SUMMARY
The phytohormone auxin plays crucial roles in nearly every aspect of plant growth and development. The AUXIN RESPONSE FACTOR (ARF) transcription factor family regulates auxin-responsive gene expression and exhibit nuclear localization in regions of high auxin responsiveness. Here we show that the ARF7 and ARF19 proteins accumulate in micron-sized assemblies within the cytoplasm of tissues with attenuated auxin responsiveness. We found that the intrinsically disordered middle region and the folded PB1 interaction domain of ARFs drive protein assembly formation. Mutation of a single lysine within the PB1 domain abrogates cytoplasmic assemblies, promotes ARF nuclear localization, and results in an altered transcriptome and morphological defects. Our data suggest a model in which ARF nucleo-cytoplasmic partitioning regulates auxin responsiveness, thus providing a mechanism for cellular competence for auxin signaling.
Graphical Abstract

eTOC BLURB
ARF transcription factors mediate activity of the phytohormone auxin, regulating every aspect of plant development. Powers et al. determine that some ARFs undergo phase transition to form large-order cytoplasmic protein assemblies that limit auxin responsiveness in a developmentally relevant context, illustrating a strong link between condensate formation and biological function.
INTRODUCTION
Auxins regulate diverse and critical aspects of cell division and expansion (Woodward and Bartel, 2005). Normal plant growth and development requires tight control of auxin responses, which are modulated by the AUXIN RESPONSE FACTOR (ARF) family of transcription factors (Korasick et al., 2015b; Wang and Estelle, 2014). In addition to ARFs, auxin-binding TRANSPORT INHIBITOR RESPONSE1/AUXIN SIGNALING F-BOX (TIR1/AFB) proteins and the AUXIN/INDOLE-3-ACETIC ACID INDUCIBLE (Aux/IAA) repressor proteins act in the canonical auxin signaling system (Korasick et al., 2015b; Wang and Estelle, 2014). Binding of Aux/IAA repressor proteins to the ARFs represses transcription factor function under low auxin conditions (Enders and Strader, 2015). Increased auxin levels result in Aux/IAA dissociation and degradation to relieve ARF repression, which allows for auxin-regulated gene transcription (Enders and Strader, 2015).
Within Arabidopsis there are 22 ARFs, most containing three regions of homology: an N-terminal B3-type DNA-binding domain (DBD) that recognizes auxin response elements (Boer et al., 2014; Guilfoyle, 2015), a variable middle region (MR), and a C-terminal type I/II Phox and Bem1p (PB1) domain (Guilfoyle and Hagen, 2007; Korasick et al., 2014; Nanao et al., 2014). Type I/II PB1 domains contain an invariant positively-charged lysine and a cluster of negatively-charged residues on opposite domain faces, which create asymmetries in surface electrostatic potential and allow bi-directional protein-protein interactions (Guilfoyle and Hagen, 2012; Korasick et al., 2015b). Structural, biochemical, and biophysical studies demonstrate that the PB1 domain drives ARF-Aux/IAA (Han et al., 2014) and ARF-ARF interactions (Han et al., 2014; Korasick et al., 2015a; Nanao et al., 2014). The bi-directional PB1 domain interactions also facilitate multimerization of ARF5 and ARF7 in vitro (Korasick et al., 2014; Nanao et al., 2014). The ARF MR may confer either activating or repressing transcriptional activity to the ARF based on enrichment of glutamine (activating) or serine (repressing) residues (Guilfoyle and Hagen, 2007; Ulmasov et al., 1999). ARF5–8 and ARF19 likely function as transcriptional activators, whereas other ARFs likely act as repressors (Tiwari et al., 2003; Ulmasov et al., 1999). Little is known about the molecular function of the MR in ARF proteins or its role in auxin-regulated gene expression.
As a single molecule, auxin controls a vast set of distinct plant processes. However, it is unclear how auxin, as a single master regulator distributed across the entire plant, coordinates growth behavior in a spatially distinct manner. Recent work revealed the importance of the ARF PB1 domain for protein-protein interaction (Korasick et al., 2014; Nanao et al., 2014); however, data supporting ARF multimerization and its contribution to auxin responses in planta have remained largely elusive. In this work, we present data indicating that in root tissues with dampened auxin responses, the Class-A ARF proteins ARF7 and ARF19 form cytoplasmic assemblies. Based on the results of our stoichiometry measurements and in accord with formal definitions introduced in the literature (Banani et al., 2017; Shin and Brangwynne, 2017), we define the cytoplasmic assemblies as non-stoichiometric ARF protein condensates that form in a concentration-dependent manner and are characterized by either liquid- or solid-like material properties, depending on their age. We identify roles for the intrinsically disordered MR and the folded bipartite PB1 domain in the formation of these protein assemblies. Disruption of ARF assembly formation results in morphological defects and altered transcriptional auxin responsiveness, suggesting a model in which ARF nucleo-cytoplasmic partitioning mediated by the formation of cytoplasmic protein assemblies i.e., non-stoichiometric ARF condensates, regulates auxin transcriptional responses in a cell-specific manner. This is critical for proper plant growth and development.
RESULTS
ARF proteins localize to cytoplasmic assemblies in certain cell types
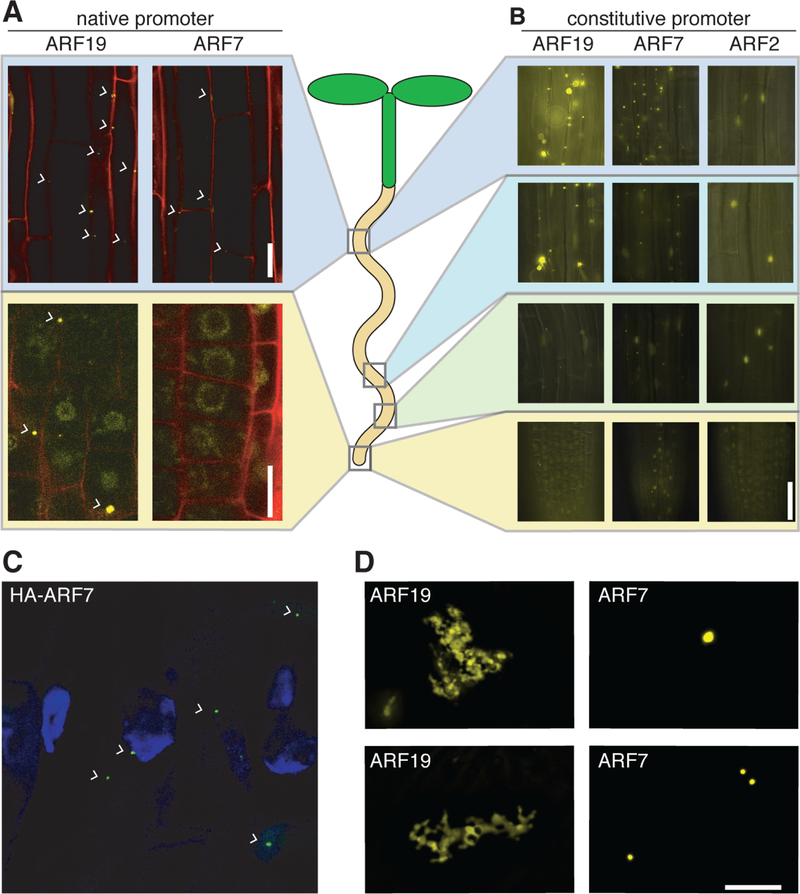
Crystallographic and biochemical studies of ARF proteins revealed that the PB1 domain drives the formation of large multimeric complexes in vitro (Korasick et al., 2014; Nanao et al., 2014). To investigate whether ARF proteins multimerize in planta, we examined Arabidopsis plants expressing ARF19:ARF19-Venus and ARF7:ARF7-Venus. ARF7 and ARF19 are Class-A activating transcription factors that serve as a nexus in several signaling pathways that regulate plant growth, lateral root development, apical dominance, and tropic responses (Chandler, 2016). Unexpectedly, we observed the accumulation of ARF7-Venus and ARF19-Venus in cytoplasmic protein assemblies displaying punctate fluorescence in mature cells of the upper root (Figure 1A and S1A). Further analysis within the root revealed that localization to cytoplasmic assemblies diminishes in actively growing cells in lower portions of the root (Figure 1A and S1A). Consistent with previous reports (Okushima et al., 2007), ARF7-Venus and ARF19-Venus localized to the nucleus in meristem cells near the root tip (Figure 1A). Seedlings expressing UBQ10 promoter-driven YFP-tagged ARF19 or 35S promoter-driven ARF7-GFP (Okushima et al., 2007), both with constitutive expression, displayed similar localization (Figure 1B), suggesting that these tissue-specific differences in localization were not due to gene expression levels. The same localization pattern in these lines suggests that formation of cytoplasmic ARF7 and ARF19 assemblies depends on cell type, irrespective of promoter. Further, cytoplasmic assemblies were found in the upper root of 35S promoter-driven ARF7-HA (Figure 1C), suggesting that these are not the result of tagging with a fluorescent protein. Cytoplasmic assemblies for both ARF7 and ARF19 were micron-sized (Figure 1D).
Figure 1. Activating ARF localization is dependent on cell type.
(A) Confocal microscopy images of upper root and root tip sections from 5d-old ARF19:ARF19-Venus (ARF19), ARF7:ARF7-Venus (ARF7) seedlings counterstained with propidium iodide (red signal), taken from upper root and root tip sections. Bar = 20 µm.
(B) Fluorescence microscopy images from upper root, intermediate, and root tip sections of 5d-old UBQ10:YFP-ARF19 (ARF19), 35S:ARF7-GFP (ARF7; (Okushima et al., 2007)), and UBQ10:YFP-ARF2 (ARF2) Arabidopsis seedlings. Bar = 50 µm.
(C) Confocal image of immunostained condensates in 35S:ARF7-HA seedlings counterstained with DAPI (blue signal).
(D) High-resolution deconvolution microscopy images of YFP-ARF19 and ARF7-GFP cytoplasmic protein assemblies. Bar = 10 µm.
Both ARF7 and ARF19 are transcriptional activators; whereas most ARF family members are considered to be transcriptional repressors (Tiwari et al., 2003; Ulmasov et al., 1999). We examined localization of the repressor ARF2 using UBQ10:YFP-ARF2 lines and found no cytoplasmic protein assemblies, with YFP-ARF2 nuclear localization in all examined cell types (Figure 1A). The DBD and PB1 domains are highly conserved among these three proteins. ARF7/19 and ARF2 protein sequences primarily differ in the enrichment of either glutamine (ARF7 and ARF19) or serine (ARF2) in their MRs, consistent with the possibility that the MR may contribute to assembly formation.
The ARF19 PB1 domain is necessary for protein assembly formation
We hypothesized that ARF7 and ARF19 cytoplasmic localization is driven by formation of the large protein assemblies we observed in our reporter lines (Figures 1 and 2C), leading us to examine which ARF protein features might promote protein assembly. A conserved lysine (K962) and a cluster of negatively charged residues leads to asymmetries in surface electrostatic potentials that drive ARF-ARF PB1 domain interactions and ARF PB1 multimer formation in vitro (Korasick et al., 2014; Nanao et al., 2014). Mutation of the corresponding lysine in ARF5 (Nanao et al., 2014) and ARF7 (Korasick et al., 2015a; Korasick et al., 2014) abrogates PB1 protein-protein interaction and disrupts PB1 domain multimerization in vitro. Accordingly, we hypothesized that the PB1 domain plays a similar role in planta and may be involved in promoting ARF-ARF interactions to facilitate protein assembly formation. To test this, we generated a YFP-ARF19 reporter line with a K962A mutation to disrupt PB1 domain interactions (UBQ10:YFP-ARF19K962A). We elected to use the UBQ10 mild constitutive promoter in these experiments to normalize gene expression and protein accumulation across tissues. The YFP-ARF19K962A protein localized to the nucleus in upper root tissues, whereas wild-type YFP-ARF19 formed cytoplasmic punctate structures in those same tissues (Figure 2C). These data suggest that the asymmetry in the PB1 domain surface electrostatic potential, which enables ARF19 multimerization, is an important contributor to cytoplasmic ARF19 assembly formation in planta.
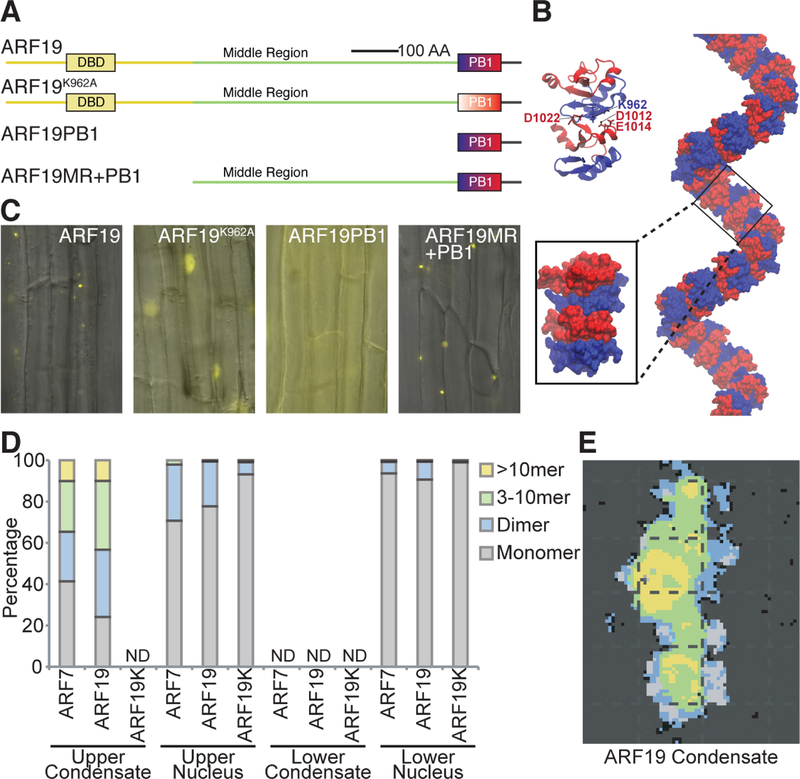
Figure 2. The ARF19 PB1 domain and middle region (MR) are required for protein assembly formation.
