Abstract
The coronavirus disease 2019 (COVID-19) pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has affected millions of people worldwide, igniting an unprecedented effort from the scientific community to understand the biological underpinning of COVID19 pathophysiology. In this Review, we summarize the current state of knowledge of innate and adaptive immune responses elicited by SARS-CoV-2 infection and the immunological pathways that likely contribute to disease severity and death. We also discuss the rationale and clinical outcome of current therapeutic strategies as well as prospective clinical trials to prevent or treat SARS-CoV-2 infection.
The coronavirus disease 2019 (COVID-19) pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has affected millions of people worldwide, igniting an unprecedented effort from the scientific community to understand the biological underpinning of COVID19 pathophysiology. In this Review, we summarize the current state of knowledge of innate and adaptive immune responses elicited by SARS-CoV-2 infection and the immunological pathways that likely contribute to disease severity and death. We also discuss the rationale and clinical outcome of current therapeutic strategies as well as prospective clinical trials to prevent or treat SARS-CoV-2 infection.
Main Text
Introduction
The recent emergence and rapid global spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the resulting coronavirus disease 2019 (COVID-19) poses an unprecedented health crisis that was declared a pandemic by the World Health Organization (WHO) on March 11, 2020. The origin of SARS-CoV-2 was traced to the city of Wuhan in the province of Hubei, China, where a cluster of viral pneumonia cases was first detected, many in connection with the Huanan Seafood Wholesale Market. China reported this outbreak to the WHO on December 31, 2019 and soon after identified the causative pathogen as a betacoronavirus with high sequence homology to bat coronaviruses (CoVs) using angiotensin-converting enzyme 2 (ACE2) receptor as the dominant mechanism of cell entry (Lu et al., 2020a, Wan et al., 2020b). Following a likely zoonotic spillover, human-to-human transmission events were confirmed with clinical presentations ranging from no symptoms to mild fever, cough, and dyspnea to cytokine storm, respiratory failure, and death. SARS-CoV-2 is also closely related to SARS (retrospectively named SARS-CoV-1) and Middle Eastern respiratory syndrome (MERS) CoVs, causing zoonotic epidemic and local outbreaks in 2003 and 2012, respectively (de Wit et al., 2016). While SARS-CoV-2 is not as lethal as SARS-CoV-1 or MERS-CoV (Fauci et al., 2020), the considerable spread of the current pandemic has brought tremendous pressure and disastrous consequences for public health and medical systems worldwide.
The scientific response to the crisis has been extraordinary, with a plethora of COVID-19 studies posted in preprint servers in an attempt to rapidly unravel the pathogenesis of COVID-19 and potential therapeutic strategies. In response, trainees and faculty members of the Precision Immunology Institute at the Icahn School of Medicine at Mount Sinai (PrIISM) have initiated an institutional effort to critically review the preprint literature (Vabret et al., 2020), together with peer-reviewed articles published in traditional journals, and summarize the current state of science on the fast-evolving field of COVID-19 immunology. We thematically focus on the innate and adaptive immune responses to SARS-CoV-2 and related CoVs, clinical studies and prognostic laboratory correlates, current therapeutic strategies, prospective clinical trials, and vaccine approaches.
Innate Immune Sensing of SARS-CoV-2
Innate immune sensing serves as the first line of antiviral defense and is essential for immunity to viruses. To date, our understanding of the specific innate immune response to SARS-CoV-2 is extremely limited. However, the virus-host interactions involving SARS-CoV-2 are likely to recapitulate many of those involving other CoVs, given the shared sequence homology among CoVs and the conserved mechanisms of innate immune signaling. In the case of RNA viruses such as SARS-CoV-2, these pathways are initiated through the engagement of pattern-recognition receptors (PRRs) by viral single-stranded RNA (ssRNA) and double-stranded RNA (dsRNA) via cytosolic RIG-I like receptors (RLRs) and extracellular and endosomal Toll-like receptors (TLRs). Upon PRR activation, downstream signaling cascades trigger the secretion of cytokines. Among these, type I/III interferons (IFNs) are considered the most important for antiviral defense, but other cytokines, such as proinflammatory tumor necrosis factor alpha (TNF-α), and interleukin-1 (IL-1), IL-6, and IL-18 are also released. Together, they induce antiviral programs in target cells and potentiate the adaptive immune response. If present early and properly localized, IFN-I can effectively limit CoV infection (Channappanavar et al., 2016, Channappanavar et al., 2019). Early evidence demonstrated that SARS-CoV-2 is sensitive to IFN-I/III pretreatment in vitro, perhaps to a greater degree than SARS-CoV-1 (Blanco-Melo et al., 2020, Lokugamage et al., 2020, Mantlo et al., 2020, Stanifer et al., 2020). However, the specific IFN-stimulated genes (ISGs) that mediate these protective effects are still being elucidated. Lymphocyte antigen 6 complex locus E (LY6E) has been shown to interfere with SARS-CoV-2 spike (S) protein-mediated membrane fusion (Pfaender et al., 2020, Zhao et al., 2020c). Likely, the IFN-induced transmembrane family (IFITM) proteins inhibit SARS-CoV-2 entry, as demonstrated for SARS-CoV-1 (Huang et al., 2011b), although their action in promoting infection has also been described for other CoVs (Zhao et al., 2014, Zhao et al., 2018).
Evasion of Innate Sensing by Coronaviruses
As these cytokines represent a major barrier to viral infection, CoVs have evolved several mechanisms to inhibit IFN-I induction and signaling. Numerous studies have demonstrated that SARS-CoV-1 suppresses IFN release in vitro and in vivo (Cameron et al., 2012, Minakshi et al., 2009, Siu et al., 2009, Wathelet et al., 2007). SARS-CoV-2 likely achieves a similar effect, as suggested by the lack of robust type I/III IFN signatures from infected cell lines, primary bronchial cells, and a ferret model (Blanco-Melo et al., 2020). In fact, patients with severe COVID-19 demonstrate remarkably impaired IFN-I signatures as compared to mild or moderate cases (Hadjadj et al., 2020). As is often the case, there are multiple mechanisms of evasion for CoVs, with viral factors antagonizing each step of the pathway from PRR sensing and cytokine secretion to IFN signal transduction (Figure 1 ).
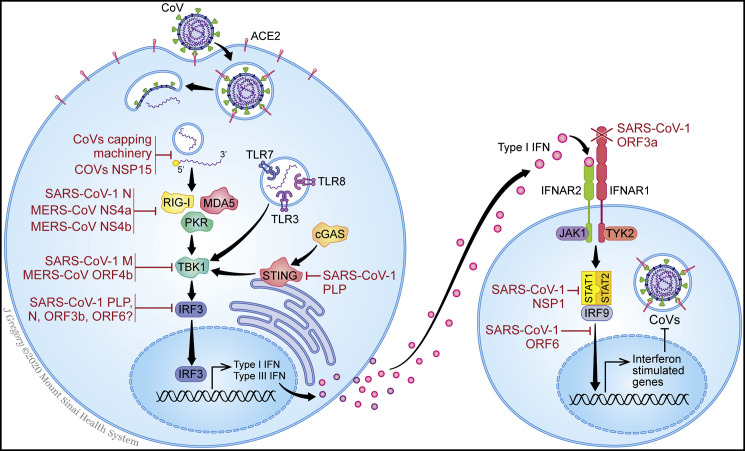
Figure 1.
Mechanisms of Host Innate Immune Response and Coronaviruses Antagonism
Overview of innate immune sensing (left) and interferon signaling (right), annotated with the known mechanisms by which SARS-CoV-1 and MERS-CoV antagonize the pathways (red).
CoV-mediated antagonism of innate immunity begins with evasion of PRR sensing. ssRNA viruses, like CoVs, form dsRNA intermediates during their replication, which can be detected by TLR3 in the endosome and RIG-I, MDA5, and PKR in the cytosol. ssRNA may also be detected by TLR7 or TLR8 and potentially RIG-I and PKR. CoVs are known to avoid PRR activation by either avoiding recognition altogether or antagonizing PRR action (Bouvet et al., 2010, Chen et al., 2009, Deng et al., 2017, Hackbart et al., 2020, Ivanov et al., 2004, Knoops et al., 2008). To evade PRRs, dsRNA is first shielded by membrane-bound compartments that form during viral replication of SARS-CoV-1 (Knoops et al., 2008). In addition, viral RNA is guanosine-capped and methylated at the 5′ end by CoVs non-structural proteins (NSPs) 10, 13, 14, and 16 (Bouvet et al., 2010, Chen et al., 2009, Ivanov et al., 2004), thereby resembling host mRNA to promote translation, prevent degradation, and evade RLR sensing. Finally, CoVs also encode an endoribonuclease, NSP15, that cleaves 5′ polyuridines formed during viral replication, which would otherwise be detected by MDA5 (Deng et al., 2017, Hackbart et al., 2020). CoVs have evolved additional strategies to impede activation of PRRs. SARS-CoV-1 N-protein prevents TRIM25 activation of RIG-I (Hu et al., 2017). Likewise, MERS-CoV NS4a, which itself binds dsRNA, impedes PKR activation (Comar et al., 2019, Rabouw et al., 2016) and inhibits PACT, an activator of RLRs (Niemeyer et al., 2013, Siu et al., 2014). Additionally, MERS-CoV NS4b antagonizes RNaseL, another activator of RLRs (Thornbrough et al., 2016). The role of other PRRs remains unclear. For example, SARS-CoV-1 papain-like protease (PLP) antagonizes STING, suggesting that self-DNA may also represent an important trigger (Sun et al., 2012). The extent to which SARS-CoV-2 homologs overlap in these functions is currently unknown.
Following activation, RLR and TLRs induce signaling cascades, leading to the phosphorylation of transcription factors, such as NF-kB and the interferon-regulatory factor family (IRF), ultimately leading to transcription of IFN and proinflammatory cytokines. Although no experimental studies have delineated the precise functions of SARS-CoV-2 proteins, proteomic studies have demonstrated interactions between viral proteins and PRR signaling cascades. SARS-CoV-2 ORF9b indirectly interacts with the signaling adaptor MAVS via its association with Tom70 (Gordon et al., 2020), consistent with prior reports that SARS-CoV-1 ORF9b suppresses MAVS signaling (Shi et al., 2014). Furthermore, SARS-CoV-2 NSP13 interacts with signaling intermediate TBK1, and NSP15 is associated with RNF41, an activator of TBK1 and IRF3 (Gordon et al., 2020). Similarly, SARS-CoV-1 M protein is known to inhibit the TBK1 signaling complex (Siu et al., 2009), as does MERS-CoV ORF4b (Yang et al., 2015). Other proteins, including SARS-CoV-1 PLP, N, ORF3b, and ORF6, block IRF3 phosphorylation and nuclear translocation (Devaraj et al., 2007, Kopecky-Bromberg et al., 2007). NF-kB is also inhibited by CoV proteins. These include SARS-CoV-1 PLP (Frieman et al., 2009) and MERS-CoV ORF4b and ORF5 (Canton et al., 2018, Menachery et al., 2017). Finally, SARS-CoV-1 NSP1 (Huang et al., 2011a, Kamitani et al., 2009) and MERS-CoV NSP1 (Lokugamage et al., 2015) initiate general inhibition of host transcription and translation, thus limiting antiviral defenses nonspecifically.
To prevent signaling downstream of IFN release, CoV proteins inhibit several steps of the signal transduction pathway that bridge the receptor subunits (IFNAR1 and IFNAR2) to the STAT proteins that activate transcription. For SARS-CoV-1, these mechanisms include IFNAR1 degradation by ORF3a (Minakshi et al., 2009), decreased STAT1 phosphorylation by NSP1 (Wathelet et al., 2007), and antagonism of STAT1 nuclear translocation by ORF6 (Frieman et al., 2007, Kopecky-Bromberg et al., 2007). However, SARS-CoV-2 ORF6 shares only 69% sequence homology with SARS-CoV-1, suggesting this function may not be conserved. In support of this notion, SARS-CoV-2 infection fails to limit STAT1 phosphorylation, unlike in SARS-CoV-1 infection (Lokugamage et al., 2020).
Imbalance between Antiviral and Proinflammatory Responses
Taken together, the multiplicity of strategies developed by pathogenic CoVs to escape immune sensing, particularly the IFN-I pathway, suggests a critical role played by the dysregulation of IFN-I response in COVID-19 pathogenicity. Concordantly, animal models of SARS-CoV-1 and MERS-CoV infection indicate that failure to elicit an early IFN-I response correlates with the severity of disease (Channappanavar et al., 2016). Perhaps more importantly, these models demonstrate that timing is key, as IFN is protective early in disease but later becomes pathologic (Channappanavar et al., 2016, Channappanavar et al., 2019). Perhaps interferon-induced upregulation of ACE2 in airway epithelia may contribute to this effect (Ziegler et al., 2020). Furthermore, while pathogenic CoVs block IFN signaling, they may actively promote other inflammatory pathways contributing to pathology. For instance, SARS-CoV-1 ORF3a, ORF8b, and E proteins enhance inflammasome activation (Chen et al., 2019, Nieto-Torres et al., 2015, Shi et al., 2019, Siu et al., 2019), leading to secretion of IL-1β and IL-18, which are likely to contribute to pathological inflammation. Similarly, SARS-CoV-2 NSP9 and NSP10 might induce IL-6 and IL-8 production, potentially by inhibition of NKRF, an endogenous NF-kB repressor (Li et al., 2020a). Collectively, these proinflammatory processes likely contribute to the “cytokine storm” observed in COVID-19 patients and substantiate a role for targeted immunosuppressive treatment regimens. Moving forward, a clear understanding of the delicate balance between antiviral and inflammatory innate immune programs will be essential to developing effective biomarkers and therapeutics for COVID-19.
Myeloid Cells
Mucosal immune responses to infectious agents are orchestrated and regulated by myeloid cells with specialized functions, which include conventional dendritic cells (cDCs), monocyte-derived DCs (moDCs), plasmacytoid DCs (pDCs), and macrophages (Guilliams et al., 2013). A growing body of evidence points to dysregulated myeloid responses that potentially drive the COVID-19 hallmark syndromes, such as acute respiratory distress syndrome (ARDS), cytokine release syndrome (CRS) and lymphopenia (Mehta et al., 2020).
Myeloid Characterization in COVID-19
Flow cytometric analyses of peripheral blood mononuclear cells (PBMCs) from symptomatic COVID-19 patients have shown a significant influx of granulocyte-macrophage colony-stimulating factor (GM-CSF)-producing, activated CD4+ T cells and CD14+HLA-DRlo inflammatory monocytes (IMs) (Giamarellos-Bourboulis et al., 2020, Zhang et al., 2020c, Zhou et al., 2020b). This matches single-cell transcriptomic (scRNA-seq) data demonstrating CD14+IL-1β+ monocytic expansion (Guo et al., 2020, Wen et al., 2020), interferon-mitogen-activated protein kinase (MAPK)-driven adaptive immune responses (Huang et al., 2020c), and IL-1β-associated inflammasome signatures (Ong et al., 2020) in peripheral blood of COVID-19 patients, although systemic levels of IL-1β detected are conspicuously low (Del Valle et al., 2020). Importantly, these immune signatures track with progression of clinical disease. scRNA-seq studies performed on pulmonary tissues of patients with severe COVID-19 disease have revealed an expansion of IMs and Ficolin-1+ monocyte-derived macrophages at the expense of tissue-resident reparative alveolar macrophages (AMs) (Liao et al., 2020). The aforementioned study also observed signatures of IFN signaling and monocyte recruitment that likely contribute to the rapid decline in alveolar patency and promote ARDS. Although most of the clinical focus has been on pulmonary damage and mononuclear phagocyte (MNP) dysfunction therein, it is increasingly clear that COVID-19 likely presents systemic challenges in other organ sites, such as the ileum and kidneys. Understanding the role of non-pulmonary myeloid cells in tissue-specific pathology associated with COVID-19 will be important.
Prior Knowledge from SARS-CoV-1, MERS-CoV, and Murine Coronaviruses
While data on COVID-19 patients continues to rapidly emerge, studies of myeloid cell dysfunction in SARS-CoV-1 and MERS-CoV can provide an important roadmap to understanding COVID-19 pathogenesis (Figure 2 ). SARS-CoV-1 infection in mouse models results in an aberrant AM phenotype that limits DC trafficking and T cell activation (Zhao et al., 2009). Additionally, YM1+ FIZZ1+ alternative macrophages can increase airway hypersensitivity, thus exacerbating SARS-associated fibrosis (Page et al., 2012). Further, as described above, murine SARS-CoV-1 studies have demonstrated that delayed IFN-I signaling and inflammatory monocytes-macrophages promote lung cytokine and chemokine levels, vascular leakage, and impaired antigen-specific T cell responses, culminating in lethal disease (Channappanavar et al., 2016). The role played by prominent IFN-producing pDCs in SARS-CoV-2 control or pathogenesis warrants investigation, as they have been shown to be critical in murine CoV (MHV) control (Cervantes-Barragan et al., 2007). Longitudinal studies in SARS-CoV-2 models are awaited, but initial phenotypic studies in humanized hACE2 mice have shown the characteristic alveolar interstitial pneumonia, with infiltration of lymphocytes and monocytes and accumulation of macrophages in the alveolar lumen (Bao et al., 2020a), which recapitulates patient findings (Xu et al., 2020c). Lastly, non-human primate (NHP) studies and patient data on SARS-CoV-1 have also shown that virus spike-specific immunoglobulin G (IgG) responses can exacerbate acute lung injury due to repolarization of alveolar macrophages into proinflammatory phenotypes and enhanced recruitment of inflammatory monocyte via CCL2 and IL-8 (Clay et al., 2012, Liu et al., 2019). However, the extent to which the antibody response contributes to disease pathophysiology remains to be confirmed.
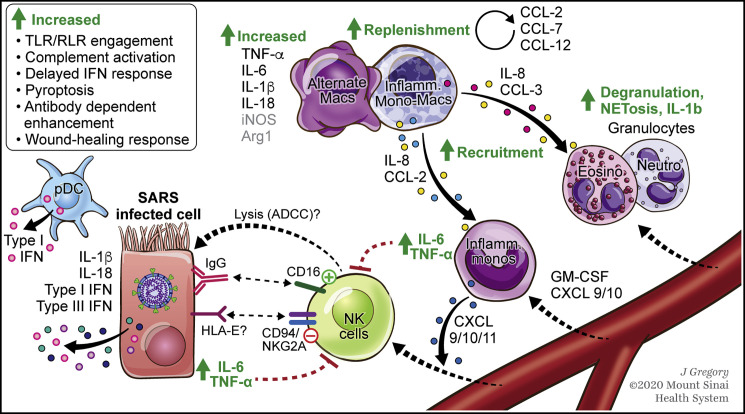
Figure 2.
SARS-CoV-2 Infection Results in Myeloid Cell Activation and Changes NK Cell Function
Based on data from preliminary COVID-19 studies and earlier studies in related coronaviruses.
IL-6, IL-1β, and IFN-I/III from infected pulmonary epithelia can induce inflammatory programs in resident (alternate) macrophages while recruiting inflammatory monocytes, as well as granulocytes and lymphocytes from circulation. Sustained IL-6 and TNF-ɑ by incoming monocytes can drive several hyperinflammation cascades. Inflammatory monocyte-derived macrophages can amplify dysfunctional responses in various ways (listed in top-left corner). The systemic CRS- and sHLH-like inflammatory response can induce neutrophilic NETosis and microthrombosis, aggravating COVID-19 severity. Other myeloid cells, such as pDCs, are purported to have an IFN-dependent role in viral control. Monocyte-derived CXCL9/10/11 might recruit NK cells from blood. Preliminary data suggest that the antiviral function of these NK cells might be regulated through crosstalk with SARS-infected cells and inflammatory monocytes.
Dashed lines indicate pathways to be confirmed. Arg1, arginase 1; iNOS, inducible-nitric oxide synthase; Inflamm., inflammatory; Mono., monocytes; Macs, macrophages; Eosino, eosinophils; Neutro, neutrophils; NETosis, neutrophil extracellular trap cell death; SHLH, secondary hemophagocytic lymphohistiocytosis.
Myeloid Cells Contribution to Pathogenic Inflammation
The initial mode of viral pathogen-associated signal (PAMP) recognition by innate cells has a major impact on downstream myeloid signaling and cytokine secretion (de Marcken et al., 2019). While macrophages are somewhat susceptible to MERS-CoV and SARS-CoV-1 infection (Perlman and Dandekar, 2005, Zhou et al., 2014), data do not suggest that they are infected by SARS-CoV-2, although one study reported ACE2 and SARS-CoV-2 nucleocapsid protein is expressed in lymph nodes and spleen-associated CD169+ macrophages of COVID-19 patients producing IL-6 (Chen et al., 2020h). Significantly elevated systemic levels of proinflammatory cytokine IL-6 have been reported in several COVID-19 patient cohorts and shown to correlate with disease severity (Mehta et al., 2020). Increased IL-6 can also be associated with higher levels of IL-2, IL-7, IFN-ɣ, and GM-CSF, as seen in secondary hemophagocytic lymphohistiocytosis. In response to viral infections, MNPs drive IL and IFN-I and IFN-III production resulting in inflammasome activation, induction of pathogenic Th1 and Th17 cell responses, recruitment of effector immune cells, and CRS pathology (Prokunina-Olsson et al., 2020, Tanaka et al., 2016). Independently, in vitro studies have demonstrated SARS-CoV-1 infection can induce intracellular stress pathways, resulting in NLRP3-dependent inflammasome activation and macrophage pyroptosis (Chen et al., 2019, Shi et al., 2019). Functional studies are required to implicate these myeloid inflammasome pathways in COVID-19 lung pathology and to assess other immunogenic pathways such as RIPK1/3-dependent necroptosis (Nailwal and Chan, 2019). In conclusion, the strength and duration of myeloid ISG)signaling potentially dictate COVID-19 disease severity, but rigorous studies are warranted to confirm this.
Lastly, more work is needed to ascertain the mechanistic role played by lung-resident and recruited granulocytes in SARS-CoV-2 control and pathogenesis (Camp and Jonsson, 2017, Flores-Torres et al., 2019). In contrast to their early protective role, neutrophil NETosis and macrophage crosstalk can drive later-stage inflammatory cascades (Barnes et al., 2020), underscoring the overall pathogenic nature of damage-sensing host responses (Figure 2).
Collectively, the current knowledge of CoVs and SARS-CoV-2 infection, in particular, points to an inadvertent collusion involving myeloid cells in COVID-19 pathogenesis, despite their critical role in early sensing and antiviral responses.
Innate Lymphoid Cells
Innate lymphoid cells (ILCs) are innate immune effector cells that lack the expression of rearranged antigen receptors (T cell receptor [TCR], B cell receptor [BCR]). The ILC family is divided into two main groups: the cytotoxic natural killer (NK) cells and the non-cytotoxic helper ILCs, which include ILC1, ILC2, and ILC3 (Vivier et al., 2018). Conventional NK cells include CD56brightCD16− NK cells and CD56dimCD16+ cells, which are specialized in cytokine production or cytotoxicity, respectively.
NK Cells Are Decreased in the Peripheral Blood of COVID-19 patients
Multiple studies have reported reduced numbers of NK cells in the peripheral blood of COVID-19 patients, which is associated with severity of the disease (Song et al., 2020, Wang et al., 2020f, Yu et al., 2020, Zheng et al., 2020b). A recent scRNA-seq analysis revealed a transcriptomic signature for NK cells that was equally represented in lungs from patients and healthy donors (Liao et al., 2020). The majority of lung NK cells are non-resident (Gasteiger et al., 2015, Marquardt et al., 2017), and CXCR3 has been shown to mediate NK cell infiltration upon influenza infection (Carlin et al., 2018). In vitro, CXCR3 ligands (CXCL9-11) are increased in SARS-CoV-2-infected human lung tissue (Chu et al., 2020), and CXCR3-ligand-producing monocytes are expanded in the lungs of COVID-19 patients (Liao et al., 2020). This suggests that the CXCR3 pathway might facilitate NK cell recruitment from the peripheral blood to the lungs in COVID-19 patients (Figure 2).
NK Cell Activation Pathways in Antiviral Immunity
NK cells express inhibitory and activating receptors that regulate their cytotoxicity. They are therefore able to induce the lysis of virus-infected cells that upregulate virus-derived proteins, as well as stress-inducible ligands, which are then recognized by NK-cell-activating receptors, such as NKp46 (Cerwenka and Lanier, 2001, Draghi et al., 2007, Duev-Cohen et al., 2016, Glasner et al., 2012). Future studies should investigate the expression of NK receptor ligands on SARS-CoV-2-infected cells in order to better understand the mechanisms underlying NK cell activation in COVID-19 disease. Further, secretion of IgG1 and IgG3 antibodies during SARS-CoV-2 infection (Amanat et al., 2020) may induce CD56dim CD16+ NK cell activation through Fc receptor recognition of antibodies either bound to surface antigens expressed on infected cells or to extracellular virions as immune complexes (Figure 2). This interaction might trigger both cytokine production by NK cells and lysis of infected cells through antibody-mediated cellular cytotoxicity (ADCC), as shown in influenza infection (Von Holle and Moody, 2019). Emerging data highlight the capacity for NK-mediated ADCC in response to naturally isolated SARS-CoV-1 anti-S IgG that crossreacts with SARS-CoV-2 S glycoprotein when transfected into Chinese hamster ovary (CHO) cells (Pinto et al., 2020). These findings suggest that triggering NK cell activation may not only contribute to the resolution of infection, but also contribute to the cytokine storm in ARDS.
Impairment of NK Cell Function in SARS-CoV-2 Infection
Ex vivo NK cells from peripheral blood of COVID-19 patients have reduced intracellular expression of CD107a, Ksp37, granzyme B, and granulysin, suggesting an impaired cytotoxicity, as well as an impaired production of chemokines, IFN-ɣ, and TNF-α (Wilk et al., 2020, Zheng et al., 2020b). Several pathways may contribute to the dysregulation of NK cells. While influenza virus infects NK cells and induces apoptosis (Mao et al., 2009), lung NK cells do not express the entry receptor for SARS-CoV-2, ACE2, and are therefore unlikely to be directly infected by SARS-CoV-2 (Travaglini et al., 2020). The majority of NK cells found in human lung display a mature CD16+KIR+CD56dim phenotype and are able to induce cell cytotoxicity in response to loss of human leukocyte antigen (HLA) class I or through Fc receptor signaling, although to a lower extent than their peripheral blood counterpart (Marquardt et al., 2017). Killer-immunoglobulin receptors (KIRs) are acquired during NK cell development alongside CD16 (FcRγIIIA) and are essential for NK cell licensing and subsequent capacity for cytolytic function (Sivori et al., 2019). Frequencies of NK cells expressing CD16 and/or KIRs are decreased in the blood following SARS-CoV-2 and SARS-CoV-1 infection, respectively (Xia et al., 2004, Wang et al., 2020d). Collectively, the data suggest either an impaired maturation of the NK compartment or migration of the mature, circulating NK cells into the lungs or other peripheral tissues of SARS-CoV-2-infected patients.
The immune checkpoint NKG2A is increased on NK cells and CD8 T cells from COVID-19 patients (Zheng et al., 2020b). NKG2A inhibits cell cytotoxicity by binding the non-classical HLA-E molecule (Braud et al., 1998, Brooks et al., 1997), and this interaction is strongly correlated with poor control of HIV-1 infection (Ramsuran et al., 2018). Genes encoding the inhibitory receptors LAG3 and TIM3 are also upregulated in NK cells from COVID-19 patients (Wilk et al., 2020, Hadjadj et al., 2020). Thus, increased immune checkpoints on NK cells might contribute to viral escape. Additionally, COVID-19 patients have higher plasma concentrations of IL-6 (Huang et al., 2020b), which significantly correlate with lower NK cell numbers (Wang et al., 2020d, Wang et al., 2020f). In vitro stimulation by IL-6 and soluble IL-6 receptor has previously revealed impaired cytolytic functions (perforin and granzyme B production) by healthy donor NK cells, which can be restored following addition of tocilizumab (IL-6R blockade) (Cifaldi et al., 2015). TNF-α is also upregulated in the plasma of COVID-19 patients (Huang et al., 2020b), and ligand-receptor interaction analysis of peripheral blood scRNA-seq data suggests that monocyte-secreted TNF-α might bind to its receptors on NK cells (Guo et al., 2020). TNF-α is known to contribute to NK cell differentiation (Lee et al., 2009), which includes downregulation of NKp46 (Ivagnès et al., 2017), though no effect of TNF-α or IL-6 on NK cell-mediated ADCC has been reported so far. Collectively, these data suggest that crosstalk with monocytes might impair NK cell recognition and killing of SARS-CoV-2-infected cells, and antibodies targeting IL-6 and TNF-signaling may benefit enhanced NK cell functions in COVID-19 patients (Figure 2).
Relevance for Helper ILCs in SARS-CoV-2 Infection
No studies, to date, have reported ILC1, ILC2, or ILC3 functions in SARS-CoV-2 infection. All three subsets are present in healthy lung (De Grove et al., 2016, Yudanin et al., 2019). ILC2s are essential for the improvement of lung function following influenza infection in mice through amphiregulin-mediated restoration of the airway epithelium and oxygen saturation (Monticelli et al., 2011). However, ILC2s also produce IL-13, contributing to the recruitment of macrophages to the lung and influenza-induced airway hyperreactivity (Chang et al., 2011). Indeed, ILCs are involved in the polarization of alveolar macrophages, either toward a M1-like phenotype (ILC1 and ILC3) or a M2-like phenotype (ILC2) (Kim et al., 2019). Given the increased IL-13 concentrations (Huang et al., 2020b) and the dysregulation of the macrophage compartment observed in COVID-19 patients, the role played by ILCs in SARS-CoV-2 infection warrants further investigation.
