Abstract
We report the results of clinical exome sequencing (CES) on >2,200 previously unpublished Saudi families as a first-tier test. The predominance of autosomal-recessive causes allowed us to make several key observations. We highlight 155 genes that we propose to be recessive, disease-related candidates. We report additional mutational events in 64 previously reported candidates (40 recessive), and these events support their candidacy. We report recessive forms of genes that were previously associated only with dominant disorders and that have phenotypes ranging from consistent with to conspicuously distinct from the known dominant phenotypes. We also report homozygous loss-of-function events that can inform the genetics of complex diseases. We were also able to deduce the likely causal variant in most couples who presented after the loss of one or more children, but we lack samples from those children. Although a similar pattern of mostly recessive causes was observed in the prenatal setting, the higher proportion of loss-of-function events in these cases was notable. The allelic series presented by the wealth of recessive variants greatly expanded the phenotypic expression of the respective genes. We also make important observations about dominant disorders; these observations include the pattern of de novo variants, the identification of 74 candidate dominant, disease-related genes, and the potential confirmation of 21 previously reported candidates. Finally, we describe the influence of a predominantly autosomal-recessive landscape on the clinical utility of rapid sequencing (Flash Exome). Our cohort’s genotypic and phenotypic data represent a unique resource that can contribute to improved variant interpretation through data sharing.
Keywords: exome, clinical genomics, candidate genes, first-tier, genomics-first, phenotypic expansion, prenatal, fetal malformation, knockout, autozygome, gonadal mosaicism, expanded carrier screening, multilocus phenotypes, hybrid phenotype, dual diagnosis
Introduction
Consanguinity is an ancient human practice that persists to this day in many parts of the world because of a multitude of incentives.1, 2 A consanguineous loop renders the genomes of the offspring “autozygous” to an extent that correlates with the degree of haplotype sharing between the parents, and this is, in turn, a function of their degree of relatedness.3 Autozygosity is a rich source of homozygous alleles, the phenotypic expression of which spans the entire spectrum from embryonic lethal to clinically benign or even advantageous.4, 5 Furthermore, the telltale sign of autozygosity, i.e., extended regions of homozygosity delimited by crossing-over events that “shrink” the founder haplotype, is an additional advantage that makes the homozygous alleles of interest readily mappable.6 It is not surprising, therefore, that Saudi Arabia, which has a consanguinity rate of >50% and a high fertility rate, represents a unique resource for the study of this phenomenon and for exploring its relevance both to basic human genetics and applied clinical genetics.2, 7
Efforts to exploit these unique characteristics have indeed revealed many important observations, not least of which was the discovery of a large number of disease-related genes that were almost always recessive.8, 9, 10, 11 However, those efforts had two key limitations. First, the discovery of disease-related genes has largely been limited to specific phenotypes. Such a selective approach, although helpful in uncovering phenotype-specific variants, is clearly inefficient in achieving the goal of having all disease-related genes identified by 2020.12 Second, the “phenotype-first” approach risks missing affected individuals that have not been evaluated by experts in the delineation of that specific phenotype, individuals that have been adequately phenotyped but not enrolled in those phenotype-specific cohorts, and individuals that lack the typical phenotype.
Large-scale sequencing of all individuals with genetic diseases by any clinician who has the clinical suspicion, even if he or she lacks the necessary phenotyping expertise, offers an attractive approach that circumvents the historical limitations outlined above. Such a large scale “genomics-first” approach built on a central database was indeed the goal of the Saudi Human Genome Program, in order to exploit the unique characteristics of the Saudi population to the fullest extent possible.13 However, it became quickly apparent that most clinicians were much more interested in clinical testing than research-based testing of their patients. Therefore, we launched a program of clinical exome sequencing (CES).14 The recent decision by the Ministry of Health to cover the cost of CES as a standard clinical test allowed clinicians from around the country to order this as a first-tier test on any individual with a suspected genetic diagnosis. Combined with the fact that all CES tests were run by a central lab, this presented an unprecedented opportunity to create the most comprehensive view to date of clinically relevant variants across the entire spectrum of genetic diseases in a highly consanguineous population.
Material and Methods
Study Population
Informed consent was obtained from all affected individuals (or their legal guardians) as a strict requirement prior to CES. The consent also covered the option to receive or not to receive “secondary findings” defined as (1) ACMG (American College of Medical Genetics and Genomics) secondary findings15 and (2) carrier status for a predefined list of “known Saudi mutations (KSMs).”16 Venous blood samples collected in EDTA tubes were collected in the majority of instances for DNA extraction. For prenatal testing, DNA was extracted from chorionic villous or amniotic fluid samples. Where family history was negative, clinicians were encouraged to consider trio CES, although the choice was also given to only request CES on the index individual and then to run a Sanger test of segregation. Similarly, clinicians were given the choice between ordering CES on multiple affected members or to request a Sanger segregation test. For molecular autopsy by proxy (see below), samples were obtained from both parents of the deceased child(ren). Phenotypic data were provided as part of a standard paper form, although clinicians were encouraged to use free text to describe additional useful information. This study was approved by the local institutional review board (IRB) (KFSHRC RAC # 2181266).
Exome Sequencing and Variant Interpretation and Confirmation
For standard CES, we used the same protocol described before.14 Positional mapping was always incorporated in the pipeline regardless of the family history, as described before.14 All indel variants and only single base-pair substitutions that fell below our predetermined quality threshold were confirmed by Sanger sequencing, as described before.14 We also offered rapid exome sequencing with a turnaround time of <48 h (Flash Exome) as per the protocol shown in Figure S1. We used the ACMG guidelines for variant interpretation17 to classify the outcome of CES into three categories:
-
(1)
Solved: instances of individuals in which we identified pathogenic or likely pathogenic variants that explain the clinical indication in the expected zygosity. In the case of molecular autopsy by proxy for a suspected recessive phenotype in the deceased child(ren), both parents had to have, in the same gene, a pathogenic or likely pathogenic variant that explains the phenotype.
-
(2)
Ambiguous: these include instances of individuals with (1) a monoallelic pathogenic or likely pathogenic gene variant that explains a recessive phenotype, (2) a variant of unknown significance (VUS) that is in an established disease-related gene and that explains the phenotype, whether or not it is present in the expected zygosity for the disease, or (3) a VUS in a candidate gene. The variants in the latter category were carefully chosen to qualify as much as possible as pathogenic or likely pathogenic if the gene were established in the Online Mendelian Inheritance in Man (OMIM) database. In addition, only genes with compelling biological candidacy (special emphasis was made on animal models, but other lines of evidence were also pursued) were highlighted.
-
(3)
Negative: all other instances.
Segregation analysis was strongly encouraged, and we only report variants for which segregation analysis was either consistent with the presumed link to the disease or unavailable; all other variants that failed the segregation test were removed. Founder variants were defined as variants that are observed with a minor allele frequency (MAF) > 0 in the Saudi population or that are present in two affected individuals who are not directly related but share the same haplotype and phenotype.
Results
The Diagnostic Yield of CES As A First-Tier Test Across Genetic Disorders in a Highly Consanguineous Population
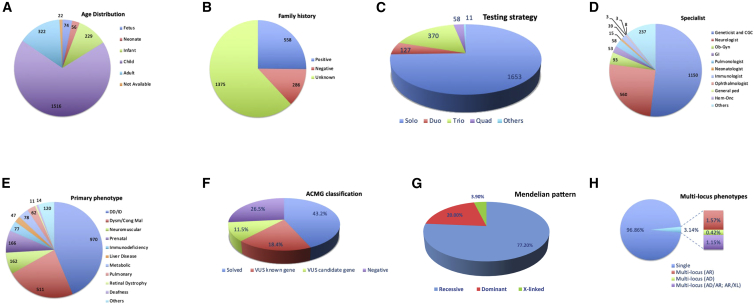
A total of 3,310 CES tests were performed on 2,219 families (1,653 solo, 127 duo, 370 trio, 58 quad, and 11 others [five or more]) (Table S1). The majority (1,653 families) had index-only CES, although 566 families had duo, trio, quad, or other combinations (Figure 1). The empowerment of clinicians from all specialties to request CES as a first-tier test for individuals with any suspected genetic disease was key to the resulting diversity of clinical indications and ordering specialties (Table S1). This is further highlighted by the observation that nearly half of the requested tests were ordered by non-geneticists (Figure 1). As shown in Table S1 and Figure 1, the phenotypes included dysmorphic, neurodevelopmental, neuromuscular, neurodegenerative, cardiac, ophthalmologic, renal, gastrointestinal, dermatologic, audiologic, hematological, surgical, prenatal, and even pharmacogenetic phenotypes. Although we did not include any CES of cancer tissues, several individuals were referred in order to rule out hereditary cancer syndromes (Table S1). The overall diagnostic yield of CES based on “solved” cases only was 43.3% (961 out of 2,219) (Figure 1).
Figure 1.
Summary of the Study Cohort Characteristics
(A) Age distribution.
(B) Family history.
(C) Testing strategy.
(D) Ordering specialty.
(E) Primary indication (prenatal in this figure includes individuals who presented for testing for carrier status).
(F) ACMG classification of variants.
(G) Mendelian patterns of pathogenic and likely pathogenic variants.
(H) Multi-locus phenotypes.
The Landscape of Disease-Related Variants in a Highly Consanguineous Population
By only considering the “solved” category, we show that autosomal-recessive variants account for the overwhelming majority (742 out of 961, 77.2%) of variants, and they were usually (730 out of 742, 98.4%) homozygous (Figure 1). Founder variants accounted for 405 (41.3%), and the remaining 575 variants (58.7%) can be considered private. Dominant variants were observed in 20% of the solved instances (192 out of 961), 25% (48 out of 192) of which were proven to be de novo, whereas parental samples were not available for the rest. On the other hand, only 3.9% (37 out of 961) of the instances were X-linked (Figure 1). Please note that the number of pathogenic or likely pathogenic variants (980) is larger than the number of solved instances (961) due to the presence of a few multi-locus phenotypes (see below). In total, we add to the literature 378 pathogenic or likely pathogenic variants not previously reported (Table S1). We also estimate the likelihood of carrier status for a predefined list of known Saudi mutations (KSMs, identified through our prior work on the local population), excluding variants related to the clinical indication, as follows: among the 1,653 simplex cases from this study, 498 individuals (30.1%) carried one KSM, 162 (9.8%) carried two KSMs, 39 (2.4%) carried three KSMs, 11 (0.7%) carried four KSMs, and one (0.06%) carried five KSMs (Figure S2). We also tested 1,006 parents and found (excluding the variants that are directly related to the condition in their tested children) that the percentage of parents who carry at least one KSM is 29.4% (19.8% carried one, 7.6% carried two, 1.7% carried three, and 0.3% carried four, which was the highest carrier status observed among parents). Furthermore, we asked whether the low percentage of multi-locus phenotypes in our study (see below) can be explained by the low probability of shared parental carrier status for more than one KSM. Indeed, by specifically interrogating couples, we found that of the 503 parental duos available to us, 49 (9.7%) shared one, and only 13 (2.6%) shared two KSMs, again after excluding the variants that are directly related to the condition in their tested child (Figure S2). No couples shared more than two KSMs. This is consistent with our observation that multi-locus phenotypes (3.1% of solved instances, or 30 out of 961) comprised dual recessive diagnoses in less than half of the cases (13 out of 30) (Figure 1 and Table S3).
High Throughput Discovery of Candidate Genes
Table S1 lists 236 genes that have no established OMIM phenotype and that we propose as candidate genes. Not surprisingly, the majority (n = 155) are recessive; many of them (n = 51) predict loss of function, then dominant (n = 74), and then X-linked (n = 7). The highest level of confidence is given to 13 genes, which we observed had mutated independently in two families with a similar phenotype (Table 1). For example, we have identified two families affected by non-syndromic intellectual disability, and each segregates a different homozygous truncating variant in ALKBH8 (MIM: 613306).18 A syndrome of skeletal dysplasia with cerebellar atrophy was similarly identified in two families, each segregating a different homozygous truncating variant in SMPD4 (MIM: 610457) (unpublished data). Abnormal cortical development with associated intellectual disability and epilepsy segregated with a different homozygous truncating variant in TP73 (MIM: 601990) in each of the two affected families. Three families with suspected neuronal ceroid lipofuscinosis-like illness (neurodegeneration with retinal involvement) were found to harbor two different candidate variants in USO1 (MIM: 603344). Other genes found to harbor two different recessive, likely deleterious variants include EIF2A (MIM: 609234), ICE1 (MIM: 617958), IQSEC3 (MIM: 612118), SMG8 (MIM: 613175), and UNC5A (MIM: 607869). Multiple congenital anomalies syndromes were also observed in individuals who harbor independent heterozygous, likely deleterious variants in MOV10 (MIM: 610742) and RASIP1 (MIM: 609623).
Table 1.