(A)Cartoon depictions of examined YFP-tagged ARF19 variants and truncations.
(B) ARF19 PB1 domain homology model with the negative (red) and positive (blue) halves of each indicated. Lys962 is at the interface on the positive face and is surrounded by a cluster of negatively charged residues from the opposing domain (Asp1012, Glu1014, Asp1022). The filamentous asymmetric unit from PDB entry 4NJ6 was used to construct a putative macromolecular filament through repetitive addition.
(C) Fluorescence microscopy images from the upper root regions of 5d-old UBQ10:YFP-ARF19, UBQ10:YFP-ARF19K962A, UBQ10:YFP-ARF19PB1, and UBQ10:YFP-ARF19MR+PB1 Arabidopsis seedlings.
(D) Quantification of ARF oligomeric state in upper root and root tip sections of ARF proteins in nuclei or condensates (n ≥ 10).
(E) Representative N&B analysis of an image of a YFP-ARF19 condensate from the upper root. Black represents background fluorescence, gray monomer, blue dimmer, green 3–10mer, and yellow >10mer. ND: not detected.
To further quantify PB1 domain interaction contributions in driving protein oligomerization and formation of the large ARF cytoplasmic assemblies, we determined ARF oligomeric state using scanning Fluorescence Correlation Spectroscopy (Scanning FCS) (Clark et al., 2016; Clark and Sozzani, 2017b). We determined the oligomeric state of UBQ10 promoter-driven YFP-ARF19 and YFP-ARF19K962A, as well as ARF7-YFP driven by its native promoter, in cytoplasmic protein assemblies and in nuclei of epidermal cells in both the upper and lower part of the root (late differentiation zone and meristematic zone). We found that ARF7 and ARF19 predominantly form higher-order oligomers (≥3) in the cytoplasmic protein assemblies, with a smaller fraction forming dimers or monomers (Figure 2D). Notably, these proteins were arranged such that higher-order oligomers were at the center of the protein assemblies, whereas ARF dimers and monomers were found at the periphery (Figure 2E). No cytoplasmic assemblies were detected (ND) in YFP-ARF19K962A, confirming the importance of PB1 domain interactions in driving oligomerization. Although many cells in upper parts of the root did not display nuclear ARF signal, when nuclear localized, both YFP-ARF7 and YFP-ARF19 were primarily monomeric in these cells, with 23–30% dimeric protein. Conversely, YFP-ARF19K962A was ~7% dimeric protein in these same cell types, suggesting that PB1 domain interactions may be important for enabling ARF dimerization in the nucleus, presumably through DBD interactions. We did not detect any cytoplasmic protein assemblies in the lower part of the root in any of the examined lines. Meanwhile, YFP-ARF7 and YFP-ARF19 were primarily monomeric in the nuclei of root meristem cells, with only 7–10% dimeric protein. YFP-ARF19K962A dimers made up less than 1% of nuclear protein in cells from the lower root. Thus, these results suggest that PB1 domain interactions are required for cytoplasmic assembly formation and contribute directly to the oligomeric states of ARF proteins within these assemblies.
The intrinsically disordered MR in ARF19 is necessary for condensate formation
To determine whether the ARF19 PB1 domain is sufficient to drive cytoplasmic assembly formation in planta, we examined the UBQ10:YFP-ARF19PB1 line, expressing the ARF19 PB1 domain (M958-to-stop) fused to YFP. The UBQ10:YFP-ARF19PB1 line displayed diffuse fluorescence consistent with cytoplasmic localization of YFP-ARF19PB1, but did not form cytoplasmic assemblies (Figure 2C). Thus, the PB1 domain alone is insufficient to drive ARF cytoplasmic assembly formation in planta. We expanded the truncated ARF19 to include the variable MR and found cytoplasmic punctate structures in lines expressing YFP-ARF19MR-PB1 (M395-to-stop), demonstrating that both the PB1 domain and the MR contribute to interactions required for cytoplasmic ARF assembly formation in planta.
The requirement of the ARF19 MR to form cytoplasmic assemblies led us to further investigate its biophysical properties. The ARF19 MR is a 563-residue low complexity region enriched in glutamine, serine, asparagine and leucine (50% of the residues) and depleted in charged residues (9%) (Figures 3A and S2). Sequence analysis predicts that the MR is an intrinsically disordered region (IDR) and is unlikely to be defined by persistent secondary and/or tertiary structures (Figure 3A) (van der Lee et al., 2014) unlike the N-terminal (1–394) and C-terminal (958–1086) regions, which are predicted to fold into DBD and PB1 domains, respectively (Figure 3A).
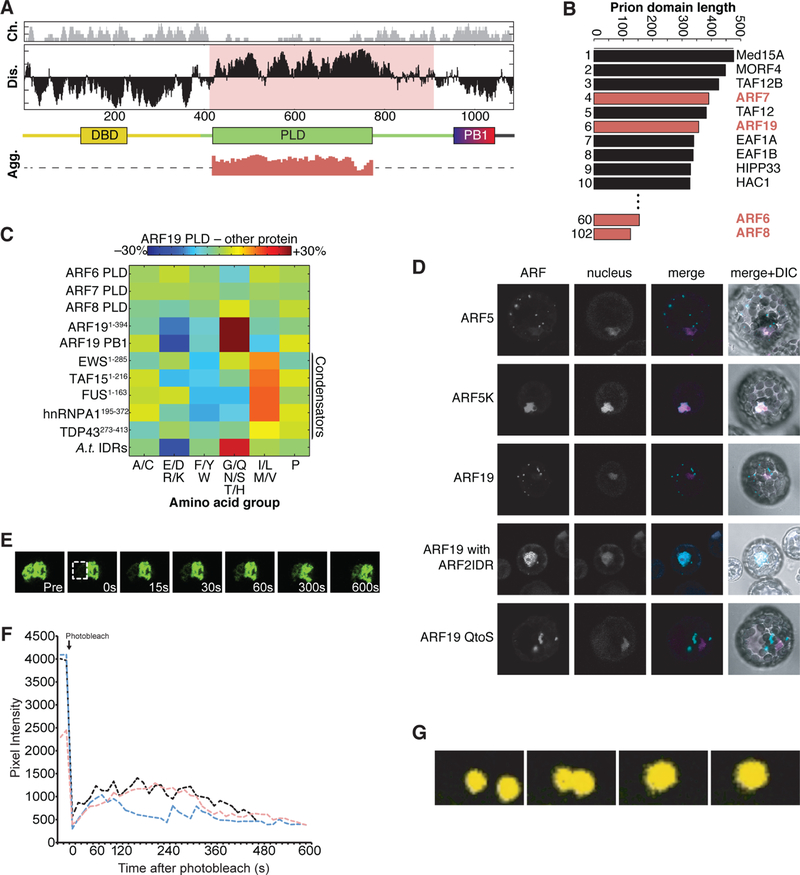
Figure 3. Sequence features of ARF19 and relationship to other condensate-forming proteins.
(A) Integrative linear sequence analysis showing local charge density (Ch.), local disorder propensity (Dis.) domain structure, and simulation-derived aggregation propensity of prion-like domain (PLD) alone (Agg.). The domain structure highlights the two known functional domains (DBD and PB1) and the PLD. The MR is predicted to be disordered and contains a PLD that has a strong propensity to self-associate.
(B) Proteome-wide analysis shows the top ten PLDs by length in the A. thaliana non-redundant proteome. See Dataset S1 for full list of Arabidopsis PRD-containing proteins.
(C) Comparative analysis of the ARF19 PLD composition in relation to different proteins. Each amino acid is assigned to one of six groups on the X-axis and the fractional difference of grouped amino acids between the ARF19 PLD and each sequence is shown. All sequence comparisons are in reference to the ARF19 PLD in relation to sequences from additional ARF PLDs, ARF19 DBD and PB1 domains, and PLD domains from known condensators. The average intrinsically disordered region (IDR) signature of disordered proteins in A. thaliana is also provided for reference.
(D) Confocal images of Wt (Col-0) protoplasts transfected with UBQ10:mVenus-ARF5, UBQ10:mVenus-ARF5K797A, UBQ10:mVenus-ARF19, UBQ10:mVenus-ARF19w/ARF2MR, and UBQ10:mVenus-ARF19QtoS. Co-transfection with a 35S:NLS-2*mCerulean marker was used to visualize nuclei.
(E) Time course of FRAP of condensate in UBQ10:YFP-ARF19 seedling. Dashed box indicates the initial photobleached region.
(F) Fluorescence recovery curve of three independent FRAP experiments on UBQ10:YFP-ARF19 condensates.
(G) Time course showing fusion of cytoplasmic condensates in root transition zone cells (containing both nuclear and condensate signal) in a seedling expressing ARF19:ARF19-Venus.
Given that the MR is necessary for assembly formation, we sought to identify additional sequence features that might explain the molecular origins of this behavior. Prion-like domains (PLDs) are often associated with the formation of biomolecular condensates as phase separation and gelation drivers, both in vitro and in vivo (Altmeyer et al., 2015; Molliex et al., 2015; Patel et al., 2015; Wang et al., 2018). We emphasize that the name “prion-like domain” is used here pure ly with respect to the sequence features used to identify the domain and does not mean this domain confers classical prion-like behavior (non-Mendelian protein-based inheritance). The ability of PLDs to drive condensate formation originates from the intrinsic behavior imparted by their amino-acid composition. PLDs are typically depleted of charged residues, enriched in polar residues, and are low-complexity domains. These features are largely consistent with our preliminary sequence analysis. Accordingly, we deployed the PLAAC algorithm (Lancaster et al., 2014) and found that it predicts that the MR of ARF19 encompasses a large putative PLD (Figure S3). Moreover, a proteome-wide analysis of non-redundant Arabidopsis proteins (15,350 proteins) uncovered only 148 proteins with PLDs greater than 100 residues in length (Dataset S1). ARF7 and ARF19 were proteins with the 4th and 6th longest PLDs on the list (Figure 3B). No PLD is identified in the ARF5 MR, but this is largely a function of the default parameters used for PLAAC – the ARF5 MR shows highly similar sequence features to other Class-A ARF MRs. No Class-B or Class-C ARF proteins, historically defined as transcriptional repressors, were identified (Dataset S1).
To further interrogate the molecular features of the PLD, we used all-atom simulations to assess the complete set of pairwise interactions of all 6-residue sub-fragments extracted from the PLD (1770 unique pairwise interactions). This provides a qualitative measure of the sequence-specific tendency of the PLD to engage in inter- and intra-molecular interactions. We found that this region shows a uniformly high propensity to undergo self-association (Figures 3A and S4).
Although PLDs and other disordered regions often display low sequence conservation as assessed by traditional sequence alignments, general sequence features such as amino acid composition are often conserved (Riback et al., 2017; Zarin et al., 2017). We compared the amino acid composition of the ARF19 PLD with PLDs from ARF6–8, all disordered regions from Arabidopsis, and five PLDs from proteins that are known to drive biomolecular condensate formation - TDP43, FUS, hnRNPA1, TAF15, and EWS (Altmeyer et al., 2015; Molliex et al., 2015; Patel et al., 2015; Wang et al., 2018). We observed nearly identical sequence composition among the ARF PLDs, which was distinct from that of the ARF19 DBD and PB1 domain (Figure 3C). Moreover, whereas the ARF19 PLD composition is unlike the average Arabidopsis IDR composition, it is similar to PLD scaffolds that drive biomolecular condensate formation, with distinct but important differences (Figure 3C).
The ARF PLDs are enriched in hydrophobic residues compared to four out of five sequences of established condensate formation drivers (Figure 3C). There is a correlation between hydrophobicity in low-complexity domains and the driving forces for self-assembly (Dao et al., 2018; Riback et al., 2017). Our analysis shows that the MRs of Class-A ARFs contain PLDs with enhanced hydrophobicity, consistent with the possibility that the ARF MR promotes biomolecular condensate formation via homotypic and heterotypic interactions with similar low-complexity sequences. Although ARF5 does not formally contain a PLD, it shows the same sequence features as the other activating ARFs, while repressive ARFs are less hydrophobic and more highly charged.
