Abstract
Pectin-rich agro-industrial residues are feedstocks with potential for sustainable biorefineries. They are generated in high amounts worldwide from the industrial processing of fruits and vegetables. The challenges posed to the industrial implementation of efficient bioprocesses are however manyfold and thoroughly discussed in this review paper, mainly at the biological level. The most important yeast cell factory platform for advanced biorefineries is currently Saccharomyces cerevisiae, but this yeast species cannot naturally catabolise the main sugars present in pectin-rich agro-industrial residues hydrolysates, in particular d-galacturonic acid and l-arabinose. However, there are non-Saccharomyces species (non-conventional yeasts) considered advantageous alternatives whenever they can express highly interesting metabolic pathways, natively assimilate a wider range of carbon sources or exhibit higher tolerance to relevant bioprocess-related stresses. For this reason, the interest in non-conventional yeasts for biomass-based biorefineries is gaining momentum. This review paper focuses on the valorisation of pectin-rich residues by exploring the potential of yeasts that exhibit vast metabolic versatility for the efficient use of the carbon substrates present in their hydrolysates and high robustness to cope with the multiple stresses encountered. The major challenges and the progresses made related with the isolation, selection, sugar catabolism, metabolic engineering and use of non-conventional yeasts and S. cerevisiae-derived strains for the bioconversion of pectin-rich residue hydrolysates are discussed. The reported examples of value-added products synthesised by different yeasts using pectin-rich residues are reviewed.
|
Key Points • Review of the challenges and progresses made on the bioconversion of pectin-rich residues by yeasts. • Catabolic pathways for the main carbon sources present in pectin-rich residues hydrolysates. • Multiple stresses with potential to affect bioconversion productivity. • Yeast metabolic engineering to improve pectin-rich residues bioconversion. |

Graphical abstract
Keywords: Pectin-rich agro-industrial residues, Non-conventional yeasts, Bioconversion, Metabolic engineering, Biorefinery, Circular bioeconomy
Introduction
Agro-industrial residues are currently in the spotlight of research and development activities worldwide; they are raw materials for the biotechnology industry, as renewable sources of carbon, nitrogen and other nutrients for microbial growth and metabolite production (Cherubini 2010; Liguori and Faraco 2016; Liu et al. 2016; Dahiya et al. 2018). The utilisation of organic waste residues as substrates to produce added-value products is environmentally friendly strategies by saving and reutilizing resources. The implementation of a circular bioeconomy based on microorganisms, in particular non-conventional (non-Saccharomyces) yeast strains with metabolic versatility and tolerance to bioprocesses-related stresses, is an important societal challenge (Leandro et al. 2006; Fletcher et al. 2016; Cristobal-Sarramian and Atzmuller 2018; Zuin et al. 2018; Rebello et al. 2018; Nielsen 2019).
Agro-industrial residues were derived from sugary materials (e.g. sugar beet, sugarcane or fruits and vegetables) and starchy feedstocks (e.g. wheat, corn, rice or potatoes) and lignocellulosic substrates (e.g. wood, straw and grasses) (Balat 2011). Pectin-rich agricultural residues and agro-food industry residues are potential feedstocks for the production of biofuels and other relevant bioproducts (Schmitz et al. 2019). Currently, a large fraction of the pectin-rich residues (e.g. sugar beet pulp and citrus peel) are dried for further use as cattle feed or put in landfills for soil improvement, although it is desirable to find new ways to convert these residues into renewable chemicals using natural or engineered microbes (Richard and Hilditch 2009; Ajila et al. 2012). The residues with the highest pectin content (sugar beet pulp, citrus peels, and apple pomace) are accumulated in high amounts worldwide from the sugar industry or the industrial processing of fruits and vegetables (Peters 2006; Balat 2011). These residues are partially pre-treated during sugar (from sugar beets) and juice (from fruits) extraction and have low lignin content which facilitates processing (Berlowska et al. 2018). Despite the difficulties inherent to the high variability of these feedstocks due to diverse geographical distribution and seasonality, they are cheap and abundant (Peters 2006; Balat 2011). However, they are interesting feedstocks for microbial fermentations, as the enzymatic hydrolysis of their component polysaccharides can be economically accomplished to yield fermentable neutral sugars (hexoses and pentoses) and d-galacturonic acid (d-GalA) (Leijdekkers et al. 2013; Cárdenas-Fernández et al. 2017; de la Torre et al. 2019).
Saccharomyces cerevisiae is currently, and by far, the most important yeast cell factory in the biotechnology industry and the major cell factory platform for the production of bioethanol and other biofuels and chemicals in advanced biorefineries (Satyanarayana and Kunze 2009; de Jong et al. 2012; Hong and Nielsen 2012; Nielsen 2019). Endogenously, S. cerevisiae can only use a very limited range of carbon sources. For this reason, genetically modified strains have been developed to also utilise pentoses and d-galacturonic acid for synthesis of novel compounds (Hong and Nielsen 2012; Benz et al. 2014; Biz et al. 2016; Yaguchi et al. 2018; Rebello et al. 2018; Protzko et al. 2018; Nielsen 2019). However, there are non-conventional species considered advantageous alternatives to S. cerevisiae since they can express highly interesting metabolic pathways (Rebello et al. 2018), efficiently assimilate a wider range of carbon sources (Do et al. 2019) or exhibit higher tolerance to relevant bioprocess-related stresses, such as the presence of a wide range of inhibitory compounds and supraoptimal temperatures (Radecka et al. 2015; Kręgiel et al. 2017; Mukherjee et al. 2017). Several non-conventional yeast species are capable of producing high concentrations of sugar alcohols (namely xylitol and arabitol) (Schirmer-Michel et al. 2009; Loman et al. 2018), lipids and single-cell oils for food or energy applications (Ratledge 2010; Taskin et al. 2016; Anschau 2017; Hicks et al. 2020), enzymes (Serrat et al. 2004; Saravanakumar et al. 2009; Sahota and Kaur 2015) and pigments (Buzzini and Martini 2000; Aksu and Eren 2005) among other added-value compounds. It should be noted that different yeast species, and even strains, significantly differ in the products synthesised and in their production rates and yields (Rodríguez Madrera et al. 2015; van Dijk et al. 2019). Non-conventional yeasts have recently been in the focus of active and relevant research, their genome sequences are being released and suitable genetic engineering tools are either available or being developed for different purposes (Mira et al. 2014; Palma et al. 2017; Nambu-Nishida et al. 2017; Lee et al. 2018; Cai et al. 2019; Protzko et al. 2019; among several other examples). Thus, it is expected that, in the near future, the currently accepted designation of “non-conventional yeast” will no longer be adequate and non-Saccharomyces strains will successfully be used in the industry (Johnson 2013a; Radecka et al. 2015; Kręgiel et al. 2017; Siripong et al. 2018). This review paper presents relevant results and discusses the potential and the current challenges of the use of yeasts for the valorisation of pectin-rich agro-industrial residues.
Pectin-rich agro-industrial residues as feedstocks for biotechnology
Pectin structure and pectin-rich biomasses
Pectin is a family of complex heteropolysaccharides and a structural component of plant cell walls (Mohnen 2008). Pectin is composed of a linear chain of α-1,4 linked d-galacturonic acid (d-GalA) molecules which represent about 70% of total weight in a homogalacturonan polymer. There are three major pectin polymers: homogalacturonan (HG), rhamnogalacturonan I (RG-I) and rhamnogalacturonan II (RG-II) (Fig. 1). More complex pectin structures, such as rhamnogalacturonan I and II, have side chains composed by neutral sugars that include l-rhamnose, l-arabinose, d-xylose, d-galactose, l-fucose and d-glucose, among others (Sakai et al. 1993). These sugars are linked to d-galacturonic acid by β-1,2 and β-1,4 glycosidic linkages (Fig. 1) (Jayani et al. 2005; Mohnen 2008). Moreover, d-galacturonic acid (d-GalA) residues can be methyl-esterified at the C6 carboxyl group and/or O-acetylated at C-2 or C-3 and neutralised by ions, like sodium, calcium or ammonium (Sakai et al. 1993; Jayani et al. 2005).
Fig. 1.
Schematic representation of the chemical structure of four pectic polysaccharides: homogalacturonan (HG), substituted HG xylogalacturonan (XGA) and rhamnogalacturonan I and II (RG-I and RG-II), based on (Mohnen 2008)
Pectin-rich biomasses, in particular the agro-food residues left after fruit or vegetable processing for juice or sugar production (e.g. apple pomace, citrus waste, and sugar beet pulp), are abundant and widely underused bioresources (Zhou et al. 2008; Mohnen 2008). Although most food waste streams contain pectin too, the residues mentioned above exhibit the highest pectin content with pectin concentrations ranging from 12 to 35% of the biomass dry weight (Müller-Maatsch et al. 2016). The low lignin content of these processed wastes is an interesting trait because lignin can impair with the enzymatic degradation of cellulose and hemicellulose, and its monomers cannot be used as carbon sources so far (Guo et al. 2009). Lignin, the most recalcitrant cell wall material, can be combusted and converted into electricity and heat (Limayem and Ricke 2012). The composition of agro-food residues and the bioavailability of their various polysaccharide fractions are highly dependent on natural variation, husbandry practices, fruit maturity and post-harvest management (Grohmann and Bothast 1994). The apparent high variability of the different pectin-rich biomasses regarding the dry-weight composition in pectin and other polysaccharides is shown in Fig. 2. Determining pectin in biomass quantitatively is actually quite challenging and the differences detected may simply result from the use of different analytic methods and sub-optimal techniques (Quemener et al. 1993; Kühnel 2011; Wikiera et al. 2015).
Fig. 2.
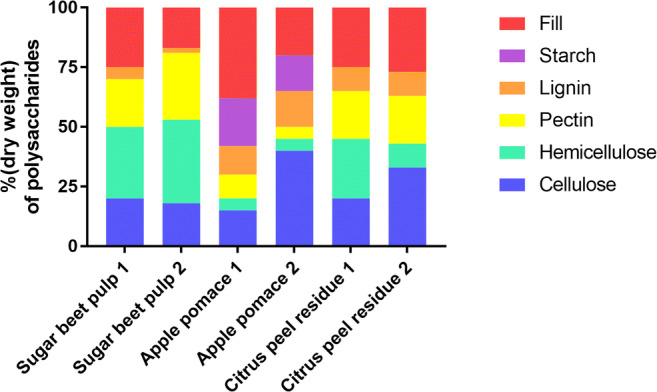
Dry-weight composition of pectin-rich residues, in particular sugar beet pulp 1 (Berlowska et al. 2018) and 2 (Edwards and Doran-Peterson 2012), apple pomace 1 (Grohmann and Bothast 1994) and 2 (Bhushan et al. 2008) and citrus peel 1 (Zhou et al. 2008) and 2 (John et al. 2017)
Pectin-rich biomass processing and composition of the resulting hydrolysates
The bioconversion of pectin-rich agro-industrial residues requires pre-treatment step(s) before microbial utilisation in order to avoid the recalcitrant material and to increase surface area to facilitate and enhance the hydrolysis step (Limayem and Ricke 2012). After pre-treatment, the enzymatic or acidic hydrolysis of cellulose, hemicellulose and pectin structures allow the release of monomeric sugars (saccharification) that will subsequently be converted into ethanol and/or other bioproducts by yeasts. Filamentous fungi, in particular Aspergillus sp., Trichoderma reesei and Neurospora crassa, naturally have degrading machinery consisting in hydrolytic and oxidative enzymes which play an important role in plant biomass degradation (Schmitz et al. 2019). A recent review details the enzyme repertoire of filamentous fungi and their specific transcriptional regulation patterns for efficient biomass degradation (Benocci et al. 2017).
Remarkably, the sugar composition of the hydrolysates obtained from the same pectin-rich agro-industrial residue is highly dependent on the pre-treatment and enzyme hydrolysis conditions used (Table 1) reducing the reproducibility of the hydrolysis process (Merz et al. 2016).
Table 1.