List of Candidate Genes with More Than One Mutational Event Identified in this Cohort
| ID | Gene | Variant(s) | Zygosity | Phenotype | Justification |
|---|---|---|---|---|---|
| 17-1192 | ALKBH8 | GenBank: NM_001301010.1:c.1660C>T:p.Arg554Ter | Hom | Nonspecific intellectual disability with positive family history | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (this gene is involved in tRNA modification, a process that is increasingly recognized as the core feature of a number of intellectual disability syndromes, most notoriously ADAT3-related, which is one of the most common forms of intellectual disability in Arabia). |
| 17-6071 | ALKBH8 | GenBank: NM_001301010.1:c.1794delC:p.Trp599GlyfsTer19 | Hom | Nonspecific intellectual disability with positive family history | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (this gene is involved in tRNA modification, a process that is increasingly recognized as the core feature of a number of intellectual disability syndromes, most notoriously ADAT3-related, which is one of the most common forms of intellectual disability in Arabia). |
| 17-1025 | EIF2A | GenBank: NM_032025.4:c.649_650insT:p.Ser217MetfsTer4 | Hom | Intellectual disability, ASD, and epilepsy | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene. This gene encodes a eukaryotic translation initiation factor that functions in the early steps of protein synthesis of a small number of specific mRNAs and is used for initiation at non-AUG codons (see PMID: 28982758). |
| 17-6950 | EIF2A | GenBank: NM_032025.4:c.1229A>C:p.Gln410Pro | Hom | Intellectual disability and intractable epilepsy | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene. This gene encodes a eukaryotic translation initiation factor that functions in the early steps of protein synthesis of a small number of specific mRNAs and is used for initiation at non-AUG codons (see PMID: 28982758). |
| 18-2383 | ICE1 | GenBank: NM_015325.2:c.3724G>T:p.Asp1242Tyr | Hom | GDD, spasticity, optic atrophy, and brain atrophy | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (ICE1 promotes a link between splicing and nonsense-mediated mRNA decay [NMD]. Mutations in other NMD-related genes result in neurodevelopmental disorders. See PMID: 29528287). |
| REQ18-0816 | ICE1 | GenBank: NM_015325.2:c.603A>C:p.Lys201Asn | Hom | GDD and nonspecific brain atrophy | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (ICE1 promotes a link between splicing and nonsense-mediated mRNA decay [NMD]. Mutations in other NMD-related genes result in neurodevelopmental disorders. See PMID: 29528287). |
| 17-5039 | IQSEC3 | GenBank: NM_015232.1:c.1144G>A:p.Gly382Ser | Hom | Speech delay, intellectual disability, ASD, and seizures (generalized tonic clonic) | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (IQSEC3 acts together with gephyrin to regulate inhibitory synapse development). |
| 17-5139 | IQSEC3 | GenBank: NM_001170738.1:c.3340C>A:p.Leu1114Met | Hom | Cognitive and speech delay | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (IQSEC3 acts together with gephyrin to regulate inhibitory synapse development). |
| 17-0049-C | MOV10 | GenBank: NM_001130079.2:c.2101_2102insA:p.Ser701TyrfsTer18 | Het | Gross motor delay, IUGR, oligohydramnios, hepatosplenomegaly, and cholestasis | (1) The nature of the variant (predicted deleterious) and (2) the nature of the gene (PLI: 0.99). Mov10 is a novel telomerase-associated protein (see PMID: 19665004). It was demonstrated that Mov10 is essential for gastrulation and for normal head, eye, and brain development (see PMID: 28662698). Consistent with the mouse model, knockdown of maternal Mov10 in Xenopus led to defects in gastrulation and the development of notochord and paraxial mesoderm and a failure to neurulate (see PMID: 29266590). |
| 18-1423 | MOV10 | GenBank: NM_001130079.2:c.2268_2269insTCGTGGA:p.Val757SerfsTer32 | Het | Large patent ductus arteriosus, scoliosis, hemivertebrae, phocomelia, and omphalocele versus gastroschisis | (1) The nature of the variant (predicted deleterious) and (2) the nature of the gene (PLI: 0.99). Mov10 is a novel telomerase-associated protein (see PMID: 19665004). It was demonstrated that Mov10 is essential for gastrulation and for normal head, eye, and brain development (see PMID: 28662698). Consistent with the mouse model, knockdown of maternal Mov10 in Xenopus led to defects in gastrulation and the development of notochord and paraxial mesoderm and a failure to neurulate (see PMID: 29266590). |
| 17-2799 | PLXNA3 | GenBank: NM_017514.5:c.1997G>A:p.Arg666His | Hemi | Epilepsy (generalized absence seizures), mild memory deficits, and history of IUGR during pregnancy | (1) The nature of the variant (predicted deleterious) and (2) the nature of the gene (PLI: 0.98. It was demonstrated that Plexin A3 is involved in semaphorin 3F-mediated oligodendrocyte precursor cell migration (see PMID: 23063687), contributes to Sema3F and Sema3A signaling, regulates the development of hippocampal axonal projections in vivo (see PMID: 11683995), mediates neuronal apoptosis during dorsal root ganglia development (see PMID: 19020035), and regulates motor axonal branch morphogenesis (see PMID: 23349787). |
| 18-1411 | PLXNA3 | GenBank: NM_017514.5:c.2623C>T:p.Arg875Trp | Hemi | Intellectual disability, autistic features, and has a twin (18-1631) with same condition | (1) The nature of the variant (predicted deleterious) and (2) the nature of the gene (PLI: 0.98). It was demonstrated that Plexin A3 is involved in semaphorin 3F-mediated oligodendrocyte precursor cell migration (see PMID: 23063687), contributes to Sema3F and Sema3A signaling, regulates the development of hippocampal axonal projections in vivo (see PMID: 11683995), mediates neuronal apoptosis during dorsal root ganglia development (see PMID: 19020035), and regulates motor axonal branch morphogenesis (see PMID: 23349787). |
| 17-6562 | PTPN12 | GenBank: NM_001131009.1:c.722C>A:p.Pro241His | Het | GDD, failure to thrive, microcephaly, and epilepsy | (1) The nature of the variant (predicted deleterious in silico) and (2) the nature of the gene (pLI 0.99). Mutations in other family members, e.g., PTPN23 and PTPN11, are identified in individuals with intellectual disability and other neurodevelopmental disorders (see PMID: 25558065, PMID: 29090338, and PMID: 28957739). |
| 17-8496 | PTPN12 | GenBank: NM_002835.3:c.2282-34A>T | Hom | GDD and nonspecific MRI findings | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (pLI 0.99). Mutations in other family members, e.g., PTPN23 and PTPN11, are identified in individuals with intellectual disability and other neurodevelopmental disorders (see PMID: 25558065, PMID: 29090338, and PMID: 28957739). |
| 18-0081 | RASIP1 | GenBank: NM_017805.2:c.2325G>C:p.Leu775Phe | Het | Microcephaly and fibular hemimelia | (1) The nature of the variant (predicted deleterious) and (2) the nature of the gene (PLI: 0.97). RASIP1 is a dynamic EPAC1-RAP1 signaling effector that controls actin bundling and restricts junction remodeling in vitro and in vivo, and it is required for stabilizing nascent patent blood vessels in both mice and zebrafish (see PMID: 23886837). It was reported that the distribution of the intersegmental arteries plays a decisive role in the formation of the definitive vertebral body anlage during the stage of resegmentation and early chondrification, and any abnormal distribution of the arteries might induce malformations (see PMID: 6797230). |
| REQ18-1889 | RASIP1 | GenBank: NM_017805.2:c.1193_1196delATGT:p.Tyr398Ter | Het | Butterfly vertebrae, bilateral fused ribs, and diaphragmatic anomaly | (1) The nature of the variant (predicted deleterious) and (2) the nature of the gene (PLI: 0.97). RASIP1 is a dynamic EPAC1-RAP1 signaling effector of that controls actin bundling and restricts junction remodeling in vitro and in vivo, and it is required for stabilizing nascent patent blood vessels in both mice and zebrafish (see PMID: 23886837). It was reported that the distribution of the intersegmental arteries plays a decisive role in the formation of the definitive vertebral body anlage during the stage of resegmentation and early chondrification and any abnormal distribution of the arteries might induce malformations (see PMID: 6797230). |
| 17-5563 | SMG8 | GenBank: NM_018149.6:c.623A>G:p.His208Arg | Hom | Microcephaly, intellectual disability, cataract, and neck hyperpigmentation | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (this is a nonsense-mediated mRNA decay factor). Mutations in the NMD complex have been shown to result in multisystem involvement (PMID: 27018474). Recessive inheritance was confirmed by parental (17-5564 and 17-5565) whole-exome sequencing. An affected sibling (17-5566) is also homozygous for this variant. |
| REQ18-0840 | SMG8 | GenBank: NM_018149.6:c.437_438insA:p.Ser146ArgfsTer13 | Hom | Growth retardation and/or short stature, microcephaly, fine motor delay, ventricular septal defect, failure to thrive, and facial dysmorphism | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (this is a nonsense-mediated mRNA decay factor). Mutations in the NMD complex have been shown to result in multisystem involvement (PMID: 27018474). |
| 17-3980 | SMPD4 | GenBank: NM_001171083.2:c.1471-28_1477del | Hom | Bilateral clenched hands and talipes, IUGR, partial absence of corpus callosum, and family history of three neonatal deaths with similar features | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) SMPD4, a bona fide sphingomyelinase in the ceramide pathway, catalyzes the hydrolysis of membrane sphingomyelin to form phosphorylcholine and ceramide (PMID: 16517606). Interestingly, inactivation or knockout of another sphingomyelinase of the ceramide pathway, SMPD3, led to growth inhibition, apoptosis, retarded development, and ultimately to a lethal, autosomal-recessive form of osteogenesis imperfecta, skeletal malformations, dwarfism, and severe chondrodysplasia as a result of the deficient sphingomyelinase activity (PMID: 16025116 and PMID: 27882938). |
| 17-6035 | SMPD4 | GenBank: NM_017751.4:c.390_406del:p.Pro131LeufsTer2 | Hom | Failure to thrive, fine and gross motor delay, brain atrophy, syndactyly, hypoplasia of phalanges, and short metacarpal bones | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) SMPD4, a bona fide sphingomyelinase in the ceramide pathway, catalyzes the hydrolysis of membrane sphingomyelin to form phosphorylcholine and ceramide (PMID: 16517606). Interestingly, inactivation or knockout of another sphingomyelinase of the ceramide pathway, SMPD3, led to growth inhibition, apoptosis, retarded development, and ultimately to a lethal, autosomal-recessive form of osteogenesis imperfecta, skeletal malformations, dwarfism, and severe chondrodysplasia as a result of the deficient sphingomyelinase activity (PMID: 16025116 and PMID: 27882938). |
| 17-4956 | TP73 | GenBank: NM_001126242.2:c.1163delT:p.Leu388HisfsTer151 | Hom | Lissencephaly and hypotonia | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the loss of p73 leads to the disappearance of the Cajal-Retzius (CR) neurons in the cortical marginal zone and hippocampal molecular layer, and this could underlie the hippocampal dysgenesis, the hydrocephalus, and the abnormalities in the pheromone sensory pathways in p73 mice (PMID: 10716451). |
| 17-6638 | TP73 | GenBank: NM_001126240.2:c.847C>T:p.Gln283Ter | Hom | Fine and gross motor delay, speech delay, cleft lip and/or palate, cortical dysplasia, pachygyria, and strabismus | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the loss of p73 leads to the disappearance of the Cajal-Retzius (CR) neurons in the cortical marginal zone and hippocampal molecular layer, and this could underlie the hippocampal dysgenesis, the hydrocephalus, and the abnormalities in the pheromone sensory pathways in p73 mice (PMID: 10716451). |
| 17-6690 | UNC5A | GenBank: NM_133369.2:c.1309C>T:p.Arg437Ter | Hom | Failure to thrive, microcephaly, GDD, hearing loss, periventricular leukomalacia, and seizures | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene (UNC-5, a transmembrane protein with immunoglobulin and thrombospondin type 1 domains) guides cell and pioneer axon migrations in C. elegans (see PMID: 1384987). |
| 17-7961 | UNC5A | GenBank: NM_133369.2:c.1550T>A:p.Ile517Asn | Hom | Congenital heart disease, failure to thrive, microcephaly, GDD, nystagmus, cafe au lait spots, and seizures. | (1) The nature of the variant (predicted deleterious, within autozygome) and (2) the nature of the gene: UNC5A belongs to a family of netrin-1 receptors thought to mediate the chemo repulsive effect of netrin-1 on specific axons (see PMID: 1384987). A loss-of-function variant of UNC5A is identified in an individual with failure to thrive, microcephaly, GDD, hearing loss, periventricular leukomalacia, and seizures (see 17-6690). |
| 17-2828-B | USO1 | GenBank: NM_003715.3:c.46G46G>C:p.Val16LeuVal161Leu | Hom | Developmental regression, retinal degeneration, and suspected NCL | (1) The nature of the variant (present within autozygome, predicted deleterious in silico) and (2) the nature of the gene: USO1, a vesicle transport factor, is a peripheral membrane protein that recycles between the cytosol and the Golgi apparatus. Mice lacking USO1 exhibit disruption of Golgi structure and early embryonic lethality (see PMID: 23185636 and MGI: 1929095). |
| 17-6599 | USO1 | GenBank: NM_003715.3:c.46G46G>C:p.Val16LeuVal161Leu | Hom | Developmental regression, retinal degeneration, and suspected NCL | (1) The nature of the variant (present within autozygome, predicted deleterious in silico) and (2) the nature of the gene: USO1, a vesicle transport factor, is a peripheral membrane protein that recycles between the cytosol and the Golgi apparatus. Mice lacking USO1 exhibit disruption of Golgi structure and early embryonic lethality (see PMID: 23185636 and MGI: 1929095). |
| 17-8522 | USO1 | GenBank: NM_003715.3:c.2514T>G:p.Ile739Arg | Hom | GDD, microcephaly, epilepsy, optic atrophy, and suspected NCL | (1) The nature of the variant (present within autozygome, predicted deleterious in silico) and (2) the nature of the gene: USO1, a vesicle transport factor, is a peripheral membrane protein that recycles between the cytosol and the Golgi apparatus. Mice lacking USO1 exhibit disruption of Golgi structure and early embryonic lethality (see PMID: 23185636 and MGI: 1929095). |
Abbreviations are as follows: Hom = homozygous, Het = heterozygous, Hemi = hemizygous, IUGR = intrauterine growth restriction, MRI = magnetic resonance imaging, NCL = neuronal ceroid lipofuscinosis, and GDD = global developmental delay
The next level of confidence is for genes where the animal model recapitulates the observed human phenotype. For example, the severe, unexplained diarrhea in the child with a truncating variant in MYO1A (MIM: 601478) seems to be recapitulated by Myo1a knockout mice that have severe defects in the brush border of their intestine.19 The child who has brain malformation and a severe neurological phenotype and harbors a homozygous truncating variant in SCYL2 (MIM: 616365) is another example wherein the candidacy of the gene is strongly supported by the corresponding knockout mouse.20 Katnal1 mutant mice display behavioral, cognitive, and structural brain anomalies, which are strikingly similar to those in the individual we report who has a truncating variant in KATNAL1 (MIM: 614764).21 Similarly, the neurological phenotype in the child with a homozygous variant in NRDC (MIM: 602651) is sufficiently recapitulated by the Nrdc−/− mouse to support the candidacy of this gene.22
The third level of evidence was available biological data on the gene of interest. For example, all oral-facial-digital (OFD) syndrome (MIM: 311200)- and Meckel-Gruber syndrome (MIM: 249000)-related genes are known to play a role in ciliary biology.23 Thus, the homozygous truncations in CCDC96 and CEP29 (MIM: 617728) were attractive candidates, respectively, given their role in ciliogenesis.24, 25, 26
In addition, our data provide high throughput “confirmation” of 64 genes (40 recessive, 21 dominant, and three X-linked) that have previously been proposed as candidates but are either not yet listed with OMIM phenotypes or have a listing that is based on a single study. These are listed along with the observed and previously reported phenotypes in Table S2. Interestingly, several of these genes were proposed as candidates from the same population, i.e., Saudis. These include TBC1D32 (MIM: 615867)-related OFD IX (MIM: 258865),27 PGAP3 (MIM: 611801)-related Toriello Carey syndrome (MIM: 615716),28 LACC1 (MIM:613409)-related arthropathy and Crohn disease (MIM: 266600),29 PITX3 (MIM: 602669)-related autosomal-recessive anterior segment dysgenesis and microphthalmia,30 PPP1R21 (MIM: 618159)-related intellectual disability,31 SMG9 (MIM: 613176)-related heart and brain malformation syndrome,32 ARMC9 (MIM: 617622)-related Joubert syndrome (MIM: 617622),33 DNAJC17 (MIM: 616844)-related syndrome of retinitis pigmentosa and immunodeficiency,34 and CLHC1-related myopathy.8 Although we have previously reported a homozygous truncating variant in PRKD1 (MIM: 605435) in a family with non-syndromic truncus arteriosus,35 OMIM currently only lists this gene in the context of a syndrome of ectodermal dysplasia with congenital heart disease on the basis of de novo, heterozygous missense changes. Here, we show that individual REQ18-1959, who has a different homozygous truncating variant and an associated, non-syndromic, complex congenital heart disease, not only provides confirmation that PRKD1 is indeed associated with recessive, non-syndromic truncus arteriosus but also expands the congenital-heart-disease phenotype associated with this gene. Although the majority of the “confirmatory” cases have phenotypes that are similar to those originally reported, some deviate significantly and are worth highlighting. These include TBC1D32 (MIM: 615867)-related holoprosencephaly, NDUFB10 (MIM: 603843)-related non-immune hydrops fetalis, and a lack of liver involvement in two brothers with LARS (MIM: 151350)-related epileptic encephalopathy, microcephaly, and global developmental delay (Table S2).
