Abstract
Background
Intercellular adhesion and biofilm production by Staphylococcus aureus makes these bacteria resistant to antimicrobial therapy. Here, Methicillin-resistant Staphylococcus aureus (MRSA) strains were characterized and the prevalence of genes encoding adhesion factors and biofilm formation was determined.
Results
All 248 MRSA isolates identified by cefoxitin disc diffusion were positive for the mecA gene. SCCmec-IV was the most frequently detected genotype (92.7%) and SCCmec-IVa was also very prevalent (84.3%). The quantitative microtiter plate assay showed that all the isolates were able to produce biofilm with levels ranging from high (21%) to moderate (46.4%) to low (32.7%).
All the strains possessed the icaD/icaA genes and produced biofilm (P < 0.05). None of the isolates possessed the bap gene. Furthermore, 94.8% of the isolates were positive for eno, 80.2% for clfA and for clfB, 78.2% for fnbA, 76.2% for ebps, 62.2% for fib, 39.9% for cna and 29.0% for fnbB. Also, nearly 69.8% of the isolates were positive for the gene sarA. All four agr groups were present: agr group 1 was predominant with 39.5%; agr group 3. agr group 2 and 3 strains carried more toxin-producing genes, and frequently produced more toxin. Sixty-six (26.6%) of the strains were multidrug resistant. All were vancomycin sensitive. Agr group I is more resistant to ciprofloxacin and gentamicin while agr group III is more resistant to erythromycin. Maximum sensitivity was to gentamicin and SXT, and they could be considered drugs of choice for controlling MRSA mediated infections in this region.
Conclusions
Biofilm development in MRSA might be an ica dependent and one needs to investigate the involvement of other global regulators, agr and sarA, and their contribution to the biofilm phenotype, as the high rate of biofilm production among the studied strains of S. aureus.
Keywords: MRSA, Antibiotics, SCCmec, Biofilm, Agr, sarA
Background
Methicillin-resistant Staphylococcus aureus (MRSA) is a serious risk to hospitalized patients worldwide and characterized by its resistance to antimicrobial treatment, and more recently to vancomycin, the drug of last resort for many strains of MRSA. In addition to its bacterial antibiotic resistance, its ability to produce biofilm, a dynamic structurally complex multilayered cellular matrix is another important complicating factor. Understanding the molecular pathogenesis of S. aureus could assist in developing novel prevention and treatment strategies. Biofilm synthesis is necessary for the survival and persistence of MRSA in its hosts and is considered to be a major virulence factor [1] and one of many, including extracellular toxins and surface structures that are effective in the induction and continuance of infection in the host [2]. Biofilm production is important during infection, providing defense against several opposing mechanisms of host and protects the microorganisms from antimicrobial agents [3]. The ability to form biofilm is a trait associated with bacterial virulence and many chronic bacterial infections [4]. Several genes are involved in the manufacture and maintenance of biofilms by staphylococci, of which the most extensively studied are the icaA and icaD (intercellular adhesion A and B) genes responsible for the synthesis of polysaccharide intercellular adhesion (PIA) that includes N-acetylglucosamine as a main component of the exopolysaccharide matrix surrounding the bacterial cells within the biofilm [5–7]. The protein components of the microbial surface recognizing adhesive matrix molecules have a high ability to interact with the host extracellular matrix proteins like collagen-binding protein (cna), fibrinogen binding protein (fib), elastin binding protein (ebpS), laminin binding protein (eno), fibronectin binding proteins A and B (fnbA and fnbB) and clumping factors A and B (clfA and clfB) [8].
Several virulence determinants of S. aureus are under the control of two genetic loci namely sarA (staphylococcal accessory regulator) and the agr quorum-sensing system. SarA might impact methicillin-resistant Staphylococcus aureus (MRSA) persistence in such infections by up regulating the expression of many virulence factors including biofilm formation to facilitate evasion of the host immune system in late phases of growth. Inhibiting the production of sarA protein might influence the down regulation of biofilm and virulence factors [9].
Suppression of the agr quorum-sensing system is required for biofilm formation. Its recurrence in established biofilms either through addition of auto-inducing peptides (AIPs) or glucose depletion triggers biofilm detachment [10–12]. Bacteria of S. aureus fall into four polymorphic agr types (agr I, agr II, agr III, and agr IV) based on the specificity of the auto- inducing peptides (AIPs) with regard to the signal receptor agr C.
There are no data on either the virulence factors of the microbial surface components recognizing adhesive matrix molecules (MSCRAMM) family or factors responsible for biofilm formation in methicillin resistance S. aureus in Palestine. This study focused on exposing the genes encoding adhesion factors and biofilm-forming capacity, and those governing antibiotic resistances in strains of MRSA isolated from Palestinian patients. It also evaluated the correlation between biofilm production and the presence of icaD, SarA and agr genes in the clinical isolates.
Results
Characterization of MRSA strains and antibiotic susceptibility
From 2015 to 2018, 248 strains of MRSA: 78 (31.5%) from infected wounds; 34 (13.7%) from blood culture; 25 (10.1%) from nasal secretions; 23 (9.3%) from urine; 88 of various other origins were collected from major hospitals in the West Bank-Palestine.
By the cefoxitin disc-diffusion Resistance test (≤22 mm), the 248 bacterial isolates were identified as being phenotypically MRSA and confirmed them as such by targeting the femA and mecA genes, which, respectively, separate susceptible S. aureus from MRSA. All of the isolates tested positive for the mecA gene by a PCR assay.
The susceptibility patterns of methicillin resistant isolates to the other antimicrobials are presented in Table 1. The cefoxitin disc-diffusion test showed that all 248 isolates were resistant to methicillin and none were resistant to vancomycin. However, sensitivity was high to varying degrees to SXT, gentamicin, clindamycin, ciprofloxacin, and erythromycin that were, respectively, 77.8, 76.6, 61.7, 55.6 and 34.3% (Table 1).
Table 1.
Frequency of antibiotic resistance of MRSA strains and biofilm
| Antibiotics | Weak biofilm producer (0.07 < ODs ≤ 0.14) | Moderate biofilm producer (0.14 < ODs ≤ 0.28) | Strong biofilm producer (0.28 < ODs) | Total (N = 248) | ||||
|---|---|---|---|---|---|---|---|---|
| S n. (%) | R n. (%) | S n. (%) | R n. (%) | S n. (%) | R n. (%) | S n. (%) | R n. (%) | |
| Cefoxitin | 0 (0%) | 81 (32.7) | 0 (0) | 115 (46.3) | 0 (0) | 52 (20.9) | 0 (0) | 248 (100) |
| Penicillin | 0 (0%) | 81 (32.6) | 0 (0) | 115 (46.3) | 0 (0) | 52 (20.9) | 0 (0) | 248 (100) |
| Ciprofloxacin | 49 (19.8%) | 32 (12.9) | 69 (27.8) | 45 (18.1) | 19 (7.6) | 32 (12.9) | 137 (55.2) | 109 (43.9) |
| Clindamycin | 50 (20.2%) | 31 (12.5) | 75 (30.2) | 40 (16.1) | 28 (11.2) | 24 (9.7) | 153 (61.6) | 95 (38.3) |
| Erythromycin | 29 (11.7%) | 52 (20.9) | 36 (14.5) | 79 (31.8) | 20 (8.0) | 32 (12.9) | 85 (34.2) | 163 (65.7) |
| Gentamycin | 69 (27.8%) | 12 (4.8) | 93 (37.5) | 22 (8.8) | 28 (11.2) | 24 (9.6) | 190 (76.6) | 58 (23.3) |
| SXT | 65 (26.2%) | 16 (6.4) | 92 (37.1) | 23 (9.2) | 36 (14.5) | 16 (6.4) | 193 (77.8) | 55 (22.1) |
| Vancomycin | 81 (32.7%) | 0 (0) | 115 (46.3) | 0 (0) | 52 (20.9) | 0 (0) | 248 (100) | 0 (0) |
Aside from cefoxitin and B-lactams antibiotics (penicillin G, amoxicillin/clavulanic acid, ceftriaxone and meropenem), the highest antibiotic resistance rates among the isolates of MRSA were for erythromycin (65.7%), ciprofloxacin (44.4%) and clindamycin (38.3%), followed by gentamycin (23.4%) and SXT (22.2%). All the isolates were susceptible to vancomycin (100%). Only, 66 (26.6%) isolates were MDR and, of 52 strongly positive biofilm-producing isolates, 20 (38.5%) were MDR and 32 (61.5%) were non-MDR.
