Abstract
Undoubtedly, viruses represent a major threat faced by human and veterinary medicines and by agronomy. The rapid evolution of viruses enables them to escape from natural immunities and from state-of-the-art antiviral treatments, with new viruses periodically emerging with deadly consequences. Viruses have also become powerful and are increasingly used tools in the field of experimental evolution. A growing body of evidence points that the evolution of viruses is mainly determined by key features such as their compacted genomes, enormous population sizes, and short generation times. In addition, RNA viruses also present large selection coefficients, antagonistic epistasis, and high mutation rates. Most of this knowledge comes from studies that have used either bacteriophages or animal viruses in cell cultures as experimental systems. However, plant viruses provide almost identical advantages for evolutionary studies and, in addition, offer an invaluable tool for studying the interplay between viruses and pluricellular hosts. Without seeking to be exhaustive, here we summarize some peculiarities of plant viruses and review recent experiments that have explored important questions on evolution, such as the role of deleterious mutation and neutrality, the effect of different transmission modes in the evolution of virulence, and the heterogeneous selective constraints imposed by multiple hosts.
Keywords: bottlenecks, fitness, host-range, neutral networks, virulence, virus evolution
Introduction
RNA viruses are the most abundant parasites infecting humans, farm animals, and cultivated plants (Domingo and Holland, 1997). Despite tremendous economical efforts, the number of eradicated viruses is quite limited. Furthermore, the perspectives for future eradications are counterbalanced by the emergence of new viruses (Murphy and Nathanson, 1994). The fact that few viruses can be effectively controlled and the pervasive emergence of new viruses are likely consequences of their remarkable ability to evolve. Immune-escaping and antiviral-resistant strains arise soon after viral populations are challenged (Althaus and Bonhoeffer, 2005) and viruses may jump the species barrier from their natural reservoir hosts to new ones (Daszak et al., 2000). Besides seeking information on these questions, the study of RNA virus evolution may help in addressing fundamental evolutionary questions. What properties make RNA viruses excellent models for experimental evolution? First, as a consequence of the lack of proofreading of their RNA-dependent RNA polymerases (RdRp), they show mutation rates within the range ∼2 × 10−5 to 1 × 10−4 per nucleotide per replication event (Drake and Holland, 1999); being variable is a key factor for pathogenicity. Second, genomes are small (for example, ranging from the 3.6 kb of bacteriophage MS2 up to the ∼30 kb of the coronaviruses), with many cases of overlapping reading frames or multifunctional proteins. Abundant molecular, functional, and structural information facilitates the analysis of the molecular basis of evolution. Third, replication rates are fast, and thus most RNA viruses reach very large population sizes after short periods of time. Plant viruses have the added value of making experiments with real hosts easy compared with experiments with animal viruses, which traditionally have been restricted to cell cultures due to the ethical and economical concerns inherent to the infection of large numbers of animals. Moreover, not only the evolution of the virus, but also the evolution of host--virus interactions, including the expression of virulence, can be studied in a natural framework. Finally, as opposed to experiments with bacterial viruses, plant viruses allow the study of complex eukaryotic multicellular hosts.
In this review, we will focus on recent studies that used plant viruses to address evolutionary questions of general interest, such as the rate and fitness effects of deleterious mutations and the role of neutrality as a source of mutational robustness, the evolution of generalist viruses, or the effect of vertical versus horizontal transmission on virulence. All these topics have also been explored using bacteriophages and/or animal viruses; to keep our review within bounds we shall not refer to this literature. Interested readers may take a look at the exhaustive review on experimental virus evolution by Elena and Sanjuán (2007).
