Abstract
To predict real-time 3D deformation field maps (DFMs) using Volumetric Cine MRI (VC-MRI) and adaptive boosting and multi-layer perceptron neural network (ADMLP-NN) for 4D target tracking.
One phase of a prior 4D-MRI is set as the prior phase, MRIprior. Principal component analysis (PCA) is used to extract three major respiratory deformation modes from the DFMs generated between the prior and remaining phases. VC-MRI at each time-step is considered a deformation of MRIprior, where the DFM is represented as a weighted linear combination of the PCA components. The PCA weightings are solved by minimizing the differences between on-board 2D cine MRI and its corresponding VC-MRI slice. The PCA weightings solved during the initial training period are used to train an ADMLP-NN to predict PCA weightings ahead of time during the prediction period. The predicted PCA weightings are used to build predicted 3D DFM and ultimately, predicted VC-MRIs for 4D target tracking. The method was evaluated using a 4D computerized phantom (XCAT) with patient breathing curves and MRI data from a real liver cancer patient. Effects of breathing amplitude change and ADMLP-NN parameter variations were assessed. The accuracy of the PCA curve prediction was evaluated. The predicted real-time 3D tumor was evaluated against the ground-truth using volume dice coefficient (VDC), center-of-mass-shift (COMS), and target tracking errors.
For the XCAT study, the average VDC and COMS for the predicted tumor were 0.92 ± 0.02and 1.06 ± 0.40 mm, respectively, across all predicted time-steps. The correlation coefficients between predicted and actual PCA curves generated through VC-MRI estimation for the 1st/2nd principal components were 0.98/0.89 and 0.99/0.57 in the SI and AP directions, respectively. The optimal number of input neurons, hidden neurons, and MLP-NN for ADMLP-NN PCA weighting coefficient prediction were determined to be 7, 4, and 10, respectively. The optimal cost function threshold was determined to be 0.05. PCA weighting coefficient and VC-MRI accuracy was reduced for increased prediction-step size. Accurate PCA weighting coefficient prediction correlated with accurate VC-MRI prediction. For the patient study, the predicted 4D tumor tracking errors in superior–inferior, anterior–posterior and lateral directions were 0.50 ± 0.47 mm, 0.40 ± 0.55 mm, and 0.28 ± 0.12 mm, respectively.
Preliminary studies demonstrated the feasibility to use VC-MRI and artificial neural networks to predict real-time 3D DFMs of the tumor for 4D target tracking.
Keywords: target tracking, real-time MRI, artificial neural networks, PCA, deep learning
1. Introduction
Respiratory target motion during radiation therapy results in intra-fraction target localization uncertainties. Lung tumors can move as much as 3 cm in the craniocaudal direction (Korreman 2015). Stereotactic body radiation therapy (SBRT) is becoming an emerging and effective treatment to treat early-stage liver and non-small cell lung cancer (NSCLC) patients with promising early outcomes (Mendez Romero et al 2006, Mendez Romero et al 2008, Fakiris et al 2009, Liu et al 2013, Chang et al 2015). SBRT uses tight planning target volume (PTV) margins and high fractionated dose to aggressively treat targets and avoid nearby critical structures. In addition, SBRT treatment times are longer than conventional treatments, making it more susceptible to localization errors caused by intra-fraction motions. Therefore, target localization, before and during treatment, is crucial for preventing tumor under-dosing and nearby healthy tissue over-dosing in SBRT.
Target tracking has been introduced in recent years as a strategy for respiratory motion management in SBRT (Hansen et al 2016). The target tracking method detects the target respiratory motion in real-time and adjusts the radiation beam position to follow the target motion to ensure the tumor is confined in the BEV during the radiation delivery. As a result, the PTV defined for the treatment can be reduced, which is beneficial for reducing the dose to the surrounding healthy tissues.
Target tracking demands the radiation beam to be synchronized with the target motion (Sharp et al 2004). Implementation of target tracking requires the incorporation of system latency in the process, which is the finite time between detection of a new target location and adjustment of the radiation beam to follow the target motion. This latency is caused by the time needed for tumor detection, beam shaping calculation, and MLC movement, and it can lead to lagging in the target tracking and consequently errors in the dose delivery. Two methods to minimize latency errors are latency error quantification into the treatment planning process and target motion prediction. Error quantification requires to redefine dose calculations to incorporate geometric errors via probability density functions (PDF). PDFs applications are limited as it assumes the dose distribution is unaffected by geometrical shifts in the medium being irradiated. As a result, internal inhomogeneities and surface curvature can cause substantial dose errors near the surface of the patient (Roland et al 2010). Conversely, the prediction methods predict future tumor position from prior and current positions, allowing the treatment system to adjust ahead of time to compensate for the latency (Krilavicius et al 2016).
The general workflow of target tracking with a prediction algorithm includes four mains steps: (1) determine current tumor position, (2) predict tumor’s next position based on current position, (3) systematically adjust treatment beam’s shape and orientation to anticipate the tumor motion. In this study, we focus on the first two steps: determine current and predict future tumor positions.
The current real-time tumor position can be determined by using real-time imaging systems. One method of real-time imaging is Calypso markers, which are electromagnetic arrays implanted into the target. Markers emit radiofrequency signals and are tracked by a detector to generate a continuous 1D signal of position with sub-millimeter levels of accuracy (Bell et al 2017). Alternatively, on-board kV/MV or MRI (Kellman et al 2009) cine slices can be used for real-time imaging. However, the real-time information is limited to 1D or 2D without volumetric information of the tumor to allow tracking of tumor motion in all three directions, which can become important for arc treatments or multiple source cobalt treatments with beams from various 3D directions. To address this, real-time volumetric imaging is under development recently for imaging modalities such as cone-beam CT (CBCT) or magnetic resonance imaging (MRI). Previous work has been done using patient prior images and deformation models for CBCT fluoroscopy that would provide fast volumetric imaging at a reduced dose (Ren et al 2008, Li et al 2010a, 2010b, Li et al 2011, Ren et al 2012, Zhang et al 2017, 2018, Gao et al 2018). We also developed volumetric-cine MRI (VC-MRI) recently for real-time volumetric MR imaging (Harris et al 2016, Harris et al 2017) for MR guided radiation therapy. VC-MRI is generated based on prior images, motion models and on-board 2D MR cine images. In comparison to fluoro CBCT, VC-MRI does not utilize ionizing radiation and provides much better soft tissue contrast.
Once the current position of the tumor is determined, the second step is to predict its future positions, as mentioned above. One method is the autoregressive integrating moving average model (ARIMA), which is a linear filtering method that predicts future respiratory signals based on a linear relationship between prior and future signals (Babu and Reddy 2014). We recently developed artificial neural networks (ANN), in particular, multi-layer perceptron neural networks (MLP-NN) to effectively predict both linear and non-linear signals from 1D patient breathing signals (Sun et al 2016). Previous prediction studies considering latency suggests predicting 100–600 ms ahead would be sufficient to account for the latency (Cho et al 2009, Liu et al 2009, Roland et al 2010, Poulsen et al 2010, Ravkilde et al 2013).
This study aims to use VC-MRI for real-time imaging and adaptive boosting and multi-layer perceptron neural network (ADMLP-NN) for VC-MRI prediction to predict real-time 3D motion of the target for 4D target tracking. ADMLP-NN has been shown to be effective in predicting future real-time position management (RPM) signals from previous ones (Sun et al 2016). Deformation field map (DFM) weighting coefficients from VC-MRI estimations are extracted and used to train an ADMLP-NN to predict future weighting coefficients. Predicted weighting coefficients are used to construct predicted DFMs and ultimately predicted VC-MRIs. VC-MRI estimation and prediction are evaluated using a 4D computerized extended-cardiac torso (XCAT) simulation of lung cancer patients and MRI data from a real liver cancer patient. For the XCAT study, motion was simulated in the anterior–posterior (AP) and superior–inferior (SI) direction based on patient-specific RPM curves. Effects of breathing amplitude change and ADMLP-NN parameter variations were assessed. The predictions accuracy is evaluated based on predicted DFM weighting coefficients and resulting predicted VC-MRIs.
2. Methods and materials
2.1. VC-MRI estimation
VC-MRI, at any time instant, is assumed to be the deformation of a prior 4D-MRI phase, MRIprior, obtained during patient simulation. On-board VC-MRI estimation can be as expressed as a function of the prior phase, MRIprior, and DFM, D, as shown in equation (1).
| (1) |
Dx, Dy and Dz are the deformation fields along the three Cartesian coordinates. D is constructed from a linear combination of the first three principal respiratory motion modes, , extracted from principal component analysis (PCA) on the patient prior 4D-MRI as seen in equation (2):
| (2) |
D0ave is the average DFM between MRIprior and other remaining prior 4D MRI phases. The coefficients, wj (j = 1,2,3), are the PCA weightings for each motion mode. PCA weighting coefficients are solved for by matching on-board sagittal MR cines with its corresponding VC-MRI slice via a data fidelity constraint.
