Summary
6S RNA binds to RNA polymerase and regulates gene expression, contributing to bacterial adaptation to environmental stresses. In this study, we examined the role of 6S RNA in murine infectivity and tick persistence of the Lyme disease spirochete Borrelia (Borreliella) burgdorferi. B. burgdorferi 6S RNA (Bb6S RNA) binds to RNA polymerase, is expressed independent of growth phase or nutrient stress in culture, and is processed by RNase Y. We found that rny (bb0504), the gene encoding RNase Y, is essential for B. burgdorferi growth, while ssrS, the gene encoding 6S RNA, is not essential, indicating a broader role for RNase Y activity in the spirochete. Bb6S RNA regulates expression of the ospC and dbpA genes encoding outer surface protein C and decorin binding protein A, respectively, which are lipoproteins important for host infection. The highest levels of Bb6S RNA are found when the spirochete resides in unfed nymphs. ssrS mutants lacking Bb6S RNA were compromised for infectivity by needle inoculation, but injected mice seroconverted, indicating an ability to activate the adaptive immune response. ssrS mutants were successfully acquired by larval ticks and persisted through fed nymphs. Bb6S RNA is one of the first regulatory RNAs identified in B. burgdorferi that controls the expression of lipoproteins involved in host infectivity.
Keywords: Borrelia burgdorferi; Lyme disease; 6S RNA; gene expression regulation, bacterial; RNA, small untranslated; Spirochaetales
Graphical Abstract

Borrelia burgdorferi, the causative agent of Lyme disease, is maintained in nature in an enzootic cycle involving a tick and a vertebrate. Bb6S RNA, a small regulatory RNA in B. burgdorferi, is processed by ribonuclease Y, binds RNA polymerase, regulates transcription, and is involved in infection of the vertebrate host and persistence in the tick vector.
Introduction
The Lyme disease agent Borrelia (Borreliella) burgdorferi navigates disparate and challenging environments as it traverses through its enzootic cycle, which alternates between Ixodes ticks and vertebrates (Radolf et al., 2012; Caimano et al., 2016). B. burgdorferi is acquired when Ixodes larvae feed on an infected animal, and the spirochete then persists in the tick midgut as the blood meal is consumed by the larvae before it molts into a nymph. The spirochete can be transmitted to and infect a new host as the nymph takes a blood meal, completing an enzootic cycle. B. burgdorferi adapts to and thrives in the different environments of the vector and host via the global regulation of gene expression (Iyer and Schwartz, 2016; Samuels and Samuels, 2016) mediated by the alternative sigma factor RpoS (σS) (Fisher et al., 2005; Caimano et al., 2007; Ouyang et al., 2008), the stringent response via guanosine tetraphosphate and pentaphosphate [(p)ppGpp] (Bugrysheva et al., 2015; Drecktrah et al., 2015), and cyclic dimeric GMP (c-di-GMP) (Rogers et al., 2009; He et al., 2011; Caimano et al., 2015). While studies of these signaling systems have begun to uncover the gene products important for infectivity and persistence in the tick, our understanding of global transcriptional regulators remains incomplete.
6S RNA is an abundant small RNA that regulates transcription in Escherichia coli, and other bacteria, by direct interaction with the σ70-containing form of RNA polymerase (RNAP), referred to as the σ70-holoenzyme (Eσ70) (Wassarman and Storz, 2000; Cavanagh and Wassarman, 2014; Steuten et al., 2014a; Steuten et al., 2014b; Burenina et al., 2015; Wassarman, 2018). Therefore, 6S RNA functions as a global regulator, in contrast to other sRNAs that typically bind to target RNAs to affect expression of a more limited set of genes (Waters and Storz, 2009; Storz et al., 2011). The defining characteristic of 6S RNAs is not their nucleotide sequence but their secondary structure, which forms a central bubble of single-stranded RNA flanked by double-stranded RNA stems and is required for 6S RNA interaction with RNAP (Wassarman and Storz, 2000; Barrick et al., 2005; Trotochaud and Wassarman, 2005; Chen et al., 2017). This structure mimics the open promoter DNA complex of transcriptional initiation sites; binding of 6S RNA to σ70-RNAP directly competes with σ70-dependent promoter binding (Wassarman and Storz, 2000). 6S RNA binding to σ70-RNAP leads to decreased expression from many σ70-dependent promoters in E. coli (Wassarman and Storz, 2000; Trotochaud and Wassarman, 2004; Trotochaud and Wassarman, 2006; Cavanagh et al., 2008; Cavanagh et al., 2010; Neusser et al., 2010). At the same time, 6S RNA binding to σ70-RNAP leads to increased expression from many σS-dependent (RpoS-dependent) promoters (Trotochaud and Wassarman, 2004; Cavanagh et al., 2008; Neusser et al., 2010). Thus, increased 6S RNA levels generally shift expression from σ70-dependent to σS-dependent promoters, although not all promoters are affected in the same manner and the assembly of sequence elements influences the response, at least for σ70-dependent promoters (Cavanagh et al., 2008). These molecular mechanisms are one example of the cellular reprogramming of gene expression to respond to environmental stresses.
In E. coli, 6S RNA increases throughout growth in log phase and early stationary phase, accumulating to high levels in late stationary phase (Wassarman and Storz, 2000). The increase in 6S RNA levels and regulated changes in gene expression are thought to contribute to adaptation to the environmental stresses associated with limited nutrients and high cell density. The effects of 6S RNA are amplified by targeting global regulators, including the transcription factors PspF (Trotochaud and Wassarman, 2006; Joly et al., 2010), Crp (Cavanagh et al., 2008; Neusser et al., 2010) and RelA (Cavanagh et al., 2010). Cells lacking 6S RNA are not compromised for growth in exponential phase but have a survival defect during long-term nutrient stress and competitive survival (Lee et al., 1985; Trotochaud and Wassarman, 2004). 6S RNA levels in some bacteria are regulated not by growth phase but by other signals, such as oxidative stress for Burkholderia cenocepacia (Peeters et al., 2010), host cell environment for Coxiella burnetii (Warrier et al., 2014) and Yersinia pestis (Yan et al., 2013), and the cell-cycle and light for cyanobacteria (Axmann et al., 2007). In addition, both Bacillus subtilis and Legionella pneumophila have two versions of 6S RNA, Bs6S-1 and Bs6S-2 and Lp6S and Lp6S-2, respectively, which are differentially expressed and regulate distinct adaptations to disparate stresses (Ando et al., 2002; Suzuma et al., 2002; Barrick et al., 2005; Trotochaud and Wassarman, 2005; Faucher et al., 2010; Weissenmayer et al., 2011; Cavanagh et al., 2012; Cavanagh and Wassarman, 2013; Burenina et al., 2014). Bs6S-1 RNA levels increase as cells enter stationary phase while Bs6S-2 RNA levels remain relatively unchanged (Ando et al., 2002; Suzuma et al., 2002; Trotochaud and Wassarman, 2005; Beckmann et al., 2011). Mutagenesis experiments showed that Bs6S-1 RNA regulates the timing of sporulation while Bs6S-2 does not (Cavanagh and Wassarman, 2013). Thus, the 6S RNAs clearly function to alter gene expression to adapt to environmental stresses but the regulatory signals, cellular responses and physiological importance of these sRNAs are varied.
More recently, 6S RNA has been suggested to play a role in virulence of bacteria, including L. pneumophila (Faucher et al., 2010), Y. pestis (Yan et al., 2013), C. burnetii (Warrier et al., 2014), and Salmonella enterica serovar Typhimurium (Ren et al., 2017), specifically where replication and stress resistance is tied to pathogenesis. However, Y. pestis is the only bacterial pathogen where 6S RNA function has been examined using an animal model (Yan et al., 2013). Expression and function of 6S RNA has not been studied in B. burgdorferi, or any other spirochete, to our knowledge. Here, we report the first study of the function of an sRNA in B. burgdorferi in the tick-mouse model of Lyme disease. We show that Bb6S RNA binds RNAP in the spirochete, is processed by RNase Y, and regulates expression of lipoproteins important for host infection.
Results
The B. burgdorferi ssrS gene (encoding Bb6S RNA), first identified by Barrick et al. (2005), is in the intergenic region between bb0187 and bb0188 on the chromosome (Fig. 1A). We experimentally determined that Bb6S RNA is 204 nucleotides by 3′ and 5′ RACE and modeled the secondary structure using mfold (Zuker, 2003) (Fig. 1B). Bb6S RNA, while having only about 35% identity with E. coli 6S RNA, is predicted to form the stem-bulge-stem structure common to all 6S RNAs that likely mimics the DNA open promoter complex during transcriptional initiation (Wassarman and Storz, 2000; Barrick et al., 2005; Trotochaud and Wassarman, 2005; Chen et al., 2017).
Figure 1.
Bb6S RNA in B. burgdorferi. (A) Genomic locus of ssrS encoding the B. burgdorferi homolog of 6S RNA (Bb6S RNA). (B) The predicted secondary structure of 6S RNA from B. burgdorferi and, for comparison, E. coli (Wassarman, 2018). The 5′ and 3′ ends of Bb6S RNA were experimentally determined by RACE. The majority of 5′ ends (5/6) mapped to a C five nucleotides downstream from the end of the bb0188 (rplT) ORF. The majority of 3′ ends (11/14) mapped to a U 24 nucleotides upstream from the start of the bb0187 ORF.
6S RNA regulates transcription in bacteria by binding to σ70-RNAP and sequestering it from binding to σ70-dependent promoters. We assayed the binding of Bb6S RNA to RNAP in B. burgdorferi by co-immunoprecipitation. Antibodies specific to B. burgdorferi RNAP (BbRNAP) are not available, but we found that antibodies specific to E. coli RNAP (EcRNAP) sufficiently cross-reacted with BbRNAP (Fig. 2A, cell lysate (1/2)). Spirochete extracts immunoprecipitated with preimmune serum or serum specific to EcRNAP antibodies were separated by SDS-PAGE, transferred to a membrane and immunoblotted with the same EcRNAP-specific serum used for immunoprecipitation. The RNAP subunits are visible in the one-half equivalent cell extract and in the anti-RNAP core immunoprecipitation (IP), but not in the preimmune serum immunoprecipitated treatment (Fig. 2A). RNA from the other portion of the cell extracts and immunoprecipitated samples were separated on a urea gel and analyzed by Northern blot to examine relative Bb6S RNA levels compared to 5S rRNA levels as a control. Bb6S RNA co-immunoprecipitated with serum specific for EcRNAP, but not the preimmune serum, indicating either a direct or an indirect interaction between Bb6S RNA and BbRNAP (Fig. 2B). 5S rRNA did not co-immunoprecipitate with either serum (Fig. 2B), supporting the specificity of the Bb6S RNA-RNAP interaction in B. burgdorferi cells. Bb6S RNA was also able to bind E. coli RNAP (σ70 holoenzyme) in vitro as demonstrated by a gel shift assay using in vitro transcribed 32P-labeled Bb6S RNA (Fig S1), suggesting a direct interaction between Bb6S RNA and RNAP. This is only the second identification of an sRNA-protein interaction in B. burgdorferi (Lybecker et al., 2010; Lybecker and Samuels, 2017).
Figure 2. Bb6S RNA binds to B. burgdorferi RNAP.
