Abstract
Due to the positive association between neoadjuvant chemotherapy (NACT) and the promising early response rates of patients with triple negative breast cancer (TNBC), including probabilities of pathological complete response, NACT is increasingly used in TNBC management. Liquid biopsy-based biomarkers with the power to diagnose the early response to NACT may support established monitoring tools, which are to a certain extent imprecise and costly. Simple serum- or urine-based analyses of non-coding RNA (ncRNA) expression may allow for fast, minimally-invasive testing and timely adjustment of the therapy regimen. The present study investigated breast cancer-related ncRNAs [microRNA (miR)-7, -9, -15a, -17, -18a, -19b, -21, -30b, -222 and -320c, PIWI-interacting RNA-36743 and GlyCCC2] in triple positive BT-474 cells and three TNBC cell lines (BT-20, HS-578T and MDA-MB-231) treated with various chemotherapeutic agents using reverse transcription-quantitative PCR. Intracellular and secreted microvesicular ncRNA expression levels were analysed using a multivariable statistical regression analysis. Chemotherapy-driven effects were investigated by analysing cell cycle determinants at the mRNA and protein levels. Serum and urine specimens from 8 patients with TNBC were compared with 10 healthy females using two-sample t-tests. Samples from the patients with TNBC were compared at two time points. Chemotherapeutic treatments induced distinct changes in ncRNA expression in TNBC cell lines and the BT-474 cell line in intra- and extracellular compartments. Serum and urine-based ncRNA expression analysis was able to discriminate between patients with TNBC and controls. Time point comparisons in the urine samples of patients with TNBC revealed a general rise in the level of ncRNA. Serum data suggested a potential association between piR-36743, miR-17, -19b and -30b expression levels and an NACT-driven complete clinical response. The present study highlighted the potential of ncRNAs as liquid biopsy-based biomarkers in TNBC chemotherapy treatment. The ncRNAs tested in the present study have been previously investigated for their involvement in BC or TNBC chemotherapy responses; however, these previous studies were restricted to patient tissue or in vitro models. The data from the present study offer novel insight into ncRNA expression in liquid samples from patients with TNBC, and the study serves as an initial step in the evaluation of ncRNAs as diagnostic biomarkers in the monitoring of TNBC therapy.
Keywords: ncRNA, triple negative breast cancer, liquid biopsy, neoadjuvant chemotherapy, therapy response, biomarker
Introduction
As the most frequent malignancy occurring in females, breast cancer (BC) represents a great threat to women's health. In 2018, the World Health Organization registered 2,088,849 BC cases and 626,679 BC-associated deaths globally (1). However, the survival rate for patients with BC has been increasing since the 1990s. One reason for this is the improvement in systemic treatment concepts (2,3). Moreover, molecular subtyping has been implemented in clinical practice as an important tool for risk-adapted therapy in patients with BC (4-7).
Triple negative breast cancer (TNBC) is classified as the most aggressive molecular subtype of BC (8). TNBCs are estimated to comprise ~15% of all cases of BC (9,10), and its occurrence is highest in younger females (9) and those of African-American ancestry (8,11). The prognosis for women diagnosed with TNBC is less favourable than for those suffering from other intrinsic subtypes (12). Characterised by the absence of hormone receptors and a lack of Her2neu (ErbB2) gene amplification, TNBCs are heterogeneous carcinomas (9) and are often poorly differentiated. Several attempts have been made to further categorise TNBC subtypes according to distinct gene expression patterns. For example, Lehmann et al (13) analysed the gene expression profiles of 587 TNBC cases and identified six distinct TNBC subtypes: Two basal-like, an immunomodulatory, a mesenchymal, a mesenchymal stem-like and a luminal androgen receptor subtype.
TNBC can be further classified into the basal-like subtype (12) or the claudin-low subtype (4). The terms 'basal-like' BC and TNBC are often used interchangeably; however, they do not describe the same condition, as not all basal-like BCs are classified as TNBC and vice versa. Most basal-like BC cases present without hormone receptor expression and Her2neu gene amplification, as in TNBC (14); in a study performed by Bertucci et al (15), ~71% of TNBCs exhibited basal-like gene expression. Characteristic basal-like markers comprise cytokeratin 5/6 and epidermal growth factor receptor (14). The claudin-low intrinsic subtype, which was described in 2007, is characterised by the high expression of epithelial-mesenchymal transition markers and cancer stem cell-like features, among other traits (4). Unlike the basal-like subtype, the claudin-low intrinsic subtype exhibits lower expression of genes associated with proliferation, such as Ki67 (4). Prat et al (4) reported that the prognosis of the claudin-low subtype was poorer than that of luminal A, but improved compared with that of the basal-like subtype.
Due to the absence of hormone receptors and the lack of Her2neu gene amplification, therapy for patients with TNBC is restricted to neoadjuvant chemotherapy (NACT), radiotherapy and surgery (16). In the case of NACT, chemotherapy is administered prior to surgery, while in adjuvant chemotherapy (ACT), surgery precedes the chemotherapy treatment. NACT is considered to be equivalent to ACT in terms of clinical outcome, and has become an established treatment in BC therapy (17-21). Additionally, fewer adverse effects were observed with NACT compared with ACT (17). The primary endpoint of NACT is defined as the pathologically determined response of the tumour in the breast and the axillary lymph nodes. Achieving pathological complete response (pCR) at the time of surgery, defined as no residual invasive or non-invasive tumour tissue in the breast and the axillary lymph nodes, represents an important surrogate marker for beneficial overall survival in these patients (22-27). Further advantages of NACT treatment can be seen in the increasing rate of breast conserving surgery and the evaluation of short-term surrogate markers (pathological, clinical and molecular) for outcome prediction (21,27,28). In this context, it is known that an early tumour response (after 1-2 cycles of NACT) is also a predictive factor for pCR, and offers the opportunity to distinguish responders from non-responders at a very early time point of therapy (27). This early determination of poor response offers the chance to alter treatment regimens and drugs with the goal of achieving an optimized response up to a pCR. Therefore, reliable measures to determine early response to NACT represent an important prerequisite for innovative personalised treatment concepts for the improvement of the clinical outcome for patients with non-responding tumours. The NACT regime for patients with TNBC includes both an anthracycline and a taxane that should be administered for 18-24 weeks (28). The addition of platinum, irrespective of breast cancer susceptibility protein status, increases pCR rates; however, it is accompanied by enhanced drug-dependent toxicity in patients with TNBC (7). A variety of diagnostic possibilities, including ultrasound (US), palpation, mammography, magnetic resonance imaging (MRI) and positron emission tomography/computed tomography, are available for response monitoring during NACT (28). Although US measurements are used routinely in the clinic to assess tumour response to NACT, few data exist concerning the accuracy of US for the prediction of pCR (29,30). Morphological reactions of solid tumours during therapy (for example, fragmentation and tumour necrosis with or without stromal reaction) may compromise the accuracy of US in discriminating between responding and non-responding tumours. A previous study performed by Chagpar et al (31) investigated the accuracy of physical examination, US and mammography in predicting residual tumour size in NACT-treated patients with BC, and found only a moderate correlation. MRI appears to be an accurate method to detect residual disease; however, the false-negative rate is increased in patients receiving chemotherapy. Furthermore, these methods depend on tumour size reduction, which can, in certain cases, take months (32).
As an early response to NACT seems to be associated with achieving pCR and an improved prognosis for patients with TNBC, prediction of therapy response is particularly important in this BC subgroup (24,33-35). A poor prognosis for patients with TNBC improves to a similarly high level if patients achieve a pCR (34). This underlines the need for more specific and sensitive predictive methods. Liquid biopsy-based non-coding RNAs (ncRNAs) in the serum and urine of patients undergoing NACT as innovative biomarkers may be a promising tool to predict early therapy response before the current standard imaging methods are employed.
ncRNAs comprise several classes, according to size, origin and function (36,37). The evidence for the involvement of ncRNAs in carcinogenesis is increasing (38). As ncRNAs are secreted by their cells of origin, they can be detected in body fluids, including serum and urine (39-41). Several previous studies have reported an association between ncRNA expression, chemotherapy response and resistance (42-45). Therefore, ncRNAs may serve as liquid-biopsy-based biomarkers to indicate early response to NACT in TNBC.
To pursue this question, the present study examined the expression of 14 ncRNAs, including 12 microRNAs (miRNA/miRs; let-7a; let-7e; miR-7, -9, -15a, -17, -18a, -19b, -21, -30b, -222 and -320c), one PIWI-interacting RNA (piRNA; piR-36743) and one transfer RNA (GlyCCC2) in a triple positive (BT-474) and three TNBC (BT-20; HS-578T; MDA-MB-231) cell lines treated with the cytostatic drugs carboplatin, epirubicin, gemcitabine and paclitaxel, both intra- and extracellularly. Analysis of cell cycle determinants [cyclin D1, DNA topoisomerase 2α (TOP2α) and targeting protein for Xklp2 (TPX2)] and RNA metabolic proteins [DEAD-box polypeptide (DDX)5 and DDX17] were performed at the mRNA and protein level to investigate drug-dependent effects on the cell cycle. Subsequently, ncRNA expression was analysed in serum and urine specimens from patients with TNBC and healthy controls. Additionally, alterations in ncRNA expression in patients with TNBC prior to NACT (t0) and immediately prior to the third cycle of therapy (t1) were investigated.
Materials and methods
Cell culture conditions and treatments
BC cell lines BT-474 and BT-20 (cat. nos. 300130 and 300131; CLS Cell Lines Service GmbH), HS-578T (cat. no. 86082104; Sigma-Aldrich; Merck KGaA) and MDA-MB-231 (cat. no. 92020424; Sigma-Aldrich; Merck KGaA) (46) were incubated in DMEM/F12 medium (Gibco; Thermo Fisher Scientific, Inc.), supplemented with 5% FBS (Gibco; Thermo Fisher Scientific, Inc.), 1% HEPES buffer (Gibco; Thermo Fisher Scientific, Inc.) and 1% 100 U/ml penicillin/streptomycin (Sigma-Aldrich; Merck KGaA). Insulin (2.5%; Insuman® rapid; Sanofi S.A.) was added to BT-474 cells. Cells were incubated under humidified conditions at 37°C and 5% CO2. The cell lines were authenticated using PCR-single-locus-technology. Mycoplasma testing is performed regularly.
Cells were treated with 2.0 μg/ml carboplatin (Abmole Biosciences Inc.), 1.0 μg/ml epirubicin (Sigma-Aldrich; Merck KGaA), 40.0 μg/ml gemcitabine (Abmole Bioscience Inc.) or 2.0 μg/ml paclitaxel (Sigma-Aldrich; Merck KGaA) in DMSO-free solution for 18 h. Untreated cells served as a control.
Acquisition of patient samples
Female patients with histologically proven, invasive TNBC as defined by the St. Gallen Consensus 2012 (47), who were subjected to NACT were included in the present study. The inclusion criteria were as follows: i) Primary diagnosis of TNBC; and ii) age of >18 years. The exclusion criteria were as follows: i) Distant metastasis at the time of diagnosis; and ii) other previous or coexistent malignancies. The present study was approved by the institutional ethical review board of the University of Freiburg (permit no. 607/16) and written informed consent was obtained from each participant. Patients (aged 40-70 years) of the Department of Gynaecology and Obstetrics at the Medical Center-University of Freiburg were enrolled between February 2017 and September 2017; additionally, 20 healthy controls (aged 23-70 years) were enrolled between March 2016 and August 2016, of whom 10 provided serum samples, and 10 provided urine samples. All patients received NACT according to standard treatment protocols [90 mg/m2 epirubicin + 600 mg/m2 cyclophosphamide (EC) ×4 (once every 3 weeks), followed by 80 mg/m2 paclitaxel (P) ×12 (weekly)] (28). BC tissue biopsies were collected prior to NACT, and evaluated via haematoxylin and eosin (H&E) and Ki67 staining by the Pathology Department of the Medical Center-University of Freiburg. Histopathological preparations of BC were graded according to the classification published by Elston and Ellis (48) on H&E-stained slides, which includes counting of mitotic cells. Furthermore, the proliferation index was measured in sections immunohistochemically stained with the Ki67 antibody clone MIB-1. Any nuclear positivity (nucleoli excluded) was counted. The percentage of positive nuclei was either calculated from counting at least 200 tumour cells or estimated semi-quantitatively, especially for percentages <10 and >30%. In total, 9 patients with TNBC were enrolled. Patient characteristics are displayed in Table I. Blood (2-9 ml) was drawn by physicians at two time points (t0, prior to NACT; t1, immediately prior to third cycle of NACT), according to standard protocols. In order to evaluate the individual response of each patient to NACT, follow-up by US was performed and interpreted, according to the RECIST criteria (30). After coagulation for 20 min, whole blood was centrifuged (10 min, 2,000 × g, 4°C). Cell-free serum specimens were transferred to a fresh test tube. Samples exhibiting haemolytic characteristics were excluded. In total, 20 ml of urine was centrifuged (15 min, 2,000 × g, 4°C) and the cell-free supernatant was transferred to fresh test tubes. Serum and urine samples were stored at -20°C. Of the nine serum and urine samples collected from patients with TNBC, only eight met quality standards and were further analysed.
Table I.
Characteristics of patients with TNBC and the control cohort.
| A, Patients with TNBC
| ||||||
|---|---|---|---|---|---|---|
| Characteristics | Serum cohort (n=8)
|
Urine cohort (n=8)
|
||||
| N | % | Mean | N | % | Mean | |
| Age | 8 | 55.4 | 8 | 53.6 | ||
| Initial tumour stage (T) | ||||||
| cT1 | 6 | 75.0 | 6 | 75.0 | ||
| cT2 | 2 | 25.0 | 2 | 25.0 | ||
| Initial nodal status (N) | ||||||
| cN0 | 7 | 87.5 | 7 | 87.5 | ||
| n.a. | 1 | 12.5 | 1 | 12.5 | ||
| Initial tumour grade | ||||||
| G2 | 4 | 50.0 | 3 | 37.5 | ||
| G3 | 4 | 50.0 | 5 | 62.5 | ||
| Lymphovascular invasion (L) | ||||||
| L0 | 5 | 62.5 | 4 | 50.0 | ||
| L1 | 1 | 12.5 | 1 | 12.5 | ||
| n.a. | 2 | 25.0 | 3 | 37.5 | ||
| Proliferation rate (MIB-1 reactivity) | 64.3 | 60.3 | ||||
| Menopausal status | ||||||
| Premenopausal | 1 | 12.5 | 2 | 25.0 | ||
| Postmenopausal | 4 | 50.0 | 4 | 50.0 | ||
| n.a. | 3 | 37.5 | 2 | 25.0 | ||
| Early responsea | 3 | 37.5 | 4 | 50.0 | ||
| cCRb | 4 | 50.0 | 4 | 50.0 | ||
| pCRc | 3 | 37.5 | 5 | 62.5 | ||
| B, Controls
| ||||
|---|---|---|---|---|
| Control cohort | Serum cohort (n=10)
|
Urine cohort (n=10)
|
||
| N | Mean | N | Mean | |
| Age | 10 | 52.0 | 10 | 49.5 |
According to RECIST criteria, defined as clinically partial response or cCR immediately prior to the third cycle of NACT, and determined by ultrasonography.
Achievement of cCR throughout the course of NACT.
Achivement of pCR. All patients displayed a histological subtype of invasive ductal carcinoma. TNBC, triple negative breast cancer; cCR, clinically complete response; pCR, pathological complete response; NACT, neoadjuvant chemotherapy; MIB-1, anti-Ki67 antibody; n.a., no information available.
Isolation of ncRNA
Intracellular ncRNA was isolated using an EURx® GeneMATRIX Universal RNA/miRNA Purification kit (EURx Sp. z o.o.) according to the manufacturer's protocol. For the isolation of secreted microvesicular extracellular ncRNA, conditioned cell culture medium from BC cells was decanted into a 15-ml tube and centrifuged at 4°C at 8,000 × g for 5 min. In total, 10 ml supernatant was transferred to a new 15-ml tube. Microvesicles from liquid samples (conditioned cell culture medium, serum and urine) were isolated via filtration. RNA was subsequently isolated using a Norgen Total RNA Purification kit (Norgen Biotek Corp.), according to the manufacturer's protocol. In total, 20 ml urine and 2 ml serum were applied. The RNA concentration was determined using a NanoDrop™ ND1000 (Thermo Fisher Scientific, Inc.) and RNA was stored at -20°C until further processing.
Reverse transcription (RT)
cDNA was generated in a total reaction volume of 20 μl using poly(A) tailing-based RT. The RT reaction mix was as follows: 4 μl RT-buffer (5X), 1 μl 2.5 μM poly A adapter/primer (Integrated DNA Technologies, Inc.), 1 μl 5 mM dNTPs (Jena Bioscience), 0.25 μl Maxima™ H Minus reverse transcriptase (Thermo Fisher Scientific, Inc.), 0.25 μl SUPERase In™ RNase inhibitor (Thermo Fisher Scientific, Inc.), 0.5 μl 10 mM ATP (New England BioLabs, Inc.), 0.25 μl poly A polymerase (New England BioLabs, Inc.) and the required amount of RNA sample. Briefly, 1,000 ng of isolated intracellular ncRNA and 50 ng of isolated microvesicular extracellular ncRNA were used for RT. Due to the small amount of ncRNA in serum and urine, a fixed volume of 5 μl was used. RT was performed at 42°C for 30 min and 85°C for 10 min. Processed cDNA was stored at 4°C. Additionally, 2,000 ng of RNA was used for the RT of mRNA. The RT reaction mix contained 5 μl RT-buffer (5X), 1 μl 5 mM dNTPs (Jena Bioscience), 1 μl RT primer (10 μM), 0.25 μl Maxima™ H Minus reverse transcriptase (Thermo Fisher Scientific, Inc.), 0.25 μl SUPERase In™ RNase inhibitor (Thermo Fisher Scientific, Inc.) and the RT was carried out in a total volume of 25 μl. The RT temperature protocol was performed as follows: 65°C for 1 min, 25°C for 10 min, 50°C for 45 min and 85°C for 10 min.
Quantitative PCR (qPCR)
In total, 1 μl cDNA at a concentration of 5 ng/μl was used for qPCR at a 1:10 dilution with the master mix. Master mix was composed of 1 μl qPCR buffer (10X), 0.5 μl 5 mM dNTPs (Jena Bioscience), 0.5 μl SYBR® Green (Jena Bioscience), 6.45 μl nuclease-free water (Analytik Jena AG), 0.05 μl HotStart Taq polymerase (Jena Bioscience) and 0.5 μl primer pair mix. The primers were designed using miRprimer (49). Sequences of specific sense and antisense primers are listed in Table SI. Duplicate analysis was performed on a LightCycler® 480 Instrument II (Roche Diagnostics) with pre-incubation at 95°C for 2 min and a two-step amplification of 40 cycles at 95°C for 5 sec and 60°C for 30 sec. Based on an extensive literature review (date, 28/02/2019) employing the academic web resource PubMed (https://www.ncbi.nlm.nih.gov/pubmed/) with a focus on (TN)BC-related ncRNAs and their involvement in chemotherapy response and resistance, the following panel of ncRNAs was selected and subsequently analysed: Let-7a; let-7e; miR-7, -9, -15a, -17, -18a, -19b, -21, -30b, -222 and -320c; piR-36743; and GlyCCC2. The following key words were used: 'miRNA' or 'piRNA', combined with 'breast cancer' (4,571 and 4,581 hits, respectively) and 'TNBC' (261 and 157 hits, respectively). The literature review on the chosen panel of ncRNAs was extended by searching for each target name individually combined with the key words 'breast cancer', 'TNBC', 'chemotherapy', 'chemoresponse' and 'chemoresistance'. The findings from this literature search are summarised in Table II.