VC-MRIs are generated by deforming MRIprior with the optimized DFM, D, via equation (1). A more detailed explanation of VC-MRI estimation can be found in Harris et al (2016).
2.2. VC-MRI prediction
VC-MRIs are predicted ahead of time using ADMLP-NN. VC-MRI prediction is based on predicting PCA weighting coefficients, , to generate predicted DFMs, Dpred, that will be used to construct predicted VCMRIs. The predicted VC-MRIs are constructed with equation (1) using the predicted DFM, Dpred, and the prior phase, MRIprior.
2.3. PCA weighting coefficient prediction using ADMLP-NN
ADMLP-NN is composed of T identical MLP-NN, which are used as weak predictors to compose a strong predictor. MLP-NNs are configured to estimate future PCA weighting coefficients from the previous coefficients.
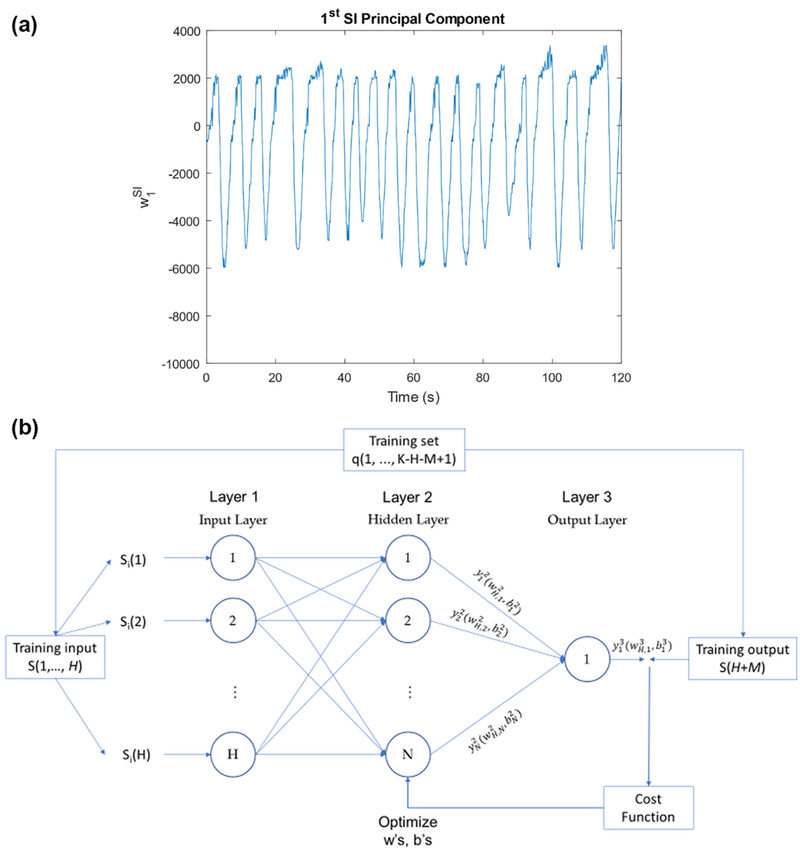
During VC-MRI estimation, the PCA weighting coefficients, wj, are tracked to generate PCA weighting coefficient curves, wj(t). Figure 1(a) shows an example for the 1st principal component in the superior–inferior (SI) direction.
Figure 1.
(a) Example of the 1st SI PCA weighting coefficient curve. (b) MLP-NN training flow chart for PCA curve prediction.
MLP-NN training divides the PCA weighting coefficient curve, wj(t), into two components separated at time index, t = K. The coefficients prior to time index K is referred to as the training signal and is used to train and determine the weights (w’s) and biases (b’s) of each MLP-NN. The coefficients after time index K are referred to as the testing signal and is used to evaluate PCA weighting coefficient predictions.
The architecture of an MLP-NN and its training flow chart is shown in figure 1(b).
MLP-NN consists of three layers (ε): input (1), hidden (2), and output (3). The input, hidden, and output layer consists of H, N, and one neuron, respectively. For each MLP-NN, the training algorithm requires, from the training signal, training sets, q, which are pairs of points consisting of a training input (for prediction) and a training output (for validation). The signals, S(1,…,H), is the training input of the first training set, q(1), used to predict the training output, S(H + M), where M is the prediction step-size. S(1) represents the PCA weighting coefficient wj(0) at t = 0 in figure 1(a). The training process continues with all training sets, where in a training signal with K points, there are q = K − H − M + 1 training sets.
The MLP-NN output, , of a neuron l in layer ε from the training set q is determined from the following equation:
| (3) |
l and j are the neuron number of the current and fore-layer. fε and is the activation function of the layer ε. and act as weights and biases to the inputs of the activation functions. Initially, the weights and biases in each MLPNN are set randomly to compute an output, . Afterwards, the network training performance is evaluated based on a cost function:
| (4) |
S(q) is the training output of training set q. The cost function is minimized by updating weights and biases via the Levenberg–Marquardt (LM) algorithm until either the pre-set maximum iteration number or cost function threshold value, G, is reached.
Furthermore, adaptive boosting (Adaboost) modifies the MLP-NN prediction algorithm by sequentially weighting identical MLP-NNs based on sample prediction error of each MLP-NN. Adaboost reduces the risk of predicting local minima and over-fitting from MLP-NN. The final predicted output, S’, of the ADMLP-NN is the weighted sum of all MLP-NN predictions:
| (5) |
is tth MLP-NN’s optimized predicted output and αt is its corresponding adaptive boosting weighting. A more detailed explanation of the adaptive boosting and ADMLP-NN can be found in Sun et al (2016).
Once the ADMLP-NN is trained, the testing signal is used to evaluate its accuracy. The signal between S(K + 1) and S(K + H) are imported to the trained ADMLP-NN to predict the first target position, S′(K + H + M). Points are continually imported to predict the next target position until the last target position, S′(K + H + M + P − 1), is predicted. P is the number of points predicted.
2.4. Simulation study using RPM signal and XCAT phantom
XCAT, a digital anthropomorphic phantom, was used to simulate a 4D extended cardiac torso for the prior 4D MRI set, on-board ground-truth VC-MRIs, and 2D cine MRIs (Segars et al 2010). A 30 mm diameter spherical tumor with uniform intensity was simulated in the middle of the lung. Both the body and tumor were simulated to move in the anterior–posterior (AP) direction and superior–inferior (SI) direction. Respiratory motions were simulated based on real patient breathing signals recorded by a real-time position management (RPM) system, where an infrared camera tracks the motion of reflective markers placed on top of a patients’ abdomen (Yan et al 2005).
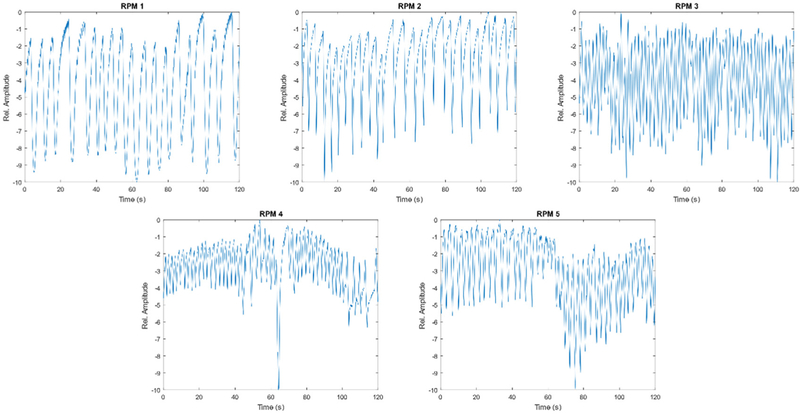
Five different two-minute long RPM signals were used to assess the effects of breathing pattern variations on VC-MRI estimation and prediction as shown in figure 2.
Figure 2.
Real-time position management (RPM) signals.
RPM 1 (top left in figure 2) represents a regular breathing pattern. On the other hand, RPM 2–5 demonstrate characteristics of baseline drift, variations in frequency and amplitude, and irregularities in breathing. Figure 2 displays the relative amplitude of the RPM signal, where peaks represent end-of-inhalation (EOI) and troughs represent end-of-exhalation (EOE). The maximum peak-to-peak amplitudes of the AP and SI motions were set to 2 and 3 cm, respectively.
