Abstract
RNA-binding proteins (RBPs) play a critical role in the regulation of various RNA processes, including splicing, cleavage and polyadenylation, transport, translation and degradation of coding RNAs, non-coding RNAs and microRNAs. Recent studies indicate that RBPs not only play an instrumental role in normal cellular processes but have also emerged as major players in the development and spread of cancer. Herein, we review the current knowledge about RNA binding proteins and their role in tumorigenesis as well as the potential to target RBPs for cancer therapeutics.
Keywords: RNA-Binding Proteins, RNA-Binding Domains, cancer therapeutics, small molecules, siRNA, peptide, aptamer
1. Introduction
A majority of eukaryotic genes are regulated at transcriptional and post-transcriptional levels, giving each one a unique expression profile under specific conditions (K. Chen & Rajewsky, 2007; Orphanides & Reinberg, 2002). Once an mRNA is transcribed, it is subjected to various processing, mainly controlled by RNA-binding proteins (RBPs) (Keene, 2007). RBPs coordinately bind to each mRNA and form ribonucleoprotein (RNP) complexes. The composition of the RNP complexes is dynamic based on the specific RNA processing. Notably, although RBPs bind to various classes of RNAs such as ribosomal RNAs, mRNAs, snRNA, snoRNA, tRNAs, and non-coding RNAs, about half of the RBPs manifest their function by binding to mRNAs and exert their distinct roles in regulating mRNA fate (Jin Zhang & Chen, 2008). Given the important role of RBPs in various cellular processes, such as transcription, splicing, mRNA stability, mRNA transport and translation, it comes as no surprise that alterations in RBP expression or RBP mutations can lead to various diseases, including cancer (Kechavarzi & Janga, 2014; Neelamraju, Gonzalez-Perez, Bhat-Nakshatri, Nakshatri, & Janga, 2018; Pereira, Billaud, & Almeida, 2017). Indeed, recent studies have found that altered expression, localization or post-translational modification of RBPs can contribute to tumorigenesis by not only increasing expression of oncogenes but also decreasing the expression of tumor suppressor genes. Thus, increasing attention has been paid to the roles of RBPs in cancer as well as the potential of targeting RBPs for cancer therapeutics. In this review, we will discuss the functional domains of various RBPs and their roles in malignant transformation. We will also discuss recent advances and new insights into targeting RBPs for cancer therapeutics.
2. Rna-Binding Domains
RBPs were initially identified due to their capability to bind various types of RNAs through an RNA-binding domain (RBD) that forms steady secondary and tertiary structures. The classical RBDs include the K-homology domain (KH), RNA recognition motif (RRM), Zinc finger domain (ZNF), Pumilio homology domain (PUM), double stranded RNA binding domain (dsRBD), and others (Figure 1) (Lunde, Moore, & Varani, 2007). Due to the recently developed high-throughput approaches, the number of RBPs has been largely increased to 1542 (Baltz et al., 2012; Castello et al., 2012; Gerstberger, Hafner, & Tuschl, 2014; Z. L. Wang et al., 2018), comprising about ~7.5% of the protein coding genes. Interestingly, among them, only a quarter of RBPs contain classical RNA-binding domains (RBDs) and the rest of them contain non-canonical RBDs that have previously uncharacterized motifs (Baltz et al., 2012). These non-canonical RBPs are unique in that they have intrinsically disordered regions that adapt their structure upon contacting to RNAs and subsequently mediate cell cycle regulation, metabolism, and signal transduction. For this section, we focus on some of the classical RNA-binding domains present in canonical RBPs. For the non-canonical RBPs, readers are referred to these recent reviews (Hentze, Castello, Schwarzl, & Preiss, 2018; Moore, Jarvelin, Davis, Bond, & Castello, 2018).
Figure 1.
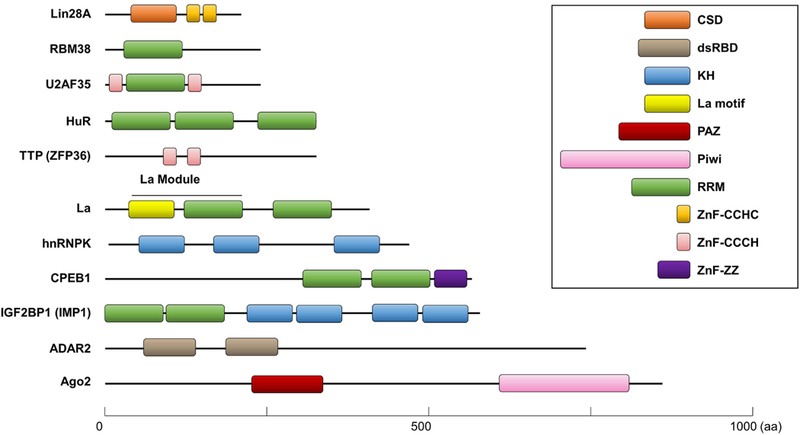
Schematic representation of RNA-binding proteins (RBPs) with various RNA-binding domains (RBDs). Each RBD is drawn as colored boxes and their lengths are shown according to their actual sizes (aa, amino acids). The common RBDs include: CSD, cold shock domain; dsRBD, double-stranded RNA-binding domain; KH, K-homology domain; La motif; PAZ, Piwi/Argonaute/Zwille domain; Piwi domain; RRM, RNA-recognition motif; ZnF, zinc fingers of the CCHC, CCCH, and ZZ type.
2.1. RNA Recognition Motif (RRM)
RNA-recognition motif (RRM), also called as ribonucleoprotein (RNP) domain, is one of the most abundant and classical RNA-binding domain present in a number of RBPs (Maris, Dominguez, & Allain, 2005). Their abundance and early discovery also make them the most studied RBD with more than 30 solved crystal structures of RRMs bound to their target RNA sequences. (Sickmier et al., 2006; X. Wang & Tanaka Hall, 2001). Each RRM is made up of about 90 amino acids and consists of two conserved sub-motifs referred to as RNP1 and RNP2. RNP1 consists of 8 amino acids with the consensus sequences, (R/K)-G-(F/Y)-(G/A)-(F/Y)-V-X-(F/Y), whereas RNP2 contains 6 amino acids with the consensus sequences, (L/I)-(F/Y)-(V/I)-X-(N/G)-L. Structurally, the RRM domain is composed of four anti-parallel β sheets stacked against two α helices, giving it the β1α1β2β3α2β4 topology with RNP1 and RNP2 motifs making the central β3 and β1 sheets, respectively. Most of the RRMs contain three conserved aromatic amino acids in the central β strands, which allow for the recognition of two nucleotides. Some RRMs lack one or more of the conserved aromatic residues forming the derivative RRM domains such as the quasi-RRM, the pseudo-RRM or U2AF homology motifs. Additional nucleotides can be recognized by the RRM using other aromatic and planar side-chains present on other β strands. In general, each RRM domain can recognize anywhere between 4 to 8 nucleotides by using β-sheet surface extensions or by using exposed loops connecting β–sheet to a β–sheet or α-helix. To achieve sequence specificity and affinity for an mRNA, RBPs generally contain more than one RRM and bind RNA cooperatively. This has been demonstrated with the structures of several tandem RRMs bound to RNA, such as HuD and nucleolin (Allain, Bouvet, Dieckmann, & Feigon, 2000; Wang & Tanaka Hall, 2001). In other cases, some RRMs do not coordinate with each other and appear to bind RNA independently. For example, the RRM1 and RRM2 in PTB are clearly independent from each other and are separated by long flexible linkers (Oberstrass et al., 2005). The different binding affinity among RRMs may be critical for target selection.
2.2. hnRNP-K-Homology domain (KH domain)
The heterogeneous nuclear ribonucleoprotein (hnRNP) K-homology (KH) domain is approximately 70 amino acids long and was initially identified in the human hnRNP-K protein more than two decades ago (Siomi, Matunis, Michael, & Dreyfuss, 1993). There are two types of KH domains: the eukaryotic type I and prokaryotic type II, both of which comprise of three α-helices and a three-stranded anti-parallel β-sheets (Grishin, 2001; Valverde, Edwards, & Regan, 2008). While both type I and type II share a minimal βααβ core, they fold differently due to the location of additional α and β elements. In the type I KH domain, the α and β elements are located at the C-terminus, whereas in the type II KH domain, these elements are located at the N-terminus. Additionally, KH domain contains a conserved “GXXG” loop motif and a variable loop. KH domain interacts with nucleotides through hydrogen bonds, electrostatic interactions, and shape complementarity. Generally, each KH domain can recognize four nucleotides through a binding cleft formed by the α1 and α2 helices linked by the “GXXG” loop on one side and the β-sheet and the variable loop on the other side. For example, Nova2 (Neuro-oncological ventral antigen 2) is a tissue-specific alternative splicing factor and contains three type I KH domains. The crystal structure of the KH3 domain in complex with RNA shows that Nova2 interacts with the single stranded UCAC motif (Lewis et al., 2000). Additionally, the KH3 domain in PCBP2 recognizes the CCCT motif whereas the KH3 domain in hnRNP K binds to TCCC motif (Du et al., 2007). Sometimes, the KH domains in the same RBP can recognize different motifs. For instance, the KH1 domain in NusA RBP recognizes AGAA motif whereas its KH2 domain recognizes CAAU motif (Beuth, Pennell, Arnvig, Martin, & Taylor, 2005). These structural analyses indicate that KH domains are versatile in recognizing a large panel of sequences; however, its affinity and specificity of interaction is low. Thus, at least two strategies are selected by KH domain containing RBPs to overcome this. The first strategy is to extend the surface of interaction with nucleic acids. For example, the binding interface in SF1/mBBP is composed of a KH domain along with a C-terminal helix, known as the QUA2 domain (Z Liu et al., 2001). As a result, SF1 specifically binds to the six nucleotide sequence 5′-UACUAAC-3′ (Berglund, Chua, Abovich, Reed, & Rosbash, 1997). The second strategy is the repetition of multiple KH domains within a single RNA binding protein, which is a common feature in most KH domain containing RBPs. In general, multiple copies of KH domains can act in synergy and interact efficiently with their target. For example, the two KH domains of NusA together show stronger binding than each separated (Beuth et al., 2005). However, there are also exceptions. FUSE-binding protein (FBP) contains four KH domains and are separated by a longer linker indicating that each domain recognizes RNA independently (Braddock, Louis, Baber, Levens, & Clore, 2002). Thus, each KH domain binds separately to its target.
2.3. Zinc-Finger domain
Discovered more than 30 years ago as classical DNA-binding proteins, zinc fingers proteins were subsequently found to bind RNAs as demonstrated by their crystal structure (Hudson, Martinez-Yamout, Dyson, & Wright, 2004; D. Lu, Searles, & Klug, 2003). Each zinc finger consists of about 25–30 amino acids and forms a ββα topology that is held together by the central Zn2+ ion. They are generally classified based on the amino acid residues coordinating the Zn2+ atom, e.g., CCHH, CHCC, CCCH, CCHC, or CCCC, with the CCHH being the most frequent type. Transcription factor TFIIIA was the first zinc finger containing protein that was initially discovered through its association with 5S rRNA (Searles, Lu, & Klug, 2000). TFIIIA contains nine zinc fingers: zinc fingers 1–3, 5 and 7–9 bind DNA, zinc fingers 4–6 contact RNA, whereas zinc finger 5 contacts both DNA and RNA. The CCHH-type zinc fingers interact directly with the major groove of DNA by forming hydrogen bonding with the Watson-Crick base pairings through the side chains of variable residues present in their α-helix. Unlike other domains that use β-sheet to recognize RNA, the zinc finger uses the exposed loops and α-helices for the recognition. Other versatile modes of interaction of zinc fingers with RNA have also been reported. For example, TIS11d, a member of the tristetraprolin (TTP) protein family contains two CCCH zinc fingers and mediates mRNA degradation via binding to an AU-rich element (ARE) in mRNA 3’-UTRs (Blackshear, 2002). Structural studies show that each zinc finger of TIS11d can specifically recognize an ‘UAU’ motif primarily through hydrogen bonds between the protein backbone and the RNA bases. MBNL1, another zinc finger protein, contains four CCCH zinc fingers and binds RNA (Pascual, Vicente, Monferrer, & Artero, 2006). The preferential binding of zinc fingers 3 and 4 of MBNL1 to ‘GC’ and ‘GCU’ RNA motifs, respectively, is mediated by stacking interactions and several hydrogen bonds involving main chains. These structure data indicate that zinc fingers can specifically bind single-stranded RNA, primarily using hydrogen bonds and aromatic-base stacking interactions. Moreover, these domains are more plastic and flexible which allows them to acquire different conformations to accommodate various sequences.
2.4. Double-stranded RNA binding domain
The double stranded RNA binding domain (dsRBD) is found in both prokaryotes and eukaryotes and controls diverse RNA related functions such as RNA interference, localization, editing and translocation. The dsRBD typically consists of 70–90 amino acids and displays a distinctive αβββα protein folding pattern (Bycroft, Grunert, Murzin, Proctor, & St Johnston, 1995; Nanduri, Carpick, Yang, Williams, & Qin, 1998; Ryter & Schultz, 1998). Notably, unlike other RBDs, most dsRBDs recognize the shape of the dsRNA rather than the RNA sequence. As observed from the crystal structure of RBP Xlrbpa complexed with synthetic dsRNA (Ryter & Schultz, 1998), the interaction mostly involves RNA’s phosphodiester backbone and its ribose 2′-OH groups, which explains why dsRBDs bind to dsRNA mainly dependent upon the structure rather than the sequence (Ryter & Schultz, 1998). In most cases, dsRBDs interact with a regular RNA A-form helix structure, composed of a major groove and two consecutive minor grooves. Additionally, dsRBDs also bind to single-stranded RNAs with bulges or mismatches (B. Tian et al., 2000). Like other RBDs, dsRBDs are often present in multiple copies in an RNA-binding protein, conferring some specificity for certain RNA secondary structures. Interestingly, despite the conservation in both sequence and structure, some dsRBDs show strong binding with various dsRNAs whereas others show weak affinity for dsRNAs. Additionally, although most dsRBDs recognize RNA shape, the solution structure of dsRBD2 of ADAR2 showed dsRNA recognition not only by shape of the RNA but also by the sequence (Stefl et al., 2010). ADAR2 is an adenosine deaminase that converts adenosine-to-inosine (A-to-I) by hydrolytic deamination in numerous mRNA and pre-mRNA transcripts.
2.5. Piwi/Argonaute/Zwille (PAZ) and Piwi domains
The PAZ domain is approximately 140 amino acids in length and is named after three proteins-Piwi, Argonaute, and Zwille. PAZ domains are commonly found in proteins involved in miRNA biogenesis, such as Dicer and Ago. In 2003, three groups determined the PAZ domain structures concurrently (Lingel, Simon, Izaurralde, & Sattler, 2003; J. J. Song et al., 2003; K. S. Yan et al., 2003). The PAZ domain structures contain two subdomains with a prominent cleft between. The larger subdomain has a fold that is reminiscent of oligonucleotide/oligosaccharide binding (OB) fold, although with slightly different topology, and the second subdomain, called by some “the appendage,” is composed of a β-hairpin followed by an α-helix. Since a majority of the OB-fold proteins bind single-stranded oligonucleotides or oligosaccharides, the initial PAZ domain structures suggested its role in nucleic acid binding. Later studies confirmed the mode of PAZ-siRNA recognition by the structures of PAZ domain in complex with 3’ overhanging ends of siRNAs (Lingel, Simon, Izaurralde, & Sattler, 2004; J. B. Ma, Ye, & Patel, 2004).
The Piwi domain is only present in Argonaute proteins and its crystal structure revealed that it clearly belongs to the RNase H family of enzymes (J. J. Song, Smith, Hannon, & Joshua-Tor, 2004), comprising a five-stranded mixed β sheet surrounded by α helices on both sides. RNase H enzymes are known to cleave the RNA strand of an RNA–DNA hybrid (W. Yang & Steitz, 1995). In addition to its core RNase H fold, Piwi domain contains two highly conserved aspartates always present on adjacent β-strands. Mutation of each of these aspartates eliminates the slicer activity of Argonaute2 (J. Liu et al., 2004; Rivas et al., 2005). Piwi domains have high affinities for ssDNA, medium affinities for RNA/DNA and dsDNA, and the lowest for dsRNA and ssRNA (J. B. Ma et al., 2005; Yuan et al., 2005).
2.6. S1 domain
The S1 domain was originally identified in ribosomal protein S1, but has since been found in a large number of RNA-associated proteins (Subramanian, 1983). The E. coli S1 protein consists of six S1 domains folded into five-stranded anti-parallel β-barrel, which are involved in RNA binding (Bycroft, Hubbard, Proctor, Freund, & Murzin, 1997). The fold in S1 domain is similar to the oligonucleotide/oligosaccharide binding (OB)-fold superfamily (Flynn & Zou, 2010; Theobald, Mitton-Fry, & Wuttke, 2003). The nucleic acid binding of S1 domain requires a two-stranded β-sheet core along with the surrounding loops and secondary structure elements (Schubert et al., 2004).
2.7. DEAD-box domain
DEAD box proteins form the largest helicase family (Fairman-Williams, Guenther, & Jankowsky, 2010) and are characterized by the presence of an Asp‑ Glu‑ Ala‑ Asp (DEAD) motif. DEAD box helicases play a central role in cellular RNA metabolism and generally function as part of larger multicomponent assemblies, such as the spliceosome or the eukaryotic translation initiation machinery (Linder & Jankowsky, 2011). Several crystal structures of helicases from the various superfamilies have been obtained and indicate that DEAD box proteins contain two covalently linked globular domains, each of which generally contains five β-strands surrounded by five α-helices, resembling the folding of the RecA ATPase (Andersen et al., 2006). There are at least 12 characteristic sequence motifs located at conserved positions, with the DEAD box located at Motif II. Structural analysis revealed that all the DEAD box proteins utilize a highly conserved mode of RNA binding (Del Campo & Lambowitz, 2009; Sengoku, Nureki, Nakamura, Kobayashi, & Yokoyama, 2006), whereby the helicase core mediates the contacts exclusively to the sugar phosphate backbone of the RNA. In addition to its helicase core, the core DEAD box proteins contain variable auxiliary domains, which are located at C- and N-terminals. These domains are thought to be critical for the diverse functions of these enzymes, e.g., allowing interaction with other proteins or with RNA targets.
3. Aberrant Expression of RBPS In Cancer
Altered RNA metabolism due to an RBP malfunction can lead to genome-wide changes in the transcriptome and proteome of the cells and subsequently, affect cell growth, proliferation, invasion and death. Thus, it is not a surprise that altered expression of RBPs is a common phenomenon during development and progression of cancers. Thus, a table is provided to briefly elucidate the alteration of these RBPs in cancer (See Table 1). In this section, we will focus on several RBP families and their emerging roles in cancer.
Table-1–
Altered RBPs in cancer and the therapeutic strategy used (if any) to target them
3.1. The RBM38 and RBM24 family proteins
The RNA-binding proteins RBM38 and RBM24 share a high degree of homology and thus constitute a family of single RRM-containing RNA-binding proteins (Y. Jiang et al., 2014; Shu, Yan, & Chen, 2006). RBM38 and RBM24 are known to regulate many aspects of mRNA metabolism, including splicing, mRNA stability and translation (Heinicke et al., 2013; Miyamoto, Hidaka, Jin, & Morisaki, 2009; Ohe et al., 2017; J Zhang et al., 2014; M. Zhang et al., 2018). Notably, RBM38 and RBM24 contain an identical RRM and prefer to bind to AU/U-rich elements in target mRNAs, suggesting that these two proteins have similar functions through modulating the same targets. For example, both RBM38 and RBM24 can be induced by tumor suppressor p53 and in turn repress p53 mRNA translation (J Zhang et al., 2011; M. Zhang et al., 2018). Other targets that can be regulated by RBM38 and RBM24 include p21 and p63 (Cho, Zhang, & Chen, 2010; Y. Jiang et al., 2014; Miyamoto et al., 2009; Shu et al., 2006; E. Xu et al., 2014; J Zhang, Jun Cho, & Chen, 2010). Despite these common features, the tissue expression profile of RBM24 differs from that of RBM38. RBM24 is mainly expressed in muscle and heart whereas RBM38 is highly expressed in lymphatic tissues, such as spleen, thymus and bone marrow. In addition, Rbm24-KO mice are embryonic lethal whereas Rbm38-null mice are bone alive but prone to hematopoietic defects and spontaneous tumors (van den Hoogenhof et al., 2017; J Yang et al., 2014; J Zhang et al., 2014; M. Zhang et al., 2018). These data indicate that both the proteins have non-redundant functions.
RBM38 has been implicated in cancer development and its expression is found to be altered in several different kinds of cancers. For example, RBM38 is suggested to be a tumor suppressor in hepatocellular carcinoma, non-small cell lung cancer and renal cell carcinoma (Ding et al., 2014; W. Huang et al., 2017; L. Yang, Zhang, Ling, & Heng, 2018). Similarly, other studies indicated a tumor suppressor function of RBM38 in breast cancers and demonstrated that RBM38 is down-regulated in breast cancers by promoter hypermethylation (Leveille et al., 2011; J. Q. Xue et al., 2014). Similarly, another study showed that high expression of RBM38 in ovarian cancer, breast cancer and glioma is associated with better patient survival (Feldstein, Ben-Hamo, Bashari, Efroni, & Ginsberg, 2012). However, we showed that RBM38 is over-expressed in canine lymphoma, which is associated with low expression of p53 (J Zhang et al., 2011). Additionally, high expression of RBM38 was found to be associated with poor prognosis in breast cancer and with malignant transformation of colorectal adenoma to carcinoma (Carvalho et al., 2009; Chin et al., 2006; Hermsen et al., 2002; Jenssen, Kuo, Stokke, & Hovig, 2002). These apparent opposing data about the role of RBM38 in tumorigenesis could be explained by the fact that RBM38 might regulate unique targets in different cancers. In line with this, RBM38 was found to regulate multiple tumor suppressors as well as oncogenes, including MDM2, GDF15, HuR, HIF1, ER, PR, ZO-1, and PTEN (Cho et al., 2015, 2010; Lou et al., 2017; L. Shi et al., 2015; E. Xu, Zhang, & Chen, 2013; T. Yin, Cho, & Chen, 2013; J Zhang et al., 2018). Thus, the role of RBM38 in tumorigenesis could be context dependent. Indeed, we showed that depending on the p53 status, RBM38 can either promote or suppress tumor formation since RBM38 represses both wild-type and mutant p53 expression via mRNA translation (J Zhang et al., 2014). By using multiple mouse models, we showed loss of RBM38 decreases the tumor penetrance of p53 heterozygous mice, presumably through enhancing wild-type p53 expression (J Zhang et al., 2014). On the contrary, in a mutant p53 background, RBM38 KO promotes tumor incidence and tumorigenesis (J Zhang et al., 2018). Taken together, RBM38 is an important RBP that plays a vital role in normal physiology and shows altered expression and function in various cancers.
The role of RBM24 in cancer is much less studied compared to RBM38. To date, the biological function of RBM24 is found to be involved in skeletal muscle differentiation by regulating p21, MyoD and myogenin expression (Y. Jiang et al., 2014; Jin, Hidaka, Shirai, & Morisaki, 2010; H. Li, Bourdelas, Carron, & Shi, 2010; Miyamoto et al., 2009). Additionally, RBM24 is found to be involved in sarcomeric assembly and cardiac contractility (Maragh et al., 2011; Poon et al., 2012). Consistent with these findings, we and others showed that loss of RBM24 leads to multiple cardiac malfunctions in mice (J Yang et al., 2014; M. Zhang et al., 2018), suggesting that RBM24 plays a critical role in the early stages of heart development. Despite its critical role in heart development, several lines of evidence suggest that RBM24 plays a role in tumorigenesis. For example, a recent study indicated RBM24 expression was found to be frequently down-regulated across 15 cancer types compared to normal tissues (Z. L. Wang et al., 2018). Consistent with this, RBM24 was found to suppress cancer progression by up-regulating miR-25 in nasopharyngeal carcinoma (W. F. Hua et al., 2016). However, we showed previously that RBM24 represses tumor suppressors, including p53 and p63 (E. Xu et al., 2014; M. Zhang et al., 2018), suggesting that RBM24 may function as an oncogene. Thus, the role of RBM24 in cancer may be complicated by its regulation of multiple targets involved in tumor suppression or promotion.
3.2. The PCBP family proteins
The PCBP family of proteins consists of hnRNP K and PCBP-1, 2, 3, 4, and belong to the KH domain superfamily of nucleic acid-binding proteins. All PCBP family proteins contain three KH domains and are unique in their high affinity for tracts of poly-cytosine (Poly-C binding specificity). Notably, PCBP1 is an intron-less gene and thought to be produced by retro-transposition of a PCBP2 splicing transcript (Makeyev, Chkheidze, & Liebhaber, 1999). Thus, PCBP1 and PCBP2 share the highest level of amino acid sequence identity (~82%) among all the members. Additionally, PCBP3 and PCBP4 share 64% and 45% homology with PCBP2, respectively. PCBP family proteins have broad functions involving transcriptional regulation and mRNA metabolism (H. S. Choi et al., 2009; Makeyev & Liebhaber, 2002). Recently, PCBP1 and PCBP2 were also identified as cytosolic iron chaperones that deliver iron to ferritin (Leidgens et al., 2013; H. Shi, Bencze, Stemmler, & Philpott, 2008). Compared with other members, the biological function of PCBP3 is much less understood. PCBP3 was found to repress mu opioid receptor gene by binding to double stranded poly(C) element (H. S. Choi et al., 2007; S. S. Kim et al., 2005) and may be involved in spermiogenesis (Chapman et al., 2013). Recently, PCBP3 was suggested to be a favorable prognostic marker for pancreatic cancer prognosis (Ger et al., 2018). Thus, we will focus on the role of PCBP1, PCBP2, PCBP4 and hnRNP K in tumorigenesis. Interestingly, although PCBP proteins share high sequence similarity, they appear to have opposing roles in tumorigenesis. PCBP1 and PCBP4 are thought to function as tumor suppressors whereas PCBP2 and hnRNP K function as oncogenes (J. Guo & Jia, 2018).
PCBP1 is shown to be down-regulated in various types of cancers, such as lung cancer, cervical cancer, breast cancer, colon cancer, and liver cancer (Pillai et al., 2003; Thakur et al., 2003; H. Wang et al., 2010; T. Zhang et al., 2010). In line with this, high expression of PCBP1 is associated with good prognosis in patients diagnosed with lung cancer and acute myeloid leukemia (Y. Liu et al., 2015; M. Zhou & Tong, 2015). The tumor suppressive function of PCBP1 is probably due to its regulation of many targets involved in cancer development. For example, one study showed that PCBP1 inhibits tumorigenesis through translationally repressing PRL-3 phosphatase (H. Wang et al., 2010). PRL-3 is a multitasking phosphatase involved in multiple steps of cancer progression, especially in cancer metastasis (Saha et al., 2001; Zeng, Hong, & Tan, 1998). In addition, PCBP1 was found to increase p27 mRNA stability and translation (H. Shi et al., 2018) but suppressed expression of MAPK1 (Y Zhang et al., 2018). However, studies from the PCBP1-null mice did not provide direct evidence that PCBP1 is a tumor suppressor. PCBP1-null mice are embryonic lethal at the pre-implantation stage whereas PCBP1-het mice are viable but prone to a postnatal weight gain defect (Ghanem et al., 2016). Later, it was found that conditional PCBP1 deficiency leads to microcytic anemia in mice and is linked to auto-inflammatory diseases (Ryu, Zhang, Protchenko, Shakoury-Elizeh, & Philpott, 2017; Z. Wang et al., 2018), both of which are known to be related to cancer.
Similar to PCBP1, several lines of evidence indicate that PCBP4 can function as a tumor suppressor. First, the PCBP4 gene is located on chromosome 3p21 and loss of chromosome 3p21 is one of the most prevalent genomic abnormalities in lung cancer (Dammann et al., 2000; Tai et al., 2006). Moreover, PCBP4 was also found to be down-regulated in several types of cancers, including lung cancer (Pio et al., 2004). Consistent with this, over-expression of PCBP4 inhibits proliferation of lung cancer cells (Castano, Vergara-Irigaray, Pajares, Montuenga, & Pio, 2008; J. Zhu & Chen, 2000). Furthermore, our group generated PCBP4-null mice and found that these mice are prone to lung cancer and lymphoma (W. Yan et al., 2016). The tumor suppressive function of PCBP4 is at least in part due to its regulation of several targets that are known to be critical for tumor suppression. For example, we showed that PCBP4 is required for the expression of tumor suppressor p53 by modulating p53 stab ility via ZFP871 (W. Yan et al., 2016). We also showed that PCBP4 is required for maintaining the stability of p21 mRNA (Scoumanne, Cho, Zhang, & Chen, 2011). Nevertheless, more studies are needed to further elucidate the role of PCBP4 in tumor suppression.
PCBP2 was found to be up-regulated in several types of cancers, such as gastric cancer, esophageal cancer, glioblastoma, and HCC (Hu, Liu, Zhang, & Huang, 2014; Luo & Zhuang, 2017; J Ye et al., 2016; X Zhang et al., 2016). Consistent with this, knockdown of PCBP2 inhibited glioma cell growth both in vitro and in nude mice (W. Han et al., 2013). Several targets of PCBP2 have been identified, which might explain the tumor promoting function of PCBP2. For example, PCBP2 was able to destabilize the mRNA of four-and-a-half LIM domain 3 (FHL3), which inhibits glioma cell growth (W. Han et al., 2013). Additionally, PCBP2 was found to enhance the binding of miR‐151‐5p and miR‐16 to ARHGDIA (rho GDP dissociation inhibitor alpha, a metastasis suppressor) and subsequently, shown to facilitate the migration and invasion of glioma cells (X. Lin et al., 2016). However, other studies indicate that PCBP2 may function as a tumor suppressor. For example, we found that PCBP2 is required for the expression of TAp73, a member of the p53 tumor suppressor family and mediates p73-dependent antioxidant defense (Ren, Zhang, Yan, Zhang, & Chen, 2016). Additionally, PCBP2 was found to form a complex with the Hippo pathway components and suppress growth in breast epithelial cells (F. Li, Bullough, Vashisht, Wohlschlegel, & Philpott, 2016). Moreover, PCBP2-null mice die between E12.5 and E15.5 with hemorrhage and edema (Ghanem et al., 2016), which also make it difficult to study the role of PCBP2 in tumorigenesis by using conventional knockout mouse models.