Table II.
Involvement of investigated ncRNAs in (TN)BC and chemoresistance.
| ncRNA | Involvement in BC | Involvement in TNBC | Involvement in chemoresistance |
|---|---|---|---|
| let-7a | Tissue ↓ (92,93); serum ↓ (94,95); plasma ↓ (96); pathological features (97); response (98); subtyping (99) | Serum ↑ (100); tissue ↑ (100); cell line ↑ (101) | DDP [c (102,103)]; DOX [m (104), c (104-106)]; EPI [t (107), c (107)]; GEM [c (108)]; PTX [t (109), c line (106,110)]; therapeutic approach (111) |
| let-7e | Metastasis (112); prognosis (113); response (114); subtyping (115) | Tissue ↓ (113) | 5-FU [t (113,115), c (116)]; CAP [t (117)]; DDP [c (118,119)]; DOX [c (114)]; PTX (119); PT [t (120)] |
| miR-7 | Tissue ↑ (85); metastasis (121,122); prognosis (67), recurrence (123); response (124) | Tissue ↑ (85); cell line ↓ (125,126) | DDP [t (127)]; DOX [c (65)]; PTX [c (128)]; TAM [t (67), c (66)]; c (129) |
| miR-9 | Metastasis (130,131); survival (132) | Tissue ↓ (133), ↓ (134); cell line ↑ (125,135); survival (136); outcome (137); response (68); neovascularization (138) | 5-FU [t (68,139)]; DDP [t (68,139)]; CPA [t (68,139)]; DOX [t (68,139)]; PTX [t (68,139), c (139)] |
| miR-15a | Plasma ↑ (140); serum ↑ (141); apoptosis (142,143); Her2neu amplification (144); prognosis (145) | Prognosis (145) | BOR [bm (146)]; DDP [c (147)]; PTX [c (148,149)]; TAM [c (150)] |
| miR-17 | Grading (151); metastasis (152); prognosis (153); recurrence (154,155); survival (156); TRA response (157) | Plasma ↓ (158); tissue, cell line ↓ (159); tissue ↓ (85); apoptosis (160); metastasis (161); progression (162) | 5-FU [c (163,164)]; CAP [p (165)]; DDP [c (163, 164,166)]; DOX [c (163)]; GEM [c (167-169)]; OXP [p (165)]; PTX [t (170), c (170-174)] |
| miR-18a | Serum ↑ (141); metastasis (152,175); recurrence (155); BC risk (176,177) | Tissue ↑ (85,134,178) | CAP [p (165)]; DDP [c (179,180)]; OXP [p (165)]; PTX [c (178,181)]; TRA [c (182)] |
| miR-19b | Serum f (183); metastasis (184,185); prognosis (184); tumourigenesis (186); tumour suppressor (187) | Cell line | (188) | 5-FU [c (189)]; CAP [p (165)]; DOX [c (190)]; OXP [p (165), c (191)]; SAR [c (192)] |
| miR-21 | Tissue ↑ (85,193-198); serum ↑ (199-202); plasma ↑ (96,203); urine ↓ (59); diagnosis (195,204,205); invasion (206); prognosis (196,207); recurrence (208,209); response (87) | Serum ↑ (100); plasma ↓ (210); tissue ↑ (100,211,212); cell line ↑ (211) | 5-FU [p (213), t (214), c (215)]; CAR [c (216)]; DDP [p (213,217), c (218-220)]; DOX [c (221,222)]; FUF [c (223)]; GEM [c (224-226)]; OXP [c (215)]; PTX [c (215,227-231)]; TOP [c (232)]; TRA [m (233), |
| miR-30b | Serum ↓ (238); blood ↑ (239); metastasis (240); non-IBC (144); recurrence (241) | DDP [c (242)]; GEF [c (243)]; TRA [c (244)]; TKI [t (245)] | |
| miR-222 | Tissue ↑ (246,247); serum ↑ (176); nodal status (194); prognosis (248); progression (249); recurrence (250); response (251,252) | Cell line ↑ (253,254); tissue ↓ (249) | DDP [c (255)]; DOX [c (256,257)]; EPI [p (251), t (252,257)]; FUF [c (258)]; GEF [c (243)]; GEM [c (259)];PTX [p (251), c (260)];TAM [c (251,261,262)] |
| miR-320c | Not yet connected to BC. | 5-FU [c (263)]; GEM [c (264)] | |
| piR-36743a | Tissue ↑ (86) | ||
| GlyCCC2 | Not yet connected to BC. Content of serum and urine exosomes (265) |
DQ598677 is an alternative name for piR-36743. BC, breast cancer; TNBC, triple negative BC; miR, microRNA; piR, PIWI-interacting RNA; ↓, downregulation; ↑, upregulation; c, cell line; m, mouse; t, tissue; bm, bone marrow; p, plasma; DDR cisplatin; DOX, doxorubicin; EPI, epirubicin; GEM, gemcitabine; PTX, paclitaxel; 5-FU, 5-fluorouracil; CAR capecitabine; PT, platinum; TAM, tamoxifen; CPA, cyclophosphamid; BOR, bortezomib; OXP, oxaliplatin; TRA, trastuzumab; SAR, saracatinib; CAR, carboplatin; FUL, fulvestrant; TOP, topotecan; GEF, gefitinib; TKI, tyrosine kinase inhibitors.
According to the ΔΔCq method (50,51), relative quantification of intracellular ncRNA expression was performed by normalising the quantification cycle (Cq) values to the geometric mean of the two endogenous controls, RNU-44 and -48 (51-54), which were determined as stably expressed using the BestKeeper software tool (55). Due to a lack of reliable endogenous controls in conditioned cell culture media, raw data of extracellular microvesicular ncRNA were normalised to the global mean of all assessed ncRNAs in this setting. Although recommended for larger data sets, this approach is commonly accepted when no suitable endogenous control can be established and can lead to more accurate performance than normalising to stable genes (52,56,57). Within the analysed serum samples, BestKeeper identified miR-26b and -191 as the most suitable endogenous controls. Similarly, miR-16 and -26b served as endogenous controls in the urine-based analysis (58,59). Additionally, 5 fmol of exogenous synthetic miRNA from Arabidopsis thaliana (ath-miR-159a) and Caenorhabditis elegans (cel-miR-39; both from biomers.net GmbH) were mixed with serum and urine samples as spike-in controls to optimise normalisation (60).
The targets of the mRNA analysis included cyclin D1, TOP2α, TPX2, DDX5 and -17. Cq values were normalised to the housekeeping gene, 5′-aminolevulinate synthase 1 (61).
Protein isolation and western blotting
Proteins from the BC cell lines were isolated from the flow-through of the RNA-binding columns used to purify RNA by the addition of 1 ml of isopropanol and subsequent centrifugation for 30 min at 12,000 × g at 4°C. The protein pellet was washed with 70% ethanol and lysed in protein lysis buffer (1% SDS, 1% triton, 1 mM EDTA, 0.1 M HEPES) for 30 min at 1,000 RPM and 50°C in an Eppendorf ThermoMixer®. Protein concentrations were determined using the bicinchoninic acid method as previously described (62). Western blotting was conducted as described by Schagger and von Jagow (63). Briefly, 10 μg/lane of protein was separated on a 10% SDS gel. Proteins were transferred onto a 0.45-μm PVDF membrane (Sigma-Aldrich; Merck KGaA) and membranes were blocked with 3% skimmed milk in PBS-Tween 20 at room temperature for 30 min. Monoclonal mouse anti-human antibodies for immunodetection were purchased from Santa Cruz Biotechnology, Inc. The following dilutions were used: Cyclin D1 (1:500; cat. no. sc-8396); DDX5 (1:5,000; cat. no. sc-365164); DDX17 (1:10,000; cat. no. sc-398168); p-Actin (1:1,000; cat. no. sc-47778); and proliferating cell nuclear antigen (PCNA; 1:1,000; cat. no. sc-56). Membranes were incubated with primary antibodies overnight at 4°C. A secondary peroxidase-conjugated anti-mouse antibody was purchased from Jackson ImmunoResearch Europe, Ltd. (cat. no. 115-036-146) and was incubated at room temperature with the membrane for 1 h (1:10,000). Bands were visualised using ECL reagent (Santa Cruz Biotechnology, Inc.). PCNA and p-Actin served as loading controls.
Statistical analysis
All statistical tests were performed in R 3.5.2 (64). Multivariable statistical analysis of in vitro results, including three independent experimental repeats, consisted of three factors: Cell line (BT-474, BT-20, HS-578T, MDA-MB-231), treatment (control, carboplatin, epirubicin, gemcitabine, paclitaxel) and compartment (intracellular, extracellular). The untreated BT-20 cell line was defined as the expected control value (intercept) in the statistical data assessment. A linear model for the log-transformed in vitro results was selected with the main effect of the cell line, three two-way interactions of the cell line and treatment, cell line and compartment, and treatment and compartment, and the three-way interaction of all three factors. In vivo results were statistically analysed by performing a two-sample t-test comparing patients with TNBC to the healthy control group. For a time point comparison (t0 vs. t1) within the TNBC group, a two-tailed paired t-test was applied. Preliminary two-tailed paired t-tests were performed for the serum data in a clinical complete response (cCR) vs. no cCR subgroup analysis, defined by the RECIST criteria (30). P<0.05 indicated a statistically significant difference.
Results
In the present study, the influence of chemotherapeutic treatment on the expression levels of 14 BC-related ncRNAs (let-7a/e, miR-7, -9, -15a, -17, -18a, -19b, -21, -30b, -222 and -320c, piR-36743 and GlyCCC2) was investigated in three TNBC (BT-20, HS-578T and MDA-MB-231) cell lines and one triple positive BC (BT-474) cell line, both intra- and extracellularly.
All examined cell lines expressed all of the 14 investigated ncRNA targets. In addition to intracellular expression, the vast majority of the panel were detectable as secreted molecules in microvesicles in cell culture medium, with the exception of miR-7, -9 and -15a in the medium of MDA-MB-231, and miR-9 in the medium of HS-578T cells. Chemotherapy treatment of cells with carboplatin, epirubicin, gemcitabine and paclitaxel resulted in distinct ncRNA expression alterations depending both on the cytotoxic agent and on the specific BC cell line (Fig. 1).
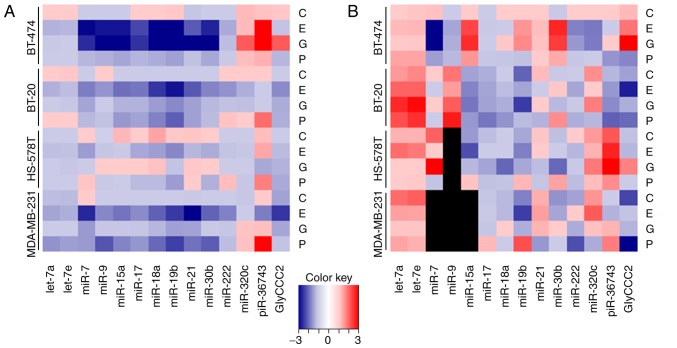
Figure 1.
Chemotherapy-driven ncRNA regulations in (triple negative) breast cancer cells. Relative expression levels (ΔΔCq) of ncRNAs in BT-474, BT-20, HS-578T and MDA-MB-231 cells treated with 2.0 μg/ml carboplatin, 1.0 μg/ml epirubicin, 40.0 μg/ml gemcitabine and 2.0 μg/ml paclitaxel for 18 h as determined by reverse transcription-quantitative PCR. Heatmaps demonstrate (A) intracellular expression levels normalised to the geometric mean of RNU-44 and -48, and (B) secreted microvesicular ncRNA levels normalised to the global mean of all investigated targets. Black squares indicate no reliably detectable ncRNA expression. ncRNAs, non-coding RNAs; miR, microRNA; piR, PIWI-interacting RNA; C, carboplatin; E, epirubicin; G, gemcitabine; P, paclitaxel.
Epirubicin
Epirubicin treatment (1.0 μg/ml) influenced intracellular and secreted microvesicular expression levels of all the ncRNAs in the panel in all the cell lines investigated. A uniform downregulation of all analysed targets, except for piR-36743 in BT-474 was observed, intracellularly. The levels of secreted microvesicular ncRNAs did not suggest a distinct regulation pattern in respect of target and cell line (Fig. 1).
Multivariable analysis confirmed the inhibitory effect of epirubicin on ncRNA expression, especially in hormone receptor positive BT-474 and TNBC cell lines BT-20 and MDA-MB-231. miR-7, -15a, -17, -18a, -19b, -21, -30b and -222 were downregulated in BT-474, BT-20 and MDA-MB-231 cell lines compared with the intercept, while reduced expression of miR-9 was restricted to BT-474 and MDA-MB-231 cells. GlyCCC2 expression was downregulated in BT-20 and MDA-MB-231 cells. piR-36743 expression decreased following epirubicin treatment in MDA-MB-231 cells, while an increase in expression was found in BT-474 cells. In HS-578T, the effect of epirubicin was limited to miR-17 and -18a expression. All estimates of multivariable analysis, including confidence intervals (CIs) and P-values are listed in Table SII.
Gemcitabine
Gemcitabine (40.0 μg/ml) predominantly triggered a decrease in the expression of ncRNAs in the intracellular compartment, except in the cases of miR-320c, piR-36743 and GlyCCC2 in BT-474 cells, where an upregu- lation was observed. The levels of secreted microvesicular ncRNAs were altered bidirectionally (Fig. 1).
In a multivariable analysis, gemcitabine treatment caused major changes in the expression levels of ncRNAs in BT-474 cells, while few detectable alterations were observed in the TNBC cell lines. Specifically, miR-7, -9, -15a, -17, -18a, -19b, -21 and -30b were downregulated compared with the intercept, while miR-320c, piR-36743 and GlyCCC2 were upregulated in BT-474 cells. In the TNBC cell lines, downregulated expression was detected for miR-17, -18a and -21 in BT-20, and miR-7 and -21 in MDA-MB-231 cells. No significant effect was detected in HS-578T cells. All estimates of multivariable analysis, including CIs and P-values are listed in Table SII.
Paclitaxel
Intracellularly, the expression levels of the ncRNAs predominantly decreased following paclitaxel (2.0 μg/ml) treatment. However, in the cases of miR-7 (HS-578T), -222 (BT-20 and HS-578T), -320c (BT-474, BT-20 and MDA-MB-231) and piR-36743 (all cell lines), upregulation was detected. The levels of secreted microvesicular ncRNAs were influenced by paclitaxel and differed among the investigated cell lines (Fig. 1).
Multivariable analysis found that paclitaxel triggered ncRNA expression fluctuations predominantly in the TNBC cell lines BT-20 and MDA-MB-231. Compared with the intercept, miR-17, -18a, -19b and -21 were downregulated in BT-20 cells, while piR-36743 was upregulated. In MDA-MB-231 cells, the expression levels of let-7a, let-7e, miR-7, -9, -15a, -17, -18a, 19b, -21, -30b and -222 were decreased, and piR-36743 was upregulated. In the HS-578T cell line, no alteration in the investigated ncRNA profiles was detected, except for the upregulation of piR-36743. Paclitaxel treatment caused the downregulation of miR-17 in BT-474 cells compared with the intercept. All estimates of multivariable analysis, including CIs and P-value are listed in Table SII.
Carboplatin
Treatment with carboplatin (2.0 μg/ml) did not influence ncRNA expression in this model (Table SII).
Intracellular vs. secreted microvesicular ncRNA expression levels
Focusing on the extracellular compartment, no conclusions can be drawn from the observation of an increase in the relative levels of ncRNA expression under control conditions compared with the intracellular expression levels for all targets, which was mirrored by significantly increased estimates in multivariable analysis (Table SIII). This phenomenon cannot be explained by biological variation but is, at least in part, observed due to different normalisation methods in intra- and extracellular analysis.
The applied model of multivariable analysis allowed the estimation of the additional impact on ncRNA expression caused by three-way interactions with the variables 'chemotherapy', 'cell line' and 'extracellular compartment'. Most of these influences were observed in the BT-474 cell line and were less frequent in the TNBC cell lines. Depending on the estimate, the aforementioned regulation of the expression of the ncRNAs will be increased (estimate >1) or alleviated (estimate <1; Table SIII). For the most part, the three-way interactions attenuated the effects presented in Table SII. This was observed for miR-15a, -17, -18a, -19b, -21, -30b, -320c and piR-36743 following gemcitabine treatment, as well as for miR-15a, -17, -30b, -320c and piR-36743 following epirubicin treatment in BT-474 cells. In the TNBC cell line HS-578T, the three-way interactions resulted in the opposite effects on miR-18a, -21 and piR-36743 following gemcitabine treatment. The same applied to miR-21 and -222 following epirubicin treatment, and for miR-17 and -19b following paclitaxel treatment in MDA-MB-231 cells (Table SIII).
While lacking significance in the aforementioned observations, three-way interaction influences estimated by multivariable analysis added to the effect of piR-36743 upregu- lation in HS-578T cells following epirubicin treatment (1.310; P=0.208) by 2.662 (P=0.024). Similarly, let-7a expression following gemcitabine treatment (0.929; P=0.760) decreased by 0.357 (P=0.036) in this cell line. Let-7e expression (0.887; P=0.619) further decreased in the same context by 0.337 (P=0.027). The reduced GlyCCC2 expression induced by paclitaxel treatment in MDA-MB-231 cells (0.772; P=0.372) was further decreased by a value of 0.277 (P= 0.029). In BT-474 cells, paclitaxel induced the downregulation of miR-9 (0.752; P=0.233), which was increased by 0.332 (P=0.023). These effects may indicate that let-7a and -7e, and GlyCCC2 may be promising liquid biomarker candidates depending on the administered chemotherapy (Table SIII). In the case of piR-36743, three-way interaction influences added to already significant findings as aforementioned. While the upregulation of piR-36743 following paclitaxel treatment in HS-578T cells (1.793; P= 0.008) increased by 3.492 (P=0.004), piR-36743 increased in MDA-MB-231 cells (2.922; P<0.001) by 2.428 (P=0.040; Table SIII).
Chemotherapy-induced ncRNA regulation pattern
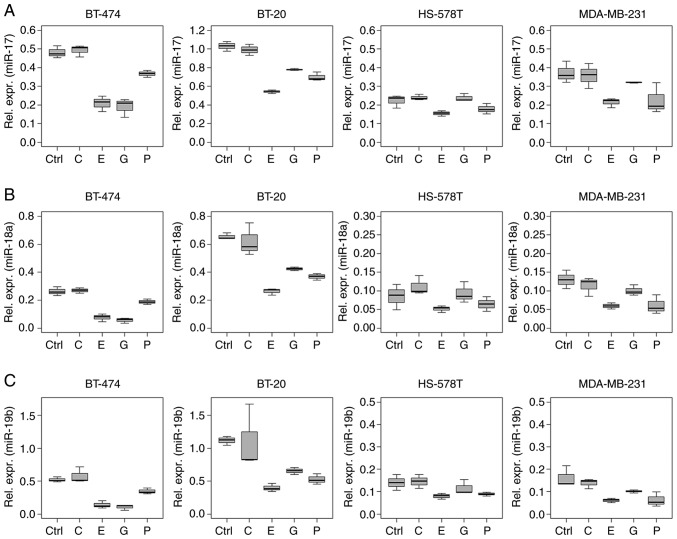
Several ncRNAs analysed in the present study showed a similar regulation pattern, especially in the intracellular analysis. This phenomenon was observed for miRNAs of the miR-17~92 cluster (miR-17, -18a and -19b), and for miR-15a and -30b (Figs. 2 and 3).
