With great interest we read the article by Gaiser et al 1 that enrichment of oral bacterial taxa in pancreatic cancer highlights the role of oral microbiota. Not coincidentally, the presence of Fusobacterium nucleatum in paired saliva and colon samples of the patients with colorectal cancer has been reported,2 3 raising interest in whether disease starts in the mouth or in the intestine.4–6 Another reason people are interested in oral microbes is their potentials serving as biomarkers for systemic diseases.3 7–9
In this study, we examined how long the collapsed bacterial community can recover to its initial state when suffering from disturbance and whether oral microbes have sufficient robustness to serve as biomarkers. We longitudinally tracked the re-assembling process of human oral biofilms after clinical scaling. Paired saliva and dental plaque samples were collected from nine subjects at 11 time points (figure 1A and online supplementary figure S1). The 16S rRNA V3–V4 regions of 169 samples were amplified and sequenced, and the generated reads were analysed using QIIME.10
Figure 1.
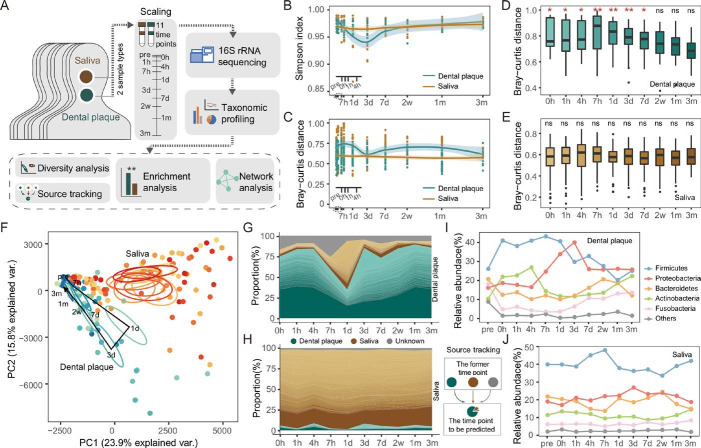
Longitudinal dynamics of in vivo human oral microbiota. (A) Overview of the study design and sample collection. (B) The alpha diversity of the dental plaque and salivary microbiota over time. The shadow around the line shows a 95% CI. (C) The Bray-Curtis distance calculated at the operational taxonomic unit (OTU) level across individual microbiota of the same time point. (D–E) The Bray-Curtis distance between microbiota of each time point and pre. Significance was measured using Wilcoxon rank-sum test (0.01>p>0.001, **; 0.05>p>0.01, *; p>0.05, ns). (F) The principal component (PC) analysis of dental plaque and saliva microbiota at each time point. The circle shows a 10% CI. (G–H) Tracing the source of OTUs in each time point. The dental plaque and saliva samples of the former time point were taken as potential sources of the latter time point. The bands of each colour indicate the top 100 most abundant OTUs. Green, brown and grey bands represent the sources of dental plaque, salivary and unknown, respectively. (I–J) The temporal changes of bacterial abundance at the phylum level.
gutjnl-2019-318977supp001.pdf (133.3KB, pdf)
The microbial diversity of dental plaques fluctuated notably over time after destruction. In contrast, there was little change in salivary samples, where the differences among individuals at the same time point were relatively small (figure 1B and online supplementary figure S2). Likewise, a wave trough appeared in dental plaque at 3 days, where the Bray-Curtis dissimilarity among different individuals was the smallest (figure 1C). At the time points of before scaling (pre) and the very early stage (0–4 hours), the differences among plaque samples were obviously larger than those of free-floating saliva. The largest community distances to the original biofilm appeared between 7 hours and 3 days rather than 0–4 hours right after scaling (figure 1D). The plaque microbiota before 7 days was significantly different from pre (Wilcoxon rank-sum test, p<0.05), while the salivary microbiota remained constant throughout the study (figure 1E).