Replication kinetics and mutation rate
As any other ribovirus that encodes an RdRp lacking proofreading activity, plant riboviruses have the potential to generate extremely variable populations. A recent study provided evidence for this high mutation rate. Malpica et al. (2002) infected Nicotiana tabacum plants, transgenic for the movement protein (MP) of Tobacco mosaic tobamovirus (TMV), with TMV and characterized the mutational spectrum for the replicating viral population at the MP locus. Since MP is provided in trans by the plant, the virus-encoded copy acts as a neutral marker wherein all sorts of mutations can be accumulated without affecting viral movement. The mutational spectrum was dominated by insertions and deletions, with only 31% being single nucleotide substitutions and with transitions and transversions arising at approximately the same rate. In nature, insertion/deletion mutations would be almost immediately lethal, hence explaining why these are found to be typically rare in natural variability studies. The genomic mutation rate, U, inferred was in the range 0.043–0.063 per replication event. However, if these estimates were corrected to account for a less efficient rescue of lethal mutants (Malpica et al., 2002), they would become twice as large (∼0.1), which although still high, is approximately sevenfold lower than the estimates previously reported for lytic riboviruses (Drake and Holland, 1999).
Despite mutation rate is still high compared to that of DNA-based microorganisms, a classic field observation is that natural plant virus populations generally exhibit limited genetic variation (García-Arenal et al., 2001), which may imply either that purifying selection may be strong or that genome replication occurs mainly by Luria's stamping machine model (Luria, 1951) rather than exponentially (French and Stenger, 2003), the two hypotheses being nonexclusive. For TMV, a positive strand virus, the RdRp copies the first positive strand into several negative strands that are used as templates to produce multiple progeny-positive strands that are encapsidated to form new virions. The bulk of positive strands are synthesized in a linear process. The implications of the linear versus exponential growth are relevant to understanding the genesis of variability. In contrast, under the stamping machine model the frequency of mutation-free genomes among the N produced within an infected cell is given by the Poisson null class e−U whereas for exponential growth this frequency depends on k, the number of replication events required to produce N viral genomes per cell, e−kU. Thus, exponential replication produces (1−e−kU)/(1−e−U) more mutant genomes per infected cell. Using the above estimates of U for TMV and assuming that N=40 per cell (Malpica et al., 2002), 339–350% more mutant genomes would be produced if replication was purely exponential. On the other hand, exponential replication has the advantage of being faster. Therefore, a tradeoff exists between the necessity of replicating faster and that of maintaining mutational load at as low a level as possible. It has been postulated that, since lysis is not the final fate of infected plant cells but instead viral particles accumulate in the cytoplasm forming densely packed bodies, selection for shorter lysis times may have been relaxed, thus giving advantage to a stamping machine replication mode (French and Stenger, 2003).
Genetic drift and viral movement
Genetic bottlenecks unavoidably occur at two different stages during the infectious cycle of plant viruses. First, it has been extensively shown that bottlenecks take place during plant virus systemic movement between leaves of the same plant, the number of viral particles being propagated on the order of tens or less for TMV (Sacristán et al., 2003) and Cucumber mosaic cucumovirus (CMV) (Li and Roossinck, 2004) or as low as four for Wheat streak mosaic tritimovirus (WSMV) (French and Stenger, 2003). Furthermore, these bottlenecks contribute to the establishment of genetically divergent populations in different parts of the plant and the generation of patched viral populations (French and Stenger, 2003; Jridi et al., 2006). Second, during aphid-mediated plant-to-plant transmission, strong bottlenecks have also been reported for CMV (Ali et al., 2006). Apparently, the bottleneck was not only due to the sampling of the donor plant during aphid feeding but also to the inoculation of the receiving plants (Ali et al., 2006).
The fitness effect of deleterious mutations
Associated with the stochastic nature of the aforementioned transmission or tissue colonization events, viral populations are continuously regenerated from a few individual genomes, and given the high mutation rate, a nonnegligible probability exists that deleterious mutations become fixed in the new population. Repeated genetic bottleneck events followed by low-fidelity genome replication have been shown to produce substantial fitness losses in several RNA viruses, in a process similar to Muller's ratchet (Elena and Sanjuán, 2007).