Composition of pectin-rich agro-industrial residues hydrolysates depending on their pre-treatment and hydrolysis
| Feedstock | Pre-treatment | Hydrolysis | Sugar composition after hydrolysis (g/100 g matter) | Reference |
|---|---|---|---|---|
| Sugar beet pulp | Steam explosion at 152 °C to 175.5 °C and 4 to 8 bar pressure | (Soluble fraction) Acid hydrolysis: 72%H2SO4 for 1 h at 30 °C and 150 rpm | Glucose 26 | Hamley-Bennett et al. (2016) |
| Arabinose 24 | ||||
| Xylose 1.6 | ||||
| Rhamnose 2.4 | ||||
| Galactose 6 | ||||
| Galacturonic acid 14 | ||||
| (insoluble fraction) Enzyme hydrolysis: 0.5 mg cellulase/g glucan 50 °C with shaking for 24 h | Glucose 10 | |||
| Arabinose 0.4 | ||||
| Xylose 0.3 | ||||
| Rhamnose 0.1 | ||||
| Galactose 0.1 | ||||
| Galacturonic acid <0.1 | ||||
| Not used | Enzyme hydrolysis: Viscozyme and Ultraflo Max (Novozymes) treatment | Total of reducing sugars 6.6 | Berlowska et al. (2017) | |
| Apple pomace | 15 g/L sulphuric acid for 16 min at 91 °C with laccase 100 units/L at 30 °C for 12 h at 90 rpm | Enzyme hydrolysis: Viscozyme and Celluclast (0.5 μL/mL, 0.038 mg/mL) together with Novozyme188 (0.05 μL/mL, 0.0024 mg/mL) | Galacturonic acid 33 | Gama et al. (2015) |
| Glucose 21 | ||||
| Arabinose 17 | ||||
| Galactose 5 | ||||
| Not used | Acid hydrolysis 1.5 g sulphuric acid/100 mL, 91 °C reaction temperature during 16 min | 18.2 g of glucose and fructose/100 g dry matter | Parmar and Rupasinghe (2012) | |
| Not used |
Acid hydrolysis 72% sulphuric acid for 45 min at room temperature and diluted with distilled water to 4% sulphuric acid, followed by autoclaving for 1 h at 121 °C |
Galacturonic acid (not quantified) | Choi et al. (2015) | |
| Glucose 25 | ||||
| Fructose 24 | ||||
| Arabinose 6 | ||||
| Sucrose 9 | ||||
| Galactose 4 | ||||
| Xylose 6 | ||||
| Rhamnose 2 | ||||
| Not used | Acid hydrolysis 2 M Trifluoroacetic acid for 2 h at 100 °C with constant shaking | Rhamnose 0.5 | Wikiera et al. (2015) | |
| Arabinose 8 | ||||
| Glucose 12 | ||||
| Galactose 4 | ||||
| Xylose 4 | ||||
| Mannose 0.7 | ||||
| Acid hydrolysis 0.2 M Trifluoroacetic acid for 72 h at 80 °C with constant shaking | Enzyme hydrolysis: Viscozyme (25 μL) incubated at 50 °C during 24 h with constant shaking | Rhamnose 0.4 | ||
| Arabinose 7 | ||||
| Glucose 11 | ||||
| Galactose 4 | ||||
| Xylose 4 | ||||
| Mannose 0.3 | ||||
| Citrus peel | 80% v/v ethanol for 20 min, filtered on a sintered glass, and dried at 40 °C for 72 h | Acid hydrolysis: 0.05 M hydrochloric acid at 85 °C | Galacturonic acid 15 | Yapo et al. (2007a) |
| Arabinose 4 | ||||
| Galactose 1 | ||||
| Glucose 1 | ||||
| Rhamnose 0.5 | ||||
| Not used | Acid hydrolysis 72% sulphuric acid for 45 min at room temperature and diluted with distilled water to 4% sulphuric acid, followed by autoclaving for 1 h at 121 °C | Orange peel | Choi et al. (2015) | |
| Galacturonic acid (not quantified) | ||||
| Glucose 36 | ||||
| Fructose 12 | ||||
| Arabinose 6 | ||||
| Sucrose 5.6 | ||||
| Galactose 3 | ||||
| Xylose 2 | ||||
| Rhamnose 2 | ||||
| Lemon peel | ||||
| Galacturonic acid (not quantified) | ||||
| Glucose 27 | ||||
| Fructose 3 | ||||
| Arabinose 5 | ||||
| Sucrose 0 | ||||
| Galactose 5 | ||||
| Xylose 3 | ||||
| Rhamnose 2 | ||||
| Steam explosion at 150 °C for 10 min and 15 kg/cm pressure | Pectinase, xylanase (5 mg/g dry matter) and β-glucosidase (2 mg/g dry matter) cocktail at 45 °C for 24 h | Galacturonic acid (not quantified) | Choi et al. (2013) | |
| Glucose 45 | ||||
| Fructose 18 | ||||
| Arabinose 3 | ||||
| Galactose 2 |
Pectin-rich biomass hydrolysates may also include growth inhibitory compounds, such as weak acids, furan derivatives and phenolic compounds generated during pre-treatment and acid hydrolysis of pectin-rich materials (Palmqvist and Hahn-Hägerdal 2000). Acetic acid and methanol are potential growth inhibitors that are likely to be present (Vendruscolo et al. 2008; Günan Yücel and Aksu 2015; Berlowska et al. 2018). These compounds have potential to affect yeast growth, fermentation kinetics and metabolite production yields, (dos Santos and Sá-Correia 2015; Cunha et al. 2019). Although, the individual toxicity of some of these compounds can be relatively low, their combined toxic effects can be additive or even synergistic (Palmqvist and Hahn-Hägerdal 2000; Teixeira et al. 2011). The average degree of methylation and acetylation of diverse pectin-rich residues is different with sugar beet exhibiting the highest acetylation degree (Table 2). Other potentially critical inhibitors are heavy metals and pesticides. They have also been detected in pectin-rich residues, mainly due to the geochemical cycles and human activities, such as intensive agriculture, waste treatment and disposal and transportation (Legrand 2005; Skrbic et al. 2010; Mukherjee et al. 2017).
Table 2.
Percentage (of total dry matter) of acetylation and methylation of different pectin-rich materials
Yeast metabolism of sugar monomers present in pectin-rich hydrolysates
The challenges
The efficient utilisation by yeasts of the mixtures of sugar monomers present in hydrolysates derived from pectin-rich residues is essential for their biotechnological valorisation. Sugar beet pulp and citrus peel hydrolysates contain predominantly the neutral sugars l-arabinose, d-glucose and d-galactose and the acidic sugar d-galacturonic acid (Micard et al. 1996; Berlowska et al. 2018). This means that the convenient yeast species/strains to be used should be able to rapidly and efficiently catabolise all the sugars present (Du et al. 2019).
The presence and simultaneous use of several sugars in pectin hydrolysates is an important challenge also due to carbon catabolite repression (CCR) regulation (Kayikci and Nielsen 2015; Gao et al. 2019). This regulation mechanism limits the efficient utilisation of multiple carbon substrates in biotechnological processes like those developed for the valorisation of pectin-rich residues. In fact, the uptake of secondary carbon sources (e.g. l-arabinose, d-galacturonic acid, d-xylose) is inhibited in the presence of the preferred substrate (d-glucose), prolonging fermentation time as the result of sequential, rather than the simultaneous, use of the carbon sources (Huisjes et al. 2012; Wu et al. 2016; Yaguchi et al. 2018; Lane et al. 2018). S. cerevisiae has a highly complex and still not fully understood network of signals and regulations, through (de)phosphorylation mechanisms depending on the presence of d-glucose in the medium which have been on the focus of extensive review papers (Gancedo 1992; Conrad et al. 2014; Kayikci and Nielsen 2015).
Moreover, pectin-rich hydrolysates contain a significant amount of d-galacturonic acid that is neither naturally used by S. cerevisiae nor by other relevant yeast species, such as Kluyveromyces marxianus, Yarrowia lipolytica, Pichia stipitis, among others. Recent efforts have been reported in order to genetically engineer S. cerevisiae to efficiently express the d-galacturonic acid catabolic pathway (Benz et al. 2014; Zhang et al. 2015; Nielsen and Keasling 2016; Matsubara et al. 2016; Biz et al. 2016; Kalia and Saini 2017; Lian et al. 2018; Protzko et al. 2018; Jeong et al. 2020). Moreover, since pectin-rich hydrolysates have significant amounts of l-arabinose, efforts have also addressed the expression of this pentose-fermentative pathway in S. cerevisiae strains (Wisselink et al. 2007; Ye et al. 2019). The pathways involved in the catabolism of d-galacturonic acid and l-arabinose, the main sugars released from pectin-rich feedstocks hydrolysis, are detailed below (Figs. 3, 4, 5). d-galacturonic acid catabolic pathway is emphasised because this acid sugar catabolism is currently the big challenge for which there is relevant recent literature. Non-conventional yeast species/strains reported in the scientific literature as naturally capable of such catabolism are also referred. Due to space limitations and the more-established catabolism of most of the other sugars present or their marginal concentrations in the hydrolysates, the corresponding pathways are not described here.
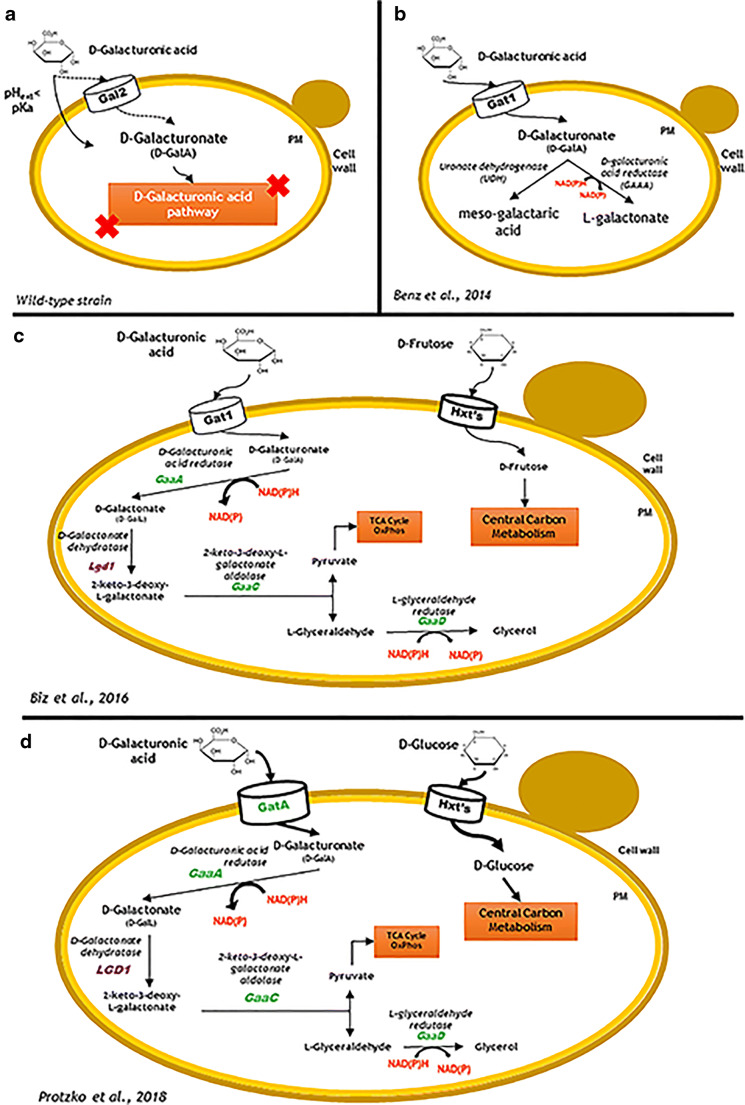
Fig. 3.
Schematic representation of S. cerevisiae strains (wild type and genetically engineered with heterologous d-galacturonic acid degradation pathways. aS. cerevisiae wild-type strain showing the basal natural uptake of d-galacturonic acid by Gal2p transporter and passive diffusion of the undissociated form through plasma membrane (PM). b Engineered S. cerevisiae strains expressing d-GalA membrane transporter Gat1 from Neurospora crassa and the uronate dehydrogenase (UDH) from Agrobacterium tumefaciens and d-galacturonic acid reductase (GAAA) from Aspergillus niger to convert d-GalA into the metabolites meso-galactaric acid and l-galactonate, (Benz et al. 2014). c Engineered S. cerevisiae strain with d-galacturonic acid plasma membrane transporters from N. crassa (GAT1) and enzymes of the d-GalA catabolic pathway GaaA, GaaB, GaaC and GaaD from A. niger (in green) and LGD1 from Trichoderma reesei (in purple); d-Fructose was used as co-substrate (Biz et al. 2016). d Engineered S. cerevisiae strains with the non-glucose repressible plasma membrane d-galacturonic acid transporter GatA from A. niger (GATA) and d-GalA catabolic pathway as in c); d-glucose was used as co-substrate (Protzko et al. 2018)
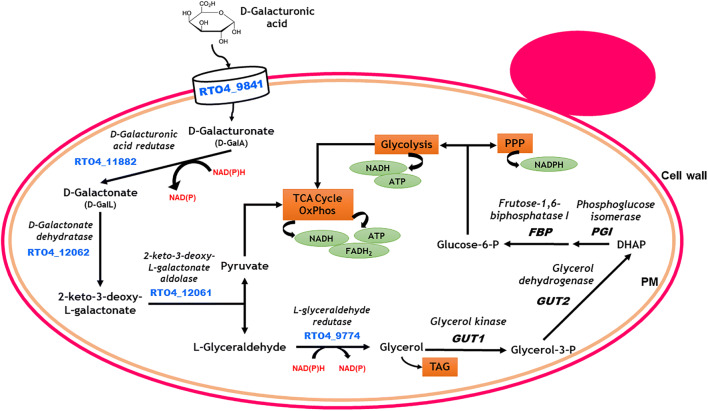
Fig. 4.
Schematic representation of the d-galacturonic acid catabolic pathway proposed for Rhodosporidium toruloides IFO0880. The genes GUT1, GUT2, FBP and PGI belong to central metabolism. TAG, triacylglycerol; PPP, pentose phosphate pathway (based on (Protzko et al. 2019)
Fig. 5.