Three different SCCmec types were detected among the isolates of MRSA that could be typed. Most of them carried SCCmec type IV (230/248, 92.7%), followed by SCCmec type I (11/248, 4.4%) and SCCmec type V (1/248, 0.4%). Also, 2.4% of the isolates could not be typed by multiplex PCR. None of the isolates carried SCCmec type II or III. The majority of the strains carrying SCCmec type IV carried SCCmec-type IVa (84.3%), followed by type IVc (4.8%), type IVd (1.2%) and type IVb (0.4%), and three patients provided two strains of type IVa/IVc (1.2%). Isolates were classified as CA-MRSA when they possessed SCCmec IV, as one of the subtypes IVa, IVb, IVc, IVd, or SCCmec type V and 231 CA-MRSA strains were found among all the isolates. Out of 12 MRSA strains with SCCmec IVc, 9 and 7 isolates were resistant to erythromycin and clindamycin with weak biofilm producers. All but one of the SCCmec IVc were sensitive to gentamycin, ciprofloxacin and SXT.
PCR screening of biofilm associated genes
Of the 248 strains of MRSA studied, 207 (83.5%) possessed the icaD gene and in 41 (16.5%) it was undetected. This percentage of icaD-negative strains was surprisingly high so detection of the icaA gene was undertaken which showed that all 41 icaD-negative strains were icaA-positive. High prevalence of icaA and icaD genes has shown a relationship to phenotypic biofilm formation.
None of the strains possessed the bap gene. The prevalence of sarA, eno, clfA, clfB, fnbA, ebps, fib, cna, and fnbB genes were 69.8, 94.8, 80.2, 80.2, 78.2, 76.2 62.2, 39.9 and 29.0%, respectively (Table 2). The frequency of clfA/B and fnbB genes of the agr group I were high at 92.9 and 52.0%, respectively. The frequency of eno, fnbA, epbS, fib and cna genes of the agr group III were also high at 97.6, 91.5, 84.1, 80.5 and 53.7%, respectively (Table 2).
Table 2.
The presence of biofilm-related genes for each agr group
| Agr group (%) | ||||||
|---|---|---|---|---|---|---|
| Biofilm Plate Titer (N = 248) | AgrI | AgrII | AgrIII | AgrIV | NT | SarA gene |
| Weak biofilm producer (n = 81, 32.7%) | 30.6 | 17.4 | 43.9 | 66.7 | 13.9 | 77.7 |
| Modertae biofilm producer (n = 115, 46.3%) | 42.9 | 56.5 | 50.0 | 22.2 | 47.2 | 73.9 |
| Strong biofilm producer (n = 52, 20.9%) | 26.5 | 26.1 | 6.1 | 11.1 | 38.9 | 48.1 |
| Total (n, %) | (98, 39.5%) | (23, 9.3%) | (82, 33.1%) | (9, 3.6%) | (36, 14.5%) | 173 |
| SarA gene (173, 69.7%) | 80.6 | 65.2 | 80.5 | 77.7 | 16.6 | |
| Adhesion Genes | ||||||
| icaD/ icaA(n = 248) | 87.7 | 87.0 | 78.0 | 88.9 | 80.6 | 58.5 |
| Eno (n = 235, 94.8%) | 92.9 | 91.3 | 97.6 | 100.0 | 94.4 | 71.5 |
| ClfA (n = 199, 80.2%) | 92.9 | 87.0 | 89.0 | 55.6 | 27.8 | 80.4 |
| ClfB (n = 199, 80.2%) | 92.9 | 87.0 | 89.0 | 55.6 | 27.8 | 80.4 |
| FnbA (n = 194, 78.2%) | 87.8 | 78.3 | 91.5 | 66.7 | 25.0 | 81.4 |
| EbpS (n = 191, 76.2%) | 82.7 | 60.9 | 84.1 | 55.6 | 55.6 | 76.4 |
| Fib (n = 154, 62.2%) | 60.2 | 73.9 | 80.5 | 66.7 | 16.7 | 83.1 |
| Cna (n = 99, 39.9%) | 46.9 | 21.7 | 53.7 | 0.0 | 11.1 | 78.8 |
| FnbB (n = 72, 29.0%) | 52.0 | 21.7 | 13.4 | 11.1 | 11.1 | 81.9 |
| Antibiotic Resistance | ||||||
| Cefoxitin/Penicillin (n = 248) | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100 |
| Erythromycin (n = 163) | 59.2 | 60.9 | 69.5 | 33.3 | 86.1 | 63.2 |
| Ciprofloxacin (n = 109) | 58.2 | 56.5 | 20.7 | 33.3 | 52.8 | 63.3 |
| Clindamycin (n = 95) | 28.6 | 43.5 | 48.8 | 11.1 | 44.4 | 65.3 |
| Gentamycin (n = 58) | 27.6 | 21.7 | 11.0 | 22.2 | 41.7 | 55.2 |
| SXT (n = 55) | 19.4 | 21.7 | 11.0 | 11.1 | 58.3 | 41.8 |
| MDR (n = 66) | 22.4 | 43.5 | 13.4 | 11.1 | 61.1 | 46.9 |
| Source of isolate | ||||||
| Wound (n = 78) | 36.7 | 34.8 | 35.4 | 22.2 | 8.3 | 80.7 |
| Blood (n = 34) | 11.2 | 17.4 | 11.0 | 11.1 | 25.0 | 52.9 |
| Nasal (n = 25) | 11.2 | 13.0 | 8.5 | 0.0 | 11.1 | 64 |
| Pus (n = 23) | 8.2 | 8.7 | 13.4 | 22.2 | 0.0 | 95.6 |
| Urine (n = 23) | 1.0 | 0.0 | 11.0 | 22.2 | 30.6 | 43.5 |
| Tissue (n = 13) | 9.2 | 8.7 | 2.4 | 0.0 | 0.0 | 61.5 |
| Sputum (n = 12) | 3.1 | 8.7 | 6.1 | 0.0 | 5.6 | 58.3 |
| Other sites (n = 40) | 19.4 | 8.7 | 12.2 | 22.2 | 19.4 | 72.5 |
The coexistence of the studied virulence genes was investigated in the 248 clinical isolates of MRSA, only nine of which had all the genes investigated. Five had only the eno gene. Despite these low prevalence rates, the other 234 strains possessed at least one other virulence gene.
Of the clinical isolates in which most virulence genes were investigated for coexistence, 56 were positive for all the genes but one, either eno (n = 29) or fnbB (n = 27) gene. Of 42 clinical isolates, 18 strains were negative for two genes, fnbB and fib genes and 24 were negative for finbB and cna genes.
Determination of biofilm production by the microtiter plate method
All the strains of MRSA produced biofilm. In the microtiter plate method for determining this and using the mean OD570 of the negative control (0.07), values between 0.07 and 0.140 (2 × the negative control value of 0.07), were considered to be strains that were weak producers, which accounted for 81 (32.7%) strains, values between 0.140 and here 0.280 (4 × the negative control value of 0.07) to be moderate producers, which accounted for 115 (46.4%) strains and values higher than 0.280 were strong producers, which accounted for 52 (21.0%) strains (Table 3).
Table 3.
