Abstract
Background
DNA transposons are ubiquitous components of eukaryotic genomes. Academ superfamily of DNA transposons is one of the least characterized DNA transposon superfamilies in eukaryotes. DNA transposons belonging to the Academ superfamily have been reported from various animals, one red algal species Chondrus crispus, and one fungal species Puccinia graminis. Six Academ families from P. graminis encode a helicase in addition to putative transposase, while some other families encode a single protein which contains a putative transposase and an XPG nuclease.
Results
Systematic searches on Repbase and BLAST searches against publicly available genome sequences revealed that several species of fungi and animals contain multiple Academ transposon families encoding a helicase. These AcademH families generate 9 or 10-bp target site duplications (TSDs) while Academ families lacking helicase generate 3 or 4-bp TSDs. Phylogenetic analysis clearly shows two lineages inside of Academ, designated here as AcademH and AcademX for encoding helicase or XPG nuclease, respectively. One sublineage of AcademH in animals encodes plant homeodomain (PHD) finger in its transposase, and its remnants are found in several fish genomes.
Conclusions
The AcademH lineage of TEs is widely distributed in animals and fungi, and originated early in the evolution of Academ DNA transposons. This analysis highlights the structural diversity in one less studied superfamily of eukaryotic DNA transposons.
Keywords: DNA transposon, Academ, Helicase, Target site duplication, Transposase, Plant homeodomain
Introduction
Transposable elements (TEs), or transposons are ubiquitous components of genomes in all three domains of life [1, 2]. TEs are traditionally classified into 2 classes: Class I retrotransposons and Class II DNA transposons [3]. Autonomous retrotransposons encode a reverse transcriptase and during the transposition, the information of RNA is transformed into DNA by reverse transcription. DNA transposons do not have a process of reverse transcription in their transposition cycle. At least 5 independent DNA-cleaving/recombining enzymes (DDE transposase or DDD/E transposase, tyrosine recombinase, serine recombinase, HUH nuclease, and Cas1 endonuclease) have been incorporated into TEs and related mobile genetic elements [4, 5]. DDE transposase or integrase is the most ubiquitous enzyme that functions as transposase of DNA transposons, as well as of long terminal repeat (LTR) retrotransposons and of retroviruses [6]. Eukaryotic DNA transposons are now classified into around 20 superfamilies [1]. Most of these superfamilies, such as Mariner/Tc1 and Harbinger/PIF1, are known to encode a DDE transposase.
DDE transposase is topologically a member of RNaseH-like fold [6]. The conserved core of the transposase domain is β1-β2-β3-α1-β4-α2/3-β5-α4-α5. Three acidic residues, DDD or DDE play the central role in the transposition. The first D is located on β1 and the second D on or just after β4. The last D or E is on or just before α4. In the case of the integrase encoded by human immunodeficiency virus type 1 (HIV-1), the distance between the second D and the last E is 35 residues. In some DNA transposons, the catalytic core domain between β5 and α4 is extended by “insertion domain.” In the case of RAG1, recombination activating gene 1, which originated from an eukaryotic DNA transposon superfamily Transib [7], the insertion domain is 264 residues in length and entirely α-helical [6]. The transposase encoded by Hermes, a member of eukaryotic DNA transposon superfamily hAT, contains a 288-aa-long insertion domain [6].
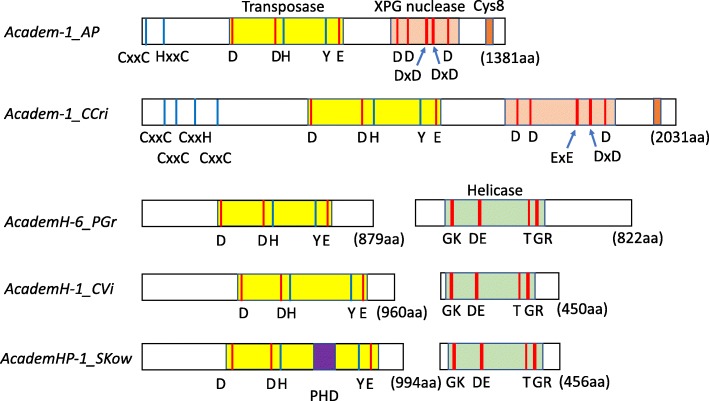
The Academ superfamily of eukaryotic DNA transposons was first described by Kapitonov and Jurka [8] from various animals. To date, Academ has been found from animals, fungi, and plants [1]. In animals, Academ is widely distributed and found from genomes of 7 phyla: Chordata, Hemichordata, Echinodermata, Annelida, Mollusca, Arthropoda, and Cnidaria. In contrast, in fungi and in plants, only one species of each group is reported to have Academ transposons: a red alga Chondrus crispus [9] and a pathogenic fungus Puccinia graminis [10], while the wide distribution of Academ in fungi was suggested [11]. The transposase domain of Academ is predicted to be a DDE transposase [12]. An entirely α-helical insertion domain was predicted between β5 and α4, as are the cases of RAG1 and Hermes. Another insertion domain was predicted between β2 and β3, unlike any other transposases. Many of Academ families encode a large protein that contains three recognizable domains, a transposase, an XPG nuclease, and a putative Cys8 zinc finger [8] (Fig. 1). These three domains can be recognized among Academ families from animals and C. crispus. The Academ families from P. graminis do not encode an XPG nuclease. Instead, they encode a superfamily II helicase as a separate protein [10] (Fig. 1). This lineage was designated as AcademH. It is not yet known whether the presence of helicase is a recently acquired characteristic specific for Academ families from P. graminis, or it is an ancient trait shared by various Academ families from diverse organisms.