Schematic representation of the initial steps of arabinose metabolism in fungi (the oxidoreductase pathway) or in bacteria (the isomerase pathway). XK, d-xylulose kinase; AI, l-arabinose isomerase; RK, l-ribulokinase; RPE, l-ribulose-5-phosphate 4-epimerase; XDH, xylitol dehydrogenase; AR, l-arabinose reductase; LAD, l-arabitol 4-dehydrogenase; LXR, l-xylulose reductase (adapted from Fonseca et al., 2007)
d-galacturonic acid pathways from fungi, expressed in S. cerevisiae, and from the oleaginous yeast Rhodosporidium toruloides
d-galacturonic acid is not catabolised by the yeast S. cerevisiae that misses the catabolic pathway (Fig. 3a). Moreover, as an acid sugar, d-galacturonic acid is more oxidised than the neutral hexose and pentose sugars. This means that its metabolism is not redox neutral as glucose metabolism and the fermentation of d-galacturonic acid requires more NADPH cofactor molecules to produce ethanol (Richard and Hilditch 2009). The d-galacturonic acid plasma membrane transporter Gat1 from N. crassa was identified and characterised and the encoding gene GAT1 successfully expressed in S. cerevisiae allowing the increased uptake of d-galacturonic acid in this yeast cell factory (Benz et al. 2014) (Fig. 3b). In fact, d-galacturonic acid uptake is poorly performed when mediated by the native Gal2 or other hexose transporters (Huisjes et al. 2012; Benz et al. 2014; Biz et al. 2016), even though d-galacturonic acid was shown to be taken up rapidly by S. cerevisiae (Souffriau et al. 2012). With the co-expression in yeast of a d-galacturonic acid reductase (from the filamentous fungus Aspergillus niger) or a uronate dehydrogenase (from the bacterium Agrobacterium tumefaciens involved in plant infection), a transporter-dependent conversion of d-galacturonic acid towards more reduced (l-galactonate) or oxidised (meso-galactaric acid) downstream metabolites was also demonstrated (Fig. 3b) (Benz et al. 2014). This heterologous co-expression, although highly relevant as proof of concept, missed the expression of the complete d-galacturonic catabolic pathway for the full catabolisation of this acid sugar.
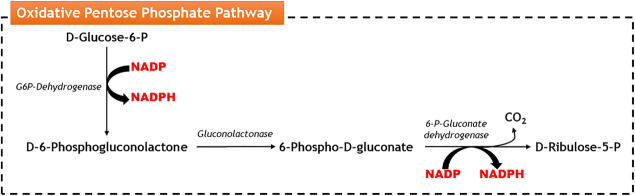
Therefore, several efforts have been made envisaging the development of a genetically engineered S. cerevisiae strain capable of efficiently using d-galacturonic acid from pectin-rich hydrolysates. For this purpose, the genes GAAA, GAAC, GAAD encoding d-galacturonic acid reductase, 2-keto-3-deoxy- l-galactonate aldolase, respectively, from A. niger and the gene LGD1 encoding d-galactonate dehydratase from T. reesei were successfully expressed in S. cerevisiae (Fig. 3c) (Biz et al. 2016). The entire d-galacturonic acid catabolic pathway from filamentous fungi comprises two NADPH-dependent enzymes: the d-galacturonate reductase and the l-glyceraldehyde reductase, for the catabolisation of d-galacturonic acid into glycerol (Biz et al. 2016) (Fig. 3c) leading to intracellular cofactor imbalance. For the efficient functioning of d-galacturonic acid catabolic pathway from filamentous fungi in S. cerevisiae, the pathway has to be coupled with NADPH regeneration steps which can be achieved through the operation of the oxidative pentose phosphate pathway (PPP). The oxidative PPP converts d-glucose-6-P into d-ribulose-5-P and CO2 with the simultaneous reduction reaction of two molecules of NADP to NADPH (Fig. 6) (Wamelink et al. 2008). This cofactor regeneration may enable the catabolisation of d-galacturonic acid in engineered S. cerevisiae strains. However, the use of co-substrates such as d-fructose or d-glucose is required although their presence leads to the delay of d-galacturonic acid catabolisation (Benz et al. 2014; Biz et al. 2016). The successful heterologous expression of the A. nigerd-galacturonic acid transporter GatA in S. cerevisiae allowed the co-uptake of d-galacturonic acid and d-glucose which could also facilitate the regeneration of redox cofactors needed for full conversion of d-galacturonic acid (Fig. 3d) (Protzko et al. 2018). A more recent study reported the expression of the efficient previously described fungal d-galacturonic acid catabolic pathway in a pentose-fermenting S. cerevisiae strain by the expression of a pentose (d-xylose and l-arabinose) catabolic pathway including genes from Pichia stipitis and Ambrosiozyma monospora, both natural pentose-fermentative yeasts (Jeong et al. 2020). Additionally, the authors made a double deletion from genes PHO13 (involved in phosphatase regulation) and ALD6 (a cytosolic aldehyde dehydrogenase required for conversion of acetaldehyde to acetate). All these genetic modifications enabled the co-consumption of more than 10 g/L of d-galacturonic acid with l-arabinose and d-xylose (Ye et al. 2019; Jeong et al. 2020).
Fig. 6.
Schematic representation of the oxidative pentose phosphate pathway (Wamelink et al. 2008)
The genome-wide and enzymatic analysis of the basidiomycete red oleaginous yeast Rhodosporidium toruloides (also known as Rhodotorula toruloides) IFO0880 revealed an efficient d-galacturonic acid metabolism, with highly active enzymes (Fig. 4), suggesting this strain as a potential industrial platform for biodiesel and carotenoid biosynthesis from pectin-rich hydrolysates (Sitepu et al. 2014; Spagnuolo et al. 2019; Protzko et al. 2019). The d-galacturonic acid metabolic pathway of R. toruloides was found to be similar to the T. reesei pathway, being the catabolic enzymes highly induced by d-galacturonic acid (Protzko et al. 2019). Moreover, R. toruloides IFO0880 was found to co-utilise d-galacturonic acid in the presence of either d-glucose or d-xylose. The final product of d-galacturonic acid catabolic pathway is glycerol that has to be used for cofactor regeneration through the oxidative pentose phosphate pathway. The study performed in R. toruloides IFO0880 also showed that the genes GUT1, encoding a glycerol kinase, and GUT2, encoding a mitochondrial glycerol 3-phosphate dehydrogenase, involved in glycerol metabolism and induced in presence of d-galacturonic acid, enabled d-galacturonic acid conversion into glycerol without the need of an additional carbon source. This study proposed that the glycerol produced could be converted into glucose-6-phosphate and, through the oxidative pentose phosphate pathway, the cofactors used in d-galacturonic acid catabolisation would be regenerated (Protzko et al. 2019). Different routes of glycerol catabolic pathways have been described in yeasts, using NAD+- or NADP+-dependent enzymes, balancing the intracellular redox power and enabling growth in respirable carbon sources (Klein et al. 2017).
l-arabinose metabolism
l-arabinose is a five-carbon sugar and, unlike other pentoses that naturally occur in the d-form such as d-xylose, l-arabinose is more common than d-arabinose in nature. Arabinose catabolic pathways include the oxidoreductase (fungal) and the isomerase (bacterial) pathways (Fig. 5). In both pathways, l-arabinose is converted into d-xylulose-5-phosphate, which is metabolised by the non-oxidative phase of the pentose phosphate pathway.
In the fungal pathway, l-arabinose reductase (AR) prefers NADPH as cofactor, whereas the sugar alcohol dehydrogenases (LAD and XDH) are strictly dependent on NAD (Seiboth and Metz 2011). Under low oxygen conditions, the availability of NAD is limited, which may cause an accumulation of l-arabitol (Loman et al. 2018). Furthermore, l-arabinose can be converted into xylitol, the common denominator between the catabolic pathways of l-arabinose and d-xylose (Fig. 5). Due to their partially overlapping pathways, there is a strong correlation between the utilisation of these two pentoses in yeasts (Seiboth and Metz 2011). The introduction of a reconstructed fungal l-arabinose oxidoredutase pathway (from T. reesei and A. monospora strains) into S. cerevisiae allowed l-arabinose utilisation and the production of substantial amounts of l-arabitol due to the severe redox imbalance resulting from the utilisation of NADPH in the reduction step catalysed by l-xylulose reductase (LXR) (Bettiga et al. 2009). In fact, l-xylulose reductase from A. monospora is NADH-dependent enzyme, contrarily to most fungi which are NADPH-dependent for this specific enzyme. However, NADH is produced in the oxidation reactions catalysed by l-arabitol-4-dehydrogenase (LAD) and xylitol dehydrogenase (XDH) improving intracellular redox balance (Bettiga et al. 2009).
Although l-arabinose fermentation by yeasts was thought to be unfeasible, several yeast species have been identified as capable of producing ethanol from l-arabinose, in particular Candida auringiensis, Candida succiphila, A. monospora, Candida sp. (YB-2248) (Dien et al. 1996), and Meyerozyma guilliermondii (Martini et al. 2016). Moreover, the successful engineering of S. cerevisiae to ferment l-arabinose, by expressing the l-arabinose isomerase pathway of the bacterial species of Lactobacillus plantarum (Fig. 5) and overexpressing the S. cerevisiae genes encoding the enzymes of the non-oxidative pentose phosphate pathway, along with extensive evolutionary engineering, resulted in ethanol production (0.43 g g−1) from l-arabinose during anaerobic growth (Wisselink et al. 2007). To increase l-arabinose fermentation rates, potential l-arabinose transporters have been identified and overexpressed in S. cerevisiae. For example, the overexpression of S. cerevisiae Gal2 led to the increase of l-arabinose fermentation rate (Becker and Boles 2003). However, this endogenous S. cerevisiae hexose transporter not only exhibits very low affinities towards pentoses but is also strongly inhibited by glucose (Gao et al. 2019). The expression of heterologous transporters with higher affinities for arabinose over glucose, in particular of Stp2 from Arabidopsis thaliana and AraT from Scheffersomyces stipitis, led to the improvement of l-arabinose fermentation, in anaerobiosis, especially at low l-arabinose concentrations. However, l-arabinose uptake through these two transporters is also inhibited by the presence of glucose (Subtil and Boles 2011).
Toxicity and possible metabolisation of compounds likely present in pectin-rich biomass hydrolysates
Multiple chemical stresses likely affecting pectin-rich biomass bioconversion
It is likely that pectin-rich residues may include variable levels of toxic compounds. Frequently, their concentrations are not always known or even considered, but these compounds may have a potential combined inhibitory effect for yeast growth and metabolism, acting in conjunction or synergistically. In particular, since pectin structures are acetylated and methyl-esterified in different positions of the d-galacturonic acid molecule, this biomass hydrolysis releases acetic acid and methanol that accumulate in the hydrolysate. The potential role of these compounds both as carbon sources and as toxicants with potential to inhibit yeast growth and fermentation is discussed below. Other toxic compounds are likely present in pectin-rich residues. This is the case for heavy metals that in small amounts are essential micronutrients for yeasts but when they reach toxic concentrations induce the generation of reactive oxygen species (ROS) leading to oxidative stress and loss of biological functions (Mukherjee et al. 2017). The pesticides (fungicides, herbicides and insecticides) used in agriculture may also be present in significant amounts, varying among countries although the maximum residual levels allowed are regulated (European Parliament 2009). Other toxic compounds, for instance phenolic compounds and furans, resulting from acid hydrolysis, may also be present (Berlowska et al. 2018). The accumulation of ethanol or other toxic metabolites are additional sources of combined chemical stresses challenging yeast performance during the bioprocess.
Acetic acid and galacturonic acid as carbon sources and toxic compounds
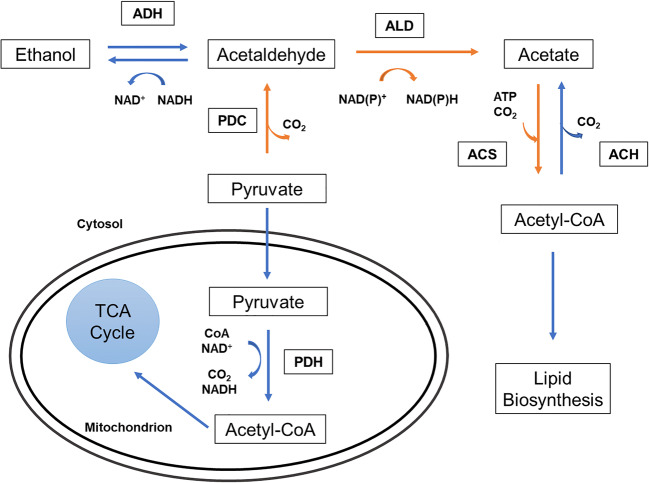
Acetic acid is present in pectin-rich residue hydrolysates at higher concentrations in sugar beet pulp hydrolysates compared with citrus peel hydrolysates (Grohmann et al. 1999; Günan Yücel and Aksu 2015), as discussed before. Acetic acid is also a yeast metabolite generated during growth and fermentation. Acetic acid is a source of carbon and energy for a large number of yeasts and can be converted into lipids (Huang et al. 2016) (Fig. 7). Most of the yeast species capable of growing in high acetic acid concentrations are oleaginous, since acetate can be assimilated and converted into acetyl-CoA, a lipid biosynthesis precursor (Spagnuolo et al. 2019). At sub-lethal concentrations, acetic acid is catabolised by several yeast species, like S. cerevisiae, Candida utilis, Torulaspora delbruecki and Dekkera anomala, its utilisation being repressed by glucose (Radecka et al. 2015). However, d-glucose and acetic acid are simultaneously catabolised in the highly tolerant Zygosaccharomyces bailii species (Rodrigues et al. 2012).
Fig. 7.