(A) Immunoblot of B. burgdorferi cell lysate one-half (1/2) or one-tenth (1/10) the equivalent used for immunoprecipitation (IP) of extracts with preimmune serum or anti-E. coli RNAP antiserum (clone WI 151). The blot was probed with the anti-E. coli RNAP antiserum (clone WI 151) to visualize B. burgdorferi RNAP core. Arrows denote proteins present in the cell extract and enriched in IP with anti-E. coli RNAP antiserum but not preimmune serum. (B) Northern blot of a B. burgdorferi total cell lysate and extracts immunoprecipitated with preimmune serum (preimmune IP) or anti-E. coli RNAP antiserum WI-151 (α-RNAP IP) separated on a urea gel and hybridized with a biotinylated RNA probe to Bb6S RNA (upper panel) or B. burgdorferi 5S rRNA (lower panel). The amount of cell lysate was 10% of the equivalent used for the immunoprecipitation.
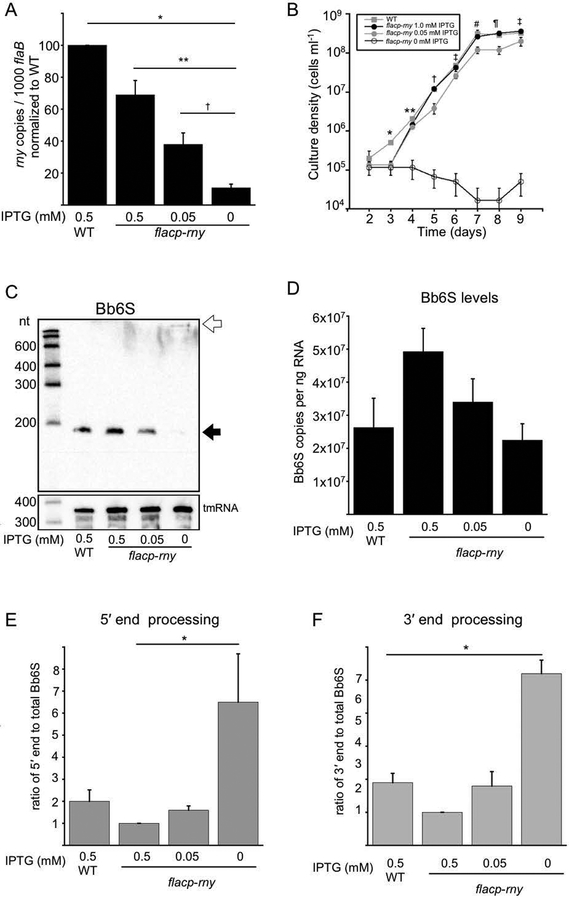
6S RNA is processed from a larger transcript in E. coli by mechanisms involving RNase BN or the endoribonucleases RNase G and RNase E (Kim and Lee, 2004; Chen et al., 2016) and further trimmed by exoribonucleases RNase T and RNase PH (Li et al., 1998). B. burgdorferi has a limited repertoire of ribonucleases, compared to E. coli and B. subtilis, and lacks genes encoding RNase G and RNase E; the only predicted endoribonuclease homologs are RNase III, RNase M5, RNase P, RNase Y, RNase Z, YbeY, and RNase HII (Fraser et al., 1997; Archambault et al., 2013; Anacker et al., 2018). The size of Bb6S RNA was assayed by Northern blot in individual RNase mutant strains to investigate Bb6S RNA processing. First, using our extant rnc mutant (Anacker et al., 2018), Bb6S RNA size and levels were largely unchanged, with only a small portion of Bb6S RNA remaining incompletely processed (Fig S2), which indicates that RNase III has a limited role in processing. Next, we focused on RNase Y since it has been postulated to functionally replace RNase E (Shahbabian et al., 2009). Multiple efforts to generate an rny deletion mutant lacking RNase Y were unsuccessful, which suggests that RNase Y is essential in B. burgdorferi. Therefore, we constructed a conditional mutant with an IPTG-inducible flacp-rny fusion (Gilbert et al., 2007). To assay the regulation of rny expression in the flacp-rny strain, cells were grown to 107 cells ml−1 in the presence of IPTG before removing the inducer. Cells were then resuspended with or without IPTG and grown for 48 h. rny expression, measured by qRT-PCR, in the flacp-rny strain in the presence of 0.5 mM IPTG is slightly less than that seen in wild type (Fig. 3A). Levels of rny mRNA decreased as the IPTG concentration was reduced in flacp-rny mutants, demonstrating IPTG-dependent rny expression (Fig. 3A). To examine the effect of rny on growth, spirochetes were treated with or without IPTG and cells were enumerated for eight days. Conditional flacp-rny mutants grown in the presence of 1.0 mM IPTG had similar growth kinetics as wild-type B. burgdorferi (Fig. 3B, gray squares and black circles). There was a slight decrease in growth rate and final cell density of flacp-rny mutants in the presence of 0.05 mM IPTG (Fig. 3B, gray circles). The flacp-rny strain failed to replicate in the absence of IPTG (Fig. 3B, open circles). The lack of growth without IPTG supports our hypothesis that the rny gene is essential in B. burgdorferi. The influence of reduced rny expression on Bb6S RNA processing was examined by Northern blot analyses. Cells were treated with or without IPTG and analyzed by Northern blot, using transfer-messenger RNA (tmRNA) as a control. In flacp-rny mutants with reduced levels of rny mRNA (Fig. 3A, 0 mM IPTG), the mature form of Bb6S RNA was dramatically reduced (Fig. 3C, filled arrow) while tmRNA levels were unchanged (Fig. 3C). To further investigate the role of RNase Y in Bb6S maturation, we used SYBR Green primers spanning the mature 5′ and 3′ ends for qRT-PCR analyses. The levels of Bb6S trended lower as IPTG was removed from the flacp-rny strain, but the differences were not significant (Fig. 3D, P > 0.05, one-way ANOVA with a Tukey’s post-hoc test). The levels of the 5′ and 3′ unprocessed ends both increased when RNase Y was depleted by removing IPTG (Fig. 3E and F, 0 mM IPTG). In the flacp-rny strain, the ratio of 5′ end to Bb6S increased sixfold (Fig. 3E) and the ratio of 3′ end to Bb6S increased about sevenfold (Fig. 3F) when IPTG was removed compared to cells with 0.5 mM IPTG. Therefore, our data suggest a role for RNase Y in the biogenesis of Bb6S RNA by processing both the 5′ and 3′ ends.
Figure 3. Bb6S RNA is processed by RNase Y.
RNase Y levels were depleted by fusing the artificially inducible promoter flacp to the rny gene, encoding RNase Y, and inserted into the rny locus on the B. burgdorferi chromosome. (A) flacp-rny spirochetes were grown in 0.5 mM IPTG to 107 cells ml−1. IPTG was removed from two-thirds of the cells, which were then placed in growth medium containing either 0 or 0.05 mM IPTG for 48 h. One-third of the flacp-rny cells remained in 0.5 mM IPTG for 48 h. Wild-type (WT) spirochetes were grown in the presence of 0.5 mM IPTG until late log phase. Levels of rny mRNA were quantified by TaqMan qRT-PCR and normalized to flaB mRNA levels relative to wild type. Values are the mean of four independent biological replicates and error bars represent the SE. * denotes a significant difference between WT and flacp-rny strains in all levels of IPTG, ** denotes a significant difference between flacp-rny in 0.5 mM IPTG and both 0.05 and 0 mM IPTG and † denotes a significant difference between 0.05 mM and 0 mM IPTG (P < 0.05 determined by one-way ANOVA with Tukey’s post-hoc test). (B) WT and flacp-rny spirochetes were grown in 0.5 mM IPTG for two days before IPTG was removed and cells were resuspended in growth medium containing 0, 0.05 or 1.0 mM IPTG. Cells were enumerated each day for the next eight days (days 2 to 9). Values are the means of three independent biological replicates and error bars represent the SE. Significance (P < 0.05) was determined by one-way ANOVA with Tukey’s post-hoc test. * denotes WT significantly different from 1.0, 0.05, and 0 mM IPTG-treated flacp-rny strains, ** denotes 0 mM IPTG-treated flacp-rny significantly different from all others and WT significantly different from 1.0 and 0.05 mM IPTG-treated flacp-rny strains, † denotes 0 mM IPTG-treated flacp-rny significantly different from all others and 0.05 mM IPTG-treated flacp-rny significantly different from WT and 1.0 mM IPTG flacp-rny strains, ‡ denotes 0 mM IPTG-treated flacp-rny significantly different from all others, # denotes 0 mM IPTG flacp-rny significantly different from WT and 1.0 mM IPTG flacp-rny and 0.05 mM IPTG flacp-rny significantly different from WT, ¶ denotes 0 mM IPTG-treated flacp-rny significantly different from WT and 0.05 mM IPTG-treated flacp-rny significantly different WT and 1.0 mM IPTG-treated flacp-rny. (C) Northern blot analyses of Bb6S RNA levels from the RNA isolated in (A) using a biotinylated RNA probe to Bb6S RNA and tmRNA as a control. The filled arrow marks the size of the mature Bb6S RNA and the open arrow indicates a large (>1000 nt) species present in the sample lacking mature Bb6S RNA, potentially representing an unprocessed Bb6S RNA transcript. (D) qRT-PCR analyses of RNA isolated from the same conditions in (A) using SYBR Green primers 6S 5F and 6S 114R (Table 1) to quantify Bb6S levels. (E) qRT-PCR analyses of the 5′ end of Bb6S using SYBR Green primers 6S U26F and 6S 24R (Table 1) expressed as a ratio of 5′ end to total Bb6S and normalized to the ratio in flacp-rny cells in 0.5 mM IPTG. * denotes significant difference between the 5′ end to Bb6S ratio in the flacp-rny strain in 0.5 mM and 0 mM IPTG (P < 0.05 by one-way ANOVA with Tukey’s post-hoc test). (F) qRT-PCR analyses of the 3′ end of Bb6S using SYBR Green primers 6S 95F and 6S D224R (Table 1) expressed as a ratio of 3′ end to total Bb6S and normalized to the ratio in flacp-rny cells in 0.5 mM IPTG. * denotes significant difference between the 3′ end to Bb6S ratio in the flacp-rny strain in 0 mM IPTG and all other values (P < 0.05 by one-way ANOVA with Tukey’s post-hoc test). Values are the mean of three independent biological replicates and error bars represent the SE in panels D, E and F.
Accumulation of 6S RNA in stationary phase is a hallmark of adaptation in E. coli (Wassarman and Storz, 2000; Cavanagh and Wassarman, 2014; Steuten et al., 2014b). To examine if B. burgdorferi follows suit, Bb6S RNA levels were assayed during growth from early log phase through late stationary phase by Northern blot analyses and qRT-PCR. Bb6S RNA levels did not significantly increase, compared to the flaA control, as spirochete cultures reached stationary phase (Fig. 4A and B). Nutrient starvation in culture has been used to simulate B. burgdorferi persistence in the tick midgut and has been shown to be an important signal for transcriptional regulation (Concepcion and Nelson, 2003; Drecktrah et al., 2015). To assess the effect of nutrient stress on Bb6S RNA levels, cells were grown to stationary phase and starved in RPMI medium for two or six hours before RNA was analyzed by Northern blot and qRT-PCR. Again, Bb6S RNA levels were not significantly affected by starvation in cultured spirochetes (Fig. 4C and D).
Figure 4. Bb6S RNA expression in vitro.
(A) Northern blot and (B) qRT-PCR analyses of Bb6S RNA and flaA mRNA levels from total RNA isolated from wild-type cultures grown at 35°C to different cell densities. (C) Northern blot and (D) qRT-PCR analyses of Bb6S RNA levels from wild-type cells grown to stationary phase (RPMI 0 h) and starved in RPMI for 2 h or 6 h. Values are the mean of three independent biological replicates and error bars represent the SE. P > 0.05 by one-way ANOVA with Tukey’s post-hoc test for comparison of Bb6S levels in both panels B and D.