Biofilm-forming capacity of 248 methicillin-resistant strains of Staphylococcus aureus (MRSA) and percentages of their adhesion genes related to antibiotics
| Biofilm | Adhesion gene | Gentamycin % | Ciprofloaxin % | SXT % | Erythromycin % | Clindamycin % | MDR % | Total % |
|---|---|---|---|---|---|---|---|---|
| Strong (n = 52, 20.9%) | FinbB | 23.1 | 23.1 | 1.9 | 15.4 | 11.5 | 7.7 | 46.2 |
| ClfA/ClfB | 32.7 | 40.4 | 11.5 | 36.5 | 28.8 | 19.2 | 69.2 | |
| Ebps | 42.3 | 46.2 | 21.2 | 44.2 | 34.6 | 32.7 | 71.2 | |
| Can | 15.4 | 19.2 | 3.8 | 15.4 | 11.5 | 7.7 | 38.5 | |
| Eno | 46.2 | 59.6 | 30.8 | 57.7 | 25.0 | 38.5 | 94.2 | |
| Icad | 42.3 | 57.7 | 26.9 | 51.9 | 36.5 | 32.7 | 88.5 | |
| FinbA | 34.6 | 44.2 | 13.5 | 38.5 | 26.9 | 21.2 | 71.2 | |
| Fib | 26.9 | 34.6 | 5.8 | 28.8 | 21.2 | 17.3 | 46.2 | |
| Moderate (n = 115, 46.3%) | FinbB | 5.2 | 18.3 | 0.9 | 17.4 | 6.1 | 6.1 | 26.1 |
| ClfA/ClfB | 13.0 | 30.4 | 13.0 | 54.8 | 28.7 | 17.4 | 81.7 | |
| Ebps | 12.2 | 31.3 | 13.9 | 53.0 | 27.0 | 18.3 | 79.1 | |
| Can | 7.0 | 7.8 | 8.7 | 28.7 | 17.4 | 8.7 | 40.9 | |
| Eno | 18.3 | 36.5 | 19.1 | 65.2 | 33.0 | 26.1 | 94.8 | |
| Icad | 13.9 | 32.2 | 17.4 | 56.5 | 29.6 | 21.7 | 82.6 | |
| FinbA | 9.6 | 28.7 | 9.6 | 50.4 | 24.3 | 12.2 | 78.3 | |
| Fib | 7.8 | 26.1 | 7.0 | 43.5 | 22.6 | 10.4 | 64.3 | |
| Weak (n = 81, 32.6%) | FinbB | 3.7 | 14.8 | 7.4 | 24.7 | 8.6 | 6.2 | 88.9 |
| ClfA/ClfB | 13.6 | 34.6 | 66.7 | 51.9 | 33.3 | 17.3 | 82.7 | |
| CLFB | 13.6 | 34.6 | 17.3 | 54.3 | 33.3 | 17.3 | 86.4 | |
| Ebps | 11.1 | 30.9 | 14.8 | 46.9 | 30.9 | 14.8 | 75.3 | |
| Can | 7.4 | 11.1 | 7.4 | 23.5 | 17.3 | 7.4 | 39.5 | |
| Eno | 12.3 | 37.0 | 17.3 | 59.3 | 37.0 | 17.3 | 95.1 | |
| Icad | 12.3 | 30.9 | 14.8 | 51.9 | 33.3 | 18.5 | 81.5 | |
| FinbA | 13.6 | 30.9 | 14.8 | 51.9 | 33.3 | 17.3 | 82.7 | |
| Fib | 9.9 | 29.6 | 8.6 | 45.7 | 28.4 | 11.1 | 69.1 |
Among the 65.7 and 43.9% of high resistance of the strains of MRSA to erythromycin and ciprofloxacin, 31.8 and 18.1% were moderate biofilm producers., respectively.
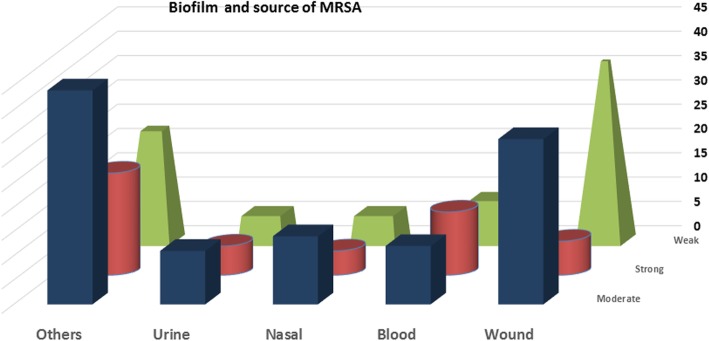
Among the 78 clinical isolates of S. aureus from wounds, 37 (47.4%) were weak producers of biofilm, 34 (43.6%) were moderate producers and 7 (9%) were strong producers. Of the 34 isolates from blood, 9 (26.5%) were weak producers, 12 (35.3%) were moderate producers and 13 (38.2%) were strong producers. Of the 23 isolates from urine, 6 (26.1%) were weak producers, eleven (5.5%) were moderate producers, and 6 (26.1%) were strong producers. Of the 25 nasal isolates, 6 (24%) were weak producers, 14 (56%) were moderate producers and 5 (20%) were strong producers. Among the 88 isolates obtained from other different clinical samples, 23 (26.1%) were weak producers, 44 (50%) were moderate producers and 21 (23.9%) were strong producers, of which half of the sputum samples were strong producers (Fig. 1).
Fig. 1.
Biofilm phenotype and the source of strains of MRSA, i. e. tissues and lesions
All the virulence genes were found in weak, moderate and strong producers of biofilm. The least frequent genes were the cna (39.9%) and fnbB (29%) genes, the percentages of which were, respectively, 39.5 and 29.6% for weak biofilm producers, 40.8 and 20.1% for moderate biofilm producers, and 38.4 and 34.6% for strong biofilm producers (Table 3).
The agr groups
Table 2 gives the agr groups of the strains of MRSA. The 248 strains separated into four agr groups with 98 (39.5%) belonging to agr-I, making it the predominant type, 23 (9.3%) belonging to agr-II, 82 (33.1%) belonging to agr-II, 9 (3.6%) belonging to agr-IV and 36 (14.5%) were negative regarding the agr PCR. There was no relationship between agr specific groups and the genes encoding MSCRAMM. Strains that belonged to the agr-I group showed higher antibiotic resistance to ciprofloxacin and gentamycin, compared to the other three agr groups. Strains that belonged to agr-III group had higher erythromycin (69.5%) and clindamycin (48.8%) resistance compared to the other agr groups. Of the 66 MDR strains, 20 belonged to the agr-I group and only 10, 11 and one belonged to the agr-II, agr-III and agr-IV groups, respectively (Table 2).
Detection of the SarA gene
The sarA gene was found in 173 (69.8%) strains of MRSA. Regarding phenotypic biofilm formation, 63 of 81 (77.7%) were weak producers of biofilm, 85 of 115 (73.9%) strains were moderate producers and 25 of 52 (48.1%) were strong producers with a significant difference (P < 0.05). There was a high prevalence of fib (83.4%), clfB and clfB (80.4% each) and fnbB (81.9%) and fnbA (81.4%). Of the strains of MRSA carrying the sarA gene, 46.9% were MDR, and 63.2, 63.3 and 65.3%, 55.2 and 41.8% were resistant to erythromycin, ciprofloxacin, clindamycin, gentamycin and SXT., respectively (Tables 2, 3).
Discussion
Biofilm production by S. aureus has been identified as the most important means of defense against host antagonistic responses. Beside enabling bacterial colonization of host tissues, it also prevents clearance of the bacteria by antimicrobial agents and host immune responses [13], leading to morbidity and mortality owing to the metastatic spread of abscesses [14]. Here the ability among strains of MRSA isolated from hospitalized- and out-patients to form biofilm was studied, combining it with their clinical molecular biological details and determining the presence of genes encoding these virulence factors and its relation to antibiotics. The SCCmec type IV was the most frequent SCCmec type among the strains. Its presence in the sporadic strains among the 92.7% and the group of strains from out-patients shows their great persistence [15]. SCCmec type IV is currently one of the most frequent nosocomial SCCmec types and is found in several countries [16, 17].
The antimicrobial resistance patterns of strains of this type varied considerably.
Here, 26.6% of the strains of MRSA with multi-resistance to more than three antibiotics were of the SCCmec type IV with 83.4% carried SCCmec type IVa and all are biofilm producers. These results indicate that the production of biofilm might be one of the crucial factors increasing resistance to commonly used antibiotics. That deserve a special comment. This higher MDR rely to protective nature of the biofilm, the bacteria growing in it are internally resistant to many antibiotics and the antibiotic resistance in the strains of the bacteria residing in biofilm could increase up to 1000 times as seen by Neupane and colleagues [18]. The main reasons for this may be difficulty in penetration of biofilm by antibiotics, slow growth rate of the bacteria and presence of antibiotic degradation mechanisms.