Similar to PCBP2, hnRNP K is also found to be up-regulated in cancers including bladder cancer, gastric cancer, melanoma, prostate cancer, nasopharyngeal cancer, and oral squamous cell carcinoma (L. C. Chen et al., 2008; X. Chen et al., 2017; Nagano et al., 2004; Wen et al., 2010; C. S. Wu et al., 2012; R. Yang et al., 2016). Consistent with these findings, hnRNP K was found to facilitate the expression of several oncogenes, such as c-Myc and Src (Adolph, Flach, Mueller, Ostareck, & Ostareck-Lederer, 2007; Evans et al., 2003). Additionally, hnRNP K is required for colon cancer survival by modulating MRPL33 (L. Liu et al., 2018). Despite clinical data suggesting that hnRNP K functions as an oncogene, studies have also showed that hnRNP K impacts tumor suppressor pathways, including the p53 pathway (Enge et al., 2009; Moumen, Masterson, O’Connor, & Jackson, 2005) and mediates tumor suppression. Indeed, hnRNP K was found to be one of genes that are frequently deleted in patients with acute myeloid leukemia (AML) (Dayyani et al., 2008; Sweetser et al., 2005), consistent with its tumor suppressive roles. Moreover, hnRNP K haploinsufficient mice are tumor prone and have reduced survival and most importantly, these mice mimic the putative haploins ufficiency observed in AML patients with 9q21.32 deletions (Gallardo et al., 2015). Together, these data suggest that hnRNP K has both oncogenic and tumor suppressive functions and may exert these functions in a tissue-specific manner.
3.4. The tristetraprolin (TIS11/TTP) family
The Tristetraprolin gene (TIS11, TTP, ZFP36, NUP475, GOS24) was initially identified as a TPA (12-O-tetradecanoylphorbol-13-acetate) inducible sequence in cultured murine fibroblasts (Q. Ma & Herschman, 1991; Varnum, Lim, Sukhatme, & Herschman, 1989). Later, it was found that TIS11 belongs to a small family of RNA binding proteins containing tandem CCCH zinc fingers (Ciais, Cherradi, & Feige, 2013). The human TIS11 family consists of three members TIS11, TIS11b (Berg36, ERF-1, ZFP36L1, BRF-1), and TIS11d (ZFP36L2, ERF-2, BRF-2). A fourth family member, Zfp36l3, is only expressed in mouse placenta, but is not detected in human placenta or other human tissues (Blackshear et al., 2005; Frederick, Ramos, & Blackshear, 2008). TIS11 family members share two highly conserved CCCH tandem zinc fingers preceded by an YKTEL motif, which is necessary for their RNA binding activities (Hudson et al., 2004; Lai, Carballo, Thorn, Kennington, & Blackshear, 2000). TIS11 family members can bind directly to AU‐rich elements within mRNAs, and then promote decay of those mRNAs and/or inhibit their translation (Baou, Jewell, & Murphy, 2009). The mRNA decay mediated by TIS11 is carried out by multiple mechanisms. For example, TIS11 and TIS11b recruit and associate with the decapping enzymes Dcp1 and Dcp2 (Lykke-Andersen & Wagner, 2005). Additionally, TIS11 associates indirectly with poly(A)‐specific ribonuclease (PARN), resulting in de‐adenylation and destabilization of target mRNAs (Lai, Kennington, & Blackshear, 2003).
Based on the studies from knockout mouse models, it seems that all TIS family proteins have a broad role in various biological processes. For example, TIS11 was first found to be involved in inflammatory disease. TIS11-null mice appeared normal at birth, but soon developed a systemic autoimmune inflammatory syndrome due to TNF-α overproduction, which can be rescued by injection of the mice with anti-TNFα antibodies shortly after birth (Taylor et al., 1996). TIS11b-null mice died mid-gestation and exhibited extraembryonic and intraembryonic vascular abnormalities and heart defects, which is due to elevated expression of vascular endothelial growth factor (VEGF)-A (Bell et al., 2006; Stumpo et al., 2004). TIS11d knock‐out mice are viable for approximately 2 weeks after birth and exhibit significant defects in hematopoiesis, resulting in anemia and thrombocytopenia (Stumpo et al., 2009). These different phenotypes observed from germline ablation of Tis11 family members suggests that they have unique role during development by modulating their own specific targets despite their highly conserved RNA binding domains.
Many targets have been identified that are regulated by the TIS11 family, most of which are involved in malignancy (Baou et al., 2009; Park, Lee, & Kang, 2018; Zanocco-Marani, 2010), suggesting a tumor suppressive function of the TIS11 family proteins. In support of this notion, TIS11 has been reported to be down-regulated in several human cancers and suppresses cancer (Milke et al., 2013; Wei et al., 2016). Taken together, these observations suggest that the TIS11 family proteins have a vital role in human cancers.
3.5. The Musashi RBPs
The Musashi (MSI) RBP was first identified in Drosophila due to its role in regulating asymmetrical division of Drosophila sensory organ precursor (SOP) cells (Nakamura, Okano, Blendy, & Montell, 1994). The Musashi gene is evolutionary conserved: mammals contain two Musashi homologs: Musashi-1 (MSI1) and Musashi-2 (MSI2) due to an earlier duplication event in vertebrates. MSI1 and MSI2 share 75% amino acid identity and contain two RRM domains that facilitate target mRNA binding. MSI1 and MSI2 are found to play an important role in guiding the appropriate differentiation of neuronal progenitor cells (S. I. Sakakibara et al., 1996; S. Sakakibara, Nakamura, Satoh, & Okano, 2001) as well as in regulating organ development for other tissue types (Sutherland et al., 2014). The Musashi proteins have also been linked to cancer (Raymond G. Fox, Park, Koechlein, Kritzik, & Reya, 2015; Kudinov, Karanicolas, Golemis, & Boumber, 2017). For example, elevated expression of MSI1 was found in gliomas (Kanemura et al., 2001) and medulloblastomas (Dat T. Vo et al., 2012). In line with this, MSI1 overexpression correlates with poor prognosis in breast cancer patients (X. Y. Wang et al., 2010) and promotes the metastasis of breast cancer cells to lungs (Oskarsson et al., 2011). Additionally, MSI1 acts as a prognostic factor in ovarian (P. xiang Chen, Li, & Yang, 2015) and colorectal cancer patients (D. Li et al., 2011). Similarly, MSI2 is also found to play a role in cancer development. Msi2 was identified in chronic myeloid leukemia (CML) as part of a translocation event that fused the RRMs of Msi2 with HoxA9 (Barbouti et al., 2003). Using CML mouse model, Ito and colleagues showed that Msi2 regulates CML disease progression by binding and suppressing Numb mRNA (T. Ito et al., 2010). Another study showed that overexpression of Msi2 in BCR-ABL1 CML mouse model, led to induction of aggressive leukemia and thus, identified MSI2 as a prognostic marker for human AML (Kharas et al., 2010). Additionally, MSI2 has been shown to regulate growth and metastasis of pancreatic, lung, bladder and colon cancer (K. Guo et al., 2017; Kudinov et al., 2016; C. Yang et al., 2016; Zong et al., 2016). Together, these studies clearly indicate that the Musashi proteins are critical modulators of oncogenic initiation and progression.
3.5. HuR
HuR (Hu antigen R) is a member of ELAV family of RNA-binding proteins, which can associate with mRNAs containing AREs in their 3’-UTRs (Brennan & Steitz, 2001; Srikantan & Gorospe, 2012). HuR protein contains two tandem RNA-recognition motifs (RRM), a hinge region and a third RRM. The hinge region in HuR can be modulated by various kinases and is involved in nucleo-cytoplasmic shuttling of the protein. In response to stress signals, such as UV irradiation and polyamine depletion, HuR is translocated from nucleus to cytosol wherein HuR binds to numerous target mRNAs and regulates their stability and/or translation (Kuwano et al., 2008; Lafarga et al., 2009; W. Wang et al., 2000; Xiaoling Yang et al., 2004; Zou et al., 2010). Additionally, HuR can directly bind to miRNAs and thus prevent the downregulation of their targets.
As the role of HuR in various cancers has been worked upon and reviewed extensively (Abdelmohsen & Gorospe, 2010; Brody & Dixon, 2018; J. Wang et al., 2013), we will discuss briefly about the function of HuR in different cancers (Also see Table 1). HuR is almost exclusively overexpressed in cancers. For example, HuR has been shown to be overexpressed in high-grade human brain tumors such as glioblastoma multiforme and medulloblastoma where it binds and stabilizes mRNAs of growth factors linked to brain tumor progression (Nabors, Gillespie, Harkins, & King, 2001). In addition, cytoplasmic expression of HuR in human breast tumors is associated with high tumor grade and poor disease-free survival (Heinonen et al., 2005). Similarly, cytoplasmic HuR correlates with increased colon cancer malignancy likely through its ability to stabilize β-catenin mRNA (López De Silanes et al., 2003). Moreover, over-expression of HuR is associated with advanced tumor grade in gastric cancer (Kang et al., 2008) and poor survival of HCC patients with early-stage disease (H. Zhu et al., 2015). Taken together, these data indicate that HuR is a critical RBP that promotes tumorigenesis and targeting HuR could be a useful cancer therapy.
4. RBPS As Therapeutic Targets For Cancer
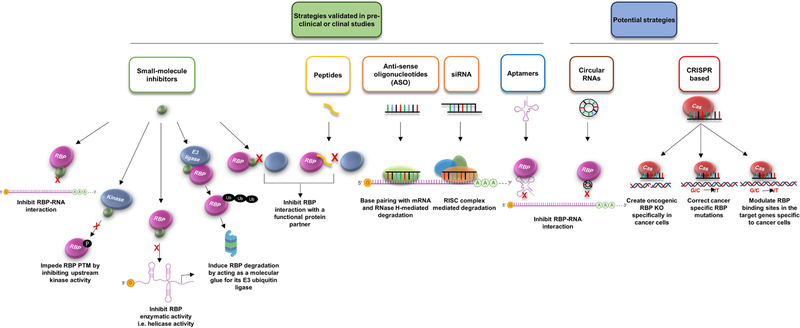
As evident from the above sections, RBPs not only play an indispensable role in normal physiology but also are vital to cancer initiation and progression. Thus, targeting RBPs for cancer therapy has gained more attention in the last decade. Recent years have seen increase in studies targeting the previously ‘undruggable’ RBPs as it has become increasingly possible to target these molecules using diverse approaches. In this section, we will review various strategies that have been used in recent years to target RBPs in cancer and other human diseases and also discuss some potential novel targeting schemes (Figure 2).
Figure 2.
Strategies that are being applied to target RBPs for cancer therapeutics. Small-molecules can be designed to inhibit the interaction of RBP with an RNA, suppress RBP’s enzymatic activity, block RBP post-translational modification (PTM), or induce RBP degradation. The Peptides-based strategy can be designed to hinder the RBP association with its functional partner(s). The oligonucleotide-based strategies include ASO, siRNA, and Aptamer, which can either mediate RBP mRNA degradation or inhibit RBP-RNA interaction. In addition, two potential strategies are included, natural or artificial circular RNAs- or CRISPR-based approaches. Circular RNAs can be designed to bind RBDs of RBPs and compete out target RNAs. CRISPR based approaches can be used to create oncogenic RBP knockout/knockin (to correct cancer-specific mutation in RBP or RBP binding sites). RISC, RNA-induced silencing complex.
4.1. Small molecules
Small molecules are by far the most commonly used compounds to target RBP function in cancers and other human diseases. Several of these molecules have shown promising anti-cancer activities in pre-clinical trials and some of them are currently being evaluated in clinical trials. Small-molecules can be used to inhibit RBP function in one of the following ways: 1) bind to the RBD and subsequently, impede RBP-RNA interaction; 2) disrupt interaction between RBP and its protein partner essential for its function in cancer; 3) prevent functional modifications of RBPs by inhibiting the activity of the enzymes involved; 4) modulate splicing of cancer causing genes by targeting core spliceosome machinery; 5) inhibit enzymatic activity of an RBP towards its target mRNA; 6) induce RBP degradation through targeting its E3 ubiquitin ligase. Here, we will provide a few examples for each strategy.
4.1.1. Compounds targeting eIF4E
The eukaryotic translation initiator factor 4E (eIF4E) is known to play an instrumental role in tumorigenesis and cancer progression and has been extensively studied as a target for cancer therapeutics. To date, several strategies have been applied to design small molecules that target eIF4E directly or indirectly for cancer management. The first approach is to target the binding of eIF4E to the cap structure. For example, ribavirin is a guanosine ribonucleoside analog that was initially found to mimic the cap structure and subsequently, compete with endogenous mRNAs for their binding with eIF4E (Kentsis, Topisirovic, Culjkovic, Shao, & Borden, 2004). Although the cap-binding mechanism of Ribavirin has been challenged by two different groups (Westman et al., 2005; Y. Yan, Svitkin, Lee, Bisaillon, & Pelletier, 2005), it showed promising pre-clinical efficacy and also potent effect in clinical trials for AML and metastatic breast cancer (Assouline et al., 2009); however, it failed to show efficacy as a single agent in multiple antitumor screens involving xenograft models. Other cap-binding agonists include 4Ei-1 and Bn7-GMP (a synthetic nucleotide derivative that blocks the binding of eIF4E to the mRNA cap) (Ghosh et al., 2009; Jia et al., 2010), both of which are still under laboratory development stage and need to be further tested. Notably, 4Ei-1 has shown potent inhibition of cap-dependent translation both in vitro and in vivo in zebrafish embryos (Ghosh et al., 2009). More importantly, in combination with other drugs, 4Ei-1 has shown potent antitumor activities in breast, lung and mesothelioma cells (E. Z. Chen et al., 2014; S. Li et al., 2013). The second approach is to target the interaction between eIF4E and eIF4G, which is known to be critical for cap-dependent translation (Martineau, Azar, Bousquet, & Pyronnet, 2013; J. D. Richter & Sonenberg, 2005). Thus, several small molecules are designed to disrupt the interaction between eIF4E and eIF4G, including 4EGI-1, 4E1RCat, and 4E2RCat (Cencic, Desforges, et al., 2011; Cencic, Hall, et al., 2011; Moerke et al., 2007). All these compounds have shown cogent anti-tumor effects on cancer cells in vitro, breast cancer xenograft models in vivo and reverted chemoresistance in MYC-driven murine lymphoma models (Cencic, Hall, et al., 2011; Moerke et al., 2007). In addition to above compounds, two more eIF4E–eIF4G inhibitors were developed, including ouabain and perillyl alcohol (Cao et al., 2014; Peffley, Sharma, Hentosh, & Buechler, 2007). Ouabain is a cardiac glycoside and can disrupt eIF4E/eIF4G association in a panel of human cancer cells (Cao et al., 2014). Perillyl alcohol is a naturally occurring monoterpene and was found to attenuate interactions between eIF4E and eIF4G in prostate cancer cells (Peffley et al., 2007). The third approach is to sequester eIF4E by modulating the activity of 4E-BP, which is known to inhibit translation initiation by competing with eIF4G for a common binding site on eIF4E. As 4EBPs are direct substrates of mTOR kinase, targeting mTOR signaling pathway is a therapeutically attractive option for the purpose of sequestering eIF4E. mTOR kinase activity inhibition by small-molecule inhibitors has shown strong anti-cancer activities. Although the first generation of mTOR inhibitors (Rapamycin, RAD001, and CCI-779) showed potential efficacy in preclinical trials, they performed poorly in clinical trials against various human cancers due to poor inhibition of 4EBP phosphorylation (Hsieh & Ruggero, 2010). Next generation of potent mTOR inhibitors such as PP242, Ku-0083784, Torin1, Palomid 529, WYE-132, and AZD8055 function as ATP-active site inhibitors on mTOR and have shown more strong inhibition of phosphorylation of all mTOR targets including 4EBPs (Chresta et al., 2010; Feldman et al., 2009; García-Martínez et al., 2009; Thoreen et al., 2009; Q. Xue et al., 2008; Yu et al., 2010). Many of these mTOR active site inhibitors and their more potent derivatives are being currently tested in clinical trials and promise a cogent therapeutic potential. The fourth approach is to block eIF4E phosphorylation by Mnks, which is known to be critical for oncogenic activity of eIF4E (Topisirovic, Ruiz-Gutierrez, & Borden, 2004; Wendel et al., 2007). MNK inhibitors such as (5-(2-(-(phenylamino)pyrimidin-4-yl)thiazol-2(3H)-one derivatives, cercosporamide, and CGP57380 have shown significant inhibition of eIF4E phosphorylation and subsequent reduction in cancer cell proliferation by inducing apoptosis. Moreover, in animal models, cercosporamide treatment blocked eIF4E phosphorylation in normal and xenograft tissues, suppressed melanoma metastasis and decreased growth of AML xenografted tumors (Altman et al., 2013; Konicek et al., 2011). Most importantly, cercosporamide also suppressed colonization of freshly explanted AML patient samples (Altman et al., 2013).
4.1.2. Compounds targeting the DEAD/H box RNA helicases
The DEAD/H box RNA helicases, such as eIF4A (also called DDX2) and DDX3, are known to modulate translation of many oncogenes and thus, are being explored as targets for cancer management. Several small molecules have been developed to target this protein family by affecting the RNA binding and helicase activity or by disrupting specific protein-protein interactions (Bhat et al., 2015; Cai et al., 2017). For example, the activity of eIF4A can be blocked by several natural compounds, including silvestrol, pateamine A, and hippuristanol. All these compounds have shown potent growth inhibition in various human cancer cell lines and in mouse xenograft models and are currently being tested at pre-clinical stage (Alachkar et al., 2013; Chu et al., 2016; Iwasaki, Floor, & Ingolia, 2016; Iwasaki et al., 2019; S. Kim et al., 2007; Kuznetsov et al., 2009; Tsumuraya et al., 2011; Wolfe et al., 2014). Both silvestrol and pateamine A act as chemical inducers to stimulate eIF4A:RNA binding, which subsequently, sequesters eIF4A and inhibits the incorporation of eIF4A into the eIF4F translation initiation complex (Bordeleau, Cencic, et al., 2006; Bordeleau et al., 2008). Unlike silvestrol and pateamine A, hippuristanol prevents free eIF4A and eIF4F-bound eIF4A from interacting with RNA and thereby, inhibits translation initiation (Bordeleau, Mori, et al., 2006). Additionally, another compound, elatol was identified as a potent inhibitor of eIF4A with in vivo antitumor activity (Peters et al., 2018). Elatol specifically inhibits eIF4A helicase activity and shows no activity against other tested ATP-hydrolyzing enzymes, including other DEAD box helicases, or kinases (Peters et al., 2018).
In addition to eIF4A, RNA helicase DDX3 was found to be an attractive target for cancer therapeutic as it promotes tumor cell proliferation (Bol, Xie, & Raman, 2015). Several compounds have been discovered to inhibit either its helicase or ATPase activity. For example, RK-33 was found to specifically bind to DDX3 and abrogate its helicase activity (Kondaskar et al., 2010). Importantly, both RK-33 and nanoparticle formation of RK-33 was found to significantly inhibit tumor growth in multiple mouse models of lung cancer (Bol, Khan, et al., 2015; Bol, Xie, et al., 2015). In addition, several ring-expanded nucleoside analogues were found to target the RNA binding site of the human DEAD-Box RNA helicase DDX3 (Radi et al., 2012). However, further studies are needed to test whether these compounds have antitumor activity. More recently, drugs targeting the helicase Brr2 have been identified (M. Ito et al., 2017; Iwatani-Yoshihara et al., 2017); however, their antitumor efficacy is yet to be determined.
4.1.3. Compounds targeting HuR
As the most extensively studied RBP in cancer, HuR (Hu antigen R) is found to be aberrantly expressed in many types of tumor and is a promising target for cancer therapeutics. In 2007, Meisner et al discovered the first small molecular inhibitors, MS-444, dehydromutactin and okicenone, which interfere with the RNA binding activity of HuR (Meisner et al., 2007). Among these three inhibitors, MS-444 was the most potent and inhibited binding of HuR to target AREs by interfering with HuR dimerization. Further work with this compound demonstrated both in vitro and in vivo antitumor activity against colorectal cancer, reverted TRAIL resistance in pancreatic cancer and reduced glioma cell survival by inhibiting HuR multimerization (Blanco et al., 2016; Filippova et al., 2017; Lang et al., 2017; Romeo et al., 2016). In addition, two chemical compounds, quercetin and b-40, were identified by their ability to inhibit the formation of HuR:ARE (TNF-α) complex and thereby, decrease TNFα mRNA stability and secreted TNFα (Chae et al., 2009; D’Agostino, Adami, & Provenzani, 2013). Nevertheless, the anti-tumor activities of these compounds need to be further tested as they function as anti-inflammatory agents (C. Guo, Yang, Jang, Zhou, & Liu, 2017). More recently, Wu et al reported that small molecule CMLD2 disrupts the interaction between HuR protein and its target mRNA, such as musashi (Xiaoqing Wu et al., 2015). CMLD-2 is a coumarin-derived molecule that was identified through high throughput screening of a library of 6,000 compounds developed by the Kansas University Chemical Methodologies and Library Development (CMLD) center. Later studies indicate that CMLD-2 exhibits cytotoxicity in a panel of cancer cell lines (R Muralidharan et al., 2017).
4.1.4. Compounds targeting Musashi proteins
Musashi proteins (MSI1 and MSI2) that are consistently over-expressed in a majority of the cancers also make an important target for therapy (Kudinov et al., 2017). Various groups have used high throughput screening assays to identify compounds that block the binding of Musashi proteins to its target RNAs. Using fluorescence polarization assay, Clingman et al. identified naturally occurring molecule oleic acid as the most potent inhibitor of Musashi binding to RNAs from two small molecule libraries that together consisted of more than 31000 compounds (Clingman et al., 2014). Oleic acid bound to the RRM1 of Musashi proteins and induced a conformational change that inhibited its binding to the target mRNA. Oleic acid was also shown to inhibit proliferation of MSI1 containing cells and provided a potential feedback loop between MSI1 activity and fatty acid biosynthesis which could be further studied and exploited to target MSI1 protein (Clingman et al., 2014). Using a smaller library, another group simultaneously identified three compounds that could inhibit Musashi association to RNA and these compounds had low IC50 (Minuesa et al., 2014). Although these compounds inhibit Musashi-RNA interaction, their effect on cell viability and proliferation has not been explored yet. Yet another group used RRM1 of MSI1 to screen for compounds that included FDA approved anti-cancer drugs and natural products. They identified the natural compound (−)-gossypol derived from cottonseed as their top hit with Ki in the nanomolar range (Lan et al., 2015). (−)-gossypol is known to have potent anti-tumor activity and has been used in several clinical trials targeting prostate cancer, NSCLC, glioblastoma, adenocortical carcinoma, CLL and other cancers either alone or in combination with other drugs.
4.1.5. Compounds targeting other RBPs
In addition to above mentioned compounds, several other compounds were also developed to target various RBPs. The RBP Lin28 regulates the maturation of the let-7 family of tumor-suppressor microRNAs by interacting with let-7 precursors through its two distinct cold shock and zinc-knuckle domains. Recently, two different groups discovered small-molecules that inhibit LIN28 binding to let-7 precursors. The first group utilized a protein/RNA FRET assay to screen for 16000 molecules and identified a molecule N-methyl-N-[3-(3-methyl[1,2,4]triazolo[4,3-b]-pyridazin-6-yl)phenyl] acetamide (referred to as 1632) that blocked the Lin28/let-7 interaction. This compound showed increased processed let-7, induced differentiation in embryonic stem cells and exhibited promising anti-tumor activities on cancer cells (Roos et al., 2016). Furthermore, Wang and colleagues used fluorescence polarization-based high throughput screen and identified two compounds that selectively blocked the LIN28-mediated let-7 processing (L Wang et al., 2018). Specifically, compound TPEN acts as a potent inhibitor of the zinc-knuckle domain of LIN28, while compound LI71 inhibits LIN28-mediated oligouridylation by interacting with the cold shock domain of LIN28. Furthermore, they showed that LI71 suppresses LIN28 in leukemia cells and embryonic stem cells. However, neither of the studies examined the effect of LIN28 inhibition on normal or cancer cell proliferation. A recent study showed that compound 1632 can inhibit cancer cell proliferation only at higher doses (Yanlian Chen et al., 2019). Moreover, compound 1632 reduced tumor xenografts in mice without affecting the body weight, indicating cancer specific effect of LIN28 inhibition.
Another recent study has identified Sam68 as a very potent target of ICG-001/CWP family of anti-tumor drugs. The authors for the first time demonstrated the role of Sam68 as a specific regulator of cancer stem cells compared to normal stem cells (Benoit et al., 2017). Sam68 was shown to be specifically overexpressed in colon, blood and breast cancer stem cells. The CWP drugs selectively inhibited self-renewal potential of patient derived leukemia stem cells as compared to healthy hematopoietic stem cells by inducing a complex formation between Sam68 and CBP leading to inhibition of the Wnt signaling pathway. More importantly, the Sam68/CBP complex formation was only induced by the drug in the cancer stem cells, thus providing specificity for killing cancer cells.
More recently, a novel approach of targeting RBPs in cancer was discovered serendipitously. Although anticancer sulfonamides such as indisulam have been used in clinical trials to treat advanced stage solid tumors, their mechanism of action was not known. Two recent studies demonstrated that indisulam increased the affinity of the E3 ubiquitin ligase DCAF15 for its substrate RBM39, resulting in the degradation of RBM39 in cancers and subsequent killing of cancer cells (T. Han et al., 2017; Uehara et al., 2017). Importantly, the treatment efficacy of indisulam was dependent on the levels of DCAF15 in patient tumors. Although indisulam has already shown modest efficacy in clinical trials of patients with metastatic disease, this compound did not advance because of its limited efficacy of response in about 10% of patients. Based on the finding that indisulam efficacy is proportional to DCAF15 expression in tumors, future clinical trials can focus on tumors expressing high levels of DCAF15 such as leukemia and lymphomas.
4.2. Oligonucleotide-based strategy
4.2.1. Antisense oligonucleotides (ASOs)
Antisense oligonucleotides (ASOs) are short (15–30bp), single stranded DNA analogs designed to bind to RNA by Watson-Crick base pairing. Upon binding to their target mRNAs, lncRNAs or miRNAs, ASOs perturb protein production by enhancing degradation of the targeted RNA (through RNaseH mediated degradation), altering RNA metabolism, or up-regulating gene expression. Being single stranded molecules, ASOs are very unstable and often need chemical modifications for stabilization. The benefit of ASOs include broad distribution and long-lasting effects. To date, six ASO drugs have been approved for the treatment of diseases such as viral infections, hyperlipidemias, and neurological diseases (Stein & Castanotto, 2017). Notably, several ASOs targeting RBPs have been designed for the treatment of neurodegenerative disorders. For example, spinal muscular atrophy (SMA) is a neurodegenerative disorder characterized by progressive loss of motor neurons, which is caused by loss or mutation of SMN1, an RBP involved in spliceosome. Interestingly, SMA patients carry a normal copy of SMN2, which is nearly identical to SMN1. However, the majority SMN2 gene products are truncated isoform excluding exon 7. Thus, an ASO, Nusinersen, was designed to prevent exon 7 exclusion by blocking the intronic splicing silencer (ISS) element in SMN2, resulting in the expression of the functional, full-length survival motor neuron protein from SMN2. In preclinical studies, Nusinersen was found to increase SMN2 levels and improved survival and neuromuscular function in SMA mice (Y. Hua et al., 2010, 2011; Rigo, Hua, Krainer, & Bennett, 2012). In December 2016, it became the first FDA approved drug for the treatment of SMA patients (Chiriboga et al., 2016; Mercuri et al., 2018). In addition to Nusinersen, ASO-based therapies are currently being tested to prevent RBP aggregation or alleviate RBP sequestration by repeat expansion in several neurodegenerative disorders (Nussbacher, Tabet, Yeo, & Lagier-Tourenne, 2019). Although there are no approved ASO drugs for cancer treatment, many attempts are being made. Indeed, ASOs against RBPs are being tested clinically and preclinically. For example, ASO against eIF4E [LY2275796] has been developed and has completed a phase I clinical trial (Hong et al., 2011). Phase II clinical trials are now under way combining ISIS EIF4E Rx with carboplatin and paclitaxel for non–small cell lung cancer (NCT01234038) or with docetaxel and prednisone for castration-resistant prostate cancer (NCT01234025). Similarly, another eIF4E ASO, ISIS 183750, showed significant tumor suppression in a mouse model and has also been tested in phase I/II clinical trials in patients with advanced colorectal cancer (Duffy et al., 2016). Additionally, ASO against Musashi was recently developed and found to effectively block tumor growth in preclinical models of pancreatic cancer (R G Fox et al., 2016). Interestingly, ASOs against Hu family proteins were also developed; however, they have not been tested for antitumor activities. To this end, ASO depletion of HuD via intrathecal injection reverts persistent pain in an animal model of antiretroviral therapy (Sanna, Peroni, Quattrone, Ghelardini, & Galeotti, 2015). Similarly, intrathecal ASO depletion of HuR attenuates mechanical allodynia in a model of autoimmune encephalomyelitis (Sanna, Quattrone, & Galeotti, 2017). Additionally, CPEB ASO treatment reduced hyperalgesic response to PGE2 or mechanical allodynia in rats (Bogen, Alessandri-Haber, Chu, Gear, & Levine, 2012; Iida et al., 2016).