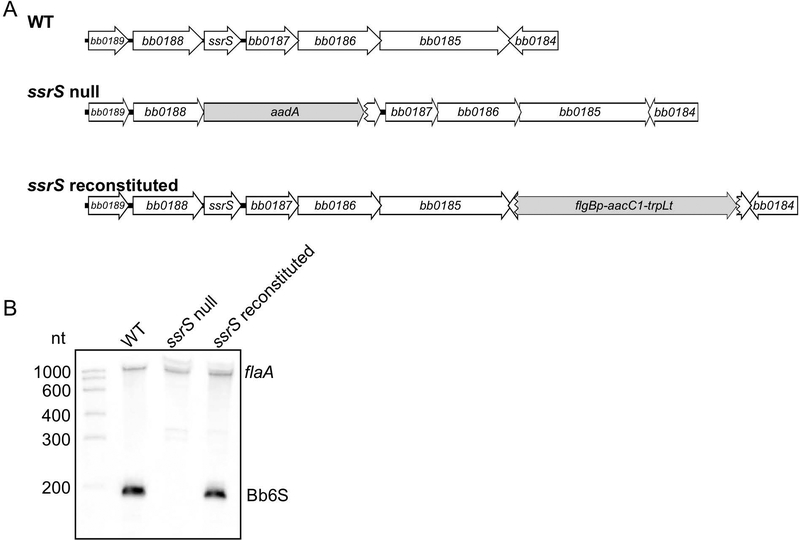
To assess the role of Bb6S RNA in gene regulation during the enzootic cycle of the Lyme disease spirochete, we replaced the ssrS gene encoding Bb6S RNA with a promoterless aadA gene, conferring streptomycin resistance, to yield an ssrS null mutant (Fig. 5A). The ssrS mutant was complemented in cis by genomic reconstitution using the gentamicin resistance gene aacC1 inserted after bb0185, the last gene of a putative operon including ssrS (Fig. 5A). The absence of Bb6S RNA in the null mutant and restoration of Bb6S RNA in the complemented strain was confirmed by Northern blot analysis (Fig. 5B). The plasmid profiles of the ssrS mutant and complemented strains were assessed by PCR analyses, and both were the same as the parental 297 wild type, except that the ssrS complemented strain lost one of the cp32s (Table S1). Additionally, expression of bb0187 and bb0188 (rplT), genes adjacent to the ssrS locus, was unchanged in the ssrS mutant and complemented strains compared to the parental strain as determined by qRT-PCR (Fig. S3), suggesting the absence of polar effects from the genetic manipulations.
Figure 5. Mutation and complementation of ssrS encoding Bb6S RNA in B. burgdorferi.
(A) The ssrS mutant was constructed by replacing the ssrS gene encoding Bb6S RNA with a promoterless streptomycin resistance gene (aadA). The ssrS mutant was complemented in cis, using a gentamicin resistance gene (aacC1) fused to a B. burgdorferi promoter (flgBp) and a B. subtilis terminator (trpLt), to generate the ssrS complemented strain (ssrS reconstituted). (B) Bb6S RNA expression in the wild-type (WT), ssrS null mutant and ssrS complemented strains analyzed by Northern blot of total RNA. Samples were separated on a 6% TBE urea gel, blotted to membrane and hybridized with biotinylated flaA and Bb6S RNA single-stranded RNA probes.
In E. coli, 6S RNA alters the relative levels of σ70-mediated and σS-mediated transcription in response to changing environmental conditions. To examine Bb6S RNA function, we assayed the RpoS-dependent synthesis of outer surface protein C, encoded by ospC, and decorin binding protein A, encoded by dbpA (Schwan et al., 1995; Yang et al., 2000; Hübner et al., 2001; Caimano et al., 2004), which are lipoproteins important for establishing infection in the mammalian host (Fischer et al., 2003; Grimm et al., 2004; Pal et al., 2004; Fingerle et al., 2007; Shi et al., 2008; Weening et al., 2008). A temperature shift from 23°C to 35°C has been experimentally used in cultured spirochetes to induce expression of some RpoS-dependent genes and mimic the transmission from ticks to mammals during the blood meal (Schwan et al., 1995; Obonyo et al., 1999; Yang et al., 2000; Ramamoorthy and Scholl-Meeker, 2001; Alverson et al., 2003; Tokarz et al., 2004). Wild-type, ssrS mutant and ssrS complemented strains were temperature shifted, grown to mid-log phase and total RNA was isolated. Expression of ospC, ospA (encoding outer surface protein A), dbpA, and flaB was assayed by Northern blot analyses. ospC and dbpA mRNA levels increased in wild type but not the ssrS mutant following a temperature shift (Fig. 6A). Expression of ospC and dbpA was restored in the ssrS complemented strain (Fig. 6A). The expression of ospA, a gene repressed, probably indirectly, by RpoS (Caimano et al., 2005), was not affected by Bb6S RNA (Fig. 6A). In addition, the effect of Bb6S RNA on the synthesis of OspC during a temperature shift was examined by immunoblot using anti-OspC antibodies, with FlaB as a control. OspC protein levels also were Bb6S RNA-dependent, as OspC levels increased in the wild type but not in the ssrS mutant (Fig. 6B). OspC induction was restored in the ssrS complemented strain (Fig. 6B). Neither rpoS transcript nor RpoS protein levels were affected by Bb6S RNA (Fig. S4). These results suggest that Bb6S RNA regulates RpoS-dependent gene expression in cultured spirochetes and controls at least a subset of the RpoS regulon, likely by affecting the sigma factor selectivity of RNAP.
Figure 6. Bb6S RNA affects ospC and dbpA gene expression.
(A) Northern blot analyses of total RNA isolated from wild-type (WT), ssrS mutant (ssrS null) and ssrS complemented (ssrS comp) strains temperature-shifted from 23°C and grown at 35°C until mid-log phase. RNA was separated on an 0.8% agarose gel, transferred to membranes and hybridized with 32P-labeled probes to ospC, ospA, dbpA and flaB mRNA (Table 1). (B) Total cell lysates from strains and conditions listed in (A) were separated by SDS-PAGE, transferred to PVDF membranes and analyzed by immunoblot using antibodies against OspC and FlaB.
To test the role of Bb6S RNA in murine infectivity, mice were intradermally injected with increasing doses (102 to 104) of wild-type, ssrS null mutant and ssrS complemented strains. Infection was determined by culturing spirochetes from ear tissues collected three weeks post inoculation and from ear tissues, bladders and tibiotarsal joints five weeks post inoculation. The ssrS mutant was compromised for infectivity in a dose-dependent manner and this defect was completely restored in the ssrS complemented strain (Table 2). Murine infectivity was also followed by seroconversion. Unexpectedly, mice inoculated with 103 ssrS mutant cells, in which live spirochetes could not be recovered (Table 2), seroconverted when examined by immunoblot using serum from the inoculated mice (Fig. 7). Thus, the ssrS null mutant stimulates an adaptive immune response, but does not establish an infection resulting in recoverable live spirochetes from distal sites.
Table 2.
Mouse infectivity of the ssrS null mutant strain.
| Route | Number of cells injected | Strain | 3 weeks | |
|---|---|---|---|---|
| Ear | Ear | |||
| Needlea | 1 × 102 | WT | 1/3 | 1/3 |
| ssrS mutant | 0/3 | 0/3 | ||
| ssrS complement | 1/3 | 1/3 | ||
| 1 × 103 | WT | 3/3 | 2/2 | |
| ssrS mutant | 0/3 | 0/2 | ||
| ssrS complement | 2/3 | 2/3 | ||
| 3 × 103 | WT | 3/3 | 3/3 | |
| ssrS mutant | 0/3 | 0/3 | ||
| ssrS complement | 3/3 | 3/3 | ||
| 1 × 104 | WT | 6/6 | 6/6 | |
| ssrS mutant | 4/6 | 4/6 | ||
| ssrS complement | 3/3 | 3/3 | ||
| Nymph biteb | WT | 3/3 | 3/3 | |
| ssrS mutant | 3/3 | 3/3 | ||
| ssrS complement | 2/2 | 2/2 | ||
| Nymph bitec | WT | 3/3 | 3/3 | |
| ssrS mutant | 3/3 | 3/3 | ||
| ssrS complement | 3/3 | 3/3 |
Intradermal injection.
Mice were infested with five nymphs each.
Mice were infested with two nymphs each.
Figure 7. Serological response of mice injected with the ssrS null mutant.
Whole cell lysates from wild-type (WT), ssrS mutant (ssrS null) and ssrS complemented (ssrS comp) B. burgdorferi, as well as E. coli (Ec) as a negative control, were separated by SDS-PAGE, transferred to membranes and incubated with mouse serum collected five weeks post-injection of 1 × 103 cells of the corresponding strains.
In nature, B. burgdorferi infects mammals via the bite of an ixodid tick. We next investigated the role of Bb6S RNA in tick-transmission using the tick-mouse model of Lyme disease (Barthold et al., 2010). Naive Ixodes larvae were allowed to feed to repletion on mice infected with 1 × 104 wild-type, ssrS null mutant or ssrS complemented spirochetes. Larvae from each mouse were tested one week post-feeding to confirm acquisition of each strain (>80% of larvae were infected for all strains). Fed larvae were allowed to molt into nymphs before nymphs infected with each strain were placed on naive mice and allowed to feed to repletion. Mice were screened three and five weeks post-infestation for infection as described above. The ssrS null mutant was able to transmit to mice by tick bite when either two or five ticks were allowed to feed per mouse, suggesting that Bb6S RNA is not absolutely required for tick transmission, at least in the tick-mouse model (Table 2).
In other bacteria, 6S RNA plays an important role in adapting to environmental stresses, including nutrient limitation (Sharma and Chatterji, 2010; Cavanagh and Wassarman, 2014; Wassarman, 2018). To gain insight into Bb6S RNA regulation, Bb6S RNA levels were followed in Ixodes ticks infected with B. burgdorferi using the tick-mouse model of Lyme disease (Barthold et al., 2010). Briefly, mice were infected with 104 wild-type spirochetes by intradermal needle inoculation. Successful infection was determined by positive cultures from ear biopsies taken three weeks post inoculation. Naive Ixodes scapularis larvae were allowed to feed to repletion on infected mice. Levels of Bb6S RNA were measured by qRT-PCR and normalized to flaB transcript levels in infected larvae (at one week and three weeks post feeding), flat nymphs (at six weeks post feeding) and fed nymphs (at one week after feeding to repletion on naive mice) as previously described (Yang et al., 2004). The amount of Bb6S RNA significantly increased after larval feeding (fed larvae, one week) to reach a maximum in unfed nymphs (Fig. 8). These data suggest Bb6S RNA functions in the persistence of B. burgdorferi in unfed ticks.
Figure 8. Bb6S RNA expression in vivo in ticks.
Bb6S RNA levels in wild type-infected ticks as measured by qRT-PCR from RNA isolated from ticks: naive larvae one week or three weeks post-feeding to repletion on an infected mouse, unfed nymphs one month after molting into nymphs, and fed nymphs one week after feeding to repletion on a naive mouse. Values are the means of at least two independent groups of ticks and error bars represent the SE. * denotes P < 0.05 as determined by one-way ANOVA with a Tukey’s post hoc test.