To test predictions regarding sequence features within the MR that are likely to drive ARF condensate formation, ARF variants were transiently expressed in Arabidopsis mesophyll protoplasts. Consistent with in planta data, UBQ10:mVenus-ARF19 localized to cytoplasmic condensates in this system (Figure 3D). ARF5, another activating ARF, was also expressed and localized to cytoplasmic condensates. Importantly, mutation of the conserved PB1 lysine in ARF5 resulted in loss of condensate formation and solely displayed nuclear localization (Figure 3D). The ARF2 protein, considered a Class-B repressing ARF, displays nuclear localization in all examined cell types in planta (Figure 1A). Further, bioinformatics analysis suggests that only the MRs of Class-A activating ARFs contain PLDs with the necessary properties for condensate formation. To test the hypothesis that MR properties confer, or at least regulate, the capacity for condensate formation, the ARF19 MR was replaced with the ARF2 MR. This MR swap was sufficient to disrupt ARF19 condensate formation (Figure 3D), suggesting that ARF19 MR sequence composition facilitates condensate formation.
Differences in the enrichment of glutamine versus serine residues within the MR have historically been used to define “activator” (Class -A) versus “repressor” (Class-B and C) ARFs. Accordingly, we speculated that the glutamine residues might be necessary for condensate formation. To test this hypothesis, we designed a mutant in which all glutamine residues in the ARF19 IDR were mutated to serine. Surprisingly this Q-to-S mutant did not prevent cytoplasmic condensate formation, but it did lead to condensates with altered morphology (Figure 3D). This result is congruent with a similar mutational approach in the PLD of the protein FUS, in which a Q-to-G mutant did not alter condensate formation but did change their material properties (Wang et al., 2018). The changes in condensate morphology indicate that these residues are important contributors to overall condensate structure, but suggest that there are additional sequence features that contribute as drivers of condensate formation. Drawing on the stickers-and-spacers formalism adapted recently by Wang et al., we propose that the glutamine and serine residues play the role of spacers whose interactions, in the form of hydrogen bonding contributions, influence the material properties and condensate morphologies. The stickers within disordered MR sequences that discriminate between activator and repressor ARFs are yet to be identified, although hydrophobicity appears to be one of the crucial discriminating features.
ARF PB1 multimerization as a means to generate the multivalence required for condensate formation
The bipartite structured PB1 domain and the intrinsically disordered MR are important for formation of cytoplasmic ARF protein assemblies, but how do these domains promote assembly? Concatenation of the asymmetric unit in the ARF7 PB1 domain crystal structure (Korasick et al., 2014) reveals a putative PB1 domain filament (Figure 2B). This filament has an open twisted helical structure, with macroscopic features similar to a previously solved PB1 filament (p62/SQSTM1) (Ciuffa et al., 2015) for which the full-length protein has been shown to form condensates in vivo (Sun et al., 2018). In this filament, the MR would extend outward, with the PB1 domains generating an extended oligomer with a curved helix topology. Sequence analysis suggests that the region between the PLD and PB1 domain is more charged and less hydrophobic, which could provide a flexible linker / spacer enabling solubility between the PB1 domain and the PLD that drives self-associations through its hydrophobic residues. The variable region between the two PB1 domain halves and the nature of the electrostatic interactions may engender the filament with flexibility.
Consistent with a filament model, the cytoplasmic ARF19 protein assemblies encompass a mixture of oligomeric states and adopt complex morphologies (Figure 1D and 2E) that move within the cytoplasm (Movie S1). The assemblies also display apparent void volumes, suggesting that they may either interact with or envelop other macromolecules or assemblies (Figure 1D). Conversely, ARF7 protein assemblies are smaller and adopt spherical morphologies (Figure 1D), consistent with the possibility that the ARF7 protein assembly is less rigid and suggesting that ARF19 and ARF7 may accumulate in distinct protein assemblies.
To better understand the material properties of these condensates, we performed fluorescence recovery after photobleaching (FRAP) experiments. Liquid-like condensates display rapid internal reorganization and a corresponding rapid fluorescent recovery on the order of seconds, whereas more solid-like condensates exhibit little or no recovery. After photobleaching YFP-ARF19 condensates from mature sections of the root, a limited amount of recovery occurs within 180 s (Figures 3E and F). This slow recovery rate suggests that these condensates have more solid-like properties, with the small amount of recovery likely being contributed to by exchange and/or addition of monomeric and dimeric protein at the condensate periphery (Figure 2E). In addition, YFP-ARF19 condensates in mature root tissues fuse (Movie S1) while retaining the distinct ultrastructure of each pre-fusion body; in contrast, the “younger” mVenus-ARF19 condensates in root cells at the zone transitioning from primarily nuclear signal to the zone containing condensates form larger spherical bodies upon fusion (Figure 3G), consistent with the possibility that the material properties of condensates change over time.
From these data, a model emerges for condensate formation. PB1 domain association forms a semi-stable filament, which generates the multivalence of MR bristles that enable lateral associations. The MR may interact with other proteins, such as other PLD-containing proteins or other activating ARF filaments, to act as an extended supramolecular scaffold for higher-order organization. In our model, cytoplasmic sequestration by biomolecular condensates should reduce the effective concentration of soluble activating ARFs, which in turn is expected to have a significant impact on transcriptional behavior in the form of reduced auxin sensitivity. To test this hypothesis, we next examined how cytoplasmic ARF assembly/condensate formation impacts global gene expression patterns.
Nucleo-cytoplasmic partitioning is sufficient to regulate auxin transcriptional responsiveness
Auxin is a critical and widely dispersed plant hormone, yet it imparts distinct functions on different plant tissues. Reporters driven by the synthetic auxin response element DR5 are commonly used to examine auxin-induced gene transcription; however, these reporters are not equally induced by auxin in every cell type (Figure 4A and S5). This unequal DR5 induction suggests the possibility that not every cell responds equally to auxin. In roots, only actively growing cells near the root tip displayed an increase in auxin-responsive DR5 activation following a 2-hour auxin treatment (Figures 4A and S5). This DR5 auxin-responsive region corresponds to the same tissues in which ARF7 and ARF19 were nuclear-localized (Figure 1). DR5 activation was not observed in tissues in which the ARF7 and ARF19 proteins were primarily confined in cytoplasmic assemblies in the upper root (Figure 4A). To determine whether the lack of DR5 transcriptional activation could be explained by differences in auxin perception and subsequent Aux/IAA protein degradation, we examined auxin-induced degradation of the DII-VENUS and mDII-VENUS reporters of Aux/IAA repressor protein stability (Brunoud et al., 2012). We found that in both root tip and upper root cells, a 2-hour auxin treatment promoted DII-VENUS degradation (Figure 4A), demonstrating that cells in which ARF7 and ARF19 localized to cytoplasmic assemblies retain auxin perception and subsequent repressor degradation. Thus, it is likely that DR5 induction differences between upper and lower root tissues are not caused by differences in auxin perception. Overall, tissue-specific cytoplasmic partitioning of ARF7 and ARF19 correlates with dampened auxin transcriptional responses and is consistent with a model in which activating ARF localization regulates auxin responsiveness.
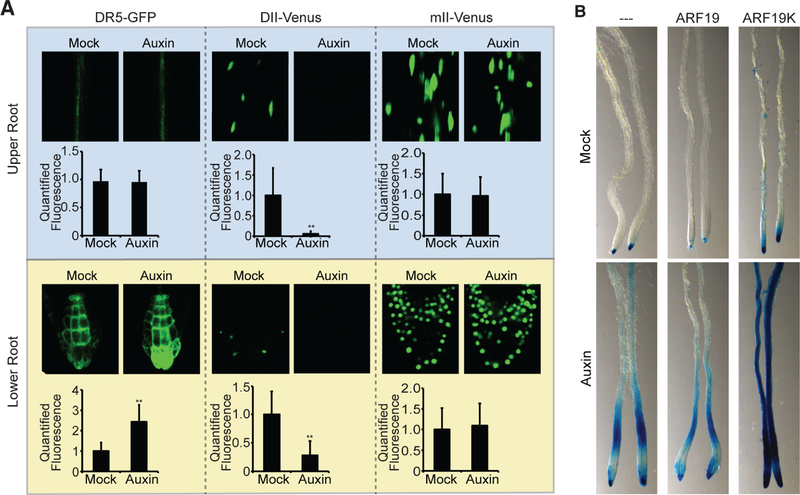
Figure 4. Auxin responsiveness corresponds with ARF nucleo-cytoplasmic Partitioning.
(A) Fluorescence microscopy imaged from 5d-old DR5-GFP (Ottenschläger et al., 2003; Sabatini et al., 1999), DII-Venus (Brunoud et al., 2012), and mII-Venus (Brunoud et al., 2012) ± 2-hour treatment of 10 µM IAA. Images were taken and quantified from upper root and root tip sections (+SD; n ≥ 20).
(B) Microscopy images of primary roots from 8d-old Col-0 (---; Wt), Col-0 carrying UBQ10:YFP-ARF19 (ARF19), and Col-0 carrying UBQ10:YFP-ARF19K962A (ARF19K) with the DR5:GUS reporter (Ulmasov et al., 1997) that were treated with a mock (Ethanol) or 10 µM IAA treatment for 2 hours prior to staining.
A longstanding question in plant biology is how a seemingly simple signal transduction pathway precisely controls and tunes auxin output to regulate a wide array of developmental processes. Our observation that cytoplasmic ARF condensates form in tissues displaying dampened auxin responses led us to hypothesize that ARF nuclear-cytoplasmic partitioning acts as a mechanism to control auxin responsiveness in specific tissues. To interrogate whether the differences in DR5 activation could be caused by differential ARF nucleo-cytoplasmic partitioning, we crossed our UBQ10:YFP-ARF19 line (ARF19) and our UBQ10:YFP-ARF19K962A line (ARF19K) to DR5:GUS (Ulmasov et al., 1997). In the line expressing YFP-ARF19, in which ARF19 protein is primarily nuclear in the meristematic region and in protein condensates in differentiated tissues, auxin-induced DR5:GUS signal looked similar to wild type (Figure 4B), with maximal activity in the meristem and attenuated activity in the upper parts of the root. In contrast, auxin-induced DR5:GUS activity in the line expressing YFP-ARF19K, in which the ARF19K protein is nuclear in every cell type, was expanded into the upper, mature region of the root (Figure 4B).
To further test our hypothesis that ARF19 cytoplasmic sequestration in the upper root could lead to attenuated auxin responsiveness, we examined global auxin-responsive transcript accumulation in wild type (Wt), the UBQ10:YFP-ARF19 line (ARF19), in which YFP-ARF19 localization varied by developmental stage in the root (Figure 1A), and the UBQ10:YFP-ARF19K962A line (ARF19K), in which YFP-ARF19K962A localized constitutively to the nucleus at all examined root developmental stages (Figure 2). In the absence of auxin treatment, transcript accumulation differences between the ARF19 line and wild type were minimal (Figure 5A). Conversely, in the absence of auxin, the constitutively nuclear ARF19K line displayed over 4,500 genes with differential expression when compared to either Wt or the ARF19 line (Figures 5A and 5B). Many genes whose transcripts displayed altered accumulation in the ARF19K line were also regulated by auxin treatment in Wt (Figure 5B), including members of the LBD and GH3 gene families (Table 1, Supplemental Datasets), suggesting that auxin responses are basally elevated in the ARF19K line.
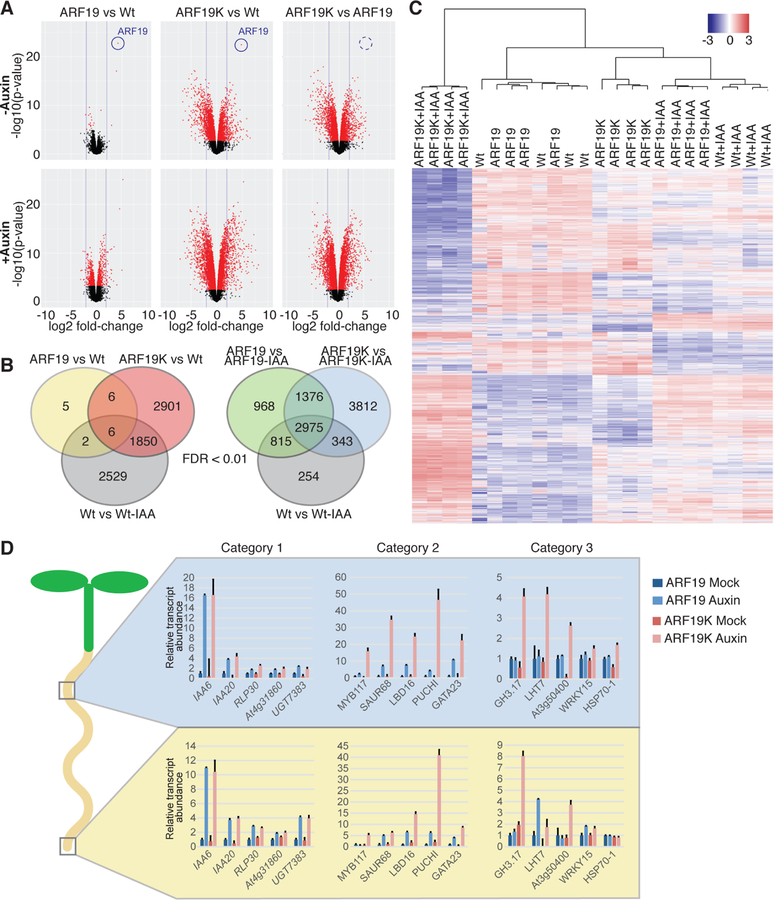
Figure 5. Disruption of ARF condensate formation results in an altered transcriptional landscape and morphological defects.