4.2.2. siRNAs
Targeted delivery of small interfering RNA (siRNA)-nanoparticles for cancer treatment was first reported in 2010 and laid a solid foundation for RNA interference therapy (Davis et al., 2010). After almost two decades of research, siRNA-based therapies have demonstrated the efficacy and clinical safety and represent one of the more clinically advanced platforms for RNA drugs (Chi, Gatti, & Papoian, 2017). In 2018, FDA approved the first siRNA-based drug, Onpattro® as treatment for symptoms of peripheral nerve disease (polyneuropathy), caused by hereditary transthyretin amyloidosis in adult patients (W. Yin & Rogge, 2019). Notably, siRNA-based therapies have also been applied to target RBPs and have shown good promise in pre-clinical studies. For example, HuR has been recognized as a potential molecular target for RNAi therapy in several types of cancers such as NSCLC and ovarian cancer (Amreddy et al., 2018; Y. H. Huang et al., 2016). HuR siRNA has been tested for its efficacy to target various cancers via different target delivery methods. For example, HuR-targeted siRNA greatly reduced HuR expression and significantly inhibited the cell proliferation in lung cancer cells but not in normal cells (Amreddy et al., 2018). To achieve targeted delivery, a folate-conjugated nanoparticle system was developed recently and showed that silencing HuR suppresses lung cancer cell proliferation and migration. The rationale to use folate-conjugated nanoparticle was to reduce non-specific toxicity to normal cells and increase the therapeutic efficacy against tumor cells as majority of cancer therapies that are currently available are limited by cytotoxicity to normal tissues (Amreddy et al., 2018). More recently, another liposome nanoparticle delivery system was developed by using transferrin as a targeting ligand and showed to reduce the tumor burden in mouse models of lung cancer (Ranganayaki Muralidharan et al., 2017). Importantly, this therapy was also able to reduce the metastasis in A549 based metastasis mouse model compared to control siRNA-based nanoparticles. Additionally, HuR targeting by siRNA has been shown to impair malignant characteristics of pancreatic ductal adenocarcinoma cells (Jimbo et al., 2015). Moreover, HuR silencing sensitizes triple negative breast cancer cells to radiotherapy due to induction of oxidative stress and DNA damage (Mehta et al., 2016). Furthermore, conjugation of siRNAs against HuR with Cy3-labeled folic acid-coated DNA dendrimer nanocarrier has shown dramatic anti-oncogenic activity in ovarian cancer (Y. H. Huang et al., 2016). In addition to HuR, siRNAs against eIF4E have also been tested for their efficacy in multiple tumors. For example, a tumor-specific siRNA against eIF4E showed potent tumor suppression and enhanced cisplatin cytotoxicity in human breast carcinoma cells (Dong et al., 2009). Additionally, siRNAs against Musashi were found to inhibit growth of a panel of cancer cells in vivo and in vitro (H. Chen et al., 2019; Sheng et al., 2016; Sureban et al., 2008).
4.4. Aptamers
Aptamers represent another strategy used for targeting RBPs in cancer, albeit less frequently. Aptamers can be either nucleic acid or peptide based and are selected on the basis of binding to a specific target (Mascini, Palchetti, & Tombelli, 2012). They fold into sequence specific three-dimensional structures that can recognize their unique target. They function similar to antibodies while also possessing several desirable properties such as smaller size, better stability and no immunogenicity (J. Zhou & Rossi, 2017). Nucleic acid aptamers consist of short, single-stranded DNA or RNA. Nucleic acid aptamers are selected using the procedure SELEX (systemic evolution of ligands by exponential enrichment), whereas peptide aptamers are selected using yeast two hybrid technique, phage display, mRNA display or other techniques (Mascini et al., 2012). The first and only aptamer approved by FDA is the anti-VEGF aptamer pegaptanib (Macugen®; Pfizer/Eyetech) for treatment of macular degeneration in 2004. Although initially designed for use in cancer treatment, it did not show efficacy in pre-clinical studies and hence has not been tested in a clinical trial for cancer. While several aptamers are in the development stages as anti-cancer agents, only two therapeutic aptamers have made it to clinical trials for the treatment of cancer (Morita, Leslie, Kameyama, Volk, & Tanaka, 2018). One of them is the aptamer AS1411 (previously called AGRO100) against the RBP nulceolin and it was the first aptamer in clinical trial for cancer treatment. Nucleolin regulates several essential cellular processes including regulation of RNA polymerase I transcription, maturation and proper folding of the pre-ribosomal RNA, mRNA translation, mRNA stability and is overexpressed in cancers (Berger, Gaume, & Bouvet, 2015). AS1411, is a 26-nucleotide DNA based aptamer that forms stable G-quadruplex structures which are resistant to nucleases. AS1411 binds to the external domain of nucleolin and has shown growth inhibition in vitro and in vivo xenograft models of breast, lung and renal cancers (Ireson & Kelland, 2006). More importantly, AS1411 has been assessed in clinical trials for safety and efficacy in advanced solid tumors (NCT00881244) and renal cell carcinoma (NCT00740441). In its initial dose-escalation phase I clinical study, the administered doses of AS1411 were well tolerated and did not show any serious side effects. Based on these results, a Phase II trial was performed in patients with refractory metastatic renal cell carcinoma, but it showed limited efficacy in unselected patients implying the need for the discovery of AS1411 responsive biomarkers in tumors. Yet another Phase II clinical trial assessed the efficacy of AS1411 in combination with anti-cancer drug cytarabine in relapsed or refractory AML patients (NCT00512083). This study found improved response rates in patients receiving the combination therapy compared to the cytarabine alone group, indicating anti-tumor effect of AS1411 in AML. In addition to nucleolin, aptamers have also been developed against the DEAD-box RNA helicase eIF4A, the RNA editing enzyme ADAR1 and the dsRNA-dependent protein kinase PKR that inhibit their helicase activity, RNA editing or dsRNA binding ability, respectively (Hallegger, Taschner, & Jantsch, 2006; Oguro, Ohtsu, Svitkin, Sonenberg, & Nakamura, 2003; Zheng & Bevilacqua, 2004). Although these aptamers have not been used as anti-cancer agents in clinical trials, based on the promising clinical efficacy of AS1411, there remains a potential for the development of these and other RBP-based aptamers for anti-cancer therapeutics.
4.5. Peptides
Therapeutic peptides are receiving increasing attention for biomedical applications. Peptides present promising opportunities in targeted drug delivery due to their high specificity, selectivity, small dimension, ease of modification, and high biocompatibility. During the last decade, the potential of peptides as therapeutics or targeting ligands has boomed in nanotechnology and cancer research. There are several types of peptides being developed for cancer treatment (Marqus, Pirogova, & Piva, 2017) and some of these peptide are designed to target RBPs. For example, Yo and colleagues developed several 4EBP-based peptides that bound eIF4E and subsequently, disrupted the interaction of eIF4E to eIF4G, leading to inhibited cap-dependent translation (Y. K. Song, Guo, Barengo, & Naora, 2009). Notably, when fused with gonadotropin-releasing hormone agonist, these peptides exerted potent antitumor effects in a mouse xenograft model of epithelial ovarian cancer without eliciting significant toxicity. Recently, we also developed a small peptide that can inhibit tumor growth in vitro and in vivo (Lucchesi, Zhang, Ma, Chen, & Chen, 2019). The rationale is based on our previous finding that RBM38 inhibits p53 mRNA translation by preventing the binding of eIF4E to the p53 mRNA (J Zhang et al., 2011). Thus, we designed synthetic peptides corresponding to the binding interface between RBM38 and eIF4E and showed that the peptides were effective in relieving RBM38-mediated repression of p53 translation. Notably, we showed that this peptide alone or together with a low dose of doxorubicin potently induced p53 expression and suppressed colony formation, tumor sphere formation and xenograft tumors in RBM38- and p53-dependent manners.
4.6. Other potential strategies to target RBPs in cancer
4.6.1. Circular RNAs (circRNAs)
Circular RNAs are naturally occurring covalently closed circle RNAs found in majority of eukaryotic cells that are formed by back-splicing of exons (Kristensen, Hansen, Venø, & Kjems, 2018). Although discovered 40 years ago, until recently they were considered the product of splicing errors (M. T. Hsu & Coca-Prados, 1979). Following recent studies that showed circRNAs are functional, conserved and that their formation is non-random (Hansen et al., 2013; Jeck et al., 2013; Memczak et al., 2013), there has been an explosion in studies describing the vital roles of circRNAs in normal physiology and disease. Several studies have described the role of circRNAs in cancer development. circRNAs function in cancers by acting as miRNA or RBP sponges, by regulating transcription and splicing, and can also undergo translation. Interestingly, one circRNA may contain several sites for a single or multiple RBPs and thus regulate the function of RBPs by acting as an RBP sponge or decoy (Kristensen et al., 2018). This phenomenon is observed in the binding of circRNAs derived from muscleblind (mbl) locus to MBL protein itself. The binding of circRNA inhibits MBL function by preventing its binding to its mRNA targets (Ashwal-Fluss et al., 2014). Another example of circRNAs acting as an RBP sponge comes from the binding of circRNA derived from PABPN1 gene to HuR. HuR has been shown to bind PABPN1 mRNA and increase its translation; however, binding of circPABPN1 to HuR prevents its binding to PABPN1 mRNA, resulting in decreased PABPN1 translation (Abdelmohsen et al., 2017). More importantly, sequestering of HuR by cicrPABPN1 may also affect other cancer relevant HuR targets such p53, c-myc and bcl-2. In addition to this, circRNAs are extremely stable and have a long half-life in cells. These properties of circRNAs make them a promising therapeutic strategy for targeting RBP functions in cancer. Natural or recombinant circRNAs capable of sequestering oncogenic RBPs can thus be used to treat cancers overexpressing the particular RBP. Although, there have been no preclinical reports that have utilized circRNAs sequestering RBPs as therapeutic molecules for cancer treatment, this is likely going to change in the near future.
4.6.2. CRISPR based strategies
In the past decade, the development of CRISPR/Cas9 system has shown potential therapeutic applications for the treatment of cancer. Several clinical trials are ongoing that are testing efficacy of CRISPR based therapy for cancer treatment. Majority of the clinical trials are immunotherapies that are targeting the negative immune regulator protein program cell death 1 (PD-1) (X. Tian et al., 2019). In addition, several studies have utilized CRISPR based therapies for the treatment of human diseases including cancer in human patient cells. For example, a recent study utilized CRISPR to correct the autoimmune disease causing IL2RA mutation in human patients and showed promising results (Roth et al., 2018). The same study also used CRISPR to replace endogenous T-cell receptor with a new T-cell receptor that assisted T-cells towards cancer antigens (Roth et al., 2018). Another study showed that RNA-targeting CRISPR system induced specific degradation of microsatellite repeat expansion RNAs, caused by dysfunctional RNA splicing in various neurodegenerative disease bearing patient cells (Batra et al., 2017). Although CRISPR/Cas9 system has not yet been used to target RBPs for cancer therapy, the above studies provide a promising future for their use in cancer therapy. CRISPR can be used to directly target an RBP or its function by different means. For example, it can be used to knockout an oncogenic RBP in cancer cells, to correct cancer specific RBP mutations or modulate RBP binding sites in mRNAs that would lead to aberrant splicing of oncogenes. Similarly, they could be used to target immune-suppressive RBPs in immunotherapies in combination with checkpoint inhibitors. Thus, although not used for targeting RBPs in preclinical cancer studies yet, CRISPR represents an encouraging approach for developing RBP-targeted cancer therapies.
5. Conclusions And Future Perspectives
RBPs control multiple cancer traits by regulating several genes that form an intricate network and multitude of studies have shown an indisputable role for RBPs in tumorigenesis. Thus, RBPs are emerging as promising targets for cancer therapy. Of particular interest, recent studies suggest a role of RBPs in cancer immunotherapy. Cancer immunotherapy works by boosting patients own immune system to attack tumor cells by using immune checkpoint inhibitors (ICIs) (Topalian, Drake, & Pardoll, 2015). The ICIs usually target the inhibitory CTLA4 and PD1 receptors present on the T-cells or PD-L1 ligand present on tumor microenvironment immune cells or overexpressed on tumor cells. Although immunotherapy has shown promising clinical outcomes in several patients, some patients still fail to respond due to development of resistance (Jenkins, Barbie, & Flaherty, 2018). Interestingly, several RBPs have been involved in pathways that lead to immunotherapy resistance either by remodeling tumor cells to evade immune response or by working in immune cells to blunt the immune cell’s efficacy to fight tumor cells. For example, the RBP MEX3B has been shown to destabilize HLA-A mRNA in tumor cells and mediate resistance to cancer immunotherapy (L. Huang et al., 2018). Similarly, the oncogenic RBP IMP3 (IGF2BP3) was shown to decrease stress-induced ligands ULBP2 and MCIB, leading to impaired recognition of transformed cells by NK cells (Schmiedel et al., 2016). In addition, ADAR1 plays a role in immune resistance by regulating the biogenesis of miR-222 and thereby ICAM1 protein expression (Galore-Haskel et al., 2015). Moreover, eIF4E has been recently shown to enhance the translation of PD-L1 in mouse tumors, which was abrogated by eFT508, the eIF4E phosphorylation inhibitor (Yichen Xu et al., 2019). Together, these studies suggest that RBPs are promising targets to increase the efficacy of immunotherapy in patients due to their role in immune suppression/evasion.
To date, only a few RBPs have been targeted for cancer therapy, by using various approaches, in preclinical and clinical studies. The major challenges include: (1) the delivery methods to target RBP in cancer need to be improved; (2) Selective targeting of cancer cells as most of the RBPs are required for the survival of both normal and cancer cells; (3) identify subsets of cancers that are highly dependent on a particular RBP for proliferation and survival; (4) the biological function of most RBPs in cancer has not been fully understood yet. For example, some RBPs work as tumor suppressors under one circumstance but as an oncogene under another circumstance. Additionally, it is in urgent need to identify critical targets of an RBP that mediate its cancer-related functions. To overcome these challenges, several strategies may be applied. For example, to develop targeted therapies that kill cancer cells, it will be important to design nanoparticle carriers for siRNA, ASOs, aptamers and peptides that recognize altered cancer-specific cell-surface markers and thus can be targeted specifically to cancer cells. Second, it will also be important to develop small molecules or peptides that inhibit the interaction of an RBP with only a subset of its targets that are cancer relevant. Third, it will be important to identify cancer-specific PTM(s) of an RBP and the kinase(s) involved in different cancer types. For example, phosphorylation of eIF4E specifically increases translation of oncogenic mRNAs and consistent with this, inhibition of eIF4E phosphorylation by small molecule inhibitors only affects the translation of oncogenic mRNAs in cancer without affecting normal cell proliferation. Fourth, the techniques such as PAR-CLIP, RIP-seq, iCLIP and others will prove valuable in defining the targets of an RBP or its interacting partners in cancer, which would lay a foundation for the development of novel cancer therapeutics. Finally, recent studies have revealed a critical role of non-canonical RBPs in regulating RNA metabolism and fate and have been implication in cancer. Thus, it is important to consider the role of these non-canonical RBPs and their cross talks with canonical RBPs. A key challenge here would be defining the targets of non-canonical RBPs in cancer as these cannot be predicted bioinformatically due to lack of available structure-function studies. In summary, considering that the number of RBPs whose functions are still not completely understood, our current knowledge about cancer-related RBPs seems to be the tip of the iceberg and reinforces the need for substantial work to be done in the future.
Acknowledgements
The authors apologize to those authors whose work they could not cite due to space constraints. This work is supported in part by the NIH grants (CA224433, CA081237, and CA195828 to X.B. Chen).
Abbreviations
- 3’UTR
3’ untranslated region
- 5’UTR
5’ untranslated region
- AML
Acute myeloid leukemia
- ARE
AU-rich element
- ASO
Antisense oligonucleotide
- circRNAs
Circular RNAs
- CML
Chronic myeloid leukemia
- dsRBD
Double-stranded RNA binding domain
- dsRNA
Double-stranded RNA
- hnRNP
heterogeneous nuclear ribonucleoprotein
- KH
K-homology
- miRNAs
MicroRNAs
- mRNAs
Messenger RNAs
- PAZ
Piwi/Argonaute/Zwille
- PCBP
poly(C)-binding protein
- PIWI
P-element Induced WImpy testis
- PUM
Pumilio homology domain
- RBD
RNA-binding domain
- RBP
RNA-binding protein
- RNP
Ribonucleoprotein
- RRM
RNA recognition motif
- siRNA
Small interfering RNA
- TTP
Tristetraprolin
- UTR
Untranslated region
- ZNF
Zinc finger
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflict of Interest Statement
The authors declare that there are no conflicts of interest.
References
- Abba MC, Sun H, Hawkins KA, Drake JA, Hu Y, Nunez MI, … Aldaz CM (2007). Breast cancer molecular s ign atures as determined by SAGE: correlation with lymph node status. Mol Cancer Res, 5(9), 881–890. 10.1158/1541-7786.MCR-07-0055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abdelmohsen K, & Gorospe M (2010). Posttranscriptional regulation of cancer traits by HuR. Wiley Interdisciplinary Reviews: RNA 10.1002/wrna.4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abdelmohsen K, Panda AC, Munk R, Grammatikakis I, Dudekula DB, De S, … Gorospe M (2017). Identification of Hu R target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biology, 14(3), 361–369. 10.1080/15476286.2017.1279788 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abdelmohsen K, Tominaga-Yamanaka K, Srikantan S, Yoon JH, Kang MJ, & Gorospe M (2012). RNA-binding protein AUF1 represses Dicer e xpression. Nucleic Acids Research, 40(22), 11531–11544. 10.1093/nar/gks930 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adolph D, Flach N, Mueller K, Ostareck DH, & Ostareck-Lederer A (2007). Deciphering the cross talk between hnRNP K and c-Src: the c-Src activation domain in hnRNP K is distinct from a second interaction site. Mol Cell Biol, 27(5), 1758–1770. 10.1128/MCB.02014-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akao Y, Marukawa O, Morikawa H, Nakao K, Kamei M, Hachiya T, & Tsujimoto Y (1995). The rck/p54 Candidate Proto-oncogene Product Is a 54-Kilodalton D-E-A-D Box Protein Differentially Expressed in Human and Mouse Tissues. Cancer Research, 55(15), 3444–3449. [PubMed] [Google Scholar]
- Alachkar H, Santhanam R, Harb JG, Lucas DM, Oaks JJ, Hickey CJ, … Marcucci G (2013). Silvestrol exhibits significant in vivo and in vitro antileukemic activities and inhibits FLT3 and miR-155 expressions in acute myeloid leukemia. J Hematol Oncol, 6, 21 10.1186/1756-8722-6-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alajez NM, Shi W, Wong D, Lenarduzzi M, Waldron J, Weinreb I, & Liu F-F (2015). Lin28b Promotes Head and Neck Cancer Progression via Modulation of the Insulin-Like Growth Factor Survival Pathway. Oncotarget, 3(12). 10.18632/oncotarget.785 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allain FH, Bouvet P, Dieckmann T, & Feigon J (2000). Molecular basis of sequence-specific recognition of pre-ribosomal RNA by nucleolin. EMBO J, 19(24), 6870–6881. 10.1093/emboj/19.24.6870 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altman JK, Szilard A, Konicek BW, Iversen PW, Kroczynska B, Glaser H, … Platanias LC (2013). Inhibition of Mnk kinase activity by cercosporamide and suppressive effects on acute myeloid leukemia precursors. Blood, 121(18), 3675–3681. 10.1182/blood-2013-01-477216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amin EM, Liu Y, Deng S, Tan KS, Chudgar N, Mayo MW, … Jones DR (2017). The RNA -editing enzyme ADAR promotes lung adenocarcinoma migration and invasion by stabilizing FAK. Science Signaling, 10(497). 10.1126/scisignal.aah3941 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amreddy N, Babu A, Panneerselvam J, Srivastava A, Muralidharan R, Chen A, … Ramesh R (2018). Chemo -biologic combinatorial drug delivery using folate receptor-targeted dendrimer nanoparticles for lung cancer treatment. Nanomedicine: Nanotechnology, Biology, and Medicine, 14(2), 373–384. 10.1016/j.nano.2017.11.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen CBF, Ballut L, Johansen JS, Chamieh H, Nielsen KH, Oliveira CLP, … Andersen GR (2006). Structure of the e xon junction core complex with a trapped DEAD-Box ATPase bound to RNA. Science, 313(5795), 1968–1972. 10.1126/science.1131981 [DOI] [PubMed] [Google Scholar]
- Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, … Kadener S (2014). Circ RNA Biogenesis competes with Pre-mRNA splicing. Molecular Cell, 56(1), 55–66. 10.1016/j.molcel.2014.08.019 [DOI] [PubMed] [Google Scholar]
- Assouline S, Culjkovic B, Cocolakis E, Rousseau C, Beslu N, Amri A, … Borden KL (2009). Molecular targeting o f the oncogene eIF4E in acute myeloid leukemia (AML): a proof-of-principle clinical trial with ribavirin. Blood, 114(2), 257–260. 10.1182/blood-2009-02-205153 [DOI] [PubMed] [Google Scholar]
- Baltz AG, Munschauer M, Schwanhausser B, Vasile A, Murakawa Y, Schueler M, … Landthaler M (2012). The mRNA -bound proteome and its global occupancy profile on protein-coding transcripts. Mol Cell, 46(5), 674–690. 10.1016/j.molcel.2012.05.021 [DOI] [PubMed] [Google Scholar]
- Baou M, Jewell A, & Murphy JJ (2009). TIS11 family proteins and their roles in posttranscriptional gene regulation. J Biomed Biotechnol, 2009, 634520 10.1155/2009/634520 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbouti A, Hoglund M, Johansson B, Lassen C, Nilsson P, Hagemeijer A, … Fioretos T (2003). Advances in Brief A Novel Gene , MSI2 , Encoding a Putative RNA-binding Protein Is Recurrently Rearranged at Disease Progression of Chronic Myeloid Leukemia and Forms a Fusion Gene with HOXA9 as a Result of the Cryptic t ( 7 ; 17 )( p15 ; q23 ) 1. Cancer Research, 63(6), 1202–1206. [PubMed] [Google Scholar]
- Baskin R, Woods NT, Mendoza-Fandiño G, Forsyth P, Egan KM, & Monteiro ANA (2015). Functional analysis of the 11q23.3 glioma susceptibility locus implicates PHLDB1 and DDX6 in glioma susceptibility. Scientific Reports, 5 10.1038/srep17367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batra R, Nelles DA, Pirie E, Blue SM, Marina RJ, Wang H, … Yeo GW (2017). Elimination of Toxic Microsatellite Repeat Expansion RNA by RNA-Targeting Cas9. Cell, 170(5), 899–912.e10 10.1016/j.cell.2017.07.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bava FA, Eliscovich C, Ferreira PG, Miñana B, Ben-Dov C, Guigó R, … Méndez R (2013). CPEB1 coordinates alternative 3′-UTR formation with translational regulation. Nature, 495(7439), 121–125. 10.1038/nature11901 [DOI] [PubMed] [Google Scholar]
- Beachy SH, Onozawa M, Chung YJ, Slape C, Bilke S, Francis P, … Aplan PD (2012). Enforced e xp ression of Lin 28b leads to impaired T-cell development, release of inflammatory cytokines, and peripheral T-cell lymphoma. Blood, 120(5), 1048–1059. 10.1182/blood-2012-01-401760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bechara EG, Sebestyén E, Bernardis I, Eyras E, & Va lcárcel J (2013). RBM5, 6, and 10 differentially regulate NUMB alternative splicing to control cancer cell proliferation. Molecular Cell, 52(5), 720–733. 10.1016/j.molcel.2013.11.010 [DOI] [PubMed] [Google Scholar]
- Beghini A, Ripamonti CB, Peterlongo P, Roversi G, Cairoli R, Morra E, & Larizza L (2000). RNA hyperediting and alternative splicing of hematopoietic cell phosphatase (PTPN6) gene in acute myeloid leukemia. Human Molecular Genetics, 9(15), 2297–2304. 10.1093/oxfordjournals.hmg.a018921 [DOI] [PubMed] [Google Scholar]
- Bell SE, Sanchez MJ, Spasic-Boskovic O, Santalucia T, Gambardella L, Burton GJ, … Turner M (2006). The RNA binding protein Zfp36l1 is require d for normal vascularisation and post-transcriptionally regulates VEGF expression. Dev Dyn, 235(11), 3144–3155. 10.1002/dvdy.20949 [DOI] [PubMed] [Google Scholar]
- Benoit YD, Mitchell RR, Risueño RM, Orlando L, Tanasijevic B, Boyd AL, … Bhatia M (2017). Sam68 Allows Selective Targeting of Human Cancer Stem Cells. Cell Chemical Biology, 24(7), 833–844.e9 10.1016/j.chembiol.2017.05.026 [DOI] [PubMed] [Google Scholar]
- Berger CM, Gaume X, & Bouvet P (2015). The roles of nucleolin subcellular localization in cancer. Biochimie 10.1016/j.biochi.2015.03.023 [DOI] [PubMed] [Google Scholar]
- Berglund JA, Chua K, Abovich N, Reed R, & Rosbash M (1997). The splicing factor BBP interacts specifically with the pre-mRNA branchpoint sequence UACUAAC. Cell, 89(5), 781–787. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/9182766 [DOI] [PubMed] [Google Scholar]
- Beuth B, Pennell S, Arnvig KB, Martin SR, & Taylor IA (2005). Structure of a Mycobacterium tuberculosis NusA -RNA complex. EMBO J, 24(20), 3576–3587. 10.1038/sj.emboj.7600829 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhat M, Robichaud N, Hulea L, Sonenberg N, Pelletier J, & Topisirovic I (2015). Targeting the translation machinery in cancer. Nature Reviews Drug Discovery 10.1038/nrd4505 [DOI] [PubMed] [Google Scholar]
- Bielli P, Busà R, Paronetto MP, & Sette C (2011). The RNA-binding protein Sam68 is a multifunctional player in human cancer. Endocrine-Related Cancer, 18(4), R91–R102. 10.1530/erc-11-0041 [DOI] [PubMed] [Google Scholar]
- Biewenga P, Buist MR, Moerland PD, van Themaat EVL, van Kampen AHC, ten Kate FJW, & Baas F (2008). Gene expression in early stage cervical cancer. Gynecologic Oncology, 108(3), 520–526. 10.1016/j.ygyno.2007.11.024 [DOI] [PubMed] [Google Scholar]
- Blackshear PJ (2002). Tristetraprolin and other CCCH tandem zinc-finger proteins in the regulation of mRNA turnover. Biochem Soc Trans, 30(Pt 6), 945–952. [DOI] [PubMed] [Google Scholar]
- Blackshear PJ, Phillips RS, Ghosh S, Ramos SB, Richfield EK, & Lai WS (2005). Zfp 36l3, a rodent X chromosome gene encoding a placenta-specific member of the Tristetraprolin family of CCCH tandem zinc finger proteins. Biol Reprod, 73(2), 297–307. 10.1095/biolreprod.105.040527 [DOI] [PubMed] [Google Scholar]