To address our hypothesis that Bb6S RNA functions in adapting to environmental stresses during persistence in the tick, we followed the acquisition and persistence of the ssrS null mutant through the tick life cycle. Naive larvae were fed to repletion on mice infected with wild-type, ssrS null mutant or ssrS complemented B. burgdorferi, and B. burgdorferi DNA was quantified by qPCR from fed larvae one week post-detachment, unfed nymphs and fed nymphs to determine spirochete loads (genomic equivalents). We found the absence of ssrS had little effect on spirochete loads in the tick; however, there was a slight decrease of B. burgdorferi numbers in fed larvae and unfed nymphs, although only the decrease in nymphs was significant (P < 0.05) and the spirochete load was restored after the nymphs fed (Fig. 9B). Examination of fed larval midguts by immunofluorescence microscopy showed no obvious morphological differences in ticks infected with the ssrS mutant compared to wild-type or ssrS complemented strains (Fig. S5). Thus, the ssrS null mutant successfully persists throughout the larval and nymph stages of the tick life cycle in the animal model of Lyme disease.
Figure 9. Persistence of the ssrS null mutant in ticks.
Quantification of spirochetes in ticks that had fed on mice infected with wild-type (black circles), ssrS null mutant (white circles) or ssrS complemented (gray circles) strains. Total DNA was isolated from larvae that had fed to repletion (fed larvae) or after larvae had molted to nymphs (unfed nymphs) or one week after nymphs had fed to repletion on uninfected mice (fed nymphs). The number of B. burgdorferi genome equivalents per tick was determined by qPCR using TaqMan primers/probe to flaB. Data were analyzed using one-way ANOVA with a Tukey’s post hoc test where * indicates P < 0.05.
Discussion
An sRNA was recently shown to be expressed in the noncoding sRNA transcriptomes of B. burgdorferi (Arnold et al., 2016; Popitsch et al., 2017; Drecktrah et al., 2018) from the region of the genome predicted to encode Bb6S RNA (Barrick et al., 2005). In this study, we present experimental evidence to establish that this sRNA is indeed the canonical 6S RNA: we demonstrate specific binding of Bb6S RNA to RNAP in cells, describe the processing of Bb6S RNA by RNase Y, and examine the role of Bb6S RNA in murine infectivity and persistence in the tick. This is the first study to examine the role and expression of an sRNA in B. burgdorferi through the tick-mouse model of Lyme disease. Furthermore, only a few 6S RNAs have been shown to bind RNAP in vivo; besides for Bb6S RNA, these include the 6S RNAs from E. coli (Wassarman and Storz, 2000), B. subtilis (Trotochaud and Wassarman, 2005) and L. pneumonia (Faucher et al., 2010). We determined that Bb6S RNA is 204 nucleotides and that the 3′ end matches that predicted by Barrick et al. (2005) although the 5′ end differs slightly (Fig. 1B). Our modeling of the Bb6S RNA suggests a secondary structure conserved among bacteria with a single-stranded central bubble flanked by double-stranded stem regions (Barrick et al., 2005; Trotochaud and Wassarman, 2005; Chen et al., 2017), although the upstream stem is shorter than that of E. coli 6S RNA (Fig. 1B).
Many bacteria, such as E. coli, accumulate 6S RNA to adapt to environmental stresses as cells enter stationary phase (Wassarman and Storz, 2000; Sharma and Chatterji, 2010; Cavanagh and Wassarman, 2014; Wassarman, 2018). Bb6S RNA levels did not significantly change in response to any environmental signals examined in culture, including growth phase, even to late stationary phase, and nutrient stress (Fig. 4), which are known to affect 6S RNA levels in some other bacteria (Wassarman and Storz, 2000; Cavanagh and Wassarman, 2014; Steuten et al., 2014b). Similarly, the RNA-seq studies that have identified expression of ssrS (Bb6S) also found no dependence on growth phase or temperature (Arnold et al., 2016; Popitsch et al., 2017). The lack of Bb6S RNA accumulation in stationary phase suggests its regulation may be similar to Bs6S-2 RNA from B. subtilis, which also does not accumulate in a growth phase-dependent manner (Suzuma et al., 2002; Trotochaud and Wassarman, 2005). However, Bb6S RNA levels increased in the tick as fed larvae molted into unfed nymphs and then decreased after nymphs fed to repletion (Fig. 8), which suggests a role in persistence in the tick (Caimano et al., 2016). Furthermore, we have recently shown that RelBbu and the stringent response upregulate Bb6S RNA (Drecktrah et al., 2018), implying that the mechanism of Bb6S RNA accumulation in the tick may involve (p)ppGpp and nutrient deprivation. However, we found the absence of ssrS had little effect on persistence in the tick throughout the larval and nymph stages (Fig. 9). While there is a slight reduction in genomic equivalents of the ssrS null mutant observed in unfed nymphs that is restored in fed nymphs in the tick-mouse model, this phenotype may have more severe consequences for B. burgdorferi in competition with other microbes in the complex environment of the tick midgut in nature. A number of studies have found global transcriptomic changes mediated by signaling pathways, in addition to those mediated by RelBbu (Drecktrah et al., 2015), to be important for persistence in the tick (Caimano et al., 2016; Samuels and Samuels, 2016), including the alternative sigma factor RpoS (Dunham-Ems et al., 2012) and the Hk1/Rrp1 two-component system regulating c-di-GMP levels (Caimano et al., 2011; He et al., 2011; Kostick et al., 2011; Sultan et al., 2011). In E. coli, 6S RNA causes an increase in relA transcription and, subsequently, (p)ppGpp levels (Cavanagh et al., 2010), so there may be positive feedback regulation. Whether the RpoS or c-di-GMP regulatory systems influence Bb6S RNA levels is currently unknown.
The biogenesis of 6S RNA in E. coli involves transcription from two promoters and processing of a long transcript that extends into the ygfA gene downstream of ssrS. RNase E and RNase G, along with exoribonucleases, process the pre-6S RNA to the mature form (Kim and Lee, 2004; Chae et al., 2011). In B. burgdorferi, ygfA is not adjacent to ssrS and the genome lacks homologs of RNase E and RNase G (Fraser et al., 1997; Archambault et al., 2013; Anacker et al., 2018). We found that processing of both the 5′ and 3′ ends of Bb6S RNA requires the endoribonuclease RNase Y (Fig. 3E and F). When rny expression was repressed in the inducible strain, the overall levels of Bb6S RNA were not significantly changed (Fig. 3D), but the mature Bb6S RNA was largely absent while a considerably longer transcript (>1000 nt) containing the Bb6S RNA sequence was visualized (Fig. 3C, open arrow). These results suggest that Bb6S RNA is also processed from a pre-Bb6S RNA transcript. RNase Y degrades mRNA transcripts (Durand et al., 2012; Lehnik-Habrink et al., 2011; Chen et al., 2013) and has not been previously implicated in 6S RNA processing in any other bacteria. However, RNase Y is thought to functionally replace RNase E in Gram-positive bacteria (Shahbabian et al., 2009). Our data further support the hypothesis that RNase Y is the counterpart of RNase E, at least for Bb6S RNA processing. Additionally, we found rny to be an essential gene in B. burgdorferi. Our multiple attempts to generate an rny null mutant were unsuccessful, so, to circumvent this obstacle, we replaced the native rny promoter with the IPTG-inducible promoter flacp (Gilbert et al., 2007). The flacp-rny cells failed to grow once IPTG was removed, but cell numbers and tmRNA levels remained constant for four and two days, respectively, suggesting that a lack of RNase Y is not immediately lethal (Fig. 3B and C). These results add rny to a short list of genes, including bamA (Lenhart and Akins, 2010), dedA (Liang et al., 2010), rrp2 (Groshong et al., 2012), resT (Bandy et al., 2014), bb0028 (Dunn et al., 2015), ftsH (Chu et al., 2016), and oppDF (Groshong et al., 2017), that have been shown to be essential in B. burgdorferi using artificially regulated gene expression. ssrS null mutants are viable, which suggests that RNase Y has a crucial role(s) in B. burgdorferi other than Bb6S RNA maturation, possibly in mRNA turnover.
Bb6S RNA regulates transcript levels of genes with RpoS-dependent promoters that encode proteins known to be essential for host infection (ospC) and dissemination (dbpA), at least in vitro (Fig. 6). This regulation is reminiscent of the canonical 6S RNA function observed in E. coli, although RpoS and its regulon have a different function in B. burgdorferi. In most bacteria, RpoS serves to regulate gene expression to adapt to environmental stresses, such as entering stationary growth phase, while RpoS in B. burgdorferi controls the expression of genes that encode proteins crucial for transmission to and infection of the vertebrate host (Hübner et al., 2001; Caimano et al., 2004; Fisher et al., 2005; Caimano et al., 2007; Ouyang et al., 2008) and, thus, RpoS has been termed the “gatekeeper” of the enzootic cycle (Caimano et al., 2007). Regulation of the RpoS pathway is complex, and includes transcriptional and post-transcriptional mechanisms (Samuels, 2011), and our data suggest that Bb6S regulates levels of at least some RpoS-dependent transcripts (ospC and dbpA), but not RpoS protein levels (Fig. S4B), illuminating yet another factor influencing the RpoS regulon.
The phenotypes of 6S RNA mutants in other bacteria are often subtle (Wassarman and Storz, 2000; Trotochaud and Wassarman, 2004; Trotochaud and Wassarman, 2006; Cavanagh et al., 2012; Cavanagh and Wassarman, 2013); Lp6S mutants of L. pneumophila have an intracellular replication defect only uncovered using a competition assay (Faucher et al., 2010). Needle inoculation with up to 3 × 103 cells of the ssrS null mutant was not infectious as determined by reisolation of spirochetes from murine tissues (Table 2), but the inoculated mice seroconverted (Fig. 7). These data imply that the ssrS mutant was present long enough to trigger an adaptive immune response, although the extent of survival during the infection is unclear. Some non-infectious B. burgdorferi mutants, including ospC (Tilly et al., 2007), guaAB (Jewett et al., 2009), plzA (Kostick-Dunn et al., 2018), and lp36− strains (Jewett et al., 2007), illicit no adaptive immune response in mice. While only vlsE mutants (Bankhead and Chaconas, 2007) and lmp1 mutants (Yang et al., 2009), which lack surface-exposed lipoproteins involved in evading host immunity, seroconverted in the absence of infection. An htrA mutant strain, lacking a virulence-associated protease, elicited a weak immune response without being able to establish infection (Ye et al., 2016). The murine immune system reacted to considerably fewer antigenic proteins in the ssrS null mutant compared to either the wild-type or ssrS complemented strains, which could be due to Bb6S RNA regulating expression of proteins targeted by the adaptive immune system and, possibly, involved in host interactions, but these questions await further investigation. The host infectivity phenotype of the ssrS null mutant can be overcome by increasing the number of inoculated spirochetes to 104 or by transmission via infected nymph bite. These results suggest that Bb6S RNA is not absolutely required for host infection and dissemination, which is not surprising given the subtle phenotypes of ssrS mutants in other bacteria. B. burgdorferi adaptation in the tick or tick factors present during feeding may mitigate the infectivity defect during tick-to-mammal transmission, and demonstrates the importance of the route of infection using animal models to study pathogens.
Bb6S RNA joins a growing list of 6S RNAs that bind RNAP to regulate transcription as bacteria adapt to environmental stress, including that encountered during infection of hosts. Here, we provide the first description of 6S RNA in any spirochete and demonstrate a role for RNase Y in Bb6S maturation.