(A) Volcano plots displaying pairwise transcript accumulation differences between UBQ10:YFP-ARF19 (ARF19) and Wt (Col-0), between UBQ10:YFP-ARF19K962A (ARF19K) and Wt, and between ARF19K and ARF19 lines, each ± 10 µM IAA treatment. Transcripts whose accumulation differences are significant (FDR ≤ 0.01) are represented in red.
(B) Venn diagrams showing the overlap between the data sets of differentially expressed genes (FDR <0.01).
(C) Hierarchical clustering of genes displaying differential expression among samples (FDR <0.01).
(D) Relative transcript abundance (+SE; n = 3) of selected genes obtained by Nanostring analysis on upper root and lower root sections from 7d-old mock and auxin-treated UBQ10:YFP-ARF19 and UBQ10:YFP-ARF19K962A seedlings. Category 1 – auxin responsive in upper and lower root of ARF19 and ARF19K, Category 2 – auxin responsive in upper and lower root of ARF19 and ARF19K with a higher amplitude of change in auxin-treated ARF19K, Category 3 – auxin responsive only in the lower root of ARF19, but in both the upper and lower root of ARF19K + auxin responsive in only ARF19K.
See Figures S6 and S7 for RNASeq quality assessments. See Dataset S2 for pairwise comparisons of mock-treated samples, Dataset S3 for pairwise comparisons of auxin-treated samples, and Dataset S4 for comparisons between mock and auxin treatment within each line.
Table 1.
Selected genes from RNASeq Datasets
| Gene | AGI | ARF19K-Wt | Wt | Mock vs IAA ARF19 | ARF19K |
|---|---|---|---|---|---|
| PUCHI | At5g18560 | −1.12 | 0.56 | 2.31 | 5.87 |
| CLE5 | At2g31083 | −2.37 | 0.47 | 1.62 | 5.81 |
| RGF9 | At5g64770 | −2.43 | 1.5 | 2.4 | 5.17 |
| ACS5 | At5g65800 | −2.43 | 0.53 | 1.88 | 7.27 |
| IAA20 | AT2G46990 | −0.55 | 2.73 | 2.96 | 4.8 |
| CKX3 | At5g56970 | 0.81 | 0.32 | 3.02 | 5.21 |
| SAUR68 | At1g29490 | −2.56 | 2.98 | 4.55 | 6.37 |
| ABCG40 | At1g15520 | 0.86 | 1.74 | 2.49 | 5.76 |
| MYB105 | At1g69560 | −2.32 | −0.59 | 1.36 | 5.42 |
| SMXL8 | At2g40130 | −1.2 | −0.82 | 1.78 | 4.06 |
| LBD16 | AT2G42430 | 2.5 | 3.2 | 3.1 | 1.7 |
| GH3.5 | AT4G27260 | 2.4 | 2.9 | 4.38 | 3.2 |
| LBD17 | AT2G42440 | 2.2 | 3.5 | 4.19 | 2.6 |
| PIN5 | AT5G16530 | −4.3 | −1.7 | −2.45 | 2.23 |
| NPF2.4 | AT3G45700 | −3.5 | −2.4 | −3.12 | −0.9 |
| YUC2 | AT4G13260 | −3 | −1.14 | −2.32 | −3.5 |
| At3g50400 | −1.84 | 0.54 | 0.17 | 4.18 | |
| CYP76C5 | At1g33730 | 0.92 | −0.85 | 0.79 | 5.07 |
| GH3.17 | At1g28130 | 1.16 | 0.37 | 0.24 | 1.2 |
| RAS1 | At1g09950 | 0.37 | 0.6 | 0.24 | 2 |
| MYB117 | At1g26780 | −0.46 | −0.22 | 0.51 | 4.37 |
| KAT1 | At1g04710 | 0.08 | −0.13 | −0.04 | 4.01 |
| At2g40250 | 0.93 | −0.22 | −0.33 | 3.99 | |
| BDG1 | At1g64670 | −1.06 | 0.42 | 0.08 | 1.88 |
| ICS2 | At1g18870 | −1.34 | −0.2 | −0.09 | 3.9 |
| At3g49270 | 2.86 | −0.04 | −0.09 | 3.82 | |
| At4g31860 | 0.54 | 1.65 | 1.71 | 1.56 | |
| LHT7 | AT4G35180 | 0.99 | 2.48 | 2.29 | 2.39 |
| PLP7 | AT3G54950 | −0.34 | 1.56 | 1.78 | 1.87 |
| WRKY15 | AT2G23320 | 0.69 | 1.41 | 1.8 | 1.52 |
| NACO53 | AT3G10500 | 0.51 | 1.74 | 1.55 | 1.44 |
| HMA5 | AT1G63440 | −0.27 | 1.75 | 1.99 | 1.74 |
| RLP30 | AT3G05360 | 0.95 | 1.99 | 1.83 | 1.73 |
| PLA-1(BETA)2 | AT4G16820 | 1.99 | 3.31 | 3.47 | 3.51 |
| PUB36 | AT3G61390 | 0.05 | 2.86 | 2.68 | 2.66 |
| UGT73B3 | AT4G34131 | 0.14 | 4.02 | 3.77 | 4.13 |
| WOX11 | AT3G03660 | 2.47 | 3.27 | 4.37 | 3.03 |
| LBD18 | AT2G45420 | 2.33 | 1.65 | 2.67 | 1.21 |
| LBD19 | AT2G45410 | 2.25 | −0.68 | 3.75 | 3.29 |
| IAA6 | AT1G52830 | 0.09 | 3.07 | 4.14 | 5.64 |
| GATA23 | AT5G26930 | 0.03 | 2.31 | 2.65 | 4 |
| At4g30180 | −1.25 | 0.34 | 0.59 | 5.87 | |
| At5g17740 | 1.89 | 0.82 | 1.48 | 5.19 | |
| At1g66610 | 0.99 | 0.57 | 0.43 | 5.01 | |
| TMK2 | At1g24650 | 1.09 | −0.06 | 1.82 | 5.00 |
| HSP26.5 | At1g52560 | 3.12 | 0.44 | 1.59 | 4.57 |
| At1g11112 | −0.58 | 0.16 | 0.48 | 4.46 |
Consistent with the ARF19 role in mediating auxin responses, UBQ10-driven expression of YFP-ARF19 resulted in mildly elevated auxin-regulated transcriptional responses compared to Wt when treated with auxin (Figure 5A). Strikingly, auxin treatment of the ARF19K line resulted in dramatically altered auxin-responsive transcript accumulation compared to auxin-treated Wt or ARF19 line samples (Figures 5A, 5B and 5C), with over 3,800 genes displaying auxin-responsive transcript accumulation in ARF19K, but not Wt or ARF19 (Figures 5B and 5C). Further, the amplitude of auxin-responsive gene expression was increased in the ARF19K line (Figure 5C, Table 1), including several genes in the ARF7 or ARF19 gene regulatory network (Lavenus et al., 2015). Because the PB1 domain is also important for interactions between ARFs and the Aux/IAA repressors, it is possible that some changes in transcript accumulation observed in the ARF19K line are caused by disruption of repressor interaction. We would expect that complete uncoupling of this interaction would result in basal altered regulation of auxin-responsive genes as well as loss of altered transcript accumulation in response to auxin treatment. We do observe some auxin-responsive genes displaying basal accumulation consistent with potential loss of Aux/IAA repression (Figure 5B; Table 1); however, the large number of auxin-responsive genes in the ARF19K line combined with the elevated amplitude of auxin response of these transcripts in the ARF19K line suggests that Aux/IAAs can bind and repress ARF19K962A activity.
The results obtained from global transcript profiling prompted us to further explore changes in transcript level of a subset of genes in upper root and root tip sections in response to auxin treatment. 7d-old ARF19 and ARF19K seedlings were treated with either a mock or an auxin treatment for two hours prior to excision a 2 mm upper root section and 2 mm root tip sections (Figure 5D). Examined transcripts were grouped into three categories: 1) transcripts that were indistinguishable in the ARF19 and ARF19K lines, 2) transcripts displaying increased amplitude of auxin response in the ARF19K line in the upper root section, and 3) transcripts that displayed auxin induction in the upper roots only of the ARF19K, but not the ARF19 line (Figure 5D). These data suggest that for at least some transcripts, auxin responsiveness in the upper root is possible only when the ARF19 protein is constitutively nuclear. There are 23 ARF transcription factor family members in Arabidopsis; these additional ARFs likely also contribute to auxin-responsive changes in gene expression in the upper root sections for Category 1 and Category 2 transcripts. Similar to these Class 3 genes, DR5 activity in the upper root is also greatly affected in ARF19K (Figure 4B), raising the possibility that DR5 is more reflective of ARF19 (and possibly ARF7) activity than that of other ARFs. Overall, these data support a model in which a subset of auxin pathway gene targets are affected by the nucleo-cytoplasmic partitioning of the ARF19 protein.
The major shift in transcript accumulation observed in the ARF19K line, combined with the increased amplitude of auxin responses, suggests that ARF nuclear-cytoplasmic partitioning acts as a mechanism to control auxin responsiveness. We therefore examined the morphological effects of expressing YFP-ARF19 (ARF19) and YFP-ARF19K962A (ARF19K) in seedlings. ARF19K lines demonstrated morphological defects associated with altered auxin transcriptional responsiveness, including reduced root length, reduced cotyledon area, and increased root hair lengths (Figures 6A–D). The phenotypes observed in the ARF19K lines are commonly associated with changes in auxin response and demonstrate that disruption of ARF19 cytoplasmic condensate formation is sufficient to alter normal plant growth and development. Our data support a model in which cytoplasmic condensates control the nuclear-cytoplasmic partitioning of activating ARFs, which in turn tunes the cellular transcriptional state.
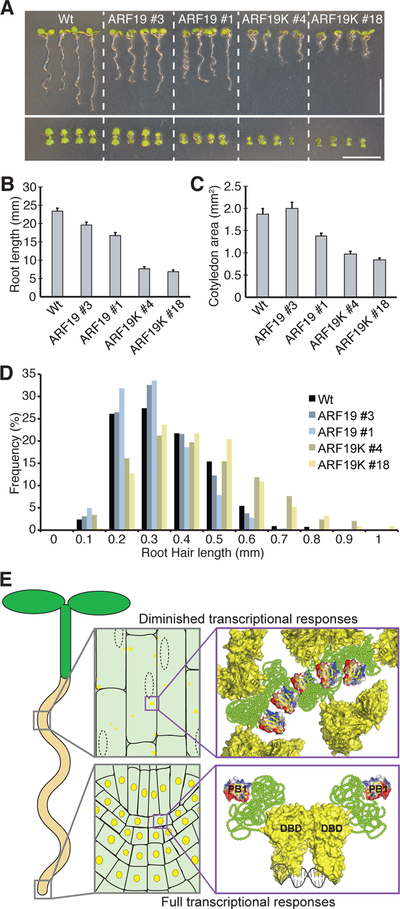
Figure 6. ARF nucleo-cytoplasmic partitioning regulates auxin responsiveness and disruption of ARF condensate formation results in morphological defects.
(A) Photos of 8d-old seedlings from wild-type (Wt; Col-0), two independent UBQ10:YFP-ARF19 (ARF19) lines, and two independent UBQ10:YFP-ARF19K962A (ARF19K) lines. Bar = 1 cm.
(B) Mean root lengths (+SE; n ≥ 15) of 8d-old seedlings.
(C) Mean cotyledon area (+SE; n ≥ 30) of 8d-old seedlings.
(D) Histograms of root hair lengths from 6d-old seedlings.
(E) In cells displaying dampened auxin responsiveness, activating ARF proteins localize to cytoplasmic biomolecular assemblies; protein assembly formation is driven by the PB1 domain and MR IDR. Conversely, ARF proteins localize to the nucleus in actively growing, highly auxin-responsive tissues, such as the root tip. Localization of these transcription factors to the cytoplasm (in cells with low auxin responsiveness) or nucleus (in cells with high auxin responsiveness) may confer cellular competence for auxin response.
DISCUSSION
Abrogation of ARF assembly formation results in massive gene transcription changes as well as gross morphological defects (Figures 6A–D), suggesting the importance of ARF condensates in regulating normal plant growth and development. This result is consistent with observed variations in ARF localization corresponding to auxin transcriptional responsiveness (Figures 1 and 4). Together, these data provide evidence for a model in which activating ARF proteins are present in the nucleus of actively growing tissues, but are sequestered in cytoplasmic biomolecular condensates in tissues where the plant no longer needs to respond to auxin (Figure 6E). This model also implies that regulatory mechanisms exist in the nucleus to prevent ARF protein condensation. Because auxin is transported from aerial tissues to the root tip (Woodward and Bartel, 2005), these regulatory mechanisms could provide the means to reduce auxin responses in auxin-transmitting tissues in which auxin responses would be undesirable. Importantly, assuming condensates are reversible, the release of ARFs could result in their localization to the nucleus in circumstances when these tissues need to respond to auxin, such as during the development of lateral root primordia or in response to various stimuli. In this way, all plants cells would be able to convert back into auxin-responsive tissues, conferring morphological plasticity to the plant.