T Cell Responses
T cells play a fundamental role in viral infections: CD4 T cells provide B cell help for antibody production and orchestrate the response of other immune cells, whereas CD8 T cells kill infected cells to reduce the viral burden. However, dysregulated T cell responses can result in immunopathology. To better understand the role of T cell responses in SARS-CoV-2 infection, the pursuit of two major questions is imperative: (1) what is the contribution of T cells to initial virus control and tissue damage in the context of COVID-19, and (2) how do memory T cells established thereafter contribute to protective immunity upon reinfection? Some tentative answers are beginning to emerge.
Overall Reduction of CD4 and CD8 T Cell Counts in Peripheral Blood
Similar to earlier observations about SARS-CoV-1 infection (He et al., 2005), several current reports emphasize the occurrence of lymphopenia with drastically reduced numbers of both CD4 and CD8 T cells in moderate and severe COVID-19 cases (Figure 3 ) (Chen et al., 2020c, Nie et al., 2020b, Wang et al., 2020d, Zeng et al., 2020, Zheng et al., 2020b). The extent of lymphopenia—most striking for CD8 T cells in patients admitted to the intensive care unit (ICU)—seemingly correlates with COVID-19-associated disease severity and mortality (Chen et al., 2020c, Diao et al., 2020, Liu et al., 2020b, Liu et al., 2020c, Tan et al., 2020a, Wang et al., 2020d, Wang et al., 2020f, Zeng et al., 2020, Zhou et al., 2020c). Patients with mild symptoms, however, typically present with normal or slightly higher T cell counts (Liu et al., 2020a, Thevarajan et al., 2020). The cause of peripheral T cell loss in moderate to severe COVID-19, though a phenomenon also observed in other viral infections, remains elusive, and direct viral infection of T cells, in contrast to MERS-CoV (Chu et al., 2016), has not been reported.
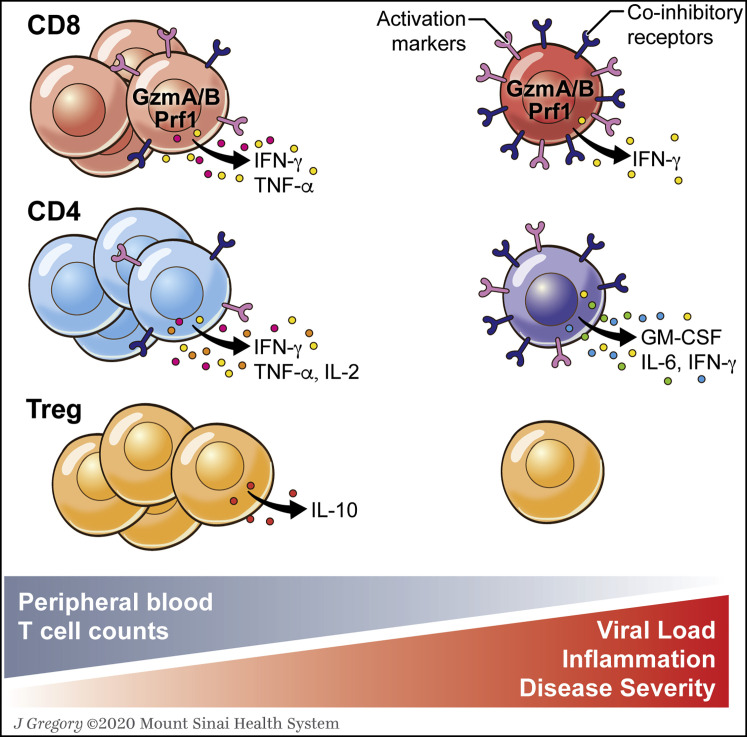
Figure 3.
Working Model for T Cell Responses to SARS-CoV-2: Changes in Peripheral Blood T Cell Frequencies and Phenotype
A decrease in peripheral blood T cells associated with disease severity and inflammation is now well documented in COVID-19. Several studies report increased numbers of activated CD4 and CD8 T cells, which display a trend toward an exhausted phenotype in persistent COVID-19, based on continuous and upregulated expression of inhibitory markers as well as potential reduced polyfunctionality and cytotoxicity. In severe disease, production of specific inflammatory cytokines by CD4 T cells has also been reported. This working model needs to be confirmed and expanded on in future studies to assess virus-specific T cell responses both in peripheral blood and in tissues. In addition, larger and more defined patient cohorts with longitudinal data are required to define the relationship between disease severity and T cell phenotype.
IL, interleukin; IFN, interferon; TNF, tumor necrosis factor; GM-CSF, granulocyte-macrophage colony-stimulating factor; GzmA/B, granzyme A/granzyme B; Prf1, perforin.
Several mechanisms likely contribute to the reduced number of T cells in the blood, including effects from the inflammatory cytokine milieu. Indeed, lymphopenia seems to correlate with serum IL-6, IL-10, and TNF-α (Diao et al., 2020, Wan et al., 2020a), while convalescent patients were found to have restored bulk T cell frequencies paired with overall lower proinflammatory cytokine levels (Chen et al., 2020f, Diao et al., 2020, Liu et al., 2020a, Liu et al., 2020b, Zheng et al., 2020b). Cytokines such as IFN-I and TNF-α may inhibit T cell recirculation in blood by promoting retention in lymphoid organs and attachment to endothelium (Kamphuis et al., 2006, Shiow et al., 2006). However, in an autopsy study examining the spleens and hilar lymph nodes of six patients who succumbed to COVID-19, Chen et al. observed extensive cell death of lymphocytes and suggested potential roles for IL-6 as well as Fas-FasL interactions (Chen et al., 2020h). In support of this hypothesis, the IL-6 receptor antagonist tocilizumab was found to increase the number of circulating lymphocytes (Giamarellos-Bourboulis et al., 2020). T cell recruitment to sites of infection may also reduce their presence in the peripheral blood compartment. scRNA-seq analysis of bronchoalveolar lavage (BAL) fluid of COVID-19 patients revealed an increase in CD8 T cell infiltrate with clonal expansion (Liao et al., 2020). Likewise, post-mortem examination of a patient who succumbed to ARDS following SARS-CoV-2 infection showed extensive lymphocyte infiltration in the lungs (Xu et al., 2020c). However, another study that examined post-mortem biopsies from four COVID-19 patients only found neutrophilic infiltration (Tian et al., 2020a). Further studies are therefore needed to better determine the cause and impact of the commonly observed lymphopenia in COVID-19 patients.
Induction of Antiviral T Cell Responses
Available information about SARS-CoV-1-specific T cell immunity may serve as an orientation for further understanding of SARS-CoV-2 infection. Immunogenic T cell epitopes are distributed across several SARS-CoV-1 proteins (S, N, and M, as well as ORF3), although CD4 T cell responses were more restricted to the S protein (Li et al., 2008). In SARS-CoV-1 survivors, the magnitude and frequency of specific CD8 memory T cells exceeded that of CD4 memory T cells, and virus-specific T cells persisted for at least 6–11 years, suggesting that T cells may confer long-term immunity (Ng et al., 2016, Tang et al., 2011). Limited data from viremic SARS patients further indicated that virus-specific CD4 T cell populations might be associated with a more severe disease course, since lethal outcomes correlated with elevated Th2 cell (IL-4, IL-5, IL-10) serum cytokines (Li et al., 2008). However, the quality of CD4 T cell responses needs to be further characterized to understand associations with disease severity. Few studies have thus far characterized specific T cell immunity in SARS-CoV-2 infection. In 12 patients recovering from mild COVID-19, robust T cell responses specific for viral N, M, and S proteins were detected by IFN-γ ELISPOT, weakly correlated with neutralizing antibody concentrations (similar to convalescent SARS-CoV-1 patients; Li et al., 2008), and subsequently contracted with only N-specific T cells detectable in about one-third of the cases post recovery (Ni et al., 2020). In a second study, PBMCs from COVID-19 patients with moderate to severe ARDS were analyzed by flow cytometry approximately 2 weeks after ICU admission (Weiskopf et al., 2020). Both virus-specific CD4 and CD8 T cells were detected in all patients at average frequencies of 1.4% and 1.3%, respectively, and very limited phenotyping according to CD45RA and CCR7 expression status characterized these cells predominantly as either CD4 Tcm (central memory) or CD8 Tem (effector memory) and Temra (effector memory RA) cells. This study is notable for the use of large complementary peptide pools comprising 1,095 SARS-Cov-2 epitopes (overlapping 15-mers for S protein as well as computationally predicted HLA-I- and -II-restricted epitopes for all other viral proteins) as antigen-specific stimuli that revealed a preferential specificity of both CD4 and CD8 T cells for S protein epitopes, with the former population modestly increasing over ∼10–30 days after initial onset of symptoms. A caveat, however, pertains to the identification of specific T cells by induced CD69 and CD137 co-expression, since upregulation of CD137 by CD4 T cells, in contrast to CD154, may preferentially capture regulatory T cells (Treg) (Bacher et al., 2016). Further analyses of S protein-specific T cells by ELISA demonstrated robust induction of IFN-γ, TNF-α, and IL-2 concomitant with lower levels of IL-5, IL-13, IL-9, IL-10, and IL-22. A third report focused on S-specific CD4 T cell responses in 18 patients with mild, severe, or critical COVID-19 using overlapping peptide pools and induced CD154 and CD137 co-expression as a readout for antiviral CD4 T cells. Such cells were present in 83% of cases and presented with enhanced CD38, HLA-DR, and Ki-67 expression indicative of recent in vivo activation (Braun et al., 2020). Of note, the authors also detected low frequencies of S-reactive CD4 T cells in 34% of SARS-CoV-2 seronegative healthy control donors. However, these CD4 T cells lacked phenotypic markers of activation and were specific for C-terminal S protein epitopes that are highly similar to endemic human CoVs, suggesting that crossreactive CD4 memory T cells in some populations (e.g., children and younger patients that experience a higher incidence of hCoV infections) may be recruited into an amplified primary SARS-CoV-2-specific response (Braun et al., 2020). Similarly, endemic CoV-specific CD4 T cells were previously shown to recognize SARS-CoV1 determinants (Gioia et al., 2005). How previous infections with endemic CoV may affect immune responses to SARS-CoV-2 will need to be further investigated.
Finally, in general accordance with the above findings on the induction of SARS-CoV-2-specific T cells, using TCR sequencing (TCR-seq), Huang et al. and Liao et al. reported greater TCR clonality of peripheral blood (Huang et al., 2020c) as well as BAL T cells (Liao et al., 2020) in patients with mild versus severe COVID-19. Moving forward, a comprehensive identification of immunogenic SARS-CoV-2 epitopes recognized by T cells (Campbell et al., 2020), as well as further studies on convalescent patients who recovered from mild and severe disease, will be particularly important.
T Cell Contribution to COVID-19 Hyperinflammation
While the induction of robust T cell immunity is likely essential for efficient virus control, dysregulated T cell responses may cause immunopathology and contribute to disease severity in COVID-19 patients (Figure 3). This is suggested in a study by Zhou et al., which reported a significantly increased PBMC frequency of polyclonal GM-CSF+ CD4 T cells capable of prodigious ex vivo IL-6 and IFN-γ production only in critically ill COVID-19 patients (Zhou et al., 2020c). Of note, GM-CSF+ CD4 T cells have been previously implicated in inflammatory autoimmune diseases, such as multiple sclerosis or juvenile rheumatoid arthritis, and high levels of circulating GM-CSF+ CD4 T cells were found to be associated with poor outcomes in sepsis (Huang et al., 2019). Additionally, two studies observed reduced frequencies of Treg cells in severe COVID-19 cases (Chen et al., 2020c, Qin et al., 2020). Since Treg cells have been shown to help resolve ARDS inflammation in mouse models (Walter et al., 2018), a loss of Tregs might facilitate the development of COVID-19 lung immunopathology. Similarly, a reduction of γδ-T cells, a subset of T cells with apparent protective antiviral function in influenza pneumonia (Dong et al., 2018, Zheng et al., 2013), has been reported in severely sick COVID-19 patients (Guo et al., 2020, Lei et al., 2020b).
Phenotype and Function of T Cell Subsets in COVID-19
Currently, little is known about specific phenotypical and/or functional T cell changes associated with COVID-19. In the majority of preprints and peer-reviewed studies, there are reports of increased presence of activated T cells (Figure 3) characterized by expression of HLA-DR, CD38, CD69, CD25, CD44, and Ki-67 (Braun et al., 2020, Ni et al., 2020, Guo et al., 2020, Liao et al., 2020, Thevarajan et al., 2020, Yang et al., 2020a, Zheng et al., 2020a). Generally, independent of COVID-19 disease severity, CD8 T cells seem to be more activated than CD4 T cells (Qin et al., 2020, Thevarajan et al., 2020, Yang et al., 2020a), a finding that echoes stronger CD8 than CD4 T cell responses during SARS-CoV-1 (Li et al., 2008). Furthermore, in a case study of 10 COVID-19 patients, Diao et al. showed that levels of PD-1 increased from prodromal to symptomatic stages of the disease (Diao et al., 2020). PD-1 expression is commonly associated with T cell exhaustion, but it is important to emphasize that PD-1 is primarily induced by TCR signaling; it is thus also expressed by activated effector T cells (Ahn et al., 2018).
In addition, several studies reported higher expression of various co-stimulatory and inhibitory molecules such as OX-40 and CD137 (Zhou et al., 2020c), CTLA-4 and TIGIT (Zheng et al., 2020a), and NKG2a (Zheng et al., 2020b). Reduced numbers of CD28+ CD8 T cells (Qin et al., 2020) as well as larger frequencies of PD-1+ TIM3+ CD8 T cells in ICU patients were also reported (Zhou et al., 2020c). Expression of most of these markers was found to be higher in CD8 than in CD4 T cells, and levels tended to increase in severe versus non-severe cases, which may be due to differences in viral load. Cellular functionality was shown to be impaired in CD4 and CD8 T cells of critically ill patients, with reduced frequencies of polyfunctional T cells (producing more than one cytokine) as well as generally lower IFN-γ and TNF-α production following restimulation with phorbol myristate acetate (PMA) and ionomycin (Chen et al., 2020c, Zheng et al., 2020a, Zheng et al., 2020b). Similarly, Zheng et al. reported that CD8 T cells in severe COVID-19 appear less cytotoxic and effector-like with reduced CD107a degranulation and granzyme B (GzmB) production (Zheng et al., 2020b). In contrast, a different study found that both GzmB and perforin were increased in CD8 T cells of severely sick patients (Zheng et al., 2020a). In accordance with the latter observation, when compared to a moderate disease group, convalescent patients with resolved severe SARS-CoV-1 infection had significantly higher frequencies of polyfunctional T cells, with CD4 T cells producing more IFN-γ, TNF-α, and IL-2 and CD8 T cells producing more IFN-γ, TNF-α, and CD107a, respectively (Li et al., 2008). However, given the vigorous dynamics of acute T cell responses and potential differences in sample timing throughout disease course, these observations are not necessarily mutually exclusive. Accordingly, RNA sequencing (RNA-seq) data by Liao et al. showed that CD8 T cells in the BAL fluid of severe COVID-19 patients express cytotoxic genes such as GZMA, GZMB, and GZMK at higher levels, while KLRC1 and XCL1 are enriched in mild cases (Liao et al., 2020).
In summary, T cells in severe COVID-19 seem to be more activated and may exhibit a trend toward exhaustion based on continuous expression of inhibitory markers such as PD-1 and TIM-3 as well as overall reduced polyfunctionality and cytotoxicity. Conversely, recovering patients were shown to have an increase in follicular helper CD4 T cells (TFH) as well as decreasing levels of inhibitory markers along with enhanced levels of effector molecules such as Gzm A, GzmB, and perforin (Thevarajan et al., 2020, Yang et al., 2020a, Zheng et al., 2020b). Collectively, these studies provide a first glimpse into T cell dynamics in acute SARS-CoV-2 infection, but any conclusions have to be tempered at this stage on account of significant limitations in many of the current investigations.
B Cell Responses
Acute B Cell and Antibody Responses
The humoral immune response is critical for the clearance of cytopathic viruses and is a major part of the memory response that prevents reinfection. SARS-CoV-2 elicits a robust B cell response, as evidenced by the rapid and near-universal detection of virus-specific IgM, IgG and IgA, and neutralizing IgG antibodies (nAbs) in the days following infection. The kinetics of the antibody response to SARS-CoV-2 are now reasonably well described (Huang et al., 2020a).
Similar to SARS-CoV-1 infection (Hsueh et al., 2004), seroconversion occurs in most COVID-19 patients between 7 and 14 days after the onset of symptoms, and antibody titers persist in the weeks following virus clearance (Figure 4 ) (Haveri et al., 2020, Lou et al., 2020, Okba et al., 2020, Tan et al., 2020b, Wölfel et al., 2020, Wu et al., 2020b, Zhao et al., 2020a). Antibodies binding the SARS-CoV-2 internal N protein and the external S glycoprotein are commonly detected (Amanat et al., 2020, Ju et al., 2020, To et al., 2020). The receptor binding domain (RBD) of the S protein is highly immunogenic, and antibodies binding this domain can be potently neutralizing, blocking virus interactions with the host entry receptor, ACE2 (Ju et al., 2020, Wu et al., 2020b). Anti-RBD nAbs are detected in most tested patients (Ju et al., 2020, To et al., 2020, Wu et al., 2020b). Although crossreactivity to SARS-CoV-1 S and N proteins and to MERS-CoV S protein was detected in plasma from COVID-19 patients, no crossreactivity was found to the RBD from SARS-CoV-1 or MERS-CoV. In addition, plasma from COVID-19 patients did not neutralize SARS-CoV-1 or MERS-CoV (Ju et al., 2020).
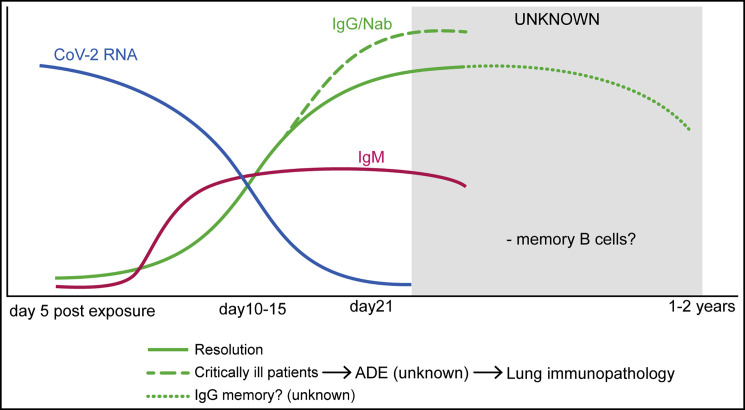
Figure 4.
Antibody-Mediated Immunity in SARS-CoV-2
Virus-specific IgM and IgG are detectable in serum between 7 and 14 days after the onset of symptoms. Viral RNA is inversely correlated with neutralizing antibody titers. Higher titers have been observed in critically ill patients, but it is unknown whether antibody responses somehow contribute to pulmonary pathology. The SARS-CoV-1 humoral response is relatively short lived, and memory B cells may disappear altogether, suggesting that immunity with SARS-CoV-2 may wane 1–2 years after primary infection.
RBD-specific CD19+IgG+ memory B cells were single-cell sorted from a cohort of eight COVID-19 donors between days 9 and 28 after the onset of symptoms (Ju et al., 2020). From their antibody gene sequences, 209 SARS-CoV-2-specific monoclonal antibodies were produced. The monoclonal antibodies had a diverse repertoire, relatively low or no somatic mutations, and variable binding reactivity, with dissociation constants reaching 10−8 to 10−9, similar to antibodies isolated during acute infections. Two potent neutralizing SARS-CoV-2 RBD-specific monoclonal antibodies were characterized that did not crossreact with the RBD of SARS-CoV-1 or MERS-CoV (Ju et al., 2020). Together, these results demonstrate that antibody mediated neutralization is virus specific and likely driven by binding of epitopes within the RBD.
B Cell Memory: Development and Lifespan
The B cell response to a virus serves not only to protect from the initial challenge, but also to offer extended immunity against reinfection. Following resolution of an infection, plasma cells formed during the acute and convalescent phases continue to secrete antibodies, giving rise to serological memory. Memory B cells that are also formed during the primary infection constitute the second arm of B cell memory. Memory B cells can quickly respond to a reinfection by generating new high-affinity plasma cells. Long-term protection is achieved through the induction of long-lived plasma cells and memory B cells.
There is great interest in understanding the lifespan of B cell memory responses to SARS-CoV-2. Protection from reinfection has direct medical and social consequences as the world works to develop vaccination strategies and resume normal activities. In COVID-19 patients, evidence of near-universal seroconversion and the lack of substantial descriptions of reinfection point to a robust antibody response, which, along with the T cell memory response, would offer protection to reinfection. Indeed, a case study of a single patient described induction of CD38HiCD27Hi antibody-secreting cells (ASCs), concomitant with an increase in circulating follicular T helper cells (Tfh) cells (Thevarajan et al., 2020), and a scRNA-seq study of PBMCs from critically ill and recently recovered individuals revealed a plasma cell population (Guo et al., 2020). In addition, IgG memory cells specific to the RBD have been identified in the blood of COVID-19 patients (Ju et al., 2020). Consistent with the development of immunity after COVID-19 infection, a recent study of SARS-CoV-2 infection in rhesus macaques found that two macaques that had resolved the primary infection were resistant to reinfection 28 days later (Bao et al., 2020b).
Due to the timing of this outbreak, it is not yet possible to know the nature and extent of long-term memory responses, but lessons may again be learned from other human CoVs. In the case of the human CoV 229E, specific IgG and nAbs are rapidly induced but wane in some individuals around a year after infection, with some residual protection to reinfection (Callow et al., 1990, Reed, 1984). The lifespan of the humoral response following SARS-CoV-1 infection is also relatively short, with the initial specific IgG and nAb response to SARS-CoV-1 diminishing 2–3 years after infection and nearly undetectable in up to 25% of individuals (Cao et al., 2007, Liu et al., 2006). A long-term study following 34 SARS-CoV-1-infected healthcare workers over a 13-year period also found that virus-specific IgG declined after several years, but the authors observed detectable virus-specific IgG 12 years after infection (Guo et al., 2020). In the case of MERS-CoV, antibodies were detected in six of seven volunteers tested 3 years after infection (Payne et al., 2016).
IgG specific to SARS-CoV-2 trimeric spike protein was detectable in serum up to 60 days after symptom onset, but IgG titers began decreasing by 8 weeks post symptom onset (Adams et al., 2020). Long-term protection from reinfection may also be mediated by reactive memory B cells. A study that analyzed SARS-CoV-1 S protein-specific IgG memory cells at 2, 4, 6, and 8 months post infection found that S-specific IgG memory B cells decreased progressively about 90% from 2 to 8 months after infection (Traggiai et al., 2004). A further retrospective study of 23 individuals found no evidence of circulating SARS-CoV-1-specific IgG+ memory B cells 6 years after infection (Tang et al., 2011). This is in contrast to the memory T cell response, which was robustly detected based on induced IFN-γ production (Tang et al., 2011).
Studies of common CoVs SARS-CoV-1 and MERS-CoV indicate that virus-specific antibody responses wane over time and, in the case of common CoVs, result in only partial protection from reinfection. These data suggest that immunity to SARS-CoV-2 may diminish following a primary infection, and further studies will be required to determine the degree of long-term protection (Figure 4).
Consequences of the B Cell Response: Protection versus Enhancement
Several studies have demonstrated that high virus-specific antibody titers to SARS-CoV-2 are correlated with greater neutralization of virus in vitro and are inversely correlated with viral load in patients (Figure 4) (Okba et al., 2020, Wölfel et al., 2020, Zhao et al., 2020a). Despite these indications of a successful neutralizing response in the majority of individuals, higher titers are also associated with more severe clinical cases (Li et al., 2020b, Okba et al., 2020, Zhao et al., 2020a, Zhou et al., 2020a), suggesting that a robust antibody response alone is insufficient to avoid severe disease (Figure 4).
This was also observed in the previous SARS-CoV-1 epidemic, where neutralizing titers were found to be significantly higher in deceased patients compared to patients who had recovered (Zhang et al., 2006). This has led to concerns that antibody responses to these viruses may contribute to pulmonary pathology via antibody-dependent enhancement (ADE) (Figure 4). This phenomenon is observed when non-neutralizing virus-specific IgG facilitate entry of virus particles into Fc-receptor (FcR) expressing cells, particularly macrophages and monocytes, leading to inflammatory activation of these cells (Taylor et al., 2015). A study in SARS-CoV-1-infected rhesus macaques found that anti-S IgG contributed to severe acute lung injury (ALI) and massive accumulation of monocytes and macrophages in the lung (Liu et al., 2019). Furthermore, serum containing anti-S Ig from SARS-CoV-1 patients enhanced the infection of SARS-CoV-1 in human monocyte-derived macrophages in vitro (Yip et al., 2014). ADE was also reported with a monoclonal antibody isolated from a patient with MERS-CoV (Wan et al., 2020c). Somewhat reassuringly, there was no evidence of ADE mediated by sera from rats vaccinated with SARS-CoV-2 RBD in vitro (Quinlan et al., 2020) nor in macaques immunized with an inactivated SARS-CoV-2 vaccine candidate (Gao et al., 2020c).
As of now, there is no evidence that naturally developed antibodies toward SARS-CoV-2 contribute to the pathological features observed in COVID-19. However, this possibility should be considered when it comes to experimental design and development of therapeutic strategies. Importantly, in all of the descriptions of ADE as it relates to CoV, the FcR was necessary to trigger the antibody-mediated pathology. High-dose intravenous immunoglobulin (IVIg), which may blunt ADE, has been trialed in COVID-19 patients (Cao et al., 2020b, Shao et al., 2020), but further studies are needed to determine the extent to which IVIg is safe or beneficial in SARS-CoV-2 infection. Vaccine trials will need to consider the possibility of antibody-driven pathology upon antigen rechallenge; strategies using F(ab) fragments or engineered Fc monoclonal antibodies may prove particularly beneficial in this setting (Amanat and Krammer, 2020).
Predictors of COVID-19 Disease Risk and Severity
With the rapidly growing number of cases in the first few months, numerous reports on predictors of COVID-19 severity with small cohorts were released. These offered clinicians and immunologists the first understanding of the clinical course and pathological processes that are associated with the novel SARS-CoV-2 infection. This section highlights key findings from those studies, with a major focus on the immune factors associated with disease risk or severity.
Susceptibility and Risk Biomarkers
There are currently limited known risk factors for susceptibility to COVID-19, although this has been evaluated in several studies. Zhao et al. compared the ABO blood group distribution in a cohort of 2,173 COVID-19 patients to that of healthy controls from the corresponding regions (Zhao et al., 2020b). They found blood group A to be associated with a higher risk for acquiring COVID-19 when compared to non-A blood groups; blood group O had the lowest risk for the infection. Another study demonstrated an identical association (Zietz and Tatonetti, 2020), and similar results have been previously described for other viruses (Lindesmith et al., 2003), including SARS-CoV-1 (Cheng et al., 2005a).
Several large collaborative efforts are currently underway to generate, share, and analyze genetic data to understand the links between human genetic variation and COVID-19 susceptibility and severity, the most prominent of which is the COVID-19 Host Genetics Initiative (covid19hg.org). These studies are supported by previous observations on SARS-CoV-1 that followed the 2003 outbreak, which have identified significant associations between genetic variants and immune phenotypes (Chan et al., 2007, Wang et al., 2011, Zhao et al., 2011). Although identifying such polymorphisms and their associated genes and pathways for SARS-CoV-2 will require large cohorts, several studies, which remain to be tested in clinical trials, have already highlighted genetic polymorphisms that may potentially impact susceptibility. These studies have focused on genetic variants that may impact the expression or function of genes important in viral entry, namely ACE2 (SARS-CoV-2 receptor) and TMPRSS2 (spike protein activator) (Asselta et al., 2020, Cao et al., 2020c, Renieri et al., 2020, Stawiski et al., 2020). Cao et al. identified variants that are potentially expression quantitative trait loci (eQTL) of ACE2 (i.e., they may potentially alter ACE2 gene expression) and analyzed their frequencies in different populations (Cao et al., 2020c). Stawiski et al. listed variants that may be critical in ACE2 binding and thereby its function and compared the frequencies of these variants within different populations (Stawiski et al., 2020).
While there are several limitations to these studies, the major question is whether the utility of these biomarkers is replicable in large populations with COVID-19 clinical outcomes data and in targeted or large-scale genomic analyses that are currently underway. In addition, these studies will reveal the potential associations between genetic variants and susceptibility in a gene or loci agnostic fashion.