Recently, de la Iglesia and Elena (2007) have extended these observations to Tobacco etch potyvirus (TEV). In this experiment, bottlenecks of size one were performed by inoculating the nonsystemic host Chenopodium quinoa and by isolating the viral progeny contained in one randomly chosen local necrotic lesion. Local necrotic lesions, similar to what happens in the lytic plaques formed by bacteriophages in a bacterial loan, are generated from a single infectious unit. Figure 1 shows the dynamics of fitness decline for the independent mutation accumulation lineages evolved in this experiment. After 11 consecutive transfers, all lineages suffered significant fitness losses, and one became extinct. The per day average fitness decline was Uds̄=5.2%, where Ud stands for the deleterious genomic mutation rate per transfer and s̄ for the average mutational effect. In a similar experiment with WSMV, a phylogenetically related virus, Hall et al. (2001) observed that the number of fixed mutations increased with bottleneck passages. Therefore, if purifying selection is not efficient because of small population size, the mutations accumulated are mostly deleterious.
Figure 1.
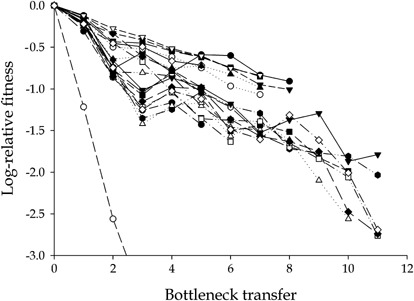
Evolution of TEV fitness upon serial bottleneck transfers. Each symbol and line represents a different mutation accumulation lineage. Fitness was expressed as the ratio between the Malthusian parameter estimated for each clone and the value obtained to the ancestral TEV. (From de la Iglesia and Elena (2007)).
Although informative, mutation accumulation experiments suffer from the problem that the number of mutations fixed per clone and its molecular nature remain unknown and the fraction of lethal mutations is undetectable. Therefore, inferences are only possible for the distribution of accumulated deleterious (but viable) effects. A straightforward way to avoid this problem is to create a collection of single-nucleotide substitution mutants by site-directed mutagenesis on an infectious cDNA and then measure fitness for each member of the collection to infer the statistical properties of the distribution of mutational fitness effects. This approach was recently undertaken by Carrasco et al. (2007a, 2007b) to explore the distribution of mutational effects on the fitness of TEV. These authors created a collection of 66 clones each carrying a different, randomly chosen, single nucleotide substitution. The fitness of each clone was measured by competition experiments against a surrogated wild type. Table 1 shows a summary of the results of this experiment. Most mutations were lethal, with nonlethal mutations causing s̄=51% fitness reduction. Given the structural compactness and functional nonredundancy of RNA viruses, it is not unexpected that most mutations will either be lethal or have a strong deleterious fitness effect.
Table 1.
Proportion of lethal, deleterious, and neutral effects on fitness
| Proportion (%) | Average effect (%) | |
|---|---|---|
| Lethal | 40.9 | −100 |
| Deleterious | 36.4 | −51.1±4.3 |
| Neutral | 22.7 |
[From Carrasco et al. (2007b)].
Combining the estimate of Uds̄ obtained by de la Iglesia and Elena (2007) and the estimate of s̄ obtained by Carrasco et al. (2007b), it is possible to infer an estimate for TEV deleterious mutation rate as Ud=0.1, a value that is consistent with the one reported for TMV (Malpica et al., 2002).
Neutrality and mutational robustness
When the mutation rate is so high that most newly generated individuals are genetically contaminated, as it appears to be the case for RNA viruses, fitness can be maximized by reducing the impact of mutations on fitness, that is, making organisms more robust, even at the cost of a lower replication rate (van Nimwegen et al., 1999; Wilke et al., 2001). However, whether robustness is an evolved property or intrinsic to genetic systems is largely unknown, because the mechanistic basis for robustness would depend on the peculiarities of each system (Elena et al., 2006). One of the simplest situations in which mutational robustness evolves in a population is represented by the quasi-species model of molecular evolution (Schuster and Swetina, 1988). In a molecular quasi-species, selection and mutation interact in a subtle way: selection should maximize the average replication rate of the swarm of genotypes interconnected by mutation rather than favoring any one genotype that has the fastest replication. However, since a faster replication also implies the production of more mutant genomes with lower fitness, as a consequence, mutation would act as a selective agent that shapes the entire genome so that it becomes robust with respect to mutation. In other words, populations would move on genotypic space toward regions of higher neutrality (van Nimwegen et al., 1999). Thus, in a highly mutagenic environment, a quasi-species occupying a low but neutral and highly connected (that is, flat) region in the fitness landscape should outcompete a quasi-species located at a higher but narrower fitness peak in which most surrounding mutants are unfit (van Nimwegen et al., 1999; Wilke et al., 2001). This phenomenon has been called the quasi-species effect or, more recently, the survival of the flattest (Wilke et al., 2001), and despite its interest, it lacked strong empirical support.