Identification of Unusual Mutational Mechanisms
The discovery of recessive forms of genes previously associated only with dominant disorders has the potential to unravel interesting mutational mechanisms for the associated phenotypes.36 Table 2 lists several recessive forms of genes that had only been described in the context of dominant phenotypes. The recessive phenotypes ranged from identical, e.g., for SLC4A10 (MIM: 605556)-related intellectual disability and epilepsy; to earlier onset, e.g., for POU4F3 (MIM: 602460)-related prelingual hearing loss; to more severe, e.g., for PITX3 (MIM: 602669)-related eye malformation and global developmental delay; to completely distinct, e.g., for MYH11 (MIM: 160745)-related megacystis, microcolon, and intestinal hypoperistalsis. One remarkably distinct recessive phenotype that is worth highlighting is ATP6V1B2 (MIM: 606939)-related cutis laxa syndrome that is identical to the one we described previously in siblings with a homozygous ATP6V1E1 (MIM: 108746) variant, even though dominant ATP6V1B2 (MIM: 606939) variants are known to cause a distinct phenotype known as Zimmermann-Laband syndrome 1 (MIM: 135500).37 Although parental gonadal mosaicism is known to occur in consanguineous families,38, 39, 40 the family of first-cousin parents and three children who have classical Rubinstein-Taybi syndrome (MIM: 180849) and share the same truncating variant in CREBBP (MIM: 600140) is a reminder that CES in apparently recessive pedigrees should be carefully reviewed for that possibility. More unusual is the observation of parental gonadal mosaicism in GNAO1 (MIM: 139311) in two families in our cohort who have GNAO1-related epileptic encephalopathy. This increases to three the number of families with gonadal mosaicism for this recently described gene, and only one family had no documented gonadal mosaicism.41 De novo point mutations in the context of autosomal-recessive disorders have very rarely been reported.39 In our cohort, we show that one individual with Joubert syndrome (MIM: 614615) harbored a compound-heterozygous mutation in C5orf42 (MIM: 614615) with one inherited variant, whereas the other has arisen de novo, although we cannot rule out paternal gonadal mosaicism. In addition, we show that one infant with cholestatic liver failure harbored a homozygous truncating variant in BAAT (MIM: 602938), a gene that has been implicated in the context of cholestatic liver disease as a potential modifier on the basis of a single missense variant.
Table 2.
List of Genes Previously Reported Only in the Context of Dominant Disorders but Were Found to Harbor Likely Deleterious Recessive Forms of Genes in this Cohort
| ID | Gene | Variant(s) | Zygosity | Dominant phenotype | Observed recessive phenotype | Phenotype comparison |
|---|---|---|---|---|---|---|
| 17-6650 | ATP6V1B2 | GenBank: NM_001693.3:c.1001T>C:p.Ile334Thr | Hom | Zimmermann-Laband syndrome 2 | Distinct cutis laxa syndrome identical to that described for ATP6V1E1 (PMID: 27023906) | Distinct |
| 16-2726 | ATP6V1B2 | GenBank: NM_001693.3:c.1001T>C:p.Ile334Thr | Hom | Zimmermann-Laband syndrome 2 | Distinct cutis laxa syndrome identical to that described for ATP6V1E1 (PMID: 27023906) | Distinct |
| REQ18-1700 | BAAT | GenBank: NM_001127610.1:c.2T>A:p.? | Hom | Cholestatic liver disease | Cholestatic liver disease | Similar |
| 17-5309 | MYH11 | GenBank: NM_002474.2:c.3424C>T:p.Arg1142Ter | Hom | Aortic aneurysm and familial thoracic 4 | Megacystis, bilateral hydronephrosis, absent stomach, borderline polyhydramnios, microcolon, and intestinal hypoperistalsis | Distinct |
| 17-0807 | MYH11 | GenBank: NM_022844.2:c.1033+1G>A | Hom | Aortic aneurysm and familial thoracic 4 | Megacystis | Distinct |
| 18-1351 | NOTCH3 | GenBank: NM_000435.3:c.1790G>C:p.Cys597Ser | Hom | CADASIL (cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy) | Neuropsychiatric disturbances, migraine headache, periventricular leukomalacia and temporal lobe pole hyperintensity. Suspected CADASIL | Similar |
| 17-5492 | PITX3 | GenBank: NM_005029.3:c.640_656del:p.Ala214ArgfsTer42 | Hom | Anterior segment dysgenesis | GDD, microphthalmia, blindness, cataracts, and spastic quadriplegia | More severe |
| 17-3180 | POU4F3 | GenBank: NM_002700.2:c.488delA:p.His163ProfsTer41 | Hom | Hearing loss (onset between the second and sixth decades) | Bilateral, prelingual, profound sensorineural hearing loss | More severe |
| 16-3003 | PPM1D | GenBank: NM_003620.3:c.939_940delTG:p.Ser313ArgfsTer28 | Hom | Intellectual developmental disorder with gastrointestinal difficulties and high pain threshold | GDD and epilepsy | Similar |
| 17-4393 | SLC4A10 | GenBank: NM_022058.3:c.2773-2A>C | Hom | Complex partial epilepsy and mental retardation | Intellectual disability and epilepsy | Similar |
Abbreviations are as follows: Hom = homozygous and GDD = global developmental delay.
Patterns of Mutations in Prenatal Cases
Despite a comparable diagnostic yield in pre- (30 out of 65, 46.2%) and post-natal cases, we note that an even higher percentage of prenatal instances are autosomal recessive (87%), and the majority of these (65.4%) represent homozygous probable loss of function (Table 3). The severe, and often lethal, prenatal presentation of these loss-of-function events often represents a phenotypic expansion of the corresponding genes (Table 3). One example is the homozygous truncating variant of MYO9A (MIM: 604875) in a fetus with severe hydrocephalus that appears to mirror the phenotype observed in Myo9 knockout mice.42 This is in contrast to the neuromuscular phenotype observed in the few reported instances of bi-allelic missense variants in this gene. Half of the individuals referred for prenatal genetic counseling (80 out of 162, 49.4%) came from couples with a history of one or more deceased children (or pregnancy losses) for whom no DNA was available for testing. Table S8 shows that this approach of “molecular autopsy by proxy”43 provided a compelling etiology in the majority of instances (45 out of 80, 56.3%). As expected by the nature of this approach, all the variants were recessive. Again, loss-of-function events were common in this sub-cohort that had lethal phenotypes (28 out 45, 62.2%).
Table 3.