Routine Bloodwork Biomarkers
Several routine blood and serological parameters have been suggested to stratify patients who might be at higher risk for complications to aid in allocation of healthcare resources in the pandemic (Table 1 ). Serologic markers from routine bloodwork were reported by comparing patients with mild or moderate symptoms to those with severe symptoms. This includes different acute phase proteins, such as SAA (serum amyloid protein) and C-reactive protein (CRP) (Ji et al., 2020). Interestingly, elevations in CRP appear to be unique to COVID-19 patients when compared to other viral infections. Other consistently reported markers in non-survivors are increased procalcitonin (PCT) and IL-6 levels (Huang et al., 2020d), as well as increased serum urea, creatinine, cystatin C, direct bilirubin, and cholinesterase (Xiang et al., 2020a). Overall, inflammatory markers are common in severe cases of COVID-19 and appear to correlate with the severity of the symptoms and clinical outcome. Moreover, the extensive damage that occurs in specific organs of severe COVID-19 patients is possibly related to differences in the expression of ACE2 (Figure 5 ) (Du et al., 2020).
Table 1.
Routine Blood and Immunological Prognostic Biomarkers in COVID-19 Patients
| Biomarker | Purpose | |
|---|---|---|
| Routine Bloodwork | Lymphocyte count | Predicted the disease severity and the outcomes of hospitalized patients (Tan et al., 2020a). Prognostic value was confirmed in numerous studies (Huang et al., 2020d, Liu et al., 2020b, Song et al., 2020, Wang et al., 2020f, Wynants et al., 2020, Yan et al., 2020b, Yang et al., 2020b, Zhang et al., 2020c, Zhao et al., 2020d). Decreased continuously in non-surviving patients (Wang et al., 2020b). |
| N/L | Patients with N/L ≥3.13 were reported to be more likely to develop severe illness and to require ICU admission (Liu et al., 2020c). N/L on admission was a risk factor for short-term progression of patients with moderate pneumonia to severe pneumonia (Feng et al., 2020). Confirmed to be of prognostic value in COVID-19 in several studies (Song et al., 2020, Wynants et al., 2020, Zhou et al., 2020d). |
|
| CRP | Proposed as an early biomarker of disease progression (Ji et al., 2020). Even in early stages, CRP levels were positively correlated with lung lesions and reflected disease severity (Wang, 2020). Confirmed in numerous studies (Fu et al., 2020, Huang et al., 2020d, Wynants et al., 2020, Yan et al., 2020b, Zhao et al., 2020d, Zhou et al., 2020b). Predicted the risk of acute myocardial injury (Liu et al., 2020g, Xu et al., 2020a). | |
| LDH | Higher in severe cases than in mild cases (Xiang et al., 2020a). Widely proposed to have prognostic value in COVID-19 (Huang et al., 2020d, Wynants et al., 2020, Yan et al., 2020b, Zhao et al., 2020d). | |
| D-dimer (and coagulation parameters) | Predicted severity independently of other variables (Zhou et al., 2020d). Elevated levels and disseminated intravascular coagulation are found in non-survivors (Wang et al., 2020b). Identified patients at risk for acute cardiac injury (Liu et al., 2020g). Other coagulation parameters, such as fibrin degradation product levels, longer prothrombin time, and activated partial thromboplastin time, were also associated with poor prognosis (Tang et al., 2020a). | |
| SAA | SAA was proposed to be used as an auxiliary index for diagnosis as it was elevated in 80% of the patients in a small cohort (Ji et al., 2020). | |
| NT-proBNP (N-terminal pro B type natriuretic peptide) | NT-proBNP was an independent risk factor of in-hospital death in patients with severe COVID-19 (Gao et al., 2020b). | |
| Platelet count | High platelet-to-lymphocyte ratio was associated with worse outcome (Qu et al., 2020). Thrombocytopenia was associated with poor outcome and with incidence of myocardial injury in COVID-19 (Liu et al., 2020h, Shi et al., 2020). | |
| Immunological | CD4+, CD8+, and NK cell counts | Lower CD4+, CD8+, and NK cells in PBMCs correlated with severity of COVID-19 (Nie et al., 2020b). Validated by several studies (Wang et al., 2020f, Zheng et al., 2020b). |
| PD-1 and Tim-3 expression on T cells | Increasing PD-1 and Tim-3 expression on T cells could be detected as patients progressed from prodromal to overtly symptomatic stages (Diao et al., 2020). Expression was higher in infected patients versus healthy controls and in ICU versus non-ICU patients in both CD4 and CD8 T cells (Zhou et al., 2020b). | |
| phenotypic changes in peripheral blood monocytes | The presence of a distinct population of monocytes with high forward scatter (CD11b+, CD14+, CD16+, CD68+, CD80+, CD163+, and CD206+, which secrete IL-6, IL-10, and TNF-α) was identified in patients requiring prolonged hospitalization and ICU admission (Zhang et al., 2020c). CD14+CD16+IL-6+ monocytes are increased in ICU patients (Zhou et al., 2020b). | |
| IP-10, MCP-3, and IL-1ra | IP-10, MCP-3, and IL-1ra were, among 48 examined cytokines, the only ones that closely associated with disease severity and outcome of COVID-19 in a study by Yang et al. (Yang et al., 2020b). | |
| IL-6 | Associated with disease severity (hospitalization and ICU admission) and poor prognosis (Chen et al., 2020g, Huang et al., 2020b, Liu et al., 2020b, Liu et al., 2020f, Wang et al., 2020b). Increased levels were associated with higher risk of respiratory failure (Yao et al., 2020b). | |
| IL-8 | Positively correlated with disease severity (Chen et al., 2020e, Gong et al., 2020), with severe cases showing the highest IL-8 levels. | |
| IL-10 | Increased in severe or critical patients as compared to mild patients (Gong et al., 2020, Zhou et al., 2020d) without a statistically significant difference between severe and critical cases (Gong et al., 2020). | |
| IL-2R | Associated with disease severity in a study that, among other cytokines, also associated ferroprotein levels, PCT levels, and eosinophil counts with COVID-19 severity (Gong et al., 2020). | |
| IL-1β | CD14+IL-1β+ monocytes are abundant in early-recovery patients as shown in a single-cell RNA-seq analysis and thought to be associated with cytokine storm (Wen et al., 2020). IL-1β did not correlate with disease severity in a cross-sectional study with mild, severe, and critical patients (Gong et al., 2020). | |
| IL-4 | IL-4 was associated with impaired lung lesions (Fu et al., 2020), but some reports point to a potential mediator effect (Wen et al., 2020). | |
| IL-18 | In modeling immune cell interaction between DCs and B cells in late recovery COVID-19 patients, IL-18 was found to be important in B cell production of antibodies, which suggests its importance in recovery (Wen et al., 2020). | |
| GM-CSF | GM-CSF+IFN-γ+ T cells are higher in ICU than in non-ICU patients. CD14+CD16+GM-CSF+ monocytes are higher in COVID-19 patients as compared to healthy controls (Zhou et al., 2020b). | |
| IL-2 and IFN-γ | IL-2 and IFN-γ levels were shown to be increased in severe cases (Liu et al., 2020b). | |
| anti-SARS-CoV-2 antibody levels | Prolonged SARS-CoV-2 IgM positivity could be utilized as a predictive factor for poor recovery (Fu et al., 2020). Higher anti-SARS-CoV-2 IgG levels and higher N/L were more commonly found in severe cases (Zhang et al., 2020a). |
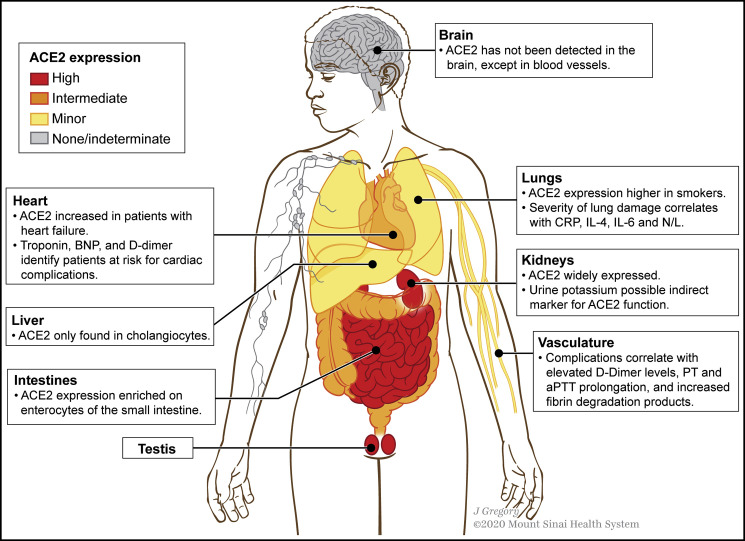
Figure 5.
ACE2 Expression in Organs and Systems Most Frequently Implicated in COVID-19 Complications
The gastrointestinal tract, kidneys, and testis have the highest ACE2 expressions. In some organs, different cell types have remarkably distinct expressions; e.g., in the lungs, alveolar epithelial cells have higher ACE2 expression levels than bronchial epithelial cells; in the liver, ACE2 is not expressed in hepatocytes, Kupffer cells, or endothelial cells but is detected in cholangiocytes, which can explain liver injury to some extent. Furthermore, ACE2 expression is enriched on enterocytes of the small intestine compared to the colon.
ACE2, angiotensin-converting enzyme 2; BNP, B-type natriuretic peptide; CRP, C-reactive protein; IL, interleukin; N/L, neutrophil-to-lymphocyte ratio; PT, prothrombin time; aPTT, activated partial thromboplastin time.
Lymphopenia is the most frequently described prognostic marker in COVID-19 (Table 1), and it appears to predict morbidity and mortality even at early stages (Fei et al., 2020). Tan et al. proposed a prognostic model based on lymphocyte counts at two time points: patients with less than 20% lymphocytes at days 10–12 from the onset of symptoms and less than 5% at days 17–19 had the worst outcomes in this study (Tan et al., 2020a). Wynants et al. compared predictors of disease severity across seven studies (>1,330 patients), highlighting CRP, neutrophil-to-lymphocyte ratio (N/L), and lactate dehydrogenase (LDH) as the most significant predictive biomarkers (Wynants et al., 2020). Furthermore, a meta-analysis of 30 COVID-19 studies with a total of 53,000 patients also attempted to identify early-stage patients with poor prognosis (Zhao et al., 2020d). The most consistent findings across the different studies were elevated levels of CRP, LDH, and D-dimer, as well as decreased blood platelet and lymphocyte counts (Yan et al., 2020b, Zhou et al., 2020d). Systemic and pulmonary thrombi have been reported with activation of the extrinsic coagulation cascade, involving dysfunctional endothelium and monocytic infiltration (Poor et al., 2020, Varga et al., 2020); thrombocytopenia and elevated D-dimer levels may be indicative of these coagulopathies in COVID-19 patients with important therapeutic implications (Fogarty et al., 2020, Poor et al., 2020).
Immunological Biomarkers in the Peripheral Blood
Immunological biomarkers are particularly important, as immunopathology has been suggested as a primary driver of morbidity and mortality with COVID-19. Several cytokines and other immunologic parameters have been correlated with COVID-19 severity (Table 1). Most notably, elevated IL-6 levels were detected in hospitalized patients, especially critically ill patients, in several studies and are associated with ICU admission, respiratory failure, and poor prognosis (Chen et al., 2020g, Huang et al., 2020b, Liu et al., 2020f). Increased IL-2R, IL-8, IL-10, and GM-CSF have been associated with disease severity as well, but studies are limited, and further studies with larger cohorts of patients are needed to indicate predictive power (Gong et al., 2020, Zhou et al., 2020b). Conflicting results regarding IL-1β and IL-4 have been reported (Fu et al., 2020, Gong et al., 2020, Wen et al., 2020). Although elevated cytokine concentrations have been widely described in COVID-19 patients, the vast majority (including IL-6, IL-10, IL-18, CTACK, and IFN-γ) do not seem to have prognostic value, because they do not always differentiate moderate cases from severe cases (Yang et al., 2020b). This stratification was possible with IP-10, MCP-3, and IL-1ra. While there are reports that levels of IL-6 at first assessment might predict respiratory failure (Herold et al., 2020), other publications with longitudinal analyses demonstrated that IL-6 increases fairly late during the disease’s course, consequently compromising its prognostic value at earlier stages (Zhou et al., 2020a).
Liu et al. developed a web-based tool using k-means clustering to predict prognosis in terms of death or hospital discharge of COVID-19 patients using age, comorbidities (binary), and baseline log helper T cell count (TH), log suppressor T cell count (TS), and log TH/TS ratio (Liu et al., 2020e). Total T cell, helper T cell, and suppressor T cell counts were significantly lower and the TH/TS ratio was significantly higher in patients who died from infection as compared to patients who were discharged.
Importantly, most serological and immunological changes observed in severe cases are associated with disease severity but cannot necessarily serve as predictive factors, as they may not have utility in early identification of patients at higher risk. Discovery of truly predictive biomarkers and potential drivers of hyperinflammatory processes requires comprehensive profiling of asymptomatic and mild cases and longitudinal studies that are limited to date. Confounding variables including age, gender, and comorbidities may dramatically affect associations observed. In addition, direct correlation with patient viral load will be important to provide a greater understanding of underlying causes of morbidity and mortality in COVID-19 and the contribution of viral infectivity, hyperinflammation, and host tolerance (Medzhitov et al., 2012).
In summary, lymphopenia, increases in proinflammatory markers and cytokines, and potential blood hypercoagulability characterize severe COVID-19 cases with features reminiscent of cytokine release syndromes. This correlates with a diverse clinical spectrum ranging from asymptomatic to severe and critical cases. During the incubation period and early phase of the disease, leukocyte and lymphocyte counts are normal or slightly reduced. After SARS-CoV-2 binds to ACE2 overexpressing organs, such as the gastrointestinal tracts and kidneys, increases in non-specific inflammation markers are observed. In more severe cases, a marked systemic release of inflammatory mediators and cytokines occurs, with corresponding worsening of lymphopenia and potential atrophy of lymphoid organs, impairing lymphocyte turnover (Terpos et al., 2020).
Antivirals
Antivirals are a class of small molecules that function as inhibitors of one or more stages of a virus life cycle. Because of similarities between different virus replication mechanisms, some antivirals can be repurposed against various viral infections. Currently, most of the available antiviral drugs tested against SARS-CoV-2 are small molecules previously developed against SARS-CoV-1, MERS-CoV, or other RNA and DNA viruses.
Broad-Spectrum Antivirals
A number of small molecules with known antiviral activity against other human RNA viruses are being evaluated for efficacy in treating SARS-CoV-2. The ribonucleoside analog β-D-N4-hydroxycytidine (NHC) reduced viral titers and lung injury in mice infected with SARS-CoV-2 via introduction of mutations in viral RNA (Sheahan et al., 2017). Further, an inhibitor of host DHODH, a rate-limiting enzyme in pyrimidine synthesis, was able to inhibit SARS-CoV-2 growth in vitro with greater efficacy than remdesivir or chloroquine (Wang et al., 2020e, Xiong et al., 2020). Merimepodib, a non-competitive inhibitor of the enzyme inosine-5′-monophosphate dehydrogenase (IMPDH), involved in host guanosine biosynthesis, is able to suppress SARS-CoV-2 replication in vitro (Bukreyeva et al., 2020). Finally, N-(2-hydroxypropyl)-3-trimethylammonium chitosan chloride (HTCC), which was previously shown to efficiently reduce infection by the less-pathogenic human CoV HCoV-NL63, was also found to inhibit MERS-CoV and pseudotyped SARS-CoV-2 in human airway epithelial cells (Milewska et al., 2020).
Protease Inhibitors
Much of the antiviral computational and experimental data currently available for SARS-CoV-2 focus on targeting the 3CL or Main protease (Mpro). Two prominent drug candidates targeting the SARS-CoV-2 Mpro were designed and synthesized by analyzing the substrate binding pocket of Mpro (Dai et al., 2020). The X-ray crystal structures of the novel inhibitors in complex with SARS-CoV-2 Mpro were resolved at 1.5 Å. Both compounds showed good pharmacokinetic activity in vitro, and one exhibited limited toxicity in vivo (Dai et al., 2020). Multiple studies also aimed to repurpose protease inhibitors to reduce SARS-CoV-2 titers. Nine existing HIV protease inhibitors (nelfinavir, lopinavir, ritonavir, saquinavir, atazanavir, tipranavir, amprenavir, darunavir, and indinavir) were evaluated for their antiviral activity in Vero cells infected with SARS-CoV-2 (Yamamoto et al., 2020), and nelfinavir was the most potent at inhibiting viral replication.
RdRp Inhibitors
The CoV RNA-dependent RNA polymerase (RdRp) catalyzes the synthesis of viral RNA (Gao et al., 2020a), making it essential for viral replication and a prime target for antiviral inhibitors. Remdesivir, an adenosine triphosphate analog, inhibits RdRp by binding to RNA strands and preventing additional nucleotides from being added, thereby terminating viral RNA transcription (Figure 6 A) (Agostini et al., 2018). Remdesivir has been previously shown to be effective against MERS-CoV and SARS-CoV-1 infections in animal models (Sheahan et al., 2017, de Wit et al., 2020). Similarly, a study investigated the efficacy of remdesivir treatments on 12 rhesus macaques with SARS-CoV-2 infections (Williamson et al., 2020). Macaques treated with remdesivir showed a reduction in lung viral loads and pneumonia symptoms but no reduction in virus shedding. This study does provide evidence that if administered early enough, remdesivir may be effective at treating SARS-CoV-2 infections.
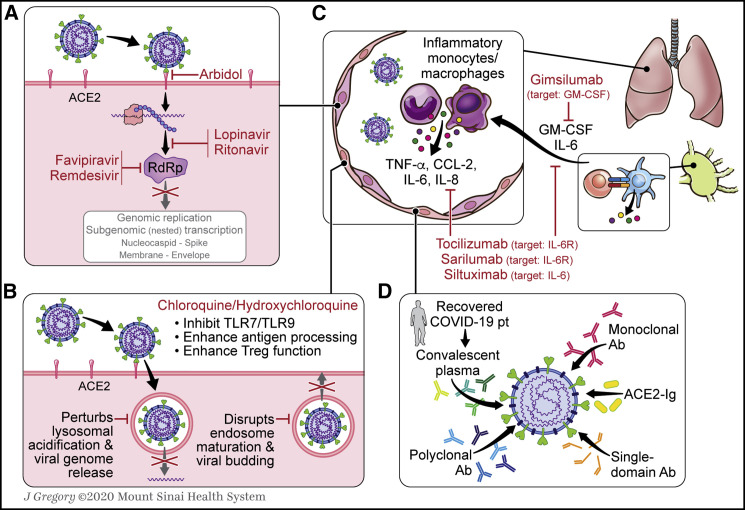
Figure 6.
Available Therapeutic Options to Manage COVID-19 Immunopathology and to Deter Viral Propagation
(A) Rdrp inhibitors (remdesivir, favipiravir), protease inhibitors (lopinavir/ritonavir), and antifusion inhibitors (arbidol) are currently being investigated in their efficacy in controlling SARS-CoV-2 infections.
(B) CQ and HCQ increase the pH within lysosomes, impairing viral transit through the endolysosomal pathway. Reduced proteolytic function within lysosomes augments antigen processing for presentation on MHC complexes and increases CTLA4 expression on Tregs.
(C) Antagonism of IL-6 signaling pathway and of other cytokine-/chemokine-associated targets has been proposed to control COVID-19 CRS. These include secreted factors like GM-CSF that contribute to the recruitment of inflammatory monocytes and macrophages.
(D) Several potential sources of SARS-CoV-2 neutralizing antibodies are currently under investigation, including monoclonal antibodies, polyclonal antibodies, and convalescent plasma from recovered COVID-19 patients.
GM-CSF, granulocyte-macrophage colony-stimulating factor; CQ, chloroquine; HCQ, hydroxychloroquine; RdRp, RNA-dependent RNA polymerase.
Antiviral Clinical Trials
A large number of clinical trials using experimental antiviral drugs are currently underway. A small proportion of them are aimed at repurposing existing antivirals, including arbidol (umifenovir), a broad-spectrum antiviral that blocks viral fusion; lopinavir/ritonavir (LPV/r), a combination of anti-HIV protease inhibitors; favipiravir, an RdRp inhibitor used to treat severe influenza infections (Hayden and Shindo, 2019); and remdesivir (Figure 6A). Chen et al. conducted a multicenter, randomized priority trial on 240 patients with confirmed COVID-19 infection to test favipiravir or arbidol (Chen et al., 2020b). Favipiravir was suggested to significantly improve symptom relief. However, the interpretation of this study is limited by a short clinical recovery window of 7 days, only 100 of 236 patients with confirmed COVID-19, and the lack of a control group.
LPV/r has previously shown efficacy in treating SARS-Cov-1 (Chu et al., 2004), prompting an early SARS-Cov-2 clinical trial (Li et al., 2020c). 44 patients were enrolled in a trial investigating the efficacy and safety of LPV/r (n = 21 patients), arbidol (n = 16), or control (n = 7) as treatment for mild to moderate COVID-19. At day 14 of treatment, 76.2%, 62.4%, and 71.4% of patients had a positive to negative conversion in the LPV/r, arbidol, and control groups, respectively, with no statistical significance between groups. A randomized controlled trial (RCT) with 200 severe COVID-19 patients did not observe a significant benefit of LPV/r either (Cao et al., 2020a). However, a study that looked at the impact of earlier administration of LPV/r treatment showed that when treatment of LPV/r was started within 10 days of symptom onset, a shorter duration of virus shedding was observed. Thus, timing of LPV/r administration may be critical to its efficacy (Yan et al., 2020a).
In a multicenter clinical study assessing the compassionate use of remdesivir in severe COVID-19 patients, 53 patients across several countries were treated with remdesivir for 10 days (Grein et al., 2020). 68% of the 53 patients who received remdesivir showed clinical improvement assessed through improved oxygen support or extubations. Without a proper control group, limited conclusions can be drawn with regards to the efficacy of remdesivir from this study. The measured 68% clinical improvement may be in line with average clinical improvement across patients treated with standard of care (Li et al., 2020c). A small RCT in China with 237 severe COVID-19 patients randomized 2:1 to remdesivir versus placebo demonstrated no significant benefit in time to clinical improvement (Wang et al., 2020g). Almost simultaneously, preliminary results from a larger National Institute of Allergy and Infectious Diseases (NIAID) RCT with more than 1,000 patients were announced with remdesevir to be associated with quicker time to recovery: 11 days compared with 15 days (Ledford, 2020). A non-significant benefit in mortality was also noted, and the trial was stopped early to allow access to remdesivir in the placebo arm. Complete safety data and full publication are awaited, but this study offers encouraging results and has resulted in an FDA emergency use authorization for remdesivir in hospitalized COVID-19 patients.
Therapeutic Immunomodulation for COVID-19 Treatment
Chloroquine: Modes of Action and Immunological Impact
Chloroquine (CQ) and its derivative hydroxychloroquine (HCQ) have gained traction as possible therapeutics for COVID-19. Both drugs are used as antimalarial agents and as immunomodulatory therapies for rheumatologic diseases. However, the application of CQ and HCQ to COVID-19 stems from their past use as antivirals (Savarino et al., 2003), including for SARS-CoV-1 (Keyaerts et al., 2004, Vincent et al., 2005). CQ and HCQ interfere with lysosomal activity and have been reported to have immunomodulatory effects. CQ augments antigen processing for MHC class I and II presentation, directly inhibits endosomal TLR7 and TLR9, and enhances the activity of regulatory T cells (Garulli et al., 2008, Lo et al., 2015, Schrezenmeier and Dörner, 2020, Thomé et al., 2013a, Thomé et al., 2013b). Early studies involving in vitro infection of host cells with SARS-CoV-2 demonstrated that both CQ and HCQ significantly impact endosomal maturation, resulting in increased sequestration of virion particles within endolysosomes. However, there has been conflicting evidence whether CQ is more potent than HCQ in reducing viral load (Liu et al., 2020d, Wang et al., 2020b, Yao et al., 2020a). Notably, one group reported that treatment of infected cells with HCQ before and during infection significantly reduced viral load, suggesting that combined prophylactic and therapeutic HCQ use yields maximum efficacy (Clementi et al., 2020). To better understand host immune responses to treatment, one group compared bulk transcriptomic changes in primary PBMCs treated with HCQ for 24 h to PBMCs from confirmed SARS-CoV-2 positive patients and controls, followed by a comparison of HCQ-treated primary macrophages to BAL and postmortem lung biopsies from COVID-19 patients (Corley et al., 2020). Across all comparisons, there was minimal overlap between host differential gene expression and genes altered by in vitro HCQ treatment. Thus, the potential mechanistic action of HCQ in the context of SARS-CoV-2 remains poorly defined.
Evaluation of HCQ Efficacy in Clinical Trials
Despite the apparent widespread use of HCQ and CQ to treat COVID-19 (Figure 6B), few controlled clinical trials have been performed so far and thus the potential benefits of these drugs for COVID-19 remains controversial. One of the earliest trials (2020-000890-25) was a single-arm, open-label trial of 600 mg daily HCQ in 20 COVID-19 patients. They reported that HCQ alone, or in combination with the antibiotic azithromycin (AZ), reduced viral load by day 6 (Gautret et al., 2020a). A follow-up trial in 80 patients treated with HCQ and AZ reported that 93% of patients had a negative PCR result on day 8 of treatment, and 81.3% were discharged within 10 days of treatment. However, it is important to note that both trials had no control arms (Gautret et al., 2020b). Rigorous statistical analyses by others that accounted for the patients excluded from the original analysis found limited evidence for HCQ monotherapy (Hulme et al., 2020, Lover, 2020). A double-blind RCT assessed HCQ monotherapy in the treatment of mild COVID-19 (ChiCTR2000029559) (Chen et al., 2020i). A total of 62 patients were enrolled; the treatment arm received 400 mg HCQ daily over 5 days. By day 6, patients who received HCQ had clinical resolution on average 1 day earlier than controls; no patients progressed to severe disease compared to four patients in the control arm. In a smaller RCT that treated 30 patients with mild COVID-19 (NCT04261517) with 400 mg HCQ for 7 days, there were no significant differences in the number of patients with negative PCR results on day 7 (all but one positive), median duration of hospitalization, time to fever resolution, or progression of disease on chest computed tomography (CT) (Chen et al., 2020d). The largest RCT to date enrolled 150 patients with mild COVID-19 across 16 centers in an open-label trial of HCQ and standard of care (ChiCTR2000029868). There were no significant differences between groups in conversion to negative SARS-CoV-2 RT-PCR result on day 28 or rate of symptom resolution; there were significantly more adverse events in the HCQ arm, though largely non-serious; they reported some evidence for faster normalization of C-reactive protein in the patients who received HCQ plus standard of care, but this finding was not significant (Tang et al., 2020b). A meta-analysis including most of the studies described here found no clinical benefits to patients receiving standard of care plus an HCQ regimen (Shamshirian et al., 2020).
Two studies have assessed HCQ efficacy in severe COVID-19. In a prospective study of 11 patients who had received 600 mg HCQ over 10 days with AZ on days 1–5, there were several patients with worsening clinical status and one death; 8 of 10 patients had a positive PCR result on day 10 (Molina et al., 2020). An ongoing double-blind RCT of patients with severe COVID-19 (NCT04323527) randomized 81 patients into high-dose HCQ (600 mg 2× per day for 10 days) or low-dose (450 mg/day for 5 days) treatment groups (Borba et al., 2020). Recruitment into the high-dose arm was halted prematurely due to poor safety outcomes. There was no significant difference in negative PCR results on day 4 or need for mechanical ventilation on day 6. Taken together, the clinical trials performed thus far to evaluate the efficacy of HCQ ± AZ for COVID-19 have not demonstrated clear evidence of clinical benefit in patients with severe disease. A search of ClinicalTrials.gov on April 27, 2020 found 140 clinical trials investigating HCQ. This number is rapidly growing, indicating the heightened interest in this therapeutic and pressing need for evidence-based recommendations.
Corticosteroids for COVID-19 Therapy
Because of their anti-inflammatory activity, corticosteroids (CSs) are an adjuvant therapy for ARDS and cytokine storm. However, the broad immunosuppression mediated by CS does raise the possibility that treatment could interfere with the development of a proper immune response against the virus. A meta-analysis of 5,270 patients with MERS-CoV, SARS-CoV-1, or SARS-CoV-2 infection found that CS treatment was associated with higher mortality rate (Yang et al., 2020c). A more recent meta-analysis of only SARS-CoV-2 infection assessed 2,636 patients and found no mortality difference associated with CS treatment, including in a subset of patients with ARDS (Gangopadhyay et al., 2020). Other studies have reported associations with delayed viral clearance and increased complications in SARS and MERS patients (Sanders et al., 2020). In fact, the interim guidelines updated by the WHO on March 13, 2020 advise against giving systemic corticosteroids for COVID-19 (World Health Organization, 2020a). Yet, new data from COVID-19 are conflicting.