Figure 2.
Expression profile of miR-17~92 cluster miRNAs. Relative expression levels (ΔCq) of the miR-17~92 cluster miRNAs (A) miR-17, (B) -18a and (C) -19b in BT-474, BT-20, HS-578T and MDA-MB-231 breast cancer cells treated with 2.0 μg/ml carboplatin, 1.0 μg/ml epirubicin, 40.0 μg/ml gemcitabine and 2.0 μg/ml paclitaxel for 18 h as determined by reverse transcription-quantitative PCR. Data are normalised to the geometric mean of RNU-44 and -48. Untreated cells served as the control. Boxplots indicate median (thick line), first and third quartile (box lines) and maximal/minimal value (upper and lower line). Y-axis scaling can deviate to improve readability. miR/miRNA, microRNA; C, carboplatin; E, epirubicin; G, gemcitabine; P paclitaxel; Ctrl, control.
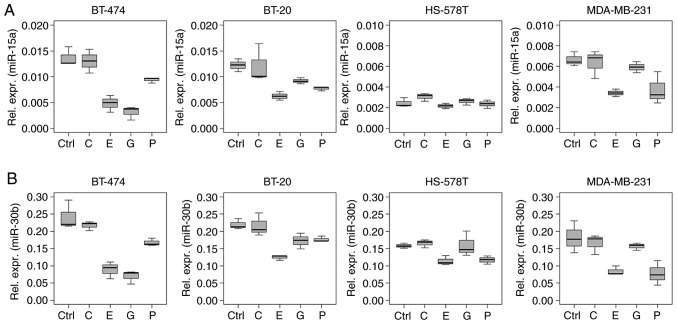
Figure 3.
Chemotherapy-driven regulation of miR-15a and -30b in (triple negative) breast cancer cells. Relative expression levels (ΔCq) of (A) miR-15a and (B) -30b in BT-474, BT-20, HS-578T and MDA-MB-231 cells treated with 2.0 μg/ml carboplatin, 1.0 μg/ml epirubicin, 40.0 μg/ml gemcitabine and 2.0 μg/ml paclitaxel for 18 h, normalised to the geometric mean of RNU-44 and -48, as determined by reverse transcription-quantitative PCR. Untreated cells served as the control. Boxplots indicate median (thick line), first and third quartile (box lines) and maximal/minimal value (upper and lower line). Y-axis scaling can deviate to improve readability. miR, microRNA; C, carboplatin; E, epirubicin; G, gemcitabine; P, paclitaxel; Ctrl, control.
Markers of proliferation
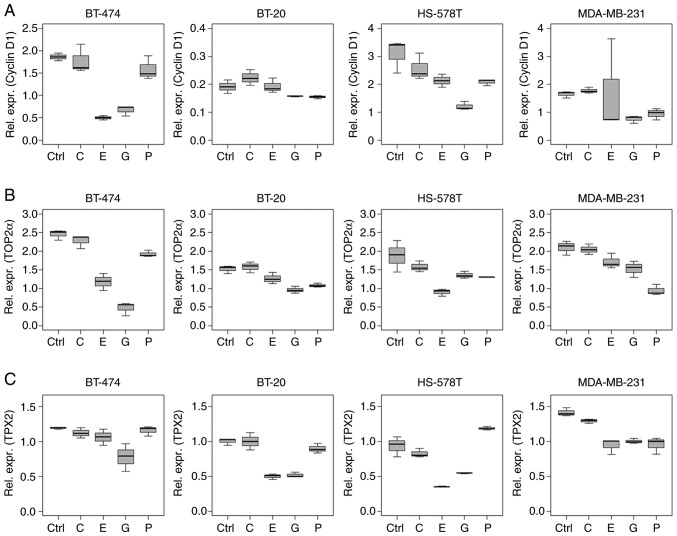
Cyclin D1 mRNA expression levels reduced after treatment with epirubicin and gemcitabine (Fig. 4A). Multivariable analysis indicated a decrease in cyclin D1 expression levels in BT-474 and HS-578T cells (epirubicin, P<0.001 and P=0.010, respectively), and in BT-474, HS-578T and MDA-MB-231 cells (gemcitabine, P=0.002, P<0.001 and P=0.016, respectively) compared with the intercept. A reduction was also observed in paclitaxel-treated HS-578T cells (P=0.007). Carboplatin did not influence cyclin D1 mRNA expression in this setting. All estimates of multivariable analysis, including CIs and P-values are listed in Table SIV.
Figure 4.
Influence of chemotherapy treatment on cell cycle determinants in (triple negative) breast cancer cell lines. Relative mRNA expression levels (Δ Cq) of (A) cyclin D1, (B) TOP2α and (C) TPX2 in BT-474, BT-20, HS-578T and MDA-MB-231 cells under control conditions, and treated with 2.0 μg/ml carboplatin, 1.0 μg/ml epirubicin, 40.0 μg/ml gemcitabine and 2.0 μg/ml paclitaxel for 18 h. Data are normalised to 5′-aminolevulinate synthase 1 and values were determined by reverse transcription-quantitative PCR. Boxplots indicate median (thick line), first and third quartile (box lines) and maximal/minimal value (upper and lower line). Y-axis scaling can deviate to improve readability. TPX2, targeting protein for Xklp2; TOP2α, DNA topoisomerase 2α; C, carboplatin; E, epirubicin; G, gemcitabine; P, paclitaxel; Ctrl, control.
TOP2α mRNA levels were reduced following chemotherapy, except following carboplatin treatment in all the cell lines tested (Fig. 4B). P-values, estimates and CIs are listed in Table SIV.
TPX2 mRNA level alterations were similar to those of cyclin D1, with reductions in expression caused by epirubicin (BT-20, HS-578T and MDA-MB-231; P<0.001) and gemcitabine (all cell lines; P<0.001) treatment (Fig. 4C). Paclitaxel treatment reduced TPX2 levels in BT-474 and MDA-MB-231 cells (P<0.001), but increased TPX2 expression in HS-578T cells (P=0.001; Table SIV).
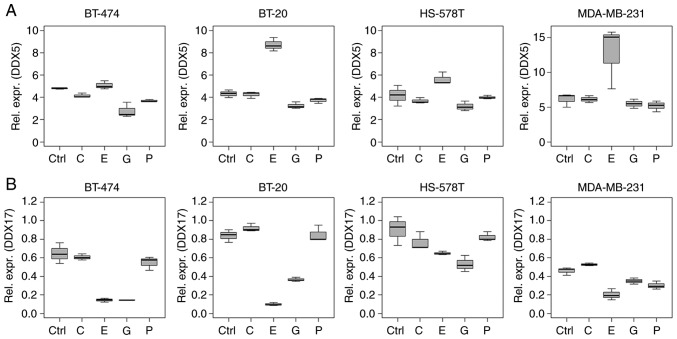
While epirubicin treatment induced differential regulation of DDX5 and -17 mRNA levels, DDX17 was regulated in a similar manner as the miR-17~92 cluster (Figs. 2 and 5). Specifically, DDX5 mRNA was upregulated by epirubicin compared with the intercept (BT-20 and MDA-MB-231, P<0.001) and downregulated in BT-474 by gemcitabine (P=0.034). By contrast, DDX17 mRNA was downregulated following epirubicin treatment in all the cell lines tested (all P<0.001). Downregulation was also observed following gemcitabine treatment in BT-20, BT-474, HS-578T (P<0.001) and MDA-MB-231 cells (P=0.049), as well as by paclitaxel in MDA-MB-231cells (P=0.006; Table SIV).
Figure 5.
Influence of chemotherapy treatment on DDX proteins in (triple negative) breast cancer cell lines. Relative mRNA expression levels (ΔCq) of (A) DDX5 and (B) DDX17 in BT-474, BT-20, HS-578T and MDA-MB-231 cells under control conditions, and treated with 2.0 μg/ml carboplatin, 1.0 μg/ml epirubicin, 40.0 μg/ml gemcitabine and 2.0 μg/ml paclitaxel for 18 h. Data are normalised to 5′-aminolevulinate synthase 1. Values were determined using reverse transcription-quantitative PCR. Boxplots indicate median (thick line), first and third quartile (box lines) and maximal/minimal value (upper and lower line). Y-axis scaling can deviate to improve readability. DDX, DEAD-box polypeptide; C, carboplatin; E, epirubicin; G, gemcitabine; P, paclitaxel; Ctrl, control.
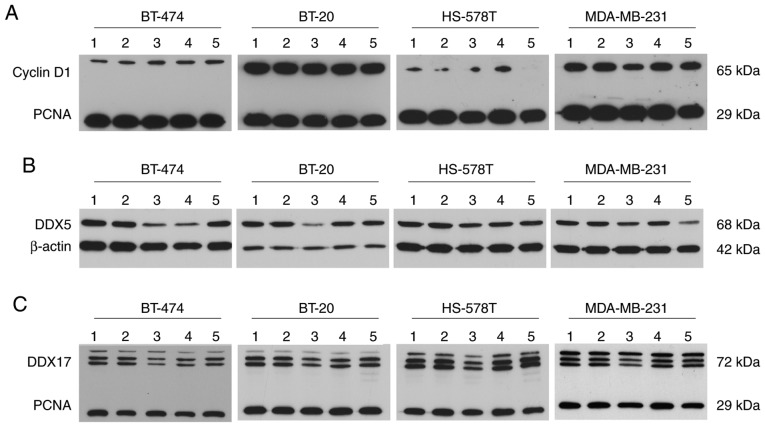
At the protein level, cyclin D1 was detected at a molecular weight of ~65 kDa. This immunoprecipitated protein may represent a ubiquitylated form of cyclin D1 and was detected in all investigated cell lines. The expression of cyclin D1 in HS-578T cells, however, was present at modest levels. Compared with the control conditions, no regulation of cyclin D1 was observed, except for a modest reduction caused by epirubicin treatment in MDA-MB-231 cells (Fig. 6A). Western blotting revealed differential DDX5 regulation following epirubicin treatment at the protein level compared with mRNA regulation; DDX5 was downregulated at the protein level. Following gemcitabine treatment, DDX5 protein levels also declined in BT-474 and HS-578T cells, while they declined in MDA-MB-231 cells following paclitaxel treatment. A reduced DDX17 protein level caused by epirubicin is consistent with reduced DDX17 mRNA. In BT-474 and BT-20 cells, gemcitabine led to lower DDX17 protein expression; paclitaxel had the same effect in MDA-MB-231 cells (Fig. 6B and C).
Figure 6.
Chemotherapy treatment influences DDX5 and -17 protein levels in (triple negative) breast cancer cells. Protein expression of (A) DDX5, (B) DDX17 and (C) cyclin D1 in BT-474, BT-20, HS-578T and MDA-MB-231 cells treated with 2.0 μg/ml carboplatin (lane 2), 1.0 μg/ml epirubicin (lane 3), 40.0 μg/ml gemcitabine (lane 4) and 2.0 μg/ml paclitaxel (lane 5) for 18 h, as determined by western blotting compared with untreated cells (lane 1). β-actin and PCNA served as loading control. DDX, DEAD-box polypeptide; PCNA, proliferating cell nuclear antigen.
In vivo pilot study
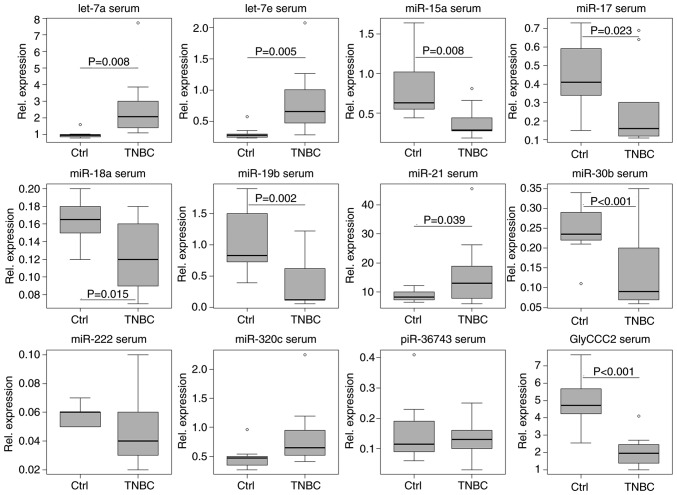
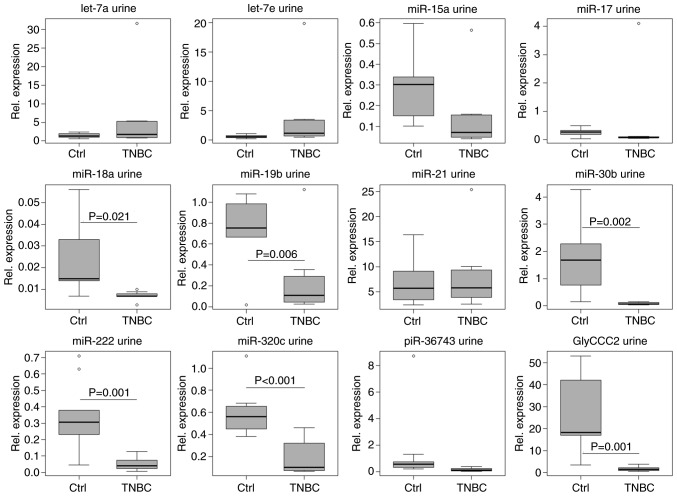
In serum, untreated patients with TNBC (t0) could be clearly distinguished from healthy controls by the expression levels of the vast majority of ncRNAs investigated, with the exception of miR-222, -320c and piR-36743 (Fig. 7). A two-sample t-test confirmed upregulated let-7a (P=0.008, SE=0.662), -7e (P=0.005, SE=0.179) and miR-21 (P=0.039, SE=4.090) expression levels in serum from patients with TNBC compared with the control group. Decreased expression was observed for miR-15a (P=0.008, SE=0.154), -17 (P=0.023, SE=0.087), -18a (P=0.015, SE=0.015), -19b (P=0.002, SE=0.209), -30b (P<0.001, SE=0.031) and GlyCCC2 (P<0.001, SE=0.623) in TNBC. In urine, untreated patients with TNBC could be distinguished from healthy women by decreased levels of miR-18a (P=0.021, SE=0.006), -19b (P=0.006, SE=0.159), -30b (P=0.002, SE=0.484), -222 (P=0.001, SE=0.071), -320c (P<0.001, SE=0.087) and GlyCCC2 expression (P=0.001, SE=5.803; Fig. 8).
Figure 7.
ncRNA expression in serum distinguishes patients with TNBC from controls. Relative expression levels (ΔCq) of microvesicular ncRNAs (let-7a/e, miR-15a, -17, -18a,-19b, -21, -30b, -222 and -320c, piR-36743 and GlyCCC2) in the serum of 8 patients with TNBC and 10 healthy controls, normalised to the geometric mean of miR-26b and -191, as determined by reverse transcription-quantitative PCR. Let-7a/e and miR-21 expression levels were increased in patients with TNBC compared to controls (two-tailed t-test; P<0.05). miR-15a, -17, -18a, -19b, -30b and GlyCCC2 were decreased. Boxplots indicate median (thick line), first and third quartile (box lines), maximal/minimal value (upper and lower line) and ° (moderate outlier). Y-axis scaling can deviate to improve readability. ncRNAs, non-coding RNAs; miR, microRNA; TNBC, triple negative breast cancer; piR, PIWI-interacting RNA; Ctrl, control.
Figure 8.
ncRNA expression in urine can distinguish patients with TNBC from controls. Relative expression levels (ΔCq) of microvesicular ncRNAs (let-7a/e, miR-15a, -17, -18a, -19b, -21, -30b, -222 and -320c, piR-36743 and GlyCCC2) in the urine of 8 patients with TNBC and 10 healthy controls, normalised to the geometric mean of miR-16 and -26b, as determined by reverse transcription-quantitative PCR. miR-18a, -19b, -30b, -222 and -320c, and GlyCCC2 expression levels were decreased in patients with TNBC compared to controls (two-tailed t-test; P<0.05). Boxplots indicate median (thick line), first and third quartile (box lines), maximal/minimal value (upper and lower line) and ° (moderate outlier). Y-axis scaling can deviate to improve readability. ncRNA, non-coding RNA; TNBC, triple negative breast cancer; miR, microRNA; piR, PIWI-interacting RNA; Ctrl, control.
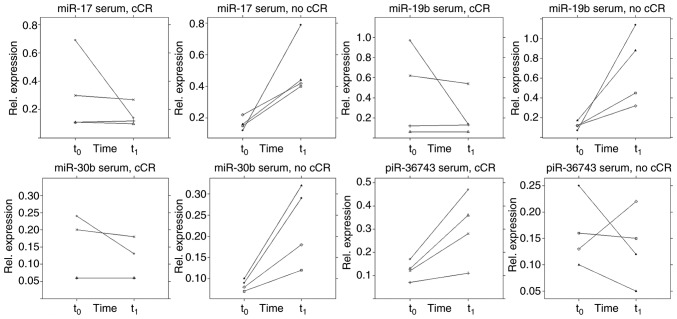
In the present study, no clear pattern was observed when dividing the TNBC group into early responder and non-responder [early response as defined by the achievement of a clinical partial response (cPR) according to RECIST criteria (30) immediately prior to the third cycle of NACT]. Also, no association was observed regarding pCR as an outcome (data not shown). However, a trend was discovered in the serum of patients receiving a cCR during NACT (Fig. 9). In this respect, those patients (n=4) showed a different ncRNA regulation pattern immediately prior to the third cycle of NACT (q) compared with patients (n=4) who did not achieve a cCR during NACT. A two-tailed paired t-test estimated a difference from t0 to t1 between the two groups in terms of serum expression of miR-17 (P=0.029, SE=0.173), -19b (P=0.030, SE= 0.282), -30b (P=0.011, SE=0.048) and piR-36743 (P=0.028, SE=0.072). However, these results remain preliminary due the small sample size. By contrast, no specific pattern was detected within the TNBC group using the urine specimens. A general rise of ncRNA expression from t0 to t1 was evident (Fig. 10).
Figure 9.
NACT influences ncRNA expression in the serum of patients with TNBC. Relative expression levels (ΔCq) of microvesicular ncRNAs (miR-17, -19b. -30b and piR-36743) in the serum of 4 patients with TNBC who achieved a cCR during NACT and 4 patients with TNBC who did not (no cCR), normalised to the geometric mean of miR-26b and -191, as determined by reverse transcription-quantitative PCR. Samples were taken at two time points: t0 and t1. Y-axis scaling can deviate to improve readability. ncRNA, non-coding RNA; TNBC, triple negative breast cancer; miR, microRNA; cCR, clinical complete response; NACT, neoadjuvant chemotherapy; piR, PIWI-interacting RNA; t0, prior to NACT; t1 immediately prior to the third cycle of therapy.
Figure 10.