- Blanco FF, Preet R, Aguado A, Vishwakarma V, Stevens LE, Vyas A, … Dixon DA (2016). Impact of HuR inhibition by the small molecule MS-444 on colorectal cancer cell tumorigenesis. Oncotarget, 7(45). 10.18632/oncotarget.12189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bla xall BC, Dwyer-Nield LD, Bauer AK, Bohlmeyer TJ, Malkinson AM, & Port JD (2000). Differential e xpression and localizatio n of the mRNA binding proteins, AU-rich element mRNA binding protein (AUF1) and Hu antigen R (Hu R), in neoplastic lung tissue. Molecular Carcinogenesis, 28(2), 76–83. [DOI] [PubMed] [Google Scholar]
- Bogen O, Alessandri-Haber N, Chu C, Gear RW, & Levine JD (2012). Generation of a Pain Memory in the Primary Afferent Nociceptor Triggered by PKC Activation of CPEB. Journal of Neuroscience, 32(6), 2018–2026. 10.1523/jneurosci.5138-11.2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bol GM, Khan R, Heerma van Voss MR, Tantravedi S, Korz D, Kato Y, & Raman V (2015). PLGA nanoparticle formulation of RK-33: an RNA helicase inhibitor against DDX3. Cancer Chemother Pharmacol, 76(4), 821–827. 10.1007/s00280-015-2851-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bol GM, Vesuna F, Xie M, Zeng J, Aziz K, Gandhi N, … Raman V (2015). Targeting DDX3 with a small molecule inhibitor for lung cancer therapy. EMBO Mol Med, 7(5), 648–669. 10.15252/emmm.201404368 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bol GM, Xie M, & Raman V (2015). DDX3, a potential target for cancer treatment. Mol Cancer, 14, 188 10.1186/s12943-015-0461-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bordeleau ME, Cencic R, Lindqvist L, Oberer M, Northcote P, Wagner G, & Pelletier J (2006). RNA -mediated sequestration of the RNA helicase eIF4A by Pateamine A inhibits translation initiation. Chem Biol, 13(12), 1287–1295. 10.1016/j.chembiol.2006.10.005 [DOI] [PubMed] [Google Scholar]
- Bordeleau ME, Mori A, Oberer M, Lindqvist L, Chard LS, Higa T, … Pelletier J (2006). Functional characterization of IRESes by an inhibitor of the RNA helicase eIF4A. Nat Chem Biol, 2(4), 213–220. 10.1038/nchembio776 [DOI] [PubMed] [Google Scholar]
- Bordeleau ME, Robert F, Gerard B, Lindqvist L, Chen SM, Wendel HG, … Pelletier J (2008). Therapeutic supp ression of translation initiat ion modulates chemosensitivity in a mouse lymphoma model. J Clin Invest, 118(7), 2651–2660. 10.1172/JCI34753 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Botlagunta M, Vesuna F, Mironchik Y, Raman A, Lisok A, Winnard P, … Raman V (2008). Oncogenic role of DDX3 in b reast cancer biogenesis. Oncogene, 27(28), 3912–3922. 10.1038/onc.2008.33 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boukakis G, Patrinou-Georgoula M, Lekarakou M, Valavanis C, & Guialis A (2010). Deregulated expression of hnRNP A/B proteins in human non -small cell lung cancer: Parallel assessment of protein and mRNA levels in paired tumour/non-tumour tissues. BMC Cancer, 10 10.1186/1471-2407-10-434 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braddock DT, Louis JM, Baber JL, Levens D, & Clore GM (2002). Structure and dynamics of KH domains from FBP bound to single-stranded DNA. Nature, 415(6875), 1051–1056. 10.1038/4151051a [DOI] [PubMed] [Google Scholar]
- Braeutigam C, Rago L, Rolke A, Waldmeier L, Christofori G, & Winter J (2014). The RNA -binding protein Rbfox2: An essential regulator of EMT-driven alternative splicing and a mediator of cellular invasion. Oncogene, 33(9), 1082–1092. 10.1038/onc.2013.50 [DOI] [PubMed] [Google Scholar]
- Brennan CM, & Steitz JA (2001). HuR and mRNA stability. Cell Mol Life Sci, 58(2), 266–277. 10.1007/PL00000854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brody JR, & Dixon DA (2018). Complex HuR function in pancreatic cancer cells. Wiley Interdisciplinary Reviews: RNA, 9(3). 10.1002/wrna.1469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Budinska E, Popovici V, Tejpar S, D’Ario G, Lapique N, Sikora KO, … Delorenzi M (2013). Gene expression patterns unveil a new level of molecular heterogeneity in colorectal cancer. Journal of Pathology, 231(1), 63–76. 10.1002/path.4212 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burrows C, Abd Latip N, Lam SJ, Carpenter L, Sawicka K, Tzolovsky G, … Blagden SP (2010). The RNA binding protein Larp1 regulates cell division, apoptosis and cell migration. Nucleic Acids Research, 38(16), 5542–5553. 10.1093/nar/gkq294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Busa R, Paronetto MP, Farini D, Pierantozzi E, Botti F, Angelini DF, … Sette C (2007). The RNA -binding protein Sam68 contributes to proliferation and survival of human prostate cancer cells. Oncogene, 26(30), 4372–4382. 10.1038/sj.onc.1210224 [DOI] [PubMed] [Google Scholar]
- Bycroft M, Grunert S, Murzin AG, Proctor M, & St Johnston D (1995). NMR solution structure of a dsRNA binding domain from Drosophila staufen protein reveals homology to the N-terminal domain of ribosomal protein S5. EMBO J, 14(14), 3563–3571. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/7628456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bycroft M, Hubbard TJ, Proctor M, Freund SM, & Murzin AG (1997). The solution structure of the S1 RNA binding domain: a member of an ancient nucleic acid - binding fold. Cell, 88(2), 235–242. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/9008164 [DOI] [PubMed] [Google Scholar]
- Cai W, Xiong Chen Z, Rane G, Satendra Singh S, Choo Z, Wang C, … Prem Kumar A (2017). Wanted DEAD/ H or Alive: Helicases Winding Up in Cancers. J Natl Cancer Inst, 109(6). 10.1093/jnci/djw278 [DOI] [PubMed] [Google Scholar]
- Calderone V, Gallego J, Fernandez-Miranda G, Garcia-Pras E, Maillo C, Berzigotti A, … Mendez R (2016). Sequential Functions of CPEB1 and CPEB4 Regulate Pathologic Expression of Vascular Endothelial Growth Factor and Angiogenesis in Chronic Liver Disease. Gastroenterology, 150(4), 982–997.e30 10.1053/j.gastro.2015.11.038 [DOI] [PubMed] [Google Scholar]
- Cao J, He L, Lin G, Hu C, Dong R, Zhang J, … Yang B (2014). Cap-dependent translation initiation factor, e IF4E, is the target for Ouabain-mediated inhibition of HIF-1alpha. Biochem Pharmacol, 89(1), 20–30. 10.1016/j.bcp.2013.12.002 [DOI] [PubMed] [Google Scholar]
- Carvalho B, Postma C, Mongera S, Hopmans E, Diskin S, Van De Wiel MA, … Meijer GA (2009). Multiple putative oncogenes at the chromosome 20q amplicon contribute to colorectal adenoma to carcinoma progression. Gut, 58(1), 79–89. 10.1136/gut.2007.143065 [DOI] [PubMed] [Google Scholar]
- Castano Z, Vergara-Irigaray N, Pajares MJ, Montuenga LM, & Pio R (2008). Expression of alpha CP −4 inhibits cell cycle progression and suppresses tumorigenicity of lung cancer cells. Int J Cancer, 122(7), 1512–1520. 10.1002/ijc.23236 [DOI] [PubMed] [Google Scholar]
- Castello A, Fischer B, Eichelbaum K, Horos R, Beckmann BM, Strein C, … Hentze MW (2012). Insights into RNA biology from an atlas of mammalian mRNA - binding proteins. Cell, 149(6), 1393–1406. 10.1016/j.cell.2012.04.031 [DOI] [PubMed] [Google Scholar]
- Causevic M, Hislop RG, Kernohan NM, Carey FA, Kay RA, Steele RJC, & Fuller-Pace FV (2001). Over expression and poly-ubiquitylation of the DEAD-box RNA helicase p68 in colorectal tumours. Oncogene, 20(53), 7734–7743. 10.1038/sj/onc/1204976 [DOI] [PubMed] [Google Scholar]
- Cencic R, Desforges M, Hall DR, Kozakov D, Du Y, Min J, … Pelletier J (2011). Blocking eIF4E-eIF4G interaction as a strategy to impair coronavirus replication. J Virol, 85(13), 6381–6389. 10.1128/JVI.00078-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cencic R, Hall DR, Robert F, Du Y, Min J, Li L, … Pelletier J (2011). Reversing chemoresistance by small molecule inhibition of the translation init iation complex eIF4F. Proc Natl Acad Sci U S A, 108(3), 1046–1051. 10.1073/pnas.1011477108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cerutti L, Mian N, & Bateman A (2000). Domains in gene silencing and cell differentiation proteins: the novel PAZ domain and redefinition of the Piwi domain. Trends Biochem Sci, 25(10), 481–482. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/11050429 [DOI] [PubMed] [Google Scholar]
- Chae MJ, Sung HY, Kim EH, Lee M, Kwak H, Chae CH, … Park WY (2009). Chemical inhibitors destabilize HuR binding to the AU-rich element of TNF-alpha mRNA. Exp Mol Med, 41(11), 824–831. 10.3858/emm.2009.41.11.088 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan THM, Lin CH, Qi L, Fei J, Li Y, Yong KJ, … Chen L (2014). A disrupted RNA editing balance mediated by ADARs (Adenosine DeAminases that act on RNA) in human hepatocellular carcinoma. Gut, 63(5), 832–843. 10.1136/gutjnl-2012-304037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan THM, Qamra A, Tan KT, Guo J, Yang H, Qi L, … Chen L (2016). ADAR-Mediated RNA Editing Predicts Progression and Prognosis of Gastric Cancer. Gastroenterology, 151(4), 637–650.e10 10.1053/j.gastro.2016.06.043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapman KM, Powell HM, Chaudhary J, Shelton JM, Richardson JA, Richardson TE, & Hamra FK (2013). Linking spermatid ribonucleic acid (RNA) binding protein and retrogene diversity to reproductive success. Mol Cell Proteomics, 12(11), 3221–3236. 10.1074/mcp.M113.030585 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y. Bin, Liao XY, Zhang JB, Wang F, Qin H. De, Zhang L, … Jia WH (2017). ADAR2 functions as a tumor suppressor via editing IGFBP7 in esophageal squamous cell carcinoma. International Journal of Oncology, 50(2), 622–630. 10.3892/ijo.2016.3823 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen CN, Hsieh FJ, Cheng YM, Lee PH, & Chang KJ (2004). Expression of Eukaryotic Initiation Factor 4E in Gastric Adenocarcinoma and Its Association with Clinical Outcome. Journal of Surgical Oncology, 86(1), 22–27. 10.1002/jso.20037 [DOI] [PubMed] [Google Scholar]
- Chen EZ, Jacobson BA, Patel MR, Okon AM, Li S, Xiong K, … Kratzke RA (2014). Small -molecule inhibition of oncogenic eukaryotic protein translation in mesothelioma cells. Invest New Drugs, 32(4), 598–603. 10.1007/s10637-014-0076-7 [DOI] [PubMed] [Google Scholar]
- Chen H, Liu J, Wang H, Cheng Q, Zhou C, Chen X, & Ye F (2019). Inhibition of RNA -Binding Protein Musashi-1 Suppresses Malignant Properties and Reverses Paclitaxel Resistance in Ovarian Carcinoma. Journal of Cancer, 10(6), 1580–1592. 10.7150/jca.27352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen JY, Liu LP, & Xu JF (2017). Decrease of RBM4 indicates poor prognosis in patients with hepatocellular carcin oma after hepatectomy. OncoTargets and Therapy, 10, 339–345. 10.2147/OTT.S125250 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen K, & Rajewsky N (2007). The evolution of gene regulation by transcription factors and microRNAs. Nature Reviews Genetics 10.1038/nrg1990 [DOI] [PubMed] [Google Scholar]
- Chen LC, Hsueh C, Tsang NM, Liang Y, Chang KP, Hao SP, … Chang YS (2008). Heterogeneous ribonucleoprotein k and thymidine phosphorylase are independent prognostic and therapeutic markers for nasopharyngeal carcinoma. Clin Cancer Res, 14(12), 3807–3813. 10.1158/1078-0432.CCR-08-0155 [DOI] [PubMed] [Google Scholar]
- Chen L, Li Y, Lin CH, Chan THM, Chow RKK, Song Y, … Guan XY (2013). Recoding RNA editing of AZIN1 predisposes to hepatocellular carcinoma. Nature Medicine, 19(2), 209–216. 10.1038/nm.3043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen P. xiang, Li Q. yan, & Yang Z (2015). Musashi-1 Expression is a Prognostic Factor in Ovarian Adenocarcinoma and Correlates with ALDH −1 Expression. Pathology and Oncology Research, 21(4), 1133–1140. 10.1007/s12253-015-9943-6 [DOI] [PubMed] [Google Scholar]
- Chen X, Gu P, Xie R, Han J, Liu H, Wang B, … Huang J (2017). Heterogeneous nuclear ribonucleoprotein K is associated with poor prognosis and regulates proliferation and apoptosis in bladder cancer. J Cell Mol Med, 21(7), 1266–1279. 10.1111/jcmm.12999 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y, Xie C, Zheng X, Nie X, Wang Z, Liu H, & Zhao Y (2019). LIN28/ let-7 /PD-L1 Pathway as a Target for Cancer Immunotherapy. Cancer Immunology Research, 7(3), 487–497. 10.1158/2326-6066.cir-18-0331 [DOI] [PubMed] [Google Scholar]
- Cheng Y, Jin Z, Agarwal R, Ma K, Yang J, Ibrahim S, … Mori Y (2012). LARP7 is a potential tumor suppressor gene in gastric cancer. Laboratory Investigation, 92(7), 1013–1019. 10.1038/labinvest.2012.59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung N, Chan LC, Thompson A, Cleary ML, & So CWE ( 2007). Protein arginine-methyltransferase-dependent oncogenesis. Nature Cell Biology, 9(10), 1208–1215. 10.1038/ncb1642 [DOI] [PubMed] [Google Scholar]
- Chi X, Gatti P, & Papoian T (2017). Safety of antisense oligonucleotide and siRNA-based therapeutics. Drug Discovery Today 10.1016/j.drudis.2017.01.013 [DOI] [PubMed] [Google Scholar]
- Chin K, De Vries S, Fridlyand J, Spellman PT, Roydasgupta R, Kuo WL, … Gray JW (2006). Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell, 10(6), 529–541. 10.1016/j.ccr.2006.10.009 [DOI] [PubMed] [Google Scholar]
- Chiriboga CA, Swoboda KJ, Darras BT, Iannaccone ST, Montes J, De Vivo DC, … Bishop KM (2016). Results from a phase 1 study of nusinersen (ISIS-SMN(Rx)) in children with spinal muscular atrophy. Neurology, 86(10), 890–897. 10.1212/WNL.0000000000002445 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho SJ, Teng IF, Zhang M, Yin T, Jung YS, Zhang J, & Chen X (2015). Hypo xia -inducible factor 1 alpha is regulated by RBM38, a RNA-binding protein and a p53 family target, via mRNA translation. Oncotarget, 6(1), 305–316. 10.18632/oncotarget.2786 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho SJ, Zhang J, & Chen X (2010). RNPC1 modulates the RNA-binding activity of, and cooperates with, HuR to regulate p21 mRNA stability. Nucleic Acids Res, 38(7), 2256–2267. 10.1093/nar/gkp1229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi HS, Hwang CK, Song KY, Law PY, Wei LN, & Loh HH (2009). Poly(C)-binding proteins as transcriptional regulators of gene expression. Biochem Biophys Res Commun, 380(3), 431–436. 10.1016/j.bbrc.2009.01.136 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi HS, Kim CS, Hwang CK, Song KY, Law PY, Wei LN, & Loh HH (2007). Novel function of the poly(C)-binding protein alpha CP3 as a transcriptional repressor of the muopioid receptor gene. FASEB J, 21(14), 3963–3973. 10.1096/fj.07-8561com [DOI] [PubMed] [Google Scholar]
- Choi S, Sa M, Cho N, Kim KK, & Park S-H (2019). Rbfox2 dissociation from stress granules suppresses cancer progression. Experimental & Molecular Medicine, 51(4), 49 10.1038/s12276-019-0246-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chresta CM, Davies BR, Hickson I, Harding T, Cosulich S, Critchlow SE, … Pass M (2010). AZD8055 is a potent, selective, and orally bioavailable ATP -competitive mammalian target of rapamycin kinase inhibitor with in vitro and in vivo antitumor activity. Cancer Research 10.1158/0008-5472.CAN-09-1751 [DOI] [PubMed] [Google Scholar]
- Chu J, Galicia-Vázquez G, Cencic R, Mills JR, Katigbak A, Porco JA, & Pelletier J (2016). CRISPR -Mediated Drug-Target Validation Reveals Selective Pharmacological Inhibition of the RNA Helicase, eIF4A. Cell Reports, 15(11), 2340–2347. 10.1016/j.celrep.2016.05.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciais D, Cherradi N, & Feige JJ (2013). Multiple functions of tristetraprolin/TIS11 RNA -binding proteins in the regulation of mRNA biogenesis and degradation. Cell Mol Life Sci, 70(12), 2031–2044. 10.1007/s00018-012-1150-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark EL, Coulson A, Dalgliesh C, Rajan P, Nicol SM, Fleming S, … Robson CN (2008). The RNA helicase p68 is a novel androgen receptor coactivator involved in splicing and is overexpressed in prostate cancer. Cancer Research, 68(19), 7938–7946. 10.1158/0008-5472.CAN-08-0932 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clingman CC, Deveau LM, Hay SA, Genga RM, Shandilya SM, Massi F, & Ryder SP (2014). Allosteric inhibition of a stem cell RNA-binding protein by an intermediary metabolite. Elife, 3 10.7554/eLife.02848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clower CV, Chatterjee D, Wang Z, Cantley LC, Vander Heiden MG, & Krainer AR (2010). The alternative splicing repressors hnRNP A1/A2 and PTB influence pyruvate kinase isoform expression and cell metabolism. Proceedings of the National Academy of Sciences, 107(5), 1894–1899. 10.1073/pnas.0914845107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Comtesse N, Keller A, Diesinger I, Bauer C, Kayser K, Huwer H, … Meese E (2007). Frequent overexpression of the genes FXR1, CLAPM1 and EIF4G located on amplicon 3q26–27 in squamous cell carcinoma of the lung. International Journal of Cancer, 120(12), 2538–2544. 10.1002/ijc.22585 [DOI] [PubMed] [Google Scholar]
- Crew JP, Fuggle S, Bicknell R, Cranston DW, Benedetti A. de, & Harris AL (2002). Eukaryotic initiation factor-4E in superficial and muscle invasive bladder cancer and its correlation with vascular endothelial growth factor expression and tumour progression. British Journal of Cancer 10.1054/bjoc.1999.0894 [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Agostino VG, Adami V, & Provenzani A (2013). A novel high throughput biochemical assay to evaluate the HuR protein -RNA complex formation. PLoS One, 8(8), e72426 10.1371/journal.pone.0072426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’A mbrogio A, Nagaoka K, & Richter JD (2013). Translational control of cell growth and malignancy by the CPEBs. Nature Reviews Cancer 10.1038/nrc3485 [DOI] [PubMed] [Google Scholar]
- Dammann R, Li C, Yoon JH, Chin PL, Bates S, & Pfeifer GP (2000). Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nat Genet, 25(3), 315–319. 10.1038/77083 [DOI] [PubMed] [Google Scholar]
- David CJ, Chen M, Assanah M, Canoll P, & Manley JL (2010). HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature, 463(7279), 364–368. 10.1038/nature08697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis ME, Zuckerman JE, Choi CHJ, Seligson D, Tolcher A, Alabi CA, … Ribas A (2010). Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature, 464(7291), 1067–1070. 10.1038/nature08956 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dayyani F, Wang J, Yeh JR, Ahn EY, Tobey E, Zhang DE, … Sweetser DA (2008). Loss of TLE1 and TLE4 from the del(9q) commonly deleted region in AML cooperates with AML1-ETO to affect myeloid cell proliferation and survival. Blood, 111(8), 4338–4347. 10.1182/blood-2007-07-103291 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Degrauwe N, Schlumpf TB, Janiszewska M, Martin P, Cauderay A, Provero P, … Stamenkovic I (2016). The RNA Binding Protein IMP2 Preserves Glioblastoma Stem Cells by Preventing let-7 Target Gene Silencing. Cell Reports, 15(8), 1634–1647. 10.1016/j.celrep.2016.04.086 [DOI] [PubMed] [Google Scholar]
- Del Campo M, & Lambowitz AM (2009). Structure of the Yeast DEAD box protein Mss116p reveals two wedges that crimp RNA. Mol Cell, 35(5), 598–609. 10.1016/j.molcel.2009.07.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deloria AJ, Höflmayer D, Kienzl P, Łopatecka J, Sampl S, Klimpfinger M, … Holzmann K (2016). Epithelial splicing regulatory protein 1 and 2 paralogues correlate with splice signatures and favorable outcome in human colorectal cancer. Oncotarget, 7(45). 10.18632/oncotarget.12070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding C, Cheng S, Yang Z, Lv Z, Xiao H, Du C, … Zheng S (2014). Long non-coding RNA HOTAIR promotes cell migration and invasion via down-regulation of RNA binding motif protein 38 in hepatocellular carcinoma cells. International Journal of Molecular Sciences, 15(3), 4060–4076. 10.3390/ijms15034060 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong K, Wang R, Wang X, Lin F, Shen JJ, Gao P, & Zhang HZ (2009). Tumor-specific RNAi targeting eIF4E suppresses tumor growth, induces apoptosis and enhances cisplatin cytotoxicity in human breast carcinoma cells. Breast Cancer Research and Treatment, 113(3), 443–456. 10.1007/s10549-008-9956-x [DOI] [PubMed] [Google Scholar]
- Doyle GA, Bourdeau-heller JM, Coulthard S, Meisner LF, & Ross J (2000). Advances in Brief Amplification in Human Breast Cancer of a Gene Encoding a c-myc. Cancer Research, 2756–2759. [PubMed] [Google Scholar]
- Du Z, Lee JK, Fenn S, Tjhen R, Stroud RM, & James TL (2007). X-ray crystallographic and NMR studies of protein-protein and protein-nucleic acid interactions involving the KH domains from human poly(C)-binding protein-2. RNA, 13(7), 1043–1051. 10.1261/rna.410107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duffy AG, Makarova-Rusher OV, Ulahannan SV, Rahma OE, Fioravanti S, Walker M, … Greten TF (2016). Modulation of tumor eIF4E by antisense inhibition: A phase I/II translational clinical trial of ISIS 183750-an antisense oligonucleotide against eIF4E-in combination with irinotecan in solid tumors and irinotecan-refractory colorectal cancer. Int J Cancer, 139(7), 1648–1657. 10.1002/ijc.30199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunford A, Weinstock DM, Savova V, Schumacher SE, Cleary JP, Yoda A, … Lane AA (2017). Tumor-suppressor genes that escape from X-inactivation contribute to cancer sex bias. Nature Genetics, 49(1), 10–16. 10.1038/ng.3726 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Enge M, Bao W, Hedstrom E, Jackson SP, Moumen A, & Selivanova G (2009). MDM 2-dependent downregulation of p21 and hnRNP K provides a switch between apoptosis and growth arrest induced by pharmacologically activated p53. Cancer Cell, 15(3), 171–183. 10.1016/j.ccr.2009.01.019 [DOI] [PubMed] [Google Scholar]
- Evans JR, Mitchell SA, Spriggs KA, Ostrowski J, Bomsztyk K, Ostarek D, & Willis AE (2003). Members of the poly (rC) binding protein family stimulate the activity of the c-myc internal ribosome entry segment in vitro and in vivo. Oncogene, 22(39), 8012–8020. 10.1038/sj.onc.1206645 [DOI] [PubMed] [Google Scholar]
- Fagoonee S, Picco G, Orso F, Arrigoni A, Longo DL, Forni M, … Altruda F (2016). The RNA -binding protein ESRP1 promotes human colorectal cancer progression. Oncotarget, 8(6). 10.18632/oncotarget.14318 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fairman-Williams ME, Guenther UP, & Jankowsky E (2010). SF1 and SF2 helicases: family matters. Curr Opin Struct Biol, 20(3), 313–324. 10.1016/j.sbi.2010.03.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan S, Ramalingam SS, Kauh J, Xu Z, Khuri FR, & Sun SY (2009). Phosphorylated eukaryotic translation initiation factor 4 (eIF4E) is elevated in human cancer tissues. Cancer Biology and Therapy, 8(15). 10.4161/cbt.8.15.8960 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feldman ME, Apsel B, Uotila A, Loewith R, Knight ZA, Ruggero D, & Shokat KM (2009). Active-site inhibitors of mTOR target rapamycin-resistant outputs of mTORC1 and mTORC2. PLoS Biology, 7(2), 0371–0383. 10.1371/journal.pbio.1000038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feldstein O, Ben-Hamo R, Bashari D, Efroni S, & Ginsberg D (2012). RBM 38 is a direct transcriptional target of E2F1 that limits E2F1-induced proliferation. Mol Cancer Res, 10(9), 1169–1177. 10.1158/1541-7786.MCR-12-0331 [DOI] [PubMed] [Google Scholar]
- Filippova N, Yang X, Ananthan S, Sorochinsky A, Hackney JR, Gentry Z, … Nabors LB (2017). Hu antigen R (HuR) multimerization contributes to glioma disease progression. Journal of Biological Chemistry 10.1074/jbc.M117.797878 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flynn RL, & Zou L (2010). Oligonucleotide/oligosaccharide-binding fold proteins: a growing family of genome guardians. Crit Rev Biochem Mol Biol, 45(4), 266–275. 10.3109/10409238.2010.488216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox RG, Lytle NK, Jaquish DV, Park FD, Ito T, Bajaj J, … Reya T (2016). Image-based detection and targeting of therapy resistance in pancreatic adenocarcinoma. Nature, 534(7607), 407–411. 10.1038/nature17988 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox RG, Park FD, Koechlein CS, Kritzik M, & Reya T (2015). Musashi Signaling in Stem Cells and Cancer. Annual Review of Cell and Developmental Biology, 31(1), 249–267. 10.1146/annurev-cellbio-100814-125446 [DOI] [PubMed] [Google Scholar]
- Frederick ED, Ramos SB, & Blackshear PJ (2008). A unique C-terminal repeat domain maintains the cytosolic localization of the placenta-specific tristetraprolin family member ZFP36L3. J Biol Chem, 283(21), 14792–14800. 10.1074/jbc.M801234200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fritzell K, Xu L. Di, Lagergren J, & Öhman M (2018). ADARs and editing: The role of A -to-I RNA modification in cancer progression. Seminars in Cell and Developmental Biology 10.1016/j.semcdb.2017.11.018 [DOI] [PubMed] [Google Scholar]
- Fu K, Sun X, Wier EM, Hodgson A, Liu Y, Sears CL, & Wan F (2016). Sam68/KHDRBS1 is critical for colon tumorigenesis by regulating genotoxic stress-induced NF-κB activation. ELife, 5(2016JULY). 10.7554/eLife.15018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu L, Qin Y-R, Ming X-Y, Zuo X-B, Diao Y-W, Zhang L-Y, … Guan X-Y (2017). RNA editing of SLC22A3 drives early tumor invasion and metastasis in familial esophageal cancer . Proceedings of the National Academy of Sciences, 114(23), E4631–E4640. 10.1073/pnas.1703178114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuller-Pace FV (2013). DEAD box RNA helicase functions in cancer. RNA Biology 10.4161/rna.23312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fumagalli D, Gacquer D, Rothé F, Lefort A, Libert F, Brown D, … Detours V (2015). Principles Governing A -to-I RNA Editing in the Breast Cancer Transcriptome. Cell Reports, 13(2), 277–289. 10.1016/j.celrep.2015.09.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furic L, Rong L, Larsson O, Koumakpayi IH, Yoshida K, Brueschke A, … Sonenberg N (2010). eIF4E phosphorylation promotes tumorigenesis and is associated with prostate cancer progression. Proceedings of the National Academy of Sciences, 107(32), 14134–14139. 10.1073/pnas.1005320107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galeano F, Rossetti C, Tomaselli S, Cifaldi L, Lezzerini M, Pezzullo M, … Gallo A (2013). ADAR2 -editing activity inhibits glioblastoma growth through the modulation of the CDC14B/Skp2/p21/p27 axis. Oncogene, 32(8), 998–1009. 10.1038/onc.2012.125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallardo M, Lee HJ, Zhang X, Bueso-Ramos C, Pageon LR, McArthur M, … Post SM (2015). hn RNP K Is a Haploinsufficient Tumor Suppressor that Regulates Proliferation and Differentiation Programs in Hematologic Malignancies. Cancer Cell, 28(4), 486–499. 10.1016/j.ccell.2015.09.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galore-Haskel G, Nemlich Y, Greenberg E, Ashkenazi S, Hakim M, Itzhaki O, … Markel G (2015). A novel immune resistance mechanism of melanoma cells controlled by the ADAR1 enzyme. Oncotarget, 6(30). 10.18632/oncotarget.4905 [DOI] [PMC free article] [PubMed] [Google Scholar]
- García-Martínez JM, Moran J, Clarke RG, Gray A, Cosulich SC, Chresta CM, & Alessi DR (2009). Ku-0063794 is a specific inhibitor of the mammalian target of rapamycin (mTOR). The Biochemical Journal, 421(1), 29–42. 10.1042/BJ20090489 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ger M, Kaupinis A, Petrulionis M, Kurlinkus B, Cicenas J, Sileikis A, … Strupas K (2018). Proteomic Identificat ion of FLT3 and PCBP3 as Potential Prognostic Biomarkers for Pancreatic Cancer. Anticancer Res, 38(10), 5759–5765. 10.21873/anticanres.12914 [DOI] [PubMed] [Google Scholar]