aList of Prenatal Cases with Available DNA from the Fetus
| ID | Result | Gene | Variant(s) | Zygosity | LOF | Observed Fetal Phenotype | Phenotype Category |
|---|---|---|---|---|---|---|---|
| 17-0422 | Solved | ALG9 | GenBank: NM_001077690.1:c.430T>C:p.Cys144Arg | Hom | – | Cardiomegaly, pericardial effusion, a thick nuchal fold, oligohydramnios, hydronephrosis, and a small bladder | Typical |
| 17-6079 | Solved | ATP11C | GenBank: NM_173694.4:c.1675+2T>C | Hemi | y | Fetus with hydrops fetalis. | Distinct prenatal presentation |
| 17-3514 | Solved | BBS1 | GenBank: NM_024649.4:c.830+2T>C | Hom | y | Severe oligohydramnios and enlarged echogenic kidney with cyst bilaterally, most likely ARPKD | Typical |
| 17-1195 | Ambiguous (candidate gene) | BRAP | GenBank: NM_006768.4:c.199C>T:p.Arg67Ter | Hom | y | Non-immune hydrops fetalis | NA (first case) |
| 16-3155 | Solved | CANT1 | GenBank: NM_138793.3:c.902_906dup:(p.Ser303AlafsTer21) | Hom | y | Skeletal dysplasia | Typical |
| REQ18-1150 | Solved | CC2D2A | GenBank: NM_001080522.2:c.4531T>C:p.Trp1511Arg | Hom | – | Enlarged kidneys and occipital encephalocele | Typical |
| 18-2311 | Solved | CEP55 | GenBank: NM_001127182.1:c.807T>G:p.Tyr269Ter | Hom | y | Renal dysplasia, anhydraminos, and hydranencephaly | Typical |
| 17-2713 | Ambiguous (VUS) | CHRNA1 | GenBank: NM_000079.3:c.254T>C:p.Leu85Pro | Hom | – | Non-immune hydrops fetalis | Typical |
| 17-0633 | Ambiguous (VUS) | CHRNG | GenBank: NM_005199.5:c.1180C>G:p.Pro394Ala | Hom | – | Club foot, oligohydramnios, cystic hygroma and/or increased nuchal thickness, skin edema, and fixed flexion in an upper limb | Typical |
| 17-5074 | Solved | CNTNAP1 | GenBank: NM_003632.2:c.1903C>T:p.Arg635Ter | Hom | y | Polyhydramnios, absent stomach, and a mother with history of IUFDs | Typical |
| 17-3078 | Solved | COL1A1 | GenBank: NM_000088.3:c.2399G>A:p.Gly800Glu | De novo | – | Skeletal dysplasia | Typical |
| REQ18-4214 | Solved | ECHS1 | GenBank: NM_004092.3:c.88+5G>A: | Het (carrier) | y | Normal (tested with the parents, who had a history of neonatal deaths with severe lactic acidosis but no available DNA. See “Molecular Autopsy by Proxy” for details.) | NA |
| 17-3698 | Solved | ERCC5 | GenBank: NM_001204425.1:c.3789delT:p.Asp1263GlufsTer24 | Hom | y | Fetus with a dilated third ventricle, dilated cisternae magna, hypertelorism, micrognathia, generalized skin edema, moderately enlarged heart, hydrothorax, bilateral talipes, bilateral clenched hands, and fixed position (hip flexed and upper limbs flexed) | Distinct prenatal presentation |
| 17-5348 | Solved | ETFA | GenBank: NM_000126.3:c.883-1_885delGACA | Hom | y | Pericardial effusion, echogenic kidney, hydrocephalus, and a small chest | Distinct prenatal presentation |
| 18-1181 | Solved | FLVCR2 | GenBank: NM_001195283.1:c.596T>A:p.Val199Glu | Hom | – | Hydrocephalus, dilated cisterna magna, hypoplastic cerebellum, and brain atrophy | Typical |
| 17-3511 | Ambiguous (candidate gene) | FRMD6 | GenBank: NM_001267046.1:c.262G>A:p.Ala88Thr | De novo | – | IUGR and microcephaly. | N/A (first case) |
| 17-0224 | Solved | FTO | GenBank: NM_001080432.3:c.871C>T:p.Gln291Ter | Hom | y | Dandy-Walker malformation, IUGR, and polyhydramnios | Typical |
| 18-2474 | Solved | GUSB | GenBank: NM_000181.4:c.307C>T:p.Arg103Trp | Hom | – | Fetus with non-immune hydrops | Typical |
| 18-2404 | Solved | HSPG2 | GenBank: NM_001291860.1:c.12462delT:p.Leu4155CysfsTer30 | Hom | y | Skeletal dysplasia | Typical |
| 17-0425 | Ambiguous (candidate gene) | IL6ST | GenBank: NM_001190981.1:c.841C>T:p.Arg281Ter | Hom | y | Brain malformation, short long bones, kidney malformation, and unilateral MCDK | NA (first case) |
| 17-0409 | Solved | KIF19 | GenBank: NM_153209.3:c.788G>A:p.Arg263His; NM_153209.3:c.1906T>G:p.Tyr636Asp | Het (CMP) | – | abnormal “strawberry” head shape, absent corpus callosum, and brain ventriculomegaly | NA (first case; see PMID: 28749478) |
| 17-0807 | Solved | MYH11 | GenBank: NM_022844.2:c.1033+1G>A | Hom | y | Megacystis | Distinct prenatal presentation |
| 17-5309 | Solved | MYH11 | GenBank: NM_002474.2:c.3424C>T:p.Arg1142Ter | Hom | y | Megacystis, bilateral hydronephrosis, absent stomach, borderline polyhydramnios, microcolon, and intestinal hypoperistalsis | Distinct prenatal presentation |
| 17-2355 | Solved | MYO9A | GenBank: NM_006901.4:c.1537C>T:p.Arg513Ter | Hom | y | Hydrocephalus | Distinct prenatal presentation |
| 17-4600 | Solved | MYSM1 | GenBank: NM_001085487.2:c.1168G>T:p.Glu390Ter | Hom | y | Non-immune hydrops fetalis | Distinct prenatal presentation |
| 17-2505 | Solved | NDUFB10 | GenBank: NM_004548.2:c.373_375delCAG:p.Gln125del | Hom | – | Non-immune hydrops fetalis and died after birth | Distinct prenatal presentation |
| 17-4671 | Solved | PEX5 | GenBank: NM_000319.4:c.1554T>G:p.Asn518Lys | Hom | – | Multicystic kidney, dilated cisterna magna, ventriculomegaly, and a positive family history | Distinct prenatal presentation |
| 17-6282 | Solved | PHGDH | GenBank: NM_006623.3:c.1030C>T:p.Arg344Ter | Hom | y | Non-immune hydrops fetalis | Phenotypic expansion |
| 17-5922 | Solved | PIEZO1 | GenBank: NM_001142864.4:c.5013_5016delGGCG:p.Ala1672CysfsTer59 | Hom | y | Non-immune hydrops | Typical |
| 17-2819 | Solved | PIEZO1 | GenBank: NM_001142864.4:c.6372G>C:p.Trp2124Cys | Hom | – | Hydrops fetalis | Typical |
| 17-4408 | Ambiguous (VUS) | POLG | GenBank: NM_001126131.1:c.2606G>A:p.Arg869Gln | Hom | – | Polyhydramnios, contractures, scoliosis, and kyphosis | Distinct prenatal presentation |
| 18-1761 | Solved | POMT1 | GenBank: NM_001077366.1:c.118+1G>T | Hom | y | Fetus with severe ventriculomegaly | Typical |
| 17-5897-B | Solved | PTPN11 | GenBank: NM_002834.4:c.781C>T:p.Leu261Phe | De novo | – | Non-immune hydrops fetalis | Phenotypic expansion |
| 17-1571 | Ambiguous (VUS) | RTEL1 | GenBank: NM_001283009.1:c.3730T>C:p.Cys1244Arg | Hom | – | Fetus with severe IUGR and oligohydramnios | Phenotypic expansion |
| REQ18-3059 | Ambiguous (VUS) | SMAD6 | GenBank: NM_005585.4:c.818A>G:p.Glu273Gly | Hom | – | Severe non-immune hydrops fetalis with a family history of a previously affected pregnancy | Distinct prenatal presentation |
| 17-3980 | Ambiguous (candidate gene) | SMPD4 | GenBank: NM_001171083.2:c.1471-28_1477del | Hom | y | Bilateral clenched hands and talipes, IUGR, partial absence of the corpus callosum, and a family history of three neonatal deaths with similar features | NA (first case) |
| 17-4710 | Solved | STRA6 | GenBank: NM_001142617.1:c.1594C>T:p.Arg532Ter | Hom | y | Hypoplastic left heart syndrome | Distinct prenatal presentation |
| 17-3069-A | Ambiguous (candidate gene) | STXBP3 | GenBank: NM_007269.4:c.622G>A:p.Ala208Thr | Hom | – | Arthrogryposis multiplex congenita | NA (first case) |
| 17-0889 | Solved | TBC1D32 | GenBank: NM_152730.6:c.1372+1G>T | Hom | y | Previous termination of 15/32 with holoprosencephaly, cyclops, cleft lip, ventricular septal defect, agenesis of the corpus callosum, club foot, and a sandal gap (no sample) | Phenotypic expansion |
| 17-0286 | Solved | TMEM130 | GenBank: NM_001134450.1:c.23G>A:p.Arg8His | Hom | – | Pericardial effusion, enlarged kidney, occipital encephalocele, and suspected to have Meckel-Gruber syndrome | Typical |
| 17-4015 | Ambiguous (VUS) | WDR34 | GenBank: NM_052844.3:c.1061C>T:p.Thr354Met | Hom | – | Fetus with IUGR and skeletal dysplasia | Typical |
Abbreviations are as follows: LOF = loss of function, y = yes, Hom = homozygous, Het = heterozygous, Hemi = hemizygous, IUGR = intrauterine growth restriction, ARKPD = autosomal-recessive polycystic kidney disease, VUS = variant of unknown significance, IUFD = intrauterine fetal death, NA = not applicable, MCDK = multicystic dysplastic kidney, and CMP = compound.
Negative instances can be found in Table S7.
Allelic Series and Phenotypic Expansion
Table 4 lists instances where the observed phenotypes differ noticeably from those reported in the literature (multi-locus phenotypes have been excluded). A few instances are worth highlighting. We have previously reported an IGFBP7 (MIM: 602867)-related syndrome characterized by retinal artery microaneurysm, supravalvular pulmonic stenosis, and potential liver involvement.44 Individual 17-0163 presented with ductal plate malformation with resulting liver cirrhosis, ischemic stroke, and aortic coarctation, features that add to the phenotypic delineation of the syndrome. Phenotypic expansion was not limited to recessive disorders, however, as exemplified by individual 17-4116. This individual harbors a truncating variant in STAT3 (MIM: 102582) (no truncating variants have been reported in this gene to our knowledge), and this might explain the unexpectedly severe brain vascular phenotype in the form of Moyamoya disease. Similarly, the individual with a de novo truncating variant in TCF12 (MIM: 600480) completely lacks craniosynostosis, although he does have the developmental delay component of TCF12-related craniosynostosis 3.
Table 4.
List of Individuals Whose Phenotypes Represent an Expansion of the Published Phenotypes
| ID | Result | Gene | Variant(s) | Zygosity | Phenotypic Expansion Elements |
|---|---|---|---|---|---|
| REQ18-2038 | Pathogenic or likely pathogenic | ACO2 | GenBank: NM_001098.2:c.1187C>T:p.Ser396Leu | Hom | Severe neonatal hypotonia and SMA-like presentation |
| 17-6786 | Pathogenic or likely pathogenic | ADAT3 | GenBank: NM_138422.4:c.430G>A:p.Val144Met | Hom | Tetralogy of Fallot |
| 17-9070 | Pathogenic or likely pathogenic | ATAD3A | GenBank: NM_018188.4:c.1410+1G>A | Hom | Severe fetal presentation |
| 17-6079 | Pathogenic or likely pathogenic | ATP11C | GenBank: NM_173694.4:c.1675+2T>C | Hemi | Non-immune hydrops fetalis |
| 17-8295 | Pathogenic or likely pathogenic | BEAN1 | GenBank: NM_001136106.4:c.180C>G:p.Tyr60Ter | Het | Intellectual disability and epilepsy |
| 17-2136 | Pathogenic or likely pathogenic | CACNA1G | GenBank: NM_198384.2:c.4759+1G>A | Het | Microcephaly |
| 17-2687 | Pathogenic or likely pathogenic | CCNO | GenBank: NM_021147.4:c.964delC:p.Leu322CysfsTer5 | Hom | Infertility |
| 17-2688 | Pathogenic or likely pathogenic | CCNO | GenBank: NM_021147.4:c.964delC:p.Leu322CysfsTer5 | Het | Infertility |
| 17-2584 | Pathogenic or likely pathogenic | CHRNG | GenBank: NM_005199.5:c.1366_1367delAG:p.His457LeufsTer2 | Hom | Significant brain involvement |
| 17-1482 | Pathogenic or likely pathogenic | COL4A1 | GenBank: NM_001303110.1:c.761delT:p.Phe254SerfsTer60 | Het | Moyamoya disease |
| 17-3698 | Pathogenic or likely pathogenic | ERCC5 | GenBank: NM_001204425.1:c.3789delT:p.Asp1263GlufsTer24 | Hom | Severe fetal presentation |
| 17-0163 | Pathogenic or likely pathogenic | IGFBP7 | GenBank: NM_001553.2:c.830-1G>A | Hom | Ischemic stroke, coarctation of aorta, and ductal plate malformation with resulting liver cirrhosis |
| 17-4391 | Pathogenic or likely pathogenic | KDM1A | GenBank: NM_015013.3:c.782_783insA:p.Ile262TyrfsTer7 | Het | Lack of a cleft |
| 17-2505 | Pathogenic or likely pathogenic | NDUFB10 | GenBank: NM_004548.2:c.373_375delCAG:p.Gln125del | Hom | Fetal presentation with non-immune hydrops fetalis |
| 18-0283 | Pathogenic or likely pathogenic | PARS2 | GenBank: NM_152268.3:c.283G>A:p.Val95Ile | Hom | Lack of lactic acidemia and liver involvement (see also 17-8126 and 17-3573 in Table S1. These individuals have the same variant and also lack lactic acidemia and liver involvement.) |
| 17-7203 | Pathogenic or likely pathogenic | PEX16 | GenBank: NM_004813.2:c.954delG:p.Arg318GlyfsTer39 | Hom | Lack of the typical neonatal presentation and presence of hypertonia instead |
| REQ18-1959 | Pathogenic or likely pathogenic | PRKD1 | GenBank: NM_002742.2:c.2554G>T:p.Glu852Ter | Hom | Much more severe and complex congenital heart disease (pulmonary atresia, ventricular septal defect, major aortopulmonary collateral artery, patent ductus arteriosus, and hypertrophic cardiomyopathy; on prostaglandin with biventricular hypertrophic cardiomyopathy) |
| 18-0876 | Pathogenic or likely pathogenic | RARS2 | GenBank: NM_020320.4:c.1123G>A:p.Val375Met | Hom | Lack of pontocerebellar hypoplasia |
| 17-5069 | Pathogenic or likely pathogenic | SCN5A | GenBank: NM_001099405.1:c.5776C>T:p.Arg1926Ter | Het | Significant skeletal-muscle involvement and no evidence of cardiac involvement |
| 17-4396 | Pathogenic or likely pathogenic | SMAD3 | GenBank: NM_001145103.1:c.72delG:p.Arg25GlyfsTer47 | Het | Severe intracranial vessel disease |
| 17-1888 | Pathogenic or likely pathogenic | SMARCAL1 | GenBank: NM_001127207.1:c.1824delT:p.Phe608LeufsTer26 | Hom | Significant brain involvement |
| 17-1889 | Pathogenic or likely pathogenic | SMARCAL1 | GenBank: NM_001127207.1:c.1824delT:p.Phe608LeufsTer26 | Hom | Significant brain involvement |
| 17-4116 | Pathogenic or likely pathogenic | STAT3 | GenBank: NM_003150.3:c.1454delA:p.Asn485ThrfsTer6 | Het | Moyamoya disease |
| 17-4750 | Pathogenic or likely pathogenic | SURF1 | GenBank: NM_003172.3:c.588+1G>A | Hom | SMA-like presentation |
| 17-4613 | Pathogenic or likely pathogenic | TCF12 | GenBank: NM_001306220.2:c.493G>T:p.Glu165Ter | De novo | Lack of craniosynostosis |
| 17-6229 | Pathogenic or likely pathogenic | VIPAS39 | GenBank: NM_001193316.1:c.373A>T:p.Arg125Ter | Hom | Basal ganglia abnormalities |
Abbreviations are as follows: Hom = homozygous, Het = heterozygous, Hemi = hemizygous, and SMA = spinal muscular atrophy.