A 10-phase 4D MRI was simulated as the prior 4D MRI from one cycle from each respective RPM signal. The MRI volume of each phase was composed of 256 × 256 × 100 voxels, with each voxel measuring 1.875 × 1.875 × 3 mm. The EOE phase of the prior 4D MRI was selected as MRIprior. On-board sagittal MR cines at the location corresponding to the central slice of the tumor in the MRIprior were extracted from the simulated on-board VC-MRIs, which were used as ‘ground-truth’ VC-MRIs. On-board 2D cines and VC-MRIs were generated at ~8 frames s−1 from the entire RPM signal. For 2 min of RPM signal, 1001 cine frames were generated, each with a resolution of 1.875 × 1.875 mm and slice thickness of 3 mm. For the XCAT study, region of interests (ROIs) surrounding the tumor in the sagittal cine frames were used to estimate VC-MRIs and obtain PCA weighting coefficients.
To evaluate the effects of potential patient breathing pattern changes from simulation to treatment, three on-board patient scenarios were simulated for each RPM signal:
Scenario 1: No breathing pattern change.
Scenario 2: For both body and tumor volume, the peak-to-peak amplitude of the SI and AP motion were decreased to 2 cm and 1.2 cm respectively.
Scenario 3: For both body and tumor, the peak-to-peak amplitude of the SI and AP motion were increased to 4 cm and 2.8 cm, respectively.
Additionally, on-board patient breathing hysteresis were examined for RPM 1 Scenario 1, where the AP and SI motion were simulated to be out-of-phase by 10% and 20%.
2.5. Patient study using liver cancer patient data
The VC-MRI prediction method was evaluated using one liver patient data, scanned on a GE scanner under an institutional review board-approved protocol. Details of image acquisition and 4D MRI reconstruction can be found in the previous publications (Cai et al 2011, Liu et al 2014, Harris et al 2016). To briefly summarize, a balanced steady state free precession (bSSFP) imaging acquisition technique was used to acquire 2D axial and sagittal images for retrospective sorting and 10-phase 4D MRI generation. Each phase was composed of 256 × 256 × 40 voxels. The EOI phase was used as MRIprior. 2D cine MRI was acquired continuously (3 frames s−1) for 1 min along each orientation after the 4D MRI scan. Both the prior 4D MRI and on-board 2D cine MRI have a resolution of 1.875 × 1.875 mm and a slice thickness of 5 mm. For the patient study, the entire sagittal cine frame was used to estimate VC-MRIs and obtain PCA weighting coefficients.
2.6. Optimization of ADMLP-NN parameters
ADMLP-NN was optimized by fine-tuning and evaluating the parameters, T, H, N, G, and M, at different values for the XCAT study shown in table 1.
Table 1.
ADMLP-NN default and testing parameters.
| ADMLP-NN parameter | |
|---|---|
| Number of MLP-NNs (T) | 2, 5, 8, 10, 12 |
| Number of input neurons (H) | 3, 5, 7, 9, 11 |
| Number of hidden neurons (N) | 1, 2, 3, 4, 5 |
| Cost function threshold (G) | 0, 0.05, 0.1, 0.15, 0.2 |
| Prediction step-size (M) | 1, 2, 3, 4, 5 |
Optimized ADMLP-NN parameters are used to evaluate VC-MRI prediction accuracy for the XCAT and patient study.
2.7. Evaluation methods
The predicted principle PCA weighting coefficient, wpredj, was evaluated against the estimated (true) weighting coefficients, wj, at every time-step via normalized cross-correlation (NCC) (equation (11)) and normalized root-mean-square-error (NRMSE) (equation (12)):
| (6) |
| (7) |
For the XCAT study, estimated and predicted VC-MRI tumors were evaluated against the ground-truth VCMRI tumor. Results were reported as an average across predicted time-steps with the corresponding standard deviations (STD). Tumors were contoured by an in-house MATLAB (MathWorks, Natick, MA) code based on threshold voxel values and preset ROIs. Volume dice coefficient (VDC) (equation (13)) and center-of mass-shift (COMS) (equation (14)) was used to assess the accuracy of the predicted/estimated tumor.
| (8) |
V is the volume of the tumor contoured in the predicted/estimated image and V0 is the volume of the tumor contoured in the ground-truth image.
| (9) |
Δx, Δy, and Δz, are the center-of-mass distances from V to V0.
For the patient study, a previously developed ROI feature-based motion tracking method (Cai et al 2011) was used to calculate and compare target tracking based on VC-MRI estimation, prediction, and 2D cine images acquired. The 2D sagittal cine was used to track the target motion along the AP and SI directions, and the 2D axial cine was used to track the target motion along the lateral direction. The average target tracking curves over the prediction time (15 s) from the 2D cine images were used as the reference to evaluate the accuracy of the target tracking curve extracted from VC-MRI estimation and prediction.
3. Results
3.1. XCAT result
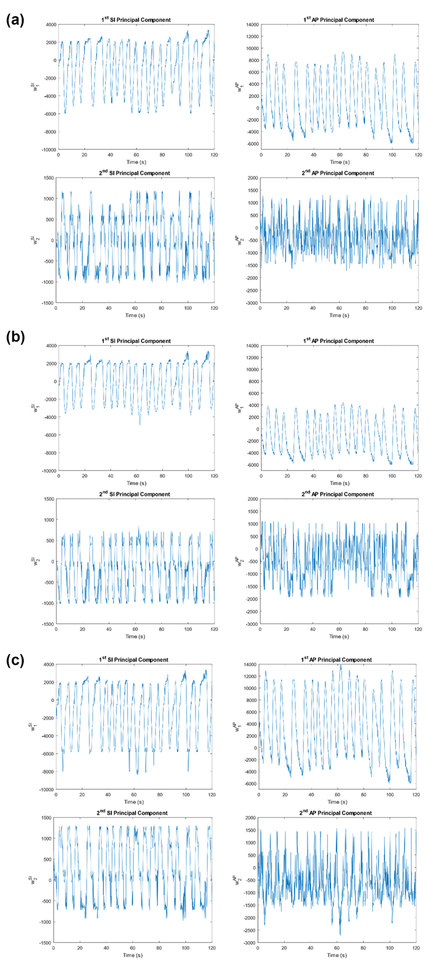
The RPM-XCAT simulations were used to optimize ADMLP-NN parameters for PCA weighting coefficient predictions. Figure 3 shows the PCA weighting coefficient curves for RPM 1 and all three scenarios generated from VC-MRI estimation. For the XCAT study, the lateral component was not evaluated as there was no motion simulated in the lateral direction. The first 90 s of each PCA curve was used to train the ADMLP-NN to predict the last 30 s.
Figure 3.
RPM 1 PCA weighting curves generated from VC-MRI estimation for (a) Scenario 1, (b) Scenario 2, and (c) Scenario 3.
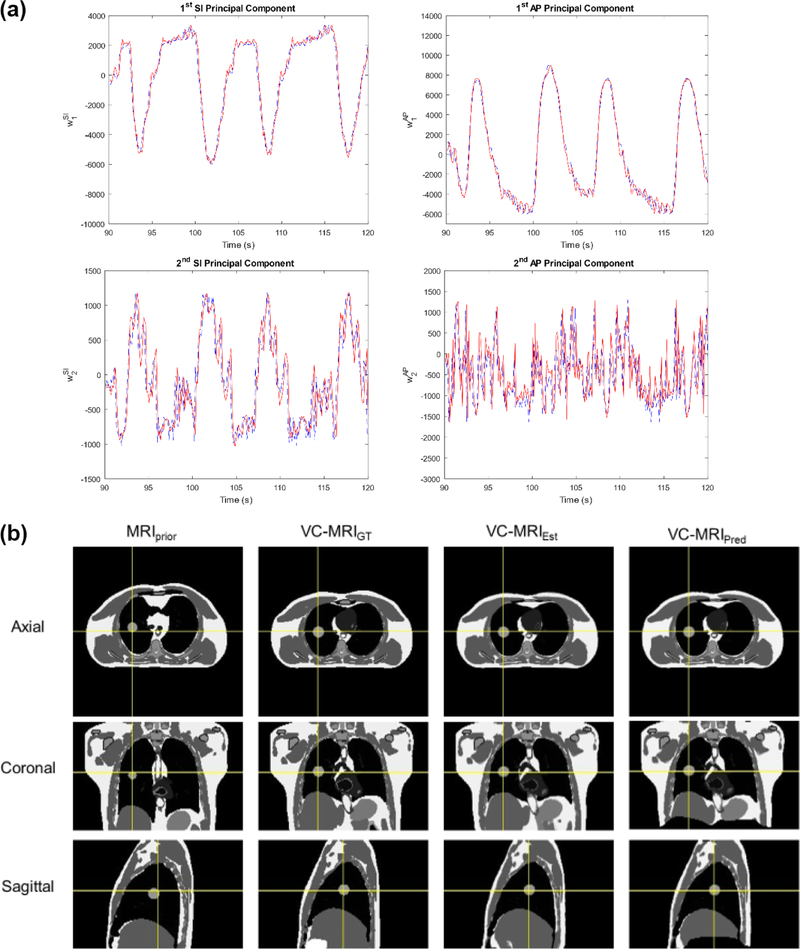
Figure 4(a) shows RPM 1 Scenario 1 predicted PCA weighting coefficients curves (red) plotted with the estimated/true curves (blue) for prediction step-size, M = 1 (120 ms). In contrast to VC-MRI estimation, predicted DFMs are constructed from the first two principal motion modes rather than three as the 3rd component is less predictable and excluding it has minimal effects on the final VC-MRI. Figure 4(b) displays the prior MRI at the EOE phase, ground-truth VC-MRI, estimated VC-MRI, and predicted VC-MRI at the EOI phase.