- Gerhauser C, Favero F, Risch T, Simon R, Feuerbach L, Assenov Y, … Weischenfeldt J (2018). Molecular Evolution of Early-Onset Prostate Cancer Identifies Molecular Risk Markers and Clinical Trajectories. Cancer Cell, 34(6), 996–1011.e8 10.1016/j.ccell.2018.10.016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerstberger S, Hafner M, & Tuschl T (2014). A census of human RNA-binding proteins. Nat Rev Genet, 15(12), 829–845. 10.1038/nrg3813 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghanem LR, Kromer A, Silverman IM, Chatterji P, Traxler E, Penzo-Mendez A, … Liebhaber SA (2016). The Poly(C) Binding Protein Pcbp2 and Its Retrotransposed Derivative Pcbp1 Are Independently Essential to Mouse Development. Mol Cell Biol, 36(2), 304–319. 10.1128/MCB.00936-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghosh B, Benyumov AO, Ghosh P, Jia Y, Avdulov S, Dahlberg PS, … Wagner CR (2009). Nontoxic chemical interdiction of the epithelial-to-mesenchymal transition by targeting cap-dependent translation. ACS Chem Biol, 4(5), 367–377. 10.1021/cb9000475 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghoshal A, Rodrigues LC, Gowda CP, Elcheva IA, Liu Z, Abraham T, & Spiegelman VS (2019). Extracellular vesicle-dependent effect of RNA-binding protein IGF2BP1 on melanoma metastasis. Oncogene 10.1038/s41388-019-0797-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Golan-Gerstl R, Cohen M, Shilo A, Suh SS, Bakàcs A, Coppola L, & Karni R (2011). Splicing factor hn RNP A2/ B1 regulates tumor suppressor gene splicing and is an oncogenic driver in glioblastoma. Cancer Research, 71(13), 4464–4472. 10.1158/0008-5472.CAN-10-4410 [DOI] [PubMed] [Google Scholar]
- Gou Q, Gao L, Nie X, Pu W, Zhu J, Wang Y, … Peng Y (2018). Long noncoding RNA AB074169 inhibits cell proliferation via modulation of KHSRP-mediated CDKN1a expressionin papillary thyroid carcinoma. Cancer Research, 78(15), 4163–4174. 10.1158/0008-5472.CAN-17-3766 [DOI] [PubMed] [Google Scholar]
- Gouble A, Grazide S, Meggetto F, Mercier P, Delsol G, & Morello D (2002). A new player in oncogenesis: AUF1/hnRNPD overe xpression leads to tumorigenesis in transgenic mice. Cancer Research, 62(5), 1489–1495. [PubMed] [Google Scholar]
- Graff JR, Konicek BW, Lynch RL, Dumstorf CA, Dowless MS, Mcnulty AM, … Carter JH (2009). eIF4E activation is commonly Elevated in advanced human prostate cancers and significantly related to reduced patient survival. Cancer Research, 69(9), 3866–3873. 10.1158/0008-5472.CAN-08-3472 [DOI] [PubMed] [Google Scholar]
- Grishin NV (2001). KH domain: one motif, two folds. Nucleic Acids Res, 29(3), 638–643. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/11160884 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu W, Pan F, & Singer RH (2009). Blocking -catenin binding to the ZBP1 promoter represses ZBP1 e xp ression, leading to increased proliferation and migration of metastatic breast-cancer cells. Journal of Cell Science, 122(11), 1895–1905. 10.1242/jcs.045278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guan X, Chen S, Liu Y, Wang L. li, Zhao Y, & Zong ZH (2018). PUM1 promotes ovarian cancer proliferation, migration and invasion. Biochemical and Biophysical Research Communications, 497(1), 313–318. 10.1016/j.bbrc.2018.02.078 [DOI] [PubMed] [Google Scholar]
- Guha M, Srinivasan S, Raman P, Jiang Y, Kaufman BA, Taylor D, … Avadhani NG (2018). Aggressive triple negat ive breast cancers have unique molecular signature on the basis of mitochondrial genetic and functional defects. Biochimica et Biophysica Acta - Molecular Basis of Disease, 1864(4), 1060–1071. 10.1016/j.bbadis.2018.01.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gumireddy K, Li A, Kossenkov AV, Sakurai M, Yan J, Li Y, … Huang Q (2016). The mRNA -edited form of GA BRA 3 suppresses GA BRA 3-mediated Akt activation and breast cancer metastasis. Nature Communications, 7 10.1038/ncomms10715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo C, Yang RJ, Jang K, Zhou XL, & Liu YZ (2017). Protective Effects of Pretreatment with Quercetin Against Lipopolysaccharide-Induced Apoptosis and the Inhibition of Osteoblast Differentiation via the MAPK and Wnt/beta-Catenin Pathways in MC3T3-E1 Cells. Cell Physiol Biochem, 43(4), 1547–1561. 10.1159/000481978 [DOI] [PubMed] [Google Scholar]
- Guo J, & Jia R (2018). Splicing factor poly(rC)-binding protein 1 is a novel and distinctive tumor suppressor. J Cell Physiol, 234(1), 33–41. 10.1002/jcp.26873 [DOI] [PubMed] [Google Scholar]
- Guo K, Cui J, Quan M, Xie D, Jia Z, Wei D, … Xie K (2017). The novel KLF4/MSI2 signaling pathway regulates growth and metastasis of pancreatic cancer. Clinical Cancer Research, 23(3), 687–696. 10.1158/1078-0432.CCR-16-1064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo X, Wu Y, & Hartley RS (2009). MicroRNA-125a represses cell growth by targeting HuR in breast cancer. RNA Biology, 6(5), 575–583. 10.4161/rna.6.5.10079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hallegger M, Taschner A, & Jantsch MF (2006). RNA aptamers binding the double-stranded RNA-binding domain. Rna, 12(11), 1993–2004. 10.1261/rna.125506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamano R, Miyata H, Yamasaki M, Sugimura K, Tanaka K, Kurokawa Y, … Doki Y (2012). High expression of Lin 28 is associated with tumour aggressiveness and poor prognosis of patients in oesophagus cancer. British Journal of Cancer, 106(8), 1415–1423. 10.1038/bjc.2012.90 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han L, Diao L, Yu S, Xu X, Li J, Zhang R, … Liang H (2015). The Genomic Landscape and Clinical Relevance of A -to-I RNA Editing in Human Cancers. Cancer Cell 10.1016/j.ccell.2015.08.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han S-W, Kim H-P, Shin J-Y, Jeong E-G, Lee W-C, Kim KY, … Kim T-Y (2014). RNA editing in RHOQ promotes invasion potential in colorectal cancer. The Journal of Experimental Medicine, 211(4), 613–621. 10.1084/jem.20132209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han T, Gora lski M, Gaskill N, Capota E, Kim J, Ting TC, … Nijhawan D (2017). Anticancer sulfonamides target splicing by inducing RBM 39 degradation via recruitment to DCAF15. Science, 356(6336). 10.1126/science.aal3755 [DOI] [PubMed] [Google Scholar]
- Han W, Xin Z, Zhao Z, Bao W, Lin X, Yin B, … Peng X (2013). RNA-binding protein PCBP2 modulates glioma growth by regulating FHL3. J Clin Invest, 123(5), 2103–2118. 10.1172/JCI61820 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen TB, Jensen TI, Clausen BH, Bra msen JB, Finsen B, Damgaard CK, & Kjems J (2013). Natural RNA circles function as efficient micro RNA sponges. Nature, 495(7441), 384–388. 10.1038/nature11993 [DOI] [PubMed] [Google Scholar]
- Harvey SE, Xu Y, Lin X, Gao XD, Qiu Y, Ahn J, … Cheng C (2018). Coregulation of alternative splicing by hnR NPM and ESRP1 during EMT. Rna, 24(10), 1326–1338. 10.1261/rna.066712.118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayakawa A, Saitoh M, & Miyazawa K (2017). Dual roles for epithelial splicing regulatory proteins 1 (ESRP1) and 2 (ESRP2) in cancer progression. In Advances in Experimental Medicine and Biology (Vol. 925, pp. 33–40). 10.1007/5584_2016_50 [DOI] [PubMed] [Google Scholar]
- He L, Zhou X, Qu C, Hu L, Tang Y, Zhang Q, … Hong J (2014). Musashi2 predicts poor prognosis and invasion in hepatocellular carcinoma by driving epit helial-mesenchymal transition. Journal of Cellular and Molecular Medicine, 18(1), 49–58. 10.1111/jcmm.12158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He N, Jahchan NS, Hong E, Li Q, Bayfield MA, Maraia RJ, … Zhou Q (2008). A La-Related Protein Modulates 7SK snRNP Integrity to Suppress P-TEFb-Dependent Transcriptional Elongation and Tumorigenesis. Molecular Cell, 29(5), 588–599. 10.1016/j.molcel.2008.01.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He X, Li W, Liang X, Zhu X, Zhang L, Huang Y, … Chen Z (2018). IGF2BP2 overe xpression indicates poor survival in patients with acute myelocytic leukemia. Cellular Physiology and Biochemistry, 51(4), 1945–1956. 10.1159/000495719 [DOI] [PubMed] [Google Scholar]
- Heerma van Voss MR, van Diest PJ, & Raman V (2017). Targeting RNA helicases in cancer: The translation trap. Biochimica et Biophysica Acta - Reviews on Cancer 10.1016/j.bbcan.2017.09.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heerma Van Voss MR, Vesuna F, Bol GM, Afzal J, Tantravedi S, Bergman Y, … Raman V (2018). Targeting mitochondrial translation by inhibiting DDX3: A novel radiosensitization strategy for cancer treatment. Oncogene, 37(1), 63–74. 10.1038/onc.2017.308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinicke LA, Nabet B, Shen S, Jiang P, van Zalen S, Cieply B, … Carstens RP (2013). The RNA Binding Protein RBM38 (RNPC1) Regulates Splicing during Late Erythroid Differentiation. PLoS ONE, 8(10), e78031 10.1371/journal.pone.0078031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinonen M, Bono P, Narko K, Chang SH, Lundin J, Joensuu H, … Ristimäki A (2005). Cytoplasmic HuR expression is a prognostic factor in invasive ductal breast carcinoma. Cancer Research, 65(6), 2157–2161. 10.1158/0008-5472.CAN-04-3765 [DOI] [PubMed] [Google Scholar]
- Heise T, Kota V, Brock A, Morris AB, Rodriguez RM, Zierk AW, … Sommer G (2016). The La protein counteract scisplatin-induced cell death by stimulating protein synthesis of anti-apoptotic factor Bcl2. Oncotarget, 7(20). 10.18632/oncotarget.8819 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentze MW, Castello A, Schwarzl T, & Preiss T (2018). A brave new world of RNA -binding proteins. Nat Rev Mol Cell Biol, 19(5), 327–341. 10.1038/nrm.2017.130 [DOI] [PubMed] [Google Scholar]
- Hermsen M, Postma C, Baak J, Weiss M, Rapallo A, Sciutto A, … Meijer G (2002). Colorectal adenoma to carcinoma progres sion follows multiple pathways of chromosomal instability. Gastroenterology, 123(4), 1109–1119. 10.1053/gast.2002.36051 [DOI] [PubMed] [Google Scholar]
- Hernández J, Bechara E, Schlesinger D, Delgado J, Serrano L, & Va lcárcel J (2016). Tumor suppressor properties of the splicing regulatory factor RBM 10. RNA Biology, 13(4), 466–472. 10.1080/15476286.2016.1144004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodson DJ, Janas ML, Galloway A, Bell SE, Andrews S, Li CM, … Turner M (2010). Deletion of the RNA -binding proteins ZFP36L1 and ZFP36L2 leads to perturbed thymic development and Tlymphoblastic leukemia. Nat Immunol, 11(8), 717–724. 10.1038/ni.1901 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holcik M, & Korneluk RG (2002). Functional Characterization of the X-Linked Inhibitor of Apoptosis (XIAP) Internal Ribosome Entry Site Element: Role of La Autoantigen in XIAP Translation. Molecular and Cellular Biology, 20(13), 4648–4657. 10.1128/mcb.20.13.4648-4657.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong DS, Kurzrock R, Oh Y, Wheler J, Naing A, Brail L, … Simon G (2011). A phase 1 dose escalation, pharmacokinetic, and pharmacodynamic evaluation of eIF-4E antisense oligonucleotide LY2275796 in patients with advanced cancer. Clin Cancer Res, 17(20), 6582–6591. 10.1158/1078-0432.CCR-11-0430 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hopkins TG, Mura M, Al-Ashtal HA, Lahr RM, Abd-Latip N, Sweeney K, … Blagden SP (2016). The RNA-binding protein LARP1 is a post-transcriptional regulator of survival and tumorigenesis in ovarian cancer. Nucleic Acids Research, 44(3), 1227–1246. 10.1093/nar/gkv1515 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsieh AC, Costa M, Zollo O, Davis C, Feldman ME, Testa JR, … Ruggero D (2010). Genetic Dissection of the Oncogenic mTOR Pathway Reveals Druggable Addiction to Translational Control via 4EBP-eIF4E. Cancer Cell, 17(3), 249–261. 10.1016/j.ccr.2010.01.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsieh AC, & Ruggero D (2010). Targeting eukaryotic translation init iation factor 4E (eIF4E) in cancer. Clinical Cancer Research 10.1158/1078-0432.CCR-10-0433 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu KF, Shen MR, Huang YF, Cheng YM, Lin SH, Chow NH, … Ho CL (2015). Overe xpression of the RNA-binding proteins Lin 28B and IGF2BP3 (IMP3) is associated with chemoresistance and poor disease outcome in ovarian cancer. British Journal of Cancer, 113(3), 414–424. 10.1038/bjc.2015.254 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu MT, & Coca-Prados M (1979). Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells [24]. Nature 10.1038/280339a0 [DOI] [PubMed] [Google Scholar]
- Hu CE, Liu YC, Zhang HD, & Huang GJ (2014). The RNA -binding protein PCBP2 facilitates gastric carcinoma growth by targeting miR-34a. Biochem Biophys Res Commun, 448(4), 437–442. 10.1016/j.bbrc.2014.04.124 [DOI] [PubMed] [Google Scholar]
- Hua WF, Zhong Q, Xia TL, Chen Q, Zhang MY, Zhou AJ, … Zeng MS (2016). RBM24 suppresses cancer progression by upregulating miR-25 to target MALAT1 in nasopharyngeal carcinoma. Cell Death Dis, 7(9), e2352 10.1038/cddis.2016.252 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hua Y, Sahashi K, Hung G, Rigo F, Passini MA, Bennett CF, & Krainer AR (2010). Antisense correction of SMN2 splicing in the CNS rescues necrosis in a type III SMA mouse model. Genes Dev, 24(15), 1634–1644. 10.1101/gad.1941310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hua Y, Sahashi K, Rigo F, Hung G, Horev G, Bennett CF, & Krainer AR (2011). Peripheral SMN restoration is essential for long-term rescue of a severe spinal muscular atrophy mouse model. Nature, 478(7367), 123–126. 10.1038/nature10485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang JS, Chao CC, Su TL, Yeh SH, Chen DS, Chen CT, … Jou YS (2004). Diverse cellular transformation capability of overe xpressed genes in human hepatocellular carcinoma. Biochemical and Biophysical Research Communications, 315(4), 950–958. 10.1016/j.bbrc.2004.01.151 [DOI] [PubMed] [Google Scholar]
- Huang L, Malu S, McKenzie JA, Andrews MC, Talukder AH, Tieu T, … Hwu P (2018). The RNA -binding Protein MEX3B mediates resistance to cancer immunotherapy by downregulating HLA-A expression. Clinical Cancer Research, 24(14), 3366–3376. 10.1158/1078-0432.CCR-17-2483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang R. sheng, Zheng Y. liang, Li C, Ding C, Xu C, & Zhao J (2018). Micro RNA −485–5p suppresses growth and metastasis in non-small cell lung cancer cells by targeting IGF2BP2. Life Sciences, 199, 104–111. 10.1016/j.lfs.2018.03.005 [DOI] [PubMed] [Google Scholar]
- Huang W, Wei XL, Ni W, Cao M, Meng L, & Yang H (2017). The e xp ression of RNA -binding protein RBM 38 decreased in renal cell carcinoma and represses renal cancer cell proliferation, migration, and invasion. Tumour Biol, 39(5), 1010428317701635 10.1177/1010428317701635 [DOI] [PubMed] [Google Scholar]
- Huang YH, Peng W, Furuuchi N, Ge rhart J, Rhodes K, Mukherjee N, … Sawicki JA (2016). Delivery of Therapeuti cs Targeting the mRNA-Binding Protein HuR Using 3DNA Nanocarriers Suppresses Ovarian Tumor Growth. Cancer Res, 76(6), 1549–1559. 10.1158/0008-5472.CAN-15-2073 [DOI] [PubMed] [Google Scholar]
- Hudson BP, Martinez-Yamout MA, Dyson HJ, & Wright PE (2004). Recognition of the mRNA AU-rich element by the zinc finger domain of TIS11d. Nature Structural and Molecular Biology, 11(3), 257–264. 10.1038/nsmb738 [DOI] [PubMed] [Google Scholar]
- Hüttelmaier S, Zenklusen D, Lederer M, Dictenberg J, Lorenz M, Meng XH, … Singer RH (2005). Spatial regulation of β-actin translation by Src-dependent phosphorylation of ZBP1. Nature, 438(7067), 512–515. 10.1038/nature04115 [DOI] [PubMed] [Google Scholar]
- Ibrahimpasic T, Xu B, Landa I, Dogan S, Middha S, Seshan V, … Ganly I (2017). Geno mic alterations in fatal forms of non-anaplastic thyroid cancer: Identification of MED12 and RBM 10 as novel thyroid cancer genes associated with tumor virulence. Clinical Cancer Research, 23(19), 5970–5980. 10.1158/1078-0432.CCR-17-1183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iida T, Yi H, Liu S, Huang W, Kanda H, Lubarsky DA, & Hao S (2016). Spinal CPEB-mt ROS-CBP signaling pathway contributes to perineural HIV gp120 with ddC-related neuropathic pain in rats. Exp Neurol, 281, 17–27. 10.1016/j.expneurol.2016.04.012 [DOI] [PubMed] [Google Scholar]
- Ioannidis P, Mahaira L, Papadopoulou A, Teixeira MR, Heim S, Andersen JA, … Trangas T (2003). 8q 24 copy number gains and expression of the c-myc mRNA stabilizing protein CRD-BP in primary breast carcinomas. International Journal of Cancer, 104(1), 54–59. 10.1002/ijc.10794 [DOI] [PubMed] [Google Scholar]
- Ioannidis P, Trangas T, Dimitriadis E, Samiotaki M, Kyriazoglou I, Tsiapalis CM, … Pandis N (2001). C-MYC and IGF-II mRNA-binding protein (CRD-BP/IMP-1) in benign and malignant mesenchymal tumors. International Journal of Cancer, 94(4), 480–484. 10.1002/ijc.1512 [DOI] [PubMed] [Google Scholar]
- Ireson CR, & Kelland LR (2006). Discovery and development of anticancer aptamers. Molecular Cancer Therapeutics, 5(12), 2957–2962. 10.1158/1535-7163.MCT-06-0172 [DOI] [PubMed] [Google Scholar]
- Ishii H, Saitoh M, Sakamoto K, Kondo T, Katoh R, Tanaka S, … Miyazawa K (2014). Epithelial splicing regulatory proteins 1 (ESRP1) and 2 (ESRP2) suppress cancer cell motility via different mechanisms. Journal of Biological Chemistry, 289(40), 27386–27399. 10.1074/jbc.M114.589432 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishimaru D, Ramalingam S, Sengupta TK, Bandyopadhyay S, Dellis S, Tholanikunnel BG, … Spicer EK (2009). Regulation of Bcl-2 Exp ression by HuR in HL60 Leukemia Cells and A431 Carcinoma Cells. Molecular Cancer Research, 7(8), 1354–1366. 10.1158/1541-7786.mcr-08-0476 [DOI] [PubMed] [Google Scholar]
- Ito M, Iwatani M, Ya mamoto T, Tanaka T, Kawamoto T, Morishita D, … Maezaki H ( 2017). Discovery of spiro[indole-3,2′-pyrrolidin]-2(1H)-one based inhibitors targeting Brr2, a core component of the U5 snRNP. Bioorganic and Medicinal Chemistry, 25(17), 4753–4767. 10.1016/j.bmc.2017.07.017 [DOI] [PubMed] [Google Scholar]
- Ito T, Kwon HY, Zimdahl B, Congdon KL, Blum J, Lento WE, … Reya T (2010). Regulation of myeloid leukaemia by the cell -fate determinant Musashi. Nature, 466(7307), 765–768. 10.1038/nature09171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito Y, Narita N, Nomi N, Sugimoto C, Takabayashi T, Yamada T, … Fujieda S (2015). Suppression of Poly(rC)-Binding Protein 4 (PCBP4) reduced cisplatin resistance in human maxillary cancer cells. Sci Rep, 5, 12360 10.1038/srep12360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki S, Floor SN, & Ingolia NT (2016). Rocaglates convert DEAD-box protein eIF4A into a sequence-selective translational repressor. Nature, 534(7608), 558–561. 10.1038/nature17978 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki S, Iwasaki W, Takahashi M, Sakamoto A, Watanabe C, Shichino Y, … Ingolia NT (2019). The Translation Inhibitor Rocaglamide Targets a Bimolecular Cavity between eIF4A and Polypurine RNA. Molecular Cell, 73(4), 738–748.e9 10.1016/j.molcel.2018.11.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwatani-Yoshihara M, Ito M, Klein MG, Yamamoto T, Yonemori K, Tanaka T, … Kawamoto T (2017). Discovery of Allosteric Inhibitors Targeting the Spliceosomal RNA Helicase Brr2. Journal of Medicinal Chemistry 10.1021/acs.jmedchem.7b00461 [DOI] [PubMed] [Google Scholar]
- Janiszewska M, Suvà ML, Riggi N, Houtkooper RH, Auwerx J, Clément-Schatlo V, … Stamenkovic I (2012). Imp2 controls oxidative phosphorylation and is crucial for preservin glioblastoma cancer stem cells. Genes and Development, 26(17), 1926–1944. 10.1101/gad.188292.112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, … Sharpless NE (2013). Circular RNAs are abundant, conserved, and associated with ALU repeats. Rna, 19(2), 141–157. 10.1261/rna.035667.112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenkins RW, Barbie DA, & Flaherty KT (2018). Mechanisms of resistance to immune checkpoint inhibitors. British Journal of Cancer, 118(1), 9–16. 10.1038/bjc.2017.434 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenssen TK, Kuo WP, Stokke T, & Hovig E (2002). Associations between gene expressions in breast cancer and patient survival. Human Genetics, 111(4–5), 411–420. 10.1007/s00439-002-0804-5 [DOI] [PubMed] [Google Scholar]
- Jeong HM, Han J, Lee SH, Park H-J, Lee HJ, Choi J-S, … Kwon MJ (2017). ESRP1 is overe xp ressed in ovarian cancer and promotes switching from mesenchymal to epithelial phenotype in ovarian cancer cells. Oncogenesis, 6(10), e389 10.1038/oncsis.2017.87 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, … Müller-Tidow C (2003). MALAT-1, a novel noncoding RNA, and thymosin β4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene, 22(39), 8031–8041. 10.1038/sj.onc.1206928 [DOI] [PubMed] [Google Scholar]
- Ji X, Lu H, Zhou Q, & Luo K (2014). LARP7 suppresses P-TEFb activity to inhibit breast cancer progression and metastasis. ELife, 3 10.7554/elife.02907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia Y, Chiu TL, Amin EA, Polunovsky V, Bitterman PB, & Wagner CR (2010). Design, synthesis and evaluation of analogs of init iation factor 4E (eIF4E) cap -binding antagonist Bn7-GMP. Eur J Med Chem, 45(4), 1304–1313. 10.1016/j.ejmech.2009.11.054 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang L, Gu ZH, Yan ZX, Zhao X, Xie YY, Zhang ZG, … Chen SJ (2015). Exome sequencing identifies somatic mutations of DDX3X in natural killer/T-cell lymphoma. Nature Genetics, 47(9), 1061–1066. 10.1038/ng.3358 [DOI] [PubMed] [Google Scholar]
- Jiang Y, Zhang M, Qian Y, Xu E, Zhang J, & Chen X (2014). Rbm24, an RNA-binding protein and a target of p53, regulates p21 expression via mRNA stability. J Biol Chem, 289(6), 3164–3175. 10.1074/jbc.M113.524413 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jimbo M, Blanco FF, Huang Y-H, Telonis AG, Screnci BA, Cosma GL, … Brody JR (2015). Targeting the mRNA-binding protein Hu R impairs malignant characteristics of pancreatic ductal adenocarcinoma cells. Oncotarget, 6(29). 10.18632/oncotarget.4743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin D, Hidaka K, Shirai M, & Morisaki T (2010). RNA-binding motif protein 24 regulates myogenin e xpression and promotes myogenic differentiation. Genes Cells, 15(11), 1158–1167. 10.1111/j.1365-2443.2010.01446.x [DOI] [PubMed] [Google Scholar]
- Jønson L, Christiansen J, Hansen TVO, Vikeså J, Ya mamoto Y, & Nielsen FC (2014). IMP3 RNP Safe Houses Prevent miRNA -Directed HM GA2 mRNA Decay in Cancer and Development. Cell Reports, 7(2), 539–551. 10.1016/j.celrep.2014.03.015 [DOI] [PubMed] [Google Scholar]
- Kanemura Y, Yamasaki M, Kanemura Y, Mori K, Fujikawa H, Hayashi H, … Okano H (2001). Musashi1, an evolutionarily conserved neural RNA -binding protein, is a versatile marker of human glioma cells in determining their cellular origin, malignancy, and proliferative activity. Differentiation, 68(2–3), 141–152. 10.1046/j.1432-0436.2001.680208.x [DOI] [PubMed] [Google Scholar]
- Kang MJ, Ryu BK, Lee MG, Han J, Lee JH, Ha TK, … Chi SG (2008). NF-κB Activates Transcription of the RNA-Binding Factor Hu R, via PI3K-AKT Signaling, to Promote Gastric Tumorigenesis. Gastroenterology, 135(6). 10.1053/j.gastro.2008.08.009 [DOI] [PubMed] [Google Scholar]
- Kato M, Goto Y, Matsushita R, Kurozumi A, Fukumoto I, Nishikawa R, … Seki N (2015). MicroRNA −26a/b directly regulate La-related protein 1 and inhibit cancer cell invasion in prostate cancer. International Journal of Oncology, 47(2), 710–718. 10.3892/ijo.2015.3043 [DOI] [PubMed] [Google Scholar]
- Kato T, Hayama S, Ya mabuki T, Ishikawa N, Miyamoto M, Ito T, … Daigo Y (2007). Increased expression of insulin -like growth factor-II messenger RNA-binding protein 1 is associated with tumor progression in patients with lung cancer. Clinical Cancer Research, 13(2 I), 434–442. 10.1158/1078-0432.CCR-06-1297 [DOI] [PubMed] [Google Scholar]
- Kechavarzi B, & Janga SC (2014). Dissecting the expression landscape of RNA-binding proteins in human cancers. Genome Biol, 15(1), R14 10.1186/gb-2014-15-1-r14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keene JD (2007). RNA regulons: Coordination of post-transcriptional events. Nature Reviews Genetics 10.1038/nrg2111 [DOI] [PubMed] [Google Scholar]
- Kentsis A, Topisirovic I, Culjkovic B, Shao L, & Borden KL (2004). Ribavirin suppresses eIF4E-mediated oncogenic transformation by physical mimicry of the 7-methyl guanosine mRNA cap. Proc Natl Acad Sci U S A, 101(52), 18105–18110. 10.1073/pnas.0406927102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kharas MG, Lengner CJ, Al-Shahrour F, Bullinger L, Ball B, Zaidi S, … Daley GQ (2010). Musashi-2 regulates normal hematopoiesis and promotes aggressive myeloid leukemia. Nature Medicine, 16(8), 903–908. 10.1038/nm.2187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim S, Hwang BY, Su BN, Chai H, Mi Q, Kinghorn AD, … Swanson SM (2007). Silvestrol, a potential antican cer rocaglate derivative from Aglaia foveolata, induces apoptosis in LNCaP cells through the mitochondrial/apoptosome pathway without activation of executioner caspase-3 or −7. Anticancer Res, 27(4B), 2175–2183. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/17695501 [PMC free article] [PubMed] [Google Scholar]
- Kim SS, Pandey KK, Choi HS, Kim SY, La w PY, Wei LN, & Loh HH (2005). Poly(C) binding protein family is a transcription factor in mu -opioid receptor gene expression. Mol Pharmacol, 68(3), 729–736. 10.1124/mol.105.012245 [DOI] [PubMed] [Google Scholar]
- King CE, Cuatrecasas M, Castells A, Sepulveda AR, Lee JS, & Rustgi AK (2011). LIN28B promotes colon cancer progression and metastasis. Cancer Research, 71(12), 4260–4268. 10.1158/0008-5472.CAN-10-4637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kondaskar A, Kondaskar S, Kumar R, Fishbein JC, Muvarak N, Lapidus RG, … Ho smane RS (2010). Novel, Broad Spectrum Anti-Cancer Agents Containing the Tricyclic 5:7:5-Fused Diimidazodiazepine Ring System. ACS Med Chem Lett, 2(3), 252–256. 10.1021/ml100281b [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konicek BW, Stephens JR, McNulty AM, Robichaud N, Peery RB, Dumstorf CA, … Graff JR (2011). Therapeutic inhibition of MAP kinase interacting kinase blocks eukaryotic in itiat ion factor 4E phosphorylation and suppresses outgrowth of experimental lung metastases. Cancer Research, 71(5), 1849–1857. 10.1158/0008-5472.CAN-10-3298 [DOI] [PubMed] [Google Scholar]
- Koso H, Yi H, Sheridan P, Miyano S, Ino Y, Todo T, & Watanabe S (2016). Identification of RNA -binding protein LARP4B as a tumor suppressor in glioma. Cancer Research, 76(8), 2254–2264. 10.1158/0008-5472.CAN-15-2308 [DOI] [PubMed] [Google Scholar]