Moreover, the high resistance of the strains of MRSA to erythromycin, ciprofloxacin,of which were moderate biofilm producers with higher rate of studied adhesion genes, particularly, eno gene. This agreed with the high prevalence of drug resistance presented in a study done in Iran where the resistance of strains of MRSA to ciprofloxacin, erythromycin and gentamicin was 51.28, 87.18 and 71.8%, respectively [19]. It seems that misuse and overuse of some antibiotics, including gentamicin, clindamycin, ciprofloxacin and erythromycin, have caused a high prevalence of resistance to them in this region, showing that empirical treatment of infections of strains of MRSA at Palestinian hospitals with these antibiotics may not be effective and they should not be used and considered first-line drugs for the treatment of infections of MRSA in the local population. Appropriate measures are needed to prevent treatment failures. All the strains were susceptible to vancomycin and more than two third of the strains were susceptible to trimethoprim sulfamethoxazole. Vancomycin is reported to be the most effective antibiotic for Gram-positive bacteria, including MRSA, but reduced susceptibility to both antibiotics has been reported in some studies [20, 21]. Vancomycin and other glycopeptides have remained the last options for eradication of infections caused by S. aureus. The data presented here also showed all the strains, producing biofilm, were sensitive to vancomycin. This is consistent with other researchers’ recommendation that vancomycin, which is a very expensive drug, is the last antibiotic option and should be used sparingly.
The data presented here agree with susceptibility rates in other countries [22]. Wang and colleagues [23] reported a 78.6% susceptibility rate to trimethoprim-sulfamethoxazole among strains of MRSA, which is of concern and emphasizes the need for persistent monitoring of the development of antimicrobial resistance by strains of S. aureus that leads to community- and hospital-acquired infections. Here, we report a high rate of SXT resistance (22.1%), which in the future could increase as a consequence of horizontal transmissibility of the dfrK gene, encoding for trimethoprim-resistance.
Here, phenotypic and genotypic evaluations, PCR and crystal violet (CV) staining assays, were combined to detect biofilm production in strains of S. aureus.. All the strains were biofilm producers with variation in biofilm mass. To understand the molecular mechanism of biofilm production by strains of MRSA, in addition to the icaD/icaA, sarA and agr group genes, the frequency of nine selected genes involved in biofilm production were detected. Many studies have shown the role and requirement of the intracellular adhesion locus (ica) in biofilm production [24, 25]. The icaA and icaD genes determine the ability of strains of S. aureus to produce biofilm by mediating the synthesis of PIA which suggest that the ica locus would be a good target in the therapy of implant infections. There was a 100% agreement between the genotypes and phenotypes of strains where all the strains having icaD/icaA and producing biofilm, which agreed with the findings of Liberto and colleagues [26] and support those of Namvar and colleagues [27], who reported that strains of S. aureus had no ability to produce biofilm, unless they were positive for the icaD gene. Similar observations were reported by Grinholc and colleagues [28], who found that 91% of strains of MRSA possessed the icaD gene. Contrarily, Arciola and colleagues [29], detected icaA and icaD genes in only 61% of strains. The relatively low percentage of icaD positive strains described by Arciola and colleagues [29] resulted from the method of detection they used, in which primers complementary to the sequence of the icaD gene from Staphylococcus epidermidis, rather than primers complementary to the sequences of the icaD and icaA genes from S. aureus were used. There was no difference in the distribution of the ica genes in strongly and weakly virulent strains, which agreed with the findings of others [5, 30, 31]. The PIA mediates intercellular adherence and accumulation of multilayer biofilms. In our study the ica operon was present in all MRSA strains but strains differed in biofilm mass. It is suggested that these strains also used other systems to form biofilm such as protein A (SpA) or fibronectin binding proteins.
Other contradictory published data stated that some strains, in spite of the presence of the ica locus, do not produce biofilm [25]. Recently, it has become evident that the presence of PIA is not essential for the production of biofilm in many strains of MRSA [32].
The ability to produce biofilm varied among the strains of MRSA and also greatly among the other different genotypes of S. aureus where an increasing number of different adhesion molecules have been found. The frequency of eno, clfA/clfB, fnbA, ebps, fib, cna, and fnbB genes was found to be 94.8, 80.2, 78.2, 76.2, 62.2, 39.9 and 29.0%, respectively (Table 2). While in other studies [33], showed the frequency of eno, clfA/clfB, fnbA, ebps, fib, cna and fnbB genes in strains of MRSA was 79, 97, 64, 12, 76, 56, and 51%., respectively. Yang and colleagues [34], showed the prevalence of genes associated with biofilm in the ST59-SCCmec IV-t437 clone were icaA (100.0%), icaD (97.3%), fnbpA (100.0%), fnbpB (0), clfA (100%), clfB (100%), cna (2.7%), bbp (0), ebpS (88.5%). This explains the discrepancies between studies, which is related to differences in the frequency of clones among different countries. The gene bap, whose protein was, probably, the first protein shown to have a role in biofilm production in S. aureus, was not tracked in our study. It is said to be absent in all strains, which agrees with the study by Serray and colleagues [35]. However, the absence of bap indicates that the ica-dependent mechanism, might be primarily responsible for adhesion and biofilm production in strains as suggested by Vautor and colleagues [36].
The fnbA and fnbB genes appear to be essential to the invasion and adhesion of bacteria and might be correlated with their biofilm-producing ability. In this study, a low percentage (29.0%) of occurrence of the fnbB gene was observed.
However, Arciola and colleagues [29], found a high occurrence of this gene (99.5%). This could partly be ascribed to the different region of the locus analyzed by the couple of primers. However, the fnbA gene was detected in 76.2% of the strains, which is similar to what was observed by Ikawaty and colleagues [28]. There is a significant difference between strains from blood and those from wounds regarding the presence of the fnbB gene. About a third of the strains (35.9%) from wounds carried the gene. However, only 14.7% of the strains from blood carried the fnbB gene. A comparative analysis between strains of MRSA and strains of MSSA showed that the fnbpA gene was more likely to be present in strains of MRSA whereas the fnbB gene was more likely to be present in strains of MSSA [34]. However, other studies did not find a correlation between methicillin resistance and the prevalence of genes associated with biofilm [37]. This disagreement may be owing to the specific clonal complexes of strains that might contain an exclusive combination of surface-associated and regulatory genes [38].
This study indicated that the clfA and the clfB genes were present in 82.7% of the strains and constituted the bound coagulase of S. aureus. This study showed that the strains from all sources except urine (43.5%) had a high percentage of, both, the clfA and the clfB genes.
Elastin is the main component of elastic fibers, which are proteins that provide strength and flexibility to connective tissue and is highly expressed in the lung, skin and blood vessels and widely expressed at low levels in most mammalian tissues [39]. Elastin binding protein of S. aureus (EbpS), said to facilitate binding of bacteria to elastin rich host, extracellular cell matrix (ECM) [40]. EbpS is a cell-surface molecule the mediates binding of a bacterial cell to soluble elastin peptides and tropoelastin [39]. The presence of the ebpS gene was found in 76.2% of the strains studied here. Another gene, shown to have a crucial role in binding to extracellular matrix, fibrinogen (fib), was also detected in 62.2% of the strains. This agreed with the findings of Pereyra and colleagues [41] who reported higher percentages of 90 and 71.7%, respectively. This contradicted the findings of [35], where the fib and ebpS genes were detected at a rates of 5.66 and 9.34% of strains. The difference in the prevalence of these genes is probably owing to the distribution of variants of the genotype of S. aureus in different countries. The incidence of cna was 39.9% in the strains of MRSA studied here. This agreed with the findings of Nashev and colleagues in Italy [36], and in Bulgaria [29], who reported of similar rates of 46.7% and occurrence of this gene (11.32%) was reported by serray and colleagues [35].
The expression of several virulence factors of S. aureus was shown to be controlled by certain genetic loci, particularly, the staphylococcal accessory regulator, which consists of the sarA gene and accessory gene regulator (agr) locus (Jarraud et al., 2002). In studies done by other colleagues [42–44] all their strains of MRSA harboring the icaADBC genes were positive for the sarA gene, which was contradicted by this study.
The ica genes are regulated by multiple genes such as the sarA, and agr genes. They might interact with each other and regulate biofilm production. The sarA gene has an effect on many virulence genes of S. aureus and appears to be a master controller of biofilm production, promoting synthesis of fibronectin and fibrinogen binding proteins and also toxins for tissue spread while repressing expression of protein A and the four major extracellular proteases governed by the SspA, SspB, Aur, and ScpA genes [2, 45]. Here, about two third of the strains genotypically possessed the sarA gene and phenotypically produced biofilm and the fnbB gene was common among strains that were strong producers of biofilm (34.6%) suggesting the importance of PIA-independent biofilm production in these strains. Interestingly in this study, a higher rate of the strains of MRSA possessed the fnbA, fnbB and the fib gene which were also positive for the sarA gene.