Recent studies have shed light on the properties of biomolecular condensates and their functional roles in biological processes (Altmeyer et al., 2015; Boke et al., 2016; Cho et al., 2018; Freeman Rosenzweig et al., 2017; Kato et al., 2012; Kroschwald et al., 2018; Molliex et al., 2015; Patel et al., 2015; Riback et al., 2017; Sabari et al., 2018; Wang et al., 2018; Woodruff et al., 2017). Our results illustrate a clear link between condensate formation and biological function. ARF19 sequestration through condensate formation leads to a shift in the transcriptional landscape (Figure 5). Further, ARF condensate morphology (Figure 1D) suggests the presence of additional components. Condensates formed through PLDs can recruit other PLD-containing proteins, although the rules governing recruitment specificity remain unclear (Kato et al., 2012; Lin et al., 2015; Wang et al., 2018). Given the extent of the changes in cell-wide transcriptional activity in ARF19K (Figure 5), one possible interpretation is that the ARF condensates sequester other PLD-containing proteins, thereby causing a fundamental shift in transcriptional regulation by sequestering these additional factors in the cytoplasm.
Recent interest on the role of condensate formation in transcriptional regulation (Boija et al., 2018; Cho et al., 2016; Cho et al., 2018; Chong et al., 2018; Hnisz et al., 2017; Lu et al., 2018; Sabari et al., 2018) have predominantly focused on condensates as drivers of recruitment of factors to distinct genomic loci. Our work here demonstrates an orthogonal but complementary route for transcriptional regulation in that key transcription factors can be kept away from the nucleus by sequestration into cytoplasmic condensates. The solid-like nature of the ARF condensates is fully consistent with a sequestration model for transcriptional reprogramming. The use of solid-like reversible assemblies as a means of spatial and temporal regulation has been invoked in various contexts (Boke et al., 2016; Olzscha et al., 2011; Riback et al., 2017; Si et al., 2003). However, we cannot rule out the possibility that the large IDRs associated with activating ARFs may also contribute to transcriptional regulation by facilitating recruitment and smaller-scale condensates at the sites of transcription (Boija et al., 2018; Cho et al., 2018; Sabari et al., 2018).
Our FCS data are inconsistent with a simple two-stage model for phase separation for condensate assembly (Posey et al., 2018). In such a model, the oligomeric state of an individual ARF protein would be expected to be monomeric (in the dilute phase) or fully networked (in the condensate). In contrast, the distribution of discrete oligomer sizes in a concentric radial manner is consistent with a model in which PB1-mediated polymerization facilitates the formation of higher-order oligomers, which are then capable of mediating condensate formation through the requisite multivalence. Indeed, our Numbers & Brightness analysis (Figures 2D and 2E) provides unique and convincing quantification of the non-stoichiometric nature of ARF assemblies, which are thought to be a defining hallmark of condensates. In our model the MR is necessary for condensate formation, but its driving force for assembly is relatively weak. This predicts that when expressed in isolation, the MR should form spherical condensates but only when highly over-expressed. We tested this hypothesis through overexpression of MR alone, and in agreement with our predictions, we observed small spherical assemblies (Figure S1B).
As a central regulator of multiple essential plant processes, the auxin response pathway requires unique and finely tuned auxin outputs. Our work identifies ARF nucleo-cytoplasmic partitioning as a mechanism to regulate cellular competence to respond to auxin. Formation of ARF assemblies mediated by PB1 domain interactions and the intrinsically disordered MR drives this partitioning. Importantly, activating ARFs localize to the nucleus in tissues with high auxin transcriptional responsiveness, but are sequestered in cytoplasmic condensates in tissues that are no longer actively growing (Figure 6E). These observations suggest that differential localization may confer cellular competence to respond to this critical hormone. The distinct developmental regulation of ARF cytoplasmic-nuclear partitioning suggests regulation of this process. We further postulate that ARF partitioning might provide a mechanism for other inputs (i.e., other hormones or environmental stimuli) to affect cellular auxin responsiveness. Further research will be necessary to determine the factors involved in regulating the formation and release of activating ARFs from cytoplasmic assemblies.
STAR Methods
CONTACT FOR REAGENT AND RESOURCE SHARING
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Lucia Strader (strader@wustl.edu).
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Arabidopsis
All Arabidopsis lines were in the Columbia (Col-0) background, which was used as the wild type (Wt). Seeds were surface-sterilized (Last and Fink, 1988), suspended in 0.1% agar, and stratified for 2 d at 4°C to promote uniform germination. After stratification, seeds were plated on plant nutrient (PN) media (Haughn and Somerville, 1986) solidified with 0.6% agar and supplemented with 0.5% (w/v) sucrose (PNS) at 22°C under continuous illumination.
METHOD DETAILS
Plant Growth and phenotypic assays
To examine root elongation in Col-0 and transgenic lines, root lengths were measured from seedlings atop horizontally-incubated media after 7 days of growth at 22°C under continuous illumination.
To examine cotyledon size, cotyledons were excised from 7d-old seedlings, mounted, imaged, and area measured using Image J software (NIH).
To examine root hair lengths, vertically-grown 6d-old seedlings were imaged using a dissecting microscope and root hair lengths from the 4-mm root section closest to the root-shoot junction were measured using Image J software.
Vector construction and plant transformation
ARF19 was amplified from cDNA using Pfx Platinum (Life Technologies) and 5`-CACCCATATGAAAGCTCCATCAAATGG-3` and 5`-CTCGAGCTATCTGTTGAAAGAAGCTGC-3`. The resultant PCR product was cloned into pENTR/D-TOPO (Life Technologies) to create pENTR-ARF19. The ARF19 PB1 domain was amplified from cDNA using 5`-CACCCATATGCGAACATATACAAAGGTTC-3` and 5`-CTCGAGCTATCTGTTGAAAGAAGCTGC-3` and the resultant PCR product cloned into pENTR/D-TOPO to create pENTR-ARF19PB1. The ARF19 MR and PB1 domain were amplified from cDNA using 5`-CACCCATATGCCTTGGATGGGAGAAGAC-3` and 5`-CTCGAGCTATCTGTTGAAAGAAGCTGC-3` and the resultant PCR product cloned into pENTR/D-TOPO to create pENTR-ARF19MR-PB1. ARF2 was amplified from cDNA using Pfx Platinum and 5`-CACCCATATGGCGAGTTCGGAGGTTTCAATG-3` and 5`-CTCGAGTTAAGAGTTCCCAGCGCTGG-3` and the resultant PCR product cloned into pENTR/D-TOPO to create pENTR-ARF2. Site-directed mutagenesis of pENTR-ARF19 was performed using the QuikChange Lightning Multi Site-Directed Mutagenesis Kit (Agilent) using 5`-CAACGAATGCGAACATATACAGCGGTTAGAGATCTTTCTCTC-3` and 5`-GAGAGAAAGATCTCTAACCGCTGTATATGTTCGCATTCGTTG-3` to create pENTR-ARF19K962A.
ARF19 and ARF2 cDNA and variants were recombined from the pENTR vectors into the UBQ10:YFP-GW plasmid (Michniewicz et al., 2015) using LR Clonase (Invitrogen). Recombinant plasmids were transformed into Agrobacterium tumefaciens strain GV3101 (Koncz and Schell, 1986) and used to transform Columbia (Col-0) plants via the floral dip method. Transformants were selected by surface sterilizing T1 seeds and plating on plant nutrient media [PN (Haughn and Somerville, 1986)] supplemented with 10µg/mL Basta. Subsequent generations were tested to identify lines homozygous for the transgene.
Protoplast vector construction, isolation, and transfection
ARF5 was amplified from cDNA using Pfx Platinum (Life Technologies) and 5’ – CACCGGTCGACGGATGATGGCTTC 3’ and 5’ – GCGGCCGTTATGAAACAGAAG 3’. The resultant PCR product was cloned into pENTR/D-TOPO (Life Technologies) to create pENTR-ARF5. Site-directed mutagenesis of pENTR-ARF5 was performed using the QuikChange Lightning Multi Site-Directed Mutagenesis Kit (Agilent) using 5’ – CGAACCTACACTGCGGTTCAAAAAACC 3’ and 5’-GGTTTTTTGAACCGCAGTGTAGGTTCG 3’ to create pENTR-ARF5K797A. The ARF2 and ARF19 Q to S IDRs were synthesized as gBlocks by Integrated DNA Technologies with a 5’ XmaI and 3’ SpeI restriction site. These gBlocks were cloned into pENTR/D-TOPO to create pENTR-ARF2 IDR and pENTR-ARF19 Q to S IDR. pENTR-ARF2 IDR, pENTR-ARF19 Q to S, and pENTR-ARF19 were digested with XmaI and SpeI and the ARF2 and ARF19 Q to S IDRs were ligated into ARF19 using the Quick Ligation Kit (NEB) to create pENTR-ARF19w/ARF2IDR and pENTR-ARF19 Q to S.
ARF5 and ARF19 cDNA and variants were recombined from the pENTR vectors into a UBQ10:mVenus-GW plasmid for transient expression in protoplasts. Protoplasts were isolated from 3–5 week-old Wt (Col-0) plants grown under long day conditions (16 hr light 8 hr dark) according to the Tape-Arabidopsis Sandwich method (Wu et al., 2009). Protoplasts were transfected as previously described (Wu et al., 2009) and incubated for 16 hr before imaging.
Microscopy
For fluorescence microscopy of plant lines, 5d-old seedlings were mounted in water under a coverslip and imaged though a x40 or x100 oil immersion lens and YFP filter set using a Leica DM6 B upright microscope.
For confocal images of plant lines, 5d-old seedlings were counterstained with propidum iodide (10µg/mL) and mounted in water for imaging through a 40× lens on a Zeiss LSM510 laser scanning confocal microscope equipped with a Meta detector. DR5-GFP, DII-Venus, and mII-Venus lines were treated with EtOH or 10 uM IAA for 2 hr prior to imaging. Changes in fluorescence were quantified using ImageJ.
For confocal images of protoplasts, protoplasts were placed in glass bottom dishes (Ted Pella) 16 hr post-transfection and imaged through a x60 oil immersion lens on a Nikon A1Si laser scanning confocal microscope.
For imaging of 35S:ARF7-HA seedlings, 5d-old seedlings were fixed as previously described (Sauer et al., 2006). Seedlings were then transferred to a 6-well plate and washed 3× with 1X PBS. 2% Driselase was added to wells and plates were incubated at 37°C for 60 min. Seedlings were washed 3× with 1X PBS and then incubated in 3% IGEPAL for 60 min at RT. Seedlings were washed 3× with 1X PBS and 5% BSA was added to each sample and incubated for 60 min at RT. The BSA was removed and HA-Tag (C29F4) Rabbit mAb antibody (Cell Signaling Technology) was added to wells and incubated at 4 °C O/N. Seedlings were washed 3× with 1X PBS before incubation with DAPI solution for 30 min at RT in the dark. Seedlings were washed 3× with 1X PBS and mounted on slides. Imaging was performed using a Nikon A1Si laser scanning confocal microscope.
FRAP
FRAP experiments were performed on condensates from the pUBQ10:YFP-ARF19 line using a Nikon A1Si laser scanning confocal microscope. Regions encompassing approximately half the condensate were chosen for initial photobleaching. The fluorescence recovery was imaged at 15s intervals for a total of 10 min.
Number and Brightness (N&B) analysis
Confocal images were collected using a Zeiss LSM 710. 3-day-old pARF7:ARF7g-YFP, UBQ10:YFP-ARF19, and UBQ10:YFP-ARF19K962A seedlings were counterstained with propidium iodide to visualize cell walls. 12-bit raster scans of a 256×256 pixel region of interest with a pixel size of 100 nm were acquired using a pixel dwell time of 8.19 µsec. We analyzed raster scan data using the SimFCS software (Clark and Sozzani, 2017a; Digman et al., 2005). Software parameters, specifically the S-factor, were calibrated using 35S:GFP and UBQ10:YFP lines. The region of monomer brightness was determined using the 35S:GFP and UBQ10:YFP lines by selecting all of the fluorescence-positive pixels with a cursor. For the ARF lines, a cursor of the appropriate size was placed at the monomer brightness determined using the monomeric controls. Then, it was assumed that the region at 2*monomer brightness was dimer, 3*monomer brightness trimer, and etc. To simplify the analysis, 3–10mers and >10mers were grouped together.
Proteome Wide Analysis
We analyzed the set of non-redundant (reviewed) proteins from A. thaliana (15350 proteins) as assembled in the UniProt proteome release UP000006548 (Feb 2017) (UniProt, 2015). We identified disordered regions across all proteins using the IUPred algorithm, with a disorder threshold of 0.4 and a gap-filling algorithm that removed short gaps (1–2 residues) between disordered or ordered regions (Dosztányi et al., 2005a, b; Holehouse et al., 2017). In previous work, we found broadly similar results when disordered prediction was performed using a consensus method when compared to using IUPred alone (Holehouse et al., 2017). We used the complete set of disordered regions from A. thaliana as a reference to compare properties obtained when analyzing the ARF proteins of interest (Figure S2).