Fig. 1.
Schematic structures of proteins encoded by Academ transposons. Encoded proteins are shown as open boxes, and inside them, protein domains are shown as colored boxes. Conserved residues are shown as bars in red and blue. Protein lengths are shown in parentheses. Transposase and helicase proteins of AcademH families are encoded in opposite directions
In this study, many families of AcademH from other fungi and animals were characterized. No intact AcademH transposons were found from vertebrates, but some fish genomes still contain remnants of AcademH transposons. AcademH shows 9 or 10-bp target site duplications (TSDs), although other Academ shows 3 or 4-bp TSDs. The sequence comparison and phylogenetic analysis revealed two independent lineages with different TSD length and protein composition inside of the Academ DNA transposons.
Results
Academ families encoding a superfamily II helicase
Manual inspection of Repbase entries revealed that besides 6 AcademH families from P. graminis, Academ-1_ADi, Academ-2_ADi, Academ-3_ADi from the coral Acropora digitifera, and Academ-2_CGi from the Pacific oyster Crassostrea gigas also encode a superfamily II helicase protein. Using the helicase protein sequences from these families as queries, Censor search [13] against published genome sequences was performed. It led to the characterization of AcademH families encoding a helicase protein from 7 species of basidiomycetes fungi (Laccaria bicolor, Puccinia coronata, Puccinia horiana, Puccinia striiformis, Puccinia triticina, Serpula lacrymans), one species of fungi in Mucoromycotina (Lobosporangium transversale), and another oyster Crassostrea virginica, in addition to more families from the three species above (Table 1 and Supplementary Dataset S1). Non-autonomous DNA transposons showing similarity in terminal regions with AcademH families were also found from three cnidarians (Exaiptasia pallida, Orbicella faveolata, and Stylophora pistillata) and the Yesso scallop Mizuhopecten yessoensis (Table 1 and Supplementary Dataset S1).
Table 1.
AcademH distribution
| Classification | Organism | AcademH families |
|---|---|---|
| Fungi/Basidiomycota | ||
| -Pucciniomycetes/Pucciniales | Puccinia coronata | AcademH-1_PCor to 17_PCor, AcademH-14N1_PCor, AcademH-N1_PCor to N13_PCor |
| Puccinia graminis | AcademH-1_PG to 6_PG, AcademH-N2_PGr to N6_PGr, AcademH-2B_PGr, AcademH-N3B_PGr | |
| Puccinia horiana | AcademH-1_PHor, AcademH-N1_PHor to N25_PHor | |
| Puccinia sorghi | AcademH-N1_PSor | |
| Puccinia striiformis | AcademH-1_PSt to 18_PSt, AcademH-N1_PSt to N12_PSt | |
| Puccinia triticina 1–1 BBBD Race 1 | AcademH-1_PTrit to 6_PTrit, AcademH-1B_PTrit, AcademH-N1_PTrit to N20_PTrit | |
| Melampsora larici-populina | AcademH-N1_MLP | |
| -Agaricomycetes/Auriculariales | Exidia glandulosa HHB12029 | AcademH-1_ExGl |
| -Agaricomycetes/Atheliales | Fibularhizoctonia sp. CBS 109695 | AcademH-1_FiCBS |
| -Agaricomycetes/Agaricales | Gymnopilus dilepis | AcademH-1_GyDi |
| Hebeloma cylindrosporum h7 | AcademH-1_HeCy | |
| Laccaria bicolor S238N-H82 | AcademH-1_LB, AcademH-2_LB | |
| Panaeolus cyanescens | AcademH-1_PaCy, AcademH-2_PaCy | |
| -Agaricomycetes/Amylocorticiales | Plicaturopsis crispa FD-325 SS-3 | AcademH-1_PlCr to 3_PlCr |
| -Agaricomycetes/Boletales | Serpula lacrymans | AcademH-1_SLL |
| -Agaricomycetes/Hymenochaetales | Schizopora paradoxa | AcademH-1_ScPa, AcademH-2_ScPa |
| -Agaricomycetes/Polyporales | Dichomitus squalens LYAD-421 SS1 | AcademH-1_DiSq |
| Trametes cinnabarina | AcademH-1_TrCi, AcademH-2_TrCi | |
| Trametes pubescens | AcademH-1_TrPu | |
| -Exobasidiomycetes/Tilletiales | Tilletia caries | AcademH-1_TiCa, AcademH-2_TiCa |
| Tilletia indica | AcademH-1_TiIn | |
| Fungi/Ascomycota | ||
| -Pezizomycetes/Pezizales | Ascobolus immersus RN42 | AcademH-1_AsIm, AcademH-2_AsIm |
| Fungi/Mucoromycota | ||
| -Mortierellomycetes/Mortierellales | Lobosporangium transversale | AcademH-1_LoTr |
| Mortierella verticillata NRRL 6337 | AcademH-1_MoVe, AcademH-2_MoVe | |
| Metazoa/Porifera | ||