Different levels of sarA expression in clinical isolates of S. aureus have been related to differences in extracellular protease production [46] and that sarA can directly and positively regulate levels of fnbA transcription [47]. On the other hand, Pozzi and colleagues [48] reported that biofilm production in strains of MSSA mainly occurs via PIA synthesis while in strains of MRSA it is related more to adhesion owing to the fnbB gene. So further investigation and studies are needed.
Strains producing biofilm have a very high tendency to exhibit antimicrobial multidrug resistance. However, 46.9% of the strains with the sarA gene were MDR, and more than halfwere shown to be resistant to erythromycin, ciprofloxacin, clindamycin and gentamycin This makes sarA an attractive target for antimicrobial drug development [49, 50]. Surprisingly, most of the isolates from wounds and pus were sarA positive.
To date, strains of S. aureus have been classified into four main groups, agr -I to agr-IV, according to differences in their agr genes, (Jarraud et al., 2002). The central role of the agr-encoded quorum-sensing system in the regulation of virulence makes it an attractive target for antimicrobial drug development. However, mutations in the agr gene or interference with agr gene activity by a cross-inhibiting agr pheromone can promote the production of colonization factors like MSCRAMMs and biofilm development [6]. All four agr groups were found among the strains studied here, with agr group I in large proportion and more than half of the samples from wounds belonged to this group.
Previous studies also found the agr group I to be the predominant type [51]. Here, 14.5%strains could not be typed by the same method, possibly, owing to a deletion in the agr locus. It is noteworthy that the strains in agr group III had a greater number of the fnbA, ebps, cna, eno and fib genes, and most of the toxin producing strains also belonged to agr group III while the strains in agr group-I had a greater number of fnbB, clfA and clfB. Regarding the relationship between agr group-III and biofilm production, the data revealed that strains belonging to agr group-III had a greater number of weak and moderate producers of biofilm compared to those belonging to agr group-I, which, interestingly, had more and stronger biofilm-producing strains. Also, the strains belonging to agr group-III had higher antibiotic resistance to erythromycin and clindamycin compared to those belonging to agr group-I, which carried a greater number of strains resistant to ciprofloxacin (58.2%), gentamycin (27.6%) and SXT (19.4%). The presence of the combination of genes studied here, where 3.9% of the strains possessed all the genes examined and including the icaA and icaD genes, could mean that they might have a selective advantage, e. g. a good genetic capacity for adherence and better colonization of hosts. Moreover, the coexistence of icaA, icaD, agr and sarA and eight MSCRAMM genes in 11.7%of the strains agree with the findings of Tristan and colleagues [52]. The most common biofilm gene combination among the strains of MRSA was that of agr, sarA, eno, clfA/clfB, fnbA, ebps and fib genes. The mechanism of multidrug resistance is said to result from close cell to cell contact in the biofilm that makes the transfer of plasmids containing MDR genes among them easier, which limits the therapeutic options, creating an economic and social burden to the healthcare system. Biofilm development is a very complicated process that involves numerous factors. The present survey study is a first step. It provides preliminary results for further detailed future studies. One limitation of the study, is the inability to use control S. aureus strains that lack each of the gene tested in this study. Mutants defective in either; IcaA, IcaD, agr, saR, each of the genes that code for the MSCRAAM proteins. This way, biofilm development by the mutant would be measured directly to that produced by the tested isolates.
Conclusion
The present study revealed that MRSA strains isolated from clinical materials from hospitalized patients produced biofilm and possessed icaA and icaD genes, with differed biofilm mass, indicating that these strains may also use other system to form biofilm. The high rate of biofilm production among the strains of S. aureus and high rate of drug resistance among the biofilm producing strains, detection of biofilm adhesion genes indicating staphylococcal virulence markers and showing that the burden of MRSA in the Palestinian West Bank region was high. Further, clinical strains of S. aureus and the ability of several strains of MRSA to produce biofilm in the absence of sarA and agr genes needs further investigation to clarify the mechanism underlying the production of biofilm independent of the activity of the sarA and agr genes. On the basis of the antimicrobial susceptibility testing, aworrisome increase in erythrocyte and ciprofloxacin resistance was observed, which deserves future attention.
Methods
Clinical strains
A Total of 248 strains of MRSA were isolated from patients admitted to four Palestinian hospitals located in, Jerusalem, Ramallah, Bethlehem and Nablus. The study period was between November 2015 and April 2018. Most came from the Al-Makassed Islamic Charitable Society Hospital in Jerusalem. MRSA ATCC 4300 and S. epidermidis ATCC 12228 were reference strains provided by Dr. Adham abu Taha of the Palestinian Al Najah University. All the strains were stored at − 80 °C in brain-heart infusion (BHI) (Himedia, Mumbai, India) plus 25% glycerol (EMPROVE, Darmstadt, Germany). This study was approved by the Research Ethical Committee of Al-Quds University. Written and informed consents were sent for the participating hospitals and clinics.
Identification of isolates
All isolates were identified by classic microbiological methods: colony morphology; mannitol fermentation; Gram staining, catalase test; coagulase test. Antibiotic susceptibility was determined by the disc-diffusion method (Oxoid, Basingstoke, UK).
The antibiotics used in this study were cefoxitin (30 μg), penicillin (10 U), amoxicillin/clavulanic acid (10 μg), ceftriaxone (30 μg), meropenem (10 μg), erythromycin (15 μg), ciprofloxacin (5 μg), clindamycin (10 μg), gentamicin (10 μg), SXT (25 μg) and vancomycin (30 μg). Apart from β-lactam, multi drug resistance (MDR) for MRSA was defined as resistance to at least three of the antimicrobial agents.
Isolates were classified as susceptible or resistant to methicillin according to the criteria of the Performance Standards for Antimicrobial Susceptibility Testing (2002). Methicillin resistant strains of S. aureus were detected by the disk-diffusion method, using a cefoxitin (FOX) disk (30 μg) on Mueller-Hinton agar plates according to the Clinical Laboratory Standards Institute (CLSI) guidelines [53]. An infection was considered healthcare-associated if the date of the infection occurred on or after the third day of admission to an inpatient facility.
Genomic DNA extraction
Genomic DNA was extracted from overnight fresh cultures on Trypticase Soy Broth (TSB), using either a ‘Nucleospin’ DNA extraction kit (Macherey-Nagel, Germany) [54] or a Presto Mini gDNA Bacteria Kit (Geneaid).
Molecular typing
Detection of the mecA gene and SCCmec typing by PCR
The mecA gene and femA endogenous control gene were amplified in the same reaction. The primers used to amplify the mecA gene were mecA1F (5′-GTAGAAATGACTGAACGTCCGATAA-3′) and mecA2R (5′-CCAATTCCACATTGTTTCGGTCTAA-3′) [16]. The primers used to amplify the femA gene were femA GFEMAR-1(5′-AAAAAAGCACATAACAAGCG-‘3) and femA GFEMAR-2 (5’-GATAAAGAAGAAACCAGCAG-‘3) [55]. Each reaction used 1 μM of each primer and 2 μl of DNA, and was performed in Thermo Scientific Reddy Mix PCR mater Mix conc 2X in a final volume of 25 μl. The thermal cycling program for detecting both genes was: one cycle of initial denaturation at 95 °C for 15 min; 34 cycles of denaturation at 95 °C for 30 s; annealing at 58 °C for 30 s; extension at 72 °C for 1 min; a final extension at 72 °C for 5 min.
The amplified products (femA: 132 bp and mecA: 310 bp) were resolved in a 2.5% agarose gel. The fragments were stained with ethidium bromide and visualized and photographed using a gel documentation system. A 100 bp ladder was run as a molecular weight marker. Isolates that were confirmed to be methicillin sensitive by the disk diffusion method and then by the absence of the mecA gene were excluded from this study.
Exposing the existence of SCCmec types and subtypes I, II, III, IVa, IVb, IVc, IVd, and V of all the isolates of MRSA was done by the multiplex PCR assay described by Boye and colleagues [56], which used 9 pairs of primers that are unique and specific for the above mentioned SCCmec types and subtypes. Exposing the existence of SCCmec subtype IV was done by the multiple PCR assay described by Zhang et al. [57]. Isolates unable to be typed were designated NT. Amplification was performed as described by Hadyeh and colleagues (2019).