- Kristensen LS, Hansen TB, Venø MT, & Kjems J (2018). Circular RNAs in cancer: Opportunities and challenges in t he field. Oncogene 10.1038/onc.2017.361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kudinov AE, Deneka A, Nikonova AS, Beck TN, Ahn Y-H, Liu X, … Boumber Y (2016). Musashi-2 (MSI2) supports TGF-β signaling and inhibits claudins to promote non-small cell lung cancer (NSCLC) metastasis. Proceedings of the National Academy of Sciences, 113(25), 6955–6960. 10.1073/pnas.1513616113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kudinov AE, Karanicolas J, Golemis EA, & Boumber Y (2017). Musashi RNA -binding proteins as cancer drivers and novel therapeutic targets. Clinical Cancer Research 10.1158/1078-0432.CCR-16-2728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuwano Y, Kim HH, Abdelmohsen K, Pullmann R, Martindale JL, Yang X, & Gorospe M (2008). MKP −1 mRNA Stabilization and Translational Control by RNA-Binding Proteins HuR and NF90. Molecular and Cellular Biology, 28(14), 4562–4575. 10.1128/mcb.00165-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuznetsov G, Xu Q, Rudolph-Owen L, Tendyke K, Liu J, Towle M, … Littlefield BA (2009). Potent in vitro and in vivo anticancer activities of des-methyl, des-amino pateamine A, a synthetic analogue of marine natural product pateamine A. Mol Cancer Ther, 8(5), 1250–1260. 10.1158/1535-7163.MCT-08-1026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lafarga V, Cuadrado A, Lopez de Silanes I, Bengoechea R, Fernandez-Capetillo O, & Nebreda AR (2009). p 38 Mitogen-Activated Protein Kinase- and HuR-Dependent Stabilization of p21Cip1 mRNA Mediates the G1/S Checkpoint. Molecular and Cellular Biology, 29(16), 4341–4351. 10.1128/MCB.00210-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai WS, Carballo E, Thorn JM, Kennington EA, & Blackshear PJ (2000). Interactions of CCCH zinc finger proteins with mRNA. Binding of tristetraprolin -related zinc finger proteins to Au-rich elements and destabilization of mRNA. J Biol Chem, 275(23), 17827–17837. 10.1074/jbc.M001696200 [DOI] [PubMed] [Google Scholar]
- Lai WS, Kennington EA, & Blackshear PJ (2003). Tristetraprolin and its family members can promote the cell-free deadenylation of AU-rich element-containing mRNAs by poly(A) ribonuclease. Mol Cell Biol, 23(11), 3798–3812. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/12748283 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lan L, Appelman C, Smith AR, Yu J, Larsen S, Marquez RT, … Xu L (2015). Natural product (−)-gossypol inhibits colon cancer cell growth by targeting RNA-binding protein Musashi-1. Mol Oncol, 9(7), 1406–1420. 10.1016/j.molonc.2015.03.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lang M, Berry D, Passecker K, Mesteri I, Bhuju S, Ebner F, … Gasche C (2017). Hu R small-molecule inhibitor elicits differential effects in adenomatosis polyposis and colorectal carcinogenesis. Cancer Research, 77(9), 2424–2438. 10.1158/0008-5472.CAN-15-1726 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen JE, Nathan V, Osborne JK, Farro w RK, Deb D, Sullivan JP, … Minna JD (2016). ZEB1 drives epithe lial-to-mesenchymal transition in lung cancer. Journal of Clinical Investigation, 126(9), 3219–3235. 10.1172/JCI76725 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee S, Kopp F, Chang TC, Sataluri A, Chen B, Sivakumar S, … Mendell JT (2016). Noncoding RNA NORAD Regulate s Geno mic Stability by Sequestering PUMILIO Proteins. Cell, 164(1–2), 69–80. 10.1016/j.cell.2015.12.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leidgens S, Bullough KZ, Shi H, Li F, Shakoury-Elizeh M, Yabe T, … Philpott CC (2013). Each member of the poly-r(C)-binding protein 1 (PCBP) family e xhib its iron chaperone activity toward ferritin. J Biol Chem, 288(24), 17791–17802. 10.1074/jbc.M113.460253 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leontieva OV, & Ionov Y (2009). RNA-binding motif protein 35A is a novel tumor suppressor for colorectal cancer. Cell Cycle, 8(3), 490–497. 10.4161/cc.8.3.7679 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leveille N, Elkon R, Davalos V, Manoharan V, Hollingworth D, Oude Vrielink J, … Agami R (2011). Selective inhib ition of microRNA accessibility by RBM 38 is required for p53 activity. Nat Commun, 2, 513 10.1038/ncomms1519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis HA, Musunuru K, Jensen KB, Edo C, Chen H, Darnell RB, & Burley SK (2000). Sequence -specific RNA binding by a Nova KH domain: implications for paraneoplastic disease and the fragile X syndrome. Cell, 100(3), 323–332. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/10676814 [DOI] [PubMed] [Google Scholar]
- Li BDL, McDonald JC, Nassar R, & De Benedetti A (1998). Clinical outcome in stage I to III breast carcinoma and elF4E overe xpression. In Annals of Surgery (Vol. 227, pp. 756–763). 10.1097/00000658-199805000-00016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D, Peng X, Yan D, Tang H, Huang F, Yang Y, & Peng Z (2011). Msi-1 is a predictor of survival and a novel therapeutic target in colon cancer. Annals of Surgical Oncology, 18(7), 2074–2083. 10.1245/s10434-011-1567-9 [DOI] [PubMed] [Google Scholar]
- Li F, Bullough KZ, Vashisht AA, Wohlschlegel JA, & Philpott CC (2016). Poly(rC)-Binding Protein 2 Regulates Hippo Signaling To Control Growth in Breast Epithelial Cells. Mol Cell Biol, 36(16), 2121–2131. 10.1128/MCB.00104-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li HY, Bourdelas A, Carron C, & Shi DL (2010). The RNA-binding protein Seb4/ RBM 24 is a direct target of MyoD and is required for myogenesis during Xenopus early development. Mech Dev, 127(5–6), 281–291. 10.1016/j.mod.2010.03.002 [DOI] [PubMed] [Google Scholar]
- Li J, He M, Xu W, & Huang S (2019). LINC01354 interacting with hnRNP-D contributes to the proliferation and metastasis in colorectal cancer through activating Wnt/β-catenin signaling pathway. Journal of Experimental and Clinical Cancer Research , 38(1), 161 10.1186/s13046-019-1150-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S, Jia Y, Jacobson B, Mc Cauley J, Kratzke R, Bitterman PB, & Wagner CR (2013). Treatment of breast and lung cancer cells with a N-7 benzyl guanosine monophosphate tryptamine phosphoramidate pronucleotide (4Ei-1) results in chemosensitization to gemcitabine and induced eIF4E proteasomal degradation. Molecular Pharmaceutics, 10(2), 523–531. 10.1021/mp300699d [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang S, Zhou Y, Chen Y, Ke G, Wen H, & Wu X (2014). Decreased e xpression of EIF4A1 after preoperative brachytherapy predicts better tumor-specific survival in cervical cancer. International Journal of Gynecological Cancer, 24(5), 908–915. 10.1097/IGC.0000000000000152 [DOI] [PubMed] [Google Scholar]
- Lin F, Wang R, Shen JJ, Wang X, Gao P, Dong K, & Zhang HZ (2008). Knockdown of RCK/p54 e xpression by RNAi inhibits proliferation of human colorectal cancer cells in vitro and in vivo. Cancer Biology and Therapy, 7(10), 1669–1676. 10.4161/cbt.7.10.6660 [DOI] [PubMed] [Google Scholar]
- Lin JC, Lee YC, Liang YC, Fann YC, Johnson KR, & Lin YJ (2017). The impact of the RBM 4-initiated splicing cascade on modulating the carcinogenic signature of colorectal cancer cells. Scientific Reports, 7 10.1038/srep44204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin X, Yang B, Liu W, Tan X, Wu F, Hu P, … Han W (2016). Interplay between PCBP2 and miRNA modulates ARHGDIA expression and function in glioma migration and invasion. Oncotarget, 7(15), 19483–19498. 10.18632/oncotarget.6869 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linder P, & Jankowsky E (2011). From unwinding to clamping - the DEAD box RNA helicase family. Nat Rev Mol Cell Biol, 12(8), 505–516. 10.1038/nrm3154 [DOI] [PubMed] [Google Scholar]
- Lingel A, Simon B, Izaurralde E, & Sattler M (2003). Structure and nucleic-acid binding of the Drosophila Argonaute 2 PAZ domain. Nature, 426(6965), 465–469. 10.1038/nature02123 [DOI] [PubMed] [Google Scholar]
- Lingel A, Simon B, Izaurralde E, & Sattler M (2004). Nucleic acid 3’-end recognition by the Argonaute2 PAZ domain. Nat Struct Mol Biol, 11(6), 576–577. 10.1038/nsmb777 [DOI] [PubMed] [Google Scholar]
- Liu J, Carmell MA, Rivas FV, Marsden CG, Thomson JM, Song JJ, … Hannon GJ (2004). Argonaute2 is the catalytic engine of mammalian RNAi. Science, 305(5689), 1437–1441. 10.1126/science.1102513 [DOI] [PubMed] [Google Scholar]
- Liu L, Luo C, Luo Y, Chen L, Liu Y, Wang Y, … Feng Y (2018). MRPL33 and its splicing regulator hnRNPK are required for mitochondria function and implicated in tumor progression. Oncogene, 37(1), 86–94. 10.1038/onc.2017.314 [DOI] [PubMed] [Google Scholar]
- Liu Y, Gai L, Liu J, Cui Y, Zhang Y, & Feng J (2015). Exp ression of poly(C)-binding protein 1 (PCBP1) in NSCLC as a negative regulator of EMT and its clinical value. Int J Clin Exp Pathol, 8(6), 7165–7172. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/26261610 [PMC free article] [PubMed] [Google Scholar]
- Liu Z, Luyten I, Bottomley MJ, Messias AC, Houngninou-Molango S, Sprangers R, … Sattler M (2001). Structural basis for recognition of the intron branch site RNA by splicing factor 1. Science, 294(5544), 1098–1102. 10.1126/science.1064719 [DOI] [PubMed] [Google Scholar]
- Liu Z, Wu G, Lin C, Guo H, Xu J, & Zhao T (2018). IGF2BP1 over-expression in skin squamous cell carcinoma cells is essential for cell growth. Biochemical and Biophysical Research Communications, 501(3), 731–738. 10.1016/j.bbrc.2018.05.057 [DOI] [PubMed] [Google Scholar]
- Lomnytska MI, Becker S, Gemoll T, Lundgren C, Habermann J, Olsson A, … Auer G (2012). Impact of genomic stability on protein expression in endometrioid endometrial cancer. British Journal of Cancer, 106(7), 1297–1305. 10.1038/bjc.2012.67 [DOI] [PMC free article] [PubMed] [Google Scholar]
- López De Silanes I, Fan J, Yang X, Zonderman AB, Potapova O, Pizer ES, & Gorospe M (2003). Role of the RNA -binding protein Hu R in colon carcinogenesis. Oncogene, 22(46), 7146–7154. 10.1038/sj.onc.1206862 [DOI] [PubMed] [Google Scholar]
- Lou P, Li C, Shi L, Xia TS, Zhou W, Wu J, … Ding Q (2017). RNPC1 enhances progesterone receptor functions by regulating its mRNA stability in breast cancer. Oncotarget, 8(10), 16387–16400. 10.18632/oncotarget.12016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu D, Searles MA, & Klug A (2003). Crystal structure of a zinc-finger-RNA complex reveals two modes of molecular recognition. Nature, 426(6962), 96–100. 10.1038/nature02088 [DOI] [PubMed] [Google Scholar]
- Lu H, Liu J, Liu S, Zeng J, Ding D, Carstens RP, … Guo W (2013). Exo70 isoform switching upon epithelial-mesenchymal transition mediates cancer cell invasion. Developmental Cell, 27(5), 560–573. 10.1016/j.devcel.2013.10.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lucchesi CA, Zhang J, Ma B, Chen M, & Chen X (2019). Dis ruption of the RBM38-eIF4E complex with a synthetic peptide PEP8 increases p53 expression. Cancer Research, 79(4), 807–818. 10.1158/0008-5472.CAN-18-2209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lunde BM, Moore C, & Varani G (2007). RNA-binding proteins: Modular design for efficient function. Nature Reviews Molecular Cell Biology, 8(6), 479–490. 10.1038/nrm2178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo K, & Zhuang K (2017). High expression of PCBP2 is associated with progression and poor prognosis in patients with glioblastoma. Biomed Pharmacother, 94, 659–665. 10.1016/j.biopha.2017.07.103 [DOI] [PubMed] [Google Scholar]
- Lykke-Andersen J, & Wagner E (2005). Recruitment and activation of mRNA decay enzymes by two ARE-mediated decay activation domains in the proteins TTP and BRF −1. Genes Dev, 19(3), 351–361. 10.1101/gad.1282305 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma JB, Ye K, & Patel DJ (2004). Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature, 429(6989), 318–322. 10.1038/nature02519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma JB, Yuan YR, Meister G, Pei Y, Tuschl T, & Patel DJ (2005). Structural basis for 5’-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature, 434(7033), 666–670. 10.1038/nature03514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma Q, & Herschman HR (1991). A corrected sequence for the predicted protein from the mitogen -inducible TIS11 primary response gene. Oncogene, 6(7), 1277–1278. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/1861870 [PubMed] [Google Scholar]
- Maas S, Patt S, Schrey M, & Rich A (2002). Underediting of glutamate receptor Glu R-B mRNA in malignant gliomas. Proceedings of the National Academy of Sciences, 98(25), 14687–14692. 10.1073/pnas.251531398 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madison BB, Liu Q, Zhong X, Hahn CM, Lin N, Emmett MJ, … Rustgi AK (2013). LIN28B promotes growth and tumorigenesis of the intestinal epithelium via Let - 7. Genes and Development, 27(20), 2233–2245. 10.1101/gad.224659.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makeyev AV, Chkheidze AN, & Liebhaber SA (1999). A set of highly conserved RNA-binding proteins, alphaCP-1 and alphaCP-2, implicated in mRNA stabilization, are coexpressed from an intronless gene and its intron-containing paralog. J Biol Chem, 274(35), 24849–24857. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/10455157 [DOI] [PubMed] [Google Scholar]
- Makeyev AV, & Liebhaber SA (2002). The poly(C)-binding proteins: a multiplicity of functions and a search for mechanis ms. RNA, 8(3), 265–278. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/12003487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malz M, Weber A, Singer S, Riehmer V, Bissinger M, Riener MO, … Breuhahn K (2009). Overe xp ression of far upstream element binding proteins: A mechanis m regulating proliferation and migration in liver cancer cells. Hepatology, 50(4), 1130–1139. 10.1002/hep.23051 [DOI] [PubMed] [Google Scholar]
- Maragh S, Miller RA, Bessling SL, Mc Gaughey DM, Wessels MW, de Graaf B, … Mc Callion AS (2011). Identi fication of RNA binding motif proteins essential for cardiovascular development. BMC Dev Biol, 11, 62 10.1186/1471-213X-11-62 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maraia RJ, Mattijssen S, Cruz-Gallardo I, & Conte MR (2017). The La and related RNA-binding proteins (LARPs): structures, functions, and evolving perspectives. Wiley Interdisciplinary Reviews: RNA 10.1002/wrna.1430 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maris C, Dominguez C, & Allain FH (2005). The RNA recognition motif, a plastic RNA -binding platform to regulate post-transcriptional gene expression. FEBS J, 272(9), 2118–2131. 10.1111/j.1742-4658.2005.04653.x [DOI] [PubMed] [Google Scholar]
- Marqus S, Pirogova E, & Piva TJ (2017). Evaluation of the use of therapeutic peptides for cancer treatment. Journal of Biomedical Science, 24(1). 10.1186/s12929-017-0328-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martineau Y, Azar R, Bousquet C, & Pyronnet S (2013). Anti-oncogenic potential of the eIF4E-binding proteins. Oncogene, 32(6), 671–677. 10.1038/onc.2012.116 [DOI] [PubMed] [Google Scholar]
- Mascini M, Palchetti I, & Tombelli S (2012). Nucleic acid and peptide aptamers: Fundamentals and bioanalytical aspects. Angewandte Chemie - International Edition 10.1002/anie.201006630 [DOI] [PubMed] [Google Scholar]
- Matter N, Herrlich P, & König H (2002). Signal-dependent regulation of splicing via phosphorylation of Sam68. Nature, 420(6916), 691–695. 10.1038/nature01153 [DOI] [PubMed] [Google Scholar]
- Mazurek A, Luo W, Krasnitz A, Hicks J, Scott Powers R, & Stillman B (2012). DDX5 regulates DNA replication and is required for cell proliferation in a subset of breast cancer cells. Cancer Discovery, 2(9), 812–825. 10.1158/2159-8290.CD-12-0116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMullen ER, Gonzalez ME, Skala SL, Tran M, Thomas D, Djomehri SI, … Kleer CG (2018). CCN6 regulates IGF2BP2 and HM GA2 signaling in metaplastic carcinomas of the breast. Breast Cancer Research and Treatment, 172(3), 577–586. 10.1007/s10549-018-4960-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehta M, Basalingappa K, Griffith JN, Andrade D, Babu A, Amreddy N, … Munshi A (2016). HuR silencing elicits oxidative stress and DNA damage and sensitizes human triple-negative breast cancer cells to radiotherapy. Oncotarget, 7(40). 10.18632/oncotarget.11706 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meisner NC, Hintersteiner M, Mueller K, Bauer R, Seifert JM, Naegeli HU, … Auer M (2007). Identification and mechanistic characterization of low-molecular-weight inhibitors for HuR. Nature Chemical Biology, 3(8), 508–515. 10.1038/nchembio.2007.14 [DOI] [PubMed] [Google Scholar]
- Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, … Rajewsky N (2013). Circular RNAs are a large class of animal RNAs with regulatory potency. Nature, 495(7441), 333–338. 10.1038/nature11928 [DOI] [PubMed] [Google Scholar]
- Mercier I, Casimiro MC, Zhou J, Wang C, Plymire C, Bryant KG, … Lisanti MP (2009). Genetic ablation of caveolin-1 drives estrogen-hypersensitivity and the development of DCIS-like mammary lesions. American Journal of Pathology, 174(4), 1172–1190. 10.2353/ajpath.2009.080882 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mercuri E, Darras BT, Chiriboga CA, Day JW, Campbell C, Connolly AM, … Group CS (2018). Nusinersen ve rsus Sham Control in Later-Onset Spinal Muscular Atrophy. N Engl J Med, 378(7), 625–635. 10.1056/NEJMoa1710504 [DOI] [PubMed] [Google Scholar]
- Miles WO, Lembo A, Volorio A, Brachtel E, Tian B, Sgroi D, … Dyson N (2016). Alternative polyadenylation in t riple-negative breast tumors allows NRAS and c-JUN to bypass PUMILIO posttranscriptional regulation. Cancer Research, 76(24), 7231–7241. 10.1158/0008-5472.CAN-16-0844 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miles WO, Tschöp K, Herr A, Ji JY, & Dyson NJ (2012). Pumilio facilitates miRNA regulation of the E2F3 oncogene. Genes and Development, 26(4), 356–368. 10.1101/gad.182568.111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milke L, Schulz K, Weigert A, Sha W, Schmid T, & Brune B (2013). Depletion of tristetraprolin in breast cancer ce lls increases interleukin-16 expression and promotes tumor infiltration with monocytes/macrophages. Carcinogenesis, 34(4), 850–857. 10.1093/carcin/bgs387 [DOI] [PubMed] [Google Scholar]
- Minuesa G, Antczak C, Shum D, Radu C, Bhinder B, Li Y, … Kharas MG (2014). A 1536 -well fluorescence polarization assay to screen for modulators of the MUSASHI family of RNA-binding proteins. Comb Chem High Throughput Screen, 17(7), 596–609. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/24912481 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyaji K, Nakagawa Y, Matsumoto K, Yoshida H, Morikawa H, Hongou Y, … Akao Y (2003). Over expression of a DEAD bo x/RNA helicase protein, rck/p54, in human hepatocytes from patients with hepatitis C virus -related chronic hepatitis and its implication in hepatocellular carcinogenesis. Journal of Viral Hepatitis, 10(4), 241–248. 10.1046/j.1365-2893.2003.00447.x [DOI] [PubMed] [Google Scholar]
- Miyamoto S, Hidaka K, Jin D, & Morisaki T (2009). RNA-binding proteins Rbm38 and Rbm24 regulate myogenic differentiation via p21 -dependent and –independent regulatory pathways. Genes to Cells, 14(11), 1241–1252. 10.1111/j.1365-2443.2009.01347.x [DOI] [PubMed] [Google Scholar]
- Mizutani A, Koinuma D, Seimiya H, & Miyazono K (2016). The Arkadia-ESRP2 axis suppresses tumor progression: Analyses in clear-cell renal cell carcinoma. Oncogene, 35(27), 3514–3523. 10.1038/onc.2015.412 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Modelska A, Turro E, Russell R, Beaton J, Sbarrato T, Spriggs K, … Le Quesne J (2015). The malignant phenotype in breast cancer is driven by eIf4A1-mediated changes in the translational landscape. Cell Death and Disease, 6(1). 10.1038/cddis.2014.542 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moerke NJ, Aktas H, Chen H, Cantel S, Reibarkh MY, Fahmy A, … Wagner G (2007). Small-molecule inhibition of the interaction between the translation initiat ion factors eIF4E and eIF4G. Cell, 128(2), 257–267. 10.1016/j.cell.2006.11.046 [DOI] [PubMed] [Google Scholar]
- Molenaar JJ, Domingo-Fernández R, Ebus ME, Lindner S, Koster J, Drabek K, … Schulte JH (2012). LIN28B induces neuroblastoma and enhances MYCN levels via let-7 suppression. Nature Genetics, 44(11), 1199–1206. 10.1038/ng.2436 [DOI] [PubMed] [Google Scholar]
- Mongroo PS, Noubissi FK, Cuatrecasas M, Kalabis J, King CE, Johnstone CN, … Rustgi AK (2011). IMP −1 displays cross-talk with K-Ras and modulates colon cancer cell survival through the novel proapoptotic protein CYFIP2. Cancer Research, 71(6), 2172–2182. 10.1158/0008-5472.CAN-10-3295 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore S, Jarvelin AI, Davis I, Bond GL, & Castello A (2018). Expanding horizons: new roles for non -canonical RNA-binding proteins in cancer. Curr Opin Genet Dev, 48, 112–120. 10.1016/j.gde.2017.11.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morita Y, Leslie M, Kameyama H, Volk DE, & Tanaka T (2018). Aptamer therapeutics in cancer: Current and future. Cancers 10.3390/cancers10030080 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moumen A, Masterson P, O’Connor MJ, & Jackson SP (2005). hnRNP K: an HDM2 target and transcriptional coactivator of p 53 in response to DNA damage. Cell, 123(6), 1065–1078. 10.1016/j.cell.2005.09.032 [DOI] [PubMed] [Google Scholar]
- Müller S, Bley N, Glaß M, Busch B, Rousseau V, Misiak D, … Hüttelmaier S (2018). IGF2BP1 enhances an aggressive tumor cell phenotype by impairing miRNA - directed downregulation of oncogenic factors. Nucleic Acids Research, 46(12), 6285–6303. 10.1093/nar/gky229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mura M, Hopkins TG, Michael T, Abd-Latip N, Weir J, Aboagye E, … Blagden SP (2015). LARP1 post-transcriptionally regulates mTOR and contributes to cancer progression. Oncogene, 34(39), 5025–5036. 10.1038/onc.2014.428 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muralidharan R, Babu A, Amreddy N, Srivastava A, Chen A, Zhao YD, … Ramesh R (2017). Tumor-targeted Nanoparticle Delivery of HuR siRNA Inhibits Lung Tumor Growth In Vit ro and In Vivo By Disrupting the Oncogenic Activity of the RNA -binding Protein HuR . Molecular Cancer Therapeutics, 16(8), 1470–1486. 10.1158/1535-7163.mct-17-0134 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muralidharan R, Mehta M, Ahmed R, Roy S, Xu L, Aube J, … Munshi A (2017). HuR-targeted small molecule inhibitor e xhib its cytotoxic ity towards human lung cancer cells. Sci Rep, 7(1), 9694 10.1038/s41598-017-07787-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nabors LB, Gillespie GY, Harkins L, & King PH (2001). HuR, a RNA stability factor, is e xpressed in malignant brain tumors and binds to adenine- and uridine-rich elements within the 3′ untranslated regions of cytokine and angiogenic factor mRNAs. Cancer Research, 61(5), 2154–2161. [PubMed] [Google Scholar]
- Nadiminty N, Tummala R, Lou W, Zhu Y, Shi XB, Zou JX, … Gao AC (2012). MicroRNA let −7c is downregulated in prostate cancer and suppresses prostate cancer growth. PLoS ONE, 7(3). 10.1371/journal.pone.0032832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagano K, Masters JR, Akpan A, Yang A, Corless S, Wood C, … Naaby-Hansen S (2004). Differential protein synthesis and expression levels in normal and neoplastic human prostate cells and their regulation by type I and II interferons. Oncogene, 23(9), 1693–1703. 10.1038/sj.onc.1207297 [DOI] [PubMed] [Google Scholar]
- Nagaoka K, Fujii K, Zhang H, Usuda K, Watanabe G, Ivshina M, & Richter JD (2016). CPEB1 mediates epithelial-to-mesenchyme transition and breast cancer metastasis. Oncogene, 35(22), 2893–2901. 10.1038/onc.2015.350 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakagawa Y, Morikawa H, Hirata I, Shiozaki M, Matsumoto A, Maemura K, … Akao Y (1999). Overe xpression of rck/ p54, a DEAD box protein, in human colorectal tumours. British Journal of Cancer, 80(5–6), 914–917. 10.1038/sj.bjc.6690441 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura M, Okano H, Blendy JA, & Montell C (1994). Musashi, a neural RNA-binding protein required for drosophila adult e xterna l sensory organ development. Neuron, 13(1), 67–81. 10.1016/0896-6273(94)90460-X [DOI] [PubMed] [Google Scholar]
- Nakatake M, Monte-Mor B, Debili N, Casadevall N, Ribrag V, Solary E, … Plo I (2012). JAK2 V617F negatively regulates p53 stabilizat ion by enhancing MDM 2 via la expression in myeloproliferative neoplasms. Oncogene, 31(10), 1323–1333. 10.1038/onc.2011.313 [DOI] [PubMed] [Google Scholar]
- Nanduri S, Carpick BW, Yang Y, Williams BR, & Qin J (1998). Structure of the double-stranded RNA-binding domain of the protein kinase PKR reveals the molecular basis of its dsRNA-mediated activation. EMBO J, 17(18), 5458–5465. 10.1093/emboj/17.18.5458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nathan CAO, Franklin S, Abreo FW, Nassar R, De Benedetti A, & Glass J (1999). Analysis of surgical margins with the molecular marker eIF4E: A prognostic factor in patients with head and neck cancer. Journal of Clinical Oncology, 17(9), 2909–2914. [DOI] [PubMed] [Google Scholar]
- Neelamraju Y, Gonzalez-Perez A, Bhat-Nakshatri P, Nakshatri H, & Janga SC (2018). Mutational landscape of RNA-binding proteins in human cancers. RNA Biology, 15(1), 115–129. 10.1080/15476286.2017.1391436 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nemlich Y, Greenberg E, Ortenberg R, Besser MJ, Barshack I, Jacob-Hirsch J, … Markel G (2013). Micro RNA -mediated loss of ADAR1 in metastatic melanoma promotes tumor growth. Journal of Clinical Investigation 10.1172/JCI62980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen LH, Robinton DA, Seligson MT, Wu L, Li L, Rakheja D, … Zhu H (2014). Lin28b is sufficient to drive liver cancer and necessary for its maintenance in murine models. Cancer Cell, 26(2), 248–261. 10.1016/j.ccr.2014.06.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nussbacher JK, Tabet R, Yeo GW, & Lagier-Tourenne C (2019). Disruption of RNA Metabolism in Neurological Diseases and Emerging Therapeutic Interventions. Neuron 10.1016/j.neuron.2019.03.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oberstrass FC, Auweter SD, Erat M, Hargous Y, Henning A, Wenter P, … Allain FH (2005). Structure of PT B bound to RNA: specific binding and implications for splicing regulation. Science, 309(5743), 2054–2057. 10.1126/science.1114066 [DOI] [PubMed] [Google Scholar]
- Oguro A, Ohtsu T, Svitkin YV, Sonenberg N, & Nakamura Y (2003). RNA aptamers to in itiation factor 4A helicase hinder cap-dependent translation by blocking ATP hydrolysis. Rna, 9(4), 394–407. 10.1261/rna.2161303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohe K, Yoshida M, Nakano-Kobayashi A, Hosokawa M, Sako Y, Sakuma M, … Hagiwara M (2017). RBM 24 promotes U1 snRNP recognition of the mutated 5′ splice site in the IKBKAP gene of familial dysautonomia. Rna, 23(9), 1393–1403. 10.1261/rna.059428.116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orphanides G, & Reinberg D (2002). A unified theory of gene expression. Cell, 108(4), 439–451. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/11909516 [DOI] [PubMed] [Google Scholar]
- Ortiz-Zapater E, Pineda D, Martínez-Bosch N, Fernández-Miranda G, Iglesias M, Alameda F, … Navarro P (2012). Key contribution of CPEB4mediated translational control to cancer progression. Nature Medicine, 18(1), 83–90. 10.1038/nm.2540 [DOI] [PubMed] [Google Scholar]