Figure 4.
(a) ADMLP-NN (M = 1 (120 ms)) predicted PCA weighting coefficient curve (red) and estimated/true curve (blue) for RPM 1 Scenario. (b) RPM 1 Scenario 1—Prior MRI (MRIprior) at end-of-expiration (EOE) phase, ground-truth on-board VC-MRI (VC-MRIGT), estimated on-board VC-MRI (VC-MRIEst), and predicted VC-MRI (VC-MRIPred) at end-of-inspiration (EOI) phase.
3.1.1. Effects of parameters in the ADMLP-NN network
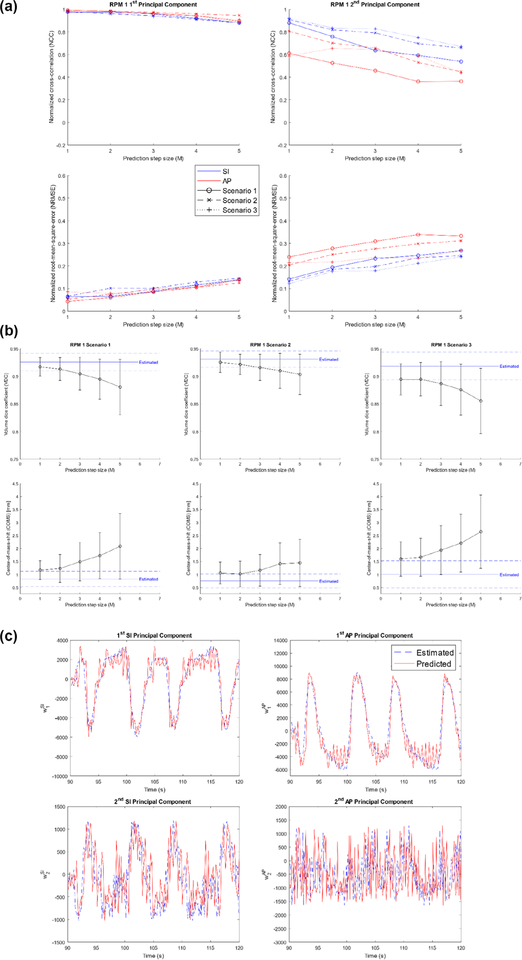
Figure 5(a) shows the NCC and NRMSE of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of prediction step-size. Figure 5(b) shows the VDC and COMS of RPM 1’s estimated and predicted VC-MRIs as a function of prediction step-size for different scenarios. Figure 5(c) shows RPM 1 Scenario 1 predicted PCA weighting coefficients curves (red) for prediction step-size, M = 4 (480 ms), plotted with the estimated/true curves (blue).
Figure 5.
(a) NCC (top) and NRMSE (bottom) of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of prediction step-sizes. (b) VDC (top) and COMS (bottom) for RPM 1’s estimated (blue; dashed—STD) and predicted (black) VC-MRIs as a function of prediction step-sizes for different scenarios. (c) ADMLP-NN (M = 4 (480 ms)) predicted PCA weighting coefficient curve (red) and estimated/true curve (blue) for RPM 1 Scenario 1.
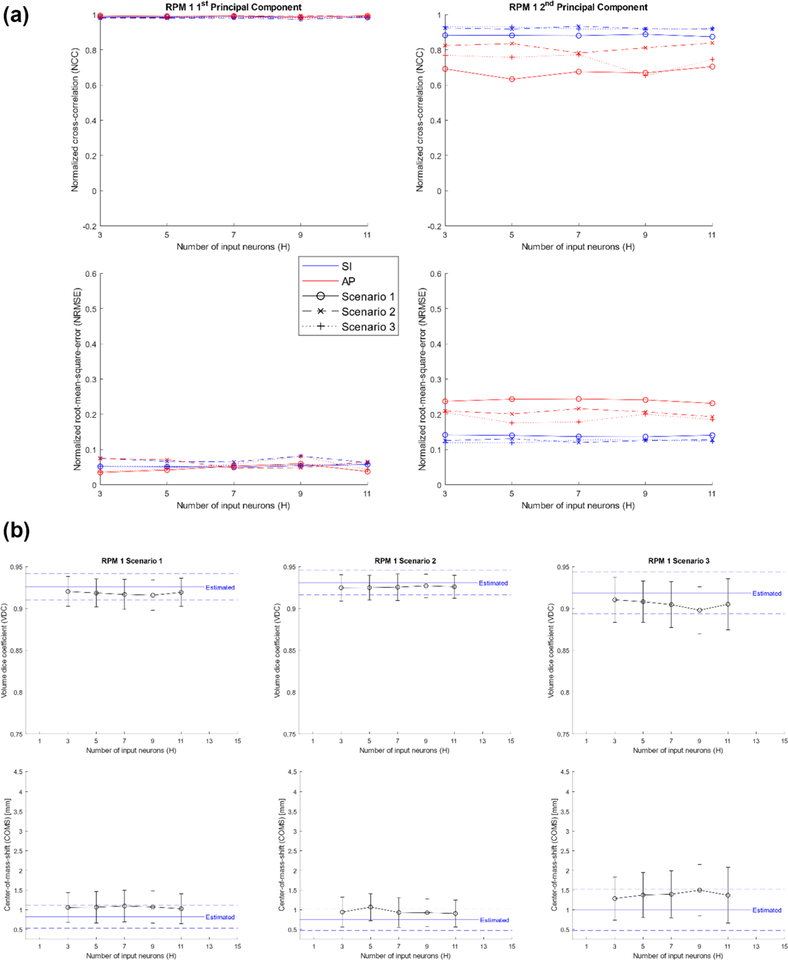
Figure 6(a) shows the NCC and NRMSE of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of input neurons. Figure 6(b) shows the VDC and COMS of RPM 1’s estimated and predicted VC-MRIS as a function of input neurons for different scenarios. From figure 6, the optimal number of input neurons was experimentally determined to be H = 7.
Figure 6.
(a) NCC (top) and NRMSE (bottom) of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of input neurons. (b) VDC (top) and COMS (bottom) for RPM 1’s estimated (blue; dashed—STD) and predicted (black) VC-MRIs as a function of input neurons for different scenarios.