Detection of biofilm genes
Simplex and multiplex PCRs were used to detect the following genes in all the isolates of MRSA: bap (encoding biofilm-associated protein); ebpS (encoding elastin-binding protein); eno (encoding laminin-binding protein); fib (encoding fibrinogen-binding protein); fnbA (encoding fibronectin-binding protein A); fnbB (encoding fibronectin-binding protein B); clfA and clfB (encoding clumping factors A and B); cna (encoding collagen-binding protein). The specific primers and PCR thermal profiles used for these genes were as described by others [3, 35, 52]. The amplified products cna: 423 bp; ebpS: 652 bp; eno: 302 bp; fnbA: 127 bp; fnbB: 524 bp; fib: 404 bp; bap: 971 bp; clfA: 292 bp and clfB: 205 bp were resolved in a 2.5% agarose gels.
Detection of icaD and icaA genes
The presence of icaD DNA was detected as described by Gowrishankar and colleagues [58]. The specific forward primer was icaD (5’ATG GTC AAG CCC AGA CAG AG3′) and the specific reverse primer was icaD (5’CGT GTT TTC AAC ATT TAA TGC AA3’). For icaD-negative strains, detection of the icaA gene was done using the forward primer icaAF (5’ACA CTT GCT GGC GCA GTC AA 3’) and reverse primer icaAR (5’TCTGGAACCAACATCCAACA3’) as proposed by [30]. The icaD and icaA genes were amplified by a PCR to generate 188 bp and 198 bp fragments, respectively.
Determination of agr group and sarA gene
The agr typing was done by a multiplex-PCR to determine the agr allele types I to IV, using the agr group specific primers and amplification conditions as described by [30]. The agr system groups were classified based on the hyper-variable domain of the agr locus and their responding receptors separated into four major agr groups. Pan-agr, corresponding to the conserved sequences of agrB, was used in all the reactions.
Based on the agr locus polymorphism, four reverse primers were used, each specific for the amplification of a single agr group. The agr groups were identified by amplicon size: 440 bp for agr I; 572 bp for agr II; 406 bp for agr III; 588 bp for agr IV.
SarA DNA was detected, using the forward primer sarAF (5’CCCAGAAATACA ATCACTGTG’3) and reverse primer sarAR (5′ AGTGCCATTAGTGCAAAACC’3) as described by Gowrishankar and colleagues [58], which produced an amplicon of 720 bp.
Biofilm formation assay
The isolates of MRSA were tested for biofilm formation. The assay was performed in polystyrene 96-well microtiter plates that had flat-bottomed wells that were stained with crystal violet according to Stepanovic and colleagues [59]. Staphylococcus epidermidis ATCC 12228, and MRSA ATCC 43300 were used as biofilm-producing controls. Trypticase soy broth medium was used as a negative control to determine background OD. The microtiter plate method was done as described by Atshan and colleagues [60]. The amount of biofilm formed was estimated by reading the optical density (OD) at 570 (630) nm and recording the absorbance using a microplate reader (RT-2100C, Rayto, IVD). The average OD value of each triplicate of experimental samples and negative controls was calculated. Biofilm formation was separated into four categories according to [19]: 1, ODs ≤ ODc = no biofilm produced, therefore a non-producer; 2, ODc ≤ ODs ≤ 2× ODc = weak biofilm produced, therefore a weak producer; 3, 2× ODc ≤ ODs ≤ 4 × ODc = moderate biofilm produced, therefore a moderate producer; 4, 4× ODc < ODs = strong biofilm produced, therefore a strong producer, where ODc = OD of the negative control and ODs = OD of the experimental samples.
Statistical analysis
Data analysis was done using SPSS software version 20.0 (IBM, Armonk, USA). Pearson’s chi-square was used in the statistical analysis. A P value less than 0.05 was considered statistically significant.
Acknowledgements
The authors thank all members of staff of the Department of Microbiology at the Al Makassed Hospital and Al Helal hospital, who helped isolate the strains used in this study. We also thank Ms. Etaf Hadyeh in sample preparation and Dr. Lionel Schnur for proofreading and revising our manuscript. This work was supported by MERC (M33-014).
Availability of data and material
All relevant materials and data supporting the findings of this study are.
contained within the manuscript. Raw data can be requested from the.
corresponding author.
Abbreviations
- BURP
Based upon Repeat Pattern
- CA-MRSA
Community-associated Methicillin Resistant Staphylococcus aureus
- clfA and clfB
Clumping factors A and B.
- ebpS
Elastin binding protein
- eno
Laminin binding protein
- fib
Fibrinogen binding protein
- fnbA and fnbB
Fibronectin binding proteins A and B
- MDR
Multi-drug resistant
- MRSA
Methicillin resistant Staphylococcus aureus
- PCR
Polymerase chain reaction
- PVL
Panton–Valentine leucocidin
- sarA
Staphylococcal accessory regulator
- Cna
Collagen-binding protein
- SCCmec
Staphylococcal chromosome cassette mec
Authors’ contributions
KA was involved in all aspects of research design, implementation, sample collection and writing the manuscript. WQ participated in the sample analysis. ZA contributed to the manuscript revision and overall support of this study. All authors read and approved the final manuscript.
Funding
This study was supported financially by grant MERC (M33–014). The funders had no role in the design of the study and collection, analysis, or interpretation of data or in writing the manuscript.
Ethics approval and consent to participate
The study was approved by the Research Ethical Committee at Al-Quds University. Written and informed consents were sent for the participating hospitals and clinics.
Consent for publication
Not applicable.
Competing interests
All authors declare that they have no conflict of interest.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Kifaya Azmi, Email: kifaya_alkam@yahoo.com, Email: ksuleiman@staff.alquds.edu.
Walaa Qrei, Email: walaa.walid@yahoo.com.
Ziad Abdeen, Email: Zabdeen13@gmail.com.