| -Demospongiae/Haplosclerida | Amphimedon queenslandica | AcademH-1_AQ to 3_AQ, AcademH-N1_AQ to N2_AQ |
| Metazoa/Cnidaria | ||
| -Anthozoa/Scleractinia | Acropora digitifera | Academ-1_ADi to 3_ADi, AcademH-4_ADi to 7_ADi, AcademH-N1_ADi to N14_ADi |
| Orbicella faveolata | AcademH-N1_OrFa to N6_OrFa | |
| Stylophora pistillata | AcademH-1_StPi, AcademH-N1_StPi to N3_StPi | |
| -Anthozoa/Actiniaria | Exaiptasia pallida | AcademH-1_ExPa to 2_ExPa, AcademH-N1_ExPa to N3_ExPa |
| Nematostella vectensis | AcademH-N1_NV, N1A_NV, N2_NV, N2A_NV | |
| Metazoa/Priapulida | ||
| -Priapulimorpha/Priapulimorphida | Priapulus caudatus | AcademHP-1_PrCa |
| Metazoa/Mollusca | ||
| -Bivalva/Ostreoida | Crassostrea gigas | Academ-2_CGi, AcademH-1_CGi to 3_CGi, AcademH-2N1_CGi, AcademH-N1_CGi to N3_CGi, AcademH-2B_CGi |
| Crassostrea virginica | AcademH-1_CVi to 16_CVi, AcademH-2N1_CVi, AcademH-7N1_CVi, AcademH-N1_CVi to N2_CVi, AcademHP-1_CVi | |
| -Bivalva/Pectinoida | Mizuhopecten yessoensis | AcademH-N1_MiYe to N3_MiYe |
| Metazoa/Hemichordata | ||
| -Euteropneusta | Saccoglossus kowalevskii | AcademHP-1_SKow to 2_SKow, AcademH-N1_SKow, AcademH-N1B_SKow |
| Metazoa/Echinodermata | ||
| -Echinoidea/Echinoida | Strongylocentrotus purpuratus | AcademHP-1_SP |
| Metazoa/Chordata | ||
| -Leptocardii/Amphioxiformes | Branchiostoma floridae | AcademH-N1_BF |
| Branchiostoma belcheri | AcademH-1_BBe, AcademH-2_BBe | |
With two of these characterized AcademH protein sequences (AcademH-6_PGr and AcademH-1_CVi) as queries, BLASTP search against the non-redundant protein sequences (nr) at NCBI BLAST website hits many proteins from diverse fungi and animals (Supplementary Table S1). In fungi, proteins related to AcademH transposases were found from three subdivisions (Agaricomicotina, Pucciniomycotina, Ustilaginomycotina) within Basidiomycota, one subdivision (Pezisomycotina) within Ascomycota, and one subdivision (Mortierellomycotina) within Mucoromycota. Despite the report that Academ transposons are widely distributed in fungi [11], no other fungal group was revealed to contain AcademH transposons in this analysis. In animals, genomes from 9 phyla (Porifera, Cnidaria, Mollusca, Annelida, Brachiopoda, Priapulida, Chordata, Hemichordata, and Echinodermata) encode proteins related to AcademH transposases. Most of these protein sequences were encoded by single-copy, non-repetitive sequences. Basidiomycetes fungi with more than 5 protein hits and all other species were further analyzed. If their terminal inverted repeats (TIRs) longer than 10 bp and TSDs adjacent to TIRs could be detected in flanking 10,000-bp sequences, they were considered as full-length Academ transposons (Table 1 and Supplementary Dataset S2). Most of these single-copy Academ transposons encode a helicase protein. The sequence lengths, numbers of uninterrupted full-length copies, and the sequence identities to the consensus sequences are shown in Supplementary Table S2.
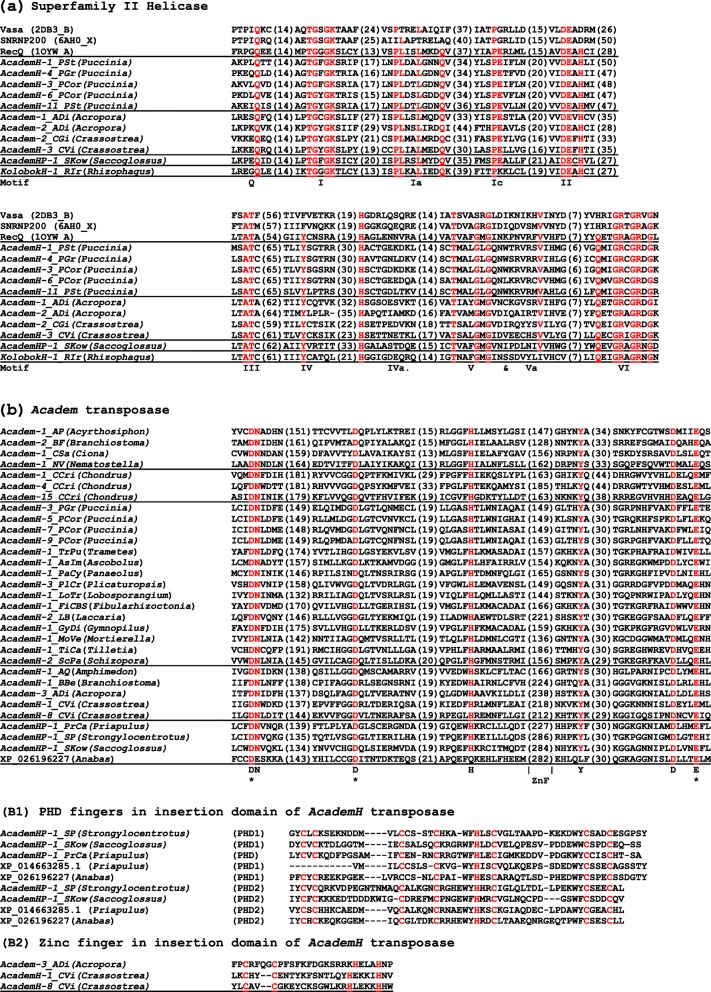
Secondary structure-based protein homology search HHpred was performed with helicase proteins encoded by AcademH DNA transposons. The top hit was RecQ DNA helicase from Escherichia coli, followed by U5 small nuclear ribonucleoprotein 200 and RNA helicase Vasa. The pairwise alignment generated by HHpred and multiple protein alignment generated by MAFFT were combined. It revealed that AcademH helicases conserve all motifs important for catalytic reactions, nucleic acid binding, and ATP binding (Fig. 2a). Censor search using helicase proteins encoded by AcademH against Repbase hit some families of KolobokH, a lineage of Kolobok DNA transposons encoding a helicase [14]. However, helicases encoded by AcademH and KolobokH are not so closely related to each other and are likely acquired independently in these two lineages of DNA transposons (data not shown). Helicases encoded by Helitron DNA transposons are Superfamily I helicases related to PIF1 helicase [15], and thus, there is little sequence similarity between helicases encoded by AcademH and Helitron.