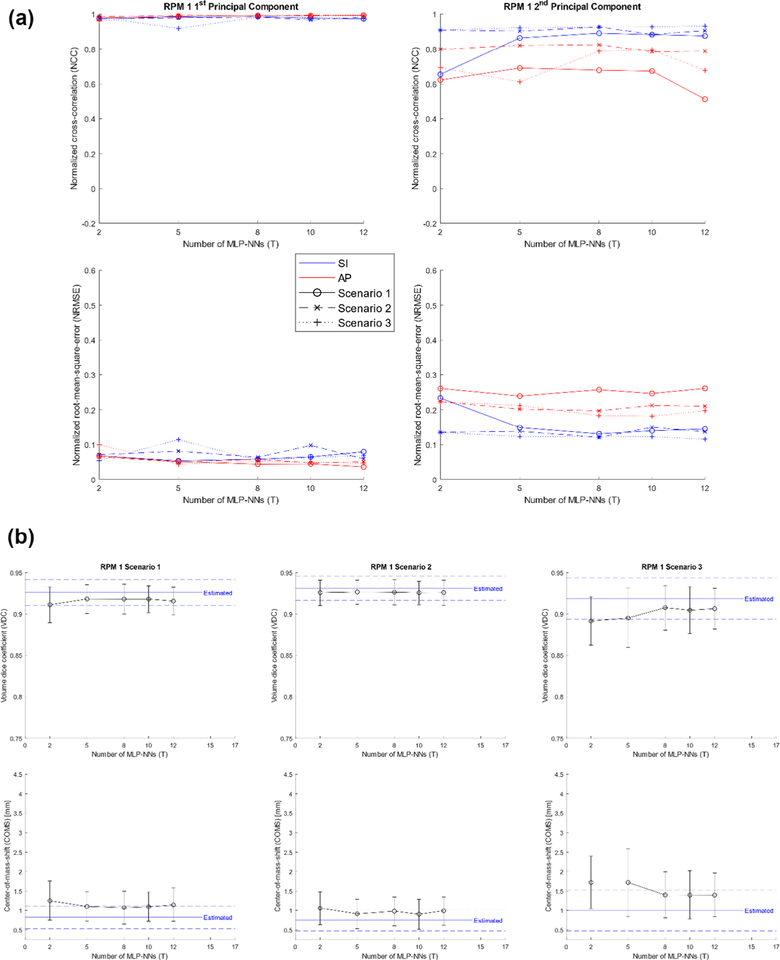
Figure 7(a) shows the NCC and NRMSE of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of MLP-NNs. Figure 7(b) shows the VDC and COMS of RPM 1’s estimated and predicted VC-MRIS as a function of MLP-NNs for different scenarios. From figure 7, the optimal number of MLP-NNs was experimentally determined to be T = 10.
Figure 7.
(a) NCC (top) and NRMSE (bottom) of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of MLP-NNs. (b) VDC (top) and COMS (bottom) for RPM 1’s estimated (blue; dashed—STD) and predicted (black) VC MRIs as a function of MLP-NNs for different scenarios.
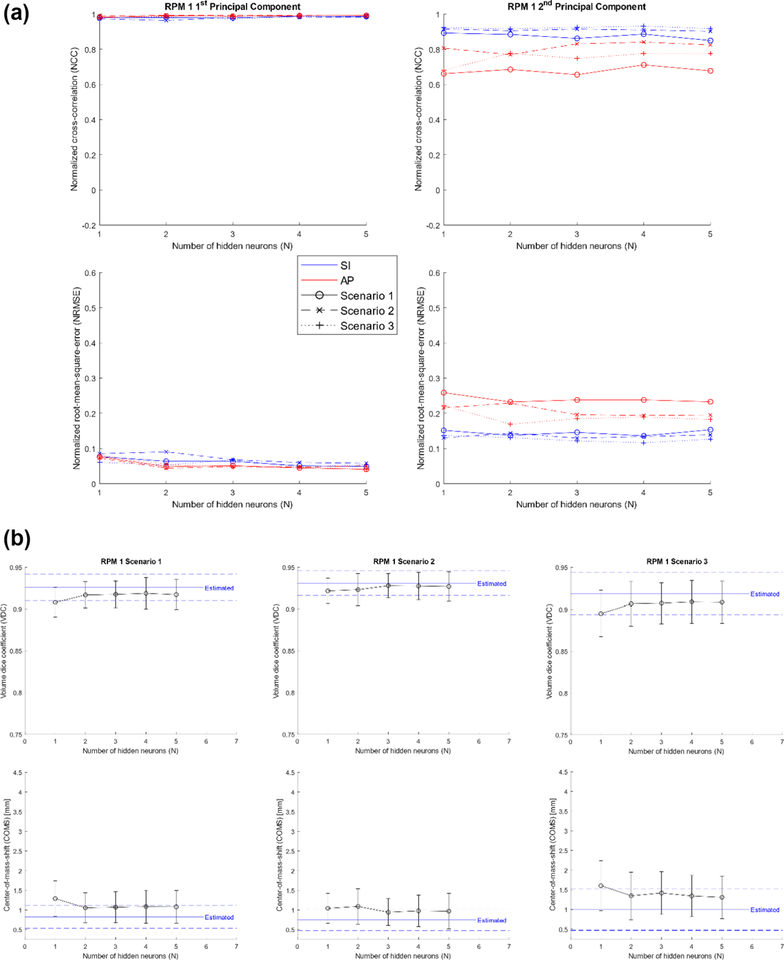
Figure 8(a) shows the NCC and NRMSE of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of hidden neurons. Figure 8(b) shows the VDC and COMS of RPM 1’s estimated and predicted VC-MRIS as a function of hidden neurons for different scenarios. From figure 8, the optimal number of hidden neurons was experimentally determined to be N = 4.
Figure 8.
NCC (top) and NRMSE (bottom) of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of hidden neurons VDC (top) and COMS (bottom) for RPM 1’s estimated (blue; dashed—STD) and predicted (black) VC-MRIs as a function of hidden neurons for different scenarios.
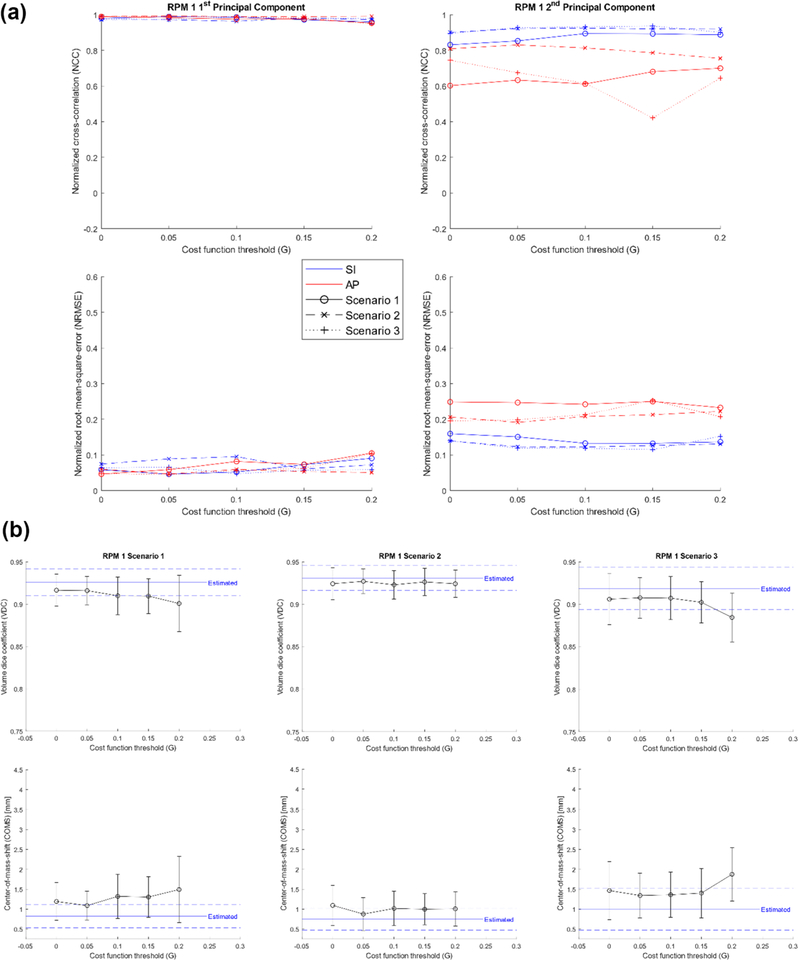
Figure 9(a) shows the NCC and NRMSE of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of cost function threshold. Figure 9(b) shows the VDC and COMS of RPM 1’s estimated and predicted VC-MRIS as a function of cost function threshold for different scenarios. From figure 9, the optimal cost function threshold was determined to G = 0.05.
Figure 9.
(a) NCC (top) and NRMSE (bottom) of RPM 1’s predicted PCA weighting coefficient curves for different scenarios as a function of cost function thresholds. (b) VDC (top) and COMS (bottom) for RPM 1’s estimated (blue; dashed—STD) and predicted (black) VC-MRIs as a function of cost function thresholds for different scenarios.