- Oskarsson T, Acharyya S, Zhang XHF, Vanharanta S, Tavazoie SF, Morris PG, … Massagué J (2011). Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs. Nature Medicine, 17(7), 867–874. 10.1038/nm.2379 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palanichamy JK, Tran TM, Howard JM, Contreras JR, Fernando TR, Sterne-Weiler T, … Rao DS (2016). RNA-binding protein IGF2BP3 targeting of oncogenic transcripts promotes hematopoietic progenitor proliferation. Journal of Clinical Investigation, 126(4), 1495–1511. 10.1172/JCI80046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park JM, Lee TH, & Kang TH (2018). Roles of Tristetraprolin in Tumorigenesis. Int J Mol Sci, 19(11). 10.3390/ijms19113384 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paronetto MP, Cappellari M, Busà R, Pedrotti S, Vita li R, Comstock C, … Sette C (2010). Alternative splicing of the cyclin D1 proto-oncogene is regulated by the RNA - binding protein Sam68. Cancer Research, 70(1), 229–239. 10.1158/0008-5472.CAN-09-2788 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pascual M, Vicente M, Monferrer L, & Artero R (2006). The Muscleblind family of proteins: an emerging class of regula tors of developmentally programmed alternative splicing. Differentiation, 74(2–3), 65–80. 10.1111/j.1432-0436.2006.00060.x [DOI] [PubMed] [Google Scholar]
- Patry C, Bouchard L, Labrecque P, Gendron D, Lemieux B, Toutant J, … Chabot B (2003). Small Interfering RNA -Mediated Reduction in Heterogeneous Nuclear Ribonucleoparticule A1/A2 Proteins Induces Apoptosis in Human Cancer Cells but not in Normal Mortal Cell Lines. Cancer Research, 63(22), 7679–7688. [PubMed] [Google Scholar]
- Paz-Yaacov N, Bazak L, Buchumenski I, Porath HT, Danan-Gotthold M, Knisbacher BA, … Levanon EY (2015). Elevated RNA Editing Activity Is a Major Contributor to Transcriptomic Diversity in Tumors. Cell Reports 10.1016/j.celrep.2015.08.080 [DOI] [PubMed] [Google Scholar]
- Paz N, Levanon EY, Amariglio N, Heimberger AB, Ram Z, Constantini S, … Rechavi G ( 2007). Altered adenosine-to-inosine RNA editing in human cancer. Genome Research, 17(11), 1586–1595. 10.1101/gr.6493107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peffley DM, Sharma C, Hentosh P, & Buechler RD (2007). Perillyl alcohol and genistein differentially regulat e PKB/Akt and 4E-BP1 phosphorylation as well as eIF4E/eIF4G interactions in human tumor cells. Arch Biochem Biophys, 465(1), 266–273. 10.1016/j.abb.2007.05.022 [DOI] [PubMed] [Google Scholar]
- Pelletier J, Graff J, Ruggero D, & Sonenberg N (2015). Targeting the eIF4F translation initiation complex: A critical nexus for cancer development. Cancer Research, 75(2), 250–263. 10.1158/0008-5472.CAN-14-2789 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng X, Xu X, Wang Y, Hawke DH, Yu S, Han L, … Mills GB (2018). A-to-I RNA Editing Contributes to Proteomic Diversity in Cancer. Cancer Cell, 33(5), 817–828.e7 10.1016/j.ccell.2018.03.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pereira B, Billaud M, & Almeida R (2017). RNA-Binding Proteins in Cancer: Old Players and New Actors. Trends in Cancer, 3(7), 506–528. 10.1016/j.trecan.2017.05.003 [DOI] [PubMed] [Google Scholar]
- Pérez-Guijarro E, Karras P, Cifdaloz M, Martínez-Herranz R, Canõn E, Granã O, … Soengas MS (2016). Lineage-specific roles of the cytoplasmic polyadenylation factor CPEB4 in the regulation of melanoma drivers. Nature Communications, 7 10.1038/ncomms13418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peters TL, Tillotson J, Yeomans AM, Wilmore S, Lemm E, Jimenez-Romero C, … Schatz JH (2018). Target-Based Screening against eIF4A1 Reveals the Marine Natural Product Elatol as a Novel Inhibitor of Translation Initiation with In Vivo Antitumor Activity. Clin Cancer Res, 24(17), 4256–4270. 10.1158/1078-0432.CCR-17-3645 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petz M, Them N, Huber H, Beug H, & Mikulits W (2012). La enhances IRES-mediated translation of laminin B1 during malignant epithelial to mesenchymal transition. Nucleic Acids Research, 40(1), 290–302. 10.1093/nar/gkr717 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pillai MR, Chacko P, Kesari LA, Jayaprakash PG, Jayaram HN, & Antony AC (2003). Exp ression of folate receptors and heterogeneous nuclear ribonucleoprotein E1 in women with human papillomavirus mediated transformation of cervical t issue to cancer. J Clin Pathol, 56(8), 569–574. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/12890803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pio R, Zudaire I, Pino I, Castano Z, Zabalegui N, Vicent S, … Montuenga LM (2004). Alpha CP −4, encoded by a putative tumor suppressor gene at 3p21, but not its alternative splice variant alpha CP-4a, is underexpressed in lung cancer. Cancer Res, 64(12), 4171–4179. 10.1158/0008-5472.CAN-03-2982 [DOI] [PubMed] [Google Scholar]
- Piskounova E, Polytarchou C, Thornton JE, Lapierre RJ, Pothoulakis C, Hagan JP, … Gregory RI (2011). Lin28A and Lin 28B inhibit let-7 Micro RNA biogenesis by distinct mechanisms. Cell, 147(5), 1066–1079. 10.1016/j.cell.2011.10.039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poon KL, Tan KT, Wei YY, Ng CP, Colman A, Korzh V, & Xu XQ (2012). RNA-binding protein RBM 24 is required for sarcomere assembly and heart contractility. Cardiovasc Res, 94(3), 418–427. 10.1093/cvr/cvs095 [DOI] [PubMed] [Google Scholar]
- Preca BT, Bajdak K, Mock K, Sundararajan V, Pfannstiel J, Maurer J, … Stemmler MP (2015). A self-enforcing CD44s/ZEB1 feedback loop maintains EMT and stemness properties in cancer cells. International Journal of Cancer, 137(11), 2566–2577. 10.1002/ijc.29642 [DOI] [PubMed] [Google Scholar]
- Pugh TJ, Weeraratne SD, Archer TC, Pomeranz Krummel DA, Auclair D, Bochicchio J, … Cho YJ (2012). Medulloblastoma exome sequencing uncovers subtype - specific somatic mutations. Nature 10.1038/nature11329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puppo M, Bucci G, Rossi M, Giovarelli M, Bordo D, Moshiri A, … Briata P (2016). miRNA-Mediated KHSRP Silencing Rewires Distinct Post-transcriptional Programs during TGF-β-Induced Epithelial-to-Mesenchymal Transition. Cell Reports, 16(4), 967–978. 10.1016/j.celrep.2016.06.055 [DOI] [PubMed] [Google Scholar]
- Radi M, Falchi F, Garbelli A, Samuele A, Bernardo V, Paolucci S, … Botta M (2012). Discovery of the first small molecule inhibitor of human DDX3 specifically designed to target the RNA binding site: towards the next generation HIV-1 inhibitors. Bioorg Med Chem Lett, 22(5), 2094–2098. 10.1016/j.bmcl.2011.12.135 [DOI] [PubMed] [Google Scholar]
- Raza F, Waldron JA, & Quesne JL (2015). Translational dysregulation in cancer: eIF4A isoforms and sequence determin ants of eIF4A dependence. Biochemical Society Transactions, 43(6), 1227–1233. 10.1042/bst20150163 [DOI] [PubMed] [Google Scholar]
- Ren C, Zhang J, Yan W, Zhang Y, & Chen X (2016). RNA-binding Protein PCBP2 Regulates p73 Expression and p73-dependent Antioxidant Defense. J Biol Chem, 291(18), 9629–9637. 10.1074/jbc.M115.712125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richard S, Voge l G, Huot ME, Guo T, Muller WJ, & Lukong KE (2008). Sam68 haploinsufficiency delays onset of mammary tumorigenesis and metastasis. Oncogene, 27(4), 548–556. 10.1038/sj.onc.1210652 [DOI] [PubMed] [Google Scholar]
- Richter JD, & Sonenberg N (2005). Regulation of cap-dependent translation by eIF4E inhibitory proteins. Nature, 433(7025), 477–480. 10.1038/nature03205 [DOI] [PubMed] [Google Scholar]
- Richter J, Schlesner M, Hoffmann S, Kreuz M, Leich E, Burkhardt B, … Siebert R (2012). Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nature Genetics, 44(12), 1316–1320. 10.1038/ng.2469 [DOI] [PubMed] [Google Scholar]
- Rigo F, Hua Y, Krainer AR, & Bennett CF (2012). Antisense-based therapy for the treatment of spinal muscular atrophy. J Cell Biol, 199(1), 21–25. 10.1083/jcb.201207087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rivas FV, Tolia NH, Song JJ, Aragon JP, Liu J, Hannon GJ, & Joshua-Tor L (2005). Purified Argonaute2 and an siRNA form recombinant human RISC. Nat Struct Mol Biol, 12(4), 340–349. 10.1038/nsmb918 [DOI] [PubMed] [Google Scholar]
- Rokavec M, Kaller M, Horst D, & Hermeking H (2017). Pan-cancer EMT-signature identifies RBM47 down-regulation during colorectal cancer progression. Scientific Reports, 7(1). 10.1038/s41598-017-04234-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Romeo C, Weber MC, Zarei M, De Cicco D, Chand SN, Lobo AD, … Brody JR (2016). Hu R Contributes to TRAIL Resistance by Restricting Death Receptor 4 Expression in Pancreatic Cancer Cells. Molecular Cancer Research, 14(7), 599–611. 10.1158/1541-7786.mcr-15-0448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roos M, Pradere U, Ngondo RP, Behera A, Allegrini S, Civenni G, … Hall J (2016). A Small-Molecule Inhibitor of Lin28. ACS Chem Biol, 11(10), 2773–2781. 10.1021/acschembio.6b00232 [DOI] [PubMed] [Google Scholar]
- Rosenwald IB, Hutzler MJ, Wang S, Savas L, & Fraire AE (2001). Expression of eukaryotic translation initiation factors 4E and 2α is increased frequently in bronchioloalveolar but not in squamous cell carcinomas of the lung. Cancer, 92(8), 2164–2171. [DOI] [PubMed] [Google Scholar]
- Ross J, Lemm I, & Berberet B (2001). Overexpression of an mRNA-binding protein in human colorectal cancer. Oncogene, 20(45), 6544–6550. 10.1038/sj.onc.1204838 [DOI] [PubMed] [Google Scholar]
- Roth TL, Puig-Saus C, Yu R, Shifrut E, Carnevale J, Li PJ, … Marson A (2018). Reprogramming human T cell function and specificity with non -viral genome targeting. Nature, 559(7714), 405–409. 10.1038/s41586-018-0326-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryter JM, & Schultz SC (1998). Molecular basis of double-stranded RNA-protein interactions: structure of a dsRNA-binding domain complexed with dsRNA. EMBO J, 17(24), 7505–7513. 10.1093/emboj/17.24.7505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryu MS, Zhang D, Protchenko O, Shakoury-Elizeh M, & Philpott CC (2017). PCBP1 and NCOA 4 regulate erythroid iron storage and heme biosynthesis. J Clin Invest, 127(5), 1786–1797. 10.1172/JCI90519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saha S, Bardelli A, Buckhaults P, Velculescu VE, Rago C, St Croix B, … Voge lstein B (2001). A phosphatase associated with metastasis of colorectal cancer. Science, 294(5545), 1343–1346. 10.1126/science.1065817 [DOI] [PubMed] [Google Scholar]
- Sakakibara SI, Imai T, Hamaguchi K, Okabe M, Aruga J, Nakajima K, … Okano H (1996). Mouse-Musashi-1, a neural RNA-Binding protein highly enriched in the mammalian CNS stem cell. Developmental Biology, 176(2), 230–242. 10.1006/dbio.1996.0130 [DOI] [PubMed] [Google Scholar]
- Sakakibara S, Nakamura Y, Satoh H, & Okano H (2001). Rna-binding protein Musashi2: developmentally regulated e xpre ssion in neural precursor cells and subpopulations of neurons in mammalian CNS. The Journal of Neuroscience : The Official Journal of the Society for Neuroscience, 21(20), 8091–8107. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/11588182 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakurai T, Isogaya K, Sakai S, Morikawa M, Morishita Y, Ehata S, … Koinuma D (2017). RNA -binding motif protein 47 inhibits Nrf2 activity to suppress tumor growth in lung adenocarcinoma. Oncogene, 36(35), 5083 10.1038/onc.2017.191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samanta S, Guru S, Elaimy AL, Amante JJ, Ou J, Yu J, … Mercurio AM (2018). IMP3 Stabilization of W NT5B mR NA Facilitates TAZ Activation in Breast Cancer. Cell Reports, 23(9), 2559–2567. 10.1016/j.celrep.2018.04.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanna MD, Peroni D, Quattrone A, Ghe lardini C, & Galeotti N (2015). Spinal Ry R2 pathway regulated by the RNA -binding protein HuD induces pain hypersensitivity in antiretroviral neuropathy. Exp Neurol, 267, 53–63. 10.1016/j.expneurol.2015.02.036 [DOI] [PubMed] [Google Scholar]
- Sanna MD, Quattrone A, & Galeotti N (2017). Silencing of the RNA-binding protein HuR attenuates hyperalgesia and motor disability in experimental autoimmune encephalomyelitis. Neuropharmacology, 123, 116–125. 10.1016/j.neuropharm.2017.06.005 [DOI] [PubMed] [Google Scholar]
- Schatz JH, Oricchio E, Wolfe AL, Jiang M, Linkov I, Maragulia J, … Wendel H-G (2011). Targeting cap-dependent translation blocks converging survival signals by AKT and PIM kinases in lymphoma. The Journal of Experimental Medicine, 208(9), 1799–1807. 10.1084/jem.20110846 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmiedel D, Tai J, Yamin R, Berhani O, Bauman Y, & Mandelboim O (2016). The RNA binding protein IMP3 facilitates tumor immune escape by downregulating the stress-induced ligands ULPB2 and MICB. ELife, 5, 1–18. 10.7554/eLife.13426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schubert M, Edge RE, Lario P, Cook MA, Strynadka NC, Mackie GA, & Mc Intosh LP (2004). Structural characterization of the RNase E S1 domain and identification of its oligonucleotide-binding and dimerization interfaces. J Mol Biol, 341(1), 37–54. 10.1016/j.jmb.2004.05.061 [DOI] [PubMed] [Google Scholar]
- Scoumanne A, Cho SJ, Zhang J, & Chen X (2011). The cyclin-dependent kinase inhibitor p21 is regulated by RNA-binding protein PCBP4 via mRNA stability. Nucleic Acids Res, 39(1), 213–224. 10.1093/nar/gkq778 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Searles MA, Lu D, & Klug A (2000). The role of the central zinc fingers of transcription factor IIIA in binding to 5 S RNA. J Mol Biol, 301(1), 47–60. 10.1006/jmbi.2000.3946 [DOI] [PubMed] [Google Scholar]
- Seetharaman S, Flemyng E, Shen J, Conte MR, & Ridley AJ (2016). The RNA-binding protein LARP4 regulates cancer cell migration and invasion. Cytoskeleton, 73(11), 680–690. 10.1002/cm.21336 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sengoku T, Nureki O, Nakamura A, Kobayashi S, & Yokoya ma S (2006). Structural basis for RNA unwinding by the DEAD-box protein Drosophila Vasa. Cell, 125(2), 287–300. 10.1016/j.cell.2006.01.054 [DOI] [PubMed] [Google Scholar]
- Senoo J, Mikata R, Kishimoto T, Hayashi M, Kusakabe Y, Yasui S, … Kato N (2018). Immunohistochemical analysis of IMP3 and p53 e xp ression in endoscopic ultrasound-guided fine needle aspiration and resected specimens of pancreatic diseases. Pancreatology 10.1016/j.pan.2017.12.010 [DOI] [PubMed] [Google Scholar]
- Shao R, Scully SJ, Yan W, Bentley B, Mueller J, Brown C, … Schwartz LM (2012). The novel lupus antigen related protein acheron enhances the development of human breast cancer. International Journal of Cancer, 130(3), 544–554. 10.1002/ijc.26015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaoyan X, Juanjuan Y, Yalan T, Ping H, Jianzhong L, & Qinian W (2013). Downregulation of EIF4A2 in non-small-cell lung cancer associates with poor prognosis. Clinical Lung Cancer, 14(6), 658–665. 10.1016/j.cllc.2013.04.011 [DOI] [PubMed] [Google Scholar]
- Shapiro IM, Cheng AW, Flytzanis NC, Balsamo M, Condeelis JS, Oktay MH, … Gertler FB (2011). An emt-driven alternative splicing program occurs in human breast cancer and modulates cellular phenotype. PLoS Genetics, 7(8). 10.1371/journal.pgen.1002218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shelton PM, Duran A, Nakanishi Y, Reina-Campos M, Kasashima H, Llado V, … Moscat J (2018). The Secretion of miR-200s by a PKCζ/ADAR2 Signaling Axis Promotes Liver Metastasis in Colorectal Cancer. Cell Reports 10.1016/j.celrep.2018.03.118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheng W, Dong M, Chen C, Li Y, Liu Q, & Dong Q (2016). Musashi2 promotes the development and progression of pancreatic cancer by down-regulating Numb protein. Oncotarget, 8(9). 10.18632/oncotarget.8736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H, Bencze KZ, Stemmler TL, & Philpott CC (2008). A cytosolic iron chaperone that delivers iron to ferrit in. Science, 320(5880), 1207–1210. 10.1126/science.1157643 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H, Li H, Yuan R, Guan W, Zhang X, Zhang S, … Wang H (2018). PCBP1 depletion promotes tumorigenesis through attenuation of p27(Kip 1) mRNA stability and translation. J Exp Clin Cancer Res, 37(1), 187 10.1186/s13046-018-0840-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi L, Xia TS, Wei XL, Zhou W, Xue J, Cheng L, … Ding Q (2015). Estrogen receptor (ER) was regulated by RNPC1 stabilizing mRNA in ER positive breast cancer. Oncotarget, 6(14), 12264–12278. 10.18632/oncotarget.3654 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimada H, Ichikawa H, Nakamura S, Katsu R, Iwasa M, Kitabayashi I, & Ohki M (2000). Analysis of genes under the downstream control of the t(8;21) fusion protein AML1-MTG8: overe xpression of the TIS11b (ERF-1, cMG1) gene induces myeloid cell proliferation in response to G-CSF. Blood, 96(2), 655–663. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/10887131 [PubMed] [Google Scholar]
- Shin S, Rossow KL, Grande JP, & Janknecht R (2007). Involvement of RNA helicases p68 and p72 in colon cancer. Cancer Research, 67(16), 7572–7578. 10.1158/0008-5472.CAN-06-4652 [DOI] [PubMed] [Google Scholar]
- Shoshan E, Mobley AK, Braeuer RR, Kamiya T, Huang L, Vasquez ME, … Bar-Eli M (2015). Reduced adenosine-to-inosine miR-455–5p editing promotes melanoma growth and metastasis. Nature Cell Biology, 17(3), 311–321. 10.1038/ncb3110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shu L, Yan W, & Chen X (2006). RNPC1, an RNA-binding protein and a target of the p 53 family, is required for maintaining the stability of the basal and stress -induced p21 transcript. Genes Dev, 20(21), 2961–2972. 10.1101/gad.1463306 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sickmier EA, Frato KE, Shen H, Paranawithana SR, Green MR, & Kielkopf CL (2006). Structural basis for p olypyrimidine tract recognition by the essential pre-mRNA splicing factor U2AF65. Mol Cell, 23(1), 49–59. 10.1016/j.molcel.2006.05.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sillars-Hardebol AH, Carvalho B, Belién JAM, De Wit M, Delis-Van Diemen PM, Tijssen M, … Fijneman RJA (2012). CSE1L, DIDO1 and RBM 39 in colorectal adenoma to carcinoma progression. Cellular Oncology, 35(4), 293–300. 10.1007/s13402-012-0088-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siomi H, Matunis MJ, Michael WM, & Dreyfuss G (1993). The pre -mRNA binding K protein contains a novel evolutionary conserved motif. Nucleic Acids Research, 21(5), 1193–1198. 10.1093/nar/21.5.1193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sommer G, Dittmann J, Kuehnert J, Reumann K, Schwartz PE, Will H, … Heise T (2011). The RNA -binding protein la contributes to cell proliferation and CCND1 expression. Oncogene, 30(4), 434–444. 10.1038/onc.2010.425 [DOI] [PubMed] [Google Scholar]
- Song JJ, Liu J, Tolia NH, Schneiderman J, Smith SK, Martienssen RA, … Joshua-Tor L (2003). The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nat Struct Biol, 10(12), 1026–1032. 10.1038/nsb1016 [DOI] [PubMed] [Google Scholar]
- Song JJ, Smith SK, Hannon GJ, & Joshua-Tor L (2004). Crystal structure of Argonaute and its implications for RISC slicer activity. Science, 305(5689), 1434–1437. 10.1126/science.1102514 [DOI] [PubMed] [Google Scholar]
- Song L, Wang L, Li Y, Xiong H, Wu J, Li J, & Li M (2010). Sam68 up-regulation correlates with, and its down-regulation inhibits, proliferation and tumourigenicity of breast cancer cells. Journal of Pathology, 222(3), 227–237. 10.1002/path.2751 [DOI] [PubMed] [Google Scholar]
- Song YK, Guo H, Barengo N, & Naora H (2009). Inhibition of ovarian cancer growth by a tumor-targeting peptide that binds eukaryotic translation init iation factor 4E. Clinical Cancer Research, 15(13), 4336–4347. 10.1158/1078-0432.CCR-08-2924 [DOI] [PubMed] [Google Scholar]
- Srikantan S, & Gorospe M (2012). HuR function in disease. Front Biosci (Landmark Ed), 17, 189–205. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/22201738 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stavraka C, & Blagden S (2015). The La-related proteins, a family with connections to cancer. Biomolecules 10.3390/biom5042701 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stefanovic B, Manojlovic Z, Vied C, Badger CD, & Stefanovic L (2019). Discovery and evaluation of inhibitor of LARP6 as specific antifibrotic compound. Scientific Reports, 9(1). 10.1038/s41598-018-36841-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stefl R, Oberstrass FC, Hood JL, Jourdan M, Zimmermann M, Skrisovska L, … Allain FH (2010). The solution structure of the ADAR2 dsRBM-RNA complex reveals a sequence-specific readout of the minor groove. Cell, 143(2), 225–237. 10.1016/j.cell.2010.09.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stein CA, & Castanotto D (2017). FDA-Approved Oligonucleotide Therapies in 2017. Molecular Therapy 10.1016/j.ymthe.2017.03.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, … Grandis JR (2011). The mutatio nal landscape of head and neck squamous cell carcinoma. Science, 333(6046), 1157–1160. 10.1126/science.1208130 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stumpo DJ, Broxmeyer HE, Ward T, Cooper S, Hangoc G, Chung YJ, … Blackshear PJ (2009). Targeted disrupt ion of Zfp36l2, encoding a CCCH tandem zinc finger RNA-binding protein, results in defective hematopoiesis. Blood, 114(12), 2401–2410. 10.1182/blood-2009-04-214619 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stumpo DJ, Byrd NA, Phillips RS, Ghosh S, Maronpot RR, Castranio T, … Blackshear PJ (2004). Chorioallantoic fusion defects and embryonic lethality resulting from disruption of Zfp 36L1, a gene encoding a CCCH tandem zinc finger protein of the Tristetraprolin family. Mol Cell Biol, 24(14), 6445–6455. 10.1128/MCB.24.14.6445-6455.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian AR (1983). Structure and functions of ribosomal protein S1. Prog Nucleic Acid Res Mol Biol, 28, 101–142. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/6348874 [DOI] [PubMed] [Google Scholar]
- Sureban SM, May R, George RJ, Dieckgraefe BK, Mc Leod HL, Ramalingam S, … Houchen CW (2008). Knockdown of RNA Binding Protein Musashi-1 Leads to Tumor Regression In Vivo. Gastroenterology, 134(5). 10.1053/j.gastro.2008.02.057 [DOI] [PubMed] [Google Scholar]
- Sutherland JM, Fraser BA, Sobinoff AP, Pye VJ, Davidson T-L, Siddall NA, … Mc Laughlin EA (2014). Developmental Expression of Musashi-1 and Musashi-2 RNA-Binding Proteins During Spermatogenesis: Analysis of the Deleterious Effects of Dysregulated Expre ssion1. Biology of Reproduction, 90(5). 10.1095/biolreprod.113.115261 [DOI] [PubMed] [Google Scholar]
- Sweetser DA, Peniket AJ, Haaland C, Blomberg AA, Zhang Y, Zaidi ST, … Daly SB (2005). Delineation of the minima l commonly deleted segment and identification of candidate tumor-suppressor genes in del(9q) acute myeloid leukemia. Genes Chromosomes Cancer, 44(3), 279–291. 10.1002/gcc.20236 [DOI] [PubMed] [Google Scholar]
- Tai AL, Mak W, Ng PK, Chua DT, Ng MY, Fu L, … Guan XY (2006). High-throughput loss-of-heterozygosity study of chromosome 3p in lung cancer using single-nucleotide polymorphism markers. Cancer Res, 66(8), 4133–4138. 10.1158/0008-5472.CAN-05-2775 [DOI] [PubMed] [Google Scholar]
- Taniguchi K, Iwatsuki A, Sugito N, Shinohara H, Kuranaga Y, Oshikawa Y, … Akao Y (2018). Oncogene RNA helicase DDX6 promotes the process of c-Myc e xpression in gastric cancer cells. Molecular Carcinogenesis, 57(5), 579–589. 10.1002/mc.22781 [DOI] [PubMed] [Google Scholar]
- Tauler J, Zudaire E, Liu H, Shih J, & Mulshine JL (2010). hnRNP A2/ B1 modulates epithelial-mesenchymal transition in lung cancer cell lines. Cancer Research, 70(18), 7137–7147. 10.1158/0008-5472.CAN-10-0860 [DOI] [PubMed] [Google Scholar]
- Taylor GA, Carballo E, Lee DM, Lai WS, Thompson MJ, Patel DD, … Blackshear PJ (1996). A pathogenetic role for TNF alpha in the syndrome of cache xia , arthritis, and autoimmunity resulting from tristetraprolin (TTP) deficiency. Immunity, 4(5), 445–454. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/8630730 [DOI] [PubMed] [Google Scholar]
- Tcherkezian J, Cargnello M, Romeo Y, Huttlin EL, Lavoie G, Gygi SP, & Roux PP (2014). Proteomic analysis of cap-dependent translation identifies LARP1 as a key regulator of 5′TOP mRNA translation. Genes and Development, 28(4), 357–371. 10.1101/gad.231407.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thakur S, Nakamura T, Calin G, Russo A, Tamburrino JF, Shimizu M, … Croce CM (2003). Regulation of BRCA1 transcription by specific single -stranded DNA binding factors. Mol Cell Biol, 23(11), 3774–3787. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/12748281 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theobald DL, Mitton-Fry RM, & Wuttke DS (2003). Nucleic acid recognition by OB-fold proteins. Annual Review of Biophysics and Biomolecular Structure, 32, 115–133. 10.1146/annurev.biophys.32.110601.142506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoreen CC, Kang SA, Chang JW, Liu Q, Zhang J, Gao Y, … Gray NS (2009). An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. Journal of Biological Chemistry, 284(12), 8023–8032. 10.1074/jbc.M900301200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian B, White RJ, Xia T, Welle S, Turner DH, Mathews MB, & Thornton CA (2000). Expanded CUG repeat RNAs form hairpins that activate the double-stranded RNA-dependent protein kinase PKR. RNA, 6(1), 79–87. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/10668800 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian X, Gu T, Patel S, Bode AM, Lee M-H, & Dong Z (2019). CRISPR/Cas9 – An evolving biological tool kit for cancer biology and oncology. Npj Precision Oncology, 3(1). 10.1038/s41698-019-0080-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong L, Luo Y, Wei T, Guo L, Wang H, Zhu W, & Zhang J (2016). KH -type splicing regulatory protein (KHSRP) contributes to tumorigenesis by promoting miR −26a maturation in small cell lung cancer. Molecular and Cellular Biochemistry, 422(1–2), 61–74. 10.1007/s11010-016-2806-y [DOI] [PubMed] [Google Scholar]
- Topalian SL, Drake CG, & Pardoll DM (2015). Immune checkpoint blockade: a common denominator approach to cancer t herapy. Cancer Cell, 27(4), 450–461. 10.1016/j.ccell.2015.03.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Topisirovic I, Ruiz-Gutierrez M, & Borden KL (2004). Phosphorylation of the eukaryotic translation initiat ion factor eIF4E contributes to its transformation and mRNA transport activities. Cancer Res, 64(23), 8639–8642. 10.1158/0008-5472.CAN-04-2677 [DOI] [PubMed] [Google Scholar]
- Trabucchi M, Briata P, Garcia -Mayoral M, Haase AD, Filipowicz W, Ramos A, … Rosenfeld MG (2009). The RNA -binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature, 459(7249), 1010–1014. 10.1038/nature08025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trojanowicz B, Brodauf L, Sekulla C, Lorenz K, Finke R, Dralle H, & Hoang-Vu C (2009). The role of AUF1 in thyroid carcinoma progression. Endocrine-Related Cancer, 16(3), 857–871. 10.1677/ERC-08-0234 [DOI] [PubMed] [Google Scholar]
- Trotta R, Vignudelli T, Candini O, Intine RV, Pecorari L, Gue rzoni C, … Calabretta B (2003). BCR/ABL activate s mdm2 mRNA translation via the La antigen. Cancer Cell, 3(2), 145–160. 10.1016/S1535-6108(03)00020-5 [DOI] [PubMed] [Google Scholar]
- Truitt ML, Conn CS, Shi Z, Pang X, Tokuyasu T, Coady AM, … Ruggero D (2015). Differential Requirements for eIF4E Dose in Normal Development and Cancer. Cell, 162(1), 59–71. 10.1016/j.cell.2015.05.049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsumuraya T, Ishikawa C, Machijima Y, Nakachi S, Senba M, Tanaka J, & Mori N (2011). Effects of hippuristanol, an inhibitor of eIF4A, on adult T-cell leukemia. Biochem Pharmacol, 81(6), 713–722. 10.1016/j.bcp.2010.12.025 [DOI] [PubMed] [Google Scholar]