Fig. 2.
Motifs of protein domains conserved in the Academ superfamily. Conserved residues are shown in red. The lengths between motifs are shown in parentheses. a Superfamily II helicase. The motif names are shown below alignment. bAcadem transposase. Conserved residues are shown below alignment. Asterisks indicate three catalytic residues. ZnF indicates the location where animal transposases encode PHD fingers or another zinc finger motif, which are shown in (B1) and (B2), respectively
Academ families without helicase often, but not always, contain 1 long open reading frame for a large protein containing three recognizable domains: a transposase, an XPG nuclease, and a putative Cys8 zinc finger (Figs. 1 and 2). Here, Academ families with XPG nuclease are designated as AcademX. In contrast, AcademH usually contain introns and encodes two proteins in opposite directions. These two proteins are encoded without overlapping. None of AcademH families encode an XPG nuclease or a Cys8 zinc finger.
Longer TSDs generated by AcademH than AcademX families
It is reported that AcademX DNA transposons generate 3-bp or 4-bp TSDs [8, 16]. In contrast, AcademH generates relatively long TSDs. Fungal AcademH families generate 9-bp TSDs with some exceptions (Fig. 3, and Supplementary Fig. S1). Animal AcademH families generate 9 or 10-bp TSDs (Fig. 4 and Supplementary Fig. S2). In the genome of coral A. digitifera, both lineages of Academ DNA transposons (AcademH and AcademX) are present. AcademH is usually inserted with 9-bp TSDs. AcademX generates 3-bp TSDs the same as previously reported AcademX DNA transposons from animals.
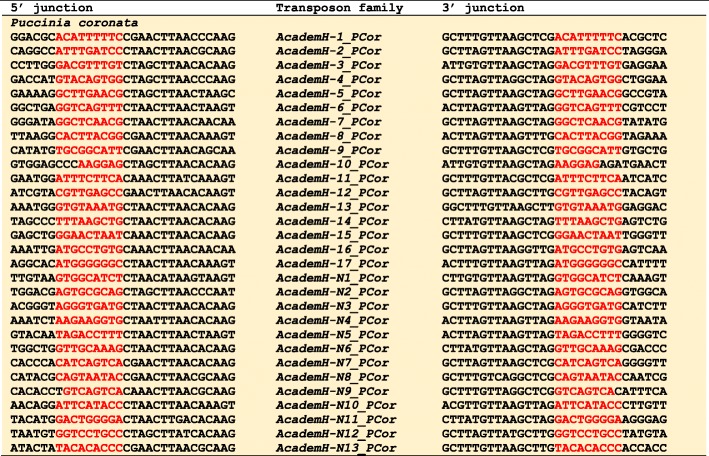
Fig. 3.
Termini and TSDs of Academ superfamily of DNA transposons from the fungus Puccinia coronata. Only one representative insertion is shown for each family. TSDs are colored in red. All termini and TSDs characterized in the study are shown in Supplementary Fig. S1
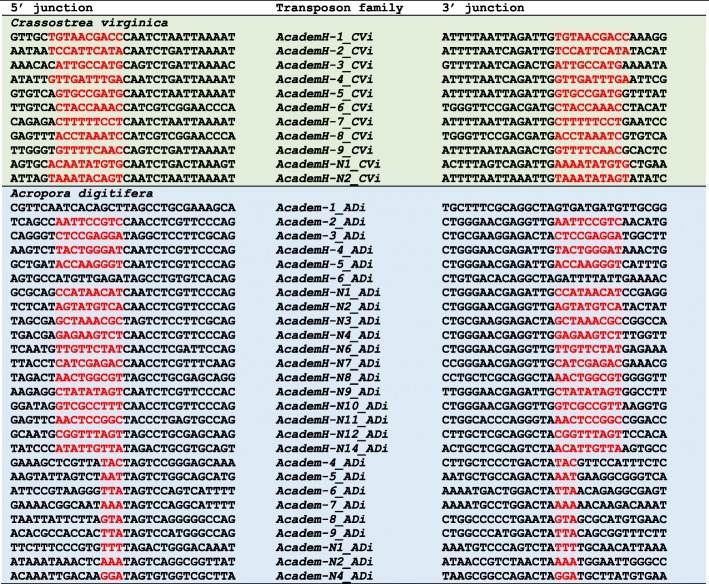
Fig. 4.