3.1.2. Predication accuracy of the ADMLP-NN network with optimized parameters
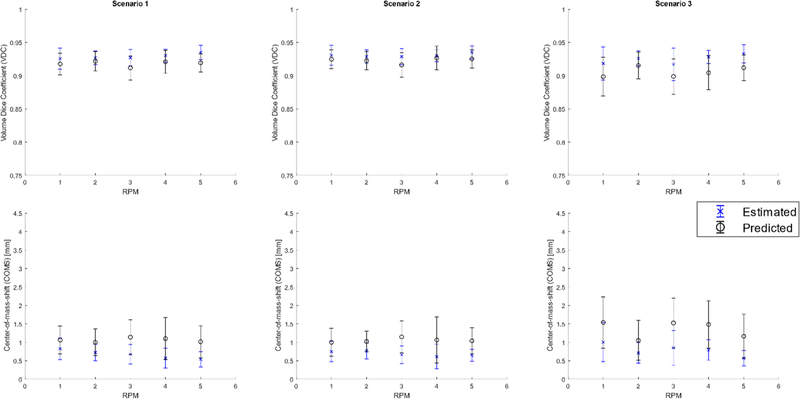
Using the optimized ADMLP-NN parameters, the predicted and estimated VC-MRIs’ VDC and COMS as a function of RPM signal is plotted in figure 10.
Figure 10.
VDC (top) and COMS (bottom) for estimated (blue) and predicted (black) VC-MRIs as a function of RPM signals for different scenarios.
3.1.3. ADMLP-NN prediction for breathing hysteresis
Table 2 shows ADMLP-NN predicted VC-MRIs’ VDC and COMS for RPM 1 Scenario 1 with no hysteresis, 10% phase shift, and 20% phase shift between AP and SI motions.
Table 2.
ADMLP-NN predicted VC-MRIs’ VDC and COMS for RPM 1 Scenario 1 with no hysteresis, 10% phase shift, and 20% phase shift between AP and SI motions.
| RPM 1 Scenario 1 | VDC | COMS (mm) |
|---|---|---|
| No hysteresis | 0.92 ± 0.02 | 1.06 ± 0.38 |
| 10% phase shift | 0.92 ± 0.02 | 1.18 ± 0.35 |
| 20% phase shift | 0.92 ± 0.01 | 1.10 ± 0.39 |
3.1.4. ADMLP-NN prediction and linear extrapolation comparison.
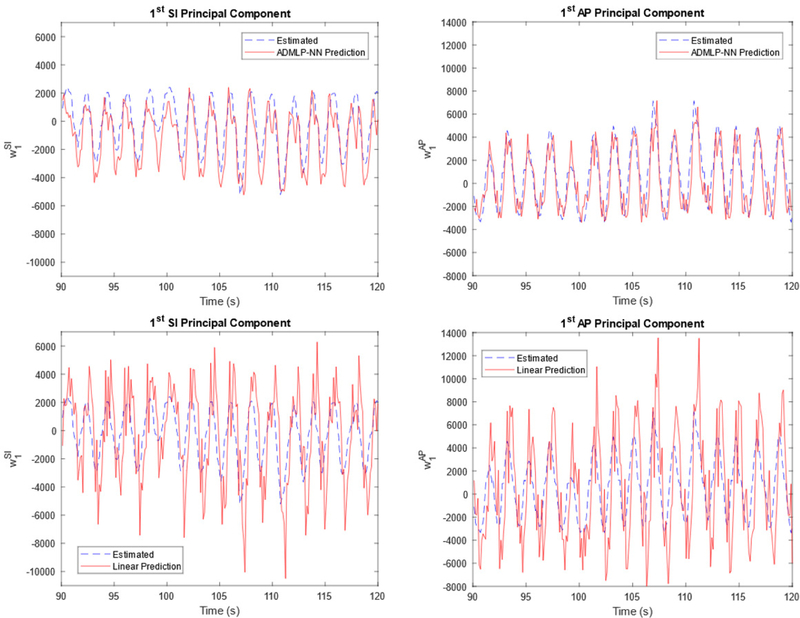
Figure 11 shows the comparison of the optimized ADMLP-NN and the linear method for predicting the 1st principal components in RPM 3 Scenario 1 with prediction step-size, M = 3 (360 ms). The estimated PCA curve was plotted as a reference to evaluate the accuracy of the prediction. Note that ADMLP-NN used 7 input neurons. Similarly, the linear model used 7 prior points to predict 3 time-steps ahead. Table 3 shows the ADMLP-NN and linear predicting methods’ prediction accuracy of the VDC and COMS of the tumor.
Figure 11.
ADMLP-NN (top) and linear (bottom) predicted (M = 3 (360 ms)) 1st principal component’s PCA weighting coefficient curve (red) and estimated/true curve (blue) for RPM 3 Scenario 1.
Table 3.
Prediction accuracy of the ADMLP-NN and linear methods for RPM 3 Scenario 1with prediction step-size, M = 3 (360 ms).
| VDC | COMS (mm) | |
|---|---|---|
| ADMLP-NN | 0.88 ± 0.03 | 1.89 ± 0.83 |
| Linear | 0.79 ± 0.09 | 4.09 ± 2.31 |
3.2. Patient result
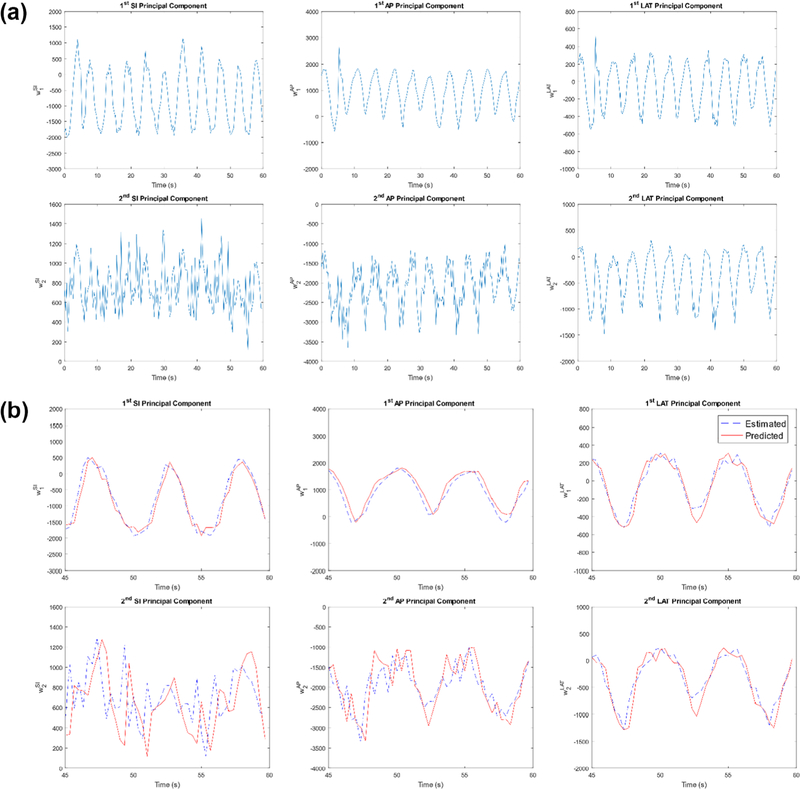
Figures 12(a) shows the PCA weighting coefficient curves for the patient study generated from VC-MRI estimation. For the patient study, the first 45 s of each PCA curve was used to train the ADMLP-NN to predict the last 15 s. Using the optimized ADMLP-NN parameters from the XCAT study, PCA weighting coefficients are predicted (red) and compared with the estimated/true curves (blue) for prediction step-size, M = 1 (330 ms), as shown in figure 12(b). Note, the cine acquisition rate differs between the XCAT and patient study, and as a result, the prediction step-size time is larger for the patient study.
Figure 12.
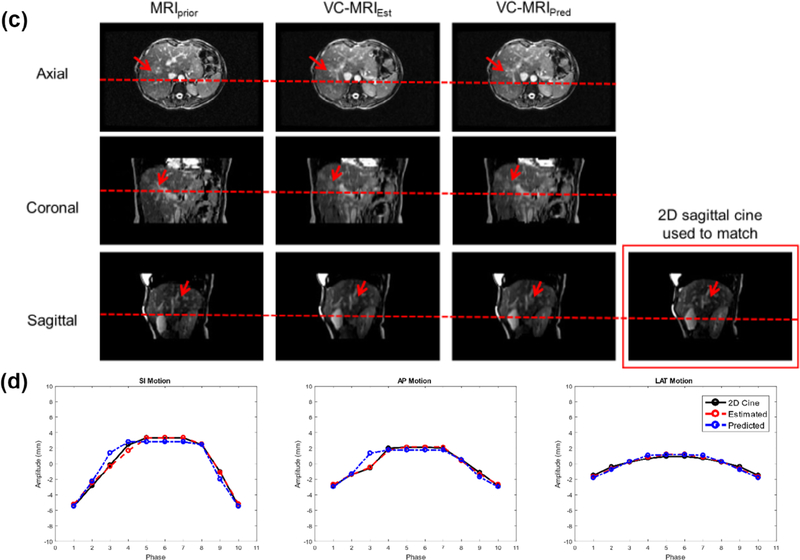
(a) Patient estimated PCA weighting curves. (b) Optimized ADMLP-NN (M = 1 (330 ms)) predicted PCA weighting coefficient curve (red) and estimated/true curve (blue) for patient study. (c) Patient—Prior MRI (MRIprior) at EOI phase, estimated on-board VC-MRI (VC-MRIEst), and predicted VC-MRI (VC-MRIPred), and 2D sagittal cine at EOE phase. (d) Liver patient tumor tracking curves based on 2D cine and VC-MRI estimation and prediction for an average cycle. Tracking along superior–inferior (SI) and anterior–posterior (AP) direction are based on sagittal cines and sagittal VC-MRI slices. Tracking along lateral direction is based on axial cines and axial VC-MRI slices.