Acetic acid metabolism in yeast. The PDH pathway is indicated by blue arrows, while the PDH bypass is indicated by orange arrows. PDH: pyruvate dehydrogenase; PDC: pyruvate decarboxylase; ALD: aldehyde dehydrogenase; ACS: acetyl-CoA synthetase; ADH: alcohol dehydrogenase (Huang et al. 2016)
Depending on the level of acetic acid-induced stress and on the tolerance of a specific yeast strain, acetic acid can act as a growth inhibitor due to the ability of the non-dissociated form (pKa 4.7) to diffuse across plasma membrane and cause toxicity when in the cytosol (Mira et al. 2010a, b ; Mira et al. 2011; Palma et al. 2018). The subsequent deprotonation of this acid in the cytosol, with a pH around neutrality, leads to the accumulation of the acetate counter-ion and cytosol acidification (Carmelo et al. 1997). The effect of a specific concentration of acetic acid is particularly drastic at pH below the pKa of the acid. To obtain a holistic view on the toxic effects and the adaptive responses of yeasts to acetic acid, the following review paper is suggested (Palma et al. 2018). The non-conventional food spoilage yeast species Z. bailii is able to thrive in acid foods and beverages due to its remarkable tolerance to weak acids at low pH (Mira et al. 2014; Palma et al. 2017). In fact, Z. bailii is able to grow at concentrations of acetic acid 3-fold higher (370–555 mM) than S. cerevisiae (80–150 mM) (Palma et al. 2015; Palma et al. 2018). The remarkable tolerance of Z. bailii to weak acids has brought to light the potential of this yeast species as an alternative cell factory for the production of high levels of weak acids (Palma and Sá-Correia 2019). Moreover, the understanding of the mechanisms underlying the tolerance to weak acids in Z. bailii sensu lato allows the identification of candidate molecular targets for the rational genome engineering for the construction of more robust S. cerevisiae strains (Mira et al. 2014; Guerreiro et al. 2016; Palma et al. 2017). Other reported acetic acid-tolerant yeast species are Pichia kudriavzevii (Dandi et al. 2013) and Candida glycerinogenes (Ji et al. 2016; Zhao et al. 2019). For efficient bioconversion of pectin-rich residues hydrolysates rich in acetic acid, the use of tolerant strains and/or a pH above this weak acid’s pKa is required.
The presence of d-galacturonic acid, even at low concentrations (up to 10 g/L) and pH 3.5 (below the pKa of the acid), in a cultivation medium with a mixture of glucose, galactose, xylose, and arabinose, mimicking pectin-rich residue hydrolysates, was reported to affect the fermentation of most of the sugars with the exception of glucose by a genetically engineered pentose-fermenting strain S. cerevisiae CEN.PK 113-7D grown under anaerobiosis (Huisjes et al. 2012). However, at pH 5, at which the concentration of the undissociated toxic form is low, sugar fermentation performance was not affected by the presence of d-galacturonic acid (Huisjes et al. 2012).
Methanol as carbon source and toxic compound
Methanol is another toxic compound likely present in pectin-rich hydrolysates. Methanol toxicity mechanisms are poorly studied but, like ethanol and other alcohols, the cell membranes are the anticipated molecular targets (van der Klei et al. 2006). Methanol can be converted into formaldehyde which is a more toxic compound (Yasokawa et al. 2010). For S. cerevisiae, 1.23 M of methanol or 1.8 mM of formaldehyde, are concentrations reported to inhibit growth without causing cell death (Yasokawa et al. 2010).
Despite S. cerevisiae inability to grow in methanol, there are several non-conventional yeasts that can efficiently use it as the sole carbon and energy source. Since methanol is an inexpensive carbon source, methylotrophic yeasts have been examined for biotechnological applications, ranging from the production of single-cell protein (SCP) and heterologous recombinant proteins to the production of number of chemical compounds (Limtong et al. 2008; Johnson 2013b; Siripong et al. 2018). The most well-known methylotrophic yeast species are Candida boidinii, Ogataea (Pichia) methanolica, Komagataella (formerly Pichia) pastoris, Ogataea minuta and Ogataea (formerly Hansenula) polymorpha, as well as Candida parapsilosis, Candida (formerly Torulopsis) glabrata and Ogataea (formerly Pichia) thermomethanolica (Limtong et al. 2008; Kurtzman and Robnett 2010; Johnson 2013b). The successful genetic modification of S. cerevisiae by expressing enzymes from Pichia pastoris methanol catabolic pathway (AOX, encoding alcohol oxidases, CAT encoding a catalase, DAS, encoding a dihydroxyacetone synthase, and DAK, encoding a dihydroxyacetone kinase) enabled the consumption by the recombinant S. cerevisiae strain of 50% of initial methanol concentration (Dai et al. 2017).
Heavy metals and agricultural pesticides as toxic compounds
Heavy metals are essential micronutrients for yeasts. However, when above concentration threshold, they induce the generation of reactive oxygen species (ROS) leading to oxidative stress with the oxidation of proteins, lipids and nucleic acids, thus affecting their biological functions (Mukherjee et al. 2017). In general, pectin can bind different heavy metals depending on their structure and natural environment (following preference: Pb2+ >> Cu2+ > Co2+ (cobalt) > Ni2+ (nickel) >> Zn2+ > Cd2+. Sugar beet biomass has preferential affinity for Cu2+and Pb2+ (Schiewer and Patil 2008). The tolerance to heavy metals is strain-dependent and the variability is large among strains of the same species to different metals (Balsalobre et al. 2003; Vadkertiová and Sláviková 2006). The pesticides (fungicides, herbicides and insecticides) used in agriculture vary among countries, but the maximum residual levels allowed are regulated in the EU and by FDA. The mechanisms of toxicity and tolerance to agricultural pesticides in yeasts are more poorly studied, although the global effects of the herbicide 2,4-D and the agricultural fungicide mancozeb, among others, have been reported (Teixeira et al. 2007; Dias et al. 2010; dos Santos 2012).
Bioconversion of carbon source mixtures: the challenges
The hydrolysates prepared from pectin-rich residues include a wide range of different carbon sources (C-sources) at variable concentrations, depending on the type of biomass and their processing conditions, as detailed above. The assimilation of usable C-sources by yeasts is strictly regulated and most of the catabolic pathways are subject to CCR (Simpson-Lavy and Kupiec 2019). This constitutes a major challenge for the efficient and economic utilisation of complex substrates in biotechnological processes since in the presence of a preferred sugar, the uptake of secondary carbon sources is inhibited and their sequential utilisation prolong the fermentation time. When d-glucose is present in the extracellular medium, the uptake and catabolism of other carbon sources is repressed in S. cerevisiae (Kayikci and Nielsen 2015; Wu et al. 2016; Lane et al. 2018). Strategies for circumventing CCR are especially important when it comes to the use of inexpensive and renewable feedstocks containing mixtures of carbon sources, such as in the case of pectin-rich residues. In fact, the separation of individual substrates is costly and impractical and for this reason, the efficient utilisation of substrate mixtures is a necessity that requires additional strain-improvement efforts (Gao et al. 2019). Efforts to enable C-sources co-utilisation include the introduction of non-native sugar transporters or catabolic pathways that are not subject to CCR or by adaptive evolution and targeted genome engineering (Papapetridis et al. 2018). Yeast strains are susceptible to CCR and in the specific case of pectin-rich biomass hydrolysates, the glucose present is used at first and galactose is expected to be consumed subsequently since the Leloir pathway, through which a molecule of d-galactose is converted into glucose-1-phosphate ready to be used in glycolysis (Sellick et al. 2008), is repressed by glucose (Huisjes et al. 2012). In the case of strains capable of using the other less easily metabolised carbon sources, they will be used sequentially. For example, the strain M. guilliermondii FTI 20037 was found to have a native ability to catabolise hexose and pentoses, but when cultivated in a mixed-sugars medium, l-arabinose is only consumed when d-glucose and d-xylose are completely depleted from the medium (Mussatto et al. 2006). However, the simultaneous co-consumption of d-glucose, d-xylose and l-arabinose by Pseudozyma hubeiensis IPM1–10 in artificial hydrolysate of lignocellulosic biomass (mixed-sugar medium) was reported leading to the production of high amounts of lipids in less time compared with single-sugar media (Tanimura et al. 2016). Very recently, a pentose-fermenting strain S. cerevisiae YE9 expressing the fungal d-galacturonic acid pathway and deleted from PHO13 and ALD6 genes (see above) was able to co-consume d-galacturonic acid, l-arabinose and d-xylose (mixed-sugar medium), showing a low susceptibility to catabolic repression (Jeong et al. 2020).
Moreover, d-glucose affects the expression of genes related to other cellular functions such as respiration, gluconeogenesis and the general stress response mechanisms (Lane et al. 2018). The repression of respiration in glucose-containing environments is known as the “Crabtree effect” (Pfeiffer and Morley 2014). The Crabtree effect is observed in S. cerevisiae that even under aerobic conditions undergoes alcoholic fermentation when glucose is present at non-limiting concentrations (Pfeiffer and Morley 2014). The fermentative, Crabtree-positive yeasts include the genera Saccharomyces, Zygosaccharomyces, Dekkera and Schizosaccharomyces while Crabtree-negative yeasts include strains belonging to the genera Pichia, Debaryomyces, Candida or Kluyveromyces (Rozpędowska et al. 2011).
It is important to notice that the use of recombinant yeasts, constructed based on the application of metabolic engineering and synthetic biology tools, has shown that when single substrates are used, several limitations to their metabolism may occur, resulting in low yield (Liu et al. 2020). For instance, when the target product has distinct chemical properties or requires long synthetic routes from starting substrates (Babel 2009). The improvement of product biosynthesis through the optimal balance of biosynthetic components can be achieved by the application of mixed substrates, changing flux distribution and cellular resources, instead of intensive genetic modifications (Liu et al. 2020).
Value-added bioproducts from pectin-rich hydrolysates by non-conventional yeasts
The interest in non-Saccharomyces yeasts is gaining momentum due to that a variety of important features they possess that are not present in the model yeast S. cerevisiae making this large group of yeast species/strains desirable cell factories for the synthesis of a wide range of added-value products (Radecka et al. 2015). These traits of metabolic versatility and yeast physiology are highly valuable for the biosynthesis of interesting added-value compounds from pectin-rich residue hydrolysates (Wagner and Alper 2016; Rebello et al. 2018). A significant genetic distance is observed in the phylogenetic tree prepared for yeasts exhibiting different capacities to catabolise pentoses (among the ascomycetous yeasts) or d-galacturonic acid (basidiomycetous yeasts, close to filamentous fungi with a similar metabolic trait) (Fig. 8).
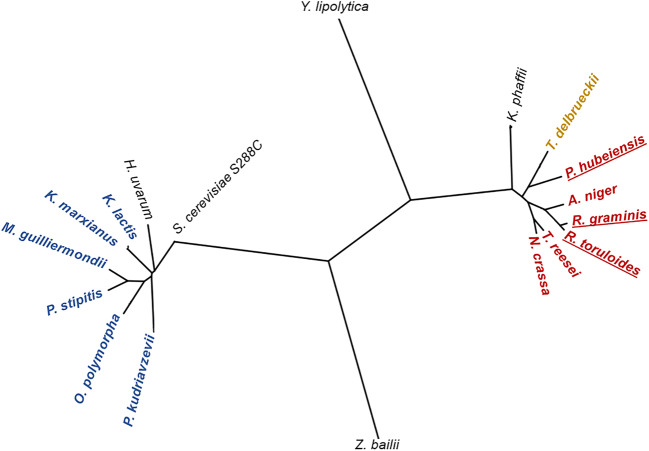
Fig. 8.
Phylogenetic tree of relevant yeasts and related filamentous fungi discussed in this work. The tree was constructed using the neighbour-joining method based on the alignment of the large subunit (26S) ribosomal DNA sequence. The sequences used were obtained from “EnsemblFungi” database. The yeasts coloured with blue (the Ascomycetous yeasts Kluyveromyces marxianus, Kluyveromyces lactis, Meyerozyma guilliermondii, Pichia stipitis, Ogataea polymorpha and Pichia kudriavzevii) are capable of utilizing d-xylose and l-arabinose as carbon sources (C-sources). Red colour represented basidiomycetous yeasts (underlined), such as Rhodosporidium toruloides, Rhodotorula graminis and Pseudozyma hubeiensis and filamentous fungi (Trichoderma reesei, Aspergillus niger and Neurospora crassa) which are able to grow in d-galacturonic acid and also in d-xylose and l-arabinose. The yeast species Torulaspora delbrueckii represented in yellow is capable to grow in d-galacturonic acid and d-xylose. The phylogenetic tree also includes (black colour) Saccharomyces cerevisiae S288C, Zygosaccharomyces bailii, Yarrowia lipolytica and Komagataella phaffii. The yeast species S. cerevisiae K. marxianus, M. guilliermondii, P. stipitis, P. kudriavzevii and T. delbrueckii are interesting bioethanol producers, while H. uvarum is also responsible for the fruity-like aromatic compounds in fermented beverages. Y. lipolytica, P. hubeiensis, R. graminis and R. toruloides are oleaginous yeasts which can convert C-sources into high concentrations and a wide range of lipids. The species K. phaffii is mainly used as cell factory for heterologous protein expression while Z. bailii exhibits a remarkable tolerance to weak acids
Although several non-conventional yeasts with potential for the bioconversion of pectin-rich wastes have received the “generally recognised as safe” (GRAS) label from FDA (Food and Drug Administration), there are several interesting species that are, unfortunately, reported opportunistic pathogens (Wirth and Goldani 2012; Johnson 2013a). The genera Cryptococcus, Candida and Rhodotorula are some of those encompassing pathogenic species, such as Cryptococcus neoformans and Cryptococcus gatii (Johnson 2013a) as well as Rhodotorula mucilaginosa, Rhodotorula glutinis and Rhodotorula minuta (Wirth and Goldani 2012). However, the potential of some of them to produce interesting metabolites is high, and therefore, they are potential sources of genetic information for the engineering of GRAS species.
There is a wide range of products synthesised by different non-conventional yeast species using pectin-rich substrates that have been reported in the literature. A summary of these examples is shown in Table 3. Most of the bioethanol in the market is produced from hexose fermentation by the yeast S. cerevisiae, namely from glucose and fructose. However, several yeast species are also able to ferment other sugars present in pectin-rich residues and were reported as bioethanol producers from that biomass. This is the case for strains of K. marxianus (Serrat et al. 2004), M. guilliermondii (Schirmer-Michel et al. 2008; Schirmer-Michel et al. 2009), Scheffersomyces (Pichia) stipitis (Günan Yücel and Aksu 2015), P. kudriavzevii (Kaur Sandhu et al. 2012), Hanseniaspora uvarum and Hanseniaspora valbyensis (Rodríguez Madrera et al. 2015).