We used the PLAAC algorithm to identify prion-like domains across the full A. thaliana proteome (Lancaster et al., 2014). As an example, the identification of the prion-like domain in ARF19 is shown in Figure S3. Although originally developed to identify regions that may facilitate non-genetic inheritance, the prion-like domains are frequently associated with proteins that drive intracellular phase separation (Banani et al., 2017; Mitrea and Kriwacki, 2016). All sequence analysis was performed using the localCIDER analysis package (Holehouse et al., 2017).
Simulations Methodology
To obtain a quantitative assessment of the local interaction behavior associated with the ARF19 prion-like domain we turned to all atom simulations. The ARF19 prion-like domain consists of 347 residues, meaning it is beyond the scope of all-atom simulations as a single sequence (for context, recent simulations of another prion-like domain were limited to a region of approximately 40 residues (Conicella et al., 2016). Instead of simulating the entire region as a single polypeptide, we subdivided the sequence into 59 non-overlapping 6-residue fragments. For each pair of unique fragments, we ran simulations (with 3 independent replicas) to assess the apparent affinity between those two fragments. Affinity was measured as the fraction of the simulation that two fragments were in direct contact (‘bound’) with one another. This yielded 1770 unique pairs, 5310 independent simulations, and approximately 16×109 Monte Carlo steps of simulation time. We also ran simulations for fragments of the same size that lacked any attractive interactions, to determine the expected affinity for a pair of fragments by random chance. This provides a normalization factor to take into account the concentration of fragments in the simulation. Using this approach allowed us to generate a 59×59 interaction matrix, quantifying the interaction between each pair of fragments (Figure S4A). Each column or row in this matrix reflects the affinity of a given fragment for all other fragments in the ARF19 prion-like domain. The mean affinity score for each fragment averaged over a sliding window of the two surrounding fragments was taken to provide an overall aggregation propensity (Figure S4B and Figure 3A). The use of a sliding window approach was applied as a qualitative measure to correct for chain connectivity, which is by definition lost in this fragmentation approach. In this way, we created a linear assessment of the tendency of the ARF19 prion-like domain to engage in interactions with itself in cis or in trans.
Simulation Details
All atom simulations were performed using the ABSINTH implicit solvent model and the CAMPARI Monte Carlo simulation engine (http://campari.sourceforge.net) (Vitalis and Pappu, 2009). The combination of ABSINTH and CAMPARI has been used to accurately describe proteins in solution to high-resolution accuracy for a number of diverse systems, including polar-rich sequences such as the ARF19 prion-like domain (Fuertes et al., 2017; Halfmann et al., 2011; Williamson et al., 2010). Conformational space is explored through a combination of backbone and sidechain torsional moves combined with rigid body rotation and translation. Each simulation was run for 1×106 steps of equilibration followed by 2 ×106 production steps, with conformations for analysis collected every 5000 steps. Three independent replicas per pair were run, and convergence assessed by comparing results from independent simulations. Simulation analysis was performed using MDTraj and CTraj (http://pappulab.wustl.edu/CTraj.html) (McGibbon et al., 2015).
Transcript analysis
For the RNASeq experiment, Col-0 (Wt), Col-0 carrying UBQ10:YFP-ARF19, and Col-0 carrying UBQ10:YFP-ARF19K962A were grown on PNS media for 5 d at 22 °C under continuous illumination. Seedlings were then transferred to liquid PN supplemented with either ethanol (Mock) or 10 µM IA A for 2h. These treatments were repeated in quadruplicate for each line. Total RNA was isolated from mock or IAA treated seedlings using the RNeasy Plant Mini Kit (Qiagen). Samples were then prepared for sequencing with the Epicentre Ribo-Zero Gold system according to manufacturer’s protocol, indexed, pooled, and sequenced across three 1×50bp lanes on a single flow-cell on an Illumina HiSeq 3000. RNA-seq reads were demultiplexed and aligned to the Ensembl release 23 (TAIR 10) top-level assembly with STAR version 2.0.4b. Gene counts were derived from the number of uniquely aligned unambiguous reads by Subread:featureCount version 1.4.5. Sequencing performance was assessed for total number of aligned reads, total number of uniquely aligned reads, and genes detected. The ribosomal fraction (Figure S6A), known junction saturation (Figure S6B), and read distribution over known gene models (Figure S6C) were quantified with RSeQC version 2.3.
All gene counts were then imported into the R/Bioconductor package EdgeR and TMM normalization size factors were calculated to adjust for samples for differences in library size. Ribosomal genes and genes not expressed in any sample greater than one count-per-million were excluded from further analysis. The TMM size factors and the matrix of counts were then imported into R/Bioconductor package Limma. Performance of the samples was assessed with a Spearman correlation matrix (Figure S7A) and Multi-Dimensional Scaling plot (Figure S7B). Weighted likelihoods based on the observed mean-variance relationship of every gene and sample were then calculated for all samples with the voomWithQualityWeights function and gene performance was assessed with plots of residual standard deviation of every gene to their average log-count with a robustly fitted trend line of the residuals. (Figure S7C). A generalized linear model was then created to test for gene level differential expression and the results were filtered for only those genes with Benjamini-Hochberg false-discovery rate adjusted p-values less than or equal to 0.01.
For each contrast extracted with Limma, global perturbations in known Gene Ontology (GO) terms and KEGG pathways were detected using the R/Bioconductor package GAGE to test for changes in expression of the reported log 2 fold-changes reported by Limma in each term versus the background log 2 fold-changes of all genes found outside the respective term. The R/Bioconductor package heatmap3 and Pathview was used to display heatmaps or annotated KEGG graphs across groups of samples for each GO term or KEGG pathway (respectively) with a Benjamini-Hochberg false-discovery rate adjusted p-value less than or equal to 0.05.
NanoStrings Analysis
For Nanostrings analysis, Col-0 carrying UBQ10:YFP-ARF19, and Col-0 carrying UBQ10:YFP-ARF19K962A were grown on PNS media for 5 d at 22 °C under continuous illumination. Seedlings were then transferred to liquid PN supplemented with either ethanol (Mock) or 10 µM IAA for 2h. 2 mm upper root and 2 mm root tip sections were dissected from the same 50 seedlings and pooled to generate a single upper root or lower root sample. These treatments were repeated in triplicate. Total RNA was isolated from samples using the RNeasy Plant Mini Kit (Qiagen). NanoString nCounter analysis was performed using 80 ng total RNA and carried out using the nCounter Digital Analyzer (NanoStrings Technologies; Seattle, WA) at the McDonnell Genome Institute at Washington University in St. Louis. In addition to 8 negative-control and 6 positive-control probes, two genes – TUB4 (AtAt5g44340) and PP2C (At1g13320) were used as references for normalization. Data was analyzed using the nSolver Analysis software.
QUANTIFICATION AND STATISTICAL ANALYSIS
Details of statistical analyses can be found in the figure legends. Microsoft Excel was used to perform statistical analyses for phenotypic assays. All error bars represent standard error of the mean. For transcript analysis, a false-discovery rate adjusted p value (FDR) of less than or equal to 0.01 was used to identify genes with significant transcript accumulation differences.
Supplementary Material
Dataset S1. Related to Figure 3. A. thaliana PLD-containing proteins
We used the PLAAC algorithm to identify prion-like domains across the full A. thaliana proteome (Lancaster et al., 2014). Analysis of PRD containing proteins using PANTHER (Mi et al., 2017) and GO Ontology (Ashburner et al., 2000) database (Released 2017–08-14) found statistical enrichment for proteins involved in mRNA binding (12.9 fold enrichment, p = 2.22E-24), transcription coactivator activity (22.95 fold enrichment, p = 5.16E-03) and transcription regulatory region DNA binding (5.15 fold enrichment, p = 9.13E-05).
Dataset S2. Related to Figure 5. Transcript accumulation differences under mock treatment.
Pairwise transcript accumulation differences between UBQ10:YFP-ARF19 (ARF19) and Wt (Col-0). Pairwise transcript accumulation differences between UBQ10:YFP-ARF19K962A (ARF19K) and Wt (Col-0). Pairwise transcript accumulation differences between UBQ10:YFP-ARF19K962A (ARF19K) and UBQ10:YFP-ARF19 (ARF19); FDR ≤ 0.01.
Dataset S3. Related to Figure 5. Transcript accumulation differences under auxin treatment
Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19 (ARF19) and auxin-treated Wt (Col-0). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19K962A (ARF19K) and auxin-treated Wt (Col-0). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19K962A (ARF19K) and auxin-treated UBQ10:YFP-ARF19 (ARF19); FDR ≤ 0.01.
Dataset S4. Related to Figure 5. Auxin-induced transcript accumulation differences in each line.
Pairwise transcript accumulation differences between auxin-treated Wt (Col-0) and mock-treated Wt (Col-0). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19 (ARF19) and mock-treated UBQ10:YFP-ARF19 (ARF19). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19K962A (ARF19K) and mock-treated UBQ10:YFP-ARF19K962A (ARF19K); FDR ≤ 0.01.
Movie S1. Related to Figure 1. ARF19 condensate movement and fusion.
Real time video of YFP-ARF19 in a cell from the upper portion of the root.
Figure S1. Related to Figures 1 and 2. Localization of ARF7-Venus and ARF19MR
(A) Confocal microscopy images of a 5d-old seedling expressing pARF7:ARF7g-Venus.
(B) Fluorescence microscopy image from the upper root regions of a 5d-old Arabdopsis seedling expressing UBQ10:YFP-ARF19MR. Scale bar = 25 µm.
Figure S2. Related to Figure 3. Comparative analysis of ARF19 prion-like domain
To quantitatively assess the sequence composition of the ARF19 PLD we compared it against all predicted disordered regions in the A. thaliana proteome. Across the 21123 discrete disordered regions the ARF19 prion-like domain was among the least charged (bottom 2%) and most enriched in the combined fraction of glutamine (Q), serine (S), asparagine (N) and leucine (L) (above 99.7% of all other IDRs).
Figure S3. Related to Figure 3. Identification of a prion-like domain in ARF19
Results of ARF19 analysis by the PLAAC algorithm. The prion-like domain is highlighted in yellow. Figure generated by the PLAAC webserver (http://plaac.wi.mit.edu).
Figure S4. Related to Figure 3. Results of simulation analysis
(A) The interaction matrix generated by simulations of all unique pairs of non-overlapping six residue fragments taken from the ARF19 prion-like domain. Attractive interaction reflects the affinity between two fragments, with any value greater than 0 reflecting a higher affinity than would expected for two fragments that did not experience any attractive interactions.
(B) The per-fragment average attractive interaction, written as an aggregation propensity. Each value uses an average sliding-window of the three surrounding fragments to capture chain connectivity. Although there is a drop towards the C-terminus of the prion-like domain, the majority of the region shows a strong tendency to undergo self-interaction. This panel is identical to the Agg. track in figure 3A, but provides quantification of the values.
Figure S5. Related to Figure 4. Auxin induction of DR5:GUS activity in root tissues
Primary roots of 8d-old DR5-GUS (Ulmasov et al., 1997) seedlings that were treated with the indicated concentration of IAA prior to staining to visualize GUS activity.
Figure S6. Related to Figure 4. RNA-seq Data Quality
(A) The summed counts-per-million of all Ensembl gene biotypes for every sample. All samples show that greater than 95% of all aligned reads aligned to protein-coding genes.
(B) Junction saturation curves for all samples across all Ensembl known exon-exon junctions randomly sampled from aligned BAM files in 5% intervals. The plateau at higher levels of sampling indicate that all samples were sequenced with enough depth to assay all expressed genes.
(C) Localization of aligned reads across a theoretical 100bp gene body. All samples have similar profiles indicating a negligible 5-prime bias with high levels of coverage from 5-prime to 3-prime.
Figure S7. Related to Figure 4. RNA-seq Sample Quality
(A) Matrix of Spearman correlations of all detected genes greater than 1 count-per-million in at least 3 samples relative to each other. Samples of the same condition have very high correlations as expected across the diagonal with no outliers.
(B) The quality of the samples in a multi-dimensional scaling plot of the leading log fold-changes. Samples for each condition cluster very tightly with each other with good cluster separation between samples of different conditions based on expression profiles.
(C) Scatter plot of the empirically derived fitted and trended mean-variance relationship across all genes and a bar plot depicting the sample specific weight for statistical model fitting and differential expression analysis.