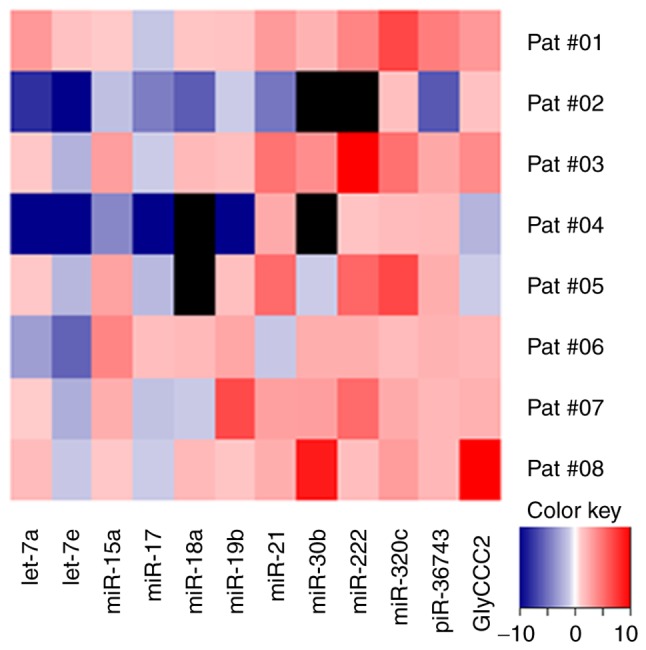
NACT influences ncRNA expression in the urine of patients with TNBC. Relative expression levels (ΔCq) of microvesicular ncRNAs in the urine of 8 patients with TNBC during NACT, normalised to the geometric mean of miR-16 and -26b, as determined by reverse transcription-quantitative PCR. Heatmap demonstrates fold change of ncRNA expression levels from t0 to t1. Black squares indicate no reliably detectable ncRNA expression. ncRNA, non-coding RNA; TNBC, triple negative breast cancer; miR, microRNA; cCR, clinical complete response; NACT, neoadjuvant chemotherapy; piR, PIWI-interacting RNA; t0, prior to NACT; t1, immediately prior to third cycle of therapy.
Discussion
The present study demonstrated that dysregulation of ncRNAs in TNBC cell lines, serum and urine can be induced by chemotherapy treatment. In vitro, the influences of chemotherapeutic treatment with epirubicin, gemcitabine and paclitaxel on the expression levels of a (TN)BC-related ncRNA panel were evaluated. Even in the extracellular compartment, influences were detected, indicating the effects of chemotherapy-driven ncRNA secretion. The expression profiles differed depending on cell line and cytostatic drug. In this in vitro model, epirubicin caused the strongest effect on ncRNA regulation.
Except for miR-7 and -9, the present study reliably detected all targets in microvesicles from liquid samples (conditioned cell culture media, serum and urine). As intracellularly, miR-7 and -9 resulted in distinct expression alterations caused by chemotherapy treatment, these ncRNAs may still be involved in therapy response mechanisms. Previous studies have demonstrated associations between BC therapy response, and the dysregulation of miR-7 and -9 in cell lines and tissue (65-68). However, these two miRNAs were not detectable in microvesicles of liquid samples in the present study, and cannot be recommended as suitable liquid biopsy marker candidates in this context.
ncRNA expression alterations occurred along with altered expression of cell cycle markers and, therefore, may be associated with it. While epirubicin exhibited the most potent effects on ncRNA regulation, carboplatin treatment did not significantly influence ncRNA expression in the present study. This observation is consistent with the present findings on the influence of chemotherapeutic treatment on cell cycle determinants. Cyclin D1, TOP2α and TPX2 were not influenced by carboplatin, indicating limitations of this in vitro model. All cells were treated with the chemotherapeutic agent for 18 h. Carboplatin may cause an inhibitory effect on cell proliferation, and possibly affect ncRNA expression, when treatment time is prolonged (69-72). In western blotting, cyclin D1 could not be detected in its native conformation, with a molecular weight of 37 kDa. However, the product detected was of 65 kDa, and may represent a dimer or a ubiquitylated form of cyclin D1. Cyclin D1 is degraded by ubiquitylation and the native protein possesses a short half-life of <30 min (73,74).
Apart from carboplatin-treated cells, mRNA expression of cell cycle determinants was reduced in most of the other samples as aforementioned. Epirubicin and gemcitabine treatment impacted proliferation and led to ncRNA expression alterations.
DDX proteins, a family of RNA helicases, are dysregulated in several types of cancer, including BC (75). Their cellular functions are highly context-dependent and influenced by posttranslational regulation (75-77). DDX5 and -17 are closely related and part of multiprotein complexes. One function of these two proteins is oestrogen receptor-α co-activation, which may not play an important role in TNBC; however, the co-activation of p53, RNA metabolism and processing may be of importance (75,78,79). Additionally, DDX proteins are involved in the processing of miRNAs (75). The interplay between DDX proteins and miRNA dysregulation may be an important feature of TNBC. In the present study, it was found that DDX5 and -17 mRNA expression in TNBC cells was influenced by chemotherapy treatment. The mRNA and protein downregulation observed supports results from previous studies that described an inhibition of cancer cell proliferation when simultaneously depleting DDX5 and -17 (80,81). The findings of the present study suggested an oncogenic function for these DDX proteins in TNBC cells. However, the mRNA upregulation of DDX5 triggered by epirubicin is of interest. As protein levels of DDX5 were reduced after epirubicin treatment, a feedback loop may be involved.
The widely studied miR-17~92 cluster exerts oncogenic functions, with its members upregulated in several types of cancer, including TNBC (82-84). Furthermore, the expression of miR-17~92 varies in different BC subtypes (85). In the present study, it was observed that miRNAs of the miR-17~92 cluster (miR-17, -18a and -19b), as well as miR-15a and -30b, were regulated in a similar manner; DDX17 mRNA displayed a similar expression profile to the miR-17~92 cluster. The present study demonstrates that the expression levels of miR-17~92 cluster miRNAs, and miR-15a and -30b, were reduced by epirubicin, gemcitabine and paclitaxel treatment in the in vitro cell models. It was also found that the miR-17~92 cluster miRNA expression levels were reduced in serum and urine samples from patients with TNBC compared with healthy controls.
piR-36743 was of interest due to the chemotherapy-driven expression alterations found in the in vitro models. piR-36743 was upregulated, while most of the other investigated ncRNAs experienced chemotherapy-dependent downregulation. The functional background of piR-36743 remains poorly understood. Hashim et al (86) found that piR-36743 (also known as DQ598677) was upregulated in BC biopsies compared with healthy breast tissue.
The present in vivo study found a clear liquid biopsy biomarker-based distinction between patients with TNBC and healthy controls. Serum and urine samples allowed patients with TNBC to be distinguished from healthy women. ncRNA dysregulation in TNBC was directed almost uniformly in both serum and urine specimens.
The comparison of microvesicular ncRNA expression levels in the urine and serum of patients with TNBC prior to NACT and immediately prior to the third cycle of therapy provided preliminary results. Urinary ncRNA expression showed a general increase from t0 to t1, except for let-7a/e and miR-17. This observation may be explained by the general effects of treatment or other factors. In serum, a different picture was observed. To evaluate the ncRNA panel for their liquid biopsy biomarker potential in TNBC therapy response evaluation, patients undergoing NACT were divided into a (early) responder and non-responder group. However, it is emphasised that due to a small sample size (8 patients with TNBC), the present study only provided a preliminary result in this respect. As aforementioned, in an attempt to visualize ncRNA differences in the serum of patients with TNBC from t0 to t1 of NACT, no clear pattern could be distinguished between the early responders and non-responders, and no association with the achievement of a pCR could be drawn. However, larger cohorts may shed further light on this issue and should be analysed to further investigate this question. It is hoped that the investigated ncRNA types of the present study may act as potential minimally-invasive biomarkers to indicate therapy response in TNBC. This hypothesis is supported as different ncRNA regulation was identified in the serum of patients with TNBC who achieved a cCR during NACT compared with those who did not. Therefore, the involvement of these ncRNAs in TNBC is likely. However, the small patient number in this pilot study is a limitation. Only studies using bigger cohorts can clarify if this ncRNA panel may serve as a liquid biopsy tool in predicting therapy response.
To date, few studies have been published connecting serum-based ncRNAs to the response of patients with BC to NACT. Liu et al (87) reported reduced miR-125b expression levels in responding vs. non-responding patients with BC before, during and after NACT. Additionally, miR-21 expression decreased throughout NACT in responding patients. Furthermore, Gu et al (88) reported that lower miR-451 expression in serum was associated with the resistance of patients with BC to NACT. Supporting this, Al-Khanbashi et al (89) reported an association between elevated serum levels of miR-451, and improved clinical and pathological response to NACT of locally advanced BC.
In the present study, expression analysis was conducted using relative quantification. As each experimental setup requires the evaluation of the most suitable reference genes, an individual normalisation for each matrix was conducted. As aforementioned, intracellular ncRNA expression was normalised to the geometric mean of RNU44 and -48, while microvesicular ncRNA in conditioned cell culture media was normalised to the global mean. In serum, miR-26b and -191 served as endogenous controls and in urine, miR-16 and -26b were used as endogenous controls. The most suitable endogenous controls were assessed using the BestKeeper software tool (55). However, different normalisation strategies can complicate the comparability of results. Nevertheless, it is essential to choose an appropriate normalisation approach according to each experiment.
Different sample matrices can represent different physiological and pathophysiological processes. Precise TNBC-related ncRNA expression profiles can be obtained from tumour tissue; however, sample acquisition remains invasive. TNBC cells can serve as a model for tumour tissue. In the case of liquid biopsies, specimens derived from the blood (for example, serum), an altered ncRNA profile could result from tumour-derived and secreted ncRNAs. However, dysregulated ncRNAs may originate from other malignant or non-malignant cells, or may originate from an immune response to the tumour. The latter may indirectly indicate the existence of a malignancy. Immune cells constitutively release exosomes, which can carry ncRNAs (90). Urine-based ncRNAs are further away still from the tumour site. It is assumed that urine-based altered ncRNA levels originate directly from tumour tissue and undergo glomerular filtration by the kidneys after secretion into the circulation. In this manner, ncRNAs may be found in urine. Similarly to serum specimens, urine-based ncRNAs may also be secreted by immune cells. Furthermore, ncRNAs can enter urine from the cells of the urinary tract (91).
The present study detected chemotherapy-driven expression alterations of a (TN)BC-related panel of ncRNAs in vitro. Furthermore, the vast majority of this panel was detectable in serum and urine specimens, and could be used to discriminate between patients with TNBC and healthy controls. Results regarding the eligibility of this panel in predicting therapy response in vivo remain preliminary due to the small sample size. The liquid biopsy potential of the investigated ncRNAs in the evaluation of the response of patients with TNBC to NACT should be further investigated in larger cohorts.
Supplementary Data
Acknowledgments
Not applicable.
Abbreviations
- BC
breast cancer
- TNBC
triple negative breast cancer
- NACT
neoadjuvant chemotherapy
- ACT
adjuvant chemotherapy
- pCR
pathological complete response
- US
ultrasound
- MRI
magnetic resonance imaging
- ncRNA
non-coding RNA, miRNA/miR, micro RNA
- piRNA/piR
PIWI-interacting RNA
- RT
reverse transcription
- qPCR
quantitative polymerase chain reaction
- Cq
quantification cycle
- cCR
complete clinical response
- CI
confidence interval
- cPR
clinical partial response
Funding
The article processing charge was funded by the open access publication fund of the Albert Ludwigs University Freiburg. No funding was received for the study.
Availability of data and materials
The datasets analysed during the current study are available from the corresponding author on reasonable request.
Authors' contributions
AR, TE, MH and MJ substantially developed the study design and experimental setup. AR, MJ, TE, KB, DW, JA, MM and SG contributed to sample collection and processing. AR, MJ, CN, and SG performed miRNA quantification analysis and data collection. GR conducted statistical analyses in extenso. AR, MH, KB, GR, MJ, DW, JA, MM and TE drafted the manuscript or revised it critically. All authors have read and approved the final manuscript.
Ethics approval and consent for publication
The present study was approved by the institutional ethical review board of the University of Freiburg (permit no. 607/16) and written informed consent was obtained from each participant.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
References
- 1.World Health Organization IAfRoC: Cancer Today. 2018 https://gco.iarc.fr/today/home. Accessed April 4, 2019.
- 2.DeSantis C, Ma J, Bryan L, Jemal A. Breast cancer statistics, 2013. CA Cancer J Clin. 2014;64:52–62. doi: 10.3322/caac.21203. [DOI] [PubMed] [Google Scholar]
- 3.Early Breast Cancer Trialists' Collaborative Group (EBCTCG) Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: An overview of the randomised trials. Lancet. 2005;365:1687–1717. doi: 10.1016/S0140-6736(05)66544-0. [DOI] [PubMed] [Google Scholar]
- 4.Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, He X, Perou CM. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010;12:R68. doi: 10.1186/bcr2635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Pérou CM, Sprlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–752. doi: 10.1038/35021093. [DOI] [PubMed] [Google Scholar]
- 6.Sprlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA. 2001;98:10869–10874. doi: 10.1073/pnas.191367098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Coates AS, Winer EP, Goldhirsch A, Gelber RD, Gnant M, Piccart-Gebhart M, Thürlimann B, Senn HJ, Panel Members. Tailoring therapies-improving the management of early breast cancer: St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 20l5. Ann Oncol. 2015;26:1533–1546. doi: 10.1093/annonc/mdv221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.American Cancer Society . Breast CancerFacts &Figures. American Cancer Society; Atlanta, GA: 2017-2018. 2017. https://www.cancer.org/content/dam/cancer-org/research/cancer-facts-and-stati-stics/breast-cancer-facts-and-figures/breast-cancer-facts-and-figures-2017-2018.pdf. Accessed April 02, 2019. [Google Scholar]
- 9.Badve S, Dabbs DJ, Schnitt SJ, Baehner FL, Decker T, Eusebi V, Fox SB, Ichihara S, Jacquemier J, Lakhani SR, et al. Basal-like and triple-negative breast cancers: A critical review with an emphasis on the implications for pathologists and oncologists. Mod Pathol. 2011;24:157–167. doi: 10.1038/modpathol.2010.200. [DOI] [PubMed] [Google Scholar]
- 10.Plasilova ML, Hayse B, Killelea BK, Horowitz NR, Chagpar AB, Lannin DR. Features of triple-negative breast cancer: Analysis of 38,813 cases from the national cancer database. Medicine (Baltimore) 2016;95:e4614. doi: 10.1097/MD.0000000000004614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Yeh J, Chun J, Schwartz S, Wang A, Kern E, Guth AA, Axelrod D, Shapiro R, Schnabel F. Clinical characteristics in patients with triple negative breast cancer. Int J Breast Cancer. 2017;2017:1796145. doi: 10.1155/2017/1796145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Anders CK, Carey LA. Biology, metastatic patterns, and treatment of patients with triple-negative breast cancer. Clin Breast Cancer. 2009;9(Suppl 2):S73–S81. doi: 10.3816/CBC.2009.s.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011;121:2750–2767. doi: 10.1172/JCI45014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Toft DJ, Cryns VL. Minireview: Basal-like breast cancer: From molecular profiles to targeted therapies. Mol Endocrinol. 2011;25:199–211. doi: 10.1210/me.2010-0164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bertucci F, Finetti P, Cervera N, Esterni B, Hermitte F, Viens P, Birnbaum D. How basal are triple-negative breast cancers? Int J Cancer. 2008;123:236–240. doi: 10.1002/ijc.23518. [DOI] [PubMed] [Google Scholar]
- 16.Harbeck N, Gnant M. Breast cancer. Lancet. 2017;389:1134–1150. doi: 10.1016/S0140-6736(16)31891-8. [DOI] [PubMed] [Google Scholar]
- 17.Mieog JS, van der Hage JA, van de Velde CJ. Neoadjuvant chemotherapy for operable breast cancer. Br J Surg. 2007;94:1189–1200. doi: 10.1002/bjs.5894. [DOI] [PubMed] [Google Scholar]
- 18.Mieog JS, van der Hage JA, van de Velde CJ. Preoperative chemotherapy for women with operable breast cancer. Cochrane Database Syst Rev. 2007:CD005002. doi: 10.1002/14651858.CD005002.pub2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fisher B, Bryant J, Wolmark N, Mamounas E, Brown A, Fisher ER, Wickerham DL, Begovic M, DeCillis A, Robidoux A, et al. Effect of preoperative chemotherapy on the outcome of women with operable breast cancer. J Clin Oncol. 1998;16:2672–2685. doi: 10.1200/JCO.1998.16.8.2672. [DOI] [PubMed] [Google Scholar]
- 20.Rastogi P, Anderson SJ, Bear HD, Geyer CE, Kahlenberg MS, Robidoux A, Margolese RG, Hoehn JL, Vogel VG, Dakhil SR, et al. Preoperative chemotherapy: Updates of National surgical adjuvant breast and bowel project protocols B-18 and B-27. J Clin Oncol. 2008;26:778–785. doi: 10.1200/JCO.2007.15.0235. [DOI] [PubMed] [Google Scholar]
- 21.van der Hage JA, van de Velde CJ, Julien JP, Tubiana-Hulin M, Vandervelden C, Duchateau L. Preoperative chemotherapy in primary operable breast cancer: Results from the European organization for research and treatment of cancer trial 10902. J Clin Oncol. 2001;19:4224–4237. doi: 10.1200/JCO.2001.19.22.4224. [DOI] [PubMed] [Google Scholar]
- 22.von Minckwitz G, Untch M, Blohmer JU, Costa SD, Eidtmann H, Fasching PA, Gerber B, Eiermann W, Hilfrich J, Huober J, et al. Definition and impact of pathologic complete response on prognosis after neoadjuvant chemotherapy in various intrinsic breast cancer subtypes. J Clin Oncol. 2012;30:1796–1804. doi: 10.1200/JCO.2011.38.8595. [DOI] [PubMed] [Google Scholar]
- 23.Berruti A, Amoroso V, Gallo F, Bertaglia V, Simoncini E, Pedersini R, Ferrari L, Bottini A, Bruzzi P, Sormani MP. Pathologic complete response as a potential surrogate for the clinical outcome in patients with breast cancer after neoadjuvant therapy: A meta-regression of 29 randomized prospective studies. J Clin Oncol. 2014;32:3883–3891. doi: 10.1200/JCO.2014.55.2836. [DOI] [PubMed] [Google Scholar]
- 24.Cortazar P, Zhang L, Untch M, Mehta K, Costantino JP, Wolmark N, Bonnefoi H, Cameron D, Gianni L, Valagussa P, et al. Pathological complete response and long-term clinical benefit in breast cancer: The CTNeoBC pooled analysis. Lancet. 2014;384:164–172. doi: 10.1016/S0140-6736(13)62422-8. [DOI] [PubMed] [Google Scholar]
- 25.Loibl S, Volz C, Mau C, Blohmer JU, Costa SD, Eidtmann H, Fasching PA, Gerber B, Hanusch C, Jackisch C, et al. Response and prognosis after neoadjuvant chemotherapy in 1,051 patients with infiltrating lobular breast carcinoma. Breast Cancer Res Treat. 2014;144:153–162. doi: 10.1007/s10549-014-2861-6. [DOI] [PubMed] [Google Scholar]
- 26.von Minckwitz G, Untch M, Nüesch E, Loibl S, Kaufmann M, Kümmel S, Fasching PA, Eiermann W, Blohmer JU, Costa SD, et al. Impact of treatment characteristics on response of different breast cancer phenotypes: Pooled analysis of the German neo-adjuvant chemotherapy trials. Breast Cancer Res Treat. 2011;125:145–156. doi: 10.1007/s10549-010-1228-x. [DOI] [PubMed] [Google Scholar]
- 27.von Minckwitz G, Blohmer JU, Costa SD, Denkert C, Eidtmann H, Eiermann W, Gerber B, Hanusch C, Hilfrich J, Huober J, et al. Response-guided neoadjuvant chemotherapy for breast cancer. J Clin Oncol. 2013;31:3623–3630. doi: 10.1200/JCO.2012.45.0940. [DOI] [PubMed] [Google Scholar]
- 28.Diagnosis and treatment of patients with primary and metastatic breast cancer. www.ago-online.de. Accessed April 2, 2019.