Codoñer et al. (2006) used two viroid species; subviral plant pathogens whose genomes are constituted by a single small noncoding RNA molecule, to test this prediction. These two viroid species, which belonged to different families and are characterized by different secondary structures, competed in the same host plants. One of the viroids, Chrysanthemum stunt pospiviroid (CSVd), is characterized by fast population growth and genetic homogeneity, whereas the other, Chrysanthemum chlorotic mottle avsunviroid (CChMVd), shows slow population growth and a high degree of variation. Furthermore, the in silico size of the neutral neighborhood, that is, the number of point mutations that do not affect the predicted RNA folding, for CChMVd was found to be twice as large as that of CSVd. As expected, under standard nonmutagenic infection conditions, the faster replicator CSVd outcompeted the slower CChMVd, being the fitness difference of ∼9% per week (Figure 2). However, as predicted by the quasi-species theory, when coinfected plants were irradiated with mutagenic doses of UVC light, CSVd was unable to outcompete the slower but more robust CChMVd (Figure 2), suggesting that the competitive advantage of CSVd vanished as a consequence of the twofold larger mutational neutral neighborhood surrounding CChMVd's optimum genotype.
Figure 2.
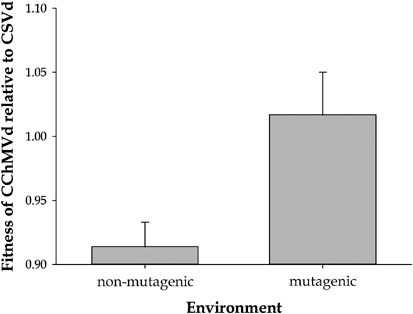
Fitness of a robust viroid, CChMVd, relative to a fit one, CSVd, in non-mutagenic and in mutagenic environmental conditions. Error bars represent standard errors. (From Codoñer et al., 2006).
Host range and the specificity of adaptation
Broadly speaking, plant viruses have highly variable host ranges: some infect only one or a few related species (that is, they are specialists) whereas others can infect a wide range of hosts from different taxonomic groups (that is, they are generalists). The advantages of generalism are not well understood; it has been suggested that selection should favor specialist parasites because there are tradeoffs limiting the fitness of generalists in any of the alternative hosts or because evolution proceeds faster with narrower niches (Woolhouse et al., 2001). It is widely accepted that adaptation to a specific environment is often coupled with fitness losses in alternative environments simply because mutations that are beneficial in the first might be deleterious in the alternative (Kawecki, 1994). This antagonistic pleiotropy could limit the range of adaptation and promotes the evolution of specialization. In a recent field study, Malpica et al. (2006) found that five different plant viruses (from four viral families) were unevenly distributed across 21 weed species, some viruses being more prevalent on a given host than on alternative hosts, regardless of the relative abundance of each host. These observations suggest that host specialization may be a successful strategy for plant viruses.
Many studies can be found in the literature in which the qualitative costs associated with viruses expanding their host range from sensitive to resistant plant genotypes had been explored. Jenner et al. (2002) went a step farther and quantified the fitness penalty on wild-type plants paid by Turnip mosaic potyvirus (TuMV) after expanding its host range from wild-type turnips to plants bearing the TuRB01 resistance gene. The fitness of three resistance-breaking (that is, generalists) TuMV genotypes, CZE1, CDN1, and v35Tunos+5570A>G, was evaluated on wild-type turnips by multiday competition assays against the wild-type specialist isolate UK1. The fitness costs associated with host range were 32.36% for CZE1, 53.91% for v35Tunos+5570A>G, and 100% for CDN1. These results support the idea that whatever mutation confers the ability to infect a new host has a strong pleiotropic effect on the ancestral host.