KEY RESOURCES TABLE
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Chemicals, Peptides, and Recombinant Proteins | ||
| Propidium Iodide | Invitrogen | P21493 |
| Indole-3-Acetic Acid (IAA) | Sigma-Aldrich | I2886 |
| Basta | GoldBio | P-165–1 |
| DAPI | Invitrogen | D21490 |
| HA-Tag (C29F4) Rabbit mAb | Cell Signaling Technologies | 3724 |
| Critical Commercial Assays | ||
| RNeasy Plant Mini Kit | Qiagen | 74904 |
| Ribo-Zero Gold Kit | Epicentre | N/A |
| QuikChange Lightning Multi Site-Directed Mutagenesis Kit | Agilent | 210518 |
| XT Codeset Gene Expression Assay | Nanostrings | |
| Deposited Data | ||
| RNASeq data | This study | GEO: GSE104385 |
| Experimental Models: Organisms/Strains | ||
| Arabidopsis: pUBQ10:YFP-ARF19 | This study | N/A |
| Arabidopsis: pUBQ10:YFP-ARF19K962A | This study | N/A |
| Arabidopsis: pUBQ10:YFP-ARF19PB1 | This study | N/A |
| Arabidopsis: pUBQ10:YFP-ARF19MR+PB1 | This study | N/A |
| Arabidopsis: p35S:ARF7-HA | Cho et al., 2014 | N/A |
| Arabidopsis: p35S:ARF7-GFP | Okushima et al., 2007 | N/A |
| Arabidopsis: pARF7:gARF7-YFP | This study | N/A |
| Arabidopsis: pUBQ10:YFP-ARF2 | This study | N/A |
| Arabidopsis: DR5-GFP | Sabatini et al., 1999 | N/A |
| Arabidopsis: DR5-GUS | Ulmasov et al., 1997 | N/A |
| Arabidopsis: DII-Venus | Brunoud et al., 2012 | N/A |
| Arabidopsis: mII-Venus | Brunoud et al., 2012 | N/A |
| Oligonucleotides | ||
| Primers used in this study, see Method Details | This study | N/A |
| Recombinant DNA | ||
| pUBQ10:YFP-GW | Michniewiczet al., 2015 | ABRC stock #CD3–1948 |
| cDNA C105352 (ARF19) | Arabidopsis Biological Resource Center (ABRC) | AY669794 |
| cDNA U12525 (ARF2) | Arabidopsis Biological Resource Center (ABRC) | BT001072 |
| cDNA U79557 (ARF5) | Arabidopsis Biological Resource Center (ABRC) | 2708483 |
| Software and Algorithms | ||
| ImageJ | NIH | https://imagej.nih.gov/ij/ |
| PLAAC algorithm | Lancaster et al., 2014 | http://plaac.wi.mit.edu |
| IUPred algorithm | Dosztányi et al., 2005 | http://iupred.enzim.hu |
| CAMPARI Monte Carlo simulation engine | Vitalis & Pappu, 2009 | http://campari.sourceforge.net) |
| MDTraj and CTraj | McGibbon et al., 2015 | http://pappulab.wustl.edu/CTraj.html |
| Spliced Transcripts Alignment to a Reference (STAR) | Dobin et al., 2012 | http://code.google.com/p/rna-star/ |
| RSeQC | Wang & Wang, 2012 | http://code.google.com/p/rseqc |
| R/Bioconductor | Gentleman et al., 2004 | https://www.bioconductor.org |
| nSolver Analysis software | Nanostrings | https://www.nanostring.com/products/analysis-software/nsolver |
| SimFCS | Laboratory for Fluorescence Dynamics | https://www.lfd.uci.edu/globals/ |
| Other | ||
HIGHLIGHTS.
Condensation of transcription factors to attenuate hormone response
Mechanism for nucleo-cytoplasmic partitioning of transcription factors
ACKNOWLEDGMENTS
We thank Hongwei Jing, Curtis Ebl, Natasha Bilkey, Ryan Emenecker, and Max Staller for helpful comments, Teva Vernoux, Tony Bishopp, and Malcolm Bennett for ARF7p:ARF7-YFP and ARF19p:ARF19-Venus seed, Doris Wagner for the 35S:NLS-2*mCerulean marker, and the ABRC for cDNA clones. This research was supported by the William H. Danforth Plant Science Fellowship Program (to S.K.P.), the US National Science Foundation (IOS-1453750 to L.C.S., MCB-1453130 to R.S., MCB-1614539 to J.M.J., and MCB-1614766 to R.V.P.), the National Institutes of Health (R01 GM112898-01 to L.C.S. and R01 5NS056114 to R.V.P.), the National Research Foundation of Korea (2017R1A2A1A17069734 to I. H.), the NSF Center for Engineering Mechanobiology (CMMI-1548571 to L.C.S.), and the St. Jude Research Collaborative fund (to R.V.P.), and the USDA-NIFA Fellowship Program (MOW-2014-01877 to D.A.K.). We thank the Genome Technology Access Center in the Department of Genetics at Washington University School of Medicine for help with genomic analysis. The Center is partially supported by NCI Cancer Center Support Grant #P30 CA91842 to the Siteman Cancer Center and by ICTS/CTSA Grant# UL1TR000448 from the National Center for Research Resources (NCRR), a component of the National Institutes of Health (NIH), and NIH Roadmap for Medical Research. This publication is solely the responsibility of the authors and does not necessarily represent the official view of either NCRR or NIH.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
DECLARATION OF INTERESTS
The authors declare no competing interests.
DATA AND SOFTWARE AVAILABILITY
The RNASeq data discussed in this publication have been deposited in NCBI’s Gene Expression Omnibus and are accessible through GEO Series accession number GSE104385.
REFERENCES
- Altmeyer M, Neelsen KJ, Teloni F, Pozdnyakova I, Pellegrino S, Grøfte M, Rask MB, Streicher W, Jungmichel S, Nielsen ML, et al. (2015). Liquid demixing of intrinsically disordered proteins is seeded by poly(ADP-ribose). Nature Communications 6, 8088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. (2000). Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genetics 25, 25–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banani SF, Lee HO, Hyman AA, and Rosen MK (2017). Biomolecular condensates: organizers of cellular biochemistry. Nature Reviews Molecular Cell Biology 18, 285–298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boer DR, Freire-Rios A, van den Berg WA, Saaki T, Manfield IW, Kepinski S, López-Vidrieo I, Franco-Zorrilla JM, de Vries SC, Solano R, et al. (2014). Structural Basis for DNA Binding Specificity by the Auxin-Dependent ARF Transcription Factors. Cell 156, 577–589. [DOI] [PubMed] [Google Scholar]
- Boija A, Klein IA, Sabari BR, Dall’Agnese A, Coffey EL, Zamudio AV, Li CH, Shrinivas K, Manteiga JC, Hannett NM, et al. (2018). Transcription Factors Activate Genes through the Phase-Separation Capacity of Their Activation Domains. Cell [DOI] [PMC free article] [PubMed]
- Boke E, Ruer M, Wuhr M, Coughlin M, Lemaitre R, Gygi SP, Alberti S, Drechsel D, Hyman AA, and Mitchison TJ (2016). Amyloid-like Self-Assembly of a Cellular Compartment. Cell 166, 637–650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brunoud G, Wells DM, Oliva M, Larrieu A, Mirabet V, Burrow AH, Beeckman T, Kepinski S, Traas J, Bennett MJ, et al. (2012). A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482, 103–106. [DOI] [PubMed] [Google Scholar]
- Chandler JW (2016). Auxin response factors. Plant Cell Environ 39, 1014–1028. [DOI] [PubMed] [Google Scholar]
- Cho WK, Jayanth N, English BP, Inoue T, Andrews JO, Conway W, Grimm JB, Spille JH, Lavis LD, Lionnet T, et al. (2016). RNA Polymerase II cluster dynamics predict mRNA output in living cells. Elife 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho WK, Spille JH, Hecht M, Lee C, Li C, Grube V, and Cisse II (2018). Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361, 412–415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chong S, Dugast-Darzacq C, Liu Z, Dong P, Dailey GM, Cattoglio C, Heckert A, Banala S, Lavis L, Darzacq X, et al. (2018). Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciuffa R, Lamark T, Tarafder AK, Guesdon A, Rybina S, Hagen WJ, Johansen T, and Sachse C (2015). The selective autophagy receptor p62 forms a flexible filamentous helical scaffold. Cell Rep 11, 748–758. [DOI] [PubMed] [Google Scholar]
- Clark NM, Hinde E, Winter CM, Fisher AP, Crosti G, Blilou I, Gratton E, Benfey PN, and Sozzani R (2016). Tracking transcription factor mobility and interaction in Arabidopsis roots with fluorescence correlation spectroscopy. Elife 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark NM, and Sozzani R (2017a). Measuring Protein Movement, Oligomerization State, and Protein-Protein Interaction in Arabidopsis Roots Using Scanning Fluorescence Correlation Spectroscopy (Scanning FCS). Methods Mol Biol 1610, 251–266. [DOI] [PubMed] [Google Scholar]
- Clark NM, and Sozzani R (2017b). Measuring Protein Movement, Oligomerization State, and Protein-Protein Interaction in Arabidopsis Roots Using Scanning Fluorescence Correlation Spectroscopy (Scanning FCS). Methods in Molecular Biology 1610, 251–266. [DOI] [PubMed] [Google Scholar]
- Conicella AE, Zerze GH, Mittal J, and Fawzi NL (2016). ALS Mutations Disrupt Phase Separation Mediated by alpha-Helical Structure in the TDP-43 Low-Complexity C-Terminal Domain. Structure 24, 1537–1549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dao TP, Kolaitis RM, Kim HJ, O’Donovan K, Martyniak B, Colicino E, Hehnly H, Taylor JP, and Castaneda CA (2018). Ubiquitin Modulates Liquid-Liquid Phase Separation of UBQLN2 via Disruption of Multivalent Interactions. Mol Cell 69, 965–978 e966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Digman MA, Brown CM, Sengupta P, Wiseman PW, Horwitz AR, and Gratton E (2005). Measuring fast dynamics in solutions and cells with a laser scanning microscope. Biophys J 89, 1317–1327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dosztányi Z, Csizmok V, Tompa P, and Simon I (2005a). IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21, 3433–3434. [DOI] [PubMed] [Google Scholar]
- Dosztányi Z, Csizmok V, Tompa P, and Simon I (2005b). The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. Journal of Molecular Biology 347, 827–839. [DOI] [PubMed] [Google Scholar]
- Enders TA, and Strader LC (2015). Auxin activity: Past, present, and future. American Journal of Botany 102, 180–196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freeman Rosenzweig ES, Xu B, Kuhn Cuellar L, Martinez-Sanchez A, Schaffer M, Strauss M, Cartwright HN, Ronceray P, Plitzko JM, Forster F, et al. (2017). The Eukaryotic CO2-Concentrating Organelle Is Liquid-like and Exhibits Dynamic Reorganization. Cell 171, 148–162 e119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuertes G, Banterle N, Ruff KM, Chowdhury A, Mercadante D, Koehler C, Kachala M, Estrada Girona G, Milles S, Mishra A, et al. (2017). Decoupling of size and shape fluctuations in heteropolymeric sequences reconciles discrepancies in SAXS vs. FRET measurements. Proc Natl Acad Sci U S A 114, E6342–E6351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guilfoyle TJ (2015). The PB1 domain in auxin response factor and Aux/IAA proteins: a versatile protein interaction module in the auxin response. Plant Cell 27, 33–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guilfoyle TJ, and Hagen G (2007). Auxin response factors. Curr Opin Plant Biol 10, 453–460. [DOI] [PubMed] [Google Scholar]
- Guilfoyle TJ, and Hagen G (2012). Getting a grasp on domain III/IV responsible for Auxin Response Factor-IAA protein interactions. Plant Sci 190, 82–88. [DOI] [PubMed] [Google Scholar]
- Halfmann R, Alberti S, Krishnan R, Lyle N, O’Donnell CW, King OD, Berger B, Pappu RV, and Lindquist S (2011). Opposing effects of glutamine and asparagine govern prion formation by intrinsically disordered proteins. Molecular Cell 43, 72–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han M, Park Y, Kim I, Kim EH, Yu TK, Rhee S, and Suh JY (2014). Structural basis for the auxin-induced transcriptional regulation by Aux/IAA17. Proc Natl Acad Sci U S A 111, 18613–18618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haughn GW, and Somerville C (1986). Sulfonylurea-resistant mutants of Arabidopsis thaliana. Mol Gen Genet 204, 430–434. [Google Scholar]
- Hnisz D, Shrinivas K, Young RA, Chakraborty AK, and Sharp PA (2017). A Phase Separation Model for Transcriptional Control. Cell 169, 13–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holehouse AS, Das RK, Ahad JN, Richardson MO, and Pappu RV (2017). CIDER: Resources to Analyze Sequence-Ensemble Relationships of Intrinsically Disordered Proteins. Biophysical Journal 112, 16–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato M, Han TW, Xie S, Shi K, Du X, Wu LC, Mirzaei H, Goldsmith EJ, Longgood J, Pei J, et al. (2012). Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell, 753–767. [DOI] [PMC free article] [PubMed]
- Koncz C, and Schell J (1986). The promoter of the TL-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Gen Genet 204, 383–396. [Google Scholar]
- Korasick DA, Chatterjee S, Tonelli M, Dashti H, Lee SG, Westfall CS, Fulton DB, Andreotti AH, Amarasinghe GK, Strader LC, et al. (2015a). Defining a two-pronged structural model for PB1 (Phox/Bem1p) domain interaction in plant auxin responses. J Biol Chem 290, 12868–12878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korasick DA, Jez JM, and Strader LC (2015b). Refining the nuclear auxin response pathway through structural biology. Curr Opin Plant Biol 27, 22–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korasick DA, Westfall CS, Lee SG, Nanao MH, Dumas R, Hagen G, Guilfoyle TJ, Jez JM, and Strader LC (2014). Molecular basis for AUXIN RESPONSE FACTOR protein interaction and the control of auxin response repression. Proc Natl Acad Sci U S A 111, 5427–5432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kroschwald S, Munder MC, Maharana S, Franzmann TM, Richter D, Ruer M, Hyman AA, and Alberti S (2018). Different Material States of Pub1 Condensates Define Distinct Modes of Stress Adaptation and Recovery. Cell Rep 23, 3327–3339. [DOI] [PubMed] [Google Scholar]
- Lancaster AK, Nutter-Upham A, Lindquist S, and King OD (2014). PLAAC: a web and command-line application to identify proteins with prion-like amino acid composition. Bioinformatics 30, 2501–2502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Last RL, and Fink GR (1988). Tryptophan-requiring mutants of the plant Arabidopsis thaliana. Science 240, 305–310. [DOI] [PubMed] [Google Scholar]