Termini and TSDs of Academ superfamily of DNA transposons from two animal species, Crassostrea virginica and Acropora digitifera. Only one representative insertion is shown for each family. TSDs are colored in red. No clear TSDs were observed for Academ-1_ADi and AcademH-6_ADi, while a short derivative of Academ-1_ADi is flanked by 9-bp TSDs (data not shown). All termini and TSDs characterized in the study are shown in Supplementary Fig. S2
Sequence comparison against reported non-autonomous TEs deposited in Repbase revealed that some of non-autonomous DNA transposons whose classification has not yet been determined are either AcademX or AcademH (Supplementary Table S3). DNA transposons with 8-bp or 9-bp TSDs show sequence similarity to AcademH termini while DNA transposons with 3-bp TSDs show sequence similarity to AcademX termini. One fungal species Melampsora larici-populina, closely related to Puccinia, and Nematostella vectensis, similarly to other cnidarian species, contain non-autonomous AcademH families (Table 1 and Supplementary Tables S2 and S3).
The presence of a pyrimidine (C or T) at the 5′ terminus and a purine (G or A) at the 3′ terminus is shared among almost all Academ families (Figs. 3 and 4). Some Academ families contain > 100-bp TIRs, represented by 526-bp TIRs of AcademH-1_LoTr and 575-bp TIRs of AcademH-16_CVi, while some have shorter than 10-bp TIRs; for example, AcademH-2_PSt and AcademH-N13_PHor have 8-bp TIRs.
AcademHP, a sublineage of AcademH with PHD zinc fingers
Although no proteins from vertebrates were hit in the first iteration of PSI-BLAST search with the transposase of AcademH-1_CVi or AcademH-6_PGr as a query, the protein sequences from the four teleost fishes were hit in the second iteration. They are from the climbing perch Anabas testudineus (XP_026195931, XP_026196227, XP_026196228, XP_026196229), the California yellowtail Seriola lalandi dorsalis (XP_023286175, XP_023286176), the spiny chromis damselfish Acanthochromis polyacanthus (XP_022063315, XP_022063316, XP_022063317, XP_022063318), and the rohu Labeo rohita (RXN19178, RXN19557). Besides these species, the genomes from a species of thornfishes Cottoperca gobio, the Siamese fighting fish Betta splendens, the bicolor damselfish Stegastes partitus, and the spotted seabass Lateolabrax maculatus contain related sequences (Supplementary Table S4). These proteins do not have all residues conserved among AcademH transposases (Fig. 2b, XP_026196227; and data not shown). Further investigation revealed that apparently intact AcademH transposons related to these proteins are present in the genomes of two deuterostomes: AcademHP-1_SP from the purple sea urchin Strongylocentrotus purpuratus and AcademHP-1_SKow from the acorn worm Saccoglossus kowalevskii (Fig. 2). These families encode 2 plant homeodomain (PHD) fingers between the second D and the last E catalytic residues (Figs. 1 and 2). One PHD finger contains 1 histidine residue sandwiched by 4 and 3 cysteine residues (Cys4-His-Cys3). PHD fingers share an ability to bind to tri-methylated lysines on histones [17], and thus, it is expected that the PHD fingers in the transposases of AcademHP families also bind to histones. Several copies of AcademHP families show 9-bp TSDs similarly to other AcademH families (Supplementary Fig. S3). One AcademHP sequence was also found as a single-copy sequence from the genome of Priapulus caudatus, although it encodes only one PHD finger (Fig. 2). Another protein encoded in the genome of P. caudatus (XP_014663285.1) contains 2 PHD fingers, although no TIRs flanked with recognizable TSDs were detected around the sequence encoding this protein. Thorough investigation revealed that other AcademH families from animals also contain a zinc finger motif between the second D and the last E catalytic residues, but they are CCHH-type (Fig. 2B2).
AcademH and AcademX, two distant linages inside of Academ superfamily
HHpred analysis with Academ transposases did not indicate any specific relationships with other transposases. The transposase domains of Academ are considered to belong to the DDE transposases, and thus to the RNaseH fold, based on Yuan and Wessler [12] which reported the conserved motifs and residues among Academ transposases. With more divergent transposases included in this analysis, fewer conserved residues are recognized (Fig. 2b). Only 7 residues, including the proposed DDE triad, are conserved among diverse Academ transposases. Compared with other DDE transposases, the first catalytic D and the second catalytic D are very distant (138–192 residues apart) in Academ transposases. The conserved G/A/E/QxxH motif following the second catalytic D residue might correspond to C/DxxH motif in MuDR, P, hAT, Kolobok and Dada, predicted to be located at the beginning of insertion domain [12].