Human Loss-of-Function Variants in GWAS Signals
The enrichment of our cohort for loss-of-function events allowed us to encounter such events in genes with only genome-wide association study (GWAS) links in the literature. In one consanguineous family of three children with non-syndromic, early-onset (<5yrs) morbid obesity, we identified a homozygous truncating variant in APOBR (MIM: 605220) (GenBank: NM_018690.3:c.1081_1087del:p.Glu361ProfsTer38). The finding of a likely null variant in HEXA (MIM: 606869) in an individual with early-onset Parkinson disease is less clear given the reported association between a carrier status for other lysosomal disorders and this adult-onset neurodegenerative disease.45
Clinical Utility of Flash Whole-Exome Sequencing
Flash whole-exome sequencing (WES) was reserved for instances when timely molecular diagnosis was needed. Typically, these included instances of fetuses approaching the legal cutoff date of termination (18.5 weeks of gestation), instances when a decision to continue or withdraw aggressive measures of life-support was needed for individuals in the neonatal intensive care unit (NICU) and pediatric intensive care unit (PICU), and instances when children required urgent decisions about liver or bone marrow transplantation. Despite the extremely rapid nature of variant interpretation in Flash WES, the overall yield was comparable to regular WES (48.1% versus 43.3%). Table S10 summarizes our experience with 54 individuals referred to us for Flash WES with various indications.
Founder Effect
Table S6 lists all the founder variants that have been encountered in two or more affected individuals who are not known to be part of the same nuclear family but share the same phenotype and haplotype. Using the broader definition of founder variants (see Material and Methods for details), we found that 41.6% of the pathogenic or likely pathogenic variants qualify as founder variants. Importantly, several of these variants could only be upgraded to pathogenic or likely pathogenic through this strong segregation. It is noteworthy that the contribution of founder variants to the overall pool of disease-causing variants has increased only slightly compared to our earlier pilot study involving ∼350 exomes (41.6% versus 32.5%). Indeed, Table S4 lists the variants from that study that were reclassified on the basis of new data from this study, and we note that only 12 could be upgraded to likely pathogenic. Several founder variants in Table S6 are worth highlighting. For example, we have previously suggested on the basis of much smaller cohorts that the ADAT3 (MIM: 615302) and C12orf57 (MIM: 615140) founder variants are of comparable frequency.14 However, this cohort clearly shows that ADAT3 is the single most common recessive cause of global developmental delay and intellectual disability in the country, accounting for 20 solved instances compared to 4 solved that were based on C12orf57. We also highlight the power of the founder effect in defining genotype-phenotype correlation for variants that would otherwise be classified as VUSs; for example, ATP6V1B2-related cutis laxa syndrome, which we observed in two “unrelated” individuals who have the same syndromic phenotype and share the same missense variant (Table 2).
Secondary Findings
The secondary findings we reported back were of two classes: (1) ACMG-recommended and (2) carrier status for KSMs (see above). The overwhelming majority of cases (1,971 out of 2,219, 88.8%) opted to receive secondary findings in the consent form. Consistent with our previous experience, the percentage of cases with ACMG secondary findings was small (n = 41, 2%), and the findings are listed in Table S5. All of these individuals have been referred for full genetic counseling on the basis of published guidelines relevant to their results, and work is underway to characterize the outcome of these referrals. On the other hand, we have identified a very high carrier frequency of Saudi founder mutations; 697 individuals carry one KSM, 238 carry two KSMs, 56 carry three KSMs, and 15 carry four or more KSMs (Figure S2). We have also extended our analysis to parents and investigated the likelihood for parents to share the carrier status of a founder variant other than the one relevant to the disease in their offspring, i.e., unbiased analysis. We found that only 62 of 503 (12.3%) couples fall in this category (Figure S2).
Discussion
Large-scale CES studies have greatly improved our understanding of the human variome and its medical and clinical relevance. These studies, nearly always from outbred populations, have nearly saturated the morbid genome for the recurrently mutated dominant genes that are linked to human phenotypes in the loss-of-function state.46 Despite their very large size, those studies suffer from the conspicuous underrepresentation of autosomal-recessive, disease-related genes. A very recent analysis of the 6,040 families from the Deciphering Developmental Disorders study estimated the contribution of recessive causes at 3.6%.47 Consanguineous populations, on the other hand, possess a much greater power to unmask recessive, disease-causing alleles, and this discrepancy increases exponentially as the allele frequency decreases.4 For example, a rare recessive allele with a frequency of 0.001 will be observed in homozygosity only once per 1,000,000 individuals in an outbred population (q2) but once per 41,494 individuals in the Saudi population that has an average inbreeding coefficient of 0.0241 (qF).16 Thus, despite the much smaller size compared to the DDD cohort, our cohort revealed a remarkably higher number of autosomal-recessive candidate genes. This large number of candidates, however, raises an interesting question about the rate at which the morbid genome is being saturated for autosomal-recessive, disease-related genes. In a previously reported pilot clinical exome study involving 347 families, a similar percentage of individuals was found to harbor potential autosomal-recessive candidate genes as in this study.14 It seems unlikely, therefore, that all autosomal-recessive disease genes will have been identified by 2020 as originally proposed by the International Rare Diseases Research Consortium (IRDiRC).12 Indeed, it seems likely that many additional thousands of affected individuals from consanguineous populations will have to be sequenced, especially in view of two observations made in this study: (1) the high contribution of “private” mutations that showed little decrement as we increased the number of tested individuals from 347 to >2,200 families and (2) the small percentage of candidate genes for which we were able to observe independent (confirmatory) mutational events. The central processing of CES was key for an internal “matchmaking” that allowed us to identify candidate genes with more than one hit, as well as to confirm founder disease-causing variants that were previously reported as candidates. To this end, we emphasize the tentative candidacy of the genes we propose on the basis of single mutational events, pending future confirmation.
A key advantage of the genomics-first approach taken in this study is made clear by the fact that the overwhelming majority of individuals with solved diagnoses (77.9%) did not carry the correct clinical diagnosis when CES was requested. This is a vivid reminder of the lack of precision in the traditional clinical approach for individuals with suspected genetic diseases and the many missed opportunities for precision clinical management. It is interesting to note that the percentage of individuals that received the correct clinical diagnosis prior to molecular confirmation did not differ significantly (p = 0.29) between those evaluated by the genetic experts (18.6%, n = 547) and non-geneticists (23.5%, n = 409). This is consistent with a prior study in which we compared a traditional clinical approach to a genomics-first approach in >300 instances of intellectual disability and/or developmental delay where we found only 16% of affected individuals had received a clinical suspicion consistent with the molecular finding.11 The limited sensitivity of clinical evaluation even by genetic experts probably reflects the established challenge of the extreme heterogeneity of Mendelian disorders. Our finding also challenges the view that genetic experts alone should request CES, because we do not see evidence that such a practice will lead to a significantly higher yield and, consequently, reduce costs. It is relevant to highlight here that our unusual study design, wherein we encouraged a “genomics-first” approach to maximize the cost-saving benefit of CES, should be taken into account when comparing our results to other studies in which affected individuals typically undergo a “reflex” CES after exhausting a series of other diagnostic tests.
Beyond the high throughput discovery and confirmation of candidate genes, our unique cohort has also provided additional insights. In particular, the high throughput creation of recessive allelic series by virtue of consanguinity loops allowed us to observe important phenotypic aspects previously unknown for the respective genes. The documentation of recessive forms of dominant genes has a significant impact on the interpretation of likely deleterious variants of dominant genes in individuals with no apparent phenotype. The tendency to classify such occurrences as “incomplete penetrance” should be carefully balanced by the growing appreciation that these might indeed represent genuinely recessive forms of genes.36 The impact on the calculation of recurrence risk cannot be overemphasized.
We have previously shown that human knockouts can inform GWAS research into complex phenotypes.48 One informative example in this study is APOBR. This gene is in the same linkage disequilibrium [LD] block as SH2B1, which has been proposed as the source of the obesity GWAS signal in 16p11.2.49, 50 However, our result suggests that APOBR might have also contributed to the obesity signal. On the other hand, lack of florid dyslipidemia in these affected children suggests that the study of APOBR in the context of complex diseases should focus on obesity rather than dyslipidemia.
The remarkable contribution of autosomal-recessive forms of genes to the disease burden in the study population presents an obvious prevention opportunity. Each of the families that had an identified recessive variant received genetic counseling that explored their future reproductive options, and we have previously shown a very high uptake for such services in the country.10 Although private mutations continue to be a significant player, it is encouraging that nearly all the founder variants listed in Table S6 have already been identified by our previous studies. This suggests that the time is ripe for the use of these variants in population-based prevention programs, which are currently in development. One advantage of specifically using these high-quality pathogenic variants is the avoidance of uncertainty associated with interpreting variants for which there is no supporting human genetics evidence; these variants can pose a significant challenge to any screening program. Inexpensive genotyping-based assays that cover all of these variants and allow for a regular update as more of these variants are identified in the population will be ideally suited for primary prevention through expanding Saudi Arabia’s current premarital carrier-screening program, which is limited to sickle cell disease and thalassemia. One can also envision their use in secondary prevention through early diagnosis by incorporating them into the existing Saudi newborn-screening program.
In conclusion, we report in a highly consanguineous population a large clinical exome study that highlights the power of a genomics-first approach to empower clinicians of all backgrounds to partake in the genomic medicine revolution. We also showcase the potential of consanguineous populations to contribute to the medical annotation of the human genome beyond individual discoveries of disease-related genes. The dataset we share through this publication can be a helpful resource for future clinical genomic studies both in outbred and inbred populations.
Declaration of Interests
Authors who are affiliated with Saudi Diagnostic Laboratories are paid employees of King Faisal Specialist Hospital International Holding Company.
Acknowledgments
We thank all those who participated in the clinical care of the study individuals beyond those who requested CES and provided phenotypic data. We acknowledge the support of the Saudi Human Genome Program as well as the Molecular Diagnostic Laboratory administration and staff. The authors declare no conflict of interest.
Published: May 23, 2019
Footnotes
Supplemental Data can be found online at https://doi.org/10.1016/j.ajhg.2019.04.011.
Appendix A: List of Candidate Genes Identified in the Study Cohort
Autosomal Recessive
AARSD1 (MIM: 613212), ABCA13 (MIM: 607807), ACBD3 (MIM: 606809), ACOT4 (MIM: 614314), ACSBG1 (MIM: 614362), AKNA (MIM: 605729), ALKBH8 (MIM: 613306), AP1B1 (MIM: 600157), ATG7 (MIM: 608760), ATRN (MIM: 603130), BCAS3 (MIM: 607470), BECN1 (MIM: 604378), BMP2K (MIM: 617648), BRAP (MIM: 604986), BTNL8 (MIM: 615606), CCDC18, CCDC67 (MIM: 617148), CCDC96, CD34 (MIM: 142230), CEP85, CEP295 (MIM: 617728), CEP131 (MIM: 613479), CFAP46, CFAP221, CHEK1 (MIM: 603078), CMYA5 (MIM: 612193), CNTF (MIM: 118945), COL6A6 (MIM: 616613), CSRP2BP (MIM: 617501), CYB5R4 (MIM: 608343), DCAF15, DEGS1 (MIM: 615843), DERL3 (MIM: 610305), DGKG (MIM: 601854), DIDO1 (MIM: 604140), DMAP1 (MIM: 605077), DMRT2 (MIM: 604935), DMTF1 (MIM: 608491), DNAJA4, DNAJC11 (MIM: 614827), DSCAML1 (MIM: 611782), E2F4 (MIM: 600659), EHBP1L1, EIF2A (MIM: 609234), EIF6 (MIM: 602912), ELFN2, EMSY (MIM: 608574), EPHB6 (MIM: 602757), ERGIC3 (MIM: 616971), FAM69B (MIM: 614543), NCOR2 (MIM: 600848), FAT3 (MIM: 612483), FOSL1 (MIM: 136515), FOXB1, GAD2 (MIM: 138275), GDI2 (MIM: 600767), GLTPD2, GTF2F2 (MIM: 189969), GTF3A (MIM: 600860), HCN2 (MIM: 603781), HSF2 (MIM: 140581), ICE1 (MIM: 617958), IL6ST (MIM: 600694), INADL (MIM: 603199), IQSEC3 (MIM: 612118), ITFG2 (MIM: 617421), ITGA4 (MIM: 192975), KDM5A (MIM: 180202), KDM6B (MIM: 611577), KIAA1024 (MIM: 618054), KIF9 (MIM: 607910), KIF19, KIF13B (MIM: 607350), LAMA5 (MIM: 601033), LETM1 (MIM: 604407), LSR (MIM: 616582), LTBP1 (MIM: 150390), LURAP1 (MIM: 616129), MANEA (MIM: 612327), MAP1B (MIM: 157129), MAP2 (MIM: 157130), MAPK8IP2 (MIM: 607755), MARK1 (MIM: 606511), MCM3 (MIM: 602693), MED18 (MIM: 612384), METTL12 (MIM: 617897), MFN1 (MIM: 608506), MICAL1 (MIM: 607129), MICAL3 (MIM: 608882), MMP10 (MIM: 185260), MTCL1 (MIM: 615766), NAA20 (MIM: 610833), NAIP (MIM: 600355), NRDC (MIM: 602651), NRP1 (MIM: 602069), NUP98 (MIM: 601021), P2RX6 (MIM: 608077), PAK1IP1 (MIM: 607811), PARD3 (MIM: 606745), PARP10 (MIM: 609564), PARP16, PHF14, PIGB (MIM: 604122), PLCH1 (MIM: 612835), PLEKHH2 (MIM: 612723), PLIN5 (MIM: 613248), PLXNA1 (MIM: 601055), PLXNA2 (MIM: 601054), PRSS8 (MIM: 600823), PTPN12 (MIM: 600079), PUS7L, RINT1 (MIM: 610089), SALL3 (MIM: 605079), SCHIP1, SCNN1D (MIM: 601328), SCYL2 (MIM: 616365), SHQ1 (MIM: 613663), SIRPB2 (MIM: 605466), SLC38A3 (MIM: 604437), SLC7A1 (MIM: 104615), SLIT1 (MIM: 603742), SLK (MIM: 616563), SMG8 (MIM: 613175), SMPD4 (MIM: 610457), SOGA1, STXBP3 (MIM: 608339), SUSD2 (MIM: 615825), TAF10 (MIM: 600475), TENM2 (MIM: 610119), THOC3 (MIM: 606929), TIMM8B (MIM: 606659), TMEM130, TMEM17 (MIM: 614950), TMEM212, TNFRSF9 (MIM: 602250), TP73 (MIM: 601990), TTLL10, TTLL6 (MIM: 610849), UGCG (MIM: 602874), UNC13B (MIM: 605836), UNC13C (MIM: 614568), UNC5A (MIM: 607869), USO1 (MIM: 603344), VAT1L, VCAM1 (MIM: 192225), VPS51 (MIM: 615738), WDR6 (MIM: 606031), WDR91 (MIM: 616303), WWTR1 (MIM: 607392), XIRP2 (MIM: 609778), XRCC5 (MIM: 194364), ZMIZ2 (MIM: 611196), ZNF280C, ZNF707, and ZYX (MIM: 602002).