One group reported no significant difference in time to viral clearance between patients who received methylprednisolone orally (mild disease) or intravenously (i.v.) (severe) and those who did not (Fang et al., 2020). Retrospective studies from groups in China report that patients who were transferred to the ICU were less likely to have received CSs (Wang et al., 2020b) and that patients with ARDS who received methylprednisolone had reduced mortality risk (Wu et al., 2020a). In contrast, another retrospective analysis found that patients who received CSs were more likely to have either been admitted to the ICU or perished, although the CS-treated group also had significantly more comorbidities (Wang et al., 2020c). A smaller observational study of 31 patients found no association between corticosteroid treatment and time to viral clearance, length of hospital stay, or symptom duration (Zha et al., 2020). A larger study of adjuvant CSs in 244 patients with critical COVID-19 found no association with 28-day mortality; subgroup analysis of patients with ARDS found no association between treatment with CSs and clinical outcomes (Lu et al., 2020b). They also found that increased dosage was significantly associated with increased mortality risk. A retrospective review of 46 patients, of whom 26 received i.v. methylprednisolone, found that early, low-dose administration significantly improved SpO2 and chest CT, time to fever resolution, and time on supplemental oxygen therapy (Wang et al., 2020h). Others have published perspectives in support of early (Lee et al., 2020) and short-term, low-dose administration (Shang et al., 2020) based on anecdotal evidence but not clinical trials. Most of the current data on CS use in COVID-19 are from observational studies and support either modest clinical benefit or no meaningful effects. Larger RCTs are necessary to understand the risks and benefits of CSs for these patients; there are 22 trials evaluating various corticosteroids registered on ClinicalTrials.gov as of April 27, 2020.
Cytokine-Directed Therapy in COVID-19
Recombinant IFN as an Antiviral Treatment
One of the first defenses of the human body against RNA viruses like SARS-CoV-2 is the release of types I and III IFNs. It is important to note that type I IFN (IFNα/β) receptors are ubiquitously expressed, so IFNα/β signaling can result in not only antiviral effects, but also the activation of immune cells that potentially exacerbate pathogenesis. In contrast, type III IFN (also known as IFNλ) signals mainly in epithelial cells, as well as in a restricted pool of immune cells. Because type III IFNs have immunomodulatory functions, subsequent signaling could induce a potent antiviral effect without enhancing pathogenic inflammation (Andreakos et al., 2017, Prokunina-Olsson et al., 2020).
Recently, there has been a growing interest in the potential therapeutic impact of modulating the IFN response to disable COVID-19 pathogenesis. Before the current pandemic, groups have studied the role of IFNs in other betacoronavirus infections. One study of 40 patients with SARS-CoV-1 infection described unresolved elevated type I IFNs and ISGs in those with poor outcomes (Cameron et al., 2007). Others report that exogenous type I IFN does not improve outcomes when given with ribavirin in patients with MERS-CoV infection (Arabi et al., 2020), suggesting that the role of IFN as a therapeutic or prophylactic option may be strain or species specific (Sheahan et al., 2020). Interestingly, a recent study by Mount Sinai virology groups revealed that type I IFN signaling is impaired in the early response to SARS-CoV-2; in vitro, SARS-CoV-2 may be more susceptible to type I IFN than SARS-CoV-1 is (Blanco-Melo et al., 2020). Based on additional evidence that IFN responses to betacoronaviruses are altered as compared to other respiratory viruses (Blanco-Melo et al., 2020, Channappanavar et al., 2016, Okabayashi et al., 2006), trials of IFN-I/III administration have been initiated (NCT04343976, NCT04331899).
Cytokine Blockade
Hyperinflammatory responses and elevated levels of inflammatory cytokines, including IL-6, -8, and -10, have been shown to correlate with COVID-19 severity (Chen et al., 2020h, Diao et al., 2020, Gong et al., 2020, Moore and June, 2020, Wan et al., 2020a, Xu et al., 2020b). The drivers of this cytokine storm remain to be established, but they are likely triggered initially by a combination of viral PAMPs and host danger signals. The heterogeneous response between patients suggests other factors are involved, possibly including the SARS-CoV-2 receptor, ACE2 (Hirano and Murakami, 2020).
Several studies have begun to report the cellular programs that may contribute to the cytokine storm detected in COVID-19 patients. One group reported that in the context of generalized lymphopenia, certain subsets of CD4 T cells that express GM-CSF and IL-6 are more abundant in severe COVID-19 patients than in COVID-19 patients who do not require intensive care (Zhou et al., 2020b). Reports that other major proinflammatory cytokines (TNF-α, IFN-ɣ, IL-2) and chemokines (CCL2, CCL3, CCL4) are elevated underscore a potentially pathogenic TH1/2 program in COVID-19 (Diao et al., 2020, Giamarellos-Bourboulis et al., 2020). Histological and single-cell analyses identified monocytes and macrophages as other potent sources of inflammatory cytokines in COVID-19 cytokine storm (Chen et al., 2020h, Giamarellos-Bourboulis et al., 2020, Law et al., 2005, Moore and June, 2020, Zhou et al., 2020b). Studies of other betacoronavirus infections, including SARS-CoV-1 and MERS-CoV, have also identified similar hyperactivation of monocytes, macrophages, and DCs as a driver of cytokine-mediated immunopathology in humans (Cheung et al., 2005, Chien et al., 2006, Huang et al., 2020c, Konig et al., 2020, Wang et al., 2005, Wong et al., 2004, Xu et al., 2020b, Zhou et al., 2020b).
Following preliminary reports of IL-6 as a critical cytokine in COVID-19-associated CRS, monoclonal antibodies that target the IL-6 signaling pathway have been proposed as therapeutic candidates (Moore and June, 2020) (Figure 6C). The commercial anti-IL-6R antibodies tocilizumab (Actemra) and sarilumab (Kevzara) and the anti-IL-6 antibody siltuximab (Sylvant) are now being tested for efficacy in managing COVID-19 CRS and pneumonia in 13 ongoing clinical trials (Table 2 ). To date, only one group has reported preliminary results from a cohort of 20 COVID-19 patients treated with a single administration of tocilizumab (400 mg, i.v.), along with lopinavir, methylprednisolone, and oxygen therapy (ChiCTR2000029765) (Xu et al., 2020b). The single observation study found recuperated lymphocyte counts in 10 of 19 patients and resolution of lung opacities in 19 of 20 patients on chest CT; 19 of 20 patients were discharged. All patients experienced an improvement in symptoms, and no subsequent pulmonary infections were reported. A second report described an association between use of tocilizumab and reduced likelihood of ICU admission and mechanical ventilation. Still, in 30 declining patients with severe COVID-19 pneumonia, this retrospective study did not report significant improvement in mortality on weighted analysis (Roumier et al., 2020). Nevertheless, these studies are encouraging, but like other treatment approaches, larger RCTs are needed.
Table 2.
Clinical Trials Evaluating the Efficacy of IL-6/IL-6R Blockade Therapy
| Clinical Trial | Intervention |
|---|---|
|
NCT04331795 (COVIDOSE) NCT04320615 (COVACTA) NCT04332913 (TOSCA) NCT04317092 (TOCOVID-19) NCT04335071 (CORON-ACT) NCT04315480 ChiCTR2000029765 |
tocilizumab |
| NCT04315298 | sarilumab |
| NCT04310228 | tocilizumab favipiravir |
| NCT04306705 (TACOS) | tocilizumab continuous renal replacement therapy standard of care |
| NCT04332094 (TOCOVID) | tocilizumab azithromycin hydroxychloroquine |
| NCT04341870 (CORIMUNO-VIRO) | sarilumab azithromycin hydroxychloroquine |
| NCT0433z638 (COV-AID) | tocilizumab siltuximab anakinra standard of care |
In addition to the IL-6 signaling pathway, other cytokine- and chemokine-associated elements, including IL-1R, GM-CSF, and the chemokine receptor CCR5, have been proposed as potential targets for blockade to manage COVID-19 CRS (Figure 6C). Finally, complement activation was shown to be overactivated in lungs of COVID-19 patients. Although results from the randomized trial are not yet published, anti-C5a monoclonal antibody therapy showed benefits in two critically ill COVID-19 patients (Gao et al., 2020d).
Neutralizing Antibodies and Convalescent Plasma Therapy for COVID-19
While vaccines are being developed to educate a person’s immune system to make their own nAbs against SARS-CoV-2, there is interest in using adoptive transfer of nAbs as a therapeutic approach (Figure 6D). This strategy has already proven to be effective against SARS-CoV-1 (Cao et al., 2010, Ho et al., 2005, ter Meulen et al., 2004, Park et al., 2020, Sui et al., 2004, Zhu et al., 2007) and MERS-CoV (Forni et al., 2015, Jia et al., 2019, Ying et al., 2015). In the case of SARS-CoV-2, these efforts are primarily centered on identifying nAbs made during natural infections or generating nAbs through animal vaccination approaches.
nAbs Derived from COVID-19 Patients
Patients who have recovered from SARS-CoV-2 infection are one potential source of nAbs (Chen et al., 2020a, Ju et al., 2020, Walls et al., 2020, Wölfel et al., 2020, Yuan et al., 2020). In an effort to obtain these nAbs, scientists sorted RBD-specific memory B cells and cloned their heavy and light variable regions to express recombinant forms of the corresponding antibodies (Chen et al., 2020a, Ju et al., 2020). Four of the antibodies produced in these studies (31B5, 32D4, P2C-2F6, P2C-1F11) showed high neutralizing potential in vitro, and all inhibited ACE2-RBD binding. Successful antibody-mediated neutralization of SARS-CoV-2 seemed to be dependent on the inhibition of ACE2/RBD binding. However, Chen et al. showed that nearly all antibodies derived from serum of 26 recovered patients bound to S1 and RBD, with only three actually inhibiting ACE2/RBD binding (Chen et al., 2020a). Of note, a SARS-CoV-1-derived neutralizing antibody (47D11) (Wang et al., 2020a) and a single-chain antibody against SARS-CoV-2 (n3130) (Wu et al., 2020c) have also been shown to neutralize SARS-CoV-2 without inhibiting ACE2/RBD binding. Thus, blocking this interaction may not be a prerequisite for an effective SARS-CoV-2 nAb. The generation of a hybridoma producing a monoclonal nAb against SARS-CoV-2 provides the potential for a therapeutic Ab that can be directly administered to patients to block ongoing infection and potentially even as a prophylactic (Figure 6D).
SARS-CoV-1 nAbs Also Neutralize SARS-CoV-2
SARS-CoV-1 and SARS-CoV-2 consensus sequences share about 80% identity (Tai et al., 2020). Thus, a wide range of SARS-CoV-1 nAbs have been tested for crossreactivity with SARS-CoV-2, as they could help speed up the development of potential COVID-19 treatments. In a recent study, antibodies were isolated from the memory B cells of an individual who recovered from SARS-CoV-1 infection. While 8 out of 25 isolated antibodies could bind SARS-CoV-2 S protein, one of them (s309; see Table 3 ) also neutralizes SARS-CoV-2 (Pinto et al., 2020). The combination of s309 with a weakly neutralizing antibody that could bind another RBD epitope led to enhanced neutralization potency. In addition, CR3022 (Table 3) was found to bind SARS-CoV-2 RBD (Tian et al., 2020b), but this antibody did not neutralize SARS-CoV-2 (Yuan et al., 2020). Computational simulations identified three amino acids that could be modified on CR3022 to enhance its binding affinity with SARS-CoV-2 RBD (Giron et al., 2020), potentially augmenting its neutralization potential.
Table 3.
Strategies to Isolate SARS-CoV-2 Neutralizing Antibodies
| Ab Source | Clone | Target | Type of Antibody | Neutralization | Inhibition of ACE2/RBD Binding | Reference |
|---|---|---|---|---|---|---|
| Derived from COVID-19 patients | 31B532D4 | RBD | human monoclonal | yes | yes | Chen et al., 2020a |
| P2C-2F6P2C-1F11 | RBD | human monoclonal | yes | yes | Ju et al., 2020 | |
| Derived from SARS-CoV-1 patients | CR3022 | RBD | human monoclonal | no | no | Tian et al., 2020b, Yuan et al., 2020, Giron et al., 2020 |
| S309 | RBD | human monoclonal | yes | no | Pinto et al., 2020 | |
| Derived from SARS-CoV-1 or MERS-CoV-1 animal models | R325R302R007 | S1 | rabbit monoclonal | yes | no | Sun et al., 2020 |
| 47D11 | S1 | recombinant human monoclonal (derived from hybridomas of immunized transgenic H2L2 mice) | yes | no | Wang et al., 2020a | |
| VHH-72-Fc | S | Fc-fusion derived from camelids VHH | yes | yes | Wrapp et al., 2020 | |
| S | polyclonal mouse antibodies | yes | N/A | Walls et al., 2020, Yuan et al., 2020 | ||
| Other | ACE2-Fc | RBD | ACE2-Fc fusion | yes | N/A | Lei et al., 2020a, Li et al., 2020d |
| RBD-Fc | ACE2 | RBD-Fc fusion | yes | N/A | Li et al., 2020d | |
| N3130 | S1 | human monoclonal single domain antibody isolated by phage display | yes | no | Wu et al., 2020c | |
| IVIg | N/A | polyclonal human IVIg | N/A | N/A | Díez et al., 2020, Shao et al., 2020 | |
| F(ab′)2 | RBD | horse polyclonal | yes | N/A | Pan et al., 2020 |
N/A, not assessed.
nAbs Derived from Animals
Animal models represent another tool to generate nAbs against SARS-CoV-2 (Table 3). In one study, the authors developed a protocol to synthetize human nanobodies, smaller antibodies that only contain a variable heavy (VH) chain as first described in camelids (Wu et al., 2020c) (Figure 6D). Another antibody isolated from camelids immunized with SARS-CoV-1 and MERS-CoV S proteins then fused to a human Fc fragment showed neutralization potential against SARS-CoV-2 (VHH-72-Fc) (Wrapp et al., 2020). Genetically modified mice with humanized antibody genes can also be used to generate therapeutic monoclonal antibodies, as successfully experimented against Ebola virus (Levine, 2019). Similar studies are now focused on the use of SARS-CoV-2 or derivatives to generate highly effective nAb in animal models, which can be directly given to infected patients, and efforts are already underway with estimates of clinical trials of pooled antibody cocktails beginning in early summer by Regeneron. Finally, another approach to nAb development is to fuse ACE2 protein and the Fc part of antibodies, as they would bind RBD and potentially be crossreactive among other CoVs (Figure 6D). Indeed, an ACE2-Fc (Lei et al., 2020a) and an RBD-Fc (Li et al., 2020d) have been shown to neutralize both SARS-CoV-1 and SARS-CoV-2 in vitro.
Convalescent Plasma Therapy
Although recombinant nAbs could provide an effective treatment, they will require a significant time investment to develop, test, and bring production to scale before becoming widely available to patients. A faster strategy consists of transferring convalescent plasma (CP) from previously infected individuals that have developed high titer nAbs that target SARS-CoV-2 (Figure 6D). Despite the current lack of appropriately controlled trials, CP therapy has been previously used and shown to be beneficial in several infectious diseases, such as the 1918 influenza pandemic (Luke et al., 2006), H1N1 influenza (Hung et al., 2011), and SARS-CoV-1 (Arabi et al., 2016). Thanks to the development of serological tests (Amanat et al., 2020, Cai et al., 2020, Xiang et al., 2020b, Zhang et al., 2020d), recovered COVID-19 patients can be screened to select plasma with high antibody titers.
Some studies and case reports on CP therapy for COVID-19 have evaluated the safety and the potential effectiveness of CP therapy in patients with severe disease (Ahn et al., 2020, Duan et al., 2020, Pei et al., 2020, Shen et al., 2020, Zhang et al., 2020b) (Table 4 ). These studies were neither controlled nor randomized, but they suggest that CP therapy is safe and can have a beneficial effect on the clinical course of disease. Further controlled trials are needed to determine the optimal timing and indication for CP therapy. CP therapy has also been proposed for prophylactic use in at-risk individuals, such as those with underlying health conditions or health care workers exposed to COVID-19 patients. The FDA has approved the use of CP to treat critically ill patients (Tanne, 2020). Determining when to administer the CP is also of great importance, as a study in SARS-CoV-1 patients showed that CP was much more efficient when given to patients before day 14 day of illness (Cheng et al., 2005b), as previously shown in influenza (Luke et al., 2006). This study also showed that CP therapy was more efficient in PCR-positive, seronegative patients. The amount of plasma and number of transfusions needed requires further investigation (Table 4).
Table 4.
Clinical Studies of Convalescent Plasma Therapy in COVID-19 Patients
| Patient Characteristics | Start of CP Therapy | Results | Reference |
|---|---|---|---|
| 5 severe patients (30–70 yo) | between 10 and 22 days after hospital admission | body temperature normalized within 3 days in 4 of 5 patients | Shen et al., 2020 |
| clinical improvement | |||
| viral loads became negative within 12 days of the transfusion | |||
| nAb titers increased | |||
| 10 severe patients (34–78 yo) | median 16.5 dpo | disappearance of clinical symptoms after 3d | Duan et al., 2020 |
| chest CT improved | |||
| elevation of lymphocyte counts in patients with lymphocytopenia. | |||
| increase in SaO2 in all patients | |||
| resolution of SARS-CoV-2 viremia in 7 patients | |||
| increase in neutralizing antibody titers in 5 patients | |||
| 4 critical patients (31–73 yo) | at degradation of symptoms, between 11 and 19 days after hospital admission |
clinical improvement | Zhang et al., 2020b |
| reduced viral load | |||
| chest CT improved | |||
| 1 moderate patient, 2 critical patients | 12 dpo, 27 dpo | viral detection negative 4 days after CP | Pei et al., 2020 |
| clinical improvement of 2 patients | |||
| 2 severe patients (67 and 71 yo) | 7 dpo or 22 dpo | clinical improvement | Ahn et al., 2020 |
| reduced viral load | |||
| chest CT improved |
yo, years old; dpo, days post onset of symptoms.
Overall, CP therapy seems to be associated with improved outcomes and appears to be safe, but RCTs are needed to confirm this. Several clinical trials are currently in progress worldwide (Belhadi et al., 2020).
Vaccine Development
The devastating effects of the pandemic spread of SARS-CoV-2 in a globally naive population has resulted in unprecedented efforts to rapidly develop, test, and disseminate a vaccine to protect against COVID-19 or to mitigate the effects of SARS-CoV-2 infection. Although vaccination has a long and successful history as an effective global health strategy, there are currently no approved vaccines to protect humans against CoVs (André, 2003). Previous work after the SARS-CoV-1 and MERS-CoV epidemics has provided a foundation on which many current efforts are currently building upon, including the importance of the S protein as a potential vaccine. Diverse vaccine platforms and preclinical animal models have been adapted to SARS-CoV-2, facilitating fast-moving and robust progress in creating and testing SARS-CoV-2 vaccine candidates. A number of vaccine candidates are already being tested in clinical trials, and more are continuing to progress toward clinical testing.
The S Protein as a Vaccine Target
Since SARS-CoV-1 first emerged, the S protein has been favored as the most promising target for vaccine development to protect against CoV infection. This particular viral protein has important roles in viral entry and in stimulating the immune response during natural infection and in vaccination studies of both SARS-CoV-1 and MERS-CoV (Du et al., 2009, Song et al., 2019, Zhou et al., 2018), which has also been confirmed for SARS-CoV-2 (Walls et al., 2020). The S protein has been found to induce robust and protective humoral and cellular immunity, including the development of nAbs and T cell-mediated immunity (Du et al., 2009). In animal models, correlates of protection against SARS-CoV-1 infection appear to be induction of nAbs against the S protein, although antibodies to other proteins have been detected, such as those against nucleoprotein (N) and ORF3a (Qiu et al., 2005, Sui et al., 2005). nAbs are also believed to protect against infection by blocking receptor binding and viral entry, which has been shown with pseudovirus-based neutralization assays (Ni et al., 2020, Nie et al., 2020a). Studies of SARS-CoV-1 indicate that T cell response against the S protein correlates with nAb titers and dominated the T cell response after natural infection, which also induced T cells active against the membrane (M) and N proteins, that memory T cell responses can persist even 11 years after infection, and that memory CD8+ T cells can protect mice from lethal challenge in the absence of memory CD4+ T cells and memory B cells (Li et al., 2008, Ng et al., 2016). RBD-specific antiviral T cell responses have also been detected in people who have recovered from COVID-19, further validating its promise as a vaccine target (Braun et al., 2020, Ni et al., 2020).
Epitope Mapping
Although the antibodies targeting the RBD of the S protein have greater potential for providing cross-protective immunity, other fragments of the S protein and additional viral proteins have been investigated as target epitopes, especially for T cells. Researchers have taken advantage of the genetic similarity between SARS-CoV-2 and SARS-CoV-1 and MERS-CoV and bioinformatics approaches to rapidly identify potential B and T cell epitopes in the S and other proteins, with many studies providing data regarding antigen presentation and antibody-binding properties and one study looking into the predicted evolution of epitopes (Ahmed et al., 2020, Baruah and Bose, 2020, Bhattacharya et al., 2020, Fast et al., 2020, Grifoni et al., 2020, Lon et al., 2020, Zheng and Song, 2020). While the S protein has been found to be the most immunodominant protein in SARS-CoV-2, the M and N proteins also contain B and T cell epitopes, including some with high conservation with SARS-CoV-1 epitopes (Grifoni et al., 2020).
Vaccine Pipeline
For SARS-CoV-1 and MERS-CoV, animal studies and phase I clinical trials of potential vaccines targeting the S protein had encouraging results, with evidence of nAb induction and induction of cellular immunity (Lin et al., 2007, Martin et al., 2008, Modjarrad et al., 2019). These findings are being translated into SARS-CoV-2 vaccine development efforts, hastening the progress drastically. The WHO provided a report in April that reported 63 vaccine candidates in preclinical testing and three in clinical testing (World Health Organization, 2020b). A recent search on May 1, 2020 on ClinicalTrials.gov revealed 10 registered vaccine candidates (Table 5 ). The University of Pittsburgh is also looking to move their microneedle array vaccine candidate containing a codon-optimized S1 subunit protein into clinical trials (Kim et al., 2020). Sanofi and GlaxoSmithKline (GSK) have recently reported their intent to collaborate and bring together Sanofi’s baculovirus expression system, which is used to produce the influenza virus vaccine, Flublok, to create an S protein vaccine adjuvanted with GSK’s AS03. Sinovac Biotech will also enter testing in a clinical trial in China after it was found to protect rhesus macaques from viral challenge without signs of detectable immunopathology (Gao et al., 2020c). Although some of these vaccine candidates are based on platforms that have been used or tested for other purposes, there remain questions regarding their safety and immunogenicity, including the longevity of any induced responses, that will require continual evaluation.
Table 5.
Vaccine Candidates Currently Registered for Clinical Trials
| Candidate | Design | Developer | Similar Strategy | ClinicalTrials.gov Identifier |
|---|---|---|---|---|
| mRNA-1273 | LNP-encapsulated mRNA for full-length S protein | ModernaTX | CMV (John et al., 2018), ZKV (Pardi et al., 2017) | NCT04283461 |
| BNT162a1, b1, b2, c2 | LNP-encapsulated mRNA vaccines with different formats of RNA and targets, two for larger S sequence and two for optimized RBD | BioNTech SE and Pfizer | NCT04368728 | |
| INO-4800 | DNA vaccine for full-length S protein | Inovio Pharmaceuticals | MERS-CoV (Modjarrad et al., 2019), HPV (Trimble et al., 2015) | NCT04336410 |
| Ad5-nCoV | adenovirus type 5 encoding full-length S protein | CanSino Biologics | EBV (Zhu et al., 2015, Zhu et al., 2017) |
NCT04313127 (phase I) NCT04341389 (phase II) |
| ChAdOx1 nCoV-19 | adenovirus encoding full-length S protein | University of Oxford | MERS-CoV (Alharbi et al., 2017), IAV (Antrobus et al., 2014) | NCT04324606 |
| COVID-19 LV-SMENP-DC | dendritic cells infected with lentivirus expressing SMENP minigenes to express COVID-19 antigens, together with activated CTLs | Shenzhen Geno-Immune Medical Institute | NCT04276896 | |
| COVID-19 aAPCs | aAPCs infected with lentivirus expressing minigenes to express COVID-19 antigens | Shenzhen Geno-Immune Medical Institute | NCT04299724 | |
| bacTRL-Spike-1 | live bacteria delivering plasmid encoding S protein | Symvivo Corporation | therapeutics reviewed (Charbonneau et al., 2020) | NCT04334980 |
| PiCoVacc | inactivated SARS-CoV-2 vaccine | Sinovac Biotech | HAV, IAV, IBV, poliovirus, rabies virus | NCT04352608 |
| SARS-CoV-2 rS | spike protein nanoparticle vaccine with or without Matrix-M adjuvant | Novavax | NCT04368988 |
aAPCs, artificial antigen-presenting cells; CMV, cytomegalovirus; EBV, Ebola virus; HAV, hepatitis A virus; HPV, human papillomavirus; IAV, influenza A virus; IBV, influenza B virus; LPN, lipid nanoparticle; ZKV, Zika virus.
Challenges
Although the development of a vaccine to protect against SARS-CoV-2 infection has progressed at an unprecedented rate and produced an impressive volume of candidates for testing, many challenges lie ahead. The prior knowledge gained after SARS-CoV-1 was first discovered in 2003, and the subsequent emergence of MERS-CoV in 2012 provided a significant jumpstart, but the progress of SARS-CoV-2 vaccine development has already far outstripped the point of the blueprint created before COVID-19 became a pandemic. While a variety of platforms are simultaneously being innovated or adapted, they each have strengths and limitations, many of which relate to the delicate balance between safety and immunogenicity. Many shortcuts have been taken and will continue to be taken due to the urgency of the ongoing COVID-19 pandemic, but significant concerns need to be addressed. One such concern involves the accumulating data supporting the initial assessment that COVID-19 is disproportionately severe in older adults. In conjunction with the large body of work related to immune senescence, these findings indicate that vaccine design should take into consideration the impact of aging on vaccine efficacy (Nikolich-Žugich, 2018). Furthermore, questions remain regarding the possibility of antibody-dependent enhancement of COVID-19, with in vitro experiments, animal studies, and two studies of COVID-19 patients supporting this possibility (Cao, 2020, Tetro, 2020, Zhang et al., 2020a, Zhao et al., 2020a). Assuming vaccine candidates that can safely induce protective immune responses are identified, additional major hurdles will be the production and dissemination of a vaccine. For some types of vaccines, large-scale production will not be as much of an issue, and infrastructure already in place to produce current Good Manufacturing Practice (cGMP)-quality biologics can be repurposed, but this will only be applicable to a subset of the candidates (Thanh Le et al., 2020). In order to address the urgent need and stem the COVID-19 pandemic, regulatory agencies need to continue to support rapid testing and progression of vaccine candidates, companies need to disseminate important findings directly and openly, and researchers need to investigate correlates of protection using in-depth immune monitoring of patients with a broad range of clinical presentations and clinical trial participants. The newly announced Accelerating COVID-19 Therapeutic Interventions and Vaccines (ACTIV) is designed to bring together numerous governmental and industry entities to help address this need.
Concluding Remarks
The rapid spread of SARS-CoV-2 and the unprecedented nature of COVID-19 has demanded an urgency in both basic science and clinical research, and the scientific community has met that call with remarkable productivity. Within months, there has been a significant generation of scientific knowledge that has shed some light on the immunology of SARS-CoV-2 infections. Studies of past coronavirus outbreaks involving SARS-CoV-1 and MERS-CoV have provided a foundation for our understanding. The pathology of severe cases of COVID-19 does indeed resemble certain immunopathologies seen in SARS-CoV-1 and MERS-CoV infections, like CRS.
However, in many other ways, immune responses to SARS-CoV-2 are distinct from those seen with other coronavirus infections. The emerging epidemiological observation that significant proportions of individuals are asymptomatic despite infection not only reflects our current understanding that SARS-CoV-2 has a longer incubation period and higher rate of transmission than other coronaviruses, but also speaks to significant differences in the host immune response. Therefore, it is imperative that immune responses against SARS-CoV-2 and mechanisms of hyperinflammation-driven pathology are further elucidated to better define therapeutic strategies for COVID-19. Here, we reviewed the recent literature and highlighted hypotheses that interrogate mechanisms for viral escape from innate sensing, for hyperinflammation associated with CRS and inflammatory myeloid subpopulations, for lymphopenia marked by T cell and NK cell dysfunction, and for correlates of protection and their duration, among others. Still, additional studies are needed to address how these immune differences across patients or between different types of coronavirus infections dictate who succumbs to disease and who remains asymptomatic. Existing studies of SARS-CoV-1 and MERS-CoV and ongoing studies of SARS-CoV-2 will likely provide a robust framework to fulfill that unmet need.
Consortia
The members of the Sinai Immunology Review Project are Manasi Agrawal, Mark Aleynick, Meriem Belabed, Graham Britton, Matthew Brown, Maria Casanova-Acebes, Jovani Catalan, Monica Centa, Andrew Charap, Andrew Chan, Steven T. Chen, Jonathan Chung, Cansu Cimen Bozkus, Evan Cody, Francesca Cossarini, Erica Dalla, Nicolas Fernandez, John Grout, Conor Gruber, Dan Fu Ruan, Pauline Hamon, Samarth Hegde, Etienne Humblin, Divya Jha, Maria Kuksin, Julia Kodysh, Andrew Leader, Rachel Levantovsky, Matthew Lin, Katherine Lindblad, Daniel Lozano-Ojalvo, Gabrielle Lubitz, Assaf Magen, Zafar Mahmood, Louise Malle, Gustavo Martinez-Delgado, Jaime Mateus-Tique, Elliot Meritt, Chang Moon, Alvaro Moreira, Justine Noel, Tim O’Donnell, Miyo Ota, Matthew D. Park, Luisanna Pia, Tamar Plitt, Venu Pothula, Jamie Redes, Ivan Reyes Torres, Emma Risson, Mark Roberto, Alfonso R. Sanchez-Paulete, Miriam Saffern, Bérengère Salomé, Myvizhi Esai Selvan, Joan Shang, Matthew Spindler, Alessandra Soares Schanoski, Maria Suprun, Jessica Tan, Michelle Tran, Verena van der Heide, Natalie Vaninov, C. Matthias Wilk, Julio Aguirre-Ghiso, Konstantina Alexandropoulos, Nina Bhardwaj, Dusan Bogunovic, Brian Brown, Judy Cho, Jeremiah Faith, Emilie Grasset, Zeynep H. Gümüş, Peter Heeger, Dirk Homann, Amir Horowitz, Ephraim Kenigsberg, Alice O. Kamphorst, Florian Krammer, Maria A. Curotto de Lafaille, Uri Laserson, Saurabh Mehandru, Miriam Merad, Robert Samstein, and Nicolas Vabret.
