholar]
- 5.Dexter C, Murray GL, Paulsen IT, Peleg AY. Community-acquired Acinetobacter baumannii : clinical characteristics, epidemiology and pathogenesis. Expert Rev Anti Infect Ther. 2015;13:567–573. doi: 10.1586/14787210.2015.1025055. [DOI] [PubMed] [Google Scholar]
- 6.Harding CM, Hennon SW, Feldman MF. Uncovering the mechanisms of Acinetobacter baumannii virulence. Nat Rev Microbiol. 2018;16:91–102. doi: 10.1038/nrmicro.2017.148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Camarena L, Bruno V, Euskirchen G, Poggio S, Snyder M. Molecular mechanisms of ethanol-induced pathogenesis revealed by RNA-sequencing. PLoS Pathog. 2010;6:e1000834. doi: 10.1371/journal.ppat.1000834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Asplund MB, Coelho C, Cordero RJ, Martinez LR. Alcohol impairs J774.16 macrophage-like cell antimicrobial functions in Acinetobacter baumannii infection. Virulence. 2013;4:467–472. doi: 10.4161/viru.25641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gandhi JA, Ekhar VV, Asplund MB, Abdulkareem AF, Ahmadi M, et al. Alcohol enhances Acinetobacter baumannii-associated pneumonia and systemic dissemination by impairing neutrophil antimicrobial activity in a murine model of infection. PLoS One. 2014;9:e95707. doi: 10.1371/journal.pone.0095707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Anstey NM, Currie BJ, Hassell M, Palmer D, Dwyer B, et al. Community-acquired bacteremic Acinetobacter pneumonia in tropical Australia is caused by diverse strains of Acinetobacter baumannii, with carriage in the throat in at-risk groups. J Clin Microbiol. 2002;40:685–686. doi: 10.1128/JCM.40.2.685-686.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Friedman ND, Kaye KS, Stout JE, McGarry SA, Trivette SL, et al. Health care-associated bloodstream infections in adults: a reason to change the accepted definition of community-acquired infections. Ann Intern Med. 2002;137:791–797. doi: 10.7326/0003-4819-137-10-200211190-00007. [DOI] [PubMed] [Google Scholar]
- 12.Sarovich DS. Microbial genome assembler pipeline. https://github.com/dsarov/MGAP-Microbial-Genome-Assembler-Pipeline Available from. Accessed: June 17, 2017.
- 13.Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–2069. doi: 10.1093/bioinformatics/btu153. [DOI] [PubMed] [Google Scholar]
- 14.Sarovich DS, Price EP. SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets. BMC Res Notes. 2014;7:618. doi: 10.1186/1756-0500-7-618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Weber BS, Ly PM, Irwin JN, Pukatzki S, Feldman MF. A multidrug resistance plasmid contains the molecular switch for type VI secretion in Acinetobacter baumannii. Proc Natl Acad Sci USA. 2015;112:9442–9447. doi: 10.1073/pnas.1502966112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.PAUP* http://paup.sc.fsu.edu/about.html Available from. Accessed: June 17, 2017.
- 17.Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 2015;32:268–274. doi: 10.1093/molbev/msu300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Didelot X, Wilson DJ. ClonalFrameML: efficient inference of recombination in whole bacterial genomes. PLoS Comput Biol. 2015;11:e1004041. doi: 10.1371/journal.pcbi.1004041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 2018;4:vey016. doi: 10.1093/ve/vey016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Holt K, Kenyon JJ, Hamidian M, Schultz MB, Pickard DJ, et al. Five decades of genome evolution in the globally distributed, extensively antibiotic-resistant Acinetobacter baumannii global clone 1. Microb Genom. 2016;2:e000052. doi: 10.1099/mgen.0.000052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Letunic I, Bork P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016;44:W242–W245. doi: 10.1093/nar/gkw290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen L, Zheng D, Liu B, Yang J, Jin Q. VFDB 2016: hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Research. 2016;44:D694–D697. doi: 10.1093/nar/gkv1239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Richmond GE, Evans LP, Anderson MJ, Wand ME, Bonney LC, et al. The Acinetobacter baumannii two-component system AdeRS regulates genes required for multidrug efflux, biofilm formation, and virulence in a strain-specific manner. mBio. 2016;7:e00430. doi: 10.1128/mBio.00430-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Weber BS, Miyata ST, Iwashkiw JA, Mortensen BL, Skaar EP, et al. Genomic and functional analysis of the type VI secretion system in Acinetobacter . PLoS One. 2013;8:e55142. doi: 10.1371/journal.pone.0055142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Inouye M, Dashnow H, Raven LA, Schultz MB, Pope BJ, et al. SRST2: Rapid genomic surveillance for public health and hospital microbiology labs. Genome Med. 2014;6:90. doi: 10.1186/s13073-014-0090-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sahl JW, Caporaso JG, Rasko DA, Keim P. The large-scale blast score ratio (LS-BSR) pipeline: a method to rapidly compare genetic content between bacterial genomes. PeerJ. 2014;2:e332. doi: 10.7717/peerj.332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Seemann T. Abricate. https://github.com/tseemann/abricate Available from. Accessed: 11th February, 2018.