Autosomal Dominant
ACAT2 (MIM: 100678), ACMSD (MIM: 608889), ACO1 (MIM: 100880), AMBRA1 (MIM: 611359), AP2B1 (MIM: 601025), BCL9 (MIM: 602597), BTBD1 (MIM: 608530), CACNA1E (MIM: 601013), CACNA2D1 (MIM: 114204), CDH8 (MIM: 610528), CDH9 (MIM: 609974), CEP89 (MIM: 615470), CHD5 (MIM: 610771), CHD6 (MIM: 616114), CREBRF (MIM: 617109), DLGAP2 (MIM: 605438), DLX1 (MIM: 600029), DSCAM (MIM: 602523), ESCO1 (MIM: 609674), EYA3 (MIM: 601655), FASN (MIM: 600212), FRMD6 (MIM: 614555), FRYL, FYTTD1 (MIM: 616933), FZD9 (MIM: 601766), GLTSCR2 (MIM: 605691), HDAC4 (MIM: 605314), HOXC4 (MIM: 142974), KDM2B (MIM: 609078), KIAA1328 (MIM: 616480), KIF5B (MIM: 602809), KLF4 (MIM: 602253), LRRC4C (MIM: 608817), MGA (MIM: 616061), MOV10 (MIM: 610742), MYO16 (MIM: 615479), MYO18A (MIM: 610067), MYO1A (MIM: 601478), MYO9B (MIM: 602129), MYT1 (MIM: 600379), NAV1 (MIM: 611628), OSBPL9 (MIM: 606737), P2RY13 (MIM: 606380), PLK2 (MIM: 607023), PLPPR3 (MIM: 610391), PML (MIM: 102578), PRKACA (MIM: 601639), PTPN12 (MIM: 600079), RASIP1 (MIM: 609623), RBM14 (MIM: 612409), RNF165, RPTOR (MIM: 607130), SEPT2 (MIM: 601506), SETDB1 (MIM: 604396), SHANK1 (MIM: 604999), SMARCC2 (MIM: 601734), SMG6 (MIM: 610963), SPEN (MIM: 613484), TBXA2R (MIM: 188070), TJP1 (MIM: 601009), TLN1 (MIM: 186745), TNMD (MIM: 300459), TOMM40 (MIM: 608061), TRIM7 (MIM: 609315), TRPV5 (MIM: 606679), UNK (MIM: 616375), USF3 (MIM: 617568), USP33 (MIM: 615146), WWP1 (MIM: 602307), XPNPEP1 (MIM: 602443), XRCC6 (MIM: 152690), ZBED6 (MIM: 613512), ZFHX3 (MIM: 104155), and ZPR1 (MIM: 603901).
X-Linked
CDR1 (MIM: 302650), KLF8 (MIM: 300286), PLXNA3 (MIM: 300022), RBBP7 (MIM: 300825), RPS6KA6 (MIM: 300303), SLITRK4 (MIM: 300562), and SMARCA1 (MIM: 300012).
Web Resources
OMIM, http://omim.org/
Supplemental Data
Table S2. List of Previously Reported Candidate Genes whose Candidacy Is Further Supported by Findings in this Cohort. Table S8. List of Couples Who Presented for Counseling with No Available DNA From the Affected Deceased Child(ren) or Fetus(es). Table S10. List of Individuals for Whom Flash WES Protocol was Implemented.
References
- 1.Alkuraya F.S. Impact of new genomic tools on the practice of clinical genetics in consanguineous populations: The Saudi experience. Clin. Genet. 2013;84:203–208. doi: 10.1111/cge.12131. [DOI] [PubMed] [Google Scholar]; Alkuraya, F.S. (2013). Impact of new genomic tools on the practice of clinical genetics in consanguineous populations: The Saudi experience. Clin. Genet. 84, 203-208. [DOI] [PubMed]
- 2.Alkuraya F.S. Genetics and genomic medicine in Saudi Arabia. Mol. Genet. Genomic Med. 2014;2:369–378. doi: 10.1002/mgg3.97. [DOI] [PMC free article] [PubMed] [Google Scholar]; Alkuraya, F.S. (2014). Genetics and genomic medicine in Saudi Arabia. Mol. Genet. Genomic Med. 2, 369-378. [DOI] [PMC free article] [PubMed]
- 3.Alkuraya F.S. Autozygome decoded. Genet. Med. 2010;12:765–771. doi: 10.1097/GIM.0b013e3181fbfcc4. [DOI] [PubMed] [Google Scholar]; Alkuraya, F.S. (2010). Autozygome decoded. Genet. Med. 12, 765-771. [DOI] [PubMed]
- 4.Alkuraya F.S. Human knockout research: New horizons and opportunities. Trends Genet. 2015;31:108–115. doi: 10.1016/j.tig.2014.11.003. [DOI] [PubMed] [Google Scholar]; Alkuraya, F.S. (2015). Human knockout research: New horizons and opportunities. Trends Genet. 31, 108-115. [DOI] [PubMed]
- 5.Alsalem A.B., Halees A.S., Anazi S., Alshamekh S., Alkuraya F.S. Autozygome sequencing expands the horizon of human knockout research and provides novel insights into human phenotypic variation. PLoS Genet. 2013;9:e1004030. doi: 10.1371/journal.pgen.1004030. [DOI] [PMC free article] [PubMed] [Google Scholar]; Alsalem, A.B., Halees, A.S., Anazi, S., Alshamekh, S., and Alkuraya, F.S. (2013). Autozygome sequencing expands the horizon of human knockout research and provides novel insights into human phenotypic variation. PLoS Genet. 9, e1004030. [DOI] [PMC free article] [PubMed]
- 6.Alkuraya F.S. Discovery of rare homozygous mutations from studies of consanguineous pedigrees. Curr. Protoc. Hum. Genet. 2012;75 doi: 10.1002/0471142905.hg0612s75. [DOI] [PubMed] [Google Scholar]; Alkuraya, F.S. (2012). Discovery of rare homozygous mutations from studies of consanguineous pedigrees. Curr. Protoc. Hum. Genet. 75. [DOI] [PubMed]
- 7.Scott E.M., Halees A., Itan Y., Spencer E.G., He Y., Azab M.A., Gabriel S.B., Belkadi A., Boisson B., Abel L., Greater Middle East Variome Consortium Characterization of Greater Middle Eastern genetic variation for enhanced disease gene discovery. Nat. Genet. 2016;48:1071–1076. doi: 10.1038/ng.3592. [DOI] [PMC free article] [PubMed] [Google Scholar]; Scott, E.M., Halees, A., Itan, Y., Spencer, E.G., He, Y., Azab, M.A., Gabriel, S.B., Belkadi, A., Boisson, B., Abel, L., et al.; Greater Middle East Variome Consortium (2016). Characterization of Greater Middle Eastern genetic variation for enhanced disease gene discovery. Nat. Genet. 48, 1071-1076. [DOI] [PMC free article] [PubMed]
- 8.Alazami A.M., Patel N., Shamseldin H.E., Anazi S., Al-Dosari M.S., Alzahrani F., Hijazi H., Alshammari M., Aldahmesh M.A., Salih M.A. Accelerating novel candidate gene discovery in neurogenetic disorders via whole-exome sequencing of prescreened multiplex consanguineous families. Cell Rep. 2015;10:148–161. doi: 10.1016/j.celrep.2014.12.015. [DOI] [PubMed] [Google Scholar]; Alazami, A.M., Patel, N., Shamseldin, H.E., Anazi, S., Al-Dosari, M.S., Alzahrani, F., Hijazi, H., Alshammari, M., Aldahmesh, M.A., Salih, M.A., et al. (2015). Accelerating novel candidate gene discovery in neurogenetic disorders via whole-exome sequencing of prescreened multiplex consanguineous families. Cell Rep. 10, 148-161. [DOI] [PubMed]
- 9.Shaheen R., Patel N., Shamseldin H., Alzahrani F., Al-Yamany R., ALMoisheer A., Ewida N., Anazi S., Alnemer M., Elsheikh M. Accelerating matchmaking of novel dysmorphology syndromes through clinical and genomic characterization of a large cohort. Genet. Med. 2016;18:686–695. doi: 10.1038/gim.2015.147. [DOI] [PubMed] [Google Scholar]; Shaheen, R., Patel, N., Shamseldin, H., Alzahrani, F., Al-Yamany, R., ALMoisheer, A., Ewida, N., Anazi, S., Alnemer, M., Elsheikh, M., et al. (2016). Accelerating matchmaking of novel dysmorphology syndromes through clinical and genomic characterization of a large cohort. Genet. Med. 18, 686-695. [DOI] [PubMed]
- 10.Alkuraya F.S. The application of next-generation sequencing in the autozygosity mapping of human recessive diseases. Hum. Genet. 2013;132:1197–1211. doi: 10.1007/s00439-013-1344-x. [DOI] [PubMed] [Google Scholar]; Alkuraya, F.S. (2013). The application of next-generation sequencing in the autozygosity mapping of human recessive diseases. Hum. Genet. 132, 1197-1211. [DOI] [PubMed]
- 11.Anazi S., Maddirevula S., Faqeih E., Alsedairy H., Alzahrani F., Shamseldin H.E., Patel N., Hashem M., Ibrahim N., Abdulwahab F. Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield. Mol. Psychiatry. 2017;22:615–624. doi: 10.1038/mp.2016.113. [DOI] [PubMed] [Google Scholar]; Anazi, S., Maddirevula, S., Faqeih, E., Alsedairy, H., Alzahrani, F., Shamseldin, H.E., Patel, N., Hashem, M., Ibrahim, N., Abdulwahab, F., et al. (2017). Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield. Mol. Psychiatry 22, 615-624. [DOI] [PubMed]
- 12.Boycott K.M., Rath A., Chong J.X., Hartley T., Alkuraya F.S., Baynam G., Brookes A.J., Brudno M., Carracedo A., den Dunnen J.T. International cooperation to enable the diagnosis of all rare genetic diseases. Am. J. Hum. Genet. 2017;100:695–705. doi: 10.1016/j.ajhg.2017.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]; Boycott, K.M., Rath, A., Chong, J.X., Hartley, T., Alkuraya, F.S., Baynam, G., Brookes, A.J., Brudno, M., Carracedo, A., den Dunnen, J.T., et al. (2017). International cooperation to enable the diagnosis of all rare genetic diseases. Am. J. Hum. Genet. 100, 695-705. [DOI] [PMC free article] [PubMed]
- 13.Group S.M., Saudi Mendeliome Group Comprehensive gene panels provide advantages over clinical exome sequencing for Mendelian diseases. Genome Biol. 2015;16:134. doi: 10.1186/s13059-015-0693-2. [DOI] [PMC free article] [PubMed] [Google Scholar]; Group, S.M.; Saudi Mendeliome Group (2015). Comprehensive gene panels provide advantages over clinical exome sequencing for Mendelian diseases. Genome Biol. 16, 134. [DOI] [PMC free article] [PubMed]
- 14.Monies D., Abouelhoda M., AlSayed M., Alhassnan Z., Alotaibi M., Kayyali H., Al-Owain M., Shah A., Rahbeeni Z., Al-Muhaizea M.A. The landscape of genetic diseases in Saudi Arabia based on the first 1000 diagnostic panels and exomes. Hum. Genet. 2017;136:921–939. doi: 10.1007/s00439-017-1821-8. [DOI] [PMC free article] [PubMed] [Google Scholar]; Monies, D., Abouelhoda, M., AlSayed, M., Alhassnan, Z., Alotaibi, M., Kayyali, H., Al-Owain, M., Shah, A., Rahbeeni, Z., Al-Muhaizea, M.A., et al. (2017). The landscape of genetic diseases in Saudi Arabia based on the first 1000 diagnostic panels and exomes. Hum. Genet. 136, 921-939. [DOI] [PMC free article] [PubMed]
- 15.Kalia S.S., Adelman K., Bale S.J., Chung W.K., Eng C., Evans J.P., Herman G.E., Hufnagel S.B., Klein T.E., Korf B.R. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): A policy statement of the American College of Medical Genetics and Genomics. Genet. Med. 2017;19:249–255. doi: 10.1038/gim.2016.190. [DOI] [PubMed] [Google Scholar]; Kalia, S.S., Adelman, K., Bale, S.J., Chung, W.K., Eng, C., Evans, J.P., Herman, G.E., Hufnagel, S.B., Klein, T.E., Korf, B.R., et al. (2017). Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): A policy statement of the American College of Medical Genetics and Genomics. Genet. Med. 19, 249-255. [DOI] [PubMed]
- 16.Abouelhoda M., Sobahy T., El-Kalioby M., Patel N., Shamseldin H., Monies D., Al-Tassan N., Ramzan K., Imtiaz F., Shaheen R., Alkuraya F.S. Clinical genomics can facilitate countrywide estimation of autosomal recessive disease burden. Genet. Med. 2016;18:1244–1249. doi: 10.1038/gim.2016.37. [DOI] [PubMed] [Google Scholar]; Abouelhoda, M., Sobahy, T., El-Kalioby, M., Patel, N., Shamseldin, H., Monies, D., Al-Tassan, N., Ramzan, K., Imtiaz, F., Shaheen, R., and Alkuraya, F.S. (2016). Clinical genomics can facilitate countrywide estimation of autosomal recessive disease burden. Genet. Med. 18, 1244-1249. [DOI] [PubMed]