One of the most remarkable examples of fast host-specific adaptation is the convergent evolution of Pelargonium flower break carmovirus (PFBV) populations adapted to C. quinoa (Rico et al., 2006). PFBV isolates maintained for a long time on quinoa leaves, by the staff of German Collection of Microorganisms and Cell Cultures, have five specific noncontiguous amino acid substitutions in the coat protein (CP), which were not present in other natural isolates. The quinoa-specific pattern of amino acids at the relevant sites is VFYII. When a Spanish isolate from geranium, with the wild-type pattern ASHMV, was mechanically inoculated on quinoa leaves, the viral population generated soon after the first passage had already fixed two of the quinoa-specific changes (ASYMI), and only four serial passages were necessary to fix the entire VFYII quinoa pattern (Rico et al., 2006). The fact that this pattern has never been found in the natural host, even partially, shows the heavy burden it may impose for viral fitness on geranium (Rico et al., 2006).
Several studies have also reported that the amount of virus genetic diversity is driven by constraints imposed by the host in as much as it is driven by its own genomic properties. For example, the amount of genetic variability in evolving CMV and TMV populations remained close to an equilibrium value throughout serial passages in a constant host (Schneider and Roossinck, 2000, 2001). However, variability was severely reduced after transferring the populations to new hosts, suggesting the existence of a bottleneck imposed by the new host, and subsequently. Variation was subsequently regenerated de novo and eventually reached a new equilibrium, which was characteristic of the new host and different from the ancestral one (Schneider and Roossinck, 2000, 2001).
Mode of transmission and virulence
Current theories for the evolution of virus virulence suggest that natural selection may favor a wide range of virulences depending on the mode of virus transmission (Ewald, 1994). When a virus is vertically transmitted (that is, from host parents to offspring), a reduction in virulence is expected because viral fitness is tightly linked to host reproductive success: the longer and healthier the host lives, the more offspring it may deliver (Lipsitch et al., 1996). By contrast, when the mode of transmission is mostly horizontal (that is, between unrelated hosts), selection would favor those viral strains with increased infectivity even at the cost of being more virulent (Lipsitch et al., 1996). Testing these predictions has been difficult because it requires having a parasite with both modes of transmission. This problem was overcome by Stewart et al. (2005) using Barley stripe mosaic hordeivirus (BSMV), a virus capable of infecting barleys, both horizontally and vertically. After four generations of horizontal transmission, on average, the virulence (for example, reduction in lifespan production of viable seeds) of BSMV increased ∼166% (Figure 3a) and the infectivity increased ∼78% (Figure 3b). In addition, a significant positive correlation was observed between virulence and infectivity, as hypothesized. By contrast, after three generations of vertical transmission, BSMV populations were ∼68% less virulent (Figure 3a) but ∼12% more efficient at transmitting vertically. These results clearly support the above expectation that virulence depends on the relative opportunities for horizontal versus vertical transmission. Two more interesting findings can be drawn from this study. First, as BSMV evolved increased horizontal infectivity, the overall rate of vertical transmission declined (Figure 3b) and the rate of horizontal transmission increased, suggesting that it might not be possible to improve simultaneously both transmission routes (Stewart et al., 2005). Second, changes in virulence were uncorrelated with changes in virus accumulation within hosts, indicating that virulence was not a direct consequence of BSMV replication and accumulation but it may be mediated by complex plant responses.
Figure 3.