- Tu HC, Schwitalla S, Qian Z, La Pier GS, Yermalovich A, Ku YC, … Daley GQ (2015). LIN28 cooperates with WNT s ignaling to drive invasive intestinal and colorectal adenocarcinoma in mice and humans. Genes and Development, 29(10), 1074–1086. 10.1101/gad.256693.114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ueda J, Matsuda Y, Ya mahatsu K, Uchida E, Naito Z, Korc M, & Ishiwata T (2014). Epithelial splicing regulatory protein 1 is a favorable prognostic factor in pancreatic cancer that attenuates pancreatic metastases. Oncogene, 33(36), 4485–4495. 10.1038/onc.2013.392 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uehara T, Minoshima Y, Sagane K, Sugi NH, Mitsuhashi KO, Ya mamoto N, … Owa T (2017). Selective degradation of splicing factor CAPERα by anticancer sulfonamides. Nature Chemical Biology, 13(6), 675–680. 10.1038/nchembio.2363 [DOI] [PubMed] [Google Scholar]
- Urbach A, Yermalovich A, Zhang J, Spina CS, Zhu H, Perez-Atayde AR, … Daley GQ (2014). Lin 28 sustains early renal progenitors and induces Wilms tumor. Genes and Development, 28(9), 971–982. 10.1101/gad.237149.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ushigome M, Ubagai T, Fukuda H, Tsuchiya N, Sugimura T, Takatsuka J, & Nakagama H (2005). Up -regulation of hn RNP A1 gene in sporadic human colorectal cancers. International Journal of Oncology, 26(3), 635–640. [PubMed] [Google Scholar]
- Valverde R, Edwards L, & Regan L (2008). Structure and function of KH domains. FEBS J, 275(11), 2712–2726. 10.1111/j.1742-4658.2008.06411.x [DOI] [PubMed] [Google Scholar]
- van den Hoogenhof MMG, van der Made I, Beqqali A, de Groot NE, Damanafshan A, van Oort RJ, … Creemers EE (2017). The RNA -binding protein Rbm38 is dispensable during pressure overload-induced cardiac remodeling in mice. PLoS One, 12(8), e0184093 10.1371/journal.pone.0184093 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Voss MRH, Vesuna F, Trumpi K, Brilliant J, Berlinicke C, de Leng W, … Raman V (2015). Identification of the DEAD box RNA helicase DDX3 as a therapeutic target in colorectal cancer. Oncotarget, 6(29). 10.18632/oncotarget.4873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanharanta S, Marney CB, Shu W, Valiente M, Zou Y, Mele A, … Massagué J (2014). Loss of the multifunctional R NA-binding protein RBM47 as a source of selectable metastatic traits in breast cancer. ELife, 3 10.7554/elife.02734 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varnum BC, Lim RW, Sukhatme VP, & Herschman HR (1989). Nucleotide sequence of a cDNA encoding TIS11, a messag e induced in Swiss 3T3 cells by the tumor promoter tetradecanoyl phorbol acetate. Oncogene, 4(1), 119–120. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/2915901 [PubMed] [Google Scholar]
- Vázquez-Chantada M, Fernández-Ramos D, Embade N, Martínez-Lopez N, Varela-Rey M, Woodhoo A, … Martínez-Chantar ML (2010). HuR/Methyl-HuR and AUF1 Regulate the MAT Expressed During Liver Proliferation, Differentiation, and Carcinogenesis. Gastroenterology, 138(5). 10.1053/j.gastro.2010.01.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venables JP, Brosseau J-P, Gadea G, Klinck R, Prinos P, Beaulieu J-F, … Chabot B (2012). RBFOX2 Is an Important Regulator of Mesenchymal Tissue-Specific Splicing in both Normal and Cancer Tissues. Molecular and Cellular Biology, 33(2), 396–405. 10.1128/mcb.01174-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venables JP, Klinck R, Koh C, Gervais-Bird J, Bramard A, Inkel L, … Elela SA (2009). Cancer-associated regulation of alternative splicing. Nature Structural and Molecular Biology, 16(6), 670–676. 10.1038/nsmb.1608 [DOI] [PubMed] [Google Scholar]
- Viswanathan SR, Powers JT, Einhorn W, Hoshida Y, Ng TL, Toffanin S, … Daley GQ ( 2009). Lin28 promotes transformation and is associated with advanced human malignancies. Nature Genetics, 41(7), 843–848. 10.1038/ng.392 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vo DT, Abdelmohsen K, Martindale JL, Qiao M, Tominaga K, Burton TL, … Penalva LOF (2012). The Oncogenic RNA-Binding Protein Musashi1 Is Regulated by HuR via mRNA Translation and Stability in Glioblastoma Cells. Molecular Cancer Research, 10(1), 143–155. 10.1158/1541-7786.mcr-11-0208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vo DT, Subramaniam D, Remke M, Burton TL, Uren PJ, Gelfond JA, … Penalva LOF (2012). The RNA-binding protein musashi1 affects medulloblastoma growth via a network of cancer-related genes and is an indicator of poor prognosis. American Journal of Pathology, 181(5), 1762–1772. 10.1016/j.ajpath.2012.07.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wampfler J, Federzoni EA, Torbett BE, Fey MF, & Tschan MP (2016). The RNA binding proteins RBM 38 and DND1 a re repressed in AML and have a novel function in APL differentiation. Leuk Res, 41, 96–102. 10.1016/j.leukres.2015.12.006 [DOI] [PubMed] [Google Scholar]
- Wang A, Bao Y, Wu Z, Zhao T, Wang D, Shi J, … Qu L (2019). Long noncoding RNA EGFR -AS1 promotes cell growth and metastasis via affecting HuR mediated mRNA stability of EGFR in renal cancer. Cell Death and Disease, 10(3). 10.1038/s41419-019-1331-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang E, Lu SX, Pastore A, Chen X, Imig J, Chun-Wei Lee S, … Aifantis I (2019). Targeting an RNA-Binding Protein Network in Acute Myeloid Leukemia. Cancer Cell, 35(3), 369–384.e7 10.1016/j.ccell.2019.01.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H, Vardy LA, Tan CP, Loo JM, Guo K, Li J, … Zeng Q (2010). PCBP1 suppresses the translation of metas tasis-associated PRL-3 phosphatase. Cancer Cell, 18(1), 52–62. 10.1016/j.ccr.2010.04.028 [DOI] [PubMed] [Google Scholar]
- Wang J, Guo Y, Chu H, Guan Y, Bi J, & Wang B (2013). Multiple functions of the RNA-binding protein HuR in cancer progression, treatment responses and prognosis. International Journal of Molecular Sciences 10.3390/ijms140510015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L, Lawrence MS, Wan Y, Stojanov P, Sougnez C, Stevenson K, … Wu CJ (2011). SF3B1 and Other Novel Cancer Genes in Chronic Lymp hocytic Leukemia . New England Journal of Medicine, 365(26), 2497–2506. 10.1056/nejmoa1109016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L, Rowe RG, Jaimes A, Yu C, Nam Y, Pearson DS, … Sliz P (2018). Small-Molecule Inhibitors Disrupt let-7 Oligouridylation and Release the Selective Blockade of let-7 Processing by LIN28. Cell Rep, 23(10), 3091–3101. 10.1016/j.celrep.2018.04.116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang R, Jiao Z, Li R, Yue H, & Chen L (2012). p68 RNA helicase promotes glioma cell proliferation in vitro and in v ivo via direct regulation of NF-κB transcription factor p50. Neuro-Oncology, 14(9), 1116–1124. 10.1093/neuonc/nos131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang SJ, Zhang C, You Y, & Shi CM (2012). Overe xpression of RNA helicase p68 protein in cutaneous squamous cell c arcinoma. Clinical and Experimental Dermatology, 37(8), 882–888. 10.1111/j.1365-2230.2012.04365.x [DOI] [PubMed] [Google Scholar]
- Wang S, Rosenwald IB, Hutzler MJ, Pihan GA, Savas L, Chen JJ, & Woda BA (1999). Expression of the eukaryotic translation in itiation factors 4E and 2α in non-Hodgkin’s lymphomas. American Journal of Pathology 10.1016/S0002-9440(10)65118-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W, Furneaux H, Cheng H, Caldwell MC, Hutter D, Liu Y, … Gorospe M (2000). Hu R Regulates p21 mRNA Stabili zation by UV Light. Molecular and Cellular Biology, 20(3), 760–769. 10.1128/mcb.20.3.760-769.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang XL, Cai HP, Ge JH, & Su XF (2012). Detection of eukaryotic translation initiation factor 4E and its clin ical significance in hepatocellular carcinoma. World Journal of Gastroenterology, 18(20), 2540–2544. 10.3748/wjg.v18.i20.2540 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, & Tanaka Hall TM (2001). Structural basis for recognition of AU-rich element RNA by the HuD protein. Nat Struct Biol, 8(2), 141–145. 10.1038/84131 [DOI] [PubMed] [Google Scholar]
- Wang XY, Penalva LOF, Yuan H, Linnoila RI, Lu J, Okano H, & Glazer RI (2010). Musashi1 regulates breast tumor cell proliferation and is a prognostic indicator of poor survival. Molecular Cancer, 9 10.1186/1476-4598-9-221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Chen D, Qian H, Tsai YS, Shao S, Liu Q, … Wang Z (2014). The Splicing Factor RBM 4 Controls Apoptosis, Proliferation, and Migration to Suppress Tumor Progression. Cancer Cell, 26(3), 374–389. 10.1016/j.ccr.2014.07.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang ZL, Li B, Luo YX, Lin Q, Liu SR, Zhang XQ, … Qu LH (2018). Comprehensive Geno mic Characterization of RNA-Binding Proteins across Human Cancers. Cell Rep, 22(1), 286–298. 10.1016/j.celrep.2017.12.035 [DOI] [PubMed] [Google Scholar]
- Wang Z, Yin W, Zhu L, Li J, Yao Y, Chen F, … Chang X (2018). Iron Drives T Helper Cell Pathogenicity by Promoting RNA-Binding Protein PCBP1-Mediated Proinflammatory Cytokine Production. Immunity, 49(1), 80–92 e7 10.1016/j.immuni.2018.05.008 [DOI] [PubMed] [Google Scholar]
- Warzecha CC, Jiang P, Amirikian K, Dittmar KA, Lu H, Shen S, … Carstens RP (2010). An ESRP -regulated splicing programme is abrogated during the epithelial-mesenchymal transition. EMBO Journal, 29(19), 3286–3300. 10.1038/emboj.2010.195 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warzecha CC, Sato TK, Nabet B, Hogenesch JB, & Carstens RP (2009). ESRP1 and ESRP2 Are Epithelial Cell -Type-Specific Regulators of FGFR2 Splicing. Molecular Cell, 33(5), 591–601. 10.1016/j.molcel.2009.01.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei ZR, Liang C, Feng D, Cheng YJ, Wang WM, Yang DJ, … Cai QP (2016). Low tristetraprolin e xpression promotes cell proliferation and predicts poor patients outcome in pancreatic cancer. Oncotarget, 7(14), 17737–17750. 10.18632/oncotarget.7397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen F, Shen A, Shanas R, Bhattacharyya A, Lian F, Hostetter G, & Shi J (2010). Higher expression of the heterog eneous nuclear ribonucleoproteink in melanoma. Ann Surg Oncol, 17(10), 2619–2627. 10.1245/s10434-010-1121-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wendel HG, Silva RL, Malina A, Mills JR, Zhu H, Ueda T, … Lowe SW (2007). Dissecting eIF4E action in tumorigenesis. Genes Dev, 21(24), 3232–3237. 10.1101/gad.1604407 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westman B, Beeren L, Grudzien E, Stepinski J, Worch R, Zuberek J, … Preiss T (2005). The antiviral drug ribavirin does not mi mic the 7-methylguanosine moiety of the mRNA cap structure in vitro. RNA, 11(10), 1505–1513. 10.1261/rna.2132505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilky BA, Kim C, Mc Carty G, Montgomery EA, Kammers K, Devine LR, … Loeb DM (2016). RNA helicase DDX3: A novel therapeutic target in Ewing sarcoma. Oncogene, 35(20), 2574–2583. 10.1038/onc.2015.336 [DOI] [PubMed] [Google Scholar]
- Witkiewicz AK, McMillan EA, Balaji U, Baek GH, Lin WC, Mansour J, … Knudsen ES (2015). Whole-e xo me sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nature Communications, 6 10.1038/ncomms7744 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfe AL, Singh K, Zhong Y, Drewe P, Rajasekhar VK, Sanghvi VR, … Wendel HG (2014). RNA G-quadruplexes cause eIF4A-dependent oncogene translation in cancer. Nature, 513(7516), 65–70. 10.1038/nature13485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wortham NC, Ahamed E, Nicol SM, Thomas RS, Periyasamy M, Jiang J, … Fuller-Pace FV (2009). The DEAD-box protein p72 regulates ERα-/oestrogen-dependent transcription and cell growth, and is associated with improved survival in ERα-positive breast cancer. Oncogene, 28(46), 4053–4064. 10.1038/onc.2009.261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu CS, Chang KP, Chen LC, Chen CC, Liang Y, Hseuh C, & Chang YS (2012). Heterogeneous ribonucleoprote in K and thymidine phosphorylase are independent prognostic and therapeutic markers for oral squamous cell carcinoma. Oral Oncol, 48(6), 516–522. 10.1016/j.oraloncology.2012.01.005 [DOI] [PubMed] [Google Scholar]
- Wu X, Lan L, Wilson DM, Marquez RT, Tsao WC, Gao P, … Xu L (2015). Identification and Validation of Novel Small Molecule Disruptors of HuR-mRNA Interaction. ACS Chemical Biology, 10(6), 1476–1484. 10.1021/cb500851u [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu X, Yang Y, Huang Y, Chen Y, Wang T, Wu S, … Zhao W (2018). RNA -binding protein AUF1 suppresses miR-122 biogenesis by down-regulating Dicer1 in hepatocellular carcinoma. Oncotarget 10.18632/oncotarget.24079 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xie M, Vesuna F, Tantravedi S, Bol GM, Van Voss MRH, Nugent K, … Raman V (2016). RK −33 radiosensitizes prostate cancer cells by blocking the RNA helicase DDX3. Cancer Research, 76(21), 6340–6350. 10.1158/0008-5472.CAN-16-0440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu E, Zhang J, & Chen X (2013). MDM 2 e xp ression is repressed by the RNA -binding protein RNPC1 via mRNA stability. Oncogene, 32(17), 2169–2178. 10.1038/onc.2012.238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu E, Zhang J, Zhang M, Jiang Y, Cho SJ, & Chen X (2014). RNA-binding protein RBM 24 regulates p63 e xpression via mRNA stability. Mol Cancer Res, 12(3), 359–369. 10.1158/1541-7786.MCR-13-0526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu M, Fang S, Song J, Chen M, Zhang Q, Weng Q, … Ji J (2018). CPEB1 mediates hepatocellular carcinoma cancer st emness and chemoresistance. Cell Death and Disease, 9(10). 10.1038/s41419-018-0974-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Y, Gao XD, Lee JH, Huang H, Tan H, Ahn J, … Cheng C (2014). Cell type-restricted activity of hnRNPM promotes breast cancer metastasis via regulating alternative splicing. Genes and Development, 28(11), 1191–1203. 10.1101/gad.241968.114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Y, Poggio M, Jin HY, Shi Z, Forester CM, Wang Y, … Ruggero D (2019). Translation control of the immune checkpoint in cancer and its therapeutic targeting. Nature Medicine, 25(2), 301–311. 10.1038/s41591-018-0321-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xue JQ, Xia TS, Liang XQ, Zhou W, Cheng L, Shi L, … Ding Q (2014). RNA-binding protein RNPC1: acting as a tumor suppressor in breast cancer. BMC Cancer, 14, 322 10.1186/1471-2407-14-322 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xue Q, Hopkins B, Perruzzi C, Udayakumar D, Sherris D, & Benjamin LE (2008). Palomid 529, a novel small-molecule drug, is a TORC1/TORC2 inhibitor that reduces tumor growth, tumor angiogenesis, and vascular permeability. Cancer Research, 68(22), 9551–9557. 10.1158/0008-5472.CAN-08-2058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yae T, Tsuchihashi K, Ishimoto T, Motohara T, Yoshikawa M, Yoshida GJ, … Nagano O (2012). Alternative splicing of CD44 mRNA by ESRP1 enhances lung colonization of metastatic cancer cell. Nature Communications, 3 10.1038/ncomms1892 [DOI] [PubMed] [Google Scholar]
- Yan KS, Yan S, Farooq A, Han A, Zeng L, & Zhou MM (2003). Structure and conserved RNA binding of the PAZ doma in. Nature, 426(6965), 468–474. 10.1038/nature02129 [DOI] [PubMed] [Google Scholar]
- Yan LX, Wu QN, Zhang Y, Li YY, Liao DZ, Hou JH, … Shao JY (2011). Knockdown of miR-21 in human breast cancer cell lines inhibits proliferation, in vitro migration and in vivo tumor growth. Breast Cancer Research, 13(1). 10.1186/bcr2803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan W, Scoumanne A, Jung YS, Xu E, Zhang J, Zhang Y, … Chen X (2016). Mice deficient in poly(C)-binding protein 4 are susceptible to spontaneous tumors through increased expression of ZFP871 that targets p53 for degradation. Genes Dev, 30(5), 522–534. 10.1101/gad.271890.115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan Y, Svitkin Y, Lee JM, Bisaillon M, & Pelletier J (2005). Ribavirin is not a functional mimic of the 7 -methyl guanosine mRNA cap. RNA, 11(8), 1238–1244. 10.1261/rna.2930805 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang C, Zhang W, Wang L, Kazobinka G, Han X, Li B, & Hou T (2016). Musashi-2 promotes migration and invasion in bladder cancer via activation of the JAK2/STAT3 pathway. Laboratory Investigation, 96(9), 950–958. 10.1038/labinvest.2016.71 [DOI] [PubMed] [Google Scholar]
- Yang J, Fan J, Li Y, Li F, Chen P, Fan Y, … Wong ST (2013). Geno me -Wide RNAi Screening Identifies Genes Inhibiting the Migration of Glioblastoma Cells. PLoS ONE , 8(4). 10.1371/journal.pone.0061915 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J, Hung LH, Licht T, Kostin S, Looso M, Khrameeva E, … Braun T (2014). RBM 24 is a major regulator of mus cle-specific alternative splicing. Dev Cell, 31(1), 87–99. 10.1016/j.devcel.2014.08.025 [DOI] [PubMed] [Google Scholar]
- Yang L, Zhang Y, Ling C, & Heng W (2018). RNPC1 inhibits non-small cell lung cancer progression via regulating miR-181a/ CASC2 axis. Biotechnology Letters, 40(3), 543–550. 10.1007/s10529-017-2504-1 [DOI] [PubMed] [Google Scholar]
- Yang R, Zeng Y, Xu H, Chen Z, Xiang M, Fu Y, … Zeng X (2016). Heterogeneous nuclear ribonucleoprotein K is overe xpressed and ass ociated with poor prognosis in gastric cancer. Oncol Rep, 36(2), 929–935. 10.3892/or.2016.4845 [DOI] [PubMed] [Google Scholar]
- Yang W, & Steitz TA (1995). Recombining the structures of HIV integrase, RuvC and RNase H. Structure, 3(2), 131–134. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/7735828 [DOI] [PubMed] [Google Scholar]
- Yang X, Lin X, Zhong X, Kaur S, Li N, Liang S, … Zhang L (2010). Double-negative feedback loop between reprogramming factor LIN28 and micro RNA let-7 regulates aldehyde dehydrogenase 1-positive cancer stem cells. Cancer Research, 70(22), 9463–9472. 10.1158/0008-5472.CAN-10-2388 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X, Wang W, Fan J, Lal A, Yang D, Cheng H, & Gorospe M (2004). Prostaglandin A2-mediated stabilization of p21 mRNA through an ERK-dependent pathway requiring the RNA-binding protein HuR. Journal of Biological Chemistry, 279(47), 49298–49306. 10.1074/jbc.M407535200 [DOI] [PubMed] [Google Scholar]
- Yao J, Caballero OL, Huang Y, Lin C, Rimoldi D, Behren A, … Zhao Q (2016). Altered Expression and Splicing of ESRP1 in Malignant Melanoma Correlates with Epithelial-Mesenchymal Status and Tumor-Associated Immune Cytolytic Activity. Cancer Immunology Research, 4(6), 552–561. 10.1158/2326-6066.cir-15-0255 [DOI] [PubMed] [Google Scholar]
- Ye J, Liang R, Bai T, Lin Y, Mai R, Wei M, … Wu F (2018). RBM 38 plays a tumor -suppressor role via stabilizing the p53-mdm2 loop function in hepatocellular carcinoma. Journal of Experimental and Clinical Cancer Research , 37(1). 10.1186/s13046-018-0852-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye J, Zhou G, Zhang Z, Sun L, He X, & Zhou J (2016). Poly (C) -binding protein 2 (PCBP2) promotes the progression of esophageal squamous cell carcinoma (ESCC) through regulating cellular proliferation and apoptosis. Pathol Res Pract, 212(8), 717–725. 10.1016/j.prp.2016.05.008 [DOI] [PubMed] [Google Scholar]
- Yin T, Cho SJ, & Chen X (2013). RNPC1, an RNA-binding protein and a p53 target, regulates macrophage inhibitory cytokine-1 (MIC-1) expression through mRNA stability. J Biol Chem, 288(33), 23680–23686. 10.1074/jbc.M113.480186 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yin W, & Rogge M (2019). Targeting RNA: A Transformative Therapeutic Strategy. Clinical and Translational Science 10.1111/cts.12624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yong H, Zhu H, Zhang S, Zhao W, Wang W, Chen C, … Liu B (2016). Prognostic value of decreased e xpression of RBM 4 in human gastric cancer. Scientific Reports, 6 10.1038/srep28222 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoshiza wa A, Fukuoka J, Shimizu S, Shilo K, Franks TJ, Hewitt SM, … Travis WD (2010). Overexpression of phospho-eIF4E is associated with survival through AKT pathway in non-small cell lung cancer. Clinical Cancer Research, 16(1), 240–248. 10.1158/1078-0432.CCR-09-0986 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu K, Shi C, Toral-Barza L, Lucas J, Shor B, Kim JE, … Gibbons JJ (2010). Beyond rapalog therapy: preclinical pharmacology and ant itumor activity of W YE-125132, an ATP-competitive and specific inhibitor of mTORC1 and mTORC2. Cancer Research, 70(2), 621–631. 10.1158/0008-5472.CAN-09-2340 [DOI] [PubMed] [Google Scholar]
- Yuan YR, Pei Y, Ma JB, Kuryavyi V, Zhadina M, Meister G, … Patel DJ (2005). Crystal structure of A. aeolicus argonaute, a site-specific DNA-guided endoribonuclease, provides insights into RISC-mediated mRNA cleavage. Mol Cell, 19(3), 405–419. 10.1016/j.molcel.2005.07.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zanocco-Marani T (2010). TIS11/TTP gene family: it’s never too late for tumor suppressors. Cell Cycle, 9(24), 4771 10.4161/cc.9.24.14217 [DOI] [PubMed] [Google Scholar]
- Zeng Q, Hong W, & Tan YH (1998). Mouse PRL-2 and PRL-3, two potentially prenylated protein tyrosine phosphatases homologous to PRL-1. Biochem Biophys Res Commun, 244(2), 421–427. 10.1006/bbrc.1998.8291 [DOI] [PubMed] [Google Scholar]
- Zhang FL, Cao JL, Xie HY, Sun R, Yang LF, Shao ZM, & Li DQ (2018). Cancer-associated MORC2-mutant M276I regulates an hnRNPM-mediated CD44 splicing switch to promote invasion and metastasis in triple-negative breast cancer. Cancer Research, 78(20), 5780–5792. 10.1158/0008-5472.CAN-17-1394 [DOI] [PubMed] [Google Scholar]
- Zhang H, Brown RL, Wei Y, Zhao P, Liu S, Liu X, … Cheng C (2019). CD44 splice isoform switching determines breast cancer stem cell state. Genes and Development, 33(3–4), 166–179. 10.1101/gad.319889.118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, & Chen X (2008). Posttranscriptional Regulation of p53 and its Targets by RNABinding Proteins. Current Molecular Medicine, 8(8), 845–849. 10.2174/156652408786733748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Cho SJ, Shu L, Yan W, Gue rrero T, Kent M, … Chen X (2011). Translational repression of p53 by RNPC1, a p53 target overexp ressed in lymphomas. Genes Dev, 25(14), 1528–1543. 10.1101/gad.2069311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Jun Cho S, & Chen X (2010). RNPC1, an RNA-binding protein and a target of the p53 family, regulates p63 expression through mRNA stability. Proc Natl Acad Sci U S A, 107(21), 9614–9619. 10.1073/pnas.0912594107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Xu E, Ren C, Yan W, Zhang M, Chen M, … Chen X (2014). Mice deficient in Rbm38, a target of the p53 fam ily, are susceptible to accelerated aging and spontaneous tumors. Proc Natl Acad Sci U S A, 111(52), 18637–18642. 10.1073/pnas.1415607112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Xu E, Ren C, Yang HJ, Zhang Y, Sun W, … Chen X (2018). Genetic Ablation of Rbm38 Promotes Lymphomage nesis in the Context of Mutant p53 by Downregulating PTEN. Cancer Res, 78(6), 1511–1521. 10.1158/0008-5472.CAN-17-2457 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang M, Zhang Y, Xu E, Mohibi S, De Anda DM, Jiang Y, … Chen X (2018). Rbm24, a target of p53, is necessary for proper expression of p53 and heart development. Cell Death and Differentiation, 25(6). 10.1038/s41418-017-0029-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang T, Huang XH, Dong L, Hu D, Ge C, Zhan YQ, … Yang XM (2010). PCBP-1 regulates alternative splicing of the CD44 gene and inhibits invasion in human hepatoma cell line HepG2 cells. Mol Cancer, 9, 72 10.1186/1476-4598-9-72 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Hua L, Yan D, Zhao F, Liu J, Zhou H, … Hu B (2016). Over expression of PCBP2 contributes to poor prognosis and enhanced cell growth in human hepatocellular carcinoma. Oncol Rep, 36(6), 3456–3464. 10.3892/or.2016.5167 [DOI] [PubMed] [Google Scholar]
- Zhang X, Wan G, Berger FG, He X, & Lu X (2011). The ATM Kinase Induces MicroRNA Biogenesis in the DNA Damage Resp onse. Molecular Cell, 41(4), 371–383. 10.1016/j.molcel.2011.01.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Meng L, Xiao L, Liu R, Li Z, & Wang YL (2018). The RNA-Binding Protein PCBP1 Functions as a Tumor Suppressor in Prostate Cancer by Inhibiting Mitogen Activated Protein Kinase 1. Cell Physiol Biochem, 48(4), 1747–1754. 10.1159/000492315 [DOI] [PubMed] [Google Scholar]
- Zhang Y, Peng L, Hu T, Wan Y, Ren Y, Zhang J, … Zhu X (2015). La-related protein 4B maintains murine MLL-AF9 leukemia stem cell self-renewal by regulating cell cycle progression. Experimental Hematology, 43(4), 309–318.e2 10.1016/j.exphem.2014.12.003 [DOI] [PubMed] [Google Scholar]
- Zhao J, Sun Y, Huang Y, Song F, Huang Z, Bao Y, … Wang Y (2017). Functional analysis reveals that RBM10 mutatio ns contribute to lung adenocarcinoma pathogenesis by deregulating splicing. Scientific Reports, 7 10.1038/srep40488 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Q, Caballero OL, Davis ID, Jonasch E, Tamboli P, Yung WKA, … Yao J (2013). Tumor-specific isoform switch of the fib roblast growth factor receptor 2 underlies the mesenchymal and malignant phenotypes of clear cell renal cell carcinomas. Clinical Cancer Research, 19(9), 2460–2472. 10.1158/1078-0432.CCR-12-3708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng X, & Bevilacqua PC (2004). Activation of the protein kinase PKR by short double -stranded RNAs with single-stranded tails. Rna, 10(12), 1934–1945. 10.1261/rna.7150804 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J, & Rossi J (2017). Aptamers as targeted therapeutics: Current potential and challenges. Nature Reviews Drug Discovery 10.1038/nrd.2016.199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou M, & Tong X (2015). Downregulated Poly-C binding protein-1 is a novel predictor associated with poor prognosis in Acute Myeloid Leukemia. Diagn Pathol, 10, 147 10.1186/s13000-015-0377-y [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Zhou ZJ, Dai Z, Zhou SL, Fu XT, Zhao YM, Shi YH, … Fan J (2013). Overexpression of HnRNP A1 promotes tumor invasion through regulating CD 44v6 and indicates poor prognosis for hepatocellular carcinoma. International Journal of Cancer, 132(5), 1080–1089. 10.1002/ijc.27742 [DOI] [PubMed] [Google Scholar]
- Zhu H, Berkova Z, Mathur R, Sehgal L, Khashab T, Tao R-H, … Samaniego F (2015). HuR Suppresses Fas Expre ssion and Correlates with Patient Outcome in Liver Cancer. Molecular Cancer Research, 13(5), 809–818. 10.1158/1541-7786.mcr-14-0241 [DOI] [PubMed] [Google Scholar]
- Zhu J, & Chen X (2000). MCG10, a novel p53 target gene that encodes a KH domain RNA -binding protein, is capable of inducing apoptosis and cell cycle arrest in G(2)-M. Mol Cell Biol, 20(15), 5602–5618. Retrieved from https://www.ncbi.nlm.nih.gov/pubmed/10891498 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zindy PJ, L’Helgoualc’h A, Bonnier D, Le Bechec A, Bourd-Boitin K, Zhang CX, … Theret N (2006). Upregulation of the tumor suppressor gene menin in hepatocellular carcinomas and its significance in fibrogenesis. Hepatology, 44(5), 1296–1307. 10.1002/hep.21367 [DOI] [PubMed] [Google Scholar]
- Zong Z, Zhou T, Rao L, Jiang Z, Li Y, Hou Z, … Chen S (2016). Musashi2 as a novel predictive biomarker for liver metastasis and poor prognosis in colorectal cancer. Cancer Medicine, 5(4), 623–630. 10.1002/cam4.624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou T, Rao JN, Liu L, Xiao L, Yu T-X, Jiang P, … Wang J-Y (2010). Polyamines Regulate the Stability of JunD mRNA by Modulating the Competitive Binding of Its 3’ Untranslated Region to HuR and AUF1. Molecular and Cellular Biology 10.1128/mcb.00807-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zucconi BE (2011). Modulation of neoplastic gene regulatory pathways by the RNA -binding factor AUF1. Frontiers in Bioscience, 16(1), 2307 10.2741/3855 [DOI] [PMC free article] [PubMed] [Google Scholar]