- 29.Marinovich ML, Houssami N, Macaskill P, von Minckwitz G, Blohmer JU, Irwig L. Accuracy of ultrasound for predicting pathologic response during neoadjuvant therapy for breast cancer. Int J Cancer. 2015;136:2730–2737. doi: 10.1002/ijc.29323. [DOI] [PubMed] [Google Scholar]
- 30.Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, Dancey J, Arbuck S, Gwyther S, Mooney M, et al. New response evaluation criteria in solid tumours: Revised RECIST guideline (version 1.1) Eur J Cancer. 2009;45:228–247. doi: 10.1016/j.ejca.2008.10.026. [DOI] [PubMed] [Google Scholar]
- 31.Chagpar AB, Middleton LP, Sahin AA, Dempsey P, Buzdar AU, Mirza AN, Ames FC, Babiera GV, Feig BW, Hunt KK, et al. Accuracy of physical examination, ultrasonography, and mammography in predicting residual pathologic tumor size in patients treated with neoadjuvant chemotherapy. Ann Surg. 2006;243:257–264. doi: 10.1097/01.sla.0000197714.14318.6f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sadeghi-Naini A, Sannachi L, Tadayyon H, Tran WT, Slodkowska E, Trudeau M, Gandhi S, Pritchard K, Kolios MC, Czarnota GJ. Chemotherapy-response monitoring of breast cancer patients using quantitative ultrasound-based Intra-tumour heterogeneities. Sci Rep. 2017;7:10352. doi: 10.1038/s41598-017-09678-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Amoroso V, Generali D, Buchholz T, Cristofanilli M, Pedersini R, Curigliano G, Daidone MG, Di Cosimo S, Dowsett M, Fox S, et al. International expert consensus on primary systemic therapy in the management of early breast cancer: Highlights of the fifth symposium on primary systemic therapy in the management of operable breast cancer, Cremona, Italy 2013. J Natl Cancer Inst Monogr. 2015;2015:90–96. doi: 10.1093/jncimonographs/lgv023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liedtke C, Mazouni C, Hess KR, Andrè F, Tordai A, Mejia JA, Symmans WF, Gonzalez-Angulo AM, Hennessy B, Green M, et al. Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol. 2008;26:1275–1281. doi: 10.1200/JCO.2007.14.4147. [DOI] [PubMed] [Google Scholar]
- 35.von Minckwitz G, Blohmer JU, Raab G, Lohr A, Gerber B, Heinrich G, Eidtmann H, Kaufmann M, Hilfrich J, Jackisch C, et al. In vivo chemosensitivity-adapted preoperative chemotherapy in patients with early-stage breast cancer: The GEPARTRIO pilot study. Ann Oncol. 2005;16:56–63. doi: 10.1093/annonc/mdi001. [DOI] [PubMed] [Google Scholar]
- 36.Hombach S, Kretz M. Non-coding RNAs: Classification, biology and functioning. Adv Exp Med Biol. 2016;937:3–17. doi: 10.1007/978-3-319-42059-2_1. [DOI] [PubMed] [Google Scholar]
- 37.Brosnan CA, Voinnet O. The long and the short of noncoding RNAs. Curr Opin Cell Biol. 2009;21:416–425. doi: 10.1016/j.ceb.2009.04.001. [DOI] [PubMed] [Google Scholar]
- 38.Anastasiadou E, Jacob LS, Slack FJ. Non-coding RNA networks in cancer. Nat Rev Cancer. 2018;18:5–18. doi: 10.1038/nrc.2017.99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Izzotti A, Carozzo S, Pulliero A, Zhabayeva D, Ravetti JL, Bersimbaev R. Extracellular MicroRNA in liquid biopsy: Applicability in cancer diagnosis and prevention. Am J Cancer Res. 2016;6:1461–1493. [PMC free article] [PubMed] [Google Scholar]
- 40.Qi P, Zhou XY, Du X. Circulating long non-coding RNAs in cancer: Current status and future perspectives. Mol Cancer. 2016;15:39. doi: 10.1186/s12943-016-0524-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Yang X, Cheng Y, Lu Q, Wei J, Yang H, Gu M. Detection of stably expressed piRNAs in human blood. Int J Clin Exp Med. 2015;8:13353–13358. [PMC free article] [PubMed] [Google Scholar]
- 42.Fanale D, Castiglia M, Bazan V, Russo A. Involvement of Non-coding RNAs in Chemo- and radioresistance of colorectal cancer. Adv Exp Med Biol. 2016;937:207–228. doi: 10.1007/978-3-319-42059-2_11. [DOI] [PubMed] [Google Scholar]
- 43.Magee P, Shi L, Garofalo M. Role of microRNAs in chemo- resistance. Ann Transl Med. 2015;3:332. doi: 10.3978/j.issn.2305-5839.2015.11.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wang YW, Zhang W, Ma R. Bioinformatic identification of chemoresistance-associated microRNAs in breast cancer based on microarray data. Oncol Rep. 2018;39:1003–1010. doi: 10.3892/or.2018.6205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Malhotra A, Jain M, Prakash H, Vasquez KM, Jain A. The regulatory roles of long non-coding RNAs in the development of chemoresistance in breast cancer. Oncotarget. 2017;8:110671–110684. doi: 10.18632/oncotarget.22577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kao J, Salari K, Bocanegra M, Choi YL, Girard L, Gandhi J, Kwei KA, Hernandez-Boussard T, Wang P, Gazdar AF, et al. Molecular profiling of breast cancer cell lines defines relevant tumor models and provides a resource for cancer gene discovery. PLoS One. 2009;4:e6146. doi: 10.1371/journal.pone.0006146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thürlimann B, Senn HJ. Panel members: Personalizing the treatment of women with early breast cancer: Highlights of the St Gallen international expert consensus on the primary therapy of Early breast cancer 2013. Ann Oncol. 2013;24:2206–2223. doi: 10.1093/annonc/mdt303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: Experience from a large study with long-term follow-up. Histopathology. 1991;19:403–410. doi: 10.1111/j.1365-2559.1991.tb00229.x. [DOI] [PubMed] [Google Scholar]
- 49.Busk PK. A tool for design of primers for microRNA-specific quantitative RT-qPCR. BMC Bioinformatics. 2014;15:29. doi: 10.1186/1471-2105-15-29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 51.Torres A, Torres K, Wdowiak P, Paszkowski T, Maciejewski R. Selection and validation of endogenous controls for microRNA expression studies in endometrioid endometrial cancer tissues. Gynecol Oncol. 2013;130:588–594. doi: 10.1016/j.ygyno.2013.06.026. [DOI] [PubMed] [Google Scholar]
- 52.Bockmeyer CL, Säuberlich K, Wittig J, Eßer M, Roeder SS, Vester U, Hoyer PF, Agustian PA, Zeuschner P, Amann K, et al. Comparison of different normalization strategies for the analysis of glomerular microRNAs in IgA nephropathy. Sci Rep. 2016;6:31992. doi: 10.1038/srep31992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chen L, Jin Y, Wang L, Sun F, Yang X, Shi M, Zhan C, Shi Y, Wang Q. Identification of reference genes and miRNAs for qRT-PCR in human esophageal squamous cell carcinoma. Med Oncol. 2017;34:2. doi: 10.1007/s12032-016-0860-7. [DOI] [PubMed] [Google Scholar]
- 54.Masè M, Grasso M, Avogaro L, D'Amato E, Tessarolo F, Graffigna A, Denti MA, Ravelli F. Selection of reference genes is critical for miRNA expression analysis in human cardiac tissue. A focus on atrial fibrillation. Sci Rep. 2017;7:41127. doi: 10.1038/srep41127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper-Excel-based tool using pair-wise correlations. Biotechnol Lett. 2004;26:509–515. doi: 10.1023/B:BILE.0000019559.84305.47. [DOI] [PubMed] [Google Scholar]
- 56.Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, Vandesompele J. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 2009;10:R64. doi: 10.1186/gb-2009-10-6-r64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.D'Haene B, Mestdagh P, Hellemans J, Vandesompele J. miRNA expression profiling: From reference genes to global mean normalization. Methods Mol Biol. 2012;822:261–272. doi: 10.1007/978-1-61779-427-8_18. [DOI] [PubMed] [Google Scholar]
- 58.Schwarzenbach H, da Silva AM, Calin G, Pantel K. Data Normalization strategies for MicroRNA quantification. Clin Chem. 2015;61:1333–1342. doi: 10.1373/clinchem.2015.239459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Erbes T, Hirschfeld M, Rücker G, Jaeger M, Boas J, Iborra S, Mayer S, Gitsch G, Stickeler E. Feasibility of urinary microRNA detection in breast cancer patients and its potential as an innovative non-invasive biomarker. BMC Cancer. 2015;15:193. doi: 10.1186/s12885-015-1190-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Marabita F, de Candia P, Torri A, Tegnér J, Abrignani S, Rossi RL. Normalization of circulating microRNA expression data obtained by quantitative real-time RT-PCR. Brief Bioinform. 2016;17:204–212. doi: 10.1093/bib/bbv056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Song W, Zhang WH, Zhang H, Li Y, Zhang Y, Yin W, Yang Q. Validation of housekeeping genes for the normalization of RT-qPCR expression studies in oral squamous cell carcinoma cell line treated by 5 kinds of chemotherapy drugs. Cell Mol Biol (Noisy-le-Grand) 2016;62:29–34. doi: 10.14715/cmb/2016.62.13.6. [DOI] [PubMed] [Google Scholar]
- 62.Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC. Measurement of protein using bicinchoninic acid. Anal Biochem. 1985;150:76–85. doi: 10.1016/0003-2697(85)90442-7. [DOI] [PubMed] [Google Scholar]
- 63.Schägger H, von Jagow G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem. 1987;166:368–379. doi: 10.1016/0003-2697(87)90587-2. [DOI] [PubMed] [Google Scholar]
- 64.R Core Team . R: A language and environment for statistical computing. R Foundation for Statistical Computing; Vienna: 2018. [Google Scholar]
- 65.Gao D, Qi X, Zhang X, Fang K, Guo Z, Li L. hsa_ circRNA_0006528 as a competing endogenous RNA promotes human breast cancer progression by sponging miR-75p and activating the MAPK/ERK signalling pathway. Mol Carcinog. 2019;58:554–564. doi: 10.1002/mc.22950. [DOI] [PubMed] [Google Scholar]
- 66.Gao D, Zhang X, Liu B, Meng D, Fang K, Guo Z, Li L. Screening circular RNA related to chemotherapeutic resistance in breast cancer. Epigenomics. 2017;9:1175–1188. doi: 10.2217/epi-2017-0055. [DOI] [PubMed] [Google Scholar]
- 67.Uhr K, Sieuwerts AM, de Weerd V, Smid M, Hammerl D, Foekens JA, Martens JWM. Association of microRNA-7 and its binding partner CDR1-AS with the prognosis and prediction of 1st-line tamoxifen therapy in breast cancer. Sci Rep. 2018;8:9657. doi: 10.1038/s41598-018-27987-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Garcia-Vazquez R, Ruiz-García E, Meneses García A, Astudillo-de la Vega H, Lara-Medina F, Alvarado-Miranda A, Mal donado-Martínez H, Gonzalez-Barrios JA, Campos-Parra AD, Rodríguez Cuevas S, et al. A microRNA signature associated with pathological complete response to novel neoadjuvant therapy regimen in triple-negative breast cancer. Tumour Biol. 2017;39:1010428317702899. doi: 10.1177/1010428317702899. [DOI] [PubMed] [Google Scholar]
- 69.Unger FT, Klasen HA, Tchartchian G, de Wilde RL, Witte I. DNA damage induced by cis- and carboplatin as indicator for in vitro sensitivity of ovarian carcinoma cells. BMC Cancer. 2009;9:359. doi: 10.1186/1471-2407-9-359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Su WC, Chang SL, Chen TY, Chen JS, Tsao CJ. Comparison of in vitro growth-inhibitory activity of carboplatin and cisplatin on leukemic cells and hematopoietic progenitors: The myelosuppressive activity of carboplatin may be greater than its antileukemic effect. Jpn. J Clin Oncol. 2000;30:562–567. doi: 10.1093/jjco/hyd137. [DOI] [PubMed] [Google Scholar]
- 71.Kuittinen T, Rovio P, Staff S, Luukkaala T, Kallioniemi A, Grénman S, Laurila M, Maenpaa J. Paclitaxel, carboplatin and 1,25D3 inhibit proliferation of endometrial cancer cells in vitro. Anticancer Res. 2017;37:6575–6581. doi: 10.21873/anticanres.12114. [DOI] [PubMed] [Google Scholar]
- 72.He Q, Liang CH, Lippard SJ. Steroid hormones induce HMG1 overexpression and sensitize breast cancer cells to cisplatin and carboplatin. Proc Natl Acad Sci USA. 2000;97:5768–5772. doi: 10.1073/pnas.100108697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Diehl JA, Zindy F, Sherr CJ. Inhibition of cyclin D1 phosphorylation on threonine-286 prevents its rapid degradation via the ubiquitin-proteasome pathway. Genes Dev. 1997;11:957–972. doi: 10.1101/gad.11.8.957. [DOI] [PubMed] [Google Scholar]
- 74.Luo H, Zhang J, Dastvan F, Yanagawa B, Reidy MA, Zhang HM, Yang D, Wilson JE, McManus BM. Ubiquitin-dependent proteolysis of cyclin D1 is associated with coxsackievirus-induced cell growth arrest. J Virol. 2003;77:1–9. doi: 10.1128/JVI.77.1.1-9.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Fuller-Pace FV. DEAD box RNA helicase functions in cancer. RNA Biol. 2013;10:121–132. doi: 10.4161/rna.23312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Mazurek A, Luo W, Krasnitz A, Hicks J, Powers RS, Stillman B. DDX5 regulates DNA replication and is required for cell proliferation in a subset of breast cancer cells. Cancer Discov. 2012;2:812–825. doi: 10.1158/2159-8290.CD-12-0116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Wang D, Huang J, Hu Z. RNA helicase DDX5 regulates microRNA expression and contributes to cytoskeletal reorganization in basal breast cancer cells. Mol Cell Proteomics. 2012;11:M111.011932. doi: 10.1074/mcp.M111.011932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Bates GJ, Nicol SM, Wilson BJ, Jacobs AM, Bourdon JC, Wardrop J, Gregory DJ, Lane DP, Perkins ND, Fuller-Pace FV. The DEAD box protein p68: A novel transcriptional coactivator of the p53 tumour suppressor. EMBO J. 2005;24:543–553. doi: 10.1038/sj.emboj.7600550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wortham NC, Ahamed E, Nicol SM, Thomas RS, Periyasamy M, Jiang J, Ochocka AM, Shousha S, Huson L, Bray SE, et al. The DEAD-box protein p72 regulates ERalpha-/oestrogen-dependent transcription and cell growth, and is associated with improved survival in ERalpha-positive breast cancer. Oncogene. 2009;28:4053–4064. doi: 10.1038/onc.2009.261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Shin S, Rossow KL, Grande JP, Janknecht R. Involvement of RNA helicases p68 and p72 in colon cancer. Cancer Res. 2007;67:7572–7578. doi: 10.1158/0008-5472.CAN-06-4652. [DOI] [PubMed] [Google Scholar]
- 81.Jalal C, Uhlmann-Schiffler H, Stahl H. Redundant role of DEAD box proteins p68 (Ddx5) and p72/p82 (Ddx17) in ribosome biogenesis and cell proliferation. Nucleic Acids Res. 2007;35:3590–3601. doi: 10.1093/nar/gkm058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Concepcion CP, Bonetti C, Ventura A. The microRNA-17-92family of microRNA clusters in development and disease. Cancer J. 2012;18:262–267. doi: 10.1097/PPO.0b013e318258b60a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Mogilyansky E, Rigoutsos I. The miR-17/92 cluster: A comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ. 2013;20:1603–1614. doi: 10.1038/cdd.2013.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Farazi TA, Horlings HM, Ten Hoeve JJ, Mihailovic A, Halfwerk H, Morozov P, Brown M, Hafner M, Reyal F, van Kouwenhove M, et al. MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res. 2011;71:4443–4453. doi: 10.1158/0008-5472.CAN-11-0608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Calvano Filho CM, Calvano-Mendes DC, Carvalho KC, Maciel GA, Ricci MD, Torres AP, Filassi JR, Baracat EC. Triple-negative and luminal A breast tumors: Differential expression of miR-18a-5p, and miR-17-5p, and miR-20a-5p. Tumour Biol. 2014;35:7733–7741. doi: 10.1007/s13277-014-2025-7. [DOI] [PubMed] [Google Scholar]
- 86.Hashim A, Rizzo F, Marchese G, Ravo M, Tarallo R, Nassa G, Giurato G, Santamaria G, Cordella A, Cantarella C, Weisz A. RNA sequencing identifies specific PIWI-interacting small non-coding RNA expression patterns in breast cancer. Oncotarget. 2014;5:9901–9910. doi: 10.18632/oncotarget.2476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Liu B, Su F, Chen M, Li Y, Qi X, Xiao J, Li X, Liu X, Liang W, Zhang Y, Zhang J. Serum miR-21 and miR-125b as markers predicting neoadjuvant chemotherapy response and prognosis in stage II/III breast cancer. Hum Pathol. 2017;64:44–52. doi: 10.1016/j.humpath.2017.03.016. [DOI] [PubMed] [Google Scholar]
- 88.Gu X, Xue JQ, Han SJ, Qian SY, Zhang WH. Circulating microRNA-451 as a predictor of resistance to neoadjuvant chemotherapy in breast cancer. Cancer Biomark. 2016;16:395–403. doi: 10.3233/CBM-160578. [DOI] [PubMed] [Google Scholar]
- 89.Al-Khanbashi M, Caramuta S, Alajmi AM, Al-Haddabi I, Al-Riyami M, Lui WO, Al-Moundhri MS. Tissue and Serum miRNA profile in locally advanced breast cancer (LABC) in response to Neo-adjuvant chemotherapy (NAC) treatment. PLoS One. 2016;11:e0152032. doi: 10.1371/journal.pone.0152032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Edgar JR. Q&A: What are exosomes, exactly? BMC Biol. 2016;14:46. doi: 10.1186/s12915-016-0268-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Ben-Dov IZ, Whalen VM, Goilav B, Max KE, Tuschl T. Cell and microvesicle urine microRNA deep sequencing profiles from healthy individuals: Observations with potential impact on biomarker studies. PLoS One. 2016;11:e0147249. doi: 10.1371/journal.pone.0147249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Mansoori B, Mohammadi A, Shirjang S, Baghbani E, Baradaran B. Micro RNA 34a and Let-7a expression in human breast cancers is associated with apoptotic expression genes. Asian Pac J Cancer Prev. 2016;17:1887–1890. doi: 10.7314/APJCP.2016.17.4.1887. [DOI] [PubMed] [Google Scholar]