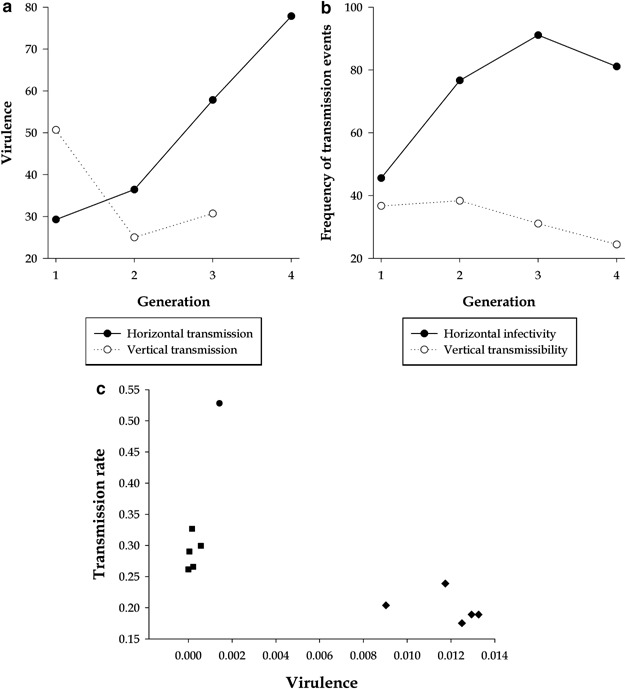
(a) Evolution of virulence in horizontally and vertically transmitted BSMV populations. (From Stewart et al., 2005). (b) Tradeoff between the efficiency of horizontal and vertical transmissions for BSMV populations evolved upon a horizontal transmission scheme. (From Stewart et al., 2005). (c) Correlation between virulence and transmission rate in CMV. Different symbols represent different CMV genotypes. (From Escriu et al., 2003).
The existence of a tradeoff between virulence and the rate of horizontal transmission has also been postulated (Ewald, 1994). The rationale for this tradeoff is that virus-induced host mortality shortens the period during which the virus can be taken up by vectors and transmitted from one host to another. Therefore, all else being equal, in a natural situation it may be costly for the virus to harm its plant host. To test this prediction, Escriu et al. (2003) performed field and laboratory studies seeking to understand the evolution of virulence (for example, reduction in lifespan biomass) and epidemiology of different CMV strains. The data gathered by these authors support the predicted negative correlation between virulence and the rate of transmission (Figure 3c: partial correlation coefficient controlling for strain type: r=−0.96, d.f.=8, P<0.001). In other words, in natural conditions, necrogenic CMV strains may jeopardize their own transmission by aphids by killing the plant, or making it less attractive to the vector insects, too quickly. However, in good agreement with the BSMV study, CMV virulence was not dependent on its accumulation level since necrogenic and nonnecrogenic variants accumulated to similar levels (Escriu et al., 2000), thus suggesting that virulence may be a side effect of the interaction between the virus and plant defense mechanisms.
Conclusions
In this contribution, we have briefly reviewed some recent advances in plant virus experimental evolution. Our goal was not to provide an exhaustive review of all the relevant literature but, instead, to illustrate the advantages of plant RNA viruses as model systems for experimental evolution studies. Traditionally, animal viruses and, in particular, bacteriophages have been widely used for testing evolutionary hypotheses, whereas plant viruses have received less attention. Here we have shown that the same questions can be also addressed using plant RNA viruses (for example, the pace and effect of deleterious mutations on fitness or the specificity of adaptation) but, in addition, new exciting questions have also been properly addressed (for example, evolution of virulence in a natural host--parasite system and the role of neutrality on evolution).
Plant RNA viruses open promising new avenues for experimental evolution. Basic evolutionary problems such as the dynamics of adaptation and genetic diversification, the evolution of viral takeover of cellular metabolism, the evolution of virulence during dynamics of multiple infection, testing gene-for-gene coevolutionary models, the evolution of mutational robustness, and coevolutionary interactions between multiple viruses infecting the same host, among others, will be tackled in the near future using plant viruses.
Acknowledgements
We thank José A Daròs for helping us in setting up the lab for plant virus work and for fruitful discussion and Fernando García-Arenal for many enlightening discussions about plant virus evolution. The technical assistance provided by Francisca de la Iglesia and Ana Cuadrado is warmly appreciated. The Ministerio de Educación y Ciencia-FEDER (Grant BFU2006-14819-C02-01/BMC), the Generalitat Valenciana (Grant ACOMP07/263) and the EMBO Young Investigator Program provided financial support.