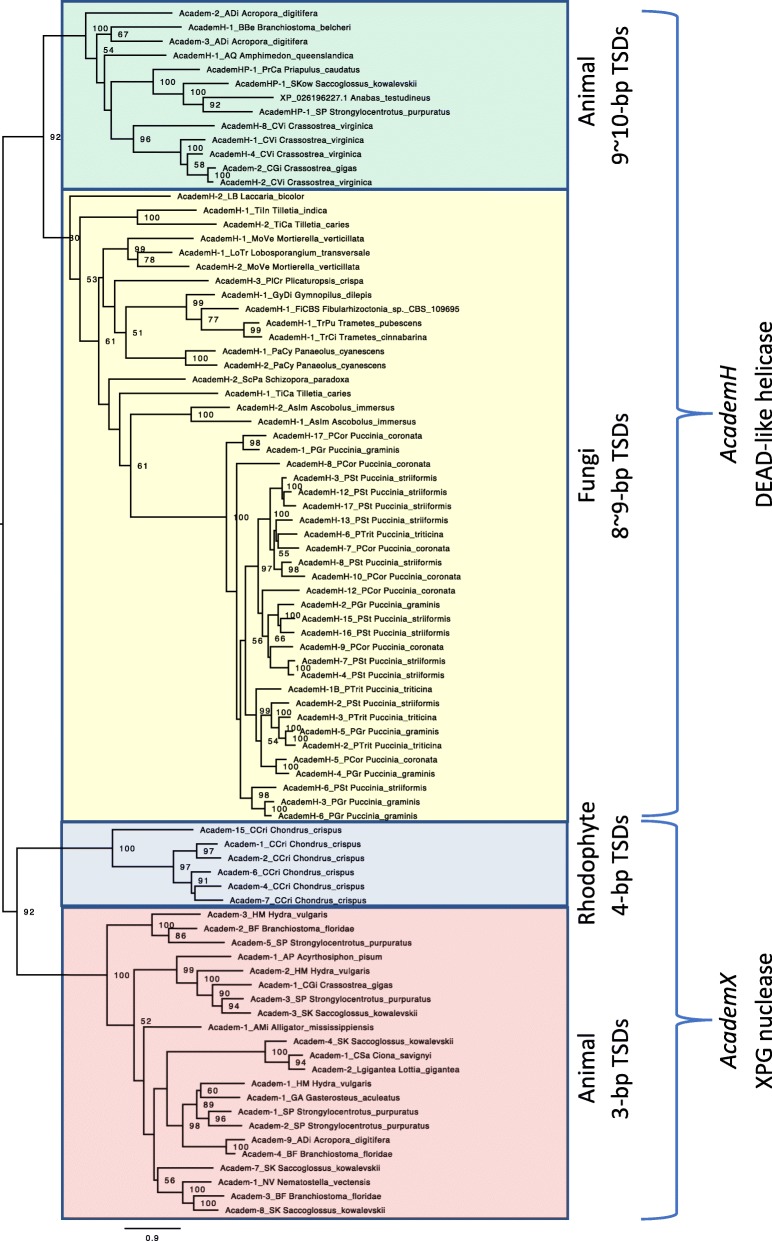
The phylogenetic analysis revealed that Academ superfamily can be classified into two large groups, AcademH and AcademX, corresponding to the protein coding ability (Fig. 5). AcademX can be further divided into two lineages, consistent with the difference in TSD length and distribution. AcademX with 3-bp TSDs are distributed among animals. AcademX with 4-bp TSDs has been found only from the red alga C. crispus. Two clusters for AcademH correspond to the AcademH from fungi and animals. The three AcademHP families with the AcademHP transposase-like protein encoded on the genome of A. testudineus (XP_026196227.1) clustered together inside of animal AcademH. AcademH transposons from closely related organisms are often clustered together, for examples, three families from Mucoromycote fungi (AcademH-1_LoTr, AcademH-1_MoVe and AcademH-2_MoVe) or five families from the oysters in the genus Crassostrea (AcademH-8_CVi, AcademH-1_CVi, AcademH-4_CVi, Academ-2_CGi, and AcdemH-2_CVi). All AcademH families from the genus Puccinia are very closely related. However, deeper phylogeny of AcademH transposases is not consistent with their host phylogeny. Considering the small number of genomes from which AcademH families were characterized, and low bootstrap supports for deeper nodes, the contribution of horizontal transfer to the AcademH evolution remains to be investigated.
Fig. 5.
Phylogeny of Academ transposases. Four lineages recognized are highlighted in different colors. Bootstrap values of 100 replicates are shown at nodes when they are > 50. Host organisms of TEs are shown after TE names
Discussion
The diversity and distribution of Academ
The Academ superfamily of DNA transposons has been found from three different groups of eukaryotes: animals, fungi and red algae. With a relatively small number of sequences, the phylogeny and structural characteristics of Academ are straightforward. The AcademX lineage encodes one large protein containing a transposase, an XPG nuclease and a putative zinc finger. It is distributed in animals and red algae. AcademX generates relatively short (3 or 4-bp) TSDs upon integration. The AcademH lineage encodes two proteins, one of which is a transposase and the other of which is a superfamily II helicase. AcademH generates relatively long (9 or 10-bp) TSDs upon integration. AcademH is distributed in animals and fungi. AcademHP is a sublineage inside of AcademH and this lineage encodes one or two PHD fingers between the second D and the last E catalytic residues. In vertebrates, the genomes of some teleost fishes keep remnants of AcademHP copies.
Functional implications for helicase in the life cycle of AcademH
The length of TSDs is one of the hallmarks of superfamilies of DNA transposons. In general, inside of the superfamily of DNA transposons, the lengths of TSDs are not so divergent [4, 18]. Almost all of superfamilies show strict restriction of TSD lengths, which allows only 1-bp difference. As rare exceptions, the hAT superfamily shows TSDs of 5, 6 or 8-bp, and the EnSpm superfamily shows TSDs of 2-bp, 3-bp, or 4-bp. In contrast, inside of the Academ superfamily, AcademX generates 3-bp or 4-bp TSDs, while AcademH generates TSDs of 9 or 10 bps in length.
AcademH families encode a superfamily II helicase related to RecQ, while AcademX families encode an XPG nuclease. Mutually exclusive presence of helicase or nuclease in Academ transposons implies the functional similarity of these two enzymes in the life cycle of Academ transposons. RecQ helicase family works for various DNA repair pathways including homologous recombination and non-homologous end joining [19]. XPG nuclease families are needed to repair DNA damages by a process called nucleotide excision repair [20]. It can be speculated that helicase and nuclease encoded by Academ transposons are coupled with cellular proteins in DNA repair pathway during the transposition of Academ transposons. DDE transposases cleave DNA at both termini of DNA transposons [21]. The difference in how DNA repair pathway is recruited to resolve the transposition intermediate might dictate the junction structures different between AcademX and AcademH.