- 93.Liu K, Zhang C, Li T, Ding Y, Tu T, Zhou F, Qi W, Chen H, Sun X. Let-7a inhibits growth and migration of breast cancer cells by targeting HMGA1. Int J Oncol. 2015;46:2526–2534. doi: 10.3892/ijo.2015.2949. [DOI] [PubMed] [Google Scholar]
- 94.Marques MM, Evangelista AF, Macedo T, Vieira RADC, Scapulatempo-Neto C, Reis RM, Carvalho AL, Silva IDCGD. Expression of tumor suppressors miR-195 and let-7a as potential biomarkers of invasive breast cancer. Clinics (Sao Paulo) 2018;73:e184. doi: 10.6061/clinics/2018/e184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Huang SK, Luo Q, Peng H, Li J, Zhao M, Wang J, Gu YY, Li Y, Yuan P, Zhao GH, Huang CZ. A Panel of serum noncoding RNAs for the diagnosis and monitoring of response to therapy in patients with breast cancer. Med Sci Monit. 2018;24:2476–2488. doi: 10.12659/MSM.909453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Khalighfard S, Alizadeh AM, Irani S, Omranipour R. Plasma miR-21, miR-155, miR-10b, and Let-7a as the potential biomarkers for the monitoring of breast cancer patients. Sci Rep. 2018;8:17981. doi: 10.1038/s41598-018-36321-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Mi Y, Liu F, Liang X, Liu S, Huang X, Sang M, Geng C. Tumor suppressor let-7a inhibits breast cancer cell proliferation, migration and invasion by targeting MAGE-A1. Neoplasma. 2019;66:54–62. doi: 10.4149/neo_2018_180302N146. [DOI] [PubMed] [Google Scholar]
- 98.Guo Q, Wen R, Shao B, Li Y, Jin X, Deng H, Wu J, Su F, Yu F. Combined Let-7a and H19 signature: A prognostic index of progression-free survival in primary breast cancer patients. J Breast Cancer. 2018;21:142–149. doi: 10.4048/jbc.2018.21.2.142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Lehmann TP, Korski K, Gryczka R, Ibbs M, Thieleman A, Grodecka-Gazdecka S, Jagodzinski PP. Relative levels of let-7a, miR-17, miR-27b, miR-125a, miR-125b and miR-206 as potential molecular markers to evaluate grade, receptor status and molecular type in breast cancer. Mol Med Rep. 2015;12:4692–4702. doi: 10.3892/mmr.2015.4002. [DOI] [PubMed] [Google Scholar]
- 100.Thakur S, Grover RK, Gupta S, Yadav AK, Das BC. Identification of Specific miRNA signature in paired sera and tissue samples of Indian Women with Triple Negative breast cancer. PLoS One. 2016;11:e0158946. doi: 10.1371/journal.pone.0158946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Ahram M, Mustafa E, Zaza R, Abu Hammad S, Alhudhud M, Bawadi R, Zihlif M. Differential expression and androgen regulation of microRNAs and metalloprotease 13 in breast cancer cells. Cell Biol Int. 2017;41:1345–1355. doi: 10.1002/cbin.10841. [DOI] [PubMed] [Google Scholar]
- 102.Yu CC, Chen YW, Chiou GY, Tsai LL, Huang PI, Chang CY, Tseng LM, Chiou SH, Yen SH, Chou MY, et al. MicroRNA let-7a represses chemoresistance and tumourigenicity in head and neck cancer via stem-like properties ablation. Oral Oncol. 2011;47:202–210. doi: 10.1016/j.oraloncology.2010.12.001. [DOI] [PubMed] [Google Scholar]
- 103.Qiu L, Zhang GF, Yu L, Wang HY, Jia XJ, Wang TJ. Novel oncogenic and chemoresistance-inducing functions of resistin in ovarian cancer cells require miRNAs-mediated induction of epithelial-to-mesenchymal transition. Sci Rep. 2018;8:12522. doi: 10.1038/s41598-018-30978-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Xiao G, Wang X, Yu Y. CXCR4/Let-7a Axis regulates metastasis and chemoresistance of pancreatic cancer cells through targeting HMGA2. Cell Physiol Biochem. 2017;43:840–851. doi: 10.1159/000481610. [DOI] [PubMed] [Google Scholar]
- 105.Serguienko A, Grad I, Wennerstrpm AB, Meza-Zepeda LA, Thiede B, Stratford EW, Myklebost O, Munthe E. Metabolic reprogramming of metastatic breast cancer and melanoma by let-7a microRNA. Oncotarget. 2015;6:2451–2465. doi: 10.18632/oncotarget.3235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Tsang WP, Kwok TT. Let-7a microRNA suppresses therapeutics-induced cancer cell death by targeting caspase-3. Apoptosis. 2008;13:1215–1222. doi: 10.1007/s10495-008-0256-z. [DOI] [PubMed] [Google Scholar]
- 107.Wu J, Li S, Jia W, Deng H, Chen K, Zhu L, Yu F, Su F. Reduced let-7a is associated with chemoresistance in primary breast cancer. PLoS One. 2015;10:e0133643. doi: 10.1371/journal.pone.0133643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Bhutia YD, Hung SW, Krentz M, Patel D, Lovin D, Manoharan R, Thomson JM, Govindarajan R. Differential processing of let-7a precursors influences RRM2 expression and chemosensitivity in pancreatic cancer: Role of LIN-28 and SET oncoprotein. PLoS One. 2013;8:e53436. doi: 10.1371/journal.pone.0053436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Lu L, Schwartz P, Scarampi L, Rutherford T, Canuto EM, Yu H, Katsaros D. MicroRNA let-7a: A potential marker for selection of paclitaxel in ovarian cancer management. Gynecol Oncol. 2011;122:366–371. doi: 10.1016/j.ygyno.2011.04.033. [DOI] [PubMed] [Google Scholar]
- 110.Lv K, Liu L, Wang L, Yu J, Liu X, Cheng Y, Dong M, Teng R, Wu L, Fu P, et al. Lin28 mediates paclitaxel resistance by modulating p21, Rb and Let-7a miRNA in breast cancer cells. PLoS One. 2012;7:e40008. doi: 10.1371/journal.pone.0040008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Yin PT, Pongkulapa T, Cho HY, Han J, Pasquale NJ, Rabie H, Kim JH, Choi JW, Lee KB. Overcoming chemoresistance in cancer via combined MicroRNA therapeutics with anticancer drugs using multifunctional magnetic Core-shell nanoparticles. ACS Appl Mater Interfaces. 2018;10:26954–26963. doi: 10.1021/acsami.8b09086. [DOI] [PubMed] [Google Scholar]
- 112.Farré PL, Scalise GD, Duca RB, Dalton GN, Massillo C, Porretti J, Grana K, Gardner K, De Luca P, De Siervi A. CTBP1 and metabolic syndrome induce an mRNA and miRNA expression profile critical for breast cancer progression and metastasis. Oncotarget. 2018;9:13848–13858. doi: 10.18632/oncotarget.24486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Aure MR, Leivonen SK, Fleischer T, Zhu Q, Overgaard J, Alsner J, Tramm T, Louhimo R, Alnaes GI, Perala M, et al. Individual and combined effects of DNA methylation and copy number alterations on miRNA expression in breast tumors. Genome Biol. 2013;14:R126. doi: 10.1186/gb-2013-14-11-r126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Lv J, Xia K, Xu P, Sun E, Ma J, Gao S, Zhou Q, Zhang M, Wang F, Chen F, et al. miRNA expression patterns in chemoresistant breast cancer tissues. Biomed Pharmacother. 2014;68:935–942. doi: 10.1016/j.biopha.2014.09.011. [DOI] [PubMed] [Google Scholar]
- 115.Oztemur Islakoglu Y, Noyan S, Aydos A, Gur Dedeoglu B. Meta-microRNA biomarker signatures to classify breast cancer subtypes. OMICS. 2018;22:709–716. doi: 10.1089/omi.2018.0157. [DOI] [PubMed] [Google Scholar]
- 116.Shan Y, Liu Y, Zhao L, Liu B, Li Y, Jia L. MicroRNA-33a and let-7e inhibit human colorectal cancer progression by targeting ST8SIA1. Int J Biochem Cell Biol. 2017;90:48–58. doi: 10.1016/j.biocel.2017.07.016. [DOI] [PubMed] [Google Scholar]
- 117.Svoboda M, Sana J, Fabian P, Kocakova I, Gombosova J, Nekvindova J, Radova L, Vyzula R, Slaby O. MicroRNA expression profile associated with response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer patients. Radiat Oncol. 2012;7:195. doi: 10.1186/1748-717X-7-195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Cai J, Yang C, Yang Q, Ding H, Jia J, Guo J, Wang J, Wang Z. Deregulation of let-7e in epithelial ovarian cancer promotes the development of resistance to cisplatin. Oncogenesis. 2013;2:e75. doi: 10.1038/oncsis.2013.39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Sorrentino A, Liu CG, Addario A, Peschle C, Scambia G, Ferlini C. Role of microRNAs in drug-resistant ovarian cancer cells. Gynecol Oncol. 2008;111:478–486. doi: 10.1016/j.ygyno.2008.08.017. [DOI] [PubMed] [Google Scholar]
- 120.Xiao M, Cai J, Cai L, Jia J, Xie L, Zhu Y, Huang B, Jin D, Wang Z. Let-7e sensitizes epithelial ovarian cancer to cisplatin through repressing DNA double strand break repair. J Ovarian Res. 2017;10:24. doi: 10.1186/s13048-017-0321-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Block I, Burton M, Sprensen KP, Andersen L, Larsen MJ, Bak M, Cold S, Thomassen M, Tan Q, Kruse TA. Association of miR-548c-5p, miR-7-5p, miR-210-3p, miR-128-3p with recurrence in systemically untreated breast cancer. Oncotarget. 2018;9:9030–9042. doi: 10.18632/oncotarget.24088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Foekens JA, Sieuwerts AM, Smid M, Look MP, de Weerd V, Boersma AW, Klijn JG, Wiemer EA, Martens JW. Four miRNAs associated with aggressiveness of lymph node-negative, estrogen receptor-positive human breast cancer. Proc Natl Acad Sci USA. 2008;105:13021–13026. doi: 10.1073/pnas.0803304105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Gong C, Tan W, Chen K, You N, Zhu S, Liang G, Xie X, Li Q, Zeng Y, Ouyang N, et al. Prognostic value of a BCSC-associated MicroRNA signature in hormone receptor-positive HER2-negative breast cancer. EBioMedicine. 2016;11:199–209. doi: 10.1016/j.ebiom.2016.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Raychaudhuri M, Bronger H, Buchner T, Kiechle M, Weichert W, Avril S. MicroRNAs miR-7 and miR-340 predict response to neoadjuvant chemotherapy in breast cancer. Breast Cancer Res Treat. 2017;162:511–521. doi: 10.1007/s10549-017-4132-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Shi Y, Ye P, Long X. Differential expression profiles of the transcriptome in breast cancer cell lines revealed by next generation sequencing. Cell Physiol Biochem. 2017;44:804–816. doi: 10.1159/000485344. [DOI] [PubMed] [Google Scholar]
- 126.Cui YX, Bradbury R, Flamini V, Wu B, Jordan N, Jiang WG. MicroRNA-7 suppresses the homing and migration potential of human endothelial cells to highly metastatic human breast cancer cells. Br J Cancer. 2017;117:89–101. doi: 10.1038/bjc.2017.156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Vera-Puente O, Rodriguez-Antolin C, Salgado-Figueroa A, Michalska P, Pernia O, Reid BM, Rosas R, Garcia-Guede A, SacristAn S, Jimenez J, et al. MAFG is a potential therapeutic target to restore chemosensitivity in cisplatin-resistant cancer cells by increasing reactive oxygen species. Transl Res. 2018;200:1–17. doi: 10.1016/j.trsl.2018.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Filipits M, Nielsen TO, Rudas M, Greil R, Stoger H, Jakesz R, Bago-Horvath Z, Dietze O, Regitnig P, Gruber-Rossipal C et al. The PAM50 risk-of-recurrence score predicts risk for late distant recurrence after endocrine therapy in postmenopausal women with endocrine-responsive early breast cancer. Clin. Cancer Res. 2014;20:1298–1305. doi: 10.1158/1078-0432.CCR-13-1845. [DOI] [PubMed] [Google Scholar]
- 129.Blume CJ, Hotz-Wagenblatt A, Hüllein J, Sellner L, Jethwa A, Stolz T, Slabicki M, Lee K, Sharathchandra A, Benner A, et al. p53-dependent non-coding RNA networks in chronic lymphocytic leukemia. Leukemia. 2015;29:2015–2023. doi: 10.1038/leu.2015.119. [DOI] [PubMed] [Google Scholar]
- 130.Liu DZ, Chang B, Li XD, Zhang QH, Zou YH. MicroRNA-9 promotes the proliferation, migration, and invasion of breast cancer cells via down-regulating FOXO1. Clin Transl Oncol. 2017;19:1133–1140. doi: 10.1007/s12094-017-1650-1. [DOI] [PubMed] [Google Scholar]
- 131.Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D, Teruya-Feldstein J, Reinhardt F, Onder TT, Valastyan S, et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol. 2010;12:247–256. doi: 10.1038/ncb2024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Sporn JC, Katsuta E, Yan L, Takabe K. Expression of MicroRNA-9 is associated with overall survival in breast cancer patients. J Surg Res. 2019;233:426–435. doi: 10.1016/j.jss.2018.08.020. [DOI] [PubMed] [Google Scholar]
- 133.Orangi E, Motovali-Bashi M. Evaluation of miRNA-9 and miRNA-34a as potential biomarkers for diagnosis of breast cancer in Iranian women. Gene. 2019;687:272–279. doi: 10.1016/j.gene.2018.11.036. [DOI] [PubMed] [Google Scholar]
- 134.Naorem LD, Muthaiyan M, Venkatesan A. Identification of dysregulated miRNAs in triple negative breast cancer: A meta-analysis approach. J Cell Physiol. 2019;234:11768–11779. doi: 10.1002/jcp.27839. [DOI] [PubMed] [Google Scholar]
- 135.Kia V, Paryan M, Mortazavi Y, Biglari A, Mohammadi-Yeganeh S. Evaluation of exosomal miR-9 and miR-155 targeting PTEN and DUSP14 in highly metastatic breast cancer and their effect on low metastatic cells. J Cell Biochem. 2019;120:5666–5676. doi: 10.1002/jcb.27850. [DOI] [PubMed] [Google Scholar]
- 136.Jang MH, Kim HJ, Gwak JM, Chung YR, Park SY. Prognostic value of microRNA-9 and microRNA-155 expression in triple-negative breast cancer. Hum Pathol. 2017;68:69–78. doi: 10.1016/j.humpath.2017.08.026. [DOI] [PubMed] [Google Scholar]
- 137.Cheng CW, Yu JC, Hsieh YH, Liao WL, Shieh JC, Yao CC, Lee HJ, Chen PM, Wu PE, Shen CY. Increased cellular levels of MicroRNA-9 and MicroRNA-221 correlate with cancer stemness and predict poor outcome in human breast cancer. Cell Physiol Biochem. 2018;48:2205–2218. doi: 10.1159/000492561. [DOI] [PubMed] [Google Scholar]
- 138.D'Ippolito E, Plantamura I, Bongiovanni L, Casalini P, Baroni S, Piovan C, Orlandi R, Gualeni AV, Gloghini A, Rossini A, et al. miR-9 and miR-200 Regulate PDGFRp-mediated endothelial differentiation of tumor cells in Triple-negative breast cancer. Cancer Res. 2016;76:5562–5572. doi: 10.1158/0008-5472.CAN-16-0140. [DOI] [PubMed] [Google Scholar]
- 139.Li X, Pan Q, Wan X, Mao Y, Lu W, Xie X, Cheng X. Methylation-associated Has-miR-9 deregulation in pacli- taxel-resistant epithelial ovarian carcinoma. BMC Cancer. 2015;15:509. doi: 10.1186/s12885-015-1509-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Pezuk JA, Miller TLA, Bevilacqua JLB, de Barros ACSD, de Andrade FEM, E Macedo LFA, Aguilar V, Claro ANM, Camargo AA, Galante PAF, Reis LFL. Measuring plasma levels of three microRNAs can improve the accuracy for identification of malignant breast lesions in women with BI-RADS 4 mammography. Oncotarget. 2017;8:83940–83948. doi: 10.18632/oncotarget.20806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Kodahl AR, Lyng MB, Binder H, Cold S, Gravgaard K, Knoop AS, Ditzel HJ. Novel circulating microRNA signature as a potential non-invasive multi-marker test in ER-positive early-stage breast cancer: A case control study. Mol Oncol. 2014;8:874–883. doi: 10.1016/j.molonc.2014.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Yang L, Zhao W, Wei P, Zuo W, Zhu S. Tumor suppressor p53 induces miR-15a processing to inhibit neuronal apoptosis inhibitory protein (NAIP) in the apoptotic response DNA damage in breast cancer cell. Am J Transl Res. 2017;9:683–691. [PMC free article] [PubMed] [Google Scholar]
- 143.Patel N, Garikapati KR, Ramaiah MJ, Polavarapu KK, Bhadra U, Bhadra MP. miR-15a/miR-16 induces mitochondrial dependent apoptosis in breast cancer cells by suppressing oncogene BMI1. Life Sci. 2016;164:60–70. doi: 10.1016/j.lfs.2016.08.028. [DOI] [PubMed] [Google Scholar]
- 144.Hamdi K, Blancato J, Goerlitz D, Islam M, Neili B, Abidi A, Gat A, Ayed FB, Chivi S, Loffredo C, et al. Circulating Cell-free miRNA expression and its association with clinicopathologic features in inflammatory and non-inflammatory breast cancer. Asian Pac J Cancer Prev. 2016;17:1801–1810. doi: 10.7314/APJCP.2016.17.4.1801. [DOI] [PubMed] [Google Scholar]
- 145.Shinden Y, Akiyoshi S, Ueo H, Nambara S, Saito T, Komatsu H, Ueda M, Hirata H, Sakimura S, Uchi R, et al. Diminished expression of MiR-15a is an independent prognostic marker for breast cancer cases. Anticancer Res. 2015;35:123–127. [PubMed] [Google Scholar]