Acknowledgments
We apologize to all authors whose work we could not cite due to space limitations. Trainee (PhD and MD/PhD or postdocs) and faculty contributing authors are listed in alphabetical order. Illustrations are by Jill K. Gregory and used with permission of Mount Sinai Health System. We would like to acknowledge funding sources including Fastgrant (M.M.), NCI Cancer Center Support Grant Supplement (M.M., R.M.S.) Burroughs Wellcome Fund (R.M.S.), NIH Director’s Early Independence Award (R.M.S.) and NIH R01AI081848 (N.V., N.B., B.D.G.).
Declaration of Interests
N.B. serves as an advisor/board member for Neon, Checkpoint Sciences, Primevax, Novartis, Array BioPharma, Roche, Avidea, Boeringer Ingelheim, Rome Therapeutics, Roswell Park, and the Parker Institute for Cancer Immunotherapy. N.B. receives research support from the Parker Insitute, Novocure, Celldex, Genentech, Oncovir, and Regeneron. M.M. serves as an advisor/board member for Celsius, Pionyr, Compugen, Myeloids and Innate pharma and ad hoc for Takeda. M.M. receives research support from Regeneron, Takeda, and Genentech. A.M. has equity in Gilead Sciences and Regeneron Pharmaceuticals.
Contributor Information
The Sinai Immunology Review Project:
Manasi Agrawal, Mark Aleynick, Meriem Belabed, Matthew Brown, Maria Casanova-Acebes, Jovani Catalan, Monica Centa, Andrew Charap, Andrew Chan, Steven T. Chen, Jonathan Chung, Cansu Cimen Bozkus, Evan Cody, Francesca Cossarini, Erica Dalla, Nicolas Fernandez, John Grout, Dan Fu Ruan, Pauline Hamon, Etienne Humblin, Divya Jha, Julia Kodysh, Andrew Leader, Matthew Lin, Katherine Lindblad, Daniel Lozano-Ojalvo, Gabrielle Lubitz, Assaf Magen, Zafar Mahmood, Gustavo Martinez-Delgado, Jaime Mateus-Tique, Elliot Meritt, Chang Moon, Justine Noel, Tim O’Donnell, Miyo Ota, Tamar Plitt, Venu Pothula, Jamie Redes, Ivan Reyes Torres, Mark Roberto, Alfonso R. Sanchez-Paulete, Joan Shang, Alessandra Soares Schanoski, Maria Suprun, Michelle Tran, Natalie Vaninov, C. Matthias Wilk, Julio Aguirre-Ghiso, Dusan Bogunovic, Judy Cho, Jeremiah Faith, Emilie Grasset, Peter Heeger, Ephraim Kenigsberg, Florian Krammer, and Uri Laserson
References
- Adams E.R., Anand R., Andersson M.I., Auckland K., Baillie J.K., Barnes E., Bell J., Berry T., Bibi S., Carroll M. Evaluation of antibody testing for SARS-Cov-2 using ELISA and lateral flow immunoassays. medRxiv. 2020 doi: 10.1101/2020.04.15.20066407. [DOI] [Google Scholar]
- Agostini M.L., Andres E.L., Sims A.C., Graham R.L., Sheahan T.P., Lu X., Smith E.C., Case J.B., Feng J.Y., Jordan R. Coronavirus Susceptibility to the Antiviral Remdesivir (GS-5734) Is Mediated by the Viral Polymerase and the Proofreading Exoribonuclease. MBio. 2018;9:e00221. doi: 10.1128/mBio.00221-18. e18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahmed S.F., Quadeer A.A., McKay M.R. Preliminary Identification of Potential Vaccine Targets for the COVID-19 Coronavirus (SARS-CoV-2) Based on SARS-CoV Immunological Studies. Viruses. 2020;12:254. doi: 10.3390/v12030254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahn E., Araki K., Hashimoto M., Li W., Riley J.L., Cheung J., Sharpe A.H., Freeman G.J., Irving B.A., Ahmed R. Role of PD-1 during effector CD8 T cell differentiation. Proc. Natl. Acad. Sci. USA. 2018;115:4749–4754. doi: 10.1073/pnas.1718217115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahn J.Y., Sohn Y., Lee S.H., Cho Y., Hyun J.H., Baek Y.J., Jeong S.J., Kim J.H., Ku N.S., Yeom J.-S. Use of Convalescent Plasma Therapy in Two COVID-19 Patients with Acute Respiratory Distress Syndrome in Korea. J. Korean Med. Sci. 2020;35:e149. doi: 10.3346/jkms.2020.35.e149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alharbi N.K., Padron-Regalado E., Thompson C.P., Kupke A., Wells D., Sloan M.A., Grehan K., Temperton N., Lambe T., Warimwe G. ChAdOx1 and MVA based vaccine candidates against MERS-CoV elicit neutralising antibodies and cellular immune responses in mice. Vaccine. 2017;35:3780–3788. doi: 10.1016/j.vaccine.2017.05.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amanat F., Krammer F. SARS-CoV-2 Vaccines: Status Report. Immunity. 2020;52:583–589. doi: 10.1016/j.immuni.2020.03.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amanat F., Stadlbauer D., Strohmeier S., Nguyen T.H.O., Chromikova V., McMahon M., Jiang K., Asthagiri Arunkumar G., Jurczyszak D., Polanco J. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 2020 doi: 10.1038/s41591-020-0913-5. Published online May 12, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- André F.E. Vaccinology: Past Achievements, Present Roadblocks and Future Promises. Vaccine. 2003;21:593–595. doi: 10.1016/s0264-410x(02)00702-8. [DOI] [PubMed] [Google Scholar]
- Andreakos E., Salagianni M., Galani I.E., Koltsida O. Interferon-λs: Front-Line Guardians of Immunity and Homeostasis in the Respiratory Tract. Front. Immunol. 2017;8:1232. doi: 10.3389/fimmu.2017.01232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antrobus R.D., Coughlan L., Berthoud T.K., Dicks M.D., Hill A.V., Lambe T., Gilbert S.C. Clinical assessment of a novel recombinant simian adenovirus ChAdOx1 as a vectored vaccine expressing conserved Influenza A antigens. Mol. Ther. 2014;22:668–674. doi: 10.1038/mt.2013.284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arabi Y.M., Al-Enezi F., Longuere K.-S., Balkhy H.H., Al-Sultan M., Al-Omari A., Al-Hameed F.M., Carson G., Shindo N., Fowler R. Feasibility of a randomized controlled trial to assess treatment of Middle East Respiratory Syndrome Coronavirus (MERS-CoV) infection in Saudi Arabia: a survey of physicians. BMC Anesthesiol. 2016;16:36. doi: 10.1186/s12871-016-0198-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arabi Y.M., Shalhoub S., Mandourah Y., Al-Hameed F., Al-Omari A., Al Qasim E., Jose J., Alraddadi B., Almotairi A., Al Khatib K. Ribavirin and Interferon Therapy for Critically Ill Patients With Middle East Respiratory Syndrome: A Multicenter Observational Study. Clin. Infect. Dis. 2020;70:1837–1844. doi: 10.1093/cid/ciz544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asselta R., Paraboschi E.M., Mantovani A., Duga S. ACE2 and TMPRSS2 variants and expression as candidates to sex and country differences in COVID-19 severity in Italy. medRxiv. 2020 doi: 10.1101/2020.03.30.20047878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bacher P., Heinrich F., Stervbo U., Nienen M., Vahldieck M., Iwert C., Vogt K., Kollet J., Babel N., Sawitzki B. Regulatory T Cell Specificity Directs Tolerance versus Allergy against Aeroantigens in Humans. Cell. 2016;167:1067–1078.e16. doi: 10.1016/j.cell.2016.09.050. [DOI] [PubMed] [Google Scholar]
- Bao L., Deng W., Huang B., Gao H., Liu J., Ren L., Wei Q., Yu P., Xu Y., Qi F. The Pathogenicity of SARS-CoV-2 in hACE2 Transgenic Mice. Nature. 2020 doi: 10.1038/s41586-020-2312-y. Published online May 7, 2020. [DOI] [PubMed] [Google Scholar]
- Bao L., Deng W., Gao H., Xiao C., Liu J., Xue J., Lv Q., Liu J., Yu P., Xu Y. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. bioRxiv. 2020 doi: 10.1101/2020.03.13.990226. [DOI] [Google Scholar]
- Barnes B.J., Adrover J.M., Baxter-Stoltzfus A., Borczuk A., Cools-Lartigue J., Crawford J.M., Daßler-Plenker J., Guerci P., Huynh C., Knight J.S. Targeting potential drivers of COVID-19: Neutrophil extracellular traps. J. Exp. Med. 2020;217:e20200652. doi: 10.1084/jem.20200652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baruah V., Bose S. Immunoinformatics-aided identification of T cell and B cell epitopes in the surface glycoprotein of 2019-nCoV. J. Med. Virol. 2020;92:495–500. doi: 10.1002/jmv.25698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belhadi D., Peiffer-Smadja N., Lescure F.-X., Yazdanpanah Y., Mentré F., Laouénan C. A brief review of antiviral drugs evaluated in registered clinical trials for COVID-19. medRxiv. 2020 doi: 10.1101/2020.03.18.20038190. [DOI] [Google Scholar]
- Bhattacharya M., Sharma A.R., Patra P., Ghosh P., Sharma G., Patra B.C., Lee S.-S., Chakraborty C. ). Development of Epitope-Based Peptide Vaccine Against Novel Coronavirus 2019 (SARS-COV-2): Immunoinformatics Approach. J. Med. Virol. 2020 doi: 10.1002/jmv.25736. Published online February 28, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanco-Melo D., Nilsson-Payant B.E., Liu W.-C., Uhl S., Hoagland D., Møller R., Jordan T.X., Oishi K., Panis M., Sachs D. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell. 2020 doi: 10.1016/j.cell.2020.04.026. Published online May 15, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borba M.G.S., de Almeida Val F., Sampaio V.S., Alexandre M.A.A., Melo G.C., Brito M., Mourão M.P.G., Brito Sousa J.D., Baía-da-Silva D.C., Guerra M.V.F. Effect of High vs Low Doses of Chloroquine Diphosphate as Adjunctive Therapy for Patients Hospitalized With Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection. JAMA Netw. Open. 2020;3:e208857. doi: 10.1001/jamanetworkopen.2020.8857. [DOI] [PubMed] [Google Scholar]
- Bouvet M., Debarnot C., Imbert I., Selisko B., Snijder E.J., Canard B., Decroly E. In vitro reconstitution of SARS-coronavirus mRNA cap methylation. PLoS Pathog. 2010;6:e1000863. doi: 10.1371/journal.ppat.1000863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braud V.M., Allan D.S., O’Callaghan C.A., Söderström K., D’Andrea A., Ogg G.S., Lazetic S., Young N.T., Bell J.I., Phillips J.H. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature. 1998;391:795–799. doi: 10.1038/35869. [DOI] [PubMed] [Google Scholar]
- Braun J., Loyal L., Frentsch M., Wendisch D., Georg P., Kurth F., Hippenstiel S., Dingeldey M., Kruse B., Fauchere F. Presence of SARS-CoV-2 reactive T cells in COVID-19 patients and healthy donors. medRxiv. 2020 doi: 10.1101/2020.04.17.20061440. [DOI] [PubMed] [Google Scholar]
- Brooks A.G., Posch P.E., Scorzelli C.J., Borrego F., Coligan J.E. NKG2A complexed with CD94 defines a novel inhibitory natural killer cell receptor. J. Exp. Med. 1997;185:795–800. doi: 10.1084/jem.185.4.795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukreyeva N., Mantlo E.K., Sattler R.A., Huang C., Paessler S., Zeldis J. The IMPDH inhibitor merimepodib suppresses SARS-CoV-2 replication in vitro. bioRxiv. 2020 doi: 10.1101/2020.04.07.028589. [DOI] [Google Scholar]
- Cai X.-f., Chen J., Hu J.-l., Long Q.-x., Deng H.-j., Fan K., Liao P., Liu B.-z., Wu G.-c., Chen Y.-k. A Peptide-Based Magnetic Chemiluminescence Enzyme Immunoassay for Serological Diagnosis of Corona virus Disease 2019. J. Infect. Dis. 2020 doi: 10.1093/infdis/jiaa243. Published online May 8, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Callow K.A., Parry H.F., Sergeant M., Tyrrell D.A. The time course of the immune response to experimental coronavirus infection of man. Epidemiol. Infect. 1990;105:435–446. doi: 10.1017/s0950268800048019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron M.J., Ran L., Xu L., Danesh A., Bermejo-Martin J.F., Cameron C.M., Muller M.P., Gold W.L., Richardson S.E., Poutanen S.M., Canadian SARS Research Network Interferon-mediated immunopathological events are associated with atypical innate and adaptive immune responses in patients with severe acute respiratory syndrome. J. Virol. 2007;81:8692–8706. doi: 10.1128/JVI.00527-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron M.J., Kelvin A.A., Leon A.J., Cameron C.M., Ran L., Xu L., Chu Y.K., Danesh A., Fang Y., Li Q. Lack of innate interferon responses during SARS coronavirus infection in a vaccination and reinfection ferret model. PLoS ONE. 2012;7:e45842. doi: 10.1371/journal.pone.0045842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camp J.V., Jonsson C.B. A Role for Neutrophils in Viral Respiratory Disease. Front. Immunol. 2017;8:550. doi: 10.3389/fimmu.2017.00550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell K.M., Steiner G., Wells D.K., Ribas A., Kalbasi A. Prediction of SARS-CoV-2 epitopes across 9360 HLA class I alleles. bioRxiv. 2020 doi: 10.1101/2020.03.30.016931. [DOI] [Google Scholar]
- Canton J., Fehr A.R., Fernandez-Delgado R., Gutierrez-Alvarez F.J., Sanchez-Aparicio M.T., García-Sastre A., Perlman S., Enjuanes L., Sola I. MERS-CoV 4b protein interferes with the NF-κB-dependent innate immune response during infection. PLoS Pathog. 2018;14:e1006838. doi: 10.1371/journal.ppat.1006838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao X. COVID-19: immunopathology and its implications for therapy. Nat. Rev. Immunol. 2020;20:269–270. doi: 10.1038/s41577-020-0308-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao W.C., Liu W., Zhang P.H., Zhang F., Richardus J.H. Disappearance of antibodies to SARS-associated coronavirus after recovery. N. Engl. J. Med. 2007;357:1162–1163. doi: 10.1056/NEJMc070348. [DOI] [PubMed] [Google Scholar]
- Cao Z., Liu L., Du L., Zhang C., Jiang S., Li T., He Y. Potent and persistent antibody responses against the receptor-binding domain of SARS-CoV spike protein in recovered patients. Virol. J. 2010;7:299. doi: 10.1186/1743-422X-7-299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao B., Wang Y., Wen D., Liu W., Wang J., Fan G., Ruan L., Song B., Cai Y., Wei M. A Trial of Lopinavir-Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020 doi: 10.1056/NEJMoa2001282. Published online March 18, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao W., Liu X., Bai T., Fan H., Hong K., Song H., Han Y., Lin L., Ruan L., Li T. High-Dose Intravenous Immunoglobulin as a Therapeutic Option for Deteriorating Patients With Coronavirus Disease 2019. Open Forum Infect. Dis. 2020;7:a102. doi: 10.1093/ofid/ofaa102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao Y., Li L., Feng Z., Wan S., Huang P., Sun X., Wen F., Huang X., Ning G., Wang W. Comparative genetic analysis of the novel coronavirus (2019-nCoV/SARS-CoV-2) receptor ACE2 in different populations. Cell Discov. 2020;6:11. doi: 10.1038/s41421-020-0147-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carlin L.E., Hemann E.A., Zacharias Z.R., Heusel J.W., Legge K.L. Natural Killer Cell Recruitment to the Lung During Influenza A Virus Infection Is Dependent on CXCR3, CCR5, and Virus Exposure Dose. Front. Immunol. 2018;9:781. doi: 10.3389/fimmu.2018.00781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cervantes-Barragan L., Züst R., Weber F., Spiegel M., Lang K.S., Akira S., Thiel V., Ludewig B. Control of coronavirus infection through plasmacytoid dendritic-cell-derived type I interferon. Blood. 2007;109:1131–1137. doi: 10.1182/blood-2006-05-023770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cerwenka A., Lanier L.L. Natural killer cells, viruses and cancer. Nat. Rev. Immunol. 2001;1:41–49. doi: 10.1038/35095564. [DOI] [PubMed] [Google Scholar]
- Chan K.Y., Ching J.C., Xu M.S., Cheung A.N., Yip S.P., Yam L.Y., Lai S.T., Chu C.M., Wong A.T., Song Y.Q. Association of ICAM3 genetic variant with severe acute respiratory syndrome. J. Infect. Dis. 2007;196:271–280. doi: 10.1086/518892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang Y.J., Kim H.Y., Albacker L.A., Baumgarth N., McKenzie A.N., Smith D.E., Dekruyff R.H., Umetsu D.T. Innate lymphoid cells mediate influenza-induced airway hyper-reactivity independently of adaptive immunity. Nat. Immunol. 2011;12:631–638. doi: 10.1038/ni.2045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Channappanavar R., Fehr A.R., Vijay R., Mack M., Zhao J., Meyerholz D.K., Perlman S. Dysregulated Type I Interferon and Inflammatory Monocyte-Macrophage Responses Cause Lethal Pneumonia in SARS-CoV-Infected Mice. Cell Host Microbe. 2016;19:181–193. doi: 10.1016/j.chom.2016.01.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Channappanavar R., Fehr A.R., Zheng J., Wohlford-Lenane C., Abrahante J.E., Mack M., Sompallae R., McCray P.B., Jr., Meyerholz D.K., Perlman S. IFN-I response timing relative to virus replication determines MERS coronavirus infection outcomes. J. Clin. Invest. 2019;130:3625–3639. doi: 10.1172/JCI126363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Charbonneau M.R., Isabella V.M., Li N., Kurtz C.B. Developing a new class of engineered live bacterial therapeutics to treat human diseases. Nat. Commun. 2020;11:1738. doi: 10.1038/s41467-020-15508-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y., Cai H., Pan J., Xiang N., Tien P., Ahola T., Guo D. Functional screen reveals SARS coronavirus nonstructural protein nsp14 as a novel cap N7 methyltransferase. Proc. Natl. Acad. Sci. USA. 2009;106:3484–3489. doi: 10.1073/pnas.0808790106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen I.Y., Moriyama M., Chang M.F., Ichinohe T. Severe acute respiratory syndrome coronavirus viroporin 3a activates the NLRP3 inflammasome. Front. Microbiol. 2019;10:50. doi: 10.3389/fmicb.2019.00050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen X., Li R., Pan Z., Qian C., Yang Y., You R., Zhao J., Liu P., Gao L., Li Z. Human monoclonal antibodies block the binding of SARS-CoV-2 spike protein to angiotensin converting enzyme 2 receptor. Cell. Mol. Immunol. 2020 doi: 10.1038/s41423-020-0426-7. Published online April 20, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen C., Zhang Y., Huang J., Yin P., Cheng Z., Wu J., Chen S., Zhang Y., Chen B., Lu M. Favipiravir versus Arbidol for COVID-19: A Randomized Clinical Trial. medRxiv. 2020 doi: 10.1101/2020.03.17.20037432. [DOI] [Google Scholar]
- Chen G., Wu D., Guo W., Cao Y., Huang D., Wang H., Wang T., Zhang X., Chen H., Yu H. Clinical and immunological features of severe and moderate coronavirus disease 2019. J. Clin. Invest. 2020 doi: 10.1172/JCI137244. Published online March 27, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J., Liu D., Liu L., Liu P., Xu Q., Xia L., Ling Y., Dan H., Song S., Zhang D. A pilot study of hydroxychloroquine in treatment of patients with common coronavirus disease-19 (COVID-19) J. Zhejiang Univ. 2020;49 doi: 10.3785/j.issn.1008-9292.2020.03.03. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen L., Liu H.G., Liu W., Liu J., Liu K., Shang J., Deng Y., Wei S. [Analysis of clinical features of 29 patients with 2019 novel coronavirus pneumonia] Zhonghua Jie He He Hu Xi Za Zhi. 2020;43:203–208. doi: 10.3760/cma.j.issn.1001-0939.2020.03.013. [DOI] [PubMed] [Google Scholar]
- Chen X., Ling J., Mo P., Zhang Y., Jiang Q., Ma Z., Cao Q., Hu W., Zou S., Chen L. Restoration of leukomonocyte counts is associated with viral clearance in COVID-19 hospitalized patients. medRxiv. 2020 doi: 10.1101/2020.03.03.20030437. [DOI] [Google Scholar]
- Chen X., Zhao B., Qu Y., Chen Y., Xiong J., Feng Y., Men D., Huang Q., Liu Y., Yang B. Detectable serum SARS-CoV-2 viral load (RNAaemia) is closely associated with drastically elevated interleukin 6 (IL-6) level in critically ill COVID-19 patients. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa449. Published online April 17, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y., Feng Z., Diao B., Wang R., Wang G., Wang C., Tan Y., Liu L., Wang C., Liu Y. The Novel Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Directly Decimates Human Spleens and Lymph Nodes. medRxiv. 2020 doi: 10.1101/2020.03.27.20045427. [DOI] [Google Scholar]
- Chen Z., Hu J., Zhang Z., Jiang S., Han S., Yan D., Zhuang R., Hu B., Zhang Z. Efficacy of hydroxychloroquine in patients with COVID-19: results of a randomized clinical trial. medRxiv. 2020 doi: 10.1101/2020.03.22.20040758. [DOI] [Google Scholar]
- Cheng Y., Cheng G., Chui C.H., Lau F.Y., Chan P.K., Ng M.H., Sung J.J., Wong R.S. ABO blood group and susceptibility to severe acute respiratory syndrome. JAMA. 2005;293:1450–1451. doi: 10.1001/jama.293.12.1450-c. [DOI] [PubMed] [Google Scholar]
- Cheng Y., Wong R., Soo Y.O.Y., Wong W.S., Lee C.K., Ng M.H.L., Chan P., Wong K.C., Leung C.B., Cheng G. Use of convalescent plasma therapy in SARS patients in Hong Kong. Eur. J. Clin. Microbiol. Infect. Dis. 2005;24:44–46. doi: 10.1007/s10096-004-1271-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung C.Y., Poon L.L.M., Ng I.H.Y., Luk W., Sia S.-F., Wu M.H.S., Chan K.-H., Yuen K.-Y., Gordon S., Guan Y., Peiris J.S. Cytokine responses in severe acute respiratory syndrome coronavirus-infected macrophages in vitro: possible relevance to pathogenesis. J. Virol. 2005;79:7819–7826. doi: 10.1128/JVI.79.12.7819-7826.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chien J.-Y., Hsueh P.-R., Cheng W.-C., Yu C.J., Yang P.-C. Temporal changes in cytokine/chemokine profiles and pulmonary involvement in severe acute respiratory syndrome. Respirology. 2006;11:715–722. doi: 10.1111/j.1440-1843.2006.00942.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu C.M., Cheng V.C., Hung I.F., Wong M.M., Chan K.H., Chan K.S., Kao R.Y., Poon L.L., Wong C.L., Guan Y., HKU/UCH SARS Study Group Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004;59:252–256. doi: 10.1136/thorax.2003.012658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu H., Zhou J., Wong B.H., Li C., Chan J.F., Cheng Z.S., Yang D., Wang D., Lee A.C., Li C. Middle East Respiratory Syndrome Coronavirus Efficiently Infects Human Primary T Lymphocytes and Activates the Extrinsic and Intrinsic Apoptosis Pathways. J. Infect. Dis. 2016;213:904–914. doi: 10.1093/infdis/jiv380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu H., Chan J.F.-W., Wang Y., Yuen T.T.-T., Chai Y., Hou Y., Shuai H., Yang D., Hu B., Huang X. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa410. Published online April 9, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cifaldi L., Prencipe G., Caiello I., Bracaglia C., Locatelli F., De Benedetti F., Strippoli R. Inhibition of natural killer cell cytotoxicity by interleukin-6: implications for the pathogenesis of macrophage activation syndrome. Arthritis Rheumatol. 2015;67:3037–3046. doi: 10.1002/art.39295. [DOI] [PubMed] [Google Scholar]
- Clay C., Donart N., Fomukong N., Knight J.B., Lei W., Price L., Hahn F., Van Westrienen J., Harrod K.S. Primary severe acute respiratory syndrome coronavirus infection limits replication but not lung inflammation upon homologous rechallenge. J. Virol. 2012;86:4234–4244. doi: 10.1128/JVI.06791-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clementi N., Criscuolo E., Diotti R.A., Ferrarese R., Castelli M., Burioni R., Clementi M., Mancini N. Combined prophylactic and therapeutic use maximizes hydroxychloroquine anti-SARS-CoV-2 effects in vitro. bioRxiv. 2020 doi: 10.1101/2020.03.29.014407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Comar C.E., Goldstein S.A., Li Y., Yount B., Baric R.S., Weiss S.R. Antagonism of dsRNA-Induced Innate Immune Pathways by NS4a and NS4b Accessory Proteins during MERS Coronavirus Infection. MBio. 2019;10:1–7. doi: 10.1128/mBio.00319-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corley M.J., Sugai C., Schotsaert M., Schwartz R.E., Ndhlovu L.C. Comparative in vitro transcriptomic analyses of COVID-19 candidate therapy hydroxychloroquine suggest limited immunomodulatory evidence of SARS-CoV-2 host response genes. bioRxiv. 2020 doi: 10.1101/2020.04.13.039263. [DOI] [Google Scholar]
- Dai W., Zhang B., Su H., Li J., Zhao Y., Xie X., Jin Z., Liu F., Li C., Li Y. Structure-based Design of Antiviral Drug Candidates Targeting the SARS-CoV-2 Main Protease. Science. 2020 doi: 10.1126/science.abb4489. Published online April 22, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Grove K.C., Provoost S., Verhamme F.M., Bracke K.R., Joos G.F., Maes T., Brusselle G.G. Characterization and Quantification of Innate Lymphoid Cell Subsets in Human Lung. PLoS ONE. 2016;11:e0145961. doi: 10.1371/journal.pone.0145961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Marcken M., Dhaliwal K., Danielsen A.C., Gautron A.S., Dominguez-Villar M. TLR7 and TLR8 activate distinct pathways in monocytes during RNA virus infection. Sci. Signal. 2019;12:eaaw1347. doi: 10.1126/scisignal.aaw1347. [DOI] [PubMed] [Google Scholar]
- de Wit E., van Doremalen N., Falzarano D., Munster V.J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016;14:523–534. doi: 10.1038/nrmicro.2016.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Wit E., Feldmann F., Cronin J., Jordan R., Okumura A., Thomas T., Scott D., Cihlar T., Feldmann H. Prophylactic and therapeutic remdesivir (GS-5734) treatment in the rhesus macaque model of MERS-CoV infection. Proc. Natl. Acad. Sci. USA. 2020;117:6771–6776. doi: 10.1073/pnas.1922083117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Del Valle D.M., Kim-Schulze S., Hsin-Hui H., Beckmann N.D., Nirenberg S., Wang B., Lavin Y., Swartz T., Madduri D., Stock A. An inflammatory cytokine signature helps predict COVID-19 severity and death. medRxiv. 2020 doi: 10.1101/2020.05.28.20115758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng X., Hackbart M., Mettelman R.C., O’Brien A., Mielech A.M., Yi G., Kao C.C., Baker S.C. Coronavirus nonstructural protein 15 mediates evasion of dsRNA sensors and limits apoptosis in macrophages. Proc. Natl. Acad. Sci. USA. 2017;114:E4251–E4260. doi: 10.1073/pnas.1618310114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devaraj S.G., Wang N., Chen Z., Chen Z., Tseng M., Barretto N., Lin R., Peters C.J., Tseng C.T.K., Baker S.C., Li K. Regulation of IRF-3-dependent innate immunity by the papain-like protease domain of the severe acute respiratory syndrome coronavirus. J. Biol. Chem. 2007;282:32208–32221. doi: 10.1074/jbc.M704870200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diao B., Wang C., Tan Y., Chen X., Liu Y., Ning L., Chen L., Li M., Liu Y., Wang G. Reduction and Functional Exhaustion of T Cells in Patients with Coronavirus Disease 2019 (COVID-19) Front. Immunol. 2020 doi: 10.3389/fimmu.2020.00827. Published online May 1, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Díez J.-M., Romero C., Gajardo R. Currently available intravenous immunoglobulin (Gamunex®-C and Flebogamma® DIF) contains antibodies reacting against SARS-CoV-2 antigens. bioRxiv. 2020 doi: 10.1101/2020.04.07.029017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong P., Ju X., Yan Y., Zhang S., Cai M., Wang H., Chen H., Hu Y., Cui L., Zhang J., He W. γδ T Cells Provide Protective Function in Highly Pathogenic Avian H5N1 Influenza A Virus Infection. Front. Immunol. 2018;9:2812. doi: 10.3389/fimmu.2018.02812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Draghi M., Pashine A., Sanjanwala B., Gendzekhadze K., Cantoni C., Cosman D., Moretta A., Valiante N.M., Parham P. NKp46 and NKG2D recognition of infected dendritic cells is necessary for NK cell activation in the human response to influenza infection. J. Immunol. 2007;178:2688–2698. doi: 10.4049/jimmunol.178.5.2688. [DOI] [PubMed] [Google Scholar]