- 17.Richards S., Aziz N., Bale S., Bick D., Das S., Gastier-Foster J., Grody W.W., Hegde M., Lyon E., Spector E., ACMG Laboratory Quality Assurance Committee Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015;17:405–424. doi: 10.1038/gim.2015.30. [DOI] [PMC free article] [PubMed] [Google Scholar]; Richards, S., Aziz, N., Bale, S., Bick, D., Das, S., Gastier-Foster, J., Grody, W.W., Hegde, M., Lyon, E., Spector, E., et al.; ACMG Laboratory Quality Assurance Committee (2015). Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405-424. [DOI] [PMC free article] [PubMed]
- 18.Monies D., Vågbø C.B., Al-Owain M., Alhomaidi S., Alkuraya F.S. Recessive Truncating Mutations in ALKBH8 Cause Severe Impairment of Wobble Uridine Modification and Intellectual Disability. American Journal of Human Genetics. 2019;104 doi: 10.1016/j.ajhg.2019.03.026. [DOI] [PMC free article] [PubMed] [Google Scholar]; Monies, D., Vagbo, C.B., Al-Owain, M., Alhomaidi, S., and Alkuraya, F.S. (2019). Recessive Truncating Mutations in ALKBH8 Cause Severe Impairment of Wobble Uridine Modification and Intellectual Disability. American Journal of Human Genetics, 104. https://doi.org/10.1016/j.ajhg.2019.03.026 [DOI] [PMC free article] [PubMed]
- 19.Tyska M.J., Mackey A.T., Huang J.-D., Copeland N.G., Jenkins N.A., Mooseker M.S. Myosin-1a is critical for normal brush border structure and composition. Mol. Biol. Cell. 2005;16:2443–2457. doi: 10.1091/mbc.E04-12-1116. [DOI] [PMC free article] [PubMed] [Google Scholar]; Tyska, M.J., Mackey, A.T., Huang, J.-D., Copeland, N.G., Jenkins, N.A., and Mooseker, M.S. (2005). Myosin-1a is critical for normal brush border structure and composition. Mol. Biol. Cell 16, 2443-2457. [DOI] [PMC free article] [PubMed]
- 20.Gingras S., Earls L.R., Howell S., Smeyne R.J., Zakharenko S.S., Pelletier S. SCYL2 protects CA3 pyramidal neurons from excitotoxicity during functional maturation of the mouse hippocampus. J. Neurosci. 2015;35:10510–10522. doi: 10.1523/JNEUROSCI.2056-14.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]; Gingras, S., Earls, L.R., Howell, S., Smeyne, R.J., Zakharenko, S.S., and Pelletier, S. (2015). SCYL2 protects CA3 pyramidal neurons from excitotoxicity during functional maturation of the mouse hippocampus. J. Neurosci. 35, 10510-10522. [DOI] [PMC free article] [PubMed]
- 21.Banks G., Lassi G., Hoerder-Suabedissen A., Tinarelli F., Simon M.M., Wilcox A., Lau P., Lawson T.N., Johnson S., Rutman A. A missense mutation in Katnal1 underlies behavioural, neurological and ciliary anomalies. Mol. Psychiatry. 2018;23:713–722. doi: 10.1038/mp.2017.54. [DOI] [PMC free article] [PubMed] [Google Scholar]; Banks, G., Lassi, G., Hoerder-Suabedissen, A., Tinarelli, F., Simon, M.M., Wilcox, A., Lau, P., Lawson, T.N., Johnson, S., Rutman, A., et al. (2018). A missense mutation in Katnal1 underlies behavioural, neurological and ciliary anomalies. Mol. Psychiatry 23, 713-722. [DOI] [PMC free article] [PubMed]
- 22.Ohno M., Hiraoka Y., Matsuoka T., Tomimoto H., Takao K., Miyakawa T., Oshima N., Kiyonari H., Kimura T., Kita T., Nishi E. Nardilysin regulates axonal maturation and myelination in the central and peripheral nervous system. Nat. Neurosci. 2009;12:1506–1513. doi: 10.1038/nn.2438. [DOI] [PubMed] [Google Scholar]; Ohno, M., Hiraoka, Y., Matsuoka, T., Tomimoto, H., Takao, K., Miyakawa, T., Oshima, N., Kiyonari, H., Kimura, T., Kita, T., and Nishi, E. (2009). Nardilysin regulates axonal maturation and myelination in the central and peripheral nervous system. Nat. Neurosci. 12, 1506-1513. [DOI] [PubMed]
- 23.Shaheen R., Szymanska K., Basu B., Patel N., Ewida N., Faqeih E., Al Hashem A., Derar N., Alsharif H., Aldahmesh M.A., Ciliopathy WorkingGroup Characterizing the morbid genome of ciliopathies. Genome Biol. 2016;17:242. doi: 10.1186/s13059-016-1099-5. [DOI] [PMC free article] [PubMed] [Google Scholar]; Shaheen, R., Szymanska, K., Basu, B., Patel, N., Ewida, N., Faqeih, E., Al Hashem, A., Derar, N., Alsharif, H., Aldahmesh, M.A., et al.; Ciliopathy WorkingGroup (2016). Characterizing the morbid genome of ciliopathies. Genome Biol. 17, 242. [DOI] [PMC free article] [PubMed]
- 24.Firat-Karalar E.N., Sante J., Elliott S., Stearns T. Proteomic analysis of mammalian sperm cells identifies new components of the centrosome. J. Cell Sci. 2014;127:4128–4133. doi: 10.1242/jcs.157008. [DOI] [PMC free article] [PubMed] [Google Scholar]; Firat-Karalar, E.N., Sante, J., Elliott, S., and Stearns, T. (2014). Proteomic analysis of mammalian sperm cells identifies new components of the centrosome. J. Cell Sci. 127, 4128-4133. [DOI] [PMC free article] [PubMed]
- 25.Nasir A., Le Bail A., Daiker V., Klima J., Richter P., Lebert M. Identification of a flagellar protein implicated in the gravitaxis in the flagellate Euglena gracilis. Sci. Rep. 2018;8:7605. doi: 10.1038/s41598-018-26046-8. [DOI] [PMC free article] [PubMed] [Google Scholar]; Nasir, A., Le Bail, A., Daiker, V., Klima, J., Richter, P., and Lebert, M. (2018). Identification of a flagellar protein implicated in the gravitaxis in the flagellate Euglena gracilis. Sci. Rep. 8, 7605. [DOI] [PMC free article] [PubMed]
- 26.Tsuchiya Y., Yoshiba S., Gupta A., Watanabe K., Kitagawa D. Cep295 is a conserved scaffold protein required for generation of a bona fide mother centriole. Nat. Commun. 2016;7:12567. doi: 10.1038/ncomms12567. [DOI] [PMC free article] [PubMed] [Google Scholar]; Tsuchiya, Y., Yoshiba, S., Gupta, A., Watanabe, K., and Kitagawa, D. (2016). Cep295 is a conserved scaffold protein required for generation of a bona fide mother centriole. Nat. Commun. 7, 12567. [DOI] [PMC free article] [PubMed]
- 27.Adly N., Alhashem A., Ammari A., Alkuraya F.S. Ciliary genes TBC1D32/C6orf170 and SCLT1 are mutated in patients with OFD type IX. Hum. Mutat. 2014;35:36–40. doi: 10.1002/humu.22477. [DOI] [PubMed] [Google Scholar]; Adly, N., Alhashem, A., Ammari, A., and Alkuraya, F.S. (2014). Ciliary genes TBC1D32/C6orf170 and SCLT1 are mutated in patients with OFD type IX. Hum. Mutat. 35, 36-40. [DOI] [PubMed]
- 28.Maddirevula S., Alsahli S., Alhabeeb L., Patel N., Alzahrani F., Shamseldin H.E., Anazi S., Ewida N., Alsaif H.S., Mohamed J.Y. Expanding the phenome and variome of skeletal dysplasia. Genet. Med. 2018;20:1609–1616. doi: 10.1038/gim.2018.50. [DOI] [PubMed] [Google Scholar]; Maddirevula, S., Alsahli, S., Alhabeeb, L., Patel, N., Alzahrani, F., Shamseldin, H.E., Anazi, S., Ewida, N., Alsaif, H.S., Mohamed, J.Y., et al. (2018). Expanding the phenome and variome of skeletal dysplasia. Genet. Med. 20, 1609-1616. [DOI] [PubMed]
- 29.Patel N., El Mouzan M.I., Al-Mayouf S.M., Adly N., Mohamed J.Y., Al Mofarreh M.A., Ibrahim N., Xiong Y., Zhao Q., Al-Saleem K.A., Alkuraya F.S. Study of Mendelian forms of Crohn’s disease in Saudi Arabia reveals novel risk loci and alleles. Gut. 2014;63:1831–1832. doi: 10.1136/gutjnl-2014-307859. [DOI] [PubMed] [Google Scholar]; Patel, N., El Mouzan, M.I., Al-Mayouf, S.M., Adly, N., Mohamed, J.Y., Al Mofarreh, M.A., Ibrahim, N., Xiong, Y., Zhao, Q., Al-Saleem, K.A., and Alkuraya, F.S. (2014). Study of Mendelian forms of Crohn’s disease in Saudi Arabia reveals novel risk loci and alleles. Gut 63, 1831-1832. [DOI] [PubMed]
- 30.Aldahmesh M.A., Khan A.O., Mohamed J., Alkuraya F.S. Novel recessive BFSP2 and PITX3 mutations: Insights into mutational mechanisms from consanguineous populations. Genet. Med. 2011;13:978–981. doi: 10.1097/GIM.0b013e31822623d5. [DOI] [PubMed] [Google Scholar]; Aldahmesh, M.A., Khan, A.O., Mohamed, J., and Alkuraya, F.S. (2011). Novel recessive BFSP2 and PITX3 mutations: Insights into mutational mechanisms from consanguineous populations. Genet. Med. 13, 978-981. [DOI] [PubMed]
- 31.Anazi S., Maddirevula S., Salpietro V., Asi Y.T., Alsahli S., Alhashem A., Shamseldin H.E., AlZahrani F., Patel N., Ibrahim N. Expanding the genetic heterogeneity of intellectual disability. Hum. Genet. 2017;136:1419–1429. doi: 10.1007/s00439-017-1843-2. [DOI] [PubMed] [Google Scholar]; Anazi, S., Maddirevula, S., Salpietro, V., Asi, Y.T., Alsahli, S., Alhashem, A., Shamseldin, H.E., AlZahrani, F., Patel, N., Ibrahim, N., et al. (2017). Expanding the genetic heterogeneity of intellectual disability. Hum. Genet. 136, 1419-1429. [DOI] [PubMed]
- 32.Shaheen R., Anazi S., Ben-Omran T., Seidahmed M.Z., Caddle L.B., Palmer K., Ali R., Alshidi T., Hagos S., Goodwin L. Mutations in SMG9, encoding an essential component of nonsense-mediated decay machinery, cause a multiple congenital anomaly syndrome in humans and mice. Am. J. Hum. Genet. 2016;98:643–652. doi: 10.1016/j.ajhg.2016.02.010. [DOI] [PMC free article] [PubMed] [Google Scholar]; Shaheen, R., Anazi, S., Ben-Omran, T., Seidahmed, M.Z., Caddle, L.B., Palmer, K., Ali, R., Alshidi, T., Hagos, S., Goodwin, L., et al. (2016). Mutations in SMG9, encoding an essential component of nonsense-mediated decay machinery, cause a multiple congenital anomaly syndrome in humans and mice. Am. J. Hum. Genet. 98, 643-652. [DOI] [PMC free article] [PubMed]
- 33.Van De Weghe J.C., Rusterholz T.D.S., Latour B., Grout M.E., Aldinger K.A., Shaheen R., Dempsey J.C., Maddirevula S., Cheng Y.-H.H., Phelps I.G., University of Washington Center for Mendelian Genomics Mutations in ARMC9, which encodes a basal body protein, cause Joubert syndrome in humans and ciliopathy phenotypes in zebrafish. Am. J. Hum. Genet. 2017;101:23–36. doi: 10.1016/j.ajhg.2017.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]; Van De Weghe, J.C., Rusterholz, T.D.S., Latour, B., Grout, M.E., Aldinger, K.A., Shaheen, R., Dempsey, J.C., Maddirevula, S., Cheng, Y.-H.H., Phelps, I.G., et al.; University of Washington Center for Mendelian Genomics (2017). Mutations in ARMC9, which encodes a basal body protein, cause Joubert syndrome in humans and ciliopathy phenotypes in zebrafish. Am. J. Hum. Genet. 101, 23-36. [DOI] [PMC free article] [PubMed]