References
- 1.Lister JL, Horswill AR. Staphylococcus aureus biofilms: recent developments in biofilm dispersal. Front Cell Infect Microbiol. 2014;4:178. doi: 10.3389/fcimb.2014.00178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cheung AL, Bayer AS, Zhang G, Gresham H, Xiong YQ. Regulation of virulence determinants in vitro and in vivo in Staphylococcus aureus. FEMS Immunol Med Microbiol. 2004;40(1):1–9. doi: 10.1016/S0928-8244(03)00309-2. [DOI] [PubMed] [Google Scholar]
- 3.Vancraeynest D, Hermans K, Haesebrouck F. Genotypic and phenotypic screening of high and low virulence Staphylococcus aureus isolates from rabbits for biofilm formation and MSCRAMMs. Vet Microbiol. 2004;103(3–4):241–247. doi: 10.1016/j.vetmic.2004.09.002. [DOI] [PubMed] [Google Scholar]
- 4.Mohamed JA, Huang DB. Biofilm formation by enterococci. J Med Microbiol. 2007;56(Pt 12):1581–1588. doi: 10.1099/jmm.0.47331-0. [DOI] [PubMed] [Google Scholar]
- 5.Rohde H, Burandt EC, Siemssen N, Frommelt L, Burdelski C, Wurster S, et al. Polysaccharide intercellular adhesin or protein factors in biofilm accumulation of Staphylococcus epidermidis and Staphylococcus aureus isolated from prosthetic hip and knee joint infections. Biomaterials. 2007;28(9):1711–1720. doi: 10.1016/j.biomaterials.2006.11.046. [DOI] [PubMed] [Google Scholar]
- 6.Fitzpatrick F, Humphreys H, O'Gara JP. The genetics of staphylococcal biofilm formation--will a greater understanding of pathogenesis lead to better management of device-related infection? Clin Microbiol Infect. 2005;11(12):967–973. doi: 10.1111/j.1469-0691.2005.01274.x. [DOI] [PubMed] [Google Scholar]
- 7.Fitzpatrick F, Humphreys H, O’Gara JP. Evidence for icaADBC-independent biofilm development mechanism in methicillin-resistant Staphylococcus aureus clinical isolates. J Clin Microbiol. 2005;43(4):1973–1976. doi: 10.1128/JCM.43.4.1973-1976.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Seo YS, Lee DY, Rayamahji N, Kang ML, Yoo HS. Biofilm-forming associated genotypic and phenotypic characteristics of Staphylococcus spp. isolated from animals and air. Res Vet Sci. 2008;85(3):433–438. doi: 10.1016/j.rvsc.2008.01.005. [DOI] [PubMed] [Google Scholar]
- 9.Balamurugan P, Praveen Krishna V, Bharath D, Lavanya R, Vairaprakash P, Adline Princy S. Staphylococcus aureus quorum regulator SarA targeted compound, 2-[(Methylamino)methyl] phenol inhibits biofilm and Down-regulates virulence genes. Front Microbiol. 2017;8:1290. doi: 10.3389/fmicb.2017.01290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Boles BR, Horswill AR. Agr-mediated dispersal of Staphylococcus aureus biofilms. PLoS Pathog. 2008;4(4):e1000052. doi: 10.1371/journal.ppat.1000052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Shopsin B, Gomez M, Montgomery SO, Smith DH, Waddington M, Dodge DE, et al. Evaluation of protein a gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J Clin Microbiol. 1999;37(11):3556–3563. doi: 10.1128/jcm.37.11.3556-3563.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Geisinger E, Chen J, Novick RP. Allele-dependent differences in quorum-sensing dynamics result in variant expression of virulence genes in Staphylococcus aureus. J Bacteriol. 2012;194(11):2854–2864. doi: 10.1128/JB.06685-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Foster TJ. Immune evasion by staphylococci. Nat Rev Microbiol. 2005;3(12):948–958. doi: 10.1038/nrmicro1289. [DOI] [PubMed] [Google Scholar]
- 14.Edwards AM, Bowden MG, Brown EL, Laabei M, Massey RC. Staphylococcus aureus extracellular adherence protein triggers TNFalpha release, promoting attachment to endothelial cells via protein a. PLoS One. 2012;7(8):e43046. doi: 10.1371/journal.pone.0043046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Biber A, Abuelaish I, Rahav G, Raz M, Cohen L, Valinsky L, et al. A typical hospital-acquired methicillin-resistant Staphylococcus aureus clone is widespread in the community in the Gaza strip. PLoS One. 2012;7(8):e42864. doi: 10.1371/journal.pone.0042864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Geha DJ, Uhl JR, Gustaferro CA, Persing DH. Multiplex PCR for identification of methicillin-resistant staphylococci in the clinical laboratory. J Clin Microbiol. 1994;32(7):1768–1772. doi: 10.1128/jcm.32.7.1768-1772.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Klevens RM, Morrison MA, Fridkin SK, Reingold A, Petit S, Gershman K, et al. Community-associated methicillin-resistant Staphylococcus aureus and healthcare risk factors. Emerg Infect Dis. 2006;12(12):1991–1993. doi: 10.3201/eid1212.060505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Neupane S, Pant ND, Khatiwada S, Chaudhary R, Banjara MR. Correlation between biofilm formation and resistance toward different commonly used antibiotics along with extended spectrum beta lactamase production in uropathogenic Escherichia coli isolated from the patients suspected of urinary tract infections visiting Shree Birendra hospital, Chhauni, Kathmandu, Nepal. Antimicrob Resist Infect Control. 2016;5:5. doi: 10.1186/s13756-016-0104-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ohadian Moghadam Solmaz, Pourmand Mohammad Reza, Aminharati Farzaneh. Biofilm formation and antimicrobial resistance in methicillin-resistant Staphylococcus aureus isolated from burn patients, Iran. The Journal of Infection in Developing Countries. 2014;8(12):1511–1517. doi: 10.3855/jidc.5514. [DOI] [PubMed] [Google Scholar]
- 20.Azimian A, Havaei SA, Fazeli H, Naderi M, Ghazvini K, Samiee SM, et al. Genetic characterization of a vancomycin-resistant Staphylococcus aureus isolate from the respiratory tract of a patient in a university hospital in northeastern Iran. J Clin Microbiol. 2012;50(11):3581–3585. doi: 10.1128/JCM.01727-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shekarabi M, Hajikhani B, Salimi Chirani A, Fazeli M, Goudarzi M. Molecular characterization of vancomycin-resistant Staphylococcus aureus strains isolated from clinical samples: a three year study in Tehran, Iran. PLoS One. 2017;12(8):e0183607. doi: 10.1371/journal.pone.0183607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Askari E, Soleymani F, Arianpoor A, Tabatabai SM, Amini A, Naderinasab M. Epidemiology of mecA-methicillin resistant Staphylococcus aureus (MRSA) in Iran: a systematic review and meta-analysis. Iran J Basic Med Sci. 2012;15(5):1010–1019. [PMC free article] [PubMed] [Google Scholar]
- 23.Wang H, Liu Y, Sun H, Xu Y, Xie X, Chen M. In vitro activity of ceftobiprole, linezolid, tigecycline, and 23 other antimicrobial agents against Staphylococcus aureus isolates in China. Diagn Microbiol Infect Dis. 2008;62(2):226–229. doi: 10.1016/j.diagmicrobio.2008.06.003. [DOI] [PubMed] [Google Scholar]
- 24.Martin-Lopez JV, Perez-Roth E, Claverie-Martin F, Diez Gil O, Batista N, Morales M, et al. Detection of Staphylococcus aureus clinical isolates harboring the Ica gene cluster needed for biofilm establishment. J Clin Microbiol. 2002;40(4):1569–1570. doi: 10.1128/JCM.40.4.1569-1570.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cramton SE, Gerke C, Schnell NF, Nichols WW, Gotz F. The intercellular adhesion (Ica) locus is present in Staphylococcus aureus and is required for biofilm formation. Infect Immun. 1999;67(10):5427–5433. doi: 10.1128/iai.67.10.5427-5433.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Liberto MC, Matera G, Quirino A, Lamberti AG, Capicotto R, Puccio R, et al. Phenotypic and genotypic evaluation of slime production by conventional and molecular microbiological techniques. Microbiol Res. 2009;164(5):522–528. doi: 10.1016/j.micres.2007.04.004. [DOI] [PubMed] [Google Scholar]
- 27.Namvar AE, Asghari B, Ezzatifar F, Azizi G, Lari AR. Detection of the intercellular adhesion gene cluster (Ica) in clinical Staphylococcus aureus isolates. GMS Hyg Infect Control. 2013;8(1):Doc03. doi: 10.3205/dgkh000203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Grinholc M, Wegrzyn G, Kurlenda J. Evaluation of biofilm production and prevalence of the icaD gene in methicillin-resistant and methicillin-susceptible Staphylococcus aureus strains isolated from patients with nosocomial infections and carriers. FEMS Immunol Med Microbiol. 2007;50(3):375–379. doi: 10.1111/j.1574-695X.2007.00262.x. [DOI] [PubMed] [Google Scholar]
- 29.Arciola CR, Baldassarri L, Montanaro L. Presence of icaA and icaD genes and slime production in a collection of staphylococcal strains from catheter-associated infections. J Clin Microbiol. 2001;39(6):2151–2156. doi: 10.1128/JCM.39.6.2151-2156.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rohde H, Knobloch JK, Horstkotte MA, Mack D. Correlation of Staphylococcus aureus icaADBC genotype and biofilm expression phenotype. J Clin Microbiol. 2001;39(12):4595–4596. doi: 10.1128/JCM.39.12.4595-4596.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rohde H, Knobloch JK, Horstkotte MA, Mack D. Correlation of biofilm expression types of Staphylococcus epidermidis with polysaccharide intercellular adhesin synthesis: evidence for involvement of icaADBC genotype-independent factors. Med Microbiol Immunol. 2001;190(3):105–112. doi: 10.1007/s00430-001-0099-5. [DOI] [PubMed] [Google Scholar]