- Du L., He Y., Zhou Y., Liu S., Zheng B.J., Jiang S. The spike protein of SARS-CoV--a target for vaccine and therapeutic development. Nat. Rev. Microbiol. 2009;7:226–236. doi: 10.1038/nrmicro2090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du M., Cai G., Chen F., Christiani D.C., Zhang Z., Wang M. Multi-omics Evaluation of Gastrointestinal and Other Clinical Characteristics of SARS-CoV-2 and COVID-19. Gastroenterology. 2020 doi: 10.1053/j.gastro.2020.03.045. Published online March 28, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duan K., Liu B., Li C., Zhang H., Yu T., Qu J., Zhou M., Chen L., Meng S., Hu Y. Effectiveness of convalescent plasma therapy in severe COVID-19 patients. PNAS. 2020;117:9490–9496. doi: 10.1073/pnas.2004168117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duev-Cohen A., Bar-On Y., Glasner A., Berhani O., Ophir Y., Levi-Schaffer F., Mandelboim M., Mandelboim O. The human 2B4 and NTB-A receptors bind the influenza viral hemagglutinin and co-stimulate NK cell cytotoxicity. Oncotarget. 2016;7:13093–13105. doi: 10.18632/oncotarget.7597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fang X., Mei Q., Yang T., Li L., Wang Y., Tong F., Geng S., Pan A. Low-dose corticosteroid therapy does not delay viral clearance in patients with COVID-19. J. Infect. 2020 doi: 10.1016/j.jinf.2020.03.039. Published online April 11, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fast E., Altman R.B., Chen B. Potential T-cell and B-cell Epitopes of 2019-nCoV. bioRxiv. 2020 doi: 10.1101/2020.02.19.955484. [DOI] [Google Scholar]
- Fauci A.S., Lane H.C., Redfield R.R. Covid-19 - Navigating the Uncharted. N. Engl. J. Med. 2020;382:1268–1269. doi: 10.1056/NEJMe2002387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fei J., Fu L., Li Y., Xiang H.-X., Xiang Y., Li M.-D., Liu F.-F., Xu D.-X., Zhao H. Reduction of lymphocyte at early stage elevates severity and death risk of COVID-19 patients: a hospital-based case-cohort study. medRxiv. 2020 doi: 10.1101/2020.04.02.20050955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng Z., Yu Q., Yao S., Luo L., Duan J., Yan Z., Yang M., Tan H., Ma M., Li T. Early Prediction of Disease Progression in 2019 Novel Coronavirus Pneumonia Patients Outside Wuhan with CT and Clinical Characteristics. medRxiv. 2020 doi: 10.1101/2020.02.19.20025296. [DOI] [Google Scholar]
- Flores-Torres A.S., Salinas-Carmona M.C., Salinas E., Rosas-Taraco A.G. Eosinophils and Respiratory Viruses. Viral Immunol. 2019;32:198–207. doi: 10.1089/vim.2018.0150. [DOI] [PubMed] [Google Scholar]
- Fogarty H., Townsend L., Cheallaigh C.N., Bergin C., Martin-Loeches I., Browne P., Bacon C.L., Gaule R., Gillett A., Byrne M. COVID-19 Coagulopathy in Caucasian patients. Br. J. Haematol. 2020 doi: 10.1111/bjh.16749. Published online April 24, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forni D., Filippi G., Cagliani R., De Gioia L., Pozzoli U., Al-Daghri N., Clerici M., Sironi M. The heptad repeat region is a major selection target in MERS-CoV and related coronaviruses. Sci. Rep. 2015;5:14480. doi: 10.1038/srep14480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frieman M., Yount B., Heise M., Kopecky-Bromberg S.A., Palese P., Baric R.S. Severe acute respiratory syndrome coronavirus ORF6 antagonizes STAT1 function by sequestering nuclear import factors on the rough endoplasmic reticulum/Golgi membrane. J. Virol. 2007;81:9812–9824. doi: 10.1128/JVI.01012-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frieman M., Ratia K., Johnston R.E., Mesecar A.D., Baric R.S. Severe acute respiratory syndrome coronavirus papain-like protease ubiquitin-like domain and catalytic domain regulate antagonism of IRF3 and NF-kappaB signaling. J. Virol. 2009;83:6689–6705. doi: 10.1128/JVI.02220-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu S., Fu X., Song Y., Li M., Pan P.-h., Tang T., Zhang C., Jiang T., Tan D., Fan X. Virologic and clinical characteristics for prognosis of severe COVID-19: a retrospective observational study in Wuhan, China. medRxiv. 2020 doi: 10.1101/2020.04.03.20051763. [DOI] [Google Scholar]
- Gangopadhyay K.K., Mukherjee J.J., Sinha B., Ghosal S. The role of corticosteroids in the management of critically ill patients with coronavirus disease 2019 (COVID-19): A meta-analysis. medRxiv. 2020 doi: 10.1101/2020.04.17.20069773. [DOI] [Google Scholar]
- Gao K., Nguyen D.D., Wang R., Wei G.-W. Machine intelligence design of 2019-nCoV drugs. bioRxiv. 2020 doi: 10.1101/2020.01.30.927889. [DOI] [Google Scholar]
- Gao L., Jiang D., Wen X.-s., Cheng X.-c., Sun M., He B., You L.-n., Lei P., Tan X.-w., Qin S. Prognostic value of NT-proBNP in patients with severe COVID-19. Respir Res. 2020;21:83. doi: 10.1186/s12931-020-01352-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao Q., Bao L., Mao H., Wang L., Xu K., Yang M., Li Y., Zhu L., Wang N., Lv Z. Rapid development of an inactivated vaccine for SARS-CoV-2. bioRxiv. 2020 doi: 10.1101/2020.04.17.046375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao T., Hu M., Zhang X., Li H., Zhu L., Liu H., Dong Q., Zhang Z., Wang Z., Hu Y. Highly pathogenic coronavirus N protein aggravates lung injury by MASP-2-mediated complement over-activation. medRxiv. 2020 doi: 10.1101/2020.03.29.20041962. [DOI] [Google Scholar]
- Garulli B., Stillitano M.G., Barnaba V., Castrucci M.R. Primary CD8+ T-cell response to soluble ovalbumin is improved by chloroquine treatment in vivo. Clin. Vaccine Immunol. 2008;15:1497–1504. doi: 10.1128/CVI.00166-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasteiger G., Fan X., Dikiy S., Lee S.Y., Rudensky A.Y. Tissue residency of innate lymphoid cells in lymphoid and nonlymphoid organs. Science. 2015;350:981–985. doi: 10.1126/science.aac9593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautret P., Lagier J.-C., Parola P., Hoang V.T., Meddeb L., Mailhe M., Doudier B., Courjon J., Giordanengo V., Vieira V.E. Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int. J. Antimicrob. Agents. 2020:105949. doi: 10.1016/j.ijantimicag.2020.105949. Published online March 20, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautret P., Lagier J.-C., Parola P., Hoang V.T., Meddeb L., Sevestre J., Mailhe M., Doudier B., Aubry C., Amrane S. Clinical and microbiological effect of a combination of hydroxychloroquine and azithromycin in 80 COVID-19 patients with at least a six-day follow up: A pilot observational study. Travel Med. Infect. Dis. 2020:101663. doi: 10.1016/j.tmaid.2020.101663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giamarellos-Bourboulis E.J., Netea M.G., Rovina N., Akinosoglou K., Antoniadou A., Antonakos N., Damoraki G., Gkavogianni T., Adami M.-E., Katsaounou P. Complex Immune Dysregulation in COVID-19 Patients With Severe Respiratory Failure. Cell Host Microbe. 2020 doi: 10.1016/j.chom.2020.04.009. Published online April 17, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gioia C., Horejsh D., Agrati C., Martini F., Capobianchi M.R., Ippolito G., Poccia F. T-Cell response profiling to biological threat agents including the SARS coronavirus. Int. J. Immunopathol. Pharmacol. 2005;18:525–530. doi: 10.1177/039463200501800312. [DOI] [PubMed] [Google Scholar]
- Giron C.C., Laaksonen A., da Silva F.L.B. On the Interactions of the Receptor-Binding Domain of SARS-CoV-1 and SARS-CoV-2 Spike Proteins with Monoclonal Antibodies and the Receptor ACE2. Virus Res. 2020;285:198021. doi: 10.1016/j.virusres.2020.198021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glasner A., Zurunic A., Meningher T., Lenac Rovis T., Tsukerman P., Bar-On Y., Yamin R., Meyers A.F., Mandeboim M., Jonjic S., Mandelboim O. Elucidating the mechanisms of influenza virus recognition by Ncr1. PLoS ONE. 2012;7:e36837. doi: 10.1371/journal.pone.0036837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gong J., Dong H., Xia S.Q., Huang Y.Z., Wang D., Zhao Y., Liu W., Tu S., Zhang M., Wang Q. Correlation Analysis Between Disease Severity and Inflammation-related Parameters in Patients with COVID-19 Pneumonia. medRxiv. 2020 doi: 10.1101/2020.02.25.20025643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordon D.E., Jang G.M., Bouhaddou M., Xu J., Obernier K., White K.M., O’Meara M.J., Rezelj V.V., Guo J.Z., Swaney D.L. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. 2020 doi: 10.1038/s41586-020-2286-9. Published online April 30, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grein J., Ohmagari N., Shin D., Diaz G., Asperges E., Castagna A., Feldt T., Green G., Green M.L., Lescure F.-X. Compassionate Use of Remdesivir for Patients with Severe Covid-19. N. Engl. J. Med. 2020 doi: 10.1056/NEJMoa2007016. Published online April 10, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grifoni A., Sidney J., Zhang Y., Scheuermann R.H., Peters B., Sette A. A Sequence Homology and Bioinformatic Approach Can Predict Candidate Targets for Immune Responses to SARS-CoV-2. Cell Host Microbe. 2020;27:671–680. doi: 10.1016/j.chom.2020.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guilliams M., Lambrecht B.N., Hammad H. Division of labor between lung dendritic cells and macrophages in the defense against pulmonary infections. Mucosal Immunol. 2013;6:464–473. doi: 10.1038/mi.2013.14. [DOI] [PubMed] [Google Scholar]
- Guo C., Li B., Ma H., Wang X., Cai P., Yu Q., Zhu L., Jin L., Jiang C., Fang J. Tocilizumab treatment in severe COVID-19 patients attenuates the inflammatory storm incited by monocyte centric immune interactions revealed by single-cell analysis. bioRxiv. 2020 doi: 10.1101/2020.04.08.029769. [DOI] [Google Scholar]
- Hackbart M., Deng X., Baker S.C. Coronavirus endoribonuclease targets viral polyuridine sequences to evade activating host sensors. Proc. Nat. Acad. Sci. USA. 2020;117:8094–8103. doi: 10.1073/pnas.1921485117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hadjadj J., Yatim N., Barnabei L., Corneau A., Boussier J., Pere H., Charbit B., Bondet V., Chenevier-Gobeaux C., Breillat P. Impaired type I interferon activity and exacerbated inflammatory responses in severe Covid-19 patients. medRxiv. 2020 doi: 10.1101/2020.04.19.20068015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haveri A., Smura T., Kuivanen S., Österlund P., Hepojoki J., Ikonen N., Pitkäpaasi M., Blomqvist S., Rönkkö E., Kantele A. Serological and molecular findings during SARS-CoV-2 infection: the first case study in Finland, January to February 2020. Euro Surveill. 2020;25:2000266. doi: 10.2807/1560-7917.ES.2020.25.11.2000266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayden F.G., Shindo N. Influenza virus polymerase inhibitors in clinical development. Curr. Opin. Infect. Dis. 2019;32:176–186. doi: 10.1097/QCO.0000000000000532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Z., Zhao C., Dong Q., Zhuang H., Song S., Peng G., Dwyer D.E. Effects of severe acute respiratory syndrome (SARS) coronavirus infection on peripheral blood lymphocytes and their subsets. Int. J. Infect. Dis. 2005;9:323–330. doi: 10.1016/j.ijid.2004.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herold T., Jurinovic V., Arnreich C., Lipworth B.J., Hellmuth J.C., von Bergwelt-Baildon M., Klein M., Weinberger T. Elevated levels of interleukin-6 and CRP predict the need for mechanical ventilation in COVID-19. J. Allergy Clin. Immunol. 2020 doi: 10.1016/j.jaci.2020.05.008. Published online May 18, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirano T., Murakami M. COVID-19: A New Virus, but a Familiar Receptor and Cytokine Release System. Immunity. 2020;52 doi: 10.1016/j.immuni.2020.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho M.-S., Chen W.-J., Chen H.-Y., Lin S.-F., Wang M.-C., Di J., Lu Y.-T., Liu C.-L., Chang S.-C., Chao C.-L. Neutralizing antibody response and SARS severity. Emerg. Infect. Dis. 2005;11:1730–1737. doi: 10.3201/eid1111.040659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsueh P.R., Huang L.M., Chen P.J., Kao C.L., Yang P.C. Chronological evolution of IgM, IgA, IgG and neutralisation antibodies after infection with SARS-associated coronavirus. Clin. Microbiol. Infect. 2004;10:1062–1066. doi: 10.1111/j.1469-0691.2004.01009.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu Y., Li W., Gao T., Cui Y., Jin Y., Li P., Ma Q., Liu X., Cao C. The Severe Acute Respiratory Syndrome Coronavirus Nucleocapsid Inhibits Type I Interferon Production by Interfering with TRIM25-Mediated RIG-I Ubiquitination. J. Virol. 2017;91:e02143. doi: 10.1128/JVI.02143-16. e16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang C., Lokugamage K.G., Rozovics J.M., Narayanan K., Semler B.L., Makino S. SARS coronavirus nsp1 protein induces template-dependent endonucleolytic cleavage of mRNAs: viral mRNAs are resistant to nsp1-induced RNA cleavage. PLoS Pathog. 2011;7:e1002433. doi: 10.1371/journal.ppat.1002433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang I.-C., Bailey C.C., Weyer J.L., Radoshitzky S.R., Becker M.M., Chiang J.J., Brass A.L., Ahmed A.A., Chi X., Dong L. Distinct patterns of IFITM-mediated restriction of filoviruses, SARS coronavirus, and influenza A virus. PLoS Pathog. 2011;7:e1001258. doi: 10.1371/journal.ppat.1001258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H., Wang S., Jiang T., Fan R., Zhang Z., Mu J., Li K., Wang Y., Jin L., Lin F. High levels of circulating GM-CSF+CD4+ T cells are predictive of poor outcomes in sepsis patients: a prospective cohort study. Cell. Mol. Immunol. 2019;16:602–610. doi: 10.1038/s41423-018-0164-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang A.T., Garcia-Carreras B., Hitchings M.D.T., Yang B., Katzelnick L., Rattigan S.M., Borgert B., Moreno C., Solomon B.D., Rodriguez-Barraquer I. A systematic review of antibody mediated immunity to coronaviruses: antibody kinetics, correlates of protection, and association of antibody responses with severity of disease. medRxiv. 2020 doi: 10.1101/2020.04.14.20065771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang C., Wang Y., Li X., Ren L., Zhao J., Hu Y., Zhang L., Fan G., Xu J., Gu X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang L., Shi Y., Gong B., Jiang L., Liu X., Yang J., Tang J., You C., Jiang Q., Long B. Blood single cell immune profiling reveals the interferon-MAPK pathway mediated adaptive immune response for COVID-19. medRxiv. 2020 doi: 10.1101/2020.03.15.20033472. [DOI] [Google Scholar]
- Huang Y., Yang R., Xu Y., Gong P. Clinical characteristics of 36 non-survivors with COVID-19 in Wuhan, China. medRxiv. 2020 doi: 10.1101/2020.02.27.20029009. [DOI] [Google Scholar]
- Hulme O.J., Wagenmakers E.-J., Damkier P., Madelung C.F., Siebner H.R., Helweg-Larsen J., Gronau Q., Benfield T.L., Madsen K.H. A Bayesian reanalysis of the effects of hydroxychloroquine and azithromycin on viral carriage in patients with COVID-19. medRxiv. 2020 doi: 10.1101/2020.03.31.20048777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hung I.F.N., To K.K.W., Lee C.-K., Lee K.-L., Chan K., Yan W.-W., Liu R., Watt C.-L., Chan W.-M., Lai K.-Y. Convalescent plasma treatment reduced mortality in patients with severe pandemic influenza A (H1N1) 2009 virus infection. Clin. Infect. Dis. 2011;52:447–456. doi: 10.1093/cid/ciq106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ivagnès A., Messaoudene M., Stoll G., Routy B., Fluckiger A., Yamazaki T., Iribarren K., Duong C.P.M., Fend L., Caignard A. TNFR2/BIRC3-TRAF1 signaling pathway as a novel NK cell immune checkpoint in cancer. OncoImmunology. 2017;7:e1386826. doi: 10.1080/2162402X.2017.1386826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ivanov K.A., Thiel V., Dobbe J.C., van der Meer Y., Snijder E.J., Ziebuhr J. Multiple enzymatic activities associated with severe acute respiratory syndrome coronavirus helicase. J. Virol. 2004;78:5619–5632. doi: 10.1128/JVI.78.11.5619-5632.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji W., Bishnu G., Cai Z., Shen X. Analysis clinical features of COVID-19 infection in secondary epidemic area and report potential biomarkers in evaluation. medRxiv. 2020 doi: 10.1101/2020.03.10.20033613. [DOI] [Google Scholar]
- Jia W., Channappanavar R., Zhang C., Li M., Zhou H., Zhang S., Zhou P., Xu J., Shan S., Shi X. Single intranasal immunization with chimpanzee adenovirus-based vaccine induces sustained and protective immunity against MERS-CoV infection. Emerg. Microbes Infect. 2019;8:760–772. doi: 10.1080/22221751.2019.1620083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- John S., Yuzhakov O., Woods A., Deterling J., Hassett K., Shaw C.A., Ciaramella G. Multi-antigenic human cytomegalovirus mRNA vaccines that elicit potent humoral and cell-mediated immunity. Vaccine. 2018;36:1689–1699. doi: 10.1016/j.vaccine.2018.01.029. [DOI] [PubMed] [Google Scholar]
- Ju B., Zhang Q., Ge X., Wang R., Sun J., Ge X., Yu J., Shan S., Zhou B., Song S. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature. 2020 doi: 10.1038/s41586-020-2380-z. Published online May 26, 2020. [DOI] [PubMed] [Google Scholar]
- Kamitani W., Huang C., Narayanan K., Lokugamage K.G., Makino S. A two-pronged strategy to suppress host protein synthesis by SARS coronavirus Nsp1 protein. Nat. Struct. Mol. Biol. 2009;16:1134–1140. doi: 10.1038/nsmb.1680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamphuis E., Junt T., Waibler Z., Forster R., Kalinke U. Type I interferons directly regulate lymphocyte recirculation and cause transient blood lymphopenia. Blood. 2006;108:3253–3261. doi: 10.1182/blood-2006-06-027599. [DOI] [PubMed] [Google Scholar]
- Keyaerts E., Vijgen L., Maes P., Neyts J., Van Ranst M. In vitro inhibition of severe acute respiratory syndrome coronavirus by chloroquine. Biochem. Biophys. Res. Commun. 2004;323:264–268. doi: 10.1016/j.bbrc.2004.08.085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J., Chang Y., Bae B., Sohn K.-H., Cho S.-H., Chung D.H., Kang H.R., Kim H.Y. Innate immune crosstalk in asthmatic airways: Innate lymphoid cells coordinate polarization of lung macrophages. J. Allergy Clin. Immunol. 2019;143:1769–1782.e11. doi: 10.1016/j.jaci.2018.10.040. [DOI] [PubMed] [Google Scholar]
- Kim E., Erdos G., Huang S., Kenniston T.W., Balmert S.C., Carey C.D., Raj V.S., Epperly M.W., Klimstra W.B., Haagmans B.L. Microneedle array delivered recombinant coronavirus vaccines: Immunogenicity and rapid translational development. EBioMedicine. 2020:102743. doi: 10.1016/j.ebiom.2020.102743. Published online April 2, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knoops K., Kikkert M., Worm S.H., Zevenhoven-Dobbe J.C., van der Meer Y., Koster A.J., Mommaas A.M., Snijder E.J. SARS-coronavirus replication is supported by a reticulovesicular network of modified endoplasmic reticulum. PLoS Biol. 2008;6:e226. doi: 10.1371/journal.pbio.0060226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konig M.F., Powell M., Staedtke V., Bai R.-Y., Thomas D.L., Fischer N., Huq S., Khalafallah A.M., Koenecke A., Xiong R. Targeting the catecholamine-cytokine axis to prevent SARS-CoV-2 cytokine storm syndrome. medRxiv. 2020 doi: 10.1101/2020.04.02.20051565. [DOI] [Google Scholar]
- Kopecky-Bromberg S.A., Martínez-Sobrido L., Frieman M., Baric R.A., Palese P. Severe acute respiratory syndrome coronavirus open reading frame (ORF) 3b, ORF 6, and nucleocapsid proteins function as interferon antagonists. J. Virol. 2007;81:548–557. doi: 10.1128/JVI.01782-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Law H.K.W., Cheung C.Y., Ng H.Y., Sia S.F., Chan Y.O., Luk W., Nicholls J.M., Peiris J.S.M., Lau Y.L. Chemokine up-regulation in SARS-coronavirus-infected, monocyte-derived human dendritic cells. Blood. 2005;106:2366–2374. doi: 10.1182/blood-2004-10-4166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ledford H. Hopes rise for coronavirus drug remdesivir. Nature. 2020 doi: 10.1038/d41586-020-01295-8. https://www.nature.com/articles/d41586-020-01295-8 [DOI] [PubMed] [Google Scholar]
- Lee J., Lee S.H., Shin N., Jeong M., Kim M.S., Kim M.J., Yoon S.R., Chung J.W., Kim T.D., Choi I. Tumor necrosis factor-alpha enhances IL-15-induced natural killer cell differentiation. Biochem. Biophys. Res. Commun. 2009;386:718–723. doi: 10.1016/j.bbrc.2009.06.120. [DOI] [PubMed] [Google Scholar]
- Lee K.-Y., Rhim J.-W., Kang J.-H. Early preemptive immunomodulators (corticosteroids) for severe pneumonia patients infected with SARS-CoV-2. Clin Exp Pediatr. 2020;63:117–118. doi: 10.3345/cep.2020.00290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lei C., Fu W., Qian K., Li T., Zhang S., Fu W., Ding M., Hu S. Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat Commun. 2020;11:2070. doi: 10.1038/s41467-020-16048-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lei L., Qian H., Yang X., Zhou X., Zhang X., Zhang D., Dai T., Guo R., Shi L., Cheng Y. The phenotypic changes of γδ T cells in COVID-19 patients. medRxiv. 2020 doi: 10.1101/2020.04.05.20046433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine M.M. Monoclonal Antibody Therapy for Ebola Virus Disease. N. Engl. J. Med. 2019;381:2365–2366. doi: 10.1056/NEJMe1915350. [DOI] [PubMed] [Google Scholar]
- Li C.K.-F., Wu H., Yan H., Ma S., Wang L., Zhang M., Tang X., Temperton N.J., Weiss R.A., Brenchley J.M. T cell responses to whole SARS coronavirus in humans. J. Immunol. 2008;181:5490–5500. doi: 10.4049/jimmunol.181.8.5490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J., Guo M., Tian X., Liu C., Wang X., Yang X., Wu P., Xiao Z., Qu Y., Yin Y. Virus-host interactome and proteomic survey of PMBCs from COVID-19 patients reveal potential virulence factors influencing SARS-CoV-2 pathogenesis. bioRxiv. 2020 doi: 10.1101/2020.03.31.019216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li K., Chen D., Chen S., Feng Y., Chang C., Wang Z., Wang N., Zhen G. Radiographic Findings and other Predictors in Adults with Covid-19. medRxiv. 2020 doi: 10.1101/2020.03.23.20041673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Xie Z., Lin W., Cai W., Wen C., Guan Y., Mo X., Wang J., Wang Y., Peng P. An exploratory randomized controlled study on the efficacy and safety of lopinavir/ritonavir or arbidol treating adult patients hospitalized with mild/moderate COVID-19 (ELACOI) medRxiv. 2020 doi: 10.1101/2020.03.19.20038984. [DOI] [Google Scholar]
- Li Y., Wang H., Tang X., Ma D., Du C., Wang Y., Pan H. Potential host range of multiple SARS-like coronaviruses and an improved ACE2-Fc variant that is potent against both SARS-CoV-2 and SARS-CoV-1. bioRxiv. 2020 doi: 10.1101/2020.04.10.032342. [DOI] [Google Scholar]
- Liao M., Liu Y., Yuan J., Wen Y., Xu G., Zhao J., Cheng L., Li J., Wang X., Wang F. Single-cell Landscape of Bronchoalveolar Immune Cells in Patients With COVID-19. Nat. Med. 2020 doi: 10.1038/s41591-020-0901-9. Published online May 12, 2020. [DOI] [PubMed] [Google Scholar]
- Lin J.T., Zhang J.S., Su N., Xu J.G., Wang N., Chen J.T., Chen X., Liu Y.X., Gao H., Jia Y.P. Safety and immunogenicity from a phase I trial of inactivated severe acute respiratory syndrome coronavirus vaccine. Antivir. Ther. (Lond.) 2007;12:1107–1113. [PubMed] [Google Scholar]
- Lindesmith L., Moe C., Marionneau S., Ruvoen N., Jiang X., Lindblad L., Stewart P., LePendu J., Baric R. Human susceptibility and resistance to Norwalk virus infection. Nat. Med. 2003;9:548–553. doi: 10.1038/nm860. [DOI] [PubMed] [Google Scholar]
- Liu W., Fontanet A., Zhang P.H., Zhan L., Xin Z.T., Baril L., Tang F., Lv H., Cao W.C. Two-year prospective study of the humoral immune response of patients with severe acute respiratory syndrome. J. Infect. Dis. 2006;193:792–795. doi: 10.1086/500469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L., Wei Q., Lin Q., Fang J., Wang H., Kwok H., Tang H., Nishiura K., Peng J., Tan Z. Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight. 2019;4:e123158. doi: 10.1172/jci.insight.123158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu B., Han J., Cheng X., Yu L., Zhang L., Wang W., Ni L., Wei C., Huang Y., Cheng Z. Persistent SARS-CoV-2 presence is companied with defects in adaptive immune system in non-severe COVID-19 patients. medRxiv. 2020 doi: 10.1101/2020.03.26.20044768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J., Li S., Liu J., Liang B., Wang X., Wang H., Li W., Tong Q., Yi J., Zhao L. Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS-CoV-2 infected patients. EBioMedicine. 2020;55:102763. doi: 10.1016/j.ebiom.2020.102763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J., Liu Y., Xiang P., Pu L., Xiong H., Li C., Zhang M., Tan J., Xu Y., Song R. Neutrophil-to-lymphocyte ratio predicts critical illness patients with 2019 coronavirus disease in the early stage. J. Transl. Med. 2020;18:206. doi: 10.1186/s12967-020-02374-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J., Cao R., Xu M., Wang X., Zhang H., Hu H., Li Y., Hu Z., Zhong W., Wang M. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov. 2020;6:16. doi: 10.1038/s41421-020-0156-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q., Fang X., Tokuno S., Chung U., Chen X., Dai X., Liu X., Xu F., Wang B., Peng P. Prediction of the clinical outcome of COVID-19 patients using T lymphocyte subsets with 340 cases from Wuhan, China: a retrospective cohort study and a web visualization tool. medRxiv. 2020 doi: 10.1101/2020.04.06.20056127. [DOI] [Google Scholar]
- Liu T., Zhang J., Yang Y., Ma H., Li Z., Zhang J., Cheng J., Zhang X., Zhao Y., Xia Z. The role of interleukin-6 in monitoring severe case of coronavirus disease 2019. EMBO Mol. Med. 2020 doi: 10.15252/emmm.202012421. Published online May 19, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y., Li J., Liu D., Song H., Chen C., Lv M., Pei X., Hu Z. Clinical features and outcomes of 2019 novel coronavirus-infected patients with cardiac injury. medRxiv. 2020 doi: 10.1101/2020.03.11.20030957. [DOI] [Google Scholar]
- Liu Y., Sun W., Guo Y., Chen L., Zhang L., Zhao S., Long D., Yu L. Association between platelet parameters and mortality in coronavirus disease 2019: Retrospective cohort study. Platelets. 2020:1–7. doi: 10.1080/09537104.2020.1754383. Published online April 16, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo B., Zhang K., Lu W., Zheng L., Zhang Q., Kanellopoulou C., Zhang Y., Liu Z., Fritz J.M., Marsh R. AUTOIMMUNE DISEASE. Patients with LRBA deficiency show CTLA4 loss and immune dysregulation responsive to abatacept therapy. Science. 2015;349:436–440. doi: 10.1126/science.aaa1663. [DOI] [PubMed] [Google Scholar]