- 34.Patel N., Aldahmesh M.A., Alkuraya H., Anazi S., Alsharif H., Khan A.O., Sunker A., Al-Mohsen S., Abboud E.B., Nowilaty S.R. Expanding the clinical, allelic, and locus heterogeneity of retinal dystrophies. Genet. Med. 2016;18:554–562. doi: 10.1038/gim.2015.127. [DOI] [PubMed] [Google Scholar]; Patel, N., Aldahmesh, M.A., Alkuraya, H., Anazi, S., Alsharif, H., Khan, A.O., Sunker, A., Al-Mohsen, S., Abboud, E.B., Nowilaty, S.R., et al. (2016). Expanding the clinical, allelic, and locus heterogeneity of retinal dystrophies. Genet. Med. 18, 554-562. [DOI] [PubMed]
- 35.Shaheen R., Al Hashem A., Alghamdi M.H., Seidahmad M.Z., Wakil S.M., Dagriri K., Keavney B., Goodship J., Alyousif S., Al-Habshan F.M. Positional mapping of PRKD1, NRP1 and PRDM1 as novel candidate disease genes in truncus arteriosus. J. Med. Genet. 2015;52:322–329. doi: 10.1136/jmedgenet-2015-102992. [DOI] [PubMed] [Google Scholar]; Shaheen, R., Al Hashem, A., Alghamdi, M.H., Seidahmad, M.Z., Wakil, S.M., Dagriri, K., Keavney, B., Goodship, J., Alyousif, S., Al-Habshan, F.M., et al. (2015). Positional mapping of PRKD1, NRP1 and PRDM1 as novel candidate disease genes in truncus arteriosus. J. Med. Genet. 52, 322-329. [DOI] [PubMed]
- 36.Monies D., Maddirevula S., Kurdi W., Alanazy M.H., Alkhalidi H., Al-Owain M., Sulaiman R.A., Faqeih E., Goljan E., Ibrahim N. Autozygosity reveals recessive mutations and novel mechanisms in dominant genes: Implications in variant interpretation. Genet. Med. 2017;19:1144–1150. doi: 10.1038/gim.2017.22. [DOI] [PubMed] [Google Scholar]; Monies, D., Maddirevula, S., Kurdi, W., Alanazy, M.H., Alkhalidi, H., Al-Owain, M., Sulaiman, R.A., Faqeih, E., Goljan, E., Ibrahim, N., et al. (2017). Autozygosity reveals recessive mutations and novel mechanisms in dominant genes: Implications in variant interpretation. Genet. Med. 19, 1144-1150. [DOI] [PubMed]
- 37.Alazami A.M., Al-Qattan S.M., Faqeih E., Alhashem A., Alshammari M., Alzahrani F., Al-Dosari M.S., Patel N., Alsagheir A., Binabbas B. Expanding the clinical and genetic heterogeneity of hereditary disorders of connective tissue. Hum. Genet. 2016;135:525–540. doi: 10.1007/s00439-016-1660-z. [DOI] [PubMed] [Google Scholar]; Alazami, A.M., Al-Qattan, S.M., Faqeih, E., Alhashem, A., Alshammari, M., Alzahrani, F., Al-Dosari, M.S., Patel, N., Alsagheir, A., Binabbas, B., et al. (2016). Expanding the clinical and genetic heterogeneity of hereditary disorders of connective tissue. Hum. Genet. 135, 525-540. [DOI] [PubMed]
- 38.Shaheen R., Alazami A.M., Alshammari M.J., Faqeih E., Alhashmi N., Mousa N., Alsinani A., Ansari S., Alzahrani F., Al-Owain M. Study of autosomal recessive osteogenesis imperfecta in Arabia reveals a novel locus defined by TMEM38B mutation. J. Med. Genet. 2012;49:630–635. doi: 10.1136/jmedgenet-2012-101142. [DOI] [PubMed] [Google Scholar]; Shaheen, R., Alazami, A.M., Alshammari, M.J., Faqeih, E., Alhashmi, N., Mousa, N., Alsinani, A., Ansari, S., Alzahrani, F., Al-Owain, M., et al. (2012). Study of autosomal recessive osteogenesis imperfecta in Arabia reveals a novel locus defined by TMEM38B mutation. J. Med. Genet. 49, 630-635. [DOI] [PubMed]
- 39.Anazi S., Al-Sabban E., Alkuraya F.S. Gonadal mosaicism as a rare cause of autosomal recessive inheritance. Clin. Genet. 2014;85:278–281. doi: 10.1111/cge.12156. [DOI] [PubMed] [Google Scholar]; Anazi, S., Al-Sabban, E., and Alkuraya, F.S. (2014). Gonadal mosaicism as a rare cause of autosomal recessive inheritance. Clin. Genet. 85, 278-281. [DOI] [PubMed]
- 40.Seidahmed M.Z., Salih M.A., Abdelbasit O.B., Alassiri A.H., Hussein K.A., Miqdad A., Samadi A., Rasheed A.A., Alorainy I.A., Shaheen R., Alkuraya F.S. Gonadal mosaicism for ACTA1 gene masquerading as autosomal recessive nemaline myopathy. Am. J. Med. Genet. A. 2016;170:2219–2221. doi: 10.1002/ajmg.a.37768. [DOI] [PubMed] [Google Scholar]; Seidahmed, M.Z., Salih, M.A., Abdelbasit, O.B., Alassiri, A.H., Hussein, K.A., Miqdad, A., Samadi, A., Rasheed, A.A., Alorainy, I.A., Shaheen, R., and Alkuraya, F.S. (2016). Gonadal mosaicism for ACTA1 gene masquerading as autosomal recessive nemaline myopathy. Am. J. Med. Genet. A. 170, 2219-2221. [DOI] [PubMed]
- 41.Kulkarni N., Tang S., Bhardwaj R., Bernes S., Grebe T.A. Progressive movement disorder in brothers carrying a GNAO1 mutation responsive to deep brain stimulation. J. Child Neurol. 2016;31:211–214. doi: 10.1177/0883073815587945. [DOI] [PubMed] [Google Scholar]; Kulkarni, N., Tang, S., Bhardwaj, R., Bernes, S., and Grebe, T.A. (2016). Progressive movement disorder in brothers carrying a GNAO1 mutation responsive to deep brain stimulation. J. Child Neurol. 31, 211-214. [DOI] [PubMed]
- 42.Abouhamed M., Grobe K., San I.V., Thelen S., Honnert U., Balda M.S., Matter K., Bähler M. Myosin IXa regulates epithelial differentiation and its deficiency results in hydrocephalus. Mol. Biol. Cell. 2009;20:5074–5085. doi: 10.1091/mbc.E09-04-0291. [DOI] [PMC free article] [PubMed] [Google Scholar]; Abouhamed, M., Grobe, K., San, I.V., Thelen, S., Honnert, U., Balda, M.S., Matter, K., and Bahler, M. (2009). Myosin IXa regulates epithelial differentiation and its deficiency results in hydrocephalus. Mol. Biol. Cell 20, 5074-5085. [DOI] [PMC free article] [PubMed]
- 43.Shamseldin H.E., Kurdi W., Almusafri F., Alnemer M., Alkaff A., Babay Z., Alhashem A., Tulbah M., Alsahan N., Khan R. Molecular autopsy in maternal-fetal medicine. Genet. Med. 2018;20:420–427. doi: 10.1038/gim.2017.111. [DOI] [PubMed] [Google Scholar]; Shamseldin, H.E., Kurdi, W., Almusafri, F., Alnemer, M., Alkaff, A., Babay, Z., Alhashem, A., Tulbah, M., Alsahan, N., Khan, R., et al. (2018). Molecular autopsy in maternal-fetal medicine. Genet. Med. 20, 420-427. [DOI] [PubMed]
- 44.Abu-Safieh L., Abboud E.B., Alkuraya H., Shamseldin H., Al-Enzi S., Al-Abdi L., Hashem M., Colak D., Jarallah A., Ahmad H. Mutation of IGFBP7 causes upregulation of BRAF/MEK/ERK pathway and familial retinal arterial macroaneurysms. Am. J. Hum. Genet. 2011;89:313–319. doi: 10.1016/j.ajhg.2011.07.010. [DOI] [PMC free article] [PubMed] [Google Scholar]; Abu-Safieh, L., Abboud, E.B., Alkuraya, H., Shamseldin, H., Al-Enzi, S., Al-Abdi, L., Hashem, M., Colak, D., Jarallah, A., Ahmad, H., et al. (2011). Mutation of IGFBP7 causes upregulation of BRAF/MEK/ERK pathway and familial retinal arterial macroaneurysms. Am. J. Hum. Genet. 89, 313-319. [DOI] [PMC free article] [PubMed]
- 45.Cookson M.R. A feedforward loop links Gaucher and Parkinson’s diseases? Cell. 2011;146:9–11. doi: 10.1016/j.cell.2011.06.031. [DOI] [PubMed] [Google Scholar]; Cookson, M.R. (2011). A feedforward loop links Gaucher and Parkinson’s diseases? Cell 146, 9-11. [DOI] [PubMed]
- 46.Study T.D.D.D., Deciphering Developmental Disorders Study Prevalence and architecture of de novo mutations in developmental disorders. Nature. 2017;542:433–438. doi: 10.1038/nature21062. [DOI] [PMC free article] [PubMed] [Google Scholar]; Study, T.D.D.D.; Deciphering Developmental Disorders Study (2017). Prevalence and architecture of de novo mutations in developmental disorders. Nature 542, 433-438. [DOI] [PMC free article] [PubMed]
- 47.Martin H.C., Jones W.D., McIntyre R., Sanchez-Andrade G., Sanderson M., Stephenson J.D., Jones C.P., Handsaker J., Gallone G., Bruntraeger M. Quantifying the contribution of recessive coding variation to developmental disorders. Science. 2018;362:1161–1164. doi: 10.1126/science.aar6731. [DOI] [PMC free article] [PubMed] [Google Scholar]; Martin, H.C., Jones, W.D., McIntyre, R., Sanchez-Andrade, G., Sanderson, M., Stephenson, J.D., Jones, C.P., Handsaker, J., Gallone, G., and Bruntraeger, M. (2018). Quantifying the contribution of recessive coding variation to developmental disorders. Science, 362. 1161-1164. [DOI] [PMC free article] [PubMed]
- 48.Maddirevula S., AlZahrani F., Anazi S., Almureikhi M., Ben-Omran T., Abdel-Salam G.M.H., Hashem M., Ibrahim N., Abdulwahab F.M., Meriki N. GWAS signals revisited using human knockouts. Genet. Med. 2018;20:64–68. doi: 10.1038/gim.2017.78. [DOI] [PubMed] [Google Scholar]; Maddirevula, S., AlZahrani, F., Anazi, S., Almureikhi, M., Ben-Omran, T., Abdel-Salam, G.M.H., Hashem, M., Ibrahim, N., Abdulwahab, F.M., Meriki, N., et al. (2018). GWAS signals revisited using human knockouts. Genet. Med. 20, 64-68. [DOI] [PubMed]
- 49.Doche M.E., Bochukova E.G., Su H.-W., Pearce L.R., Keogh J.M., Henning E., Cline J.M., Saeed S., Dale A., Cheetham T. Human SH2B1 mutations are associated with maladaptive behaviors and obesity. J. Clin. Invest. 2012;122:4732–4736. doi: 10.1172/JCI62696. [DOI] [PMC free article] [PubMed] [Google Scholar]; Doche, M.E., Bochukova, E.G., Su, H.-W., Pearce, L.R., Keogh, J.M., Henning, E., Cline, J.M., Saeed, S., Dale, A., Cheetham, T., et al. (2012). Human SH2B1 mutations are associated with maladaptive behaviors and obesity. J. Clin. Invest. 122, 4732-4736. [DOI] [PMC free article] [PubMed]
- 50.Willer C.J., Speliotes E.K., Loos R.J., Li S., Lindgren C.M., Heid I.M., Berndt S.I., Elliott A.L., Jackson A.U., Lamina C., Wellcome Trust Case Control Consortium. Genetic Investigation of ANthropometric Traits Consortium Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 2009;41:25–34. doi: 10.1038/ng.287. [DOI] [PMC free article] [PubMed] [Google Scholar]; Willer, C.J., Speliotes, E.K., Loos, R.J., Li, S., Lindgren, C.M., Heid, I.M., Berndt, S.I., Elliott, A.L., Jackson, A.U., Lamina, C., et al.; Wellcome Trust Case Control Consortium; Genetic Investigation of ANthropometric Traits Consortium (2009). Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 41, 25-34. [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S2. List of Previously Reported Candidate Genes whose Candidacy Is Further Supported by Findings in this Cohort. Table S8. List of Couples Who Presented for Counseling with No Available DNA From the Affected Deceased Child(ren) or Fetus(es). Table S10. List of Individuals for Whom Flash WES Protocol was Implemented.