Table 3.
Reported examples of bioconversion of pectin-rich residues by non-conventional yeasts
| Yeast | Pectin-rich residues | Initial sugar concentration (of total sugars in hydrolysate) | Bioproducts (final concentration or yield) | References |
|---|---|---|---|---|
| Scheffersomyces (Pichia) stipitis NRRL Y-7124 | Sugar beet pulp hydrolysate | 75 g/L | Ethanol (37.1 g/L) | Günan Yücel and Aksu (2015) |
| Pichia kudriavzevii KVMP10 | Kinnow mandarin peels hydrolysate | 79 g/L | Ethanol (34 g/L) | Kaur Sandhu et al. (2012) |
| Orange peel hydrolysate | 101 g/L | Ethanol (54 g/L) | Koutinas et al. (2016) | |
| Candida parapsilosis IFM 48375 | Orange peel hydrolysate | – | Ethanol (0.85 g EtOH/4.2 g of dry matter of orange peel) | Tsukamoto et al. (2013) |
| Candida parapsilosis NRRL Y-12969 | Orange peel hydrolysate | – | Ethanol (0.76 g EtOH/4.2 of dry matter of orange peel) | Tsukamoto et al. (2013) |
| Hanseniaspora uvarum H.u. 283 | Apple pomace hydrolysate | 36 g/kg of apple pomace | Ethanol (2.8% (w/w) of reducing sugars) Volatile fruity-like aroma compounds (esters and γ-Nonalactone) | Rodríguez Madrera et al. (2015) |
| Hanseniaspora valbyensis H.v. 43 | Apple pomace hydrolysate | 36 g/Kg of apple pomace | Ethanol (2.8% (w/w) of reducing sugars) Volatile fruity-like aroma compounds (esters and γ-Nonalactone) | Rodríguez Madrera et al. (2015) |
| Yarrowia lipolytica MYA-2613 | Apple pomace hydrolysate | 80 g/L | Lipids (25.8 g/L (C16:0; C18:0; C18:1 C20:0) | Liu et al. (2019) |
| Trichosporon cutaneum AS 2.571 | Beet pulp hydrolysate | 52 g/L | Lipids (7.2 g/L) (palmitic; stearic; oleic, linolenic) | Wang et al. (2015) |
| Trichosporon fermentans CICC 1368 | Beet pulp hydrolysate | 52 g/L | Lipids (5.8 g/L) (palmitic; stearic; oleic, linolenic) | Wang et al. (2015) |
| Cryptococcus curvatus ATCC 20509 | Beet pulp hydrolysate | 52 g/L | Lipids (6.9 g/L) (palmitic; stearic; oleic, linolenic) | Wang et al. (2015) |
| Rhodosporidium toruloides NRRL1091 | Orange peel waste | 18 g/L | Lipids (5.8 g/L) (palmitic; oleic) | Carota et al. (2020) |
| Cryptococcus laurentii UCD 68–201 | Orange peel waste | 18 g/L | Lipids (4.5 g/L) (palmitic; oleic) | |
| Rhodotorula sp. | Apple pomace hydrolysate | 40 g/L | Carotenoids (16.8 mg/100 g DCW) | Joshi et al. (2013) |
| Trichosporon penicillatum SNO-3 | Citrus peel hydrolysate (Citrus unshiu) | 23.2% (w/w) | Protopectin-solubilizing enzyme | Sakai and Okushima (1980) |
| Torula (Candida) utilis CCT3469 | Apple pomace hydrolysate | 15%(w/w) | Lignocellulosic enzymes: pectinase (25 μg/mL), manganese-dependent peroxidase (2.5 μg/mL), cellulase and xylanase (< 1 μg/mL) | Villas-Bôas et al. (2002) |
| Torula (Candida) utilis DSM 70163 | Sugar beet pulp hydrolysate | 45 g/L | Single cell protein (43% g protein/g sugar consumed) | Nigam and Vogel (1991) |
| Candida tropicalis DSM 7015 | 45 g/L | Single cell protein (39% g protein/g sugar consumed) | Nigam and Vogel (1991) | |
| Candida parapsilosis DSM 70125 | 45 g/L | Single cell protein (34% g protein/g sugar consumed) | Nigam and Vogel (1991) | |
| Candida solani ATCC 14440 | 45 g/L | Single cell protein (35% g protein/g sugar consumed) | Nigam and Vogel (1991) |
H. uvarum and H. valbyensis strains produce volatile fruity-like aroma compounds, with high acetic acid ester content, from apple pomace (Rodríguez Madrera et al. 2015). These volatile or non-volatile aromatic compounds are very valuable ingredients in chemical, food, cosmetic and pharmaceutical industries (Martínez et al. 2017) and comprise 25% (aroma compounds) of global market of food additives (Rodríguez Madrera et al. 2015).
Single-cell oil and lipids, namely fatty acids, are obtained from oleaginous yeasts, for utilisation as substitutes for vegetable oils and animal and vegetal fats (e.g. as cocoa butter) (Wang et al. 2012). The demand for biobased-fuels to replace fossil-based-products has led to an increase of biodiesel production and other oleochemical products from oleaginous yeasts (Wang et al. 2012; Anschau 2017). Yeast species, such as Y. lipolytica, Trichosporon cutaneum, Trichosporon fermentans and Cryptococcus curvatus, were reported as yeast platforms to produce different levels of fatty acids from pectin residues. Remarkably, C. curvatus can convert acetate (5 g/L, at pH 6.0) into oils (up to 50% (w/w) of lipid accumulation in the biomass) (Christophe et al. 2012) and Rhodosporidium toruloides can convert 20 g/L of acetic acid (at pH 6.0) in lipids up to 48% (w/w) of the biomass (Huang et al. 2016). R. toruloides lipids are mainly triacylglycerols (C16 and C18 fatty acids) (Singh et al. 2018) and the dried cellular biomass can be directly converted into biodiesel (Guo et al. 2019). A recent study conducted with 18 strains of oleaginous yeasts also reported the accumulation of lipids in R. toruloides NRRL 1091 and Cryptococcus laurentii UCD 68-201 (77 and 47% on a dry matter basis, respectively) from orange peel extract (Carota et al. 2020).
A Rhodotorula sp. strain, isolated from spoiled sauerkraut, was reported to grow and produce carotenoids from 50 g/L apple pomace, but the addition of 0.3% (v/v) ferrous ammonium sulphate led to the highest carotenoid concentration (Joshi et al. 2013). Oleaginous yeasts were identified as capable of growing in medium containing only d-galacturonic acid as carbon source. This is the case of the species C. laurentii, C. curvatus, Cryptococcus cf. aureus, Cryptococcus ramirezgomezianus, Leucosporidiella creatinivora, Tremella encephala, Geotrichum fermentans, R. mucilaginosa, Trichosporon dermatis and Trigonopsis variabilis which exhibit relevant genetic information related with d-galacturonic acid metabolic pathway for alternative expression in S. cerevisiae (Sitepu et al. 2014).
The production of enzymes from agro-industrial residues by yeasts is still one of the most relevant applications for these substrates, in particular for the production of pectinases (Vendruscolo et al. 2008). Single-cell protein (SCP) or yeast components can easily be produced from several agro-industrial wastes and are extremely useful for food and feed nutritional enrichment (Vendruscolo et al. 2008; Johnson and Echavarri-Erasun 2011). Torula utilis, Candida tropicalis, Candida parapsilosis and Candida solani are sources of SCP from sugar beet pulp (Nigam and Vogel 1991).
Metabolic engineering of non-conventional yeasts with potential for the bioconversion of pectin-rich residues
The application of metabolic engineering strategies to non-conventional yeasts envisages the resolution of the problems discussed in previous sections, in particular the co-utilisation of different carbon sources, the enhancement of the tolerance to the inhibitors commonly present in the hydrolysates and other bioprocess-related stresses and the improvement of, or the production of, novel bioproducts. However, the metabolic engineering of non-conventional yeasts faces several challenges such as the reduced availability of stable and high copy number of plasmids and suitable approaches for foreign DNA integration into the host’s genome (Löbs et al. 2017). For industrial bioprocesses, metabolic engineering requires genomic integration of genetic information for high stability of the expression cassette over extended cultivations, homogenous expression levels in cell population and the elimination of the selective marker (Löbs et al. 2017). Currently, there are several genome editing tools already available for metabolic engineering of non-conventional yeasts (Gupta and Shukla 2017). The CRISPR technology is allowing gene disruptions and integrations in several yeast species, such as Kluyveromyces lactis, K. marxianus, S. stipitis, Y. lipolytica, Hansenula polymorpha and P. pastoris (Weninger et al. 2015; Gao et al. 2016; Löbs et al. 2017; Raschmanová et al. 2018; Nurcholis et al. 2020). The perspectives of the metabolic engineering of non-conventional yeasts more suited to industrial bioprocesses are encouraging, supported by the increased availability of genome sequences obtained by next-generation sequencing and the development and availability of genome editing and bioinformatic tools. Among them is the YEASTRACT+ database that also provides biological information and tools for the analysis and prediction of transcription regulatory associations at the gene and genomic levels in non-conventional yeasts of biotechnological interest, in particular Z. baillii, K. lactis, K. marxianus, Y. lipolytica and K. phaffii (Monteiro et al. 2020). These developments are paradigmatic examples that the exploitation of non-Saccharomyces yeasts is gaining momentum.
From the already significant number of examples of metabolic engineering of yeasts for biomass bioconversion, only a few examples of potential interest for the bioconversion of pectin-rich biomass were reported. Oleaginous yeasts are being intensively studied due to their native mechanisms to convert carbon sources into neutral lipids and lipid-derived compounds. For example, the triacylglyceride pathway was engineered into Y. lypolytica by introducing a synthetic pathway that enhances glycolysis activity with an improvement in glycolytic NADH and an increase of approximately 25% of lipid biosynthesis from glucose (Qiao et al. 2017). Tools for the genetic engineering of the oleaginous yeast species R. toruloides to improve the production of carotenoids and lipids were recently developed (Park et al. 2018). The metabolic engineering of K. lactis by the construction of a null mutant in a single gene encoding a mitochondrial alternative internal dehydrogenase led to a metabolic shift from respiration to fermentation, increasing the rate of ethanol production (González-Siso et al. 2015). In K. marxianus, the simultaneous knockdown of the TCA cycle and the electron transport chain genes ACO2b, SDH2, RIP1 and MSS51, resulted in a 3.8-fold increase in ethyl acetate productivity from glucose (Löbs et al. 2018). The examples of genetic manipulation of non-conventional yeasts for sugar transporters are not many, but the heterologous integration of the xylose transporter gene AT5G17010 from A. thaliana into C. tropicalis resulted in a 37–73% increase in xylose uptake compared to the original strain (Jeon et al. 2013). Given that synthetic biology methods and tools are being adapted to be used in non-conventional yeasts, the construction of engineered strains with specific traits for the more efficient bioconversion of pectin-rich agro-industrial residues can be anticipated.
Concluding remarks
The valorisation of pectin-rich residues resulting from the industrial processing of fruits and vegetables for the production of value-added compounds by non-conventional yeast species is gaining momentum. The challenges posed to the industrial implementation of efficient bioprocesses are however many and thoroughly discussed in this review paper. The challenges encountered, at the biological level, range from the simultaneous effective metabolisation of C-source mixtures present in pectin-rich residue hydrolysates and the required increase of yeast robustness to cope with the multiple potential stresses encountered during specific bioprocesses, to the improvement of production of interesting and novel metabolites.
Acknowledgements
This review paper was prepared in the context of the ERANET-IB project YEASTPEC (Engineering of the yeast Saccharomyces cerevisiae for bioconversion of pectin- containing agro-industrial side-streams) funded by the Portuguese Foundation for Science and Technology (FCT) (ERA-IB-2/0003/2015). We would like to thank the research teams of project partners, Elke Nevoigt (Jacobs University, Bremen), Wolfgang Liebl (TUM, Munich) and Peter Richard (VTT, Espoo) for all the scientific interactions and discussions. Thanks are also due to reviewer no. 1 for the deep, knowledgeable and generous review of the submitted manuscript.
Author contribution
All the authors contributed to the writing and read and approved the manuscript.
Funding information
This work was funded by the Portuguese Foundation for Science and Technology (FCT) in particular the YEASTPEC project grant: ERA-IB-2/0003/2015. Funding received by the Institute for Bioengineering and Biosciences (iBB) (UIDB/04565/2020) was from FCT, as well as the PhD fellowship to LCM (DP_AEM—PhD programme—PD/BD/128035/2016).