- Lokugamage K.G., Narayanan K., Nakagawa K., Terasaki K., Ramirez S.I., Tseng C.-T.K., Makino S. Middle East Respiratory Syndrome Coronavirus nsp1 Inhibits Host Gene Expression by Selectively Targeting mRNAs Transcribed in the Nucleus while Sparing mRNAs of Cytoplasmic Origin. J. Virol. 2015;89:10970–10981. doi: 10.1128/JVI.01352-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lokugamage K.G., Hage A., Schindewolf C., Rajsbaum R., Menachery V.D. SARS-CoV-2 is sensitive to type I interferon pretreatment. bioRxiv. 2020 doi: 10.1101/2020.03.07.982264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lon J.R., Bai Y., Zhong B., Cai F., Du H. Prediction and Evolution of B Cell Epitopes of Surface Protein in SARS-CoV-2. bioRxiv. 2020 doi: 10.1101/2020.04.03.022723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lou B., Li T., Zheng S., Su Y., Li Z., Liu W., Yu F., Ge S., Zou Q., Yuan Q. Serology characteristics of SARS-CoV-2 infection since the exposure and post symptoms onset. medRxiv. 2020 doi: 10.1101/2020.03.23.20041707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lover A.A. Quantifying treatment effects of hydroxychloroquine and azithromycin for COVID-19: a secondary analysis of an open label non-randomized clinical trial (Gautret et al, 2020) medRxiv. 2020 doi: 10.1101/2020.03.22.20040949. [DOI] [Google Scholar]
- Lu R., Zhao X., Li J., Niu P., Yang B., Wu H., Wang W., Song H., Huang B., Zhu N. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565–574. doi: 10.1016/S0140-6736(20)30251-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu X., Chen T., Wang Y., Wang J., Zhang B., Li Y., Yan F. Adjuvant corticosteroid therapy for critically ill patients with COVID-19. medRxiv. 2020 doi: 10.1101/2020.04.07.20056390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luke T.C., Kilbane E.M., Jackson J.L., Hoffman S.L. Meta-analysis: convalescent blood products for Spanish influenza pneumonia: a future H5N1 treatment? Ann. Intern. Med. 2006;145:599–609. doi: 10.7326/0003-4819-145-8-200610170-00139. [DOI] [PubMed] [Google Scholar]
- Mantlo E.K., Bukreyeva N., Maruyama J., Paessler S., Huang C. Antiviral Activities of Type I Interferons to SARS-CoV-2 Infection. Antiviral Res. 2020 doi: 10.1016/j.antiviral.2020.104811. Published online April 29, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mao H., Tu W., Qin G., Law H.K., Sia S.F., Chan P.L., Liu Y., Lam K.T., Zheng J., Peiris M., Lau Y.L. Influenza virus directly infects human natural killer cells and induces cell apoptosis. J. Virol. 2009;83:9215–9222. doi: 10.1128/JVI.00805-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marquardt N., Kekäläinen E., Chen P., Kvedaraite E., Wilson J.N., Ivarsson M.A., Mjösberg J., Berglin L., Säfholm J., Manson M.L. Human Lung Natural Killer Cells Are Predominantly Comprised of Highly Differentiated Hypofunctional CD69 − CD56 dim Cells. J. Allergy Clin. Immunol. 2017;139:1321–1330. doi: 10.1016/j.jaci.2016.07.043. [DOI] [PubMed] [Google Scholar]
- Martin J.E., Louder M.K., Holman L.A., Gordon I.J., Enama M.E., Larkin B.D., Andrews C.A., Vogel L., Koup R.A., Roederer M., VRC 301 Study Team A SARS DNA vaccine induces neutralizing antibody and cellular immune responses in healthy adults in a Phase I clinical trial. Vaccine. 2008;26:6338–6343. doi: 10.1016/j.vaccine.2008.09.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medzhitov R., Schneider D.S., Soares M.P. Disease tolerance as a defense strategy. Science. 2012;335:936–941. doi: 10.1126/science.1214935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehta P., McAuley D.F., Brown M., Sanchez E., Tattersall R.S., Manson J.J., HLH Across Speciality Collaboration, UK COVID-19: consider cytokine storm syndromes and immunosuppression. Lancet. 2020;395:1033–1034. doi: 10.1016/S0140-6736(20)30628-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Menachery V.D., Mitchell H.D., Cockrell A.S., Gralinski L.E., Yount B.L., Jr., Graham R.L., McAnarney E.T., Douglas M.G., Scobey T., Beall A. MERS-CoV accessory orfs play key role for infection and pathogenesis. MBio. 2017;8:1–14. doi: 10.1128/mBio.00665-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milewska A., Chi Y., Szczepanski A., Barreto-Duran E., Liu K., Liu D., Guo X., Ge Y., Li J., Cui L. HTCC as a highly effective polymeric inhibitor of SARS-CoV-2 and MERS-CoV. bioRxiv. 2020 doi: 10.1101/2020.03.29.014183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minakshi R., Padhan K., Rani M., Khan N., Ahmad F., Jameel S. The SARS Coronavirus 3a protein causes endoplasmic reticulum stress and induces ligand-independent downregulation of the type 1 interferon receptor. PLoS ONE. 2009;4:e8342. doi: 10.1371/journal.pone.0008342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Modjarrad K., Roberts C.C., Mills K.T., Castellano A.R., Paolino K., Muthumani K., Reuschel E.L., Robb M.L., Racine T., Oh M.D. Safety and immunogenicity of an anti-Middle East respiratory syndrome coronavirus DNA vaccine: a phase 1, open-label, single-arm, dose-escalation trial. Lancet Infect. Dis. 2019;19:1013–1022. doi: 10.1016/S1473-3099(19)30266-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molina J.M., Delaugerre C., Le Goff J., Mela-Lima B., Ponscarme D., Goldwirt L., de Castro N. No evidence of rapid antiviral clearance or clinical benefit with the combination of hydroxychloroquine and azithromycin in patients with severe COVID-19 infection. Med. Mal. Infect. 2020 doi: 10.1016/j.medmal.2020.03.006. S0399-077X(20)30085-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monticelli L.A., Sonnenberg G.F., Abt M.C., Alenghat T., Ziegler C.G., Doering T.A., Angelosanto J.M., Laidlaw B.J., Yang C.Y., Sathaliyawala T. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 2011;12:1045–1054. doi: 10.1031/ni.2131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore J.B., June C.H. Cytokine release syndrome in severe COVID-19. Science. 2020;368:473–474. doi: 10.1126/science.abb8925. [DOI] [PubMed] [Google Scholar]
- Nailwal H., Chan F.K. Necroptosis in anti-viral inflammation. Cell Death Differ. 2019;26:4–13. doi: 10.1038/s41418-018-0172-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ng O.-W., Chia A., Tan A.T., Jadi R.S., Leong H.N., Bertoletti A., Tan Y.-J. Memory T cell responses targeting the SARS coronavirus persist up to 11 years post-infection. Vaccine. 2016;34:2008–2014. doi: 10.1016/j.vaccine.2016.02.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni L., Ye F., Chen M.-L., Feng Y., Deng Y.-Q., Zhao H., Wei P., Ge J., Gou M., Li X. Detection of SARS-CoV-2-Specific Humoral and Cellular Immunity in COVID-19 Convalescent Individuals. Immunity. 2020 doi: 10.1016/j.immuni.2020.04.023. Published online May 3, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nie J., Li Q., Wu J., Zhao C., Hao H., Liu H., Zhang L., Nie L., Qin H., Wang M. Establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2. Emerg. Microbes Infect. 2020;9:680–686. doi: 10.1080/22221751.2020.1743767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nie S., Zhao X., Zhao K., Zhang Z., Zhang Z., Zhang Z. Metabolic disturbances and inflammatory dysfunction predict severity of coronavirus disease 2019 (COVID-19): a retrospective study. medRxiv. 2020 doi: 10.1101/2020.03.24.20042283. [DOI] [Google Scholar]
- Niemeyer D., Zillinger T., Muth D., Zielecki F., Horvath G., Suliman T., Barchet W., Weber F., Drosten C., Müller M.A. Middle East respiratory syndrome coronavirus accessory protein 4a is a type I interferon antagonist. J. Virol. 2013;87:12489–12495. doi: 10.1128/JVI.01845-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nieto-Torres J.L., Verdiá-Báguena C., Jimenez-Guardeño J.M., Regla-Nava J.A., Castaño-Rodriguez C., Fernandez-Delgado R., Torres J., Aguilella V.M., Enjuanes L. Severe acute respiratory syndrome coronavirus E protein transports calcium ions and activates the NLRP3 inflammasome. Virology. 2015;485:330–339. doi: 10.1016/j.virol.2015.08.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikolich-Žugich J. The twilight of immunity: emerging concepts in aging of the immune system. Nat. Immunol. 2018;19:10–19. doi: 10.1038/s41590-017-0006-x. [DOI] [PubMed] [Google Scholar]
- Okabayashi T., Kariwa H., Yokota S., Iki S., Indoh T., Yokosawa N., Takashima I., Tsutsumi H., Fujii N. Cytokine regulation in SARS coronavirus infection compared to other respiratory virus infections. J. Med. Virol. 2006;78:417–424. doi: 10.1002/jmv.20556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okba N.M.A., Müller M.A., Li W., Wang C., GeurtsvanKessel C.H., Corman V.M., Lamers M.M., Sikkema R.S., de Bruin E., Chandler F.D. Severe Acute Respiratory Syndrome Coronavirus 2-Specific Antibody Responses in Coronavirus Disease 2019 Patients. Emerg. Infect. Dis. 2020;26 doi: 10.3201/eid2607.200841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ong E.Z., Chan Y.F.Z., Leong W.Y., Lee N.M.Y., Kalimuddin S., Haja Mohideen S.M., Chan K.S., Tan A.T., Bertoletti A., Ooi E.E., Low J.G.H. A dynamic immune response shapes COVID-19 progression. Cell Host Microbe. 2020 doi: 10.1016/j.chom.2020.03.021. S1931-3128(20)30185-2. Published online April 30, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Page C., Goicochea L., Matthews K., Zhang Y., Klover P., Holtzman M.J., Hennighausen L., Frieman M. Induction of alternatively activated macrophages enhances pathogenesis during severe acute respiratory syndrome coronavirus infection. J. Virol. 2012;86:13334–13349. doi: 10.1128/JVI.01689-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan X., Zhou P., Fan T., Wu Y., Zhang J., Shi X., Shang W., Fang L., Jiang X., Shi J. Immunoglobulin fragment F(ab’)2 against RBD potently neutralizes SARS-CoV-2 in vitro. bioRxiv. 2020 doi: 10.1101/2020.04.07.029884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pardi N., Hogan M.J., Pelc R.S., Muramatsu H., Andersen H., DeMaso C.R., Dowd K.A., Sutherland L.L., Scearce R.M., Parks R. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature. 2017;543:248–251. doi: 10.1038/nature21428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park T., Lee S.-Y., Kim S., Kim M.J., Kim H.G., Jun S., Kim S.I., Kim B.T., Park E.C., Park D. Spike protein binding prediction with neutralizing antibodies of SARS-CoV-2. bioRxiv. 2020 doi: 10.1101/2020.02.22.951178. [DOI] [Google Scholar]
- Payne D.C., Iblan I., Rha B., Alqasrawi S., Haddadin A., Al Nsour M., Alsanouri T., Ali S.S., Harcourt J., Miao C. Persistence of Antibodies against Middle East Respiratory Syndrome Coronavirus. Emerg. Infect. Dis. 2016;22:1824–1826. doi: 10.3201/eid2210.160706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pei S., Yuan X., Zhang Z., Yao R., Xie Y., Shen M., Li B., Chen X., Yin M. Convalescent Plasma to Treat COVID-19: Chinese Strategy and Experiences. medRxiv. 2020 doi: 10.1101/2020.04.07.20056440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlman S., Dandekar A.A. Immunopathogenesis of coronavirus infections: implications for SARS. Nat. Rev. Immunol. 2005;5:917–927. doi: 10.1038/nri1732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfaender S., Mar K.B., Michailidis E., Kratzel A., Hirt D., V’kovski P., Fan W., Ebert N., Stalder H., Kleine-Weber H. LY6E impairs coronavirus fusion and confers immune control of viral disease. bioRxiv. 2020 doi: 10.1101/2020.03.05.979260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pinto D., Park Y.-J., Beltramello M., Walls A.C., Tortorici M.A., Bianchi S., Jaconi S., Culap K., Zatta F., De Marco A. Structural and functional analysis of a potent sarbecovirus neutralizing antibody. bioRxiv. 2020 doi: 10.1101/2020.04.07.023903. [DOI] [Google Scholar]
- Poor H.D., Ventetuolo C.E., Tolbert T., Chun G., Serrao G., Zeidman A., Dangayach N.S., Olin J., Kohli-Seth R., Powell C.A. COVID-19 Critical Illness Pathophysiology Driven by Diffuse Pulmonary Thrombi and Pulmonary Endothelial Dysfunction Responsive to Thrombolysis. medRxiv. 2020 doi: 10.1101/2020.04.17.20057125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prokunina-Olsson L., Alphonse N., Dickenson R.E., Durbin J.E., Glenn J.S., Hartmann R., Kotenko S.V., Lazear H.M., O’Brien T.R., Odendall C. COVID-19 and emerging viral infections: The case for interferon lambda. J. Exp. Med. 2020;217:e20200653. doi: 10.1084/jem.20200653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin C., Zhou L., Hu Z., Zhang S., Yang S., Tao Y., Xie C., Ma K., Shang K., Wang W. Dysregulation of Immune Response in Patients with COVID-19 in Wuhan, China. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa248. Published online March 12, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu M., Shi Y., Guo Z., Chen Z., He R., Chen R., Zhou D., Dai E., Wang X., Si B. Antibody responses to individual proteins of SARS coronavirus and their neutralization activities. Microbes Infect. 2005;7:882–889. doi: 10.1016/j.micinf.2005.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qu R., Ling Y., Zhang Y.-H.-Z., Wei L.-Y., Chen X., Li X.-M., Liu X.-Y., Liu H.-M., Guo Z., Ren H. Platelet-to-lymphocyte Ratio Is Associated With Prognosis in Patients With Coronavirus disease-19. J. Med. Virol. 2020 doi: 10.1002/jmv.25767. Published online March 17, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quinlan B.D., Mou H., Zhang L., Guo Y., He W., Ojha A., Parcells M.S., Luo G., Li W., Zhong G. The SARS-CoV-2 receptor-binding domain elicits a potent neutralizing response without antibody-dependent enhancement. bioRxiv. 2020 doi: 10.1101/2020.04.10.036418. [DOI] [Google Scholar]
- Rabouw H.H., Langereis M.A., Knaap R.C.M., Dalebout T.J., Canton J., Sola I., Enjuanes L., Bredenbeek P.J., Kikkert M., de Groot R.J., van Kuppeveld F.J. Middle East Respiratory Coronavirus Accessory Protein 4a Inhibits PKR-Mediated Antiviral Stress Responses. PLoS Pathog. 2016;12:e1005982. doi: 10.1371/journal.ppat.1005982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramsuran V., Naranbhai V., Horowitz A., Qi Y., Martin M.P., Yuki Y., Gao X., Walker-Sperling V., Del Prete G.Q., Schneider D.K. Elevated HLA-A expression impairs HIV control through inhibition of NKG2A-expressing cells. Science. 2018;359:86–90. doi: 10.1126/science.aam8825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reed S.E. The behaviour of recent isolates of human respiratory coronavirus in vitro and in volunteers: evidence of heterogeneity among 229E-related strains. J. Med. Virol. 1984;13:179–192. doi: 10.1002/jmv.1890130208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renieri A., Benetti E., Tita R., Spiga O., Ciolfi A., Birolo G., Bruselles A., Doddato G., Giliberti A., Marconi C. ACE2 variants underlie interindividual variability and susceptibility to COVID-19 in Italian population. medRxiv. 2020 doi: 10.1101/2020.04.03.20047977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roumier M., Paule R., Vallee A., Ackermann F. Interleukin-6 blockade for severe COVID-19. medRxiv. 2020 doi: 10.1101/2020.04.20.20061861. [DOI] [Google Scholar]
- Sanders J.M., Monogue M.L., Jodlowski T.Z., Cutrell J.B. Pharmacologic Treatments for Coronavirus Disease 2019 (COVID-19): A Review. JAMA. 2020 doi: 10.1001/jama.2020.6019. Published online April 13, 2020. [DOI] [PubMed] [Google Scholar]
- Savarino A., Boelaert J.R., Cassone A., Majori G., Cauda R. Effects of chloroquine on viral infections: an old drug against today’s diseases? Lancet Infect. Dis. 2003;3:722–727. doi: 10.1016/S1473-3099(03)00806-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schrezenmeier E., Dörner T. Mechanisms of action of hydroxychloroquine and chloroquine: implications for rheumatology. Nat. Rev. Rheumatol. 2020;16:155–166. doi: 10.1038/s41584-020-0372-x. [DOI] [PubMed] [Google Scholar]
- Shamshirian A., Hessami A., Heydari K., Alizadeh-Navaei R., Ebrahimzadeh M.A., Ghasemian R., Aboufazeli E., Baradaran H., Karimifar K., Eftekhari A. Hydroxychloroquine Versus COVID-19: A Rapid Systematic Review and Meta-Analysis. medRxiv. 2020 doi: 10.1101/2020.04.14.20065276. [DOI] [Google Scholar]
- Shang L., Zhao J., Hu Y., Du R., Cao B. On the use of corticosteroids for 2019-nCoV pneumonia. Lancet. 2020;395:683–684. doi: 10.1016/S0140-6736(20)30361-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shao Z., Feng Y., Zhong L., Xie Q., Lei M., Liu Z., Wang C., Ji J., Li W., Liu H. Clinical Efficacy of Intravenous Immunoglobulin Therapy in Critical Patients with COVID-19: A multicenter retrospective cohort study. medRxiv. 2020 doi: 10.1101/2020.04.11.20061739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheahan T.P., Sims A.C., Graham R.L., Menachery V.D., Gralinski L.E., Case J.B., Leist S.R., Pyrc K., Feng J.Y., Trantcheva I. ). Broad-Spectrum Antiviral GS-5734 Inhibits Both Epidemic and Zoonotic Coronaviruses. Sci. Transl. Med. 2017;9:eaal3653. doi: 10.1126/scitranslmed.aal3653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheahan T.P., Sims A.C., Leist S.R., Schäfer A., Won J., Brown A.J., Montgomery S.A., Hogg A., Babusis D., Clarke M.O. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat. Commun. 2020;11:222. doi: 10.1038/s41467-019-13940-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen C., Wang Z., Zhao F., Yang Y., Li J., Yuan J., Wang F., Li D., Yang M., Xing L. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA. 2020;323:1582–1589. doi: 10.1001/jama.2020.4783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi C.-S., Qi H.-Y., Boularan C., Huang N.-N., Abu-Asab M., Shelhamer J.H., Kehrl J.H. SARS-coronavirus open reading frame-9b suppresses innate immunity by targeting mitochondria and the MAVS/TRAF3/TRAF6 signalosome. J. Immunol. 2014;193:3080–3089. doi: 10.4049/jimmunol.1303196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi C.-S., Nabar N.R., Huang N.N., Kehrl J.H. SARS-Coronavirus Open Reading Frame-8b triggers intracellular stress pathways and activates NLRP3 inflammasomes. Cell Death Discov. 2019;5:101. doi: 10.1038/s41420-019-0181-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi S., Qin M., Shen B., Cai Y., Liu T., Yang F., Gong W., Liu X., Liang J., Zhao Q. Association of Cardiac Injury With Mortality in Hospitalized Patients With COVID-19 in Wuhan, China. JAMA Cardiol. 2020 doi: 10.1001/jamacardio.2020.0950. Published online March 25, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiow L.R., Rosen D.B., Brdicková N., Xu Y., An J., Lanier L.L., Cyster J.G., Matloubian M. CD69 acts downstream of interferon-alpha/beta to inhibit S1P1 and lymphocyte egress from lymphoid organs. Nature. 2006;440:540–544. doi: 10.1038/nature04606. [DOI] [PubMed] [Google Scholar]
- Siu K.L., Kok K.H., Ng M.H.J., Poon V.K.M., Yuen K.Y., Zheng B.J., Jin D.Y. Severe acute respiratory syndrome coronavirus M protein inhibits type I interferon production by impeding the formation of TRAF3.TANK.TBK1/IKKepsilon complex. J. Biol. Chem. 2009;284:16202–16209. doi: 10.1074/jbc.M109.008227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siu K.L., Yeung M.L., Kok K.H., Yuen K.S., Kew C., Lui P.Y., Chan C.P., Tse H., Woo P.C.Y., Yuen K.Y., Jin D.Y. Middle east respiratory syndrome coronavirus 4a protein is a double-stranded RNA-binding protein that suppresses PACT-induced activation of RIG-I and MDA5 in the innate antiviral response. J. Virol. 2014;88:4866–4876. doi: 10.1128/JVI.03649-13. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Siu K.L., Yuen K.S., Castaño-Rodriguez C., Ye Z.W., Yeung M.L., Fung S.Y., Yuan S., Chan C.P., Yuen K.Y., Enjuanes L., Jin D.Y. Severe acute respiratory syndrome coronavirus ORF3a protein activates the NLRP3 inflammasome by promoting TRAF3-dependent ubiquitination of ASC. FASEB J. 2019;33:8865–8877. doi: 10.1096/fj.201802418R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sivori S., Vacca P., Del Zotto G., Munari E., Mingari M.C., Moretta L. Human NK cells: surface receptors, inhibitory checkpoints, and translational applications. Cell. Mol. Immunol. 2019;16:430–441. doi: 10.1038/s41423-019-0206-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song Z., Xu Y., Bao L., Zhang L., Yu P., Qu Y., Zhu H., Zhao W., Han Y., Qin C. From SARS to MERS, Thrusting Coronaviruses into the Spotlight. Viruses. 2019;11:59. doi: 10.3390/v11010059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song C.-Y., Xu J., He J.-Q., Lu Y.-Q. COVID-19 early warning score: a multi-parameter screening tool to identify highly suspected patients. medRxiv. 2020 doi: 10.1101/2020.03.05.20031906. [DOI] [Google Scholar]
- Stanifer M.L., Kee C., Cortese M., Triana S., Mukenhirn M., Kraeusslich H.G., Alexandrov T., Bartenschlager R., Boulant S. Critical role of type III interferon in controlling SARS-CoV-2 infection, replication and spread in primary human intestinal epithelial cells. bioRxiv. 2020 doi: 10.1101/2020.04.24.059667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stawiski E.W., Diwanji D., Suryamohan K., Gupta R., Fellouse F.A., Sathirapongsasuti J.F., Liu J., Jiang Y.-P., Ratan A., Mis M. Human ACE2 receptor polymorphisms predict SARS-CoV-2 susceptibility. bioRxiv. 2020 doi: 10.1101/2020.04.07.024752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sui J., Li W., Murakami A., Tamin A., Matthews L.J., Wong S.K., Moore M.J., Tallarico A.S.C., Olurinde M., Choe H. Potent neutralization of severe acute respiratory syndrome (SARS) coronavirus by a human mAb to S1 protein that blocks receptor association. Proc. Natl. Acad. Sci. USA. 2004;101:2536–2541. doi: 10.1073/pnas.0307140101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sui J., Li W., Roberts A., Matthews L.J., Murakami A., Vogel L., Wong S.K., Subbarao K., Farzan M., Marasco W.A. Evaluation of human monoclonal antibody 80R for immunoprophylaxis of severe acute respiratory syndrome by an animal study, epitope mapping, and analysis of spike variants. J. Virol. 2005;79:5900–5906. doi: 10.1128/JVI.79.10.5900-5906.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun L., Xing Y., Chen X., Zheng Y., Yang Y., Nichols D.B., Clementz M.A., Banach B.S., Li K., Baker S.C., Chen Z. Coronavirus papain-like proteases negatively regulate antiviral innate immune response through disruption of STING-mediated signaling. PLoS ONE. 2012;7:e30802. doi: 10.1371/journal.pone.0030802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun C., Luo C., Zhang Y., Zhang J., Yang J., Chen L., Yang J., Li J., Xie L. SARS-CoV-2 and SARS-CoV Spike-RBD Structure and Receptor Binding Comparison and Potential Implications on Neutralizing Antibody and Vaccine Development. bioRxiv. 2020 doi: 10.1101/2020.02.16.951723. [DOI] [Google Scholar]
- Tai W., He L., Zhang X., Pu J., Voronin D., Jiang S., Zhou Y., Du L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 2020 doi: 10.1038/s41423-020-0400-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan L., Wang Q., Zhang D., Ding J., Huang Q., Tang Y.-Q., Wang Q., Miao H. Lymphopenia predicts disease severity of COVID-19: a descriptive and predictive study. Signal Transduct. Target. Ther. 2020;5:33. doi: 10.1038/s41392-020-0148-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan W., Lu Y., Zhang J., Wang J., Dan Y., Tan Z., He X., Qian C., Sun Q., Hu Q. Viral Kinetics and Antibody Responses in Patients with COVID-19. medRxiv. 2020 doi: 10.1101/2020.03.24.20042382. [DOI] [Google Scholar]
- Tanaka T., Narazaki M., Kishimoto T. Immunotherapeutic implications of IL-6 blockade for cytokine storm. Immunotherapy. 2016;8:959–970. doi: 10.2217/imt-2016-0020. [DOI] [PubMed] [Google Scholar]
- Tang F., Quan Y., Xin Z.T., Wrammert J., Ma M.J., Lv H., Wang T.B., Yang H., Richardus J.H., Liu W., Cao W.C. Lack of peripheral memory B cell responses in recovered patients with severe acute respiratory syndrome: a six-year follow-up study. J. Immunol. 2011;186:7264–7268. doi: 10.4049/jimmunol.0903490. [DOI] [PubMed] [Google Scholar]
- Tang N., Li D., Wang X., Sun Z. Abnormal coagulation parameters are associated with poor prognosis in patients with novel coronavirus pneumonia. J. Thromb. Haemost. 2020;18:844–847. doi: 10.1111/jth.14768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang W., Cao Z., Han M., Wang Z., Chen J., Sun W., Wu Y., Xiao W., Liu S., Chen E. Hydroxychloroquine in patients with COVID-19: an open-label, randomized, controlled trial. medRxiv. 2020 doi: 10.1101/2020.04.10.20060558. [DOI] [Google Scholar]
- Tanne J.H. Covid-19: FDA approves use of convalescent plasma to treat critically ill patients. BMJ. 2020;368:m1256. doi: 10.1136/bmj.m1256. [DOI] [PubMed] [Google Scholar]
- Taylor A., Foo S.-S., Bruzzone R., Dinh L.V., King N.J.C., Mahalingam S. Fc receptors in antibody-dependent enhancement of viral infections. Immunol. Rev. 2015;268:340–364. doi: 10.1111/imr.12367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ter Meulen J., Bakker A.B.H., van den Brink E.N., Weverling G.J., Martina B.E.E., Haagmans B.L., Kuiken T., de Kruif J., Preiser W., Spaan W. Human monoclonal antibody as prophylaxis for SARS coronavirus infection in ferrets. Lancet. 2004;363:2139–2141. doi: 10.1016/S0140-6736(04)16506-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terpos E., Ntanasis-Stathopoulos I., Elalamy I., Kastritis E., Sergentanis T.N., Politou M., Psaltopoulou T., Gerotziafas G., Dimopoulos M.A. Hematological findings and complications of COVID-19. Am. J. Hematol. 2020 doi: 10.1002/ajh.25829. Published online April 13, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tetro J.A. Is COVID-19 receiving ADE from other coronaviruses? Microbes Infect. 2020;22:72–73. doi: 10.1016/j.micinf.2020.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thanh Le T., Andreadakis Z., Kumar A., Gómez Román R., Tollefsen S., Saville M., Mayhew S. The COVID-19 vaccine development landscape. Nat. Rev. Drug Discov. 2020;19:305–306. doi: 10.1038/d41573-020-00073-5. [DOI] [PubMed] [Google Scholar]
- Thevarajan I., Nguyen T.H.O., Koutsakos M., Druce J., Caly L., van de Sandt C.E., Jia X., Nicholson S., Catton M., Cowie B. Breadth of Concomitant Immune Responses Prior to Patient Recovery: A Case Report of Non-Severe COVID-19. Nat. Med. 2020;26:453–455. doi: 10.1038/s41591-020-0819-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomé R., Lopes S.C.P., Costa F.T.M., Verinaud L. Chloroquine: modes of action of an undervalued drug. Immunol. Lett. 2013;153:50–57. doi: 10.1016/j.imlet.2013.07.004. [DOI] [PubMed] [Google Scholar]