Experimental Procedures
Ethics Statement
All animal experiments were approved by the University of Montana Institutional Animal Care and Use committee and followed the Guidelines for the Care and Use of Laboratory Animals from the National Institutes of Health.
B. burgdorferi strains and growth conditions
Low-passage B. burgdorferi strains 297 (BbAH130) (Hübner et al., 2001) and B31–5A4 (Purser and Norris, 2000), and genetically manipulated derivatives were grown and maintained in Barbour-Stoenner-Kelly II (BSK) liquid medium, pH 7.6, containing 6% rabbit serum (Pel-Freez Biologicals) (Barbour, 1984) without gelatin unless otherwise noted. Cultures were inoculated at 1 × 103 and grown at 35°C to mid log phase (1 – 3 × 107 cells ml−1), late log phase (4 – 9 × 107 cells ml−1), stationary phase (1 – 2 × 108 cells ml−1) or late stationary phase (≥ 3 × 108 cells ml−1) before RNA and whole-cell protein lysates were collected. B. burgdorferi was starved in RPMI 1640 without L-glutamine and without serum as previously described (Drecktrah et al., 2015). Cell density was determined by enumeration using a Petroff-Hausser cell counting chamber (Samuels et al., 2018). E. coli TOP10F′ grown in lysogeny broth (Bertani, 1951) were used for cloning.
Construction of mutant strains
To generate an ssrS null strain, the gene encoding the B. burgdorferi 6S RNA (Barrick et al., 2005) was replaced with the streptomycin resistance gene aadA (Frank et al., 2003), without a promoter (Sultan et al., 2010), by homologous recombination (Samuels et al., 2018). Genomic regions upstream and downstream of ssrS (encoding Bb6S RNA) were amplified using KOD polymerase (Novagen) with the primers 6S U971F and 6S 5R+AatIIAgeI and primers 6S 109F+AatII and 6S 1325R+AgeI, respectively. PCR products were separated on an agarose gel, extracted using a Qiagen kit, poly(A)-tailed, cloned into pCR2.1-TOPO (Invitrogen), and verified by sequencing. Both constructs were digested with AatII and AgeI, and ligated together. A promoterless streptomycin resistance gene (aadA) flanked by AatII sites was ligated into the synthetic AatII site. This plasmid was linearized with AhdI and electroporated into competent B. burgdorferi 297 as previously described (Drecktrah et al., 2013; Samuels et al., 2018). Transformants were cloned in liquid BSK containing 50 μg ml−1 streptomycin in 96-well plates (Yang et al., 2004). Mutants were confirmed by PCR analysis. To complement the ssrS null strain in cis by genetic reconstitution, two segments of DNA were amplified by PCR using KOD polymerase: a region containing ssrS and upstream sequence using primers 6S U441F and 6S D1564R+AatIIAgeI and a slightly overlapping downstream region with primers 6S D1463F+AatII and 6S D2498R+AgeI. PCR products were prepared as described above, cloned into pCR-XL-TOPO and pCR2.1-TOPO, respectively, and verified by sequencing. The plasmids were digested with AatII and AgeI, separated on an agarose gel, and ligated together. A gentamicin resistance cassette with the flgB promoter from B. burgdorferi (Elias et al., 2002) and the trpL terminator from B. subtilis (Babitzke et al., 1996) (flgBp-aacC1-trpLt) was inserted into the synthetic AatII site (Drecktrah et al., 2015; Samuels et al., 2018). This ssrS-containing plasmid was linearized and transformed into competent ssrS mutant cells as described above.
The B. burgdorferi flacp-rny mutant strain was constructed by replacing the native rny promoter with the inducible promoter flacp (Gilbert et al., 2007). Briefly, primers rny U921F and rny U4R+AatII+AgeI were used to amplify the genomic region upstream of the rny gene and primers rny 1F+NdeI+AatII and rny 1000R+AgeI were used to amplify a portion of the rny gene (bb0504). The two PCR products were cloned into pCR2.1-TOPO. The plasmids were purified, digested with AatII and AgeI, and ligated together. The inducible promoter flacp was fused to the rny gene by digesting with NdeI and AatII, and ligating the promoter into the rny plasmid. The gentamicin resistance cassette flgBp-aacC1 was inserted into the synthetic AatII site. B. burgdorferi strain 297 carrying the lacI gene inserted into bbe02 on the endogenous plasmid lp28–1 (Gilbert et al., 2007) was transformed with linearized DNA containing the flacp-rny construct. Transformed cells were plated in 40 μg ml−1 gentamicin as described above. The promoter fusion was confirmed by PCR analysis.
Immunoprecipitation of RNAP from B. burgdorferi
Co-immunoprecipitation of RNAP and Bb6S RNA was by modification of a previously described protocol (Wassarman and Storz, 2000; Trotochaud and Wassarman, 2005). B. burgdorferi strain B31–5A4 was grown in 100-ml cultures to early stationary phase (1 – 2 × 108 cells ml−1) and collected by centrifugation at 3000 × g, 10 min at 4°C. Cell pellets were washed in BSA-free BSK, resuspended in 200 μl of lysis buffer (20 mM Tris-HCl, pH 8.0, 150 mM KCl, 1 mM MgCl2, 1 mM DTT) and placed in a flat-bottom tube containing 200 μl of 100 μm glass beads. Cells were lysed by 10 cycles of vortexing for 30 s and incubating on ice for 15 s; 400 μl of lysis buffer was added to the lysed cells and centrifuged at 20,800 × g for 10 min at 4°C. Antibody-protein A Sepharose (PAS) complexes were prepared by mixing 2 mg of pre-swollen PAS and 10 μl of serum generated against E. coli core RNAP (WI-151) or 10 μl of preimmune serum in Net2 buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 0.05% Triton X-100). The antibody-PAS mixture was then nutated for 2 h at 23°C and the beads were washed three times at 20,800 × g for 30 s at 4°C in Net2 buffer before addition of 100 μl of B. burgdorferi cell extract supernatant. The mixture was nutated for 2 h at 4°C to allow antibody binding. The antibody-PAS complexes were centrifuged at 20,800 × g for 30 s at 4°C and washed five times with Net 2 Buffer before final resuspension in 500 μl of Net 2 buffer. One hundred μl of the antibody-PAS complexes was removed for immunoblot analyses using antiserum against E. coli core RNAP (WI-151), as described below, and RNA was isolated from the remaining 400 μl to be analyzed by Northern blot. To remove bound protein from the PAS beads, the sample was first centrifuged at 20,800 × g for 30 s at 4°C and 20 μl of 2× SDS-PAGE loading buffer was added to the pelleted beads. The supernatant was used for immunoblot analyses. RNA was isolated by adding 50 μl of 3M NaOAc (pH 5.5), 5 μl of 10% SDS and 600 μl of phenol:chloroform:isoamyl alcohol (25:24:1), pH 8, to the sample, vortexing, and centrifuging at 20,800 × g for 5 min at 23°C, followed by ethanol precipitation.
Growth of the flacp-rny conditional mutant
flacp-rny was cultured at 35°C in the presence of 0.5 mM IPTG to 107 cells ml−1. Two-thirds of the cells were divided into two aliquots, washed twice with BSK, and grown for 48 h at 35°C in 0 or 0.05 mM IPTG. The remaining one-third of the flacp-rny cells were allowed to grow in the presence of 0.5 mM IPTG for 48 h. The parental strain was grown in the presence of 0.5 mM IPTG until late log phase and RNA was isolated using Trizol for qRT-PCR as described below. Additionally, RNA was harvested using hot phenol for Northern blot analyses as previously described (Popitsch et al., 2017).
Immunoblot analyses
Equivalent amounts of total cell lysates collected from B. burgdorferi cultures (1 – 3 × 107 cells ml−1) were analyzed by SDS-PAGE using pre-cast Novex 4–20% Tris-Glycine polyacrylamide gels (Invitrogen) and transferred to PVDF Immobilon membranes (Millipore) as previously described (Drecktrah et al., 2013). Protein levels were analyzed by incubating membranes with rabbit antibodies against OspC (1:1000) or mouse antibodies against FlaB (1:50) followed by goat anti-rabbit or goat anti-mouse HRP-linked antibodies (Bio-Rad Laboratories) (1:5000). Blots were developed by chemiluminescence (Amersham ECL Prime, GE Healthcare) and visualized using an LAS-3000 Intelligent Dark Box (Fujifilm Medical Systems USA).
Seroconversion
To determine if mice seroconverted after intradermal inoculation of wild-type, ssrS null or ssrS complemented strains, blood was collected five weeks post injection. Serum was prepared by allowing blood to clot at 23°C for 45 min, followed by centrifugation at 1500 × g at 4°C for 15 min and collection of the supernatant. Equivalent amounts of cell lysates were analyzed by an immunoblot, as described above, using serum (1:200 dilution) instead of the primary antibodies. E. coli strain DH5α cell lysate was a negative control for immunoreactivity to mouse serum.
qRT-PCR
Total RNA was isolated from B. burgdorferi cultures using TRIzol™; 2 μg of RNA was treated with TURBO DNase as previously described (Lybecker et al., 2010; Drecktrah et al., 2015). The absence of contaminating DNA was determined by PCR analysis using the primers flaB 423F and flaB 542R. One μg of total RNA from each sample was converted to cDNA using the SuperScript III kit (Invitrogen). TaqMan or SYBR Green quantitative PCR (Table 1) was performed with gene transcript copy numbers normalized to flaB or flaA copy number as previously described (Drecktrah et al., 2013; Drecktrah et al., 2015). Values represent the mean ± SEM from three independent experiments.
Table 1.