- 146.Li F, Xu Y, Deng S, Li Z, Zou D, Yi S, Sui W, Hao M, Qiu L. MicroRNA-15a/161 cluster located at chromosome 13q14 is down-regulated but displays different expression pattern and prognostic significance in multiple myeloma. Oncotarget. 2015;6:38270–38282. doi: 10.18632/oncotarget.5681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Patel N, Garikapati KR, Makani VKK, Nair AD, Vangara N, Bhadra U, Pal Bhadra M. Regulating BMI1 expression via miRNAs promote Mesenchymal to Epithelial Transition (MET) and sensitizes breast cancer cell to chemotherapeutic drug. PLoS One. 2018;13:e0190245. doi: 10.1371/journal.pone.0190245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Rodriguez-Aguayo C, Monroig PDC, Redis RS, Bayraktar E, Almeida MI, Ivan C, Fuentes-Mattei E, Rashed MH, Chavez-Reyes A, Ozpolat B, et al. Regulation of hnRNPA1 by microRNAs controls the miR-18a-K-RAS axis in chemotherapy-resistant ovarian cancer. Cell Discov. 2017;3:17029. doi: 10.1038/celldisc.2017.29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Wang L, Zhang X, Sheng L, Qiu C, Luo R. LINC00473 promotes the Taxol resistance via miR-15a in colorectal cancer. Biosci Rep. 2018;38 doi: 10.1042/BSR20180790. pii: BSR20180790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Chu J, Zhu Y, Liu Y, Sun L, Lv X, Wu Y, Hu P, Su F, Gong C, Song E, et al. E2F7 overexpression leads to tamoxifen resistance in breast cancer cells by competing with E2F1 at miR-15a/16 promoter. Oncotarget. 2015;6:31944–31957. doi: 10.18632/oncotarget.5128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Jurkovicova D, Smolkova B, Magyerkova M, Sestakova Z, Kajabova VH, Kulcsar L, Zmetakova I, Kalinkova L, Krivulcik T, Karaba M, et al. Down-regulation of traditional oncomiRs in plasma of breast cancer patients. Oncotarget. 2017;8:77369–77384. doi: 10.18632/oncotarget.20484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Zhang N, Zhang H, Liu Y, Su P, Zhang J, Wang X, Sun M, Chen B, Zhao W, Wang L, et al. SREBP1, targeted by miR-18a-5p, modulates epithelial-mesenchymal transition in breast cancer via forming a co-repressor complex with Snail and HDAC1/2. Cell Death Differ. 2019;26:843–859. doi: 10.1038/s41418-018-0158-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Yang F, Li Y, Xu L, Zhu Y, Gao H, Zhen L, Fang L. miR-17 as a diagnostic biomarker regulates cell proliferation in breast cancer. Onco Targets Ther. 2017;10:543–550. doi: 10.2147/OTT.S127723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Wang Y, Li J, Dai L, Zheng J, Yi Z, Chen L. MiR-17-5p may serve as a novel predictor for breast cancer recurrence. Cancer Biomark. 2018;22:721–726. doi: 10.3233/CBM-181228. [DOI] [PubMed] [Google Scholar]
- 155.Sueta A, Yamamoto Y, Tomiguchi M, Takeshita T, Yamamoto-Ibusuki M, Iwase H. Differential expression of exosomal miRNAs between breast cancer patients with and without recurrence. Oncotarget. 2017;8:69934–69944. doi: 10.18632/oncotarget.19482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Hesari A, Azizian M, Darabi H, Nesaei A, Hosseini SA, Salarinia R, Motaghi AA, Ghasemi F. Expression of circulating miR-17, miR-25, and miR-133 in breast cancer patients. J Cell Biochem. 2018 Nov 28; doi: 10.1002/jcb.27984. Epub ahead of print. [DOI] [PubMed] [Google Scholar]
- 157.Li H, Liu J, Chen J, Wang H, Yang L, Chen F, Fan S, Wang J, Shao B, Yin D, et al. A serum microRNA signature predicts trastuzumab benefit in HER2-positive metastatic breast cancer patients. Nat Commun. 2018;9:1614. doi: 10.1038/s41467-018-03537-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Braicu C, Raduly L, Morar-Bolba G, Cojocneanu R, Jurj A, Pop LA, Pileczki V, Ciocan C, Moldovan A, Irimie A, et al. Aberrant miRNAs expressed in HER-2 negative breast cancers patient. J Exp Cli. Cancer Res. 2018;37:257. doi: 10.1186/s13046-018-0920-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Li J, Lai Y, Ma J, Liu Y, Bi J, Zhang L, Chen L, Yao C, Lv W, Chang G, et al. miR-17-5p suppresses cell proliferation and invasion by targeting ETV1 in triple-negative breast cancer. BMC Cancer. 2017;17:745. doi: 10.1186/s12885-017-3674-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Li X, Wu B, Chen L, Ju Y, Li C, Meng S. Urokinase-type plasminogen activator receptor inhibits apoptosis in triple-negative breast cancer through miR-17/20a suppression of death receptors 4 and 5. Oncotarget. 2017;8:88645–88657. doi: 10.18632/oncotarget.20435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Cui YH, Kim H, Lee M, Yi JM, Kim RK, Uddin N, Yoo KC, Kang JH, Choi MY, Cha HJ, et al. FBXL14 abolishes breast cancer progression by targeting CDCP1 for proteasomal degradation. Oncogene. 2018;37:5794–5809. doi: 10.1038/s41388-018-0372-3. [DOI] [PubMed] [Google Scholar]
- 162.Wang ST, Liu LB, Li XM, Wang YF, Xie PJ, Li Q, Wang R, Wei Q, Kang YH, Meng R, et al. Circ-ITCH regulates triple-negative breast cancer progression through the Wnt/p-catenin pathway. Neoplasma. 2019;66:232–239. doi: 10.4149/neo_2018_180710N460. [DOI] [PubMed] [Google Scholar]
- 163.Jia J, Zhan D, Li J, Li Z, Li H, Qian J. The contrary functions of lncRNA HOTAIR/miR-17-5p/PTEN axis and Shenqifuzheng injection on chemosensitivity of gastric cancer cells. J Cell Mol Med. 2019;23:656–669. doi: 10.1111/jcmm.13970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Wu DM, Hong XW, Wang LL, Cui XF, Lu J, Chen GQ, Zheng YL. MicroRNA-17 inhibition overcomes chemoresistance and suppresses epithelial-mesenchymal transition through a DEDD-dependent mechanism in gastric cancer. Int J Biochem Cell Biol. 2018;102:59–70. doi: 10.1016/j.biocel.2018.06.007. [DOI] [PubMed] [Google Scholar]
- 165.Fan B, Shen C, Wu M, Zhao J, Guo Q, Luo Y. miR-17-92 cluster is connected with disease progression and oxaliplatin/capecitabine chemotherapy efficacy in advanced gastric cancer patients: A preliminary study. Medicine (Baltimore) 2018;97:e12007. doi: 10.1097/MD.0000000000012007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Huang FX, Chen HJ, Zheng FX, Gao ZY, Sun PF, Peng Q, Liu Y, Deng X, Huang YH, Zhao C, Miao LJ. LncRNA BLACAT1 is involved in chemoresistance of nonsmall cell lung cancer cells by regulating autophagy. Int J Oncol. 2019;54:339–347. doi: 10.3892/ijo.2018.4614. [DOI] [PubMed] [Google Scholar]
- 167.Gu J, Wang D, Zhang J, Zhu Y, Li Y, Chen H, Shi M, Wang X, Shen B, Deng X, et al. GFRa2 prompts cell growth and chemo- resistance through down-regulating tumor suppressor gene PTEN via Mir-17-5p in pancreatic cancer. Cancer Lett. 2016;380:434–441. doi: 10.1016/j.canlet.2016.06.016. [DOI] [PubMed] [Google Scholar]
- 168.Cioffi M, Trabulo SM, Sanchez-Ripoll Y, Miranda-Lorenzo I, Lonardo E, Dorado J, Reis Vieira C, Ramirez JC, Hidalgo M, Aicher A, et al. The miR-17-92 cluster counteracts quiescence and chemoresistance in a distinct subpopulation of pancreatic cancer stem cells. Gut. 2015;64:1936–1948. doi: 10.1136/gutjnl-2014-308470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Yan HJ, Liu WS, Sun WH, Wu J, Ji M, Wang Q, Zheng X, Jiang JT, Wu CP. miR-17-5p inhibitor enhances chemo- sensitivity to gemcitabine via upregulating Bim expression in pancreatic cancer cells. Dig Dis Sci. 2012;57:3160–3167. doi: 10.1007/s10620-012-2400-4. [DOI] [PubMed] [Google Scholar]
- 170.Ao X. Decreased expression of microRNA-17 and microRNA-20b promotes breast cancer resistance to taxol therapy by upregula- tion of NCOA3. Cell Death Dis. 2016;7:e2463. doi: 10.1038/cddis.2016.367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Liao XH, Xiang Y, Yu CX, Li JP, Li H, Nie Q. STAT3 is required for MiR-17-5p-mediated sensitization to chemotherapy-induced apoptosis in breast cancer cells. Oncotarget. 2017;8:15763–15774. doi: 10.18632/oncotarget.15000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Zhu H, Yang SY, Wang J, Wang L, Han SY. Evidence for miR-s17-92 and miR-134 gene cluster regulation of ovarian cancer drug resistance. Eur Rev Med Pharmacol Sci. 2016;20:2526–2531. [PubMed] [Google Scholar]
- 173.Fang Y, Xu C, Fu Y. MicroRNA-17-5p induces drug resistance and invasion of ovarian carcinoma cells by targeting PTEN signaling. J Biol Res (Thessalon) 2015;22:12. doi: 10.1186/s40709-015-0035-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Chatterjee A, Chattopadhyay D, Chakrabarti G. miR-17-5p downregulation contributes to paclitaxel resistance of lung cancer cells through altering beclin1 expression. PLoS One. 2014;9:e95716. doi: 10.1371/journal.pone.0095716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Krutilina R, Sun W, Sethuraman A, Brown M, Seagroves TN, Pfeffer LM, Ignatova T, Fan M. MicroRNA-18a inhibits hypoxia-inducible factor 1a activity and lung metastasis in basal breast cancers. Breast Cancer Res. 2014;16:R78. doi: 10.1186/bcr3693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Godfrey AC, Xu Z, Weinberg CR, Getts RC, Wade PA, DeRoo LA, Sandler DP, Taylor JA. Serum microRNA expression as an early marker for breast cancer risk in prospectively collected samples from the Sister Study cohort. Breast Cancer Res. 2013;15:R42. doi: 10.1186/bcr3428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Shidfar A, Costa FF, Scholtens D, Bischof JM, Sullivan ME, Ivancic DZ, Vanin EF, Soares MB, Wang J, Khan SA. Expression of miR-18a and miR-210 in normal breast tissue as candidate biomarkers of breast cancer risk. Cancer Prev Res (Phila) 2017;10:89–97. doi: 10.1158/1940-6207.CAPR-16-0177. [DOI] [PubMed] [Google Scholar]
- 178.Sha LY, Zhang Y, Wang W, Sui X, Liu SK, Wang T, Zhang H. MiR-18a upregulation decreases Dicer expression and confers paclitaxel resistance in triple negative breast cancer. Eur Rev Med Pharmacol Sci. 2016;20:2201–2208. [PubMed] [Google Scholar]
- 179.Xiao H, Liu Y, Liang P, Wang B, Tan H, Zhang Y, Gao X, Gao J. TP53TG1 enhances cisplatin sensitivity of non-small cell lung cancer cells through regulating miR-18a/PTEN axis. Cell Biosci. 2018;8:23. doi: 10.1186/s13578-018-0221-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Hummel R, Sie C, Watson DI, Wang T, Ansar A, Michael MZ, Van der Hoek M, Haier J, Hussey DJ. MicroRNA signatures in chemotherapy resistant esophageal cancer cell lines. World J Gastroenterol. 2014;20:14904–14912. doi: 10.3748/wjg.v20.i40.14904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Fan YX, Dai YZ, Wang XL, Ren YQ, Han JJ, Zhang H. MiR-18a upregulation enhances autophagy in triple negative cancer cells via inhibiting mTOR signaling pathway. Eur Rev Med Pharmacol Sci. 2016;20:2194–2200. [PubMed] [Google Scholar]
- 182.Zhu HY, Bai WD, Ye XM, Yang AG, Jia LT. Long non-coding RNA UCA1 desensitizes breast cancer cells to trastuzumab by impeding miR-18a repression of Yes-associated protein 1. Biochem Biophys Res Commun. 2018;496:1308–1313. doi: 10.1016/j.bbrc.2018.02.006. [DOI] [PubMed] [Google Scholar]
- 183.Li M, Zhou Y, Xia T, Zhou X, Huang Z, Zhang H, Zhu W, Ding Q, Wang S. Circulating microRNAs from the miR-106a-363 cluster on chromosome X as novel diagnostic biomarkers for breast cancer. Breast Cancer Res Treat. 2018;170:257–270. doi: 10.1007/s10549-018-4757-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Li C, Zhang J, Ma Z, Zhang F, Yu W. miR-19b serves as a prognostic biomarker of breast cancer and promotes tumor progression through PI3K/AKT signaling pathway. Onco Targets Ther. 2018;11:4087–4095. doi: 10.2147/OTT.S171043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Zhao L, Zhao Y, He Y, Mao Y. miR-19b promotes breast cancer metastasis through targeting MYLIP and its related cell adhesion molecules. Oncotarget. 2017;8:64330–64343. doi: 10.18632/oncotarget.19278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Liu M, Yang R, Urrehman U, Ye C, Yan X, Cui S, Hong Y, Gu Y, Liu Y, Zhao C, et al. MiR-19b suppresses PTPRG to promote breast tumorigenesis. Oncotarget. 2016;7:64100–64108. doi: 10.18632/oncotarget.11799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Maleki E, Ghaedi K, Shahanipoor K, Karimi Kurdistani Z. Down-regulation of microRNA-19b in hormone receptor-posi- tive/HER2-negative breast cancer. APMI. S. 2018;126:303–308. doi: 10.1111/apm.12820. [DOI] [PubMed] [Google Scholar]
- 188.Wu Q, Guo L, Jiang F, Li L, Li Z, Chen F. Analysis of the miRNA-mRNA-lncRNA networks in ER+ and ER-breast cancer cell lines. J Cell Mol Med. 2015;19:2874–2887. doi: 10.1111/jcmm.12681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Kurokawa K, Tanahashi T, Iima T, Yamamoto Y, Akaike Y, Nishida K, Masuda K, Kuwano Y, Murakami Y, Fukushima M, Rokutan K. Role of miR-19b and its target mRNAs in 5-fluo - rouracil resistance in colon cancer cells. J Gastroenterol. 2012;47:883–895. doi: 10.1007/s00535-012-0547-6. [DOI] [PubMed] [Google Scholar]
- 190.Thorne JL, Battaglia S, Baxter DE, Hayes JL, Hutchinson SA, Jana S, Millican-Slater RA, Smith L, Teske MC, Wastall LM, Hughes TA. MiR-19b non-canonical binding is directed by HuR and confers chemosensitivity through regulation of P-glycoprotein in breast cancer. Biochim Biophys Acta Gene Regul Mech. 2018;1861:996–1006. doi: 10.1016/j.bbagrm.2018.08.005. [DOI] [PubMed] [Google Scholar]
- 191.Jiang T, Ye L, Han Z, Liu Y, Yang Y, Peng Z, Fan J. miR-19b-3p promotes colon cancer proliferation and oxalipl- atin-based chemoresistance by targeting SMAD4: Validation by bioinformatics and experimental analyses. J Exp Clin Cancer Res. 2017;36:131. doi: 10.1186/s13046-017-0602-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Jin J, Sun Z, Yang F, Tang L, Chen W, Guan X. miR-19b-3p inhibits breast cancer cell proliferation and reverses saraca- tinib-resistance by regulating PI3K/Akt pathway. Arch Biochem Biophys. 2018;645:54–60. doi: 10.1016/j.abb.2018.03.015. [DOI] [PubMed] [Google Scholar]
- 193.Tsai HP, Huang SF, Li CF, Chien HT, Chen SC. Differential microRNA expression in breast cancer with different onset age. PLoS One. 2018;13:e0191195. doi: 10.1371/journal.pone.0191195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Chernyy V, Pustylnyak V, Kozlov V, Gulyaeva L. Increased expression of miR-155 and miR-222 is associated with lymph node positive status. J Cancer. 2018;9:135–140. doi: 10.7150/jca.22181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Xiong DD, Lv J, Wei KL, Feng ZB, Chen JT, Liu KC, Chen G, Luo DZ. A nine-miRNA signature as a potential diagnostic marker for breast carcinoma: An integrated study of 1,110 cases. Oncol Rep. 2017;37:3297–3304. doi: 10.3892/or.2017.5600. [DOI] [PubMed] [Google Scholar]
- 196.Bahrami A, Aledavood A, Anvari K, Hassanian SM, Maftouh M, Yaghobzade A, Salarzaee O, ShahidSales S, Avan A. The prognostic and therapeutic application of microRNAs in breast cancer: Tissue and circulating microRNAs. J Cell Physiol. 2018;233:774–786. doi: 10.1002/jcp.25813. [DOI] [PubMed] [Google Scholar]
- 197.Zhu M, Wang X, Gu Y, Wang F, Li L, Qiu X. MEG3 overexpression inhibits the tumorigenesis of breast cancer by downregulating miR-21 through the PI3K/Akt pathway. Arch Biochem Biophys. 2019;661:22–30. doi: 10.1016/j.abb.2018.10.021. [DOI] [PubMed] [Google Scholar]
- 198.Wang W, Yuan X, Xu A, Zhu X, Zhan Y, Wang S, Liu M. Human cancer cells suppress behaviors of endothelial progenitor cells through miR-21 targeting IL6R. Microvasc Res. 2018;120:21–28. doi: 10.1016/j.mvr.2018.05.007. [DOI] [PubMed] [Google Scholar]
- 199.Han JG, Jiang YD, Zhang CH, Yang YM, Pang D, Song YN, Zhang GQ. A novel panel of serum miR-21/miR-155/miR-365 as a potential diagnostic biomarker for breast cancer. Ann Surg Treat Res. 2017;92:55–66. doi: 10.4174/astr.2017.92.2.55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Yu X, Liang J, Xu J, Li X, Xing S, Li H, Liu W, Liu D, Xu J, Huang L, Du H. Identification and validation of circulating MicroRNA signatures for breast cancer early detection based on large scale tissue-derived data. J Breast Cancer. 2018;21:363–370. doi: 10.4048/jbc.2018.21.e56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Hu X, Fan J, Duan B, Zhang H, He Y, Duan P, Li X. Single-molecule catalytic hairpin assembly for rapid and direct quantification of circulating miRNA biomarkers. Anal Chim. Acta. 2018;1042:109–115. doi: 10.1016/j.aca.2018.08.037. [DOI] [PubMed] [Google Scholar]
- 202.Fan T, Mao Y, Sun Q, Liu F, Lin JS, Liu Y, Cui J, Jiang Y. Branched rolling circle amplification method for measuring serum circulating microRNA levels for early breast cancer detection. Cancer Sci. 2018;109:2897–2906. doi: 10.1111/cas.13725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Hannafon BN, Trigoso YD, Calloway CL, Zhao YD, Lum DH, Welm AL, Zhao ZJ, Blick KE, Dooley WC, Ding WQ. Plasma exosome microRNAs are indicative of breast cancer. Breast Cancer Res. 2016;18:90. doi: 10.1186/s13058-016-0753-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Wang M, Ji S, Shao G, Zhang J, Zhao K, Wang Z, Wu A. Effect of exosome biomarkers for diagnosis and prognosis of breast cancer patients. Clin Transl Oncol. 2018;20:906–911. doi: 10.1007/s12094-017-1805-0. [DOI] [PubMed] [Google Scholar]
- 205.Adhami M, Haghdoost AA, Sadeghi B, Malekpour Afshar R. Candidate miRNAs in human breast cancer biomark ers: A systematic review. Breast Cancer. 2018;25:198–205. doi: 10.1007/s12282-017-0814-8. [DOI] [PubMed] [Google Scholar]
- 206.Petrovic N, Sami A, Martinovic J, Zaric M, Nakashidze I, Lukic S, Jovanovic-Cupic S. TIMP-3 mRNA expression levels positively correlates with levels of miR-21 in in situ BC and negatively in PR positive invasive BC. Pathol Res Pract. 2017;213:1264–1270. doi: 10.1016/j.prp.2017.08.012. [DOI] [PubMed] [Google Scholar]
- 207.Jinling W, Sijing S, Jie Z, Guinian W. Prognostic value of circulating microRNA-21 for breast cancer: A systematic review and meta-analysis. Artif Cells Nanomed Biotechnol. 2017;45:1–6. doi: 10.1080/21691401.2016.1216856. [DOI] [PubMed] [Google Scholar]