The unweighted principal component analysis clearly illustrated the dynamic process that plaque microbiota greatly deviated from the original state (pre) to the biofilm deconstruction after 1 day, rebuilt after 3 days and gradually recovered to its original intact form over time (figure 1F). In contrast, the temporal changes of saliva samples were not obvious. To track the microbial replenishment process, plaque and saliva samples at the former time point served as the potential sources to predict the origin of microbial operational taxonomic units in the latter time point. We found that the microbial transfer from saliva to plaque mainly happened between 1 day and 3 days (figure 1G–H). At the time points between 7 hours and 3 days, almost all of the bacterial phyla were at their highest or lowest abundances (figure 1I). The bacterial abundance of saliva, however, only slightly fluctuated at certain time points and then returned to their original level (figure 1J).
The longitudinal gradient samples of the dental plaque were grouped into three clusters which represents the early, middle and mature phases of oral biofilm, respectively (figure 2A). When comparing each time point with pre, the change patterns of significantly changed genera (Wilcoxon test with false discovery rate correction, p<0.05) were diverse within each phase. The bacteria recovered in the early phase represented the first colonisers with quick feedback; the bacteria that underwent significant changes in the middle phase are secondary colonisers, interacting with the primary colonisers and paving the foundation for the successors; and the bacteria that reproduced continuously in the three phases but had not completely recovered at the end were members of long-term changes (online supplementary figure S3 and figure 2B). The bacteria that have undergone significant variations in the middle phase had the highest number of edges and positive correlations in the co-occurrence network analysis (r≥0.7 and p≤0.05), indicating that they may play a key role in the development of dental biofilm (figure 2C).
Figure 2.
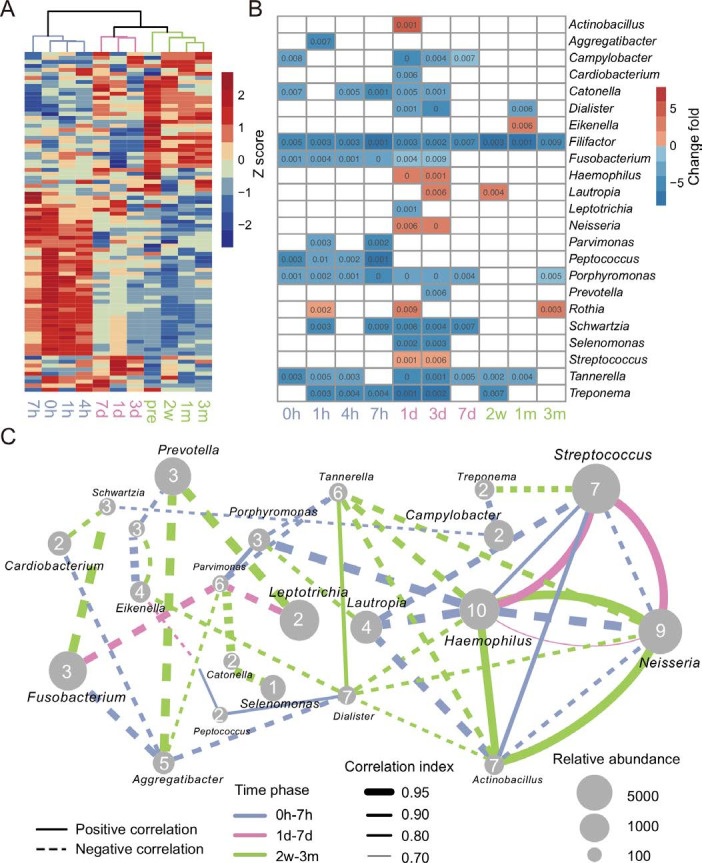
Microbial taxonomy and interaction of oral biofilms over time. (A) Clustering of the dental plaque microbiota at the genus level. (B) Significantly different genera of each time point compared with the time point pre. Wilcoxon rank-sum test with false discovery rate correction, p≤0.01. (C) Co-occurrence network of the significantly different genera. The thresholds of SparCC correlations were r≥0.7 and p≤0.05. The blue, pink and green lines represent the connection of two genera in the early phase (0 hour, 1 hour, 4 hours, 7 hours), the medium phase (1 day, 3 days, 7 days) and the late phase (2 weeks, 1 month, 3 months, pre), respectively. The dotted and solid lines indicate negative correlation and positive correlation, respectively. The thickness of the line is proportional to the value of correlation. The size of the nodes is proportional to their relative abundance, and the number marked on each node represents the degree of this node.