Oligonucleotides used in this study.a
| Name | Sequence (5′ to 3′) |
|---|---|
| 6S U971F | TTGAGGTTTCTCCTAATGTA |
| 6S 5R+AatIIAgeI | ACCGGTAGCGACGTCTTTTCATCTTCTTATTTCTAAAA |
| 6S 109F+AatII | GACGTCAGCAATTCAGAGGTTAAGA |
| 6S 1325R+AgeI | ACCGGTGGATCAATCTGATTCAAATA |
| 6S U441F | GAGAAATTTAAGGAAATCGG |
| 6S D1564R+AatIIAgeI | ACCGGTCACGACGTCAGCAAAATTAAAAGCCTTTT |
| 6S D1463F+AatII | GACGTCAAGGTGATGATATTTTGTCA |
| 6S D2498R+AgeI | ACCGGTAAGTAGGTCTATTTTCGATG |
| rpoC 3285F | TGCATCTTATGTATTACCAG |
| rpsL U149F+AatII | GACGTCTGGACATTTAATTCCTACTG |
| rpsG 385R+AgeI | ACCGGTATGCATTTAAAAGTTCGTTT |
| rny 467F | AAAGAGATGCTCAAGTTATT |
| rny 1455F+AatII | GACGTCAGAAGCTGAAATGAGATATC |
| rny D2344R+AgeI | ACCGGTCTGGATACCTTTTACAAAG |
| Northern blot probes | |
| 6S 19F | GGGAGGTTTAATTTTAAGAA |
| 6S 180R+T7 | TAATACGACTCACTATAGGCCAACATAAGAACATTTT |
| flaA 64F | GCTCAAGAGACTGATGGATTAGC |
| flaA 284R+T7 | TAATACGACTCACTATAGGCGCAGAAGGAGTAAGTAAAACGCTC |
| rrf 7F | TGGTTAAAGAAAAGAGGAAA |
| rrf 110+T7 | TAATACGACTCACTATAGGCTGGCAATAACCTACTCT |
| ssrA 53F | GGGAATCTCTTAAAACTTCT |
| ssrA 231R+T7 | TAATACGACTCACTATAGGAAGTCCCTAAAAATCAACTT |
| ospC 118R | GCCCTTTAACAGACTCATCAGCAGAATTTGCAGATGTATTCCCATCTTTCCC |
| dbpA 254R | GCCCTTTAACAGACTCATCAGCAGAATTTGCAGATGTATTCCCATCTTTCCC |
| ospA 105R | AGGCAAATCTACTGAAACGCTGTTTTTCTCGTCAAGGCTGCTAACATTTTGC |
| flaB 698R | CCGCCTTGAGAAGGTGCTGTAGCAGGTGCTGGCTGTTGAGC |
| RACE | |
| 6S 131F | AAATTCCGCCTTGAAAAATAGCGCTAAA |
| 6S 155R | AGCGCTATTTTTCAAGGCGGAATTTTTA |
| qRT-PCR (TaqMan) | |
| 6S 96F | CCAAAAGCAATTCAGAGGTT |
| 6S 183R | ACTCCCAAAAAGCCAACATA |
| 6S probe | 6-FAM-AAGAATAAAAATTCCGCCTTGAAAAATAGCGCT-TAM |
| rny 1147F | TGCGGAGAGAGCGAAATTG |
| rny 1247R | ATGGCATCTGCTATTTGAACCA |
| rny probe | 6-FAM-CGCATCACAATGAGGTGAAACCCG-TAM |
| flaB 423F | TTCTCAAAATGTAAGAACAGCTGAAGA |
| flaB 542R | TGGTTTGTCCAACATGAACTC |
| flaB probe | 6-FAM-TCACTTTCAGGGTCTCAAGCGTCTTGGAC-TAM |
| flaA 90F | GGGTTCTAAAAGGCCAGAGC |
| flaA 213R | ACCAGAAGCGCCCGAATATA |
| flaA probe | 6-FAM-TGCCGAGCTTGCAAGAGATCCAAGTTCA-TAM |
| qRT-PCR (SYBR Green) | |
| 6S U62F | TTAGCAATTGAGGATGTTGAAGCTTT |
| 6S 24R | AAATTAAACCTCCCAGGAGCTTTGA |
| 6S 5F | GCTCCTGGGAGGTTTAATTTTAAGAATT |
| 6S 114R | ACCTCTGAATTGCTTTTGGCTATC |
| 6S 95F | GCCAAAAGCAATTCAGAGGTTAAGA |
| 6S D224R | GAGAAATAAGCTGCACAATAATAATCCT |
Restriction sites are underlined and T7 promoter sequences are in bold.
Northern hybridization
Northern blots were probed with either single-stranded RNA as previously described (Lybecker et al., 2010) or oligonucleotides as previously described (Popitsch et al., 2017; Drecktrah et al., 2018). Briefly, Bb6S, flaA, 5S RNA, and tmRNA probes were constructed by PCR amplification of B. burgdorferi genomic DNA using primers 6S 19F and 6S 180R+T7, primers flaA 64F and flaA 284R+T7, primers rrf 7F and rrf 110+T7, and primers ssrA 53F and ssrA 231R+T7, respectively, followed by in vitro transcription with the MEGAScript T7 kit (Ambion) using biotin-16-UTP (Roche) according to the manufacturer’s instructions. DNA was removed from reactions with TURBO DNase (Invitrogen). RNA was precipitated twice with ammonium acetate and washed with 70% ethanol after each precipitation. Probes were separated on a precast 6% TBE urea gel (Invitrogen) and stained with acridine orange. Bands were extracted by overnight incubation in gel extraction buffer (0.5 M ammonium acetate, 1 mM EDTA, 0.1% SDS) at 37°C. Liquid containing the extracted probes was transferred to a fresh tube and precipitated by the addition of three volumes of cold 100% ethanol and incubation at −20°C overnight. RNA was pelleted by centrifugation, washed in 70% ethanol, dried, resuspended in nuclease-free water and stored at −80°C. Three μg of total RNA was separated on a 6% TBE urea gel and transferred to a BrightStar Plus membrane (Ambion). Membranes were hybridized with biotinylated RNA probes overnight at 68°C and washed; the probe was detected with streptavidin-alkaline phosphatase (Invitrogen) diluted 1:10,000. Blots were developed by incubation with CDP-Star chemiluminescent substrate (Applied Biosystems). Images were collected on a Fujifilm LAS-3000.
Oligonucleotide probes to ospC, ospA, dbpA and flaB (Table 1) were 32P-end-labeled using T4 PNK (New England Biolabs) and γ−32P-ATP (Perkin-Elmer). Total RNA was isolated from B. burgdorferi cultures by the hot phenol method (Popitsch et al., 2017). RNA was treated with DNase to remove DNA before 5 μg per sample was loaded and run on an 0.8% agarose-formaldehyde gel in 1× MOPS buffer at 70 V for 3 h. The gel was soaked in nuclease-free water for 10 min to remove the formaldehyde and RNA was transferred from the gel to Hybond XL membranes (Amersham) by capillary action in 20× SSC buffer overnight. The membrane was then UV-crosslinked, blocked in ULTRAhyb Oligo Hybridization Buffer (Invitrogen) for 1 h at 40°C and 32P-labeled probe allowed to hybridize overnight at 40°C. The blot was washed twice with 2× SSC + 0.5% SDS and covered in plastic wrap. Images were obtained with a Fujifilm FLA-3000G phosphorimager.
Identification of 5′ and 3′ ends of Bb6S RNA
The ends of Bb6S RNA were determined by 5′ and 3′ rapid amplification of cDNA ends (RACE) as previously described (Anacker et al., 2018). RNA was isolated from B. burgdorferi cultures grown to late log phase as described above, except that samples were not treated with DNase. The 5′ and 3′ RACE PCR products were separated by agarose gel electrophoresis, stained with ethidium bromide, gel-extracted (Qiagen), and cloned into pCR2.1-TOPO. DNA was isolated with a Qiagen mini-prep kit from positive clones and sequenced.
Quantification of B. burgdorferi in tick
Spirochete loads were quantified in fed larvae (one week and three weeks post feeding, groups of 5), unfed nymphs and fed nymphs (one week post feeding) as previously described (Drecktrah et al., 2015) using the DNeasy Blood/Tissue kit (Qiagen) and TaqMan qPCR with primers and probe to the flaB gene (Table 1).
Immunofluorescence microscopy of infected ticks
B. burgdorferi in infected ticks were visualized by indirect immunofluorescence microscopy as previously described (Hoon-Hanks et al., 2012; Drecktrah et al., 2015). Briefly, dissected tick midguts were fixed in acetone and B. burgdorferi was visualized using mouse anti-Borrelia antibodies (a gift from T. Schwan) followed by goat anti-mouse Alexa Fluor 488 (Invitrogen). Tick cells were stained with wheat germ agglutinin (WGA)-Alexa Fluor 594 (Invitrogen).
Mouse infectivity and tick persistence
The role of Bb6S RNA in murine infectivity was examined as previously described (Barthold et al., 2010) by intradermal injection of female C3H-HeJ mice with 1 × 102 to 1 × 104 cells of wild-type (297 clone AH130), ssrS null mutant or ssrS complemented strains. Infection was determined by culturing mouse ear biopsies in BSK containing 50 μg ml−1 rifampicin, 20 μg ml−1 phosphomycin and 2.5 μg ml−1 amphotericin B and examining cultures by dark-field microscopy for the presence of spirochetes. Five weeks post-infection, mice were sacrificed and ear, ankle and bladder tissues were collected, cultured and examined for spirochetes. Uninfected Ixodes scapularis larvae (National Tick Research and Education Resource, Oklahoma State University) were maintained in a 98% humidified chamber. To allow ticks to acquire B. burgdorferi, approximately 100 larvae per mouse were allowed to feed to repletion. To examine tick acquisition of B. burgdorferi from infected mice, larvae were dissected and processed for immunofluorescence microscopy, as described above. Spirochete loads per tick were quantified by qPCR as described above. After infected larvae molted into nymphs (about 8 weeks), three mice were infested with two or five nymphs each and allowed to feed to repletion. Murine transmission was monitored by culturing ear, ankle and bladder tissues as described above. Persistence of B. burgdorferi in ticks was followed by immunofluorescence microscopy and qPCR as described above.
RNA isolation from infected ticks
Five to ten ticks were placed in a nuclease-free tube containing 100 μl TE, pH 8.0 and crushed with a nuclease-free pestle. Two hundred μg lysozyme (2 μl of 100 mg ml−1) was added and tubes were incubated at 37°C for 10 min. RNA was isolated using a Nucleospin RNA II kit (Macherey-Nagel) according to the manufacturer’s instructions. RNA was processed for qRT-PCR as described above. Bb6S RNA levels were quantified by TaqMan qRT-PCR using the primers and probe listed in Table 1.
Supplementary Material
Acknowledgements
We are grateful to Ron Breaker and Zasha Weinberg for sequence data, Jessica Wexler for assistance with mouse blood collection, Meghan Lybecker for critical reading of the manuscript, Tom Schwan for the anti-Borrelia and anti-FlaB antibodies, George Chaconas, Mike Norgard, and Frank Yang for sharing B. burgdorferi strains, Paul Babitzke for providing the B. subtilis trpL terminator, and Melissa Anacker and Meghan Lybecker for useful discussions.
Funding
This work was supported by Public Health Service grant R01 AI051486 to DSS.
Footnotes
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Conflict of Interest Statement
The authors declare no conflict of interest.