References
- Ali A, Li H, Schneider WL, Sherman DJ, Gray S, Smith D. Analysis of genetic bottlenecks during horizontal transmission of Cucumber mosaic virus. J Virol. 2006;80:8345–8350. doi: 10.1128/JVI.00568-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Althaus CL, Bonhoeffer S. Stochastic interplay between mutation and recombination during the acquisition of drug resistance mutations in Human immunodeficiency virus type 1. J Virol. 2005;79:13572–13578. doi: 10.1128/JVI.79.21.13572-13578.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrasco P, Daròs JA, Agudelo-Romero P, Elena SF. A real-time RT-PCR assay for quantifying the fitness of Tobacco etch virus in competition experiments. J Virol Meth. 2007;139:181–188. doi: 10.1016/j.jviromet.2006.09.020. [DOI] [PubMed] [Google Scholar]
- Carrasco P, de la Iglesia F, Elena SF. The distribution of fitness and virulence effects caused by single-nucleotide substitutions in Tobacco etch virus. J Virol. 2007;81:12979–12984. doi: 10.1128/JVI.00524-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Codoñer FM, Daròs JA, Solé RV, Elena SF. The fittest versus the flattest: experimental confirmation of the quasispecies effect with subviral pathogens. PLoS Pathog. 2006;2:e136. doi: 10.1371/journal.ppat.0020136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daszak P, Cunningham AA, Hyatt AD. Emerging infectious diseases of wildlife—threats to biodiversity and human health. Science. 2000;287:443–449. doi: 10.1126/science.287.5452.443. [DOI] [PubMed] [Google Scholar]
- de la Iglesia F, Elena SF. Fitness declines in Tobacco etch virus upon serial bottleneck transfers. J Virol. 2007;81:4941–4947. doi: 10.1128/JVI.02528-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Domingo E, Holland JJ. RNA virus mutations and fitness for survival. Annu Rev Microbiol. 1997;51:151–178. doi: 10.1146/annurev.micro.51.1.151. [DOI] [PubMed] [Google Scholar]
- Drake JW, Holland JJ. Mutation rates among RNA viruses. Proc Natl Acad Sci USA. 1999;96:13910–13913. doi: 10.1073/pnas.96.24.13910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elena SF, Carrasco P, Daròs JA, Sanjuán R. Mechanisms of genetic robustness in RNA viruses. EMBO Rep. 2006;7:168–173. doi: 10.1038/sj.embor.7400636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elena SF, Sanjuán R. Virus evolution: insights from an experimental approach. Annu Rev Ecol Evol Syst. 2007;38:27–52. doi: 10.1146/annurev.ecolsys.38.091206.095637. [DOI] [Google Scholar]
- Escriu F, Fraile A, García-Arenal F. Evolution of virulence in natural populations of the satellite RNA of Cucumber mosaic virus. Phytopathology. 2000;90:480–485. doi: 10.1094/PHYTO.2000.90.5.480. [DOI] [PubMed] [Google Scholar]
- Escriu F, Fraile A, García-Arenal F. The evolution of virulence in a plant virus. Evolution. 2003;57:755–765. doi: 10.1111/j.0014-3820.2003.tb00287.x. [DOI] [PubMed] [Google Scholar]
- Ewald PW. The Evolution of Infectious Diseases. Oxford University Press: Oxford; 1994. [Google Scholar]
- French R, Stenger DC. Evolution of Wheat streak mosaic virus: dynamics of population growth within plants may explain limited variation. Annu Rev Phytopathol. 2003;41:199–214. doi: 10.1146/annurev.phyto.41.052002.095559. [DOI] [PubMed] [Google Scholar]
- García-Arenal F, Fraile A, Malpica JM. Variability and genetic structure of plant virus populations. Annu Rev Phytopathol. 2001;39:157–186. doi: 10.1146/annurev.phyto.39.1.157. [DOI] [PubMed] [Google Scholar]
- Hall JS, French R, Morris TJ, Stenger DC. Structure and temporal dynamics of populations within Wheat streak mosaic virus isolates. J Virol. 2001;75:10231–10243. doi: 10.1128/JVI.75.21.10231-10243.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenner CE, Wang X, Ponz F, Walsh JA. A fitness cost for Turnip mosaic virus to overcome host resistance. Virus Res. 2002;86:1–6. doi: 10.1016/S0168-1702(02)00031-X. [DOI] [PubMed] [Google Scholar]