- Lavenus J, Goh T, Guyomarc’h S, Hill K, Lucas M, Voss U, Kenobi K, Wilson MH, Farcot E, Hagen G, et al. (2015). Inference of the Arabidopsis lateral root gene regulatory network suggests a bifurcation mechanism that defines primordia flanking and central zones. Plant Cell 27, 1368–1388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin Y, Protter DS, Rosen MK, and Parker R (2015). Formation and Maturation of Phase-Separated Liquid Droplets by RNA-Binding Proteins. Mol Cell 60, 208–219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu H, Yu D, Hansen AS, Ganguly S, Liu R, Heckert A, Darzacq X, and Zhou Q (2018). Phase-separation mechanism for C-terminal hyperphosphorylation of RNA polymerase II. Nature 558, 318–323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGibbon RT, Beauchamp KA, Harrigan MP, Klein C, Swails JM, Hernández CX, Schwantes CR, Wang LP, Lane TJ, and Pande VS (2015). MDTraj: A Modern Open Library for the Analysis of Molecular Dynamics Trajectories. Biophysical Journal 109, 1528–1532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mi H, Huang X, Muruganujan A, Tang H, Mills C, Kang D, and Thomas PD (2017). PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res 45, D183–D189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michniewicz M, Frick EM, and Strader LC (2015). Gateway-compatible tissue-specific vectors for plant transformation. BMC Res Notes 8, 63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitrea DM, and Kriwacki RW (2016). Phase separation in biology; functional organization of a higher order. Cell Communication and Signaling 14, 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molliex A, Temirov J, Lee J, Coughlin M, Kanagaraj AP, Kim HJ, Mittag T, and Taylor JP (2015). Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nanao MH, Vinos-Poyo T, Brunoud G, Thevenon E, Mazzoleni M, Mast D, Laine S, Wang S, Hagen G, Li H, et al. (2014). Structural basis for oligomerization of auxin transcriptional regulators. Nat Commun 5, 3617. [DOI] [PubMed] [Google Scholar]
- Okushima Y, Fukaki H, Onoda M, Theologis A, and Tasaka M (2007). ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 19, 118–130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olzscha H, Schermann SM, Woerner AC, Pinkert S, Hecht MH, Tartaglia GG, Vendruscolo M, Hayer-Hartl M, Hartl FU, and Vabulas RM (2011). Amyloid-like aggregates sequester numerous metastable proteins with essential cellular functions. Cell 144, 67–78. [DOI] [PubMed] [Google Scholar]
- Ottenschläger I, Wolff P, Wolverton C, Bhalerao RP, Sandberg G, Ishikawa H, Evans M, and Palme K (2003). Gravity-regulated differential auxin transport from columella to lateral root cap cells. Proc Natl Acad Sci U S A 100, 2987–2991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel A, Lee HO, Jawerth L, Maharana S, Jahnel M, Hein MY, Stoynov S, Mahamid J, Saha S, Franzmann TM, et al. (2015). A Liquid-to-Solid Phase Transition of the ALS Protein FUS Accelerated by Disease Mutation. Cell 162, 1066–1077. [DOI] [PubMed] [Google Scholar]
- Posey AE, Holehouse AS, and Pappu RV (2018). Phase Separation of Intrinsically Disordered Proteins. Methods Enzymol 611, 1–30. [DOI] [PubMed] [Google Scholar]
- Riback JA, Katanski CD, Kear-Scott JL, Pilipenko EV, Rojek AE, Sosnick TR, and Drummond DA (2017). Stress-Triggered Phase Separation Is an Adaptive, Evolutionarily Tuned Response. Cell 168, 1028–1040 e1019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sabari BR, Dall’Agnese A, Boija A, Klein IA, Coffey EL, Shrinivas K, Abraham BJ, Hannett NM, Zamudio AV, Manteiga JC, et al. (2018). Coactivator condensation at super-enhancers links phase separation and gene control. Science 361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sabatini S, Beis D, Wolkenfelt H, Murfett J, Guilfoyle T, Malamy J, Benfey P, Leyser O, Bechtold N, Weisbeek P, et al. (1999). An Auxin-Dependent Distal Organizer of Pattern and Polarity in the Arabidopsis root. Cell 99, 463–472. [DOI] [PubMed] [Google Scholar]
- Sauer M, Paciorek T, Benkova E, and Friml J (2006). Immunocytochemical techniques for whole-mount in situ protein localization in plants. Nat Protoc 1, 98–103. [DOI] [PubMed] [Google Scholar]
- Shin Y, and Brangwynne CP (2017). Liquid phase condensation in cell physiology and disease. Science 357. [DOI] [PubMed] [Google Scholar]
- Si K, Lindquist S, and Kandel ER (2003). A neuronal isoform of the aplysia CPEB has prion-like properties. Cell 115, 879–891. [DOI] [PubMed] [Google Scholar]
- Sun D, Wu R, Zheng J, Li P, and Yu L (2018). Polyubiquitin chain-induced p62 phase separation drives autophagic cargo segregation. Cell Res 28, 405–415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiwari SB, Hagen G, and Guilfoyle T (2003). The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 15, 533–543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulmasov T, Hagen G, and Guilfoyle TJ (1999). Activation and repression of transcription by auxin-response factors. Proc Natl Acad Sci USA 96, 5844–5849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulmasov T, Murfett J, Hagen G, and Guilfoyle TJ (1997). Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9, 1963–1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- UniProt, C. (2015). UniProt: a hub for protein information. Nucleic Acids Res 43, D204–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Lee R, Buljan M, Lang B, Weatheritt RJ, Daughdrill GW, Dunker AK, Fuxreiter M, Gough J, Gsponer J, Jones DT, et al. (2014). Classification of intrinsically disordered regions and proteins. Chemical Reviews 114, 6589–6631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vitalis A, and Pappu RV (2009). ABSINTH: a new continuum solvation model for simulations of polypeptides in aqueous solutions. Journal of Computational Chemistry 30, 673–699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J, Choi JM, Holehouse AS, Lee HO, Zhang X, Jahnel M, Maharana S, Lemaitre R, Pozniakovsky A, Drechsel D, et al. (2018). A Molecular Grammar Governing the Driving Forces for Phase Separation of Prion-like RNA Binding Proteins. Cell 174, 688–699 e616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang R, and Estelle M (2014). Diversity and specificity: auxin perception and signaling through the TIR1/AFB pathway. Curr Opin Plant Biol 21, 51–58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williamson TE, Vitalis A, Crick SL, and Pappu RV (2010). Modulation of polyglutamine conformations and dimer formation by the N-terminus of huntingtin. Journal of Molecular Biology 396, 1295–1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodruff JB, Ferreira Gomes B, Widlund PO, Mahamid J, Honigmann A, and Hyman AA (2017). The Centrosome Is a Selective Condensate that Nucleates Microtubules by Concentrating Tubulin. Cell 169, 1066–1077 e1010. [DOI] [PubMed] [Google Scholar]
- Woodward AW, and Bartel B (2005). Auxin: regulation, action, and interaction. Ann Bot 95, 707–735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu FH, Shen SC, Lee LY, Lee SH, Chan MT, and Lin CS (2009). Tape-Arabidopsis Sandwich - a simpler Arabidopsis protoplast isolation method. Plant Methods 5, 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zarin T, Tsai CN, Nguyen Ba AN, and Moses AM (2017). Selection maintains signaling function of a highly diverged intrinsically disordered region. Proc Natl Acad Sci U S A 114, E1450–E1459. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Dataset S1. Related to Figure 3. A. thaliana PLD-containing proteins
We used the PLAAC algorithm to identify prion-like domains across the full A. thaliana proteome (Lancaster et al., 2014). Analysis of PRD containing proteins using PANTHER (Mi et al., 2017) and GO Ontology (Ashburner et al., 2000) database (Released 2017–08-14) found statistical enrichment for proteins involved in mRNA binding (12.9 fold enrichment, p = 2.22E-24), transcription coactivator activity (22.95 fold enrichment, p = 5.16E-03) and transcription regulatory region DNA binding (5.15 fold enrichment, p = 9.13E-05).
Dataset S2. Related to Figure 5. Transcript accumulation differences under mock treatment.
Pairwise transcript accumulation differences between UBQ10:YFP-ARF19 (ARF19) and Wt (Col-0). Pairwise transcript accumulation differences between UBQ10:YFP-ARF19K962A (ARF19K) and Wt (Col-0). Pairwise transcript accumulation differences between UBQ10:YFP-ARF19K962A (ARF19K) and UBQ10:YFP-ARF19 (ARF19); FDR ≤ 0.01.
Dataset S3. Related to Figure 5. Transcript accumulation differences under auxin treatment
Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19 (ARF19) and auxin-treated Wt (Col-0). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19K962A (ARF19K) and auxin-treated Wt (Col-0). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19K962A (ARF19K) and auxin-treated UBQ10:YFP-ARF19 (ARF19); FDR ≤ 0.01.
Dataset S4. Related to Figure 5. Auxin-induced transcript accumulation differences in each line.
Pairwise transcript accumulation differences between auxin-treated Wt (Col-0) and mock-treated Wt (Col-0). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19 (ARF19) and mock-treated UBQ10:YFP-ARF19 (ARF19). Pairwise transcript accumulation differences between auxin-treated UBQ10:YFP-ARF19K962A (ARF19K) and mock-treated UBQ10:YFP-ARF19K962A (ARF19K); FDR ≤ 0.01.
Movie S1. Related to Figure 1. ARF19 condensate movement and fusion.
Real time video of YFP-ARF19 in a cell from the upper portion of the root.
Figure S1. Related to Figures 1 and 2. Localization of ARF7-Venus and ARF19MR
(A) Confocal microscopy images of a 5d-old seedling expressing pARF7:ARF7g-Venus.
(B) Fluorescence microscopy image from the upper root regions of a 5d-old Arabdopsis seedling expressing UBQ10:YFP-ARF19MR. Scale bar = 25 µm.
Figure S2. Related to Figure 3. Comparative analysis of ARF19 prion-like domain
To quantitatively assess the sequence composition of the ARF19 PLD we compared it against all predicted disordered regions in the A. thaliana proteome. Across the 21123 discrete disordered regions the ARF19 prion-like domain was among the least charged (bottom 2%) and most enriched in the combined fraction of glutamine (Q), serine (S), asparagine (N) and leucine (L) (above 99.7% of all other IDRs).
Figure S3. Related to Figure 3. Identification of a prion-like domain in ARF19
Results of ARF19 analysis by the PLAAC algorithm. The prion-like domain is highlighted in yellow. Figure generated by the PLAAC webserver (http://plaac.wi.mit.edu).
Figure S4. Related to Figure 3. Results of simulation analysis
(A) The interaction matrix generated by simulations of all unique pairs of non-overlapping six residue fragments taken from the ARF19 prion-like domain. Attractive interaction reflects the affinity between two fragments, with any value greater than 0 reflecting a higher affinity than would expected for two fragments that did not experience any attractive interactions.
(B) The per-fragment average attractive interaction, written as an aggregation propensity. Each value uses an average sliding-window of the three surrounding fragments to capture chain connectivity. Although there is a drop towards the C-terminus of the prion-like domain, the majority of the region shows a strong tendency to undergo self-interaction. This panel is identical to the Agg. track in figure 3A, but provides quantification of the values.
Figure S5. Related to Figure 4. Auxin induction of DR5:GUS activity in root tissues
Primary roots of 8d-old DR5-GUS (Ulmasov et al., 1997) seedlings that were treated with the indicated concentration of IAA prior to staining to visualize GUS activity.
Figure S6. Related to Figure 4. RNA-seq Data Quality
(A) The summed counts-per-million of all Ensembl gene biotypes for every sample. All samples show that greater than 95% of all aligned reads aligned to protein-coding genes.
(B) Junction saturation curves for all samples across all Ensembl known exon-exon junctions randomly sampled from aligned BAM files in 5% intervals. The plateau at higher levels of sampling indicate that all samples were sequenced with enough depth to assay all expressed genes.
(C) Localization of aligned reads across a theoretical 100bp gene body. All samples have similar profiles indicating a negligible 5-prime bias with high levels of coverage from 5-prime to 3-prime.
Figure S7. Related to Figure 4. RNA-seq Sample Quality
(A) Matrix of Spearman correlations of all detected genes greater than 1 count-per-million in at least 3 samples relative to each other. Samples of the same condition have very high correlations as expected across the diagonal with no outliers.
(B) The quality of the samples in a multi-dimensional scaling plot of the leading log fold-changes. Samples for each condition cluster very tightly with each other with good cluster separation between samples of different conditions based on expression profiles.
(C) Scatter plot of the empirically derived fitted and trended mean-variance relationship across all genes and a bar plot depicting the sample specific weight for statistical model fitting and differential expression analysis.