Compliance with ethical standards
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- Ajila CM, Brar SK, Verma M, Tyagi RD, Godbout S, Valéro JR. Bio-processing of agro-byproducts to animal feed. Crit Rev Biotechnol. 2012;32:382–400. doi: 10.3109/07388551.2012.659172. [DOI] [PubMed] [Google Scholar]
- Aksu Z, Eren AT. Carotenoids production by the yeast Rhodotorula mucilaginosa: use of agricultural wastes as a carbon source. Process Biochem. 2005;40:2985–2991. doi: 10.1016/j.procbio.2005.01.011. [DOI] [Google Scholar]
- Anschau A (2017) Lipids from oleaginous yeasts: production and encapsulation. In: Nutrient Delivery. Elsevier, pp 749–794
- Babel W. The auxiliary substrate concept: from simple considerations to heuristically valuable knowledge. Eng Life Sci. 2009;9:285–290. doi: 10.1002/elsc.200900027. [DOI] [Google Scholar]
- Balat M. Production of bioethanol from lignocellulosic materials via the biochemical pathway: a review. Energy Convers Manag. 2011;52:858–875. doi: 10.1016/j.enconman.2010.08.013. [DOI] [Google Scholar]
- Balsalobre L, De Silóniz M-I, Valderrama M-J, Benito T, Larrea M-T, Peinado J-M. Occurrence of yeasts in municipal wastes and their behaviour in presence of cadmium, copper and zinc. J Basic Microbiol. 2003;43:185–193. doi: 10.1002/jobm.200390021. [DOI] [PubMed] [Google Scholar]
- Becker J, Boles E. A modified Saccharomyces cerevisiae strain that consumes L-arabinose and produces ethanol. Appl Environ Microbiol. 2003;69:4144–4150. doi: 10.1128/AEM.69.7.4144-4150.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benocci T, Victoria M, Pontes A, Zhou M, Seiboth B, De Vries RP. Biotechnology for biofuels regulators of plant biomass degradation in ascomycetous fungi. Biotechnol Biofuels. 2017;10:1–25. doi: 10.1186/s13068-017-0841-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benz J, Protzko RJ, Andrich JM, Bauer S, Dueber JE, Somerville CR. Identification and characterization of a galacturonic acid transporter from Neurospora crassa and its application for Saccharomyces cerevisiae fermentation processes. Biotechnol Biofuels. 2014;7:20. doi: 10.1186/1754-6834-7-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berlowska J, Pielech-Przybylska K, Balcerek M, Cieciura W, Borowski S, Kregiel D (2017) Integrated bioethanol fermentation/anaerobic digestion for valorization of sugar beet pulp. Energies 10: . 10.3390/en10091255
- Berlowska J, Binczarski M, Dziugan P, Wilkowska A, Kregiel D, Witonska I (2018) Sugar beet pulp as a source of valuable biotechnological products. In: Advances in Biotechnology for Food Industry. Elsevier, pp 359–392
- Bettiga M, Bengtsson O, Hahn-Hägerdal B, Gorwa-Grauslund MF. Arabinose and xylose fermentation by recombinant Saccharomyces cerevisiae expressing a fungal pentose utilization pathway. Microb Cell Factories. 2009;8:40. doi: 10.1186/1475-2859-8-40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhushan S, Kalia K, Sharma M, Singh B, Ahuja PS. Processing of apple pomace for bioactive molecules. Crit Rev Biotechnol. 2008;28:285–296. doi: 10.1080/07388550802368895. [DOI] [PubMed] [Google Scholar]
- Biz A, Sugai-Guérios MH, Kuivanen J, Maaheimo H, Krieger N, Mitchell DA, Richard P. The introduction of the fungal d-galacturonate pathway enables the consumption of d-galacturonic acid by Saccharomyces cerevisiae. Microb Cell Factories. 2016;15:144. doi: 10.1186/s12934-016-0544-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buzzini P, Martini A. Production of carotenoids by strains of Rhodotorula glutinis cultured in raw materials of agro-industrial origin. Bioresour Technol. 2000;71:41–44. doi: 10.1016/S0960-8524(99)00056-5. [DOI] [Google Scholar]
- Cai P, Gao J, Zhou Y. CRISPR-mediated genome editing in non-conventional yeasts for biotechnological applications. Microb Cell Factories. 2019;18:63. doi: 10.1186/s12934-019-1112-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cárdenas-Fernández M, Bawn M, Hamley-Bennett C, Bharat PKV, Subrizi F, Suhaili N, Ward DP, Bourdin S, Dalby PA, Hailes HC, Hewitson P, Ignatova S, Kontoravdi C, Leak DJ, Shah N, Sheppard TD, Ward JM, Lye GJ. An integrated biorefinery concept for conversion of sugar beet pulp into value-added chemicals and pharmaceutical intermediates. Faraday Discuss. 2017;202:415–431. doi: 10.1039/c7fd00094d. [DOI] [PubMed] [Google Scholar]
- Carmelo V, Santos H, Sá-Correia I. Effect of extracellular acidification on the activity of plasma membrane ATPase and on the cytosolic and vacuolar pH of Saccharomyces cerevisiae. Biochim Biophys Acta Biomembr. 1997;1325:63–70. doi: 10.1016/S0005-2736(96)00245-3. [DOI] [PubMed] [Google Scholar]
- Carota E, Petruccioli M, D’Annibale A, Gallo AM, Crognale S (2020) Orange peel waste–based liquid medium for biodiesel production by oleaginous yeasts. Appl Microbiol Biotechnol 4617–4628 . 10.1007/s00253-020-10579-y [DOI] [PubMed]
- Cherubini F. The biorefinery concept: using biomass instead of oil for producing energy and chemicals. Energy Convers Manag. 2010;51:1412–1421. doi: 10.1016/j.enconman.2010.01.015. [DOI] [Google Scholar]
- Choi IS, Kim J-H, Wi SG, Kim KH, Bae H-J. Bioethanol production from mandarin (Citrus unshiu) peel waste using popping pretreatment. Appl Energy. 2013;102:204–210. doi: 10.1016/j.apenergy.2012.03.066. [DOI] [Google Scholar]
- Choi IS, Lee YG, Khanal SK, Park BJ, Bae H-J. A low-energy, cost-effective approach to fruit and citrus peel waste processing for bioethanol production. Appl Energy. 2015;140:65–74. doi: 10.1016/j.apenergy.2014.11.070. [DOI] [Google Scholar]
- Christophe G, Deo JL, Kumar V, Nouaille R, Fontanille P, Larroche C. Production of oils from acetic acid by the oleaginous yeast Cryptococcus curvatus. Appl Biochem Biotechnol. 2012;167:1270–1279. doi: 10.1007/s12010-011-9507-5. [DOI] [PubMed] [Google Scholar]
- Conrad M, Schothorst J, Kankipati HN, Van Zeebroeck G, Rubio-Texeira M, Thevelein JM. Nutrient sensing and signaling in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev. 2014;38:254–299. doi: 10.1111/1574-6976.12065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cristobal-Sarramian A, Atzmuller D. Yeast as a production platform in bioreflneries: conversion of agricultural residues into value-added products. Agron Res. 2018;16:377–388. doi: 10.15159/AR.18.066. [DOI] [Google Scholar]
- Cunha JT, Romaní A, Costa CE, Sá-Correia I, Domingues L. Molecular and physiological basis of Saccharomyces cerevisiae tolerance to adverse lignocellulose-based process conditions. Appl Microbiol Biotechnol. 2019;103:159–175. doi: 10.1007/s00253-018-9478-3. [DOI] [PubMed] [Google Scholar]
- Dahiya S, Kumar AN, Shanthi Sravan J, Chatterjee S, Sarkar O, Mohan SV. Food waste biorefinery: sustainable strategy for circular bioeconomy. Bioresour Technol. 2018;248:2–12. doi: 10.1016/j.biortech.2017.07.176. [DOI] [PubMed] [Google Scholar]
- Dai Z, Gu H, Zhang S, Xin F, Zhang W, Dong W, Ma J, Jia H, Jiang M. Metabolic construction strategies for direct methanol utilization in Saccharomyces cerevisiae. Bioresour Technol. 2017;245:1407–1412. doi: 10.1016/j.biortech.2017.05.100. [DOI] [PubMed] [Google Scholar]
- Dandi ND, Dandi BN, Chaudhari AB. Bioprospecting of thermo- and osmo-tolerant fungi from mango pulp–peel compost for bioethanol production. Antonie Van Leeuwenhoek. 2013;103:723–736. doi: 10.1007/s10482-012-9854-4. [DOI] [PubMed] [Google Scholar]
- de Jong B, Siewers V, Nielsen J. Systems biology of yeast: enabling technology for development of cell factories for production of advanced biofuels. Curr Opin Biotechnol. 2012;23:624–630. doi: 10.1016/j.copbio.2011.11.021. [DOI] [PubMed] [Google Scholar]
- de la Torre I, Acedos MG, Esteban J, Santos VE, Ladero M (2019) Utilisation / upgrading of orange peel waste from a biological biorefinery perspective. Appl Microbiol Biotechnol 103:5975–5991. 10.1007/s00253-019-09929-2 [DOI] [PubMed]
- Dias PJ, Teixeira MC, Telo JP, Sá-Correia I. Insights into the mechanisms of toxicity and tolerance to the agricultural fungicide mancozeb in yeast, as suggested by a chemogenomic approach. Omi A J Integr Biol. 2010;14:211–227. doi: 10.1089/omi.2009.0134. [DOI] [PubMed] [Google Scholar]
- Dien BS, Kurtzman CP, Saha BC, Bothast RJ. Screening forl-arabinose fermenting yeasts. Appl Biochem Biotechnol. 1996;57–58:233–242. doi: 10.1007/BF02941704. [DOI] [PubMed] [Google Scholar]
- Do DTH, Theron CW, Fickers P. Organic wastes as feedstocks for non-conventional yeast-based bioprocesses. Microorganisms. 2019;7:1–22. doi: 10.3390/microorganisms7080229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- dos Santos SC. Yeast toxicogenomics: genome-wide responses to chemical stresses with impact in environmental health, pharmacology, and biotechnology. Front Genet. 2012;3:1–17. doi: 10.3389/fgene.2012.00063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- dos Santos SC, Sá-Correia I. Yeast toxicogenomics: lessons from a eukaryotic cell model and cell factory. Curr Opin Biotechnol. 2015;33:183–191. doi: 10.1016/j.copbio.2015.03.001. [DOI] [PubMed] [Google Scholar]
- Du C, Li Y, Zhao X, Pei X, Yuan W, Bai F, Jiang Y. The production of ethanol from lignocellulosic biomass by Kluyveromyces marxianus CICC 1727-5 and Spathaspora passalidarum ATCC MYA-4345. Appl Microbiol Biotechnol. 2019;103:2845–2855. doi: 10.1007/s00253-019-09625-1. [DOI] [PubMed] [Google Scholar]
- Edwards MC, Doran-Peterson J. Pectin-rich biomass as feedstock for fuel ethanol production. Appl Microbiol Biotechnol. 2012;95:565–575. doi: 10.1007/s00253-012-4173-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- European Parliament (2009) Directive 2009/128/EC of the European Parliament and the council of 21 October 2009 establishing a framework for community action to achieve the sustainable use of pesticides. October 309:71–86 . 10.3000/17252555.L_2009.309
- Fletcher E, Krivoruchko A, Nielsen J. Industrial systems biology and its impact on synthetic biology of yeast cell factories. Biotechnol Bioeng. 2016;113:1164–1170. doi: 10.1002/bit.25870. [DOI] [PubMed] [Google Scholar]
- Gama R, Van Dyk JS, Pletschke BI. Optimisation of enzymatic hydrolysis of apple pomace for production of biofuel and biorefinery chemicals using commercial enzymes. 3 Biotech. 2015;5:1075–1087. doi: 10.1007/s13205-015-0312-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gancedo JM. Carbon catabolite repression in yeast. Eur J Biochem. 1992;206:297–313. doi: 10.1111/j.1432-1033.1992.tb16928.x. [DOI] [PubMed] [Google Scholar]
- Gao S, Tong Y, Wen Z, Zhu L, Ge M, Chen D, Jiang Y, Yang S. Multiplex gene editing of the Yarrowia lipolytica genome using the CRISPR-Cas9 system. J Ind Microbiol Biotechnol. 2016;43:1085–1093. doi: 10.1007/s10295-016-1789-8. [DOI] [PubMed] [Google Scholar]
- Gao M, Ploessl D, Shao Z. Enhancing the co-utilization of biomass-derived mixed sugars by yeasts. Front Microbiol. 2019;9:1–21. doi: 10.3389/fmicb.2018.03264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- González-Siso MI, Touriño A, Vizoso Á, Pereira-Rodríguez Á, Rodríguez-Belmonte E, Becerra M, Cerdán ME. Improved bioethanol production in an engineered Kluyveromyces lactis strain shifted from respiratory to fermentative metabolism by deletion of NDI 1. Microb Biotechnol. 2015;8:319–330. doi: 10.1111/1751-7915.12160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grohmann K, Bothast RJ (1994) Pectin-rich residues generated by processing of citrus fruits, apples, and sugar beets. In: Enzymatic Conversion of Biomass for Fuels Production. pp 372–390
- Grohmann K, Manthey JA, Cameron RG, Buslig BS. Purification of citrus peel juice and molasses. J Agric Food Chem. 1999;47:4859–4867. doi: 10.1021/jf9903049. [DOI] [PubMed] [Google Scholar]