- Jridi C, Martin JF, Mareie-Jeanne V, Labonne G, Blanc S. Distinct viral populations differentiate and evolve independently in a single perennial host plant. J Virol. 2006;80:2349–2357. doi: 10.1128/JVI.80.5.2349-2357.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawecki TJ. Accumulation of deleterious mutations and the evolutionary cost of being a generalist. Am Nat. 1994;144:833–838. doi: 10.1086/285709. [DOI] [Google Scholar]
- Li H, Roossinck MJ. Genetic bottlenecks reduce population variation in an experimental RNA virus population. J Virol. 2004;78:10582–10587. doi: 10.1128/JVI.78.19.10582-10587.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lipsitch M, Siller S, Nowak MA. The evolution of virulence in pathogens with vertical and horizontal transmission. Evolution. 1996;50:1729–1741. doi: 10.1111/j.1558-5646.1996.tb03560.x. [DOI] [PubMed] [Google Scholar]
- Luria SE. The frequency distribution of spontaneous bacteriophage mutants as evidence for the exponential rate of phage reproduction. Cold Spring Harbor Symp Quant Biol. 1951;16:463–470. doi: 10.1101/SQB.1951.016.01.033. [DOI] [PubMed] [Google Scholar]
- Malpica JM, Fraile A, Moreno I, Obies CI, Drake JW, García-Arenal F. The rate and character of spontaneous mutation in an RNA virus. Genetics. 2002;162:1505–1511. doi: 10.1093/genetics/162.4.1505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malpica JM, Sacristán S, Fraile A, García-Arenal F. Association and host selectivity in multi-host pathogens. PLoS ONE. 2006;1:e41. doi: 10.1371/journal.pone.0000041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murphy FA, Nathanson N. The emergence of new virus diseases: an overview. Sem Virol. 1994;5:87–102. doi: 10.1006/smvy.1994.1010. [DOI] [Google Scholar]
- Rico P, Ivars P, Elena SF, Hernández C. Insights into the selective pressures restricting Pelargonium flower break virus genome variability: evidence for host adaptation. J Virol. 2006;80:8124–8132. doi: 10.1128/JVI.00603-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sacristán S, Malpica JM, Fraile A, García-Arenal F. Estimation of population bottlenecks during systemic movement of Tobacco mosaic virus in tobacco plants. J Virol. 2003;77:9906–9911. doi: 10.1128/JVI.77.18.9906-9911.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider WL, Roossinck MJ. Evolutionarily related Sindbis-like plant viruses maintain different levels of population diversity in a common host. J Virol. 2000;74:3130–3134. doi: 10.1128/JVI.74.7.3130-3134.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider WL, Roossinck MJ. Genetic diversity in RNA virus quasispecies is controlled by host-virus interactions. J Virol. 2001;75:6566–6571. doi: 10.1128/JVI.75.14.6566-6571.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schuster P, Swetina J. Stationary mutant distributions and evolutionary optimization. Bull Math Biol. 1988;50:635–660. doi: 10.1007/BF02460094. [DOI] [PubMed] [Google Scholar]
- Stewart AD, Logsdon GM, Jr, Kelley SE. An empirial study of the evolution of virulence under both horizontal and vertical transmission. Evolution. 2005;59:730–739. doi: 10.1111/j.0014-3820.2005.tb01749.x. [DOI] [PubMed] [Google Scholar]
- van Nimwegen E, Crutchfield JP, Huynen M. Neutral evolution of mutational robustness. Proc Natl Acad Sci USA. 1999;96:9716–9720. doi: 10.1073/pnas.96.17.9716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilke CO, Wang JL, Ofria C, Lenski RE, Adami C. Evolution of digital organisms at high mutation rate leads to survival of the flattest. Nature. 2001;412:331–333. doi: 10.1038/35085569. [DOI] [PubMed] [Google Scholar]
- Woolhouse MEJ, Taylor LH, Haydon DT. Population biology of multihost pathogens. Science. 2001;292:1109–1112. doi: 10.1126/science.1059026. [DOI] [PubMed] [Google Scholar]