- Thomé R., Moraes A.S., Bombeiro A.L., Farias A. dos S., Francelin C., da Costa T.A., Di Gangi R., dos Santos L.M.B., de Oliveira A.L.R., Verinaud L. Chloroquine treatment enhances regulatory T cells and reduces the severity of experimental autoimmune encephalomyelitis. PLoS ONE. 2013;8:e65913. doi: 10.1371/journal.pone.0065913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thornbrough J.M., Jha B.K., Yount B., Goldstein S.A., Li Y., Elliott R., Sims A.C., Baric R.S., Silverman R.H., Weiss S.R. Middle east respiratory syndrome coronavirus NS4b protein inhibits host RNase L activation. MBio. 2016;7:e00258. doi: 10.1128/mBio.00258-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian S., Xiong Y., Liu H., Niu L., Guo J., Liao M., Xiao S.-Y. Pathological study of the 2019 novel coronavirus disease (COVID-19) through postmortem core biopsies. Mod. Pathol. 2020 doi: 10.1038/s41379-020-0536-x. Published online April 14, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian X., Li C., Huang A., Xia S., Lu S., Shi Z., Lu L., Jiang S., Yang Z., Wu Y., Ying T. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 2020;9:382–385. doi: 10.1080/22221751.2020.1729069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- To K.K.-W., Tsang O.T.-Y., Leung W.-S., Tam A.R., Wu T.-C., Lung D.C., Yip C.C.-Y., Cai J.-P., Chan J.M.-C., Chik T.S.-H. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect. Dis. 2020;20:565–574. doi: 10.1016/S1473-3099(20)30196-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Traggiai E., Becker S., Subbarao K., Kolesnikova L., Uematsu Y., Gismondo M.R., Murphy B.R., Rappuoli R., Lanzavecchia A. An efficient method to make human monoclonal antibodies from memory B cells: potent neutralization of SARS coronavirus. Nat. Med. 2004;10:871–875. doi: 10.1038/nm1080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Travaglini K.J., Nabhan A.N., Penland L., Sinha R., Gillich A., Sit R.V., Chang S., Conley S.D., Mori Y., Seita J. A molecular cell atlas of the human lung from single cell RNA sequencing. bioRxiv. 2020 doi: 10.1101/742320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trimble C.L., Morrow M.P., Kraynyak K.A., Shen X., Dallas M., Yan J., Edwards L., Parker R.L., Denny L., Giffear M. Safety, efficacy, and immunogenicity of VGX-3100, a therapeutic synthetic DNA vaccine targeting human papillomavirus 16 and 18 E6 and E7 proteins for cervical intraepithelial neoplasia 2/3: a randomised, double-blind, placebo-controlled phase 2b trial. Lancet. 2015;386:2078–2088. doi: 10.1016/S0140-6736(15)00239-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vabret N., Samstein R., Fernandez N., Merad M., Sinai Immunology Review Project. Trainees. Faculty Advancing scientific knowledge in times of pandemics. Nat. Rev. Immunol. 2020 doi: 10.1038/s41577-020-0319-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varga Z., Flammer A.J., Steiger P., Haberecker M., Andermatt R., Zinkernagel A.S., Mehra M.R., Schuepbach R.A., Ruschitzka F., Moch H. Endothelial Cell Infection and Endotheliitis in COVID-19. Lancet. 2020;395:1417–1418. doi: 10.1016/S0140-6736(20)30937-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vincent M.J., Bergeron E., Benjannet S., Erickson B.R., Rollin P.E., Ksiazek T.G., Seidah N.G., Nichol S.T. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol. J. 2005;2:69. doi: 10.1186/1743-422X-2-69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vivier E., Artis D., Colonna M., Diefenbach A., Di Santo J.P., Eberl G., Koyasu S., Locksley R.M., McKenzie A.N.J., Mebius R.E. Innate Lymphoid Cells: 10 Years On. Cell. 2018;174:1054–1066. doi: 10.1016/j.cell.2018.07.017. [DOI] [PubMed] [Google Scholar]
- Von Holle T.A., Moody M.A. Influenza and Antibody-Dependent Cellular Cytotoxicity. Front. Immunol. 2019;10:1457. doi: 10.3389/fimmu.2019.01457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walls A.C., Park Y.-J., Tortorici M.A., Wall A., McGuire A.T., Veesler D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell. 2020;181:281–292.e6. doi: 10.1016/j.cell.2020.02.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walter J.M., Helmin K.A., Abdala-Valencia H., Wunderink R.G., Singer B.D. Multidimensional assessment of alveolar T cells in critically ill patients. JCI Insight. 2018;3:e123287. doi: 10.1172/jci.insight.123287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wan S., Yi Q., Fan S., Lv J., Zhang X., Guo L., Lang C., Xiao Q., Xiao K., Yi Z. Characteristics of lymphocyte subsets and cytokines in peripheral blood of 123 hospitalized patients with 2019 novel coronavirus pneumonia (NCP) medRxiv. 2020 doi: 10.1101/2020.02.10.20021832. [DOI] [Google Scholar]
- Wan Y., Shang J., Graham R., Baric R.S., Li F. Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J. Virol. 2020;94:e00127-20. doi: 10.1128/JVI.00127-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wan Y., Shang J., Sun S., Tai W., Chen J., Geng Q., He L., Chen Y., Wu J., Shi Z. Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry. J. Virol. 2020;94:e02015–e02019. doi: 10.1128/JVI.02015-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L. C-reactive Protein Levels in the Early Stage of COVID-19. Med. Mal. Infect. 2020 doi: 10.1016/j.medmal.2020.03.007. Published online March 31, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang C.-H., Liu C.-Y., Wan Y.-L., Chou C.-L., Huang K.-H., Lin H.-C., Lin S.-M., Lin T.-Y., Chung K.F., Kuo H.-P. Persistence of lung inflammation and lung cytokines with high-resolution CT abnormalities during recovery from SARS. Respir. Res. 2005;6:42. doi: 10.1186/1465-9921-6-42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S.F., Chen K.H., Chen M., Li W.Y., Chen Y.J., Tsao C.H., Yen M.Y., Huang J.C., Chen Y.M. Human-leukocyte antigen class I Cw 1502 and class II DR 0301 genotypes are associated with resistance to severe acute respiratory syndrome (SARS) infection. Viral Immunol. 2011;24:421–426. doi: 10.1089/vim.2011.0024. [DOI] [PubMed] [Google Scholar]
- Wang C., Li W., Drabek D., Okba N.M.A., van Haperen R., Osterhaus A.D.M.E., van Kuppeveld F.J.M., Haagmans B.L., Grosveld F., Bosch B.-J. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat. Commun. 2020 doi: 10.1038/s41467-020-16256-y. Published online May 4, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D., Hu B., Hu C., Zhu F., Liu X., Zhang J., Wang B., Xiang H., Cheng Z., Xiong Y. JAMA; China: 2020. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D., Wang J., Jiang Q., Yang J., Li J., Gao C., Jiang H., Ge L., Liu Y. No Clear Benefit to the Use of Corticosteroid as Treatment in Adult Patients with Coronavirus Disease 2019: A Retrospective Cohort Study. medRxiv. 2020 doi: 10.1101/2020.04.21.20066258. [DOI] [Google Scholar]
- Wang F., Nie J., Wang H., Zhao Q., Xiong Y., Deng L., Song S., Ma Z., Mo P., Zhang Y. Characteristics of peripheral lymphocyte subset alteration in COVID-19 pneumonia. J. Infect. Dis. 2020;221:1762–1769. doi: 10.1093/infdis/jiaa150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang M., Cao R., Zhang L., Yang X., Liu J., Xu M., Shi Z., Hu Z., Zhong W., Xiao G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30:269–271. doi: 10.1038/s41422-020-0282-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W., He J., Lie P., Huang L., Wu S., Lin Y., Liu X. The definition and risks of Cytokine Release Syndrome-Like in 11 COVID-19-Infected Pneumonia critically ill patients: Disease Characteristics and Retrospective Analysis. medRxiv. 2020 doi: 10.1101/2020.02.26.20026989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Zhang D., Du G., Du R., Zhao J., Jin Y., Fu S., Gao L., Cheng Z., Lu Q. Remdesivir in adults with severe COVID-19: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet. 2020 doi: 10.1016/S0140-6736(20)31022-9. Published online April 29, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Jiang W., He Q., Wang C., Wang B., Zhou P., Dong N., Tong Q. Early, low-dose and short-term application of corticosteroid treatment in patients with severe COVID-19 pneumonia: single-center experience from Wuhan, China. medRxiv. 2020 doi: 10.1101/2020.03.06.20032342. [DOI] [Google Scholar]
- Wathelet M.G., Orr M., Frieman M.B., Baric R.S. Severe acute respiratory syndrome coronavirus evades antiviral signaling: role of nsp1 and rational design of an attenuated strain. J. Virol. 2007;81:11620–11633. doi: 10.1128/JVI.00702-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiskopf D., Schmitz K.S., Raadsen M.P., Grifoni A., Okba N.M.A., Endeman H., van den Akker J.P.C., Molenkamp R., Koopmans M.P.G., van Gorp E.C.M. Phenotype of SARS-CoV-2-specific T-cells in COVID-19 patients with acute respiratory distress syndrome. medRxiv. 2020 doi: 10.1101/2020.04.11.20062349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen W., Su W., Tang H., Le W., Zhang X., Zheng Y., Liu X., Xie L., Li J., Ye J. Immune cell profiling of COVID-19 patients in the recovery stage by single-cell sequencing. Cell Discov. 2020 doi: 10.1038/s41421-020-0168-9. Published online May 4, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilk A.J., Rustagi A., Zhao N.Q., Roque J., Martinez-Colon G.J., McKechnie J.L., Ivison G.T., Ranganath T., Vergara R., Hollis T. A single-cell atlas of the peripheral immune response to severe COVID-19. medRxiv. 2020 doi: 10.1101/2020.04.17.20069930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williamson B., Feldmann F., Schwarz B., Meade-White K., Porter D., Schulz J., van Doremalen N., Leighton I., Yinda C.K., Perez-Perez L. Clinical benefit of remdesivir in rhesus macaques infected with SARS-CoV-2. bioRxiv. 2020 doi: 10.1101/2020.04.15.043166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wölfel R., Corman V.M., Guggemos W., Seilmaier M., Zange S., Müller M.A., Niemeyer D., Jones T.C., Vollmar P., Rothe C. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020 doi: 10.1038/s41586-020-2196-x. Published online April 1, 2020. [DOI] [PubMed] [Google Scholar]
- Wong C.K., Lam C.W.K., Wu A.K.L., Ip W.K., Lee N.L.S., Chan I.H.S., Lit L.C.W., Hui D.S.C., Chan M.H.M., Chung S.S.C., Sung J.J. Plasma inflammatory cytokines and chemokines in severe acute respiratory syndrome. Clin. Exp. Immunol. 2004;136:95–103. doi: 10.1111/j.1365-2249.2004.02415.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- World Health Organization . 2020. Clinical management of severe acute respiratory infection (SARI) when COVID-19 disease is suspected: Interim Guidance.https://www.who.int/publications-detail/clinical-management-of-severe-acute-respiratory-infection-when-novel-coronavirus-(ncov)-infection-is-suspected [Google Scholar]
- World Health Organization . 2020. Draft of the landscape of COVID-19 candidate vaccines.https://www.who.int/who-documents-detail/draft-landscape-of-covid-19-candidate-vaccines [Google Scholar]
- Wrapp D., De Vlieger D., Corbett K.S., Torres G.M., Wang N., Van Breedam W., Roose K., van Schie L., Hoffmann M., Pöhlmann S., VIB-CMB COVID-19 Response Team Structural Basis for Potent Neutralization of Betacoronaviruses by Single-Domain Camelid Antibodies. Cell. 2020;181 doi: 10.1016/j.cell.2020.04.031. Published online May 5, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C., Chen X., Cai Y., Xia J., Zhou X., Xu S., Huang H., Zhang L., Zhou X., Du C. Risk Factors Associated With Acute Respiratory Distress Syndrome and Death in Patients With Coronavirus Disease 2019 Pneumonia in Wuhan, China. JAMA Intern. Med. 2020 doi: 10.1001/jamainternmed.2020.0994. Published online March 13, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu F., Wang A., Liu M., Wang Q., Chen J., Xia S., Ling Y., Zhang Y., Xun J., Lu L. Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. medRxiv. 2020 doi: 10.1101/2020.03.30.20047365. [DOI] [Google Scholar]
- Wu Y., Li C., Xia S., Tian X., Kong Y., Wang Z., Gu C., Zhang R., Tu C., Xie Y. Identification of Human Single-Domain Antibodies against SARS-CoV-2. Cell Host Microbe. 2020 doi: 10.1016/j.chom.2020.04.023. Published online May 14, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wynants L., Van Calster B., Bonten M.M.J., Collins G.S., Debray T.P.A., De Vos M., Haller M.C., Heinze G., Moons K.G.M., Riley R.D. Prediction models for diagnosis and prognosis of covid-19 infection: systematic review and critical appraisal. BMJ. 2020;369:m1328. doi: 10.1136/bmj.m1328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia C.Q., Xu L.L., Wang Z., Qin Z.Q., Tong Z.H., Huang K.W., Xiao B., Qi M., Jiang B.Z., Wang C. The involvement of natural killer cells in the pathogenesis of severe acute respiratory syndrome. Am. J. Clin. Pathol. 2004;121:507–511. doi: 10.1309/WPK7Y2XKNF4CBF3R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiang J., Wen J., Yuan X., Xiong S., Zhou Xue., Liu C., Min Xun. Potential biochemical markers to identify severe cases among COVID-19 patients. medRxiv. 2020 doi: 10.1101/2020.03.19.20034447. [DOI] [Google Scholar]
- Xiang J., Yan M., Li H., Liu T., Lin C., Huang S., Shen C. Evaluation of Enzyme-Linked Immunoassay and Colloidal Gold- Immunochromatographic Assay Kit for Detection of Novel Coronavirus (SARS-Cov-2) Causing an Outbreak of Pneumonia (COVID-19) medRxiv. 2020 doi: 10.1101/2020.02.27.20028787. [DOI] [Google Scholar]
- Xiong R., Zhang L., Li S., Sun Y., Ding M., Wang Y., Zhao Y., Wu Y., Shang W., Jiang X. Novel and potent inhibitors targeting DHODH, a rate-limiting enzyme in de novo pyrimidine biosynthesis, are broad-spectrum antiviral against RNA viruses including newly emerged coronavirus SARS-CoV-2. bioRxiv. 2020 doi: 10.1101/2020.03.11.983056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu H., Hou K., Xu H., Li Z., Chen H., Zhang N., Xu R., Fu H., Sun R., Wen L. Acute Myocardial Injury of Patients with Coronavirus Disease 2019. medRxiv. 2020 doi: 10.1101/2020.03.05.20031591. [DOI] [Google Scholar]
- Xu X., Han M., Li T., Sun W., Wang D., Fu B., Zhou Y., Zheng X., Yang Y., Li X. Effective treatment of severe COVID-19 patients with tocilizumab. Proc. Natl. Acad. Sci. USA. 2020 doi: 10.1073/pnas.2005615117. 202005615. Published online April 29, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Z., Shi L., Wang Y., Zhang J., Huang L., Zhang C., Liu S., Zhao P., Liu H., Zhu L. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020;8:420–422. doi: 10.1016/S2213-2600(20)30076-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto N., Matsuyama S., Hoshino T., Yamamoto N. Nelfinavir inhibits replication of severe acute respiratory syndrome coronavirus 2 in vitro. bioRxiv. 2020 doi: 10.1101/2020.04.06.026476. [DOI] [Google Scholar]
- Yan D., Liu X.-Y., Zhu Y.-n., Huang L., Dan B.-t., Zhang G.-j., Gao Y.-h. Factors associated with prolonged viral shedding and impact of Lopinavir/Ritonavir treatment in patients with SARS-CoV-2 infection. Eur. Respir. J. 2020 doi: 10.1183/13993003.00799-2020. Published online May 19, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan L., Zhang H.-T., Xiao Y., Wang M., Sun C., Liang J., Li S., Zhang M., Guo Y., Xiao Y. Prediction of criticality in patients with severe Covid-19 infection using three clinical features: a machine learning-based prognostic model with clinical data in Wuhan. medRxiv. 2020 doi: 10.1101/2020.02.27.20028027. [DOI] [Google Scholar]
- Yang Y., Ye F., Zhu N., Wang W., Deng Y., Zhao Z., Tan W. Middle East respiratory syndrome coronavirus ORF4b protein inhibits type I interferon production through both cytoplasmic and nuclear targets. Sci. Rep. 2015;5:17554. doi: 10.1038/srep17554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang X., Dai T., Zhou X., Qian H., Guo R., Lei L., Zhang X., Zhang D., Shi L., Cheng Y. Analysis of adaptive immune cell populations and phenotypes in the patients infected by SARS-CoV-2. medRxiv. 2020 doi: 10.1101/2020.03.23.20040675. [DOI] [Google Scholar]
- Yang Y., Shen C., Li J., Yuan J., Wei J., Huang F., Wang F., Li G., Li Y., Xing L. Plasma IP-10 and MCP-3 levels are highly associated with disease severity and predict the progression of COVID-19. J. Allergy Clin. Immunol. 2020 doi: 10.1016/j.jaci.2020.04.027. Published online April 29, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Z., Liu J., Zhou Y., Zhao X., Zhao Q., Liu J. The effect of corticosteroid treatment on patients with coronavirus infection: a systematic review and meta-analysis. J. Infect. 2020 doi: 10.1016/j.jinf.2020.03.062. S0163-4453(20)30191-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao X., Ye F., Zhang M., Cui C., Huang B., Niu P., Liu X., Zhao L., Dong E., Song C. In Vitro Antiviral Activity and Projection of Optimized Dosing Design of Hydroxychloroquine for the Treatment of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa237. Published online March 9, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao X.H., Li T.Y., He Z.C., Ping Y.F., Liu H.W., Yu S.C., Mou H.M., Wang L.H., Zhang H.R., Fu W.J. [A pathological report of three COVID-19 cases by minimally invasive autopsies] Zhonghua Bing Li Xue Za Zhi. 2020;49:E009. doi: 10.3760/cma.j.cn112151-20200312-00193. [DOI] [PubMed] [Google Scholar]
- Ying T., Li H., Lu L., Dimitrov D.S., Jiang S. Development of human neutralizing monoclonal antibodies for prevention and therapy of MERS-CoV infections. Microbes Infect. 2015;17:142–148. doi: 10.1016/j.micinf.2014.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yip M.S., Leung N.H.L., Cheung C.Y., Li P.H., Lee H.H.Y., Daëron M., Peiris J.S.M., Bruzzone R., Jaume M. Antibody-dependent infection of human macrophages by severe acute respiratory syndrome coronavirus. Virol. J. 2014;11:82. doi: 10.1186/1743-422X-11-82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu L., Tong Y., Shen G., Fu A., Lai Y., Zhou X., Yuan Y. ). Immunodepletion with Hypoxemia: A Potential High Risk Subtype of Coronavirus Disease 2019. medRxiv. 2020 doi: 10.1101/2020.03.03.20030650. [DOI] [Google Scholar]
- Yuan M., Wu N.C., Zhu X., Lee C.-C.D., So R.T.Y., Lv H., Mok C.K.P., Wilson I.A. A Highly Conserved Cryptic Epitope in the Receptor Binding Domains of SARS-CoV-2 and SARS-CoV. Science. 2020;868:630–633. doi: 10.1126/science.abb7269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yudanin N.A., Schmitz F., Flamar A.-L., Thome J.J.C., Tait Wojno E., Moeller J.B., Schirmer M., Latorre I.J., Xavier R.J., Farber D.L. Spatial and Temporal Mapping of Human Innate Lymphoid Cells Reveals Elements of Tissue Specificity. Immunity. 2019;50:505–519.e4. doi: 10.1016/j.immuni.2019.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng Q., Li Y.-Z., Huang G., Wu W., Dong S.-Y., Xu Y. Mortality of COVID-19 is Associated with Cellular Immune Function Compared to Immune Function in Chinese Han Population. medRxiv. 2020 doi: 10.1101/2020.03.08.20031229. [DOI] [Google Scholar]
- Zha L., Li S., Pan L., Tefsen B., Li Y., French N., Chen L., Yang G., Villanueva E.V. Corticosteroid Treatment of Patients with Coronavirus Disease 2019 (COVID-19) Med. J. Aust. 2020 doi: 10.5694/mja2.50577. Published online April 8, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L., Zhang F., Yu W., He T., Yu J., Yi C.E., Ba L., Li W., Farzan M., Chen Z. Antibody responses against SARS coronavirus are correlated with disease outcome of infected individuals. J. Med. Virol. 2006;78:1–8. doi: 10.1002/jmv.20499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang B., Zhou X., Zhu C., Feng F., Qiu Y., Feng J., Jia Q., Song Q., Zhu B., Wang J. Immune phenotyping based on neutrophil-to-lymphocyte ratio and IgG predicts disease severity and outcome for patients with COVID-19. medRxiv. 2020 doi: 10.1101/2020.03.12.20035048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang B., Liu S., Tan T., Huang W., Dong Y., Chen L., Chen Q., Zhang L., Zhong Q., Zhang X. Treatment With Convalescent Plasma for Critically Ill Patients With SARS-CoV-2 Infection. Chest. 2020 doi: 10.1016/j.chest.2020.03.039. Published online March 31, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang D., Guo R., Lei L., Liu H., Wang Y., Wang Y., Dai T., Zhang T., Lai Y., Wang J. COVID-19 infection induces readily detectable morphological and inflammation-related phenotypic changes in peripheral blood monocytes, the severity of which correlate with patient outcome. medRxiv. 2020 doi: 10.1101/2020.03.24.20042655. [DOI] [Google Scholar]
- Zhang J., Liu J., Li N., Liu Y., Ye R., Qin X., Zheng R. Serological detection of 2019-nCoV respond to the epidemic: A useful complement to nucleic acid testing. medRxiv. 2020 doi: 10.1101/2020.03.04.20030916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J., Zhao J., Van Rooijen N., Perlman S. Evasion by stealth: inefficient immune activation underlies poor T cell response and severe disease in SARS-CoV-infected mice. PLoS Pathog. 2009;5:e1000636. doi: 10.1371/journal.ppat.1000636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao K., Wang H., Wu C. The immune responses of HLA-A∗0201 restricted SARS-CoV S peptide-specific CD8+ T cells are augmented in varying degrees by CpG ODN, PolyI:C and R848. Vaccine. 2011;29:6670–6678. doi: 10.1016/j.vaccine.2011.06.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X., Guo F., Liu F., Cuconati A., Chang J., Block T.M., Guo J.-T. Interferon induction of IFITM proteins promotes infection by human coronavirus OC43. Proc. Natl. Acad. Sci. USA. 2014;111:6756–6761. doi: 10.1073/pnas.1320856111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X., Sehgal M., Hou Z., Cheng J., Shu S., Wu S., Guo F., Le Marchand S.J., Lin H., Chang J., Guo J.T. Identification of Residues Controlling Restriction versus Enhancing Activities of IFITM Proteins on Entry of Human Coronaviruses. J. Virol. 2018;92:e01535-17. doi: 10.1128/JVI.01535-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J., Yuan Q., Wang H., Liu W., Liao X., Su Y., Wang X., Yuan J., Li T., Li J. Antibody Responses to SARS-CoV-2 in Patients of Novel Coronavirus Disease 2019. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa344. Published online March 28, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J., Yang Y., Huang H., Li D., Gu D., Lu X., Zhang Z., Liu L., Liu T., Liu Y. Relationship between the ABO Blood Group and the COVID-19 Susceptibility. medRxiv. 2020 doi: 10.1101/2020.03.11.20031096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X., Zheng S., Chen D., Zheng M., Li X., Li G., Lin H., Chang J., Zeng H., Guo J.-T. LY6E Restricts the Entry of Human Coronaviruses, including the currently pandemic SARS-CoV-2. bioRxiv. 2020 doi: 10.1101/2020.04.02.021469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X., Zhang B., Li P., Ma C., Gu J., Hou P., Guo Z., Wu H., Bai Y. Incidence, clinical characteristics and prognostic factor of patients with COVID-19: a systematic review and meta-analysis. medRxiv. 2020 doi: 10.1101/2020.03.17.20037572. [DOI] [Google Scholar]
- Zheng M., Song L. Novel antibody epitopes dominate the antigenicity of spike glycoprotein in SARS-CoV-2 compared to SARS-CoV. Cell. Mol. Immunol. 2020;17:536–538. doi: 10.1038/s41423-020-0385-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng J., Liu Y., Lau Y.L., Tu W. γδ-T cells: an unpolished sword in human anti-infection immunity. Cell. Mol. Immunol. 2013;10:50–57. doi: 10.1038/cmi.2012.43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng H.-Y., Zhang M., Yang C.-X., Zhang N., Wang X.-C., Yang X.-P., Dong X.-Q., Zheng Y.-T. Elevated exhaustion levels and reduced functional diversity of T cells in peripheral blood may predict severe progression in COVID-19 patients. Cell. Mol. Immunol. 2020;17:541–543. doi: 10.1038/s41423-020-0401-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng M., Gao Y., Wang G., Song G., Liu S., Sun D., Xu Y., Tian Z. Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cell. Mol. Immunol. 2020;17:533–535. doi: 10.1038/s41423-020-0402-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J., Chu H., Li C., Wong B.H.-Y., Cheng Z.-S., Poon V.K.-M., Sun T., Lau C.C.-Y., Wong K.K.-Y., Chan J.Y.-W. Active replication of Middle East respiratory syndrome coronavirus and aberrant induction of inflammatory cytokines and chemokines in human macrophages: implications for pathogenesis. J. Infect. Dis. 2014;209:1331–1342. doi: 10.1093/infdis/jit504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou Y., Jiang S., Du L. Prospects for a MERS-CoV spike vaccine. Expert Rev. Vaccines. 2018;17:677–686. doi: 10.1080/14760584.2018.1506702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou F., Yu T., Du R., Fan G., Liu Y., Liu Z., Xiang J., Wang Y., Song B., Gu X. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395:1054–1062. doi: 10.1016/S0140-6736(20)30566-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou Y., Fu B., Zheng X., Wang D., Zhao C., Qi Y., Sun R., Tian Z., Xu X., Wei H. Aberrant pathogenic GM-CSF+ T cells and inflammatory CD14+CD16+ monocytes in severe pulmonary syndrome patients of a new coronavirus. bioRxiv. 2020 doi: 10.1101/2020.02.12.945576. [DOI] [Google Scholar]
- Zhou Y., Fu B., Zheng X., Wang D., Zhao C., Qi Y., Sun R., Tian Z., Xu X., Wei H. Pathogenic T cells and inflammatory monocytes incite inflammatory storm in severe COVID-19 patients. Natl. Sci. Rev. 2020 doi: 10.1093/nsr/nwaa041. Published online March 13, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou Y., Yang Z., Guo Y., Geng S., Gao S., Ye S., Hu Y., Wang Y. A New Predictor of Disease Severity in Patients with COVID-19 in Wuhan, China. medRxiv. 2020 doi: 10.1101/2020.03.24.20042119. [DOI] [Google Scholar]
- Zhu Z., Chakraborti S., He Y., Roberts A., Sheahan T., Xiao X., Hensley L.E., Prabakaran P., Rockx B., Sidorov I.A. Potent cross-reactive neutralization of SARS coronavirus isolates by human monoclonal antibodies. Proc. Natl. Acad. Sci. USA. 2007;104:12123–12128. doi: 10.1073/pnas.0701000104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu F.C., Hou L.H., Li J.X., Wu S.P., Liu P., Zhang G.R., Hu Y.M., Meng F.Y., Xu J.J., Tang R. Safety and immunogenicity of a novel recombinant adenovirus type-5 vector-based Ebola vaccine in healthy adults in China: preliminary report of a randomised, double-blind, placebo-controlled, phase 1 trial. Lancet. 2015;385:2272–2279. doi: 10.1016/S0140-6736(15)60553-0. [DOI] [PubMed] [Google Scholar]
- Zhu F.-C., Wurie A.H., Hou L.-H., Liang Q., Li Y.-H., Russell J.B.W., Wu S.-P., Li J.-X., Hu Y.-M., Guo Q. Safety and immunogenicity of a recombinant adenovirus type-5 vector-based Ebola vaccine in healthy adults in Sierra Leone: a single-centre, randomised, double-blind, placebo-controlled, phase 2 trial. Lancet. 2017;389:621–628. doi: 10.1016/S0140-6736(16)32617-4. [DOI] [PubMed] [Google Scholar]
- Ziegler C.J.K., Allon S.J., Nyquist S.K., Mbano I.M., Miao V.N., Tzouanas C.N., Cao Y., Yousif A.S., Bals J., Hauser B.M. SARS-CoV-2 Receptor ACE2 Is an Interferon-Stimulated Gene in Human Airway Epithelial Cells and Is Detected in Specific Cell Subsets Across Tissues. Cell. 2020 doi: 10.1016/j.cell.2020.04.035. Published online April 27, 2020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zietz M., Tatonetti N.P. Testing the association between blood type and COVID-19 infection, intubation, and death. medRxiv. 2020 doi: 10.1101/2020.04.08.20058073. [DOI] [PMC free article] [PubMed] [Google Scholar]