- 208.Papadaki C, Stratigos M, Markakis G, Spiliotaki M, Mastrostamatis G, Nikolaou C, Mavroudis D, Agelaki S. Circulating microRNAs in the early prediction of disease recurrence in primary breast cancer. Breast Cancer Res. 2018;20:72. doi: 10.1186/s13058-018-1001-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Huo D, Clayton WM, Yoshimatsu TF, Chen J, Olopade OI. Identification of a circulating microRNA signature to distinguish recurrence in breast cancer patients. Oncotarget. 2016;7:55231–55248. doi: 10.18632/oncotarget.10485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Shin VY, Siu JM, Cheuk I, Ng EK, Kwong A. Circulating cell-free miRNAs as biomarker for triple-negative breast cancer. Br J Cancer. 2015;112:1751–1759. doi: 10.1038/bjc.2015.143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Fang H, Xie J, Zhang M, Zhao Z, Wan Y, Yao Y. miRNA-21 promotes proliferation and invasion of triple-negative breast cancer cells through targeting PTEN. Am J Transl Res. 2017;9:953–961. [PMC free article] [PubMed] [Google Scholar]
- 212.Dong G, Liang X, Wang D, Gao H, Wang L, Wang L, Liu J, Du Z. High expression of miR-21 in triple-negative breast cancers was correlated with a poor prognosis and promoted tumor cell in vitro proliferation. Med Oncol. 2014;31:57. doi: 10.1007/s12032-014-0057-x. [DOI] [PubMed] [Google Scholar]
- 213.Komatsu S, Ichikawa D, Kawaguchi T, Miyamae M, Okajima W, Ohashi T, Imamura T, Kiuchi J, Konishi H, Shiozaki A, et al. Circulating miR-21 as an independent predictive biomarker for chemoresistance in esophageal squamous cell carcinoma. Am J Cancer Res. 2016;6:1511–1523. [PMC free article] [PubMed] [Google Scholar]
- 214.Campayo M, Navarro A, Benitez JC, Santasusagna S, Ferrer C, Monzo M, Cirera L. miR-21, miR-99b and miR-375 combination as predictive response signature for preoperative chemoradiotherapy in rectal cancer. PLoS One. 2018;13:e0206542. doi: 10.1371/journal.pone.0206542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Gaudelot K, Gibier JB, Pottier N, Hemon B, Van Seuningen I, Glowacki F, Leroy X, Cauffiez C, Gnemmi V, Aubert S, Perrais M. Targeting miR-21 decreases expression of multi-drug resistant genes and promotes chemosensitivity of renal carcinoma. Tumour Biol. 2017;39:1010428317707372. doi: 10.1177/1010428317707372. [DOI] [PubMed] [Google Scholar]
- 216.Lin L, Tu HB, Wu L, Liu M, Jiang GN. MicroRNA-21 regulates non-small cell lung cancer cell invasion and chemo-sensitivity through SMAD7. Cell Physiol Biochem. 2016;38:2152–2162. doi: 10.1159/000445571. [DOI] [PubMed] [Google Scholar]
- 217.Szejniuk WM, Robles AI, McCulloch T, Falkmer UGI, Roe OD. Epigenetic predictive biomarkers for response or outcome to platinum-based chemotherapy in non-small cell lung cancer, current state-of-art. Pharmacogenomics J. 2019;19:5–14. doi: 10.1038/s41397-018-0029-1. [DOI] [PubMed] [Google Scholar]
- 218.Feng Y, Zou W, Hu C, Li G, Zhou S, He Y, Ma F, Deng C, Sun L. Modulation of CASC2/miR-21/PTEN pathway sensitizes cervical cancer to cisplatin. Arch Biochem Biophys. 2017;623-624:20–30. doi: 10.1016/j.abb.2017.05.001. [DOI] [PubMed] [Google Scholar]
- 219.Chan JK, Blansit K, Kiet T, Sherman A, Wong G, Earle C, Bourguignon LY. The inhibition of miR-21 promotes apoptosis and chemosensitivity in ovarian cancer. Gynecol Oncol. 2014;132:739–744. doi: 10.1016/j.ygyno.2014.01.034. [DOI] [PubMed] [Google Scholar]
- 220.Pink RC, Samuel P, Massa D, Caley DP, Brooks SA, Carter DR. The passenger strand, miR-21-3p, plays a role in mediating cisplatin resistance in ovarian cancer cells. Gynecol Oncol. 2015;137:143–151. doi: 10.1016/j.ygyno.2014.12.042. [DOI] [PubMed] [Google Scholar]
- 221.Dai X, Fang M, Li S, Yan Y, Zhong Y, Du B. miR-21 is involved in transforming growth factor |31-induced chemoresis- tance and invasion by targeting PTEN in breast cancer. Oncol Lett. 2017;14:6929–6936. doi: 10.3892/ol.2017.7007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Bose RJC, Uday Kumar S, Zeng Y, Afjei R, Robinson E, Lau K, Bermudez A, Habte F, Pitteri SJ, Sinclair R, et al. Tumor cell-derived extracellular vesicle-coated nanocarriers: An efficient theranostic platform for the cancer-specific delivery of Anti-miR-21 and imaging agents. ACS Nano. 2018;12:10817–10832. doi: 10.1021/acsnano.8b02587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.Zhou Q, Zeng H, Ye P, Shi Y, Guo J, Long X. Differential microRNA profiles between fulvestrant-resistant and tamoxifen-resistant human breast cancer cells. Anticancer Drugs. 2018;29:539–548. doi: 10.1097/CAD.0000000000000623. [DOI] [PubMed] [Google Scholar]
- 224.Wu ZH, Tao ZH, Zhang J, Li T, Ni C, Xie J, Zhang JF, Hu XC. MiRNA-21 induces epithelial to mesenchymal transition and gemcitabine resistance via the PTEN/AKT pathway in breast cancer. Tumour Biol. 2016;37:7245–7254. doi: 10.1007/s13277-015-4604-7. [DOI] [PubMed] [Google Scholar]
- 225.Dhayat SA, Mardin WA, Seggewifi J, Strose AJ, Matuszcak C, Hummel R, Senninger N, Mees ST, Haier J. MicroRNA profiling implies new markers of gemcitabine chemoresistance in mutant p53 pancreatic ductal adenocarcinoma. PLoS One. 2015;10:e0143755. doi: 10.1371/journal.pone.0143755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Dong J, Zhao YP, Zhou L, Zhang TP, Chen G. Bcl-2 upregu- lation induced by miR-21 via a direct interaction is associated with apoptosis and chemoresistance in MIA PaCa-2 pancreatic cancer cells. Arch Med Res. 2011;42:8–14. doi: 10.1016/j.arcmed.2011.01.006. [DOI] [PubMed] [Google Scholar]
- 227.Kopczynska E. Role of microRNAs in the resistance of prostate cancer to docetaxel and paclitaxel. Contemp Oncol (Pozn) 2015;19:423–427. doi: 10.5114/wo.2015.56648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Haghnavaz N, Asghari F, Komi Ali Elieh D, Shanehbandi D, Baradaran B, Kazemi T. HER2 positivity may confer resistance to therapy with paclitaxel in breast cancer cell lines. Artif Cells Nanomed Biotechnol. 2018;46:518–523. doi: 10.1080/21691401.2017.1326927. [DOI] [PubMed] [Google Scholar]
- 229.Du G, Cao D, Meng L. miR-21 inhibitor suppresses cell proliferation and colony formation through regulating the PTEN/AKT pathway and improves paclitaxel sensitivity in cervical cancer cells. Mol Med Rep. 2017;15:2713–2719. doi: 10.3892/mmr.2017.6340. [DOI] [PubMed] [Google Scholar]
- 230.Jin B, Liu Y, Wang H. Antagonism of miRNA-21 sensitizes human gastric cancer cells to paclitaxel. Cell Biochem Biophys. 2015;72:275–282. doi: 10.1007/s12013-014-0450-2. [DOI] [PubMed] [Google Scholar]
- 231.Mei M, Ren Y, Zhou X, Yuan XB, Han L, Wang GX, Jia Z, Pu PY, Kang CS, Yao Z. Downregulation of miR-21 enhances chemotherapeutic effect of taxol in breast carcinoma cells. Technol Cancer Res Treat. 2010;9:77–86. doi: 10.1177/153303461000900109. [DOI] [PubMed] [Google Scholar]
- 232.Naro Y, Ankenbruck N, Thomas M, Tivon Y, Connelly CM, Gardner L, Deiters A. Small molecule inhibition of MicroRNA miR-21 rescues chemosensitivity of renal-cell carcinoma to topotecan. J Med Chem. 2018;61:5900–5909. doi: 10.1021/acs.jmedchem.7b01891. [DOI] [PubMed] [Google Scholar]
- 233.Liu Y, Xu J, Choi HH, Han C, Fang Y, Li Y, Van der Jeught K, Xu H, Zhang L, Frieden M, et al. Targeting 17q23 amplicon to overcome the resistance to anti-HER2 therapy in HER2+ breast cancer. Nat Commun. 2018;9:4718. doi: 10.1038/s41467-018-07264-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Decker JT, Hall MS, Blaisdell RB, Schwark K, Jeruss JS, Shea LD. Dynamic microRNA activity identifies therapeutic targets in trastuzumab-resistant HER2+ breast cancer. Biotechnol Bioeng. 2018;115:2613–2623. doi: 10.1002/bit.26791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235.Bahreyni A, Alibolandi M, Ramezani M, Sarafan Sadeghi A, Abnous K, Taghdisi SM. A novel MUC1 aptamer-modified PLGA-epirubicin-PpAE-antimir-21 nanocomplex platform for targeted co-delivery of anticancer agents in vitro and in vivo. Colloids Surf B Biointerfaces. 2018;175:231–238. doi: 10.1016/j.colsurfb.2018.12.006. [DOI] [PubMed] [Google Scholar]
- 236.Vandghanooni S, Eskandani M, Barar J, Omidi Y. AS1411 aptamer-decorated cisplatin-loaded poly(lactic-co-glycolic acid) nanoparticles for targeted therapy of miR-21-inhibited ovarian cancer cells. Nanomedicine (Lond) 2018;13:2729–2758. doi: 10.2217/nnm-2018-0205. [DOI] [PubMed] [Google Scholar]
- 237.Wang W, Huang S, Yuan J, Xu X, Li H, Lv Z, Yu W, Duan S, Hu Y. Reverse multidrug resistance in human HepG2/ADR by Anti-miR-21 combined with hyperthermia mediated by functionalized gold nanocages. Mol Pharm. 2018;15:3767–3776. doi: 10.1021/acs.molpharmaceut.8b00046. [DOI] [PubMed] [Google Scholar]
- 238.Luo J, Zhao Q, Zhang W, Zhang Z, Gao J, Zhang C, Li Y, Tian Y. A novel panel of microRNAs provides a sensitive and specific tool for the diagnosis of breast cancer. Mol Med Rep. 2014;10:785–791. doi: 10.3892/mmr.2014.2274. [DOI] [PubMed] [Google Scholar]
- 239.Zhang K, Wang YW, Wang YY, Song Y, Zhu J, Si PC, Ma R. Identification of microRNA biomarkers in the blood of breast cancer patients based on microRNA profiling. Gene. 2017;619:10–20. doi: 10.1016/j.gene.2017.03.038. [DOI] [PubMed] [Google Scholar]
- 240.Croset M, Pantano F, Kan CWS, Bonnelye E, Descotes F, Alix-Panabieres C, Lecellier CH, Bachelier R, Allioli N, Hong SS, et al. MicroRNA-30 family members inhibit breast cancer invasion, osteomimicry, and bone destruction by directly targeting multiple bone metastasis-associated genes. Cancer Res. 2018;78:5259–5273. doi: 10.1158/0008-5472.CAN-17-3058. [DOI] [PubMed] [Google Scholar]
- 241.Ni Q, Stevic I, Pan C, Muller V, Oliviera-Ferrer L, Pantel K, Schwarzenbach H. Different signatures of miR-16, miR-30b and miR-93 in exosomes from breast cancer and DCIS patients. Sci Rep. 2018;8:12974. doi: 10.1038/s41598-018-31108-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Xi Z, Si J, Nan J. LncRNA MALAT1 potentiates autophagyassociated cisplatin resistance by regulating the microRNA30b/autophagyrelated gene 5 axis in gastric cancer. Int J Oncol. 2019;54:239–248. doi: 10.3892/ijo.2018.4609. [DOI] [PubMed] [Google Scholar]
- 243.Chen JC, Su YH, Chiu CF, Chang YW, Yu YH, Tseng CF, Chen HA, Su JL. Suppression of Dicer increases sensitivity to gefitinib in human lung cancer cells. Ann Surg Oncol. 2014;21(Suppl 4):S555–S563. doi: 10.1245/s10434-014-3673-y. [DOI] [PubMed] [Google Scholar]
- 244.Tormo E, Adam-Artigues A, Ballester S, Pineda B, Zazo S, Gonzalez-Alonso P, Albanell J, Rovira A, Rojo F, Lluch A, Eroles P. The role of miR-26a and miR-30b in HER2+ breast cancer trastuzumab resistance and regulation of the CCNE2 gene. Sci Rep. 2017;7:41309. doi: 10.1038/srep41309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245.Gu YF, Zhang H, Su D, Mo ML, Song P, Zhang F, Zhang SC. miR-30b and miR-30c expression predicted response to tyrosine kinase inhibitors as first line treatment in non-small cell lung cancer. Chin Med J (Engl) 2013;126:4435–4439. [PubMed] [Google Scholar]
- 246.Amini S, Abak A, Estiar MA, Montazeri V, Abhari A, Sakhinia E. Expression Analysis of MicroRNA-222 in breast cancer. Clin Lab. 2018;64:491–496. doi: 10.7754/Clin.Lab.2017.171002. [DOI] [PubMed] [Google Scholar]
- 247.Zong Y, Zhang Y, Sun X, Xu T, Cheng X, Qin Y. miR-221/222 promote tumor growth and suppress apoptosis by targeting lncRNA GAS5 in breast cancer. Biosci Rep. 2019;39 doi: 10.1042/BSR20181859. pii: BSR20181859. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 248.Hoppe R, Fan P, Buttner F, Winter S, Tyagi AK, Cunliffe H, Jordan VC, Brauch H. Profiles of miRNAs matched to biology in aromatase inhibitor resistant breast cancer. Oncotarget. 2016;7:71235–71254. doi: 10.18632/oncotarget.12103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 249.Han SH, Kim HJ, Gwak JM, Kim M, Chung YR, Park SY. MicroRNA-222 expression as a predictive marker for tumor progression in hormone receptor-positive breast cancer. J Breast Cancer. 2017;20:35–44. doi: 10.4048/jbc.2017.20.1.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Kim C, Go EJ, Kim A. Recurrence prediction using microRNA expression in hormone receptor positive breast cancer during tamoxifen treatment. Biomarkers. 2018;23:804–811. doi: 10.1080/1354750X.2018.1499131. [DOI] [PubMed] [Google Scholar]
- 251.Zhu W, Liu M, Fan Y, Ma F, Xu N, Xu B. Dynamics of circulating microRNAs as a novel indicator of clinical response to neoadjuvant chemotherapy in breast cancer. Cancer Med. 2018;7:4420–4433. doi: 10.1002/cam4.1723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 252.Chen X, Lu P, Wang DD, Yang SJ, Wu Y, Shen HY, Zhong SL, Zhao JH, Tang JH. The role of miRNAs in drug resistance and prognosis of breast cancer formalin-fixed paraffin-embedded tissues. Gene. 2016;595:221–226. doi: 10.1016/j.gene.2016.10.015. [DOI] [PubMed] [Google Scholar]
- 253.Chen J, Chen Z, Huang J, Chen F, Ye W, Ding G, Wang X. Bioinformatics identification of dysregulated microRNAs in triple negative breast cancer based on microRNA expression profiling. Oncol Lett. 2018;15:3017–3023. doi: 10.3892/ol.2017.7707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 254.Li Y, Liang C, Ma H, Zhao Q, Lu Y, Xiang Z, Li L, Qin J, Chen Y, Cho WC, et al. miR-221/222 promotes S-phase entry and cellular migration in control of basal-like breast cancer. Molecules. 2014;19:7122–7137. doi: 10.3390/molecules19067122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Zhao L, Ren Y, Tang H, Wang W, He Q, Sun J, Zhou X, Wang A. Deregulation of the miR-222-ABCG2 regulatory module in tongue squamous cell carcinoma contributes to chemoresistance and enhanced migratory/invasive potential. Oncotarget. 2015;6:44538–44550. doi: 10.18632/oncotarget.6253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Shen H, Wang D, Li L, Yang S, Chen X, Zhou S, Zhong S, Zhao J, Tang J. MiR-222 promotes drug-resistance of breast cancer cells to adriamycin via modulation of PTEN/Akt/FOXO1 pathway. Gene. 2017;596:110–118. doi: 10.1016/j.gene.2016.10.016. [DOI] [PubMed] [Google Scholar]
- 257.Wang DD, Yang SJ, Chen X, Shen HY, Luo LJ, Zhang XH, Zhong SL, Zhao JH, Tang JH. miR-222 induces Adriamycin resistance in breast cancer through PTEN/Akt/p27(kip1) pathway. Tumour Biol. 2016;37:15315–15324. doi: 10.1007/s13277-016-5341-2. [DOI] [PubMed] [Google Scholar]
- 258.Rao X, Di Leva G, Li M, Fang F, Devlin C, Hartman-Frey C, Burow ME, Ivan M, Croce CM, Nephew KP. MicroRNA- 221/222 confers breast cancer fulvestrant resistance by regulating multiple signaling pathways. Oncogene. 2011;30:1082–1097. doi: 10.1038/onc.2010.487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Wei F, Ma C, Zhou T, Dong X, Luo Q, Geng L, Ding L, Zhang Y, Zhang L, Zhang L, et al. Exosomes derived from gemcitabine-resistant cells transfer malignant phenotypic traits via delivery of miRNA-222-3p. Mol Cancer. 2017;16:132. doi: 10.1186/s12943-017-0694-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Bliss SA, Sinha G, Sandiford OA, Williams LM, Engelberth DJ, Guiro K, Isenalumhe LL, Greco SJ, Ayer S, Bryan M, et al. Mesenchymal stem cell-derived exosomes stimulate cycling quiescence and early breast cancer dormancy in bone marrow. Cancer Res. 2016;76:5832–5844. doi: 10.1158/0008-5472.CAN-16-1092. [DOI] [PubMed] [Google Scholar]
- 261.Ding J, Xu Z, Zhang Y, Tan C, Hu W, Wang M, Xu Y, Tang J. Exosome-mediated miR-222 transferring: An insight into NF-kappaB-mediated breast cancer metastasis. Exp Cell Res. 2018;369:129–138. doi: 10.1016/j.yexcr.2018.05.014. [DOI] [PubMed] [Google Scholar]
- 262.Gan R, Yang Y, Yang X, Zhao L, Lu J, Meng QH. Downregulation of miR-221/222 enhances sensitivity of breast cancer cells to tamoxifen through upregulation of TIMP3. Cancer Gene Ther. 2014;21:290–296. doi: 10.1038/cgt.2014.29. [DOI] [PubMed] [Google Scholar]
- 263.Vishnubalaji R, Hamam R, Yue S, Al-Obeed O, Kassem M, Liu FF, Aldahmash A, Alajez NM. MicroRNA-320 suppresses colorectal cancer by targeting SOX4, FOXM1, and FOXQ1. Oncotarget. 2016;7:35789–35802. doi: 10.18632/oncotarget.8937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 264.Iwagami Y, Eguchi H, Nagano H, Akita H, Hama N, Wada H, Kawamoto K, Kobayashi S, Tomokuni A, Tomimaru Y, et al. miR-320c regulates gemcitabine-resistance in pancreatic cancer via SMARCC1. Br J Cancer. 2013;109:502–511. doi: 10.1038/bjc.2013.320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 265.Li M, Zeringer E, Barta T, Schageman J, Cheng A, Vlassov AV. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Philos Trans R Soc Lond B Biol Sci. 2014;369:pii: 20130502. doi: 10.1098/rstb.2013.0502. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets analysed during the current study are available from the corresponding author on reasonable request.