The NCC and NRMSE of the PCA curves for the patient study are shown in table 4.
Table 4.
NCC and NRMSE of patient study’s PCA weighting coefficient curves.
| NCC | NRMSE | |||||
|---|---|---|---|---|---|---|
| PC | SI | AP | LAT | SI | AP | LAT |
| 1st | 0.981 | 0.987 | 0.963 | 0.090 | 0.109 | 0.096 |
| 2nd | 0.891 | 0.974 | 0.960 | 0.295 | 0.202 | 0.117 |
Figures 12(c) displays the liver patient’s prior MRI at the EOI phase, estimated VC-MRI, predicted VC-MRI, and sagittal 2D cine at the EOE phase. Figure 12(d) shows the liver patient’s tumor tracking based on estimated and predicted VC-MRI and 2D cine for average cycles in the SI, AP, and lateral directions. Table 5 shows the mean, standard deviation, and max tracking error based on VC-MRI estimation and prediction. The 2D cine images acquired were considered as the ground-truth VC-MRI slices.
Table 5.
Tumor motion tracking errors based on VC-MRI estimation and prediction in the liver patient study.
| SI | AP | LAT | ||||
|---|---|---|---|---|---|---|
| μ ± σ (mm) | Errormax (mm) | μ ± σ (mm) | Errormax (mm) | μ ± σ (mm) | Errormax (mm) | |
| Estimated | 0.21 ± 0.24 | 0.78 | 0.13 ± 0.09 | 0.25 | 0.18 ± 0.09 | 0.28 |
| Predicted | 0.50 ± 0.47 | 1.58 | 0.40 ± 0.55 | 1.90 | 0.28 ± 0.12 | 0.39 |
4. Discussion
Our previous study demonstrated the accuracy of the ADMLP-NN method for predicting the 1D RPM signal (Sun et al 2016). However, RPM signal only represents the motion of a small surface region of the body, which may not correlate well with the internal tumor motion as demonstrated in previous studies. As a result, RPM prediction cannot provide accurate information for target tracking. This study integrates several novel techniques we developed recently, including the ADMLP-NN prediction and the VC-MRI, to predict real-time 3D information of the tumor motion for 4D target tracking. To our knowledge, this is the first time the concept of 4D target tracking is proposed and the first study to demonstrate its feasibility using the state-of-the-art technologies. The method is also novel in that it uniquely predicts the PCA curves generated from the VC-MRI estimation to predict the 3D DFM. The integrated system was optimized for different parameters and its efficacy for 4D tracking was evaluated using both simulation and patient studies.
As shown in figure 3, PCA curves are reflective of the RPM signal’s pattern and amplitude. As expected, for Scenario 2 and 3, the PCA curves’ peak-to-peak amplitude decreased and increased, respectively, which correlates with the changes in the RPM curves.
In figure 5(a), the NCC and NRMSE between RPM 1’s predicted and estimated/true PCA curves as a function of prediction step-size showed that the prediction accuracy decreased with increasing prediction step-size (decreasing NCC, increasing NRMSE). In figure 5(b) the resulting predicted VC-MRI showed the same trend (decreasing VDC, increasing COMS). Relative to 120 ms (M = 1) prediction step-size, the VDC and COMS could degrade as much as 0.05 and 3 mm for prediction step-size of 600 ms (M = 5). This is understood by comparing figures 4(a) and 5(c)s’ predicted PCA weighting curves, where both predictions were made with the same parameters, except figure 5(c) shows prediction with a prediction step-size of M = 4. Increasing the prediction step-size resulted in noisier predicted PCA curves, which degraded VC-MRI prediction accuracy. Overall, stronger PCA weighting coefficient prediction (high NCC, low NRMSE) resulted in more accurate VC-MRI predictions (high VDC, low COMS). The optimal prediction step-size is highly dependent on a system’s latency and as a result, a system with short latency would be ideal for accurate VC-MRI predictions.
ADMLP-NN optimization showed increasing the number of input neurons would initially improve prediction results by providing more features that could improve the prediction algorithm; however, if the number of input neurons was more than the optimal value (H = 7), the performance of the network would degrade due to overfit and local minimum problems. Furthermore, using too few MLP-NNs (T < 10) resulted in poorer accuracy and larger uncertainties as the adaptive boosting algorithm had fewer MLP-NNs to formulate a strong predictor with smaller uncertainties. Predictions were relatively robust after N = 4 or more hidden neurons were used. Cost function threshold had the most significant effect on predictions. If the cost function threshold was too small (G < 0.05) or too large (G > 0.05), the prediction would overfit or underfit.
Predictions for the 1st principle component prediction were stronger and more robust against ADMLP-NN parameter variations than the 2nd component, which is reflective of the 1st component’s cyclic nature. Overall, cyclic patterns had more robust and accurate predictions.
VDC shared the same trends as COMS, indicating VC-MRI target volume accuracy correlated with target positioning accuracy. Figure 10 shows predicted VC-MRI are as accurate as estimated VC-MRIs for various RPM signals. Relatively, predicted VC-MRIs for RPM 3 and 4 were less accurate than the other RPM signals. This can be attributed to RPM 3’s high frequency and RPM 4’s abrupt change at t = 65 s affecting the prediction model. Additionally, Scenario 3’s predicted VC-MRIs had the poorest accuracy and largest margins of uncertainty. This is due to the absolute difference between the estimated and predicted PCA curves. NCC and NRMSE for different scenarios were similar; however, after unnormalizing the coefficients for VC-MRI construction, the error for Scenario 3 scaled more than the other scenarios, and as a result the absolute accuracy of Scenario 3’s VC-MRIs worsened. In contrast, Table 2 shows that ADMLP-NN VC-MRI predictions are unaffected by breathing hysteresis/phase shifts.
VC-MRI enables real-time volumetric MR imaging for high precision target localization during treatment. VC-MRI spatial resolution can be approximately 1 mm in-plane and 1–3 mm plane-to-plane depending on the resolution of the prior 4D MRI. Currently, VC-MRI reconstruction takes approximately two minutes, but can potentially be accelerated to high temporal resolutions of 3–5 frames s−1 via graphics card and parallel processing. Furthermore, ADMLP-NN PCA weighting coefficient prediction can improve VC-MRI optimization by initializing PCA weighting coefficients at the predicted value, which would be near the optimal value that satisfies the data fidelity constraint. ADMLP-NN training takes 10–20 s and once after, can greatly improve VC-MRI optimization speeds. After the VC-MRI is predicted, the corresponding predicted DFM can be used to automatically propagate tumor contours from planning images to obtain its on-board location, removing the need for on-board target re-localization and thus reducing system latency. Ultimately, real-time 4D target tracking by VC-MRI prediction would reduce the treatment errors and allow for margin reduction and dose escalation in SBRT treatments.
This work was carried out using simulation and limited patient study to investigate the feasibility of such a technique. Additionally, respiratory signals used in the simulation study were based on RPM signals only, which could be different from the actual respiration motion of internal organs. Future work will be designed to conduct more patient studies to further evaluate the clinical utility of the methods comprehensively.
5. Conclusion
It is feasible to predict PCA weighting coefficients using ADMLP-NN to construct predicted DFMs, which can be used to generate predicted VC-MRIs for 4D target tracking. Overall, ADMLP-NN prediction was dependent on an optimal number of input neurons. Using more hidden neurons and MLP-NNs in ADMLP-NN would improve VC-MRI prediction accuracy and minimize its uncertainty. VC-MRI prediction accuracy was sensitive to ADMLP-NN cost function threshold as too small thresholds would overfit the prediction and too large thresholds would underfit it. Increasing the prediction step size led to degradation of the prediction accuracy. Patient studies showed that VC-MRIs could be predicted 330 ms ahead of time with 0.5–2 mm accuracy in all three Cartesian directions, which can compensate for treatment system latencies to achieve 4D target tracking.