- 28.Bacterial Antimicrobial Resistance Reference Gene Database. https://www.ncbi.nlm.nih.gov/bioproject/PRJNA313047 Available from. accessed: November 2, 2017.
- 29.Ou HY, Kuang SN, He X, Molgora BM, Ewing PJ, et al. Complete genome sequence of hypervirulent and outbreak-associated Acinetobacter baumannii strain LAC-4: epidemiology, resistance genetic determinants and potential virulence factors. Sci Rep. 2015;5:8643. doi: 10.1038/srep08643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Schultz MB, Pham Thanh D, Tran do Hoan N, Wick RR, Ingle DJ, et al. Repeated local emergence of carbapenem-resistant Acinetobacter baumannii in a single hospital ward. Microb Genom. 2016;2:e000050. doi: 10.1099/mgen.0.000050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Li H, Liu F, Zhang Y, Wang X, Zhao C, et al. Evolution of carbapenem-resistant Acinetobacter baumannii revealed through whole-genome sequencing and comparative genomic analysis. Antimicrob Agents Chemother. 2015;59:1168–1176. doi: 10.1128/AAC.04609-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Jones CL, Clancy M, Honnold C, Singh S, Snesrud E, et al. Fatal outbreak of an emerging clone of extensively drug-resistant Acinetobacter baumannii with enhanced virulence. Clin Infect Dis. 2015;61:145–154. doi: 10.1093/cid/civ225. [DOI] [PubMed] [Google Scholar]
- 33.Tomaras AP, Flagler MJ, Dorsey CW, Gaddy JA, Actis LA. Characterization of a two-component regulatory system from Acinetobacter baumannii that controls biofilm formation and cellular morphology. Microbiology. 2008;154:3398–3409. doi: 10.1099/mic.0.2008/019471-0. [DOI] [PubMed] [Google Scholar]
- 34.Niu C, Clemmer KM, Bonomo RA, Rather PN. Isolation and characterization of an autoinducer synthase from Acinetobacter baumannii . J Bacteriol. 2008;190:3386–3392. doi: 10.1128/JB.01929-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Iwashkiw JA, Seper A, Weber BS, Scott NE, Vinogradov E, et al. Identification of a general O-linked protein glycosylation system in Acinetobacter baumannii and its role in virulence and biofilm formation. PLoS Pathog. 2012;8:e1002758. doi: 10.1371/journal.ppat.1002758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Russo TA, Luke NR, Beanan JM, Olson R, Sauberan SL, et al. The K1 capsular polysaccharide of Acinetobacter baumannii strain 307-0294 is a major virulence factor. Infect Immun. 2010;78:3993–4000. doi: 10.1128/IAI.00366-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kenyon JJ, Hall RM. Variation in the complex carbohydrate biosynthesis loci of Acinetobacter baumannii genomes. PLoS One. 2013;8:e62160. doi: 10.1371/journal.pone.0062160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kenyon JJ, Marzaioli AM, Hall RM, de Castro C. Structure of the K12 capsule containing 5,7-di-N-acetylacinetaminic acid from Acinetobacter baumannii isolate D36. Glycobiology. 2015;25:881–887. doi: 10.1093/glycob/cwv028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kenyon JJ, Nigro SJ, Hall RM. Variation in the OC locus of Acinetobacter baumannii genomes predicts extensive structural diversity in the lipooligosaccharide. PLoS One. 2014;9:e107833. doi: 10.1371/journal.pone.0107833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Vila J, Ruiz J, Goñi P, Jimenez de Anta T. Quinolone-resistance mutations in the topoisomerase IV parC gene of Acinetobacter baumannii . J Antimicrob Chemother. 1997;39:757–762. doi: 10.1093/jac/39.6.757. [DOI] [PubMed] [Google Scholar]
- 41.Vila J, Ruiz J, Goñi P, Marcos A, Jimenez de Anta T. Mutation in the gyrA gene of quinolone-resistant clinical isolates of Acinetobacter baumannii . Antimicrob Agents Chemother. 1995;39:1201–1203. doi: 10.1128/AAC.39.5.1201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Huovinen P. Resistance to trimethoprim-sulfamethoxazole. Clin Infect Dis. 2001;32:1608–1614. doi: 10.1086/320532. [DOI] [PubMed] [Google Scholar]