References
- Alverson J, Bundle SF, Sohaskey CD, Lybecker MC, and Samuels DS (2003) Transcriptional regulation of the ospAB and ospC promoters from Borrelia burgdorferi. Mol. Microbiol 48: 1665–1677. [DOI] [PubMed] [Google Scholar]
- Anacker ML, Drecktrah D, LeCoultre RD, Lybecker M, and Samuels DS (2018) RNase III processing of rRNA in the Lyme disease spirochete Borrelia burgdorferi. J. Bacteriol 200: e00035–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ando Y, Asari S, Suzuma S, Yamane K, and Nakamura K (2002) Expression of a small RNA, BS203 RNA, from the yocI-yocJ intergenic region of Bacillus subtilis genome. FEMS Microbiol. Lett 207: 29–33. [DOI] [PubMed] [Google Scholar]
- Archambault L, Borchert JS, Bergeron J, Snow S, and Schlax PJ (2013) Measurements of mRNA degradation in Borrelia burgdorferi. J. Bacteriol 195: 4879–4887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnold WK, Savage CR, Brissette CA, Seshu J, Livny J, and Stevenson B (2016) RNA-seq of Borrelia burgdorferi in multiple phases of growth reveals insights into the dynamics of gene expression, transcriptome architecture, and noncoding RNAs. PLoS One 11: e0164165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Axmann IM, Holtzendorff J, Voß B, Kensche P, and Hess WR (2007) Two distinct types of 6S RNA in Prochlorococcus. Gene 406: 69–78. [DOI] [PubMed] [Google Scholar]
- Babitzke P, Yealy J, and Campanelli D (1996) Interaction of the trp RNA-Binding attenuation protein (TRAP) of Bacillus subtilis with RNA: effects of the number of GAG repeats, the nucleotides separating adjacent repeats, and RNA secondary structure. J. Bacteriol 178: 5159–5163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bandy NJ, Salman-Dilgimen A, and Chaconas G (2014) Construction and characterization of a Borrelia burgdorferi strain with conditional expression of the essential telomere resolvase, ResT. J. Bacteriol 196: 2396–2404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bankhead T, and Chaconas G (2007) The role of VlsE antigenic variation in the Lyme disease spirochete: persistence through a mechanism that differs from other pathogens. Mol. Microbiol 65: 1547–1558. [DOI] [PubMed] [Google Scholar]
- Barbour AG (1984) Isolation and cultivation of Lyme disease spirochetes. Yale J. Biol. Med 57: 521–525. [PMC free article] [PubMed] [Google Scholar]
- Barrick JE, Sudarsan N, Weinberg Z, Ruzzo WL, and Breaker RR (2005) 6S RNA is a widespread regulator of eubacterial RNA polymerase that resembles an open promoter. RNA 11: 774–784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barthold SW, Cadavid D, and Philipp MT, (2010) Animal models of borreliosis In: Borrelia: Molecular Biology, Host Interaction and Pathogenesis. Samuels DSand Radolf JD(eds). Norfolk, UK: Caister Academic Press, pp. 359–411. [Google Scholar]
- Beckmann BM, Burenina OY, Hoch PG, Kubareva EA, Sharma CM, and Hartmann RK (2011) In vivo and in vitro analysis of 6S RNA-templated short transcripts in Bacillus subtilis. RNA Biol 8: 839–849. [DOI] [PubMed] [Google Scholar]
- Bertani G (1951) Studies on lysogenesis: I. The mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol 62: 293–300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bugrysheva JV, Pappas CJ, Terekhova DA, Iyer R, Godfrey HP, Schwartz I, et al. (2015) Characterization of the RelBbu regulon in Borrelia burgdorferi reveals modulation of glycerol metabolism by (p)ppGpp. PLoS One 10: e0118063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burenina OY, Elkina DA, Hartmann RK, Oretskaya TS, and Kubareva EA (2015) Small noncoding 6S RNAs of bacteria. Biochemistry (Mosc) 80: 1429–1446. [DOI] [PubMed] [Google Scholar]
- Burenina OY, Hoch PG, Damm K, Salas M, Zatsepin TS, Lechner M, et al. (2014) Mechanistic comparison of Bacillus subtilis 6S-1 and 6S-2 RNAs–commonalities and differences. RNA 20: 348–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caimano MJ, Drecktrah D, Kung F, and Samuels DS (2016) Interaction of the Lyme disease spirochete with its tick vector. Cell. Microbiol 18: 919–927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caimano MJ, Dunham-Ems S, Allard AM, Cassera MB, Kenedy M, and Radolf JD (2015) Cyclic di-GMP modulates gene expression in Lyme disease spirochetes at the tick-mammal interface to promote spirochete survival during the blood meal and tick-to-mammal transmission. Infect. Immun 83: 3043–3060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caimano MJ, Eggers CH, Gonzalez CA, and Radolf JD (2005) Alternate sigma factor RpoS is required for the in vivo-specific repression of Borrelia burgdorferi plasmid lp54-borne ospA and lp6.6 genes. J. Bacteriol 187: 7845–7852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caimano MJ, Eggers CH, Hazlett KRO, and Radolf JD (2004) RpoS is not central to the general stress response in Borrelia burgdorferi but does control expression of one or more essential virulence determinants. Infect. Immun 72: 6433–6445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caimano MJ, Iyer R, Eggers CH, Gonzalez C, Morton EA, Gilbert MA, et al. (2007) Analysis of the RpoS regulon in Borrelia burgdorferi in response to mammalian host signals provides insight into RpoS function during the enzootic cycle. Mol. Microbiol 65: 1193–1217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caimano MJ, Kenedy MR, Kairu T, Desrosiers DC, Harman M, Dunham-Ems S, et al. (2011) The hybrid histidine kinase Hk1 is part of a two-component system that is essential for survival of Borrelia burgdorferi in feeding Ixodes scapularis ticks. Infect. Immun 79: 3117–3130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavanagh AT, Chandrangsu P, and Wassarman KM (2010) 6S RNA regulation of relA alters ppGpp levels in early stationary phase. Microbiology 156: 3791–3800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavanagh AT, Klocko AD, Liu X, and Wassarman KM (2008) Promoter specificity for 6S RNA regulation of transcription is determined by core promoter sequences and competition for region 4.2 of σ70. Mol. Microbiol 67: 1242–1256. [DOI] [PubMed] [Google Scholar]
- Cavanagh AT, Sperger JM, and Wassarman KM (2012) Regulation of 6S RNA by pRNA synthesis is required for efficient recovery from stationary phase in E. coli and B. subtilis. Nucleic Acids Res 40: 2234–2246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavanagh AT, and Wassarman KM (2013) 6S-1 RNA function leads to a delay in sporulation in Bacillus subtilis. J. Bacteriol 195: 2079–2086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cavanagh AT, and Wassarman KM (2014) 6S RNA, a global regulator of transcription in Escherichia coli, Bacillus subtilis, and beyond. Annu. Rev. Microbiol 68: 45–60. [DOI] [PubMed] [Google Scholar]
- Chae H, Han K, Kim K, Park H, Lee J, and Lee Y (2011) Rho-dependent termination of ssrS (6S RNA) transcription in Escherichia coli: implication for 3′ processing of 6S RNA and expression of downstream ygfA (putative 5-formyl-tetrahydrofolate cyclo-ligase). J. Biol. Chem 286: 114–122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen H, Dutta T, and Deutscher MP (2016) Growth phase-dependent variation of RNase BN/Z affects small RNAs: regulation of 6S RNA. J. Biol. Chem 291: 26435–26442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J, Wassarman KM, Feng S, Leon K, Feklistov A, Winkelman JT, et al. (2017) 6S RNA mimics B-form DNA to regulate Escherichia coli RNA polymerase. Mol. Cell 68: 388–397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Z, Itzek A, Malke H, Ferretti JJ, and Kreth J (2013) Multiple roles of RNase Y in Streptococcus pyogenes mRNA processing and degradation. J. Bacteriol 195: 2585–2594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu C-Y, Stewart PE, Bestor A, Hansen B, Lin T, Gao L, et al. (2016) Function of the Borrelia burgdorferi FtsH homolog is essential for viability both in vitro and in vivo and independent of HflK/C. mBio 7: e00404–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Concepcion MB, and Nelson DR (2003) Expression of spoT in Borrelia burgdorferi during serum starvation. J. Bacteriol 185: 444–452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drecktrah D, Hall LS, Hoon-Hanks LL, and Samuels DS (2013) An inverted repeat in the ospC operator is required for induction in Borrelia burgdorferi. PLoS One 8: e68799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drecktrah D, Hall LS, Rescheneder P, Lybecker M, and Samuels DS (2018) The stringent response-regulated sRNA transcriptome of Borrelia burgdorferi. Front. Cell. Infect. Microbiol 8: 231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drecktrah D, Lybecker M, Popitsch N, Rescheneder P, Hall LS, and Samuels DS (2015) The Borrelia burgdorferi RelA/SpoT homolog and stringent response regulate survival in the tick vector and global gene expression during starvation. PLoS Pathog 11: e1005160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunham-Ems SM, Caimano MJ, Eggers CH, and Radolf JD (2012) Borrelia burgdorferi requires the alternative sigma factor RpoS for dissemination within the vector during tick-to-mammal transmission. PLoS Pathog 8: e1002532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunn JP, Kenedy MR, Iqbal H, and Akins DR (2015) Characterization of the β-barrel assembly machine accessory lipoproteins from Borrelia burgdorferi. BMC Microbiol 15: 70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durand S, Gilet L, and Condon C (2012) The essential function of B. subtilis RNase III is to silence foreign toxin genes. PLoS Genet 8: e1003181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elias AF, Stewart PE, Grimm D, Caimano MJ, Eggers CH, Tilly K, et al. (2002) Clonal polymorphism of Borrelia burgdorferi strain B31 MI: implications for mutagenesis in an infectious strain background. Infect. Immun 70: 2139–2150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faucher SP, Friedlander G, Livny J, Margalit H, and Shuman HA (2010) Legionella pneumophila 6S RNA optimizes intracellular multiplication. Proc. Natl. Acad. Sci. USA 107: 7533–7538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fingerle V, Goettner G, Gern L, Wilske B, and Schulte-Spechtel U (2007) Complementation of a Borrelia afzelii OspC mutant highlights the crucial role of OspC for dissemination of Borrelia afzelii in Ixodes ricinus. Int. J. Med. Microbiol 297: 97–107. [DOI] [PubMed] [Google Scholar]
- Fischer JR, Parveen N, Magoun L, and Leong JM (2003) Decorin-binding proteins A and B confer distinct mammalian cell type-specific attachment by Borrelia burgdorferi, the Lyme disease spirochete. Proc. Natl. Acad. Sci. USA 100: 7307–7312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher MA, Grimm D, Henion AK, Elias AF, Stewart PE, Rosa PA, et al. (2005) Borrelia burgdorferi σ54 is required for mammalian infection and vector transmission but not for tick colonization. Proc. Natl. Acad. Sci. USA 102: 5162–5167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frank KL, Bundle SF, Kresge ME, Eggers CH, and Samuels DS (2003) aadA confers streptomycin-resistance in Borrelia burgdorferi. J. Bacteriol 185: 6723–6727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fraser CM, Casjens S, Huang WM, Sutton GG, Clayton R, Lathigra R, et al. (1997) Genomic sequence of a Lyme disease spirochete, Borrelia burgdorferi. Nature 390: 580–586. [DOI] [PubMed] [Google Scholar]
- Gilbert MA, Morton EA, Bundle SF, and Samuels DS (2007) Artificial regulation of ospC expression in Borrelia burgdorferi. Mol. Microbiol 63: 1259–1273. [DOI] [PubMed] [Google Scholar]