- Guerreiro J, Sampaio-Marques B, Soares R, Coelho A, Leao C, Ludovico P, Sa-Correia I. Mitochondrial proteomics of the acetic acid - induced programmed cell death response in a highly tolerant Zygosaccharomyces bailii - derived hybrid strain. Microb Cell. 2016;3:65–78. doi: 10.15698/mic2016.02.477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Günan Yücel H, Aksu Z. Ethanol fermentation characteristics of Pichia stipitis yeast from sugar beet pulp hydrolysate: use of new detoxification methods. Fuel. 2015;158:793–799. doi: 10.1016/j.fuel.2015.06.016. [DOI] [Google Scholar]
- Guo G, Hsu D, Chen W-H, Chen W, Hwang W. Characterization of enzymatic saccharification for acid-pretreated lignocellulosic materials with different lignin composition. Enzym Microb Technol. 2009;45:80–87. doi: 10.1016/j.enzmictec.2009.05.012. [DOI] [Google Scholar]
- Guo M, Cheng S, Chen G, Chen J. Improvement of lipid production in oleaginous yeast Rhodosporidium toruloides by ultraviolet mutagenesis. Eng Life Sci. 2019;19:548–556. doi: 10.1002/elsc.201800203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta SK, Shukla P. Gene editing for cell engineering: trends and applications. Crit Rev Biotechnol. 2017;37:672–684. doi: 10.1080/07388551.2016.1214557. [DOI] [PubMed] [Google Scholar]
- Hamley-Bennett C, Lye GJ, Leak DJ. Selective fractionation of sugar beet pulp for release of fermentation and chemical feedstocks; optimisation of thermo-chemical pre-treatment. Bioresour Technol. 2016;209:259–264. doi: 10.1016/j.biortech.2016.02.131. [DOI] [PubMed] [Google Scholar]
- Hicks RH, Sze Y, Chuck CJ, Henk DA (2020) Enhanced inhibitor tolerance and increased lipid productivity through adaptive laboratory evolution in the oleaginous yeast Metshnikowia pulcherrima. bioRxiv 2020.02.17.952291 . 10.1101/2020.02.17.952291
- Hong K-K, Nielsen J. Metabolic engineering of Saccharomyces cerevisiae: a key cell factory platform for future biorefineries. Cell Mol Life Sci. 2012;69:2671–2690. doi: 10.1007/s00018-012-0945-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang X-F, Liu J-N, Lu L-J, Peng K-M, Yang G-X, Liu J. Culture strategies for lipid production using acetic acid as sole carbon source by Rhodosporidium toruloides. Bioresour Technol. 2016;206:141–149. doi: 10.1016/j.biortech.2016.01.073. [DOI] [PubMed] [Google Scholar]
- Huisjes EH, de Hulster E, van Dam JC, Pronk JT, van Maris AJA. Galacturonic acid inhibits the growth of Saccharomyces cerevisiae on galactose, xylose, and arabinose. Appl Environ Microbiol. 2012;78:5052–5059. doi: 10.1128/AEM.07617-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayani RS, Saxena S, Gupta R. Microbial pectinolytic enzymes: a review. Process Biochem. 2005;40:2931–2944. doi: 10.1016/j.procbio.2005.03.026. [DOI] [Google Scholar]
- Jeon WY, Shim WY, Lee SH, Choi JH, Kim JH. Effect of heterologous xylose transporter expression in Candida tropicalis on xylitol production rate. Bioprocess Biosyst Eng. 2013;36:809–817. doi: 10.1007/s00449-013-0907-5. [DOI] [PubMed] [Google Scholar]
- Jeong D, Ye S, Park H, Kim SR. Simultaneous fermentation of galacturonic acid and five-carbon sugars by engineered Saccharomyces cerevisiae. Bioresour Technol. 2020;295:122259. doi: 10.1016/j.biortech.2019.122259. [DOI] [PubMed] [Google Scholar]
- Ji H, Zhuge B, Zong H, Lu X, Fang H, Zhuge J. Role of CgHOG1 in stress responses and glycerol overproduction of Candida glycerinogenes. Curr Microbiol. 2016;73:827–833. doi: 10.1007/s00284-016-1132-7. [DOI] [PubMed] [Google Scholar]
- John I, Muthukumar K, Arunagiri A. A review on the potential of citrus waste for D-limonene, pectin, and bioethanol production. Int J Green Energy. 2017;14:599–612. doi: 10.1080/15435075.2017.1307753. [DOI] [Google Scholar]
- Johnson EA. Biotechnology of non-Saccharomyces yeasts—the basidiomycetes. Appl Microbiol Biotechnol. 2013;97:7563–7577. doi: 10.1007/s00253-013-5046-z. [DOI] [PubMed] [Google Scholar]
- Johnson EA. Biotechnology of non-Saccharomyces yeasts—the ascomycetes. Appl Microbiol Biotechnol. 2013;97:503–517. doi: 10.1007/s00253-012-4497-y. [DOI] [PubMed] [Google Scholar]
- Johnson EA, Echavarri-Erasun C (2011) Yeast biotechnology. In: The Yeasts. Elsevier, pp 21–44
- Joshi VK, Sharma R, Girdher A. Production and evaluation of biocolour (carotenoids) from Rhodotorula using apple pomace: effect of composition of different nitrogen sources and methods of cell disruption. Int J Food Ferment Technol. 2013;3:127. doi: 10.5958/2277-9396.2014.00340.7. [DOI] [Google Scholar]
- Kalia VC, Saini AK. Metabolic engineering for bioactive compounds. Singapore: Springer Singapore; 2017. [Google Scholar]
- Kaur Sandhu S, Singh Oberoi H, Singh Dhaliwal S, Babbar N, Kaur U, Nanda D, Kumar D. Ethanol production from Kinnow mandarin (Citrus reticulata) peels via simultaneous saccharification and fermentation using crude enzyme produced by Aspergillus oryzae and the thermotolerant Pichia kudriavzevii strain. Ann Microbiol. 2012;62:655–666. doi: 10.1007/s13213-011-0302-x. [DOI] [Google Scholar]
- Kayikci Ö, Nielsen J. Glucose repression in Saccharomyces cerevisiae. FEMS Yeast Res. 2015;15:fov068. doi: 10.1093/femsyr/fov068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein M, Swinnen S, Thevelein JM, Nevoigt E. Glycerol metabolism and transport in yeast and fungi: established knowledge and ambiguities. Environ Microbiol. 2017;19:878–893. doi: 10.1111/1462-2920.13617. [DOI] [PubMed] [Google Scholar]
- Koutinas M, Patsalou M, Stavrinou S, Vyrides I. High temperature alcoholic fermentation of orange peel by the newly isolated thermotolerant Pichia kudriavzevii KVMP10. Lett Appl Microbiol. 2016;62:75–83. doi: 10.1111/lam.12514. [DOI] [PubMed] [Google Scholar]
- Kręgiel D, Pawlikowska E, Antolak H (2017) Non-conventional yeasts in fermentation processes: potentialities and limitations. Old Yeasts - New Quest. 10.5772/intechopen.70404
- Kühnel S (2011) Characterization of cell wall degrading enzymes from Chrysosporium lucknowense C1 and their use to degrade sugar beet pulp. Wageningen University, NL
- Kurtzman CP, Robnett CJ. Systematics of methanol assimilating yeasts and neighboring taxa from multigene sequence analysis and the proposal of Peterozyma gen. nov., a new member of the Saccharomycetales. FEMS Yeast Res. 2010;10:353–361. doi: 10.1111/j.1567-1364.2010.00625.x. [DOI] [PubMed] [Google Scholar]
- Lane S, Xu H, Oh EJ, Kim H, Lesmana A, Jeong D, Zhang G, Tsai C-S, Jin Y-S, Kim SR. Glucose repression can be alleviated by reducing glucose phosphorylation rate in Saccharomyces cerevisiae. Sci Rep. 2018;8:2613. doi: 10.1038/s41598-018-20804-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leandro MJ, Gonçalves P, Spencer-Martins I. Two glucose/xylose transporter genes from the yeast Candida intermedia: first molecular characterization of a yeast xylose–H+ symporter. Biochem J. 2006;395:543–549. doi: 10.1042/BJ20051465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee M-H, Lin J-J, Lin Y-J, Chang J-J, Ke H-M, Fan W-L, Wang T-Y, Li W-H. Genome-wide prediction of CRISPR/Cas9 targets in Kluyveromyces marxianus and its application to obtain a stable haploid strain. Sci Rep. 2018;8:7305. doi: 10.1038/s41598-018-25366-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legrand G (2005) The correct use of pressed beet pulp. Collect “Guides Tech Irbab/Kbivb 46
- Leijdekkers AGM, Bink JPM, Geutjes S, Schols HA, Gruppen H. Enzymatic saccharification of sugar beet pulp for the production of galacturonic acid and arabinose; a study on the impact of the formation of recalcitrant oligosaccharides. Bioresour Technol. 2013;128:518–525. doi: 10.1016/j.biortech.2012.10.126. [DOI] [PubMed] [Google Scholar]
- Lian J, Mishra S, Zhao H. Recent advances in metabolic engineering of Saccharomyces cerevisiae: new tools and their applications. Metab Eng. 2018;50:85–108. doi: 10.1016/j.ymben.2018.04.011. [DOI] [PubMed] [Google Scholar]
- Liguori R, Faraco V. Biological processes for advancing lignocellulosic waste biorefinery by advocating circular economy. Bioresour Technol. 2016;215:13–20. doi: 10.1016/j.biortech.2016.04.054. [DOI] [PubMed] [Google Scholar]
- Limayem A, Ricke SC. Lignocellulosic biomass for bioethanol production: current perspectives, potential issues and future prospects. Prog Energy Combust Sci. 2012;38:449–467. doi: 10.1016/j.pecs.2012.03.002. [DOI] [Google Scholar]
- Limtong S, Srisuk N, Yongmanitchai W, Yurimoto H, Nakase T. Ogataea chonburiensis sp. nov. and Ogataea nakhonphanomensis sp. nov., thermotolerant, methylotrophic yeast species isolated in Thailand, and transfer of Pichia siamensis and Pichia thermomethanolica to the genus Ogataea. Int J Syst Evol Microbiol. 2008;58:302–307. doi: 10.1099/ijs.0.65380-0. [DOI] [PubMed] [Google Scholar]
- Liu Z, Ho S-H, Hasunuma T, Chang J-S, Ren N-Q, Kondo A. Recent advances in yeast cell-surface display technologies for waste biorefineries. Bioresour Technol. 2016;215:324–333. doi: 10.1016/j.biortech.2016.03.132. [DOI] [PubMed] [Google Scholar]
- Liu L, You Y, Deng H, Guo Y, Meng Y. Promoting hydrolysis of apple pomace by pectinase and cellulase to produce microbial oils using engineered Yarrowia lipolytica. Biomass Bioenergy. 2019;126:62–69. doi: 10.1016/j.biombioe.2019.04.025. [DOI] [Google Scholar]
- Liu N, Santala S, Stephanopoulos G. Mixed carbon substrates: a necessary nuisance or a missed opportunity? Curr Opin Biotechnol. 2020;62:15–21. doi: 10.1016/j.copbio.2019.07.003. [DOI] [PubMed] [Google Scholar]
- Löbs A-K, Schwartz C, Wheeldon I. Genome and metabolic engineering in non-conventional yeasts: current advances and applications. Synth Syst Biotechnol. 2017;2:198–207. doi: 10.1016/j.synbio.2017.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Löbs AK, Schwartz C, Thorwall S, Wheeldon I. Highly multiplexed CRISPRi repression of respiratory functions enhances mitochondrial localized ethyl acetate biosynthesis in Kluyveromyces marxianus. ACS Synth Biol. 2018;7:2647–2655. doi: 10.1021/acssynbio.8b00331. [DOI] [PubMed] [Google Scholar]
- Loman AA, Islam SMM, Ju L-K. Production of arabitol from enzymatic hydrolysate of soybean flour by Debaryomyces hansenii fermentation. Appl Microbiol Biotechnol. 2018;102:641–653. doi: 10.1007/s00253-017-8626-5. [DOI] [PubMed] [Google Scholar]
- Martínez O, Sánchez A, Font X, Barrena R. Valorization of sugarcane bagasse and sugar beet molasses using Kluyveromyces marxianus for producing value-added aroma compounds via solid-state fermentation. J Clean Prod. 2017;158:8–17. doi: 10.1016/j.jclepro.2017.04.155. [DOI] [Google Scholar]
- Martini C, Tauk-Tornisielo SM, Codato CB, Bastos RG, Ceccato-Antonini SR. A strain of Meyerozyma guilliermondii isolated from sugarcane juice is able to grow and ferment pentoses in synthetic and bagasse hydrolysate media. World J Microbiol Biotechnol. 2016;32:80. doi: 10.1007/s11274-016-2036-1. [DOI] [PubMed] [Google Scholar]
- Matsubara T, Hamada S, Wakabayashi A, Kishida M. Fermentative production of l-galactonate by using recombinant Saccharomyces cerevisiae containing the endogenous galacturonate reductase gene from Cryptococcus diffluens. J Biosci Bioeng. 2016;122:639–644. doi: 10.1016/j.jbiosc.2016.05.002. [DOI] [PubMed] [Google Scholar]
- Merz M, Appel D, Berends P, Rabe S, Blank I, Stressler T, Fischer L. Batch-to-batch variation and storage stability of the commercial peptidase preparation Flavourzyme in respect of key enzyme activities and its influence on process reproducibility. Eur Food Res Technol. 2016;242:1005–1012. doi: 10.1007/s00217-015-2606-8. [DOI] [Google Scholar]
- Micard V, Renard CMGC, Thibault J-F. Enzymatic saccharification of sugar-beet pulp. Enzym Microb Technol. 1996;19:162–170. doi: 10.1016/0141-0229(95)00224-3. [DOI] [Google Scholar]