Conclusions
The Academ superfamily of DNA transposons has 2 deep-branching lineages: AcademX and AcademH. Besides its transposase, AcademH encodes a superfamily II helicase, which may contribute to the generation of long TSDs.
Methods
Characterization of Academ DNA transposons
All genome sequences used in this study were downloaded from either of three websites: NCBI Assembly database (https://www.ncbi.nlm.nih.gov/assembly), UCSC Genome Browser (https://genome.ucsc.edu/), and OIST Marine Genomics Unit (http://marinegenomics.oist.jp/lingula/viewer/download?project_id=47), and listed in Supplementary Table S5.
Censor searches [13] using reported Academ sequences as queries against genomes were performed. Sequences showing similarity to Academ were clustered by BLASTCLUST in the NCBI Blast package. Censor searches were done with consensus sequence of each cluster and the hits with flanking sequences were extracted to characterize the complete repeat unit until TSDs were detected.
In parallel, RepeatModeler (http://www.repeatmasker.org/RepeatModeler/) and Repbase [1] were used for the initial screening of repetitive families with default parameters for all animal genomes used here except for C. gigas, C. virginica, M. yessoensis, C. teleta, P. caudatus, and B. floridae. Consensus sequences generated by RepeatModeler output with the annotation as Academ were chosen to reconstruct the second consensus sequences using the top 10 hits with the 1000-bp flanking sequences at both sides in the Censor search.
Single-copy sequences similar to AcademH families were annotated as AcademH transposons if > 10-bp TIRs and adjacent > 8-bp TSDs were detected within their 10,000-bp flanking sequences.
The consensus or single-copy representative sequences for all TE families reported here have been submitted to Repbase [1], and are also available in Supplementary Datasets S1 and S2.
Protein structure and phylogenetic analyses
Protein coding regions were predicted from consensus sequences and representative single-copy sequences with Softberry FGENESH (http://www.softberry.com/berry.phtml?topic=fgenesh&group=programs&subgroup=gfind) [22], followed by manual curation with reference to predicted mRNA sequences available at NCBI website. NCBI CD-Search (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) [23] was done to detect protein domains. HHpred (https://toolkit.tuebingen.mpg.de/tools/hhpred) [24] was used to find similar structures of respective proteins.
Multiple sequence alignment was done with MAFFT with linsi option [25]. Academ transposase domains were extracted following the definition in [12, 16]. Protein sequences with truncation or internal deletion inside of transposase domain were excluded from the analysis. The final dataset used for the phylogenetic analysis contains 86 sequences which are 319 to 541 residues in length (Supplementary Dataset S3). Maximum likelihood trees with bootstrap values of 100 replicates were constructed using PhyML [26] with the amino acid substitution model LG + G + I + F, which was chosen based on the best Akaike Information Criterion score. The phylogenetic trees were drawn with the aid of FigTree 1.3.1 (http://tree.bio.ed.ac.uk/software/figtree/).
Supplementary information
Additional file 1: Figure S1. Termini and TSDs of newly characterized families of Academ from the fungus Puccinia coronata. Figure S2. Termini and TSDs of newly characterized families of Academ from two animal species, Crassostrea virginica and Acropora digitifera. Figure S3. Termini and TSDs of AcademHP families from animals. Table S3. Non-autonomous DNA transposons newly classified as Academ. Table S4.AcademHP remnants found in teleost.
Additional file 2 : Table S1. Protein sequences showing similarity to AcademH transposases. Table S2. Characteristics of AcademH families. Table S5. Genome assembly sequences used in this study.
Additional file 3 : Data S1. Consensus sequences of multicopy Academ transposons characterized in this study. Data S2. Representative sequences of single-copy Academ transposons characterized in this study. Data S3. Protein multiple alignment of Academ transposase domains used for the phylogenetic analysis.
Acknowledgements
Not applicable.
Abbreviations
- TSD
Target site duplication
- TIR
Terminal inverted repeat
- PHD
Plant homeodomain
- TE
Transposable element
- LTR
Long terminal repeat
- RAG1
Recombination activating gene 1
Author’s contributions
KKK performed experiments and analysis, and wrote the manuscript. The author read and approved the final manuscript.
Funding
The author received no specific funding for this work.
Availability of data and materials
All data generated or analyzed in this study are included in this published article and its supplementary information files. Consensus and single-copy representative sequences of TEs are also submitted to Repbase (http://www.girinst.org/repbase/).
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The author declare that he has no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1186/s13100-020-00211-1.