- Grimm D, Tilly K, Byram R, Stewart PE, Krum JG, Bueschel DM, et al. (2004) Outer-surface protein C of the Lyme disease spirochete: a protein induced in ticks for infection of mammals. Proc. Natl. Acad. Sci. USA 101: 3142–3147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groshong AM, Dey A, Bezsonova I, Caimano MJ, and Radolf JD (2017) Peptide uptake is essential for Borrelia burgdorferi viability and involves structural and regulatory complexity of its oligopeptide transporter. mBio 8: e02047–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Groshong AM, Gibbons NE, Yang XF, and Blevins JS (2012) Rrp2, a prokaryotic enhancer-like binding protein, is essential for viability of Borrelia burgdorferi. J. Bacteriol 194: 3336–3342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He M, Ouyang Z, Troxell B, Xu H, Moh A, Piesman J, et al. (2011) Cyclic di-GMP is essential for the survival of the Lyme disease spirochete in ticks. PLoS Pathog 7: e1002133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoon-Hanks LL, Morton EA, Lybecker MC, Battisti JM, Samuels DS, and Drecktrah D (2012) Borrelia burgdorferi malQ mutants utilize disaccharides and traverse the enzootic cycle. FEMS Immunol. Med. Microbiol 66: 157–165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hübner A, Yang X, Nolen DM, Popova TG, Cabello FC, and Norgard MV (2001) Expression of Borrelia burgdorferi OspC and DbpA is controlled by a RpoN-RpoS regulatory pathway. Proc. Natl. Acad. Sci. USA 98: 12724–12729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iyer R, and Schwartz I (2016) Microarray-based comparative genomic and transcriptome analysis of Borrelia burgdorferi. Microarrays 5: 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jewett MW, Lawrence K, Bestor AC, Tilly K, Grimm D, Shaw P, et al. (2007) The critical role of the linear plasmid lp36 in the infectious cycle of Borrelia burgdorferi. Mol. Microbiol 64: 1358–1374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jewett MW, Lawrence KA, Bestor A, Byram R, Gherardini F, and Rosa PA (2009) GuaA and GuaB are essential for Borrelia burgdorferi survival in the tick-mouse infection cycle. J. Bacteriol 191: 6231–6241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joly N, Engl C, Jovanovic G, Huvet M, Toni T, Sheng X, et al. (2010) Managing membrane stress: the phage shock protein (Psp) response, from molecular mechanisms to physiology. FEMS Microbiol. Rev 34: 797–827. [DOI] [PubMed] [Google Scholar]
- Kim K, and Lee Y (2004) Regulation of 6S RNA biogenesis by switching utilization of both sigma factors and endoribonucleases. Nucleic Acids Res 32: 6057–6068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kostick JL, Szkotnicki LT, Rogers EA, Bocci P, Raffaelli N, and Marconi RT (2011) The diguanylate cyclase, Rrp1, regulates critical steps in the enzootic cycle of the Lyme disease spirochetes. Mol. Microbiol 81: 219–231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kostick-Dunn JL, Izac JR, Freedman JC, Szkotnicki LT, Oliver LD Jr., and Marconi RT (2018) The Borrelia burgdorferi c-di-GMP binding receptors, PlzA and PlzB, are functionally distinct. Front Cell Infect Microbiol 8: 213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee CA, Fournier MJ, and Beckwith J (1985) Escherichia coli 6S RNA is not essential for growth or protein secretion. J. Bacteriol 161: 1156–1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, and Stülke J (2011) RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol. Microbiol 81: 1459–1473. [DOI] [PubMed] [Google Scholar]
- Lenhart TR, and Akins DR (2010) Borrelia burgdorferi locus BB0795 encodes a BamA orthologue required for growth and efficient localization of outer membrane proteins. Mol. Microbiol 75: 692–709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Z, Pandit S, and Deutscher MP (1998) 3′ exoribonucleolytic trimming is a common feature of the maturation of small, stable RNAs in Escherichia coli. Proc. Natl. Acad. Sci. USA 95: 2856–2861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang FT, Xu Q, Sikdar R, Xiao Y, Cox JS, and Doerrler WT (2010) BB0250 of Borrelia burgdorferi is a conserved and essential inner membrane protein required for cell division. J. Bacteriol 192: 6105–6115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lybecker MC, Abel CA, Feig AL, and Samuels DS (2010) Identification and function of the RNA chaperone Hfq in the Lyme disease spirochete Borrelia burgdorferi. Mol. Microbiol 78: 622–635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lybecker MC, and Samuels DS (2017) Small RNAs of Borrelia burgdorferi: characterizing functional regulators in a sea of sRNAs. Yale J. Biol. Med 90: 317–323. [PMC free article] [PubMed] [Google Scholar]
- Neusser T, Polen T, Geissen R, and Wagner R (2010) Depletion of the non-coding regulatory 6S RNA in E. coli causes a surprising reduction in the expression of the translation machinery. BMC Genomics 11: 165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Obonyo M, Munderloh UG, Fingerle V, Wilske B, and Kurtti TJ (1999) Borrelia burgdorferi in tick cell culture modulates expression of outer surface proteins A and C in response to temperature. J. Clin. Microbiol 37: 2137–2141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ouyang Z, Blevins JS, and Norgard MV (2008) Transcriptional interplay among the regulators Rrp2, RpoN and RpoS in Borrelia burgdorferi. Microbiology 154: 2641–2658. [DOI] [PubMed] [Google Scholar]
- Pal U, Yang X, Chen M, Bockenstedt LK, Anderson JF, Flavell RA, et al. (2004) OspC facilitates Borrelia burgdorferi invasion of Ixodes scapularis salivary glands. J. Clin. Invest 113: 220–230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peeters E, Sass A, Mahenthiralingam E, Nelis H, and Coenye T (2010) Transcriptional response of Burkholderia cenocepacia J2315 sessile cells to treatments with high doses of hydrogen peroxide and sodium hypochlorite. BMC Genomics 11: 90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Popitsch N, Bilusic I, Rescheneder P, Schroeder R, and Lybecker M (2017) Temperature-dependent sRNA transcriptome of the Lyme disease spirochete. BMC Genomics 18: 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purser JE, and Norris SJ (2000) Correlation between plasmid content and infectivity in Borrelia burgdorferi. Proc. Natl. Acad. Sci. USA 97: 13865–13870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radolf JD, Caimano MJ, Stevenson B, and Hu LT (2012) Of ticks, mice and men: understanding the dual-host lifestyle of Lyme disease spirochaetes. Nat. Rev. Microbiol 10: 87–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramamoorthy R, and Scholl-Meeker D (2001) Borrelia burgdorferi proteins whose expression is similarly affected by culture temperature and pH. Infect. Immun 69: 2739–2742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren J, Sang Y, Qin R, Cui Z, and Yao Y-F (2017) 6S RNA is involved in acid resistance and invasion of epithelial cells in Salmonella enterica serovar Typhimurium. Future Microbiol 12: 1045–1057. [DOI] [PubMed] [Google Scholar]
- Rogers EA, Terekhova D, Zhang H-M, Hovis KM, Schwartz I, and Marconi RT (2009) Rrp1, a cyclic-di-GMP-producing response regulator, is an important regulator of Borrelia burgdorferi core cellular functions. Mol. Microbiol 71: 1551–1573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samuels DS (2011) Gene regulation in Borrelia burgdorferi. Annu. Rev. Microbiol 65: 479–499. [DOI] [PubMed] [Google Scholar]
- Samuels DS, Drecktrah D, and Hall LS, (2018) Genetic transformation and complementation In: Borrelia burgdorferi: Methods and Protocols. Pal U and Buyuktanir O(eds). New York, NY: Humana Press, pp. 183–200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samuels DS, and Samuels LRN (2016) Gene regulation during the enzootic cycle of the Lyme disease spirochete. For. Immunopathol. Dis. Therap 7: 205–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwan TG, Piesman J, Golde WT, Dolan MC, and Rosa PA (1995) Induction of an outer surface protein on Borrelia burgdorferi during tick feeding. Proc. Natl. Acad. Sci. USA 92: 2909–2913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shahbabian K, Jamalli A, Zig L, and Putzer H (2009) RNase Y, a novel endoribonuclease, initiates riboswitch turnover in Bacillus subtilis. EMBO J 28: 3523–3533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma UK, and Chatterji D (2010) Transcriptional switching in Escherichia coli during stress and starvation by modulation of σ70 activity. FEMS Microbiol. Rev 34: 646–657. [DOI] [PubMed] [Google Scholar]
- Shi Y, Xu Q, McShan K, and Liang FT (2008) Both decorin-binding proteins A and B are critical for the overall virulence of Borrelia burgdorferi. Infect. Immun 76: 1239–1246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steuten B, Hoch PG, Damm K, Schneider S, Köhler K, Wagner R, et al. (2014a) Regulation of transcription by 6S RNAs: insights from the Escherichia coli and Bacillus subtilis model systems. RNA Biol 11: 508–521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steuten B, Schneider S, and Wagner R (2014b) 6S RNA: recent answers–future questions. Mol. Microbiol 91: 641–648. [DOI] [PubMed] [Google Scholar]
- Storz G, Vogel J, and Wassarman KM (2011) Regulation by small RNAs in bacteria: expanding frontiers. Mol. Cell 43: 880–891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sultan SZ, Pitzer JE, Boquoi T, Hobbs G, Miller MR, and Motaleb MA (2011) Analysis of the HD-GYP domain cyclic-di-GMP phosphodiesterase reveals a role in motility and enzootic life cycle of Borrelia burgdorferi. Infect. Immun 79: 3273–3283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sultan SZ, Pitzer JE, Miller MR, and Motaleb MA (2010) Analysis of a Borrelia burgdorferi phosphodiesterase demonstrates a role for cyclic-di-guanosine monophosphate in motility and virulence. Mol. Microbiol 77: 128–142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuma S, Asari S, Bunai K, Yoshino K, Ando Y, Kakeshita H, et al. (2002) Identification and characterization of novel small RNAs in the aspS-yrvM intergenic region of the Bacillus subtilis genome. Microbiology 148: 2591–2598. [DOI] [PubMed] [Google Scholar]
- Tilly K, Bestor A, Jewett MW, and Rosa P (2007) Rapid clearance of Lyme disease spirochetes lacking OspC from skin. Infect. Immun 75: 1517–1519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tokarz R, Anderton JM, Katona LI, and Benach JL (2004) Combined effects of blood and temperature shift on Borrelia burgdorferi gene expression as determined by whole genome DNA array. Infect. Immun 72: 5419–5432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trotochaud AE, and Wassarman KM (2004) 6S RNA function enhances long-term cell survival. J. Bacteriol 186: 4978–4985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trotochaud AE, and Wassarman KM (2005) A highly conserved 6S RNA structure is required for regulation of transcription. Nat. Struct. Mol. Biol 12: 313–319. [DOI] [PubMed] [Google Scholar]
- Trotochaud AE, and Wassarman KM (2006) 6S RNA regulation of pspF transcription leads to altered cell survival at high pH. J. Bacteriol 188: 3936–3943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warrier I, Hicks LD, Battisti JM, Raghavan R, and Minnick MF (2014) Identification of novel small RNAs and characterization of the 6S RNA of Coxiella burnetii. PLoS One 9: e100147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wassarman KM (2018) 6S RNA, a global regulator of transcription. Microbiol. Spectr 6: RWR-0019–2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wassarman KM, and Storz G (2000) 6S RNA regulates E. coli RNA polymerase activity. Cell 101: 613–623. [DOI] [PubMed] [Google Scholar]
- Waters LS, and Storz G (2009) Regulatory RNAs in bacteria. Cell 136: 615–628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weening EH, Parveen N, Trzeciakowski JP, Leong JM, Höök M, and Skare JT (2008) Borrelia burgdorferi lacking DbpBA exhibits an early survival defect during experimental infection. Infect. Immun 76: 5694–5705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weissenmayer BA, Prendergast JGD, Lohan AJ, and Loftus BJ (2011) Sequencing illustrates the transcriptional response of Legionella pneumophila during infection and identifies seventy novel small non-coding RNAs. PLoS One 6: e17570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan Y, Su S, Meng X, Ji X, Qu Y, Liu Z, et al. (2013) Determination of sRNA expressions by RNA-seq in Yersinia pestis grown in vitro and during infection. PLoS One 8: e74495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X, Coleman AS, Anguita J, and Pal U (2009) A chromosomally encoded virulence factor protects the Lyme disease pathogen against host-adaptive immunity. PLoS Pathog 5: e1000326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X, Goldberg MS, Popova TG, Schoeler GB, Wikel SK, Hagman KE, et al. (2000) Interdependence of environmental factors influencing reciprocal patterns of gene expression in virulent Borrelia burgdorferi. Mol. Microbiol 37: 1470–1479. [DOI] [PubMed] [Google Scholar]
- Yang XF, Pal U, Alani SM, Fikrig E, and Norgard MV (2004) Essential role for OspA/B in the life cycle of the Lyme disease spirochete. J. Exp. Med 199: 641–648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye M, Sharma K, Thakur M, Smith AA, Buyuktanir O, Xiang X, et al. (2016) HtrA, a temperature- and stationary phase-activated protease involved in maturation of a key microbial virulence determinant, facilitates Borrelia burgdorferi infection in mammalian hosts. Infect. Immun 84: 2372–2381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31: 1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.