- 32.O’Neill E, Pozzi C, Houston P, Humphreys H, Robinson DA, Loughman A, et al. A novel Staphylococcus aureus biofilm phenotype mediated by the fibronectin-binding proteins, FnBPA and FnBPB. J Bacteriol. 2008;190(11):3835–3850. doi: 10.1128/JB.00167-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ghasemian A, Najar Peerayeh S, Bakhshi B, Mirzaee M. Comparison of biofilm formation between methicillin-resistant and methicillin-susceptible isolates of Staphylococcus aureus. Iran Biomed J. 2016;20(3):175–181. doi: 10.7508/ibj.2016.03.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yang X, Qian S, Yao K, Wang L, Liu Y, Dong F, et al. Multiresistant ST59-SCCmec IV-t437 clone with strong biofilm-forming capacity was identified predominantly in MRSA isolated from Chinese children. BMC Infect Dis. 2017;17(1):733. doi: 10.1186/s12879-017-2833-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Serray B, Oufrid S, Hannaoui I, Bourjilate F, Soraa N, Mliji M, et al. Genes encoding adhesion factors and biofilm formation in methicillin-resistant Staphylococcus aureus in Morocco. J Infect Dev Ctries. 2016;10(8):863–869. doi: 10.3855/jidc.8361. [DOI] [PubMed] [Google Scholar]
- 36.Vautor E, Magnone V, Rios G, Le Brigand K, Bergonier D, Lina G, et al. Genetic differences among Staphylococcus aureus isolates from dairy ruminant species: a single-dye DNA microarray approach. Vet Microbiol. 2009;133(1–2):105–114. doi: 10.1016/j.vetmic.2008.06.006. [DOI] [PubMed] [Google Scholar]
- 37.McCarthy H, Rudkin JK, Black NS, Gallagher L, O'Neill E, O'Gara JP. Methicillin resistance and the biofilm phenotype in Staphylococcus aureus. Front Cell Infect Microbiol. 2015;5:1. doi: 10.3389/fcimb.2015.00001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lindsay JA, Moore CE, Day NP, Peacock SJ, Witney AA, Stabler RA, et al. Microarrays reveal that each of the ten dominant lineages of Staphylococcus aureus has a unique combination of surface-associated and regulatory genes. J Bacteriol. 2006;188(2):669–676. doi: 10.1128/JB.188.2.669-676.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Park PW, Rosenbloom J, Abrams WR, Rosenbloom J, Mecham RP. Molecular cloning and expression of the gene for elastin-binding protein (ebpS) in Staphylococcus aureus. J Biol Chem. 1996;271(26):15803–15809. doi: 10.1074/jbc.271.26.15803. [DOI] [PubMed] [Google Scholar]
- 40.Otsuka T, Saito K, Dohmae S, Takano T, Higuchi W, Takizawa Y, et al. Key adhesin gene in community-acquired methicillin-resistant Staphylococcus aureus. Biochem Biophys Res Commun. 2006;346(4):1234–1244. doi: 10.1016/j.bbrc.2006.06.038. [DOI] [PubMed] [Google Scholar]
- 41.Zuniga E, Melville PA, Saidenberg AB, Laes MA, Gonsales FF, Salaberry SR, et al. Occurrence of genes coding for MSCRAMM and biofilm-associated protein bap in Staphylococcus spp. isolated from bovine subclinical mastitis and relationship with somatic cell counts. Microb Pathog. 2015;89:1–6. doi: 10.1016/j.micpath.2015.08.014. [DOI] [PubMed] [Google Scholar]
- 42.Valle J, Toledo-Arana A, Berasain C, Ghigo JM, Amorena B, Penades JR, et al. SarA and not sigmaB is essential for biofilm development by Staphylococcus aureus. Mol Microbiol. 2003;48(4):1075–1087. doi: 10.1046/j.1365-2958.2003.03493.x. [DOI] [PubMed] [Google Scholar]
- 43.Beenken KE, Blevins JS, Smeltzer MS. Mutation of sarA in Staphylococcus aureus limits biofilm formation. Infect Immun. 2003;71(7):4206–4211. doi: 10.1128/IAI.71.7.4206-4211.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Blevins JS, Elasri MO, Allmendinger SD, Beenken KE, Skinner RA, Thomas JR, et al. Role of sarA in the pathogenesis of Staphylococcus aureus musculoskeletal infection. Infect Immun. 2003;71(1):516–523. doi: 10.1128/IAI.71.1.516-523.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Dunman PM, Murphy E, Haney S, Palacios D, Tucker-Kellogg G, Wu S, et al. Transcription profiling-based identification of Staphylococcus aureus genes regulated by the agr and/or sarA loci. J Bacteriol. 2001;183(24):7341–7353. doi: 10.1128/JB.183.24.7341-7353.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Karlsson A, Arvidson S. Variation in extracellular protease production among clinical isolates of Staphylococcus aureus due to different levels of expression of the protease repressor sarA. Infect Immun. 2002;70(8):4239–4246. doi: 10.1128/IAI.70.8.4239-4246.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Blevins JS, Beenken KE, Elasri MO, Hurlburt BK, Smeltzer MS. Strain-dependent differences in the regulatory roles of sarA and agr in Staphylococcus aureus. Infect Immun. 2002;70(2):470–480. doi: 10.1128/IAI.70.2.470-480.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Pozzi C, Waters EM, Rudkin JK, Schaeffer CR, Lohan AJ, Tong P, et al. Methicillin resistance alters the biofilm phenotype and attenuates virulence in Staphylococcus aureus device-associated infections. PLoS Pathog. 2012;8(4):e1002626. doi: 10.1371/journal.ppat.1002626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mrak LN, Zielinska AK, Beenken KE, Mrak IN, Atwood DN, Griffin LM, et al. saeRS and sarA act synergistically to repress protease production and promote biofilm formation in Staphylococcus aureus. PLoS One. 2012;7(6):e38453. doi: 10.1371/journal.pone.0038453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zielinska AK, Beenken KE, Mrak LN, Spencer HJ, Post GR, Skinner RA, et al. sarA-mediated repression of protease production plays a key role in the pathogenesis of Staphylococcus aureus USA300 isolates. Mol Microbiol. 2012;86(5):1183–1196. doi: 10.1111/mmi.12048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gilot P, Lina G, Cochard T, Poutrel B. Analysis of the genetic variability of genes encoding the RNA III-activating components Agr and TRAP in a population of Staphylococcus aureus strains isolated from cows with mastitis. J Clin Microbiol. 2002;40(11):4060–4067. doi: 10.1128/JCM.40.11.4060-4067.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Tristan A, Ying L, Bes M, Etienne J, Vandenesch F, Lina G. Use of multiplex PCR to identify Staphylococcus aureus adhesins involved in human hematogenous infections. J Clin Microbiol. 2003;41(9):4465–4467. doi: 10.1128/JCM.41.9.4465-4467.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.CLSI. Performance standards for antimicrobial susceptibility testing; 27th informational supplement. CLSI document M100-S27, Wayne, Pa, USA. 2017.
- 54.NucleoSpin®. Genomic DNA from tissue Kit. Retrieved from: http://www.mn-net.com/Portals/8/attachments/Redakteure_Bio/Protocols/Genomic%20DNA/UM_gDNATissue_2017.pdf.
- 55.Mehrotra M, Wang G, Johnson WM. Multiplex PCR for detection of genes for Staphylococcus aureus enterotoxins, exfoliative toxins, toxic shock syndrome toxin 1, and methicillin resistance. J Clin Microbiol. 2000;38(3):1032–1035. doi: 10.1128/jcm.38.3.1032-1035.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Boye K, Bartels MD, Andersen IS, Moller JA, Westh H. A new multiplex PCR for easy screening of methicillin-resistant Staphylococcus aureus SCCmec types I -V Clin Microbiol Infect. 2007;13(7):725–727. doi: 10.1111/j.1469-0691.2007.01720.x. [DOI] [PubMed] [Google Scholar]
- 57.Zhang K, McClure JA, Elsayed S, Louie T, Conly JM. Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2005;43(10):5026–5033. doi: 10.1128/JCM.43.10.5026-5033.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Gowrishankar S, Kamaladevi A, Balamurugan K, Pandian SK. In vitro and in vivo biofilm characterization of methicillin-resistant Staphylococcus aureus from patients associated with pharyngitis infection. Biomed Res Int. 2016;2016:1289157. doi: 10.1155/2016/1289157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Stepanovic S, Vukovic D, Hola V, Di Bonaventura G, Djukic S, Cirkovic I, et al. Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS. 2007;115(8):891–899. doi: 10.1111/j.1600-0463.2007.apm_630.x. [DOI] [PubMed] [Google Scholar]
- 60.Atshan SS, Nor Shamsudin M, Sekawi Z, Lung LT, Hamat RA, Karunanidhi A, et al. Prevalence of adhesion and regulation of biofilm-related genes in different clones of Staphylococcus aureus. J Biomed Biotechnol. 2012;2012:976972. doi: 10.1155/2012/976972. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All relevant materials and data supporting the findings of this study are.
contained within the manuscript. Raw data can be requested from the.
corresponding author.