References
- 1.Bao W, Kojima KK, Kohany O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob DNA. 2015;6:11. doi: 10.1186/s13100-015-0041-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Siguier P, Gourbeyre E, Varani A, Ton-Hoang B, Chandler M. Everyman’s Guide to Bacterial Insertion Sequences. Microbiol Spectr. 2015;3(2):MDNA3–0030-2014. doi: 10.1128/microbiolspec.MDNA3-0030-2014. [DOI] [PubMed] [Google Scholar]
- 3.Finnegan DJ. Eukaryotic transposable elements and genome evolution. Trends Genet. 1989;5(4):103–107. doi: 10.1016/0168-9525(89)90039-5. [DOI] [PubMed] [Google Scholar]
- 4.Kojima KK. Structural and sequence diversity of eukaryotic transposable elements. Genes Genet Syst. 2019;94:233–52. [DOI] [PubMed]
- 5.Arkhipova IR. Using bioinformatic and phylogenetic approaches to classify transposable elements and understand their complex evolutionary histories. Mob DNA. 2017;8:19. doi: 10.1186/s13100-017-0103-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hickman AB, Chandler M, Dyda F. Integrating prokaryotes and eukaryotes: DNA transposases in light of structure. Crit Rev Biochem Mol Biol. 2010;45(1):50–69. doi: 10.3109/10409230903505596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kapitonov VV, Jurka J. RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLoS Biol. 2005;3(6):e181. doi: 10.1371/journal.pbio.0030181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kapitonov VV, Jurka J. Academ - a novel superfamily of eukaryotic DNA transposons. Repbase Rep. 2010;10(4):643–647. [Google Scholar]
- 9.Bao W, Jurka J. DNA transposons from the red seaweed. Repbase Rep. 2013;13(10):2271–2285. [Google Scholar]
- 10.Kojima KK, Jurka J. DNA transposons from the Puccinia graminis genome. Repbase Rep. 2015;15(8):2495–2508. [Google Scholar]
- 11.Muszewska A, Steczkiewicz K, Stepniewska-Dziubinska M, Ginalski K. Cut-and-paste transposons in Fungi with diverse lifestyles. Genome Biol Evol. 2017;9(12):3463–3477. doi: 10.1093/gbe/evx261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yuan YW, Wessler SR. The catalytic domain of all eukaryotic cut-and-paste transposase superfamilies. Proc Natl Acad Sci U S A. 2011;108(19):7884–7889. doi: 10.1073/pnas.1104208108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kohany O, Gentles AJ, Hankus L, Jurka J. Annotation, submission and screening of repetitive elements in Repbase: RepbaseSubmitter and censor. BMC Bioinformatics. 2006;7:474. doi: 10.1186/1471-2105-7-474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kapitonov VV, Jurka J. Kolobok transposons in the Glomeromycota fungus. Repbase Rep. 2014;14(7):1925–1929. [Google Scholar]
- 15.Kapitonov VV, Jurka J. Rolling-circle transposons in eukaryotes. Proc Natl Acad Sci U S A. 2001;98(15):8714–8719. doi: 10.1073/pnas.151269298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang HH, Shen YH, Xiong XM, Han MJ, Qi DW, Zhang XG. Evidence for horizontal transfer of a recently active Academ transposon. Insect Mol Biol. 2016;25(3):338–346. doi: 10.1111/imb.12225. [DOI] [PubMed] [Google Scholar]
- 17.Musselman CA, Kutateladze TG. Handpicking epigenetic marks with PHD fingers. Nucleic Acids Res. 2011;39(21):9061–9071. doi: 10.1093/nar/gkr613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kapitonov VV, Jurka J. A universal classification of eukaryotic transposable elements implemented in Repbase. Nat Rev Genet. 2008;9(5):411–412. doi: 10.1038/nrg2165-c1. [DOI] [PubMed] [Google Scholar]
- 19.Croteau DL, Popuri V, Opresko PL, Bohr VA. Human RecQ helicases in DNA repair, recombination, and replication. Annu Rev Biochem. 2014;83:519–552. doi: 10.1146/annurev-biochem-060713-035428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Scharer OD. XPG: its products and biological roles. Adv Exp Med Biol. 2008;637:83–92. doi: 10.1007/978-0-387-09599-8_9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Curcio MJ, Derbyshire KM. The outs and ins of transposition: from mu to kangaroo. Nat Rev Mol Cell Biol. 2003;4(11):865–877. doi: 10.1038/nrm1241. [DOI] [PubMed] [Google Scholar]
- 22.Solovyev V, Kosarev P, Seledsov I, Vorobyev D. Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol. 2006;7(Suppl 1):S10 1–S10 2. doi: 10.1186/gb-2006-7-s1-s10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, et al. CDD: NCBI's conserved domain database. Nucleic Acids Res. 2015;43(Database issue):D222–D226. doi: 10.1093/nar/gku1221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zimmermann L, Stephens A, Nam SZ, Rau D, Kubler J, Lozajic M, et al. A completely Reimplemented MPI bioinformatics toolkit with a new HHpred server at its Core. J Mol Biol. 2018;430(15):2237–2243. doi: 10.1016/j.jmb.2017.12.007. [DOI] [PubMed] [Google Scholar]
- 25.Katoh K, Kuma K, Toh H, Miyata T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 2005;33(2):511–518. doi: 10.1093/nar/gki198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59(3):307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Figure S1. Termini and TSDs of newly characterized families of Academ from the fungus Puccinia coronata. Figure S2. Termini and TSDs of newly characterized families of Academ from two animal species, Crassostrea virginica and Acropora digitifera. Figure S3. Termini and TSDs of AcademHP families from animals. Table S3. Non-autonomous DNA transposons newly classified as Academ. Table S4.AcademHP remnants found in teleost.
Additional file 2 : Table S1. Protein sequences showing similarity to AcademH transposases. Table S2. Characteristics of AcademH families. Table S5. Genome assembly sequences used in this study.
Additional file 3 : Data S1. Consensus sequences of multicopy Academ transposons characterized in this study. Data S2. Representative sequences of single-copy Academ transposons characterized in this study. Data S3. Protein multiple alignment of Academ transposase domains used for the phylogenetic analysis.
Data Availability Statement
All data generated or analyzed in this study are included in this published article and its supplementary information files. Consensus and single-copy representative sequences of TEs are also submitted to Repbase (http://www.girinst.org/repbase/).