Acknowledgment
This work was supported by the National Institutes of Health under Grant No. R01-CA184173.
References
- Babu CN and Reddy BE 2014. A moving-average filter based hybrid ARIMA–ANN model for forecasting time series data Appl. Soft Comput 23 27–38 [Google Scholar]
- Bell LJ, Eade T, Kneebone A, Hruby G, Alfieri F, Bromley R, Grimberg K, Barnes M and Booth JT 2017. Initial experience with intra-fraction motion monitoring using Calypso guided volumetric modulated arc therapy for definitive prostate cancer treatment J. Med. Radiat. Sci 64 25–34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai J, Chang Z, Wang Z, Segars WP and Yin F 2011. Four-dimensional magnetic resonance imaging (4D-MRI) using image-based respiratory surrogate: a feasibility study Med. Phys 38 6384–94 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang JY et al. 2015. Stereotactic ablative radiotherapy versus lobectomy for operable stage I non-small-cell lung cancer: a pooled analysis of two randomised trials Lancet Oncol 16 630–7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho B, Poulsen PR, Sloutsky A, Sawant A and Keall PJ 2009. First demonstration of combined kV/MV image-guided real-time dynamic multileaf-collimator target tracking Int. J. Radiat. Oncol. Biol. Phys 74 859–67 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fakiris AJ, McGarry RC, Yiannoutsos CT, Papiez L, Williams M, Henderson MA and Timmerman R 2009. Stereotactic body radiation therapy for early-stage nonsmall-cell lung carcinoma: four-year results of a prospective phase II study Int. J. Radiat. Oncol. Biol. Phys 75 677–82 [DOI] [PubMed] [Google Scholar]
- Gao H, Zhang Y, Ren L and Yin FF 2018. Principal component reconstruction (PCR) for cine CBCT with motion learning from 2D fluoroscopy Med. Phys 45 167–77 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen R, Ravkilde T, Worm ES, Toftegaard J, Grau C, Macek K and Poulsen PR 2016. Electromagnetic guided couch and multileaf collimator tracking on a TrueBeam accelerator Med. Phys 43 2387–98 [DOI] [PubMed] [Google Scholar]
- Harris W, Ren L, Cai J, Zhang Y, Chang Z and Yin F 2016. A technique for generating volumetric cine-magnetic resonance imaging Int. J. Radiat. Oncol. Biol. Phys 95 844–53 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris W, Yin F, Wang C, Zhang Y, Cai J and Ren L 2017. Accelerating volumetric cine MRI (VC-MRI) using undersampling for real-time 3D target localization/tracking in radiation therapy: a feasibility study Phys. Med. Biol 63 01NT01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kellman P, Chefdhotel C, Lorenz CH, Mancini C, Arai AE and Mcveigh ER 2009. High spatial and temporal resolution cardiac cine MRI from retrospective reconstruction of data acquired in real time using motion correction and resorting Magn. Reson. Med 62 1557–64 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korreman SS. Image-guided radiotherapy and motion management in lung cancer. Br. J. Radiol. 2015;88:20150100. doi: 10.1259/bjr.20150100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krilavicius T, Zliobaite I, Simonavicius H and Jaruevicius L 2016. Predicting respiratory motion for real-time tumour tracking in radiotherapy 2016 IEEE 29th Int. Symp. on Computer-Based Medical Systems (Dublin, Ireland, 20–24 June 2016) ( 10.1109/cbms.2016.52) [DOI] [Google Scholar]
- Li R, Jia X, Lewis JH, Gu X, Folkerts M, Men C and Jiang SB 2010a. Single-projection based volumetric image reconstruction and 3D tumor localization in real time for lung cancer radiotherapy Int. Conf. on Medical Image Computing and ComputerAssisted Intervention vol 6363 pp 449–56 [DOI] [PubMed] [Google Scholar]
- Li R, Jia X, Lewis JH, Gu X, Folkerts M, Men C and Jiang SB 2010b. Real-time volumetric image reconstruction and 3D tumor localization based on a single x-ray projection image for lung cancer radiotherapy Med. Phys 37 2822–6 [DOI] [PubMed] [Google Scholar]
- Li R, Lewis JH, Jia X, Zhao T, Liu W, Wuenschel S, Lamb J, Yang D, Low DA and Jiang SB 2011. On a PCA-based lung motion model Phys. Med. Biol 56 6009–30 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu E, Stenmark MH, Schipper MJ, Balter JM, Kessler ML, Caoili EM, Lee OE, Ben-Josef E, Lawrence TS and Feng M 2013. Stereotactic body radiation therapy for primary and metastatic liver tumors Transl. Oncol 6 442–6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Shi C, Lin B, Ha CS and Papanikolaou N 2009. Delivery of four-dimensional radiotherapy with TrackBeam for moving target using a dual-layer MLC: dynamic phantoms study J. Appl. Clin. Med. Phys 10 21–33 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Yin F, Chang Z, Czito BG, Palta M, Bashir MR, Qin Y and Cai J 2014. Investigation of sagittal image acquisition for 4D-MRI with body area as respiratory surrogate Med. Phys 41 101902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mendez Romero A et al. 2006. Stereotactic body radiation therapy for primary and metastatic liver tumors: a single institution phase i–ii study Acta Oncol 45 831–7 [DOI] [PubMed] [Google Scholar]
- Mendez Romero A, Wunderink W, van Os RM, Nowak PJ, Heijmen BJ, Nuyttens JJ, Brandwijk RP, Verhoef C, Ijzermans JN and Levendag PC 2008. Quality of life after stereotactic body radiation therapy for primary and metastatic liver tumors Int. J. Radiat. Oncol. Biol. Phys 70 1447–52 [DOI] [PubMed] [Google Scholar]
- Poulsen PR, Cho B, Ruan D, Sawant A and Keall PJ 2010. Dynamic multileaf collimator tracking of respiratory target motion based on a single kilovoltage imager during arc radiotherapy Int. J. Radiat. Oncol. Biol. Phys 77 600–7 [DOI] [PubMed] [Google Scholar]
- Ravkilde T, Keall PJ, Grau C, Høyer M and Poulsen PR 2013. Time-resolved dose distributions to moving targets during volumetric modulated arc therapy with and without dynamic MLC tracking Med. Phys 40 111723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren L, Chetty IJ, Zhang J, Jin JY, Wu QJ, Yan H, Brizel DM, Lee WR, Movsas B and Yin FF 2012. Development and clinical evaluation of a three-dimensional conebeam computed tomography estimation method using a deformation field map Int. J. Radiat. Oncol. Biol. Phys 82 1584–93 [DOI] [PubMed] [Google Scholar]
- Ren L, Zhang J, Thongphiew D, Godfrey DJ, Wu QJ, Zhou SM and Yin FF 2008. A novel digital tomosynthesis (DTS) reconstruction method using a deformation field map Med. Phys 35 3110–5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roland T, Mavroidis P, Shi C and Papanikolaou N 2010. Incorporating system latency associated with real-time target tracking radiotherapy in the dose prediction step Phys. Med. Biol 55 2651–68 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Segars WP, Sturgeon G, Mendonca S, Grimes J and Tsui BM 2010. 4D XCAT phantom for multimodality imaging research Med. Phys 37 4902–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp GC, Jiang SB, Shimizu S and Shirato H 2004. Prediction of respiratory tumour motion for real-time image-guided radiotherapy Phys. Med. Biol 49 425–40 [DOI] [PubMed] [Google Scholar]
- Sun W, Yin F, Jiang M, Dang J and You T 2016. Respiratory signal prediction based on adaptive boosting and multilayer perceptron neural network Int. J. Radiat. Oncol. Biol. Phys 96 E702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan H, Yin F, Zhu G, Ajlouni M and Kim JH 2005. Adaptive prediction of internal target motion using external marker motion: a technical study Phys. Med. Biol 51 31–44 [DOI] [PubMed] [Google Scholar]
- Zhang Y, Deng X, Yin FF and Ren L 2018. Image acquisition optimization of a limited-angle intrafraction verification (LIVE) system for lung radiotherapy Med. Phys 45 340–51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Yin FF, Zhang Y and Ren L 2017. Reducing scan angle using adaptive prior knowledge for a limited-angle intrafraction verification (LIVE) system for conformal arc radiotherapy Phys. Med. Biol 62 3859–82 [DOI] [PMC free article] [PubMed] [Google Scholar]