- 43.Akers KS, Mende K, Yun HC, Hospenthal DR, Beckius ML, et al. Tetracycline susceptibility testing and resistance genes in isolates of Acinetobacter baumannii-Acinetobacter calcoaceticus complex from a U.S. military hospital. Antimicrob Agents Chemother. 2009;53:2693–2695. doi: 10.1128/AAC.01405-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Farrugia DN, Elbourne LD, Hassan KA, Eijkelkamp BA, Tetu SG, et al. The complete genome and phenome of a community-acquired Acinetobacter baumannii . PLoS One. 2013;8:e58628. doi: 10.1371/journal.pone.0058628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Eveillard M, Kempf M, Belmonte O, Pailhoriès H, Joly-Guillou ML. Reservoirs of Acinetobacter baumannii outside the hospital and potential involvement in emerging human community-acquired infections. Int J Infect Dis. 2013;17:e802. doi: 10.1016/j.ijid.2013.03.021. [DOI] [PubMed] [Google Scholar]
- 46.Pailhoriès H, Belmonte O, Kempf M, Lemarié C, Cuziat J, et al. Diversity of Acinetobacter baumannii strains isolated in humans, companion animals, and the environment in Reunion Island: an exploratory study. Int J Infect Dis. 2015;37:64–69. doi: 10.1016/j.ijid.2015.05.012. [DOI] [PubMed] [Google Scholar]
- 47.Rafei R, Hamze M, Pailhoriès H, Eveillard M, Marsollier L, et al. Extrahuman epidemiology of Acinetobacter baumannii in Lebanon. Appl Environ Microbiol. 2015;81:2359–2367. doi: 10.1128/AEM.03824-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zeana C, Larson E, Sahni J, Bayuga SJ, Wu F, et al. The epidemiology of multidrug-resistant Acinetobacter baumannii: does the community represent a reservoir? Infect Control Hosp Epidemiol. 2003;24:275–279. doi: 10.1086/502209. [DOI] [PubMed] [Google Scholar]
- 49.Murray CK, Yun HC, Griffith ME, Hospenthal DR, Tong MJ. Acinetobacter infection: what was the true impact during the Vietnam conflict? Clin Infect Dis. 2006;43:383–384. doi: 10.1086/505601. [DOI] [PubMed] [Google Scholar]
- 50.Sebeny PJ, Riddle MS, Petersen K. Acinetobacter baumannii skin and soft-tissue infection associated with war trauma. Clin Infect Dis. 2008;47:444–449. doi: 10.1086/590568. [DOI] [PubMed] [Google Scholar]
- 51.Elston JW, Bannan CL, Chih DT, Boutlis CS. Acinetobacter spp. in gunshot injuries. Emerg Infect Dis. 2008;14:178–180. doi: 10.3201/eid1401.070878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Maegele M, Gregor S, Steinhausen E, Bouillon B, Heiss MM, et al. The long-distance tertiary air transfer and care of tsunami victims: injury pattern and microbiological and psychological aspects. Crit Care Med. 2005;33:1136–1140. doi: 10.1097/01.CCM.0000163269.42524.50. [DOI] [PubMed] [Google Scholar]
- 53.Popovich KJ, Snitkin ES, Hota B, Green SJ, Pirani A, et al. Genomic and epidemiological evidence for community origins of hospital-onset methicillin-resistant Staphylococcus aureus bloodstream infections. J Infect Dis. 2017;215:1640–1647. doi: 10.1093/infdis/jiw647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Chen C-T, Wang Y-C, Kuo S-C, Shih F-H, Chen T-L, et al. Community-acquired bloodstream infections caused by Acinetobacter baumannii: A matched case–control study. J Microbiol Immunol Infect. 2018;51:629–635. doi: 10.1016/j.jmii.2017.02.004. [DOI] [PubMed] [Google Scholar]
- 55.Harris G, Kuo Lee R, Lam CK, Kanzaki G, Patel GB, et al. A mouse model of Acinetobacter baumannii-associated pneumonia using a clinically isolated hypervirulent strain. Antimicrob Agents Chemother. 2013;57:3601–3613. doi: 10.1128/AAC.00944-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Chan AP, Sutton G, Depew J, Krishnakumar R, Choi Y, et al. A novel method of consensus pan-chromosome assembly and large-scale comparative analysis reveal the highly flexible pan-genome of Acinetobacter baumannii . Genome Biol. 2015;16:143. doi: 10.1186/s13059-015-0701-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Cooper RM, Tsimring L, Hasty J. Inter-species population dynamics enhance microbial horizontal gene transfer and spread of antibiotic resistance. Elife. 2017;6 doi: 10.7554/eLife.25950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Vinogradov E, Maclean L, Xu HH, Chen W. The structure of the polysaccharide isolated from Acinetobacter baumannii strain LAC-4. Carbohydr Res. 2014;390:42–45. doi: 10.1016/j.carres.2014.03.001. [DOI] [PubMed] [Google Scholar]
- 59.Zhou K, Tang X, Wang L, Guo Z, Xiao S, et al. An emerging clone (ST457) of Acinetobacter baumannii clonal complex 92 with enhanced virulence and increasing endemicity in South China. Clin Infect Dis. 2018;67:S179–S188. doi: 10.1093/cid/ciy691. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.