As the gateway and source of microbial down-transmission to the GI tract, the oral cavity is in a vital position. Our study uncovered the recovery and long-term stability of the oral microbiota after strong disturbance, and identified the key time points and phases of the most dramatic community changes and structural recovery. These findings suggest that when considering the use of oral bacteria as biomarkers to predict digestive system diseases, collection time and site should be taken into consideration. This study provides an opportunity to evaluate whether oral bacteria are suitable for disease prediction and promotes the development of non-invasive diagnostic techniques.
Acknowledgments
We thank Yifei Zhang and Haibo Zhang for sampling assistance, and the subjects for providing specimens to this study.
Footnotes
JW, ZJ and BZ contributed equally.
Contributors: FZ conceived the study. FZ and JW designed the study and prepared the manuscript. JW, ZJ and LP collected the samples and conducted the experiments. JW, ZJ, BZ and FZ analysed the data. All authors approved the final version of the manuscript.
Funding: This work was supported by grants from the National Natural Science Foundation of China (31722031, 31670119, 31870107), National Key R&D Program (2016YFC1200804) and the Key Research Program of the Chinese Academy of Sciences (KFZD-SW-219).
Competing interests: None declared.
Ethics approval: Beijing Friendship Hospital (2017-P-178-02).
Provenance and peer review: Not commissioned; externally peer reviewed.
Patient consent for publication: Not required.
References
- 1. Gaiser RA, Halimi A, Alkharaan H, et al. . Enrichment of oral microbiota in early cystic precursors to invasive pancreatic cancer. Gut 2019;68:2186–94. 10.1136/gutjnl-2018-317458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Komiya Y, Shimomura Y, Higurashi T, et al. . Patients with colorectal cancer have identical strains of Fusobacterium nucleatum in their colorectal cancer and oral cavity. Gut 2019;69:1335–7. 10.1136/gutjnl-2018-316661 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Flemer B, Warren RD, Barrett MP, et al. . The oral microbiota in colorectal cancer is distinctive and predictive. Gut 2018;67:1454–63. 10.1136/gutjnl-2017-314814 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Schmidt TS, Hayward MR, Coelho LP, et al. . Extensive transmission of microbes along the gastrointestinal tract. Elife 2019;8 10.7554/eLife.42693 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Thomas AM, Manghi P, Asnicar F, et al. . Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation. Nat Med 2019;25:667–78. 10.1038/s41591-019-0405-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Atarashi K, Suda W, Luo C, et al. . Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 2017;358:359–65. 10.1126/science.aan4526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Farrell JJ, Zhang L, Zhou H, et al. . Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 2012;61:582–8. 10.1136/gutjnl-2011-300784 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Wang J, Zheng J, Shi W, et al. . Dysbiosis of maternal and neonatal microbiota associated with gestational diabetes mellitus. Gut 2018;67:1614–25. 10.1136/gutjnl-2018-315988 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Fan X, Alekseyenko AV, Wu J, et al. . Human oral microbiome and prospective risk for pancreatic cancer: a population-based nested case–control study. Gut 2018;67:120–7. 10.1136/gutjnl-2016-312580 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Caporaso JG, Kuczynski J, Stombaugh J, et al. . QIIME allows analysis of high-throughput community sequencing data. Nat Methods 2010;7:335–6. 10.1038/nmeth.f.303 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
gutjnl-2019-318977supp001.pdf (133.3KB, pdf)