- Mira NP, Palma M, Guerreiro JF, Sá-Correia I. Genome-wide identification of Saccharomyces cerevisiae genes required for tolerance to acetic acid. Microb Cell Factories. 2010;9:79. doi: 10.1186/1475-2859-9-79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mira NP, Teixeira MC, Sá-Correia I. Adaptive response and tolerance to weak acids in Saccharomyces cerevisiae : a genome-wide view. Omi A J Integr Biol. 2010;14:525–540. doi: 10.1089/omi.2010.0072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mira NP, Henriques SF, Keller G, Teixeira MC, Matos RG, Arraiano CM, Winge DR, Sá-Correia I. Identification of a DNA-binding site for the transcription factor Haa1, required for Saccharomyces cerevisiae response to acetic acid stress. Nucleic Acids Res. 2011;39:6896–6907. doi: 10.1093/nar/gkr228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mira NP, Munsterkotter M, Dias-Valada F, Santos J, Palma M, Roque FC, Guerreiro JF, Rodrigues F, Sousa MJ, Leao C, Guldener U, Sa-Correia I. The genome sequence of the highly acetic acid-tolerant Zygosaccharomyces bailii-derived interspecies hybrid strain ISA1307, isolated from a sparkling wine plant. DNA Res. 2014;21:299–313. doi: 10.1093/dnares/dst058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohnen D. Pectin structure and biosynthesis. Curr Opin Plant Biol. 2008;11:266–277. doi: 10.1016/j.pbi.2008.03.006. [DOI] [PubMed] [Google Scholar]
- Monteiro PT, Oliveira J, Pais P, Antunes M, Palma M, Cavalheiro M, Galocha M, Godinho CP, Martins LC, Bourbon N, Mota MN, Ribeiro RA, Viana R, Sá-Correia I, Teixeira MC. YEASTRACT+: a portal for cross-species comparative genomics of transcription regulation in yeasts. Nucleic Acids Res. 2020;48:642–649. doi: 10.1093/nar/gkz859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mukherjee V, Radecka D, Aerts G, Verstrepen KJ, Lievens B, Thevelein JM. Phenotypic landscape of non-conventional yeast species for different stress tolerance traits desirable in bioethanol fermentation. Biotechnol Biofuels. 2017;10:216. doi: 10.1186/s13068-017-0899-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller-Maatsch J, Bencivenni M, Caligiani A, Tedeschi T, Bruggeman G, Bosch M, Petrusan J, Van Droogenbroeck B, Elst K, Sforza S. Pectin content and composition from different food waste streams. Food Chem. 2016;201:37–45. doi: 10.1016/j.foodchem.2016.01.012. [DOI] [PubMed] [Google Scholar]
- Mussatto SI, Silva CJSM, Roberto IC. Fermentation performance of Candida guilliermondii for xylitol production on single and mixed substrate media. Appl Microbiol Biotechnol. 2006;72:681–686. doi: 10.1007/s00253-006-0372-z. [DOI] [PubMed] [Google Scholar]
- Nambu-Nishida Y, Nishida K, Hasunuma T, Kondo A. Development of a comprehensive set of tools for genome engineering in a cold- and thermo-tolerant Kluyveromyces marxianus yeast strain. Sci Rep. 2017;7:8993. doi: 10.1038/s41598-017-08356-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen J. Yeast systems biology: model organism and cell factory. Biotechnol J. 2019;14:1800421. doi: 10.1002/biot.201800421. [DOI] [PubMed] [Google Scholar]
- Nielsen J, Keasling JD. Engineering cellular metabolism. Cell. 2016;164:1185–1197. doi: 10.1016/j.cell.2016.02.004. [DOI] [PubMed] [Google Scholar]
- Nigam P, Vogel M. Bioconversion of sugar industry by-products—molasses and sugar beet pulp for single cell protein production by yeasts. Biomass Bioenergy. 1991;1:339–345. doi: 10.1016/0961-9534(91)90014-4. [DOI] [Google Scholar]
- Nurcholis M, Lertwattanasakul N, Rodrussamee N, Kosaka T, Murata M, Yamada M. Integration of comprehensive data and biotechnological tools for industrial applications of Kluyveromyces marxianus. Appl Microbiol Biotechnol. 2020;104:475–488. doi: 10.1007/s00253-019-10224-3. [DOI] [PubMed] [Google Scholar]
- Palma M, Sá-Correia I. Physiological genomics of the highly weak-acid-tolerant food spoilage yeasts of Zygosaccharomyces bailii sensu lato. Prog Mol Subcell Biol. 2019;58:85–109. doi: 10.1007/978-3-030-13035-0_4. [DOI] [PubMed] [Google Scholar]
- Palma M, Roque F d C, Guerreiro JF, Mira NP, Queiroz L, Sá-Correia I. Search for genes responsible for the remarkably high acetic acid tolerance of a Zygosaccharomyces bailii-derived interspecies hybrid strain. BMC Genomics. 2015;16:1070. doi: 10.1186/s12864-015-2278-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palma M, Dias PJ, Roque F d C, Luzia L, Guerreiro JF, Sá-Correia I. The Zygosaccharomyces bailii transcription factor Haa1 is required for acetic acid and copper stress responses suggesting subfunctionalization of the ancestral bifunctional protein Haa1/Cup2. BMC Genomics. 2017;18:75. doi: 10.1186/s12864-016-3443-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palma M, Guerreiro JF, Sá-Correia I. Adaptive response and tolerance to acetic acid in Saccharomyces cerevisiae and Zygosaccharomyces bailii: a physiological genomics perspective. Front Microbiol. 2018;9:1–16. doi: 10.3389/fmicb.2018.00274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmqvist E, Hahn-Hägerdal B. Fermentation of lignocellulosic hydrolysates. II: inhibitors and mechanisms of inhibition. Bioresour Technol. 2000;74:25–33. doi: 10.1016/S0960-8524(99)00161-3. [DOI] [Google Scholar]
- Papapetridis I, Verhoeven MD, Wiersma SJ, Goudriaan M, van Maris AJA, Pronk JT. Laboratory evolution for forced glucose-xylose co-consumption enables identification of mutations that improve mixed-sugar fermentation by xylose-fermenting Saccharomyces cerevisiae. FEMS Yeast Res. 2018;18:1–17. doi: 10.1093/femsyr/foy056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park Y-K, Nicaud J-M, Ledesma-Amaro R. The engineering potential of Rhodosporidium toruloides as a workhorse for biotechnological applications. Trends Biotechnol. 2018;36:304–317. doi: 10.1016/j.tibtech.2017.10.013. [DOI] [PubMed] [Google Scholar]
- Parmar I, Rupasinghe HPV. Optimization of dilute acid-based pretreatment and application of laccase on apple pomace. Bioresour Technol. 2012;124:433–439. doi: 10.1016/j.biortech.2012.07.030. [DOI] [PubMed] [Google Scholar]
- Peters D. Raw materials. Adv Biochem Eng Biotechnol. 2006;105:1–30. doi: 10.1007/10_031. [DOI] [PubMed] [Google Scholar]
- Pfeiffer T, Morley A. An evolutionary perspective on the Crabtree effect. Front Mol Biosci. 2014;1:1–6. doi: 10.3389/fmolb.2014.00017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Protzko RJ, Latimer LN, Martinho Z, de Reus E, Seibert T, Benz JP, Dueber JE. Engineering Saccharomyces cerevisiae for co-utilization of d-galacturonic acid and d-glucose from citrus peel waste. Nat Commun. 2018;9:5059. doi: 10.1038/s41467-018-07589-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Protzko RJ, Hach CA, Coradetti ST, Hackhofer MA, Magosch S, Thieme N, Geiselman GM, Arkin AP, Skerker JM, Dueber JE, Benz JP (2019) Genomewide and Enzymatic Analysis Reveals Efficient D-Galacturonic Acid Metabolism in the Basidiomycete Yeast Rhodosporidium toruloides. mSystems 4: . 10.1128/mSystems.00389-19 [DOI] [PMC free article] [PubMed]
- Qiao K, Wasylenko TM, Zhou K, Xu P, Stephanopoulos G. Lipid production in Yarrowia lipolytica is maximized by engineering cytosolic redox metabolism. Nat Biotechnol. 2017;35:173–177. doi: 10.1038/nbt.3763. [DOI] [PubMed] [Google Scholar]
- Quemener B, Lahaye M, Thibault JF. Studies on the simultaneous determination of acidic and neutral sugars of plant cell wall materials by HPLC of their methyl glycosides after combined methanolysis and enzymic prehydrolysis. Carbohydr Polym. 1993;20:87–94. doi: 10.1016/0144-8617(93)90082-F. [DOI] [Google Scholar]
- Radecka D, Mukherjee V, Mateo RQ, Stojiljkovic M, Foulquié-Moreno MR, Thevelein JM. Looking beyond Saccharomyces: the potential of non-conventional yeast species for desirable traits in bioethanol fermentation. FEMS Yeast Res. 2015;15:53. doi: 10.1093/femsyr/fov053. [DOI] [PubMed] [Google Scholar]
- Raschmanová H, Weninger A, Glieder A, Kovar K, Vogl T. Implementing CRISPR-Cas technologies in conventional and non-conventional yeasts: current state and future prospects. Biotechnol Adv. 2018;36:641–665. doi: 10.1016/j.biotechadv.2018.01.006. [DOI] [PubMed] [Google Scholar]
- Ratledge C (2010) Single cell oils for the 21st century. In: Single Cell Oils, Second Edi. Elsevier, pp 3–26
- Rebello S, Abraham A, Madhavan A, Sindhu R, Binod P, Babu AK, Aneesh EM, Pandey A. Non-conventional yeast cell factories for sustainable bioprocesses. FEMS Microbiol Lett. 2018;365:1–10. doi: 10.1093/femsle/fny222. [DOI] [PubMed] [Google Scholar]
- Richard P, Hilditch S. D-galacturonic acid catabolism in microorganisms and its biotechnological relevance. Appl Microbiol Biotechnol. 2009;82:597–604. doi: 10.1007/s00253-009-1870-6. [DOI] [PubMed] [Google Scholar]
- Rodrigues F, Sousa MJ, Ludovico P, Santos H, Côrte-Real M, Leão C. The fate of acetic acid during glucose co-metabolism by the spoilage yeast Zygosaccharomyces bailii. PLoS One. 2012;7:e52402. doi: 10.1371/journal.pone.0052402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodríguez Madrera R, Pando Bedriñana R, Suárez Valles B. Production and characterization of aroma compounds from apple pomace by solid-state fermentation with selected yeasts. LWT Food Sci Technol. 2015;64:1342–1353. doi: 10.1016/j.lwt.2015.07.056. [DOI] [Google Scholar]
- Rozpędowska E, Hellborg L, Ishchuk OP, Orhan F, Galafassi S, Merico A, Woolfit M, Compagno C, Piškur J. Parallel evolution of the make–accumulate–consume strategy in Saccharomyces and Dekkera yeasts. Nat Commun. 2011;2:302. doi: 10.1038/ncomms1305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sahota PP, Kaur N. Characterization of enzyme naringinase and the production of debittered low alcoholic kinnow (Citrus raticulata blanco) beverage. Int J Adv Res. 2015;3:1220–1233. [Google Scholar]
- Sakai T, Okushima M. Microbial production of pectin from citrus peel. Appl Environ Microbiol. 1980;39:908–912. doi: 10.1128/AEM.39.4.908-912.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakai T, Sakamoto T, Hallaert J, Vandamme EJ (1993) Pectin, pectinase, and protopectinase: production, properties, and applications. Adv Appl Microbiol 213–294 [DOI] [PubMed]
- Saravanakumar D, Spadaro D, Garibaldi A, Gullino ML. Detection of enzymatic activity and partial sequence of a chitinase gene in Metschnikowia pulcherrima strain MACH1 used as post-harvest biocontrol agent. Eur J Plant Pathol. 2009;123:183–193. doi: 10.1007/s10658-008-9355-5. [DOI] [Google Scholar]
- Satyanarayana T, Kunze G. Yeast biotechnology: diversity and applications. Dordrecht: Springer Netherlands; 2009. [Google Scholar]
- Schiewer S, Patil SB. Pectin-rich fruit wastes as biosorbents for heavy metal removal: equilibrium and kinetics. Bioresour Technol. 2008;99:1896–1903. doi: 10.1016/j.biortech.2007.03.060. [DOI] [PubMed] [Google Scholar]
- Schirmer-Michel ÂC, Flôres SH, Hertz PF, Matos GS, Ayub MAZ. Production of ethanol from soybean hull hydrolysate by osmotolerant Candida guilliermondii NRRL Y-2075. Bioresour Technol. 2008;99:2898–2904. doi: 10.1016/j.biortech.2007.06.042. [DOI] [PubMed] [Google Scholar]
- Schirmer-Michel ÂC, Flôres SH, Hertz PF, Ayub MAZ. Effect of oxygen transfer rates on alcohols production by Candida guilliermondii cultivated on soybean hull hydrolysate. J Chem Technol Biotechnol. 2009;84:223–228. doi: 10.1002/jctb.2028. [DOI] [Google Scholar]
- Schmitz K, Protzko R, Zhang L, Benz JP. Spotlight on fungal pectin utilization—from phytopathogenicity to molecular recognition and industrial applications. Appl Microbiol Biotechnol. 2019;103:2507–2524. doi: 10.1007/s00253-019-09622-4. [DOI] [PubMed] [Google Scholar]
- Seiboth B, Metz B. Fungal arabinan and L-arabinose metabolism. Appl Microbiol Biotechnol. 2011;89:1665–1673. doi: 10.1007/s00253-010-3071-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sellick CA, Campbell RN, Reece RJ (2008) Galactose metabolism in yeast—structure and regulation of the leloir pathway enzymes and the genes encoding them. In: International Review of Cell and Molecular Biology. pp 111–150 [DOI] [PubMed]
- Serrat M, Bermúdez RC, Villa TG. Polygalacturonase and ethanol production in Kluyveromyces marxianus: potential use of polygalacturonase in foodstuffs. Appl Biochem Biotechnol. 2004;117:49–64. doi: 10.1385/ABAB:117:1:49. [DOI] [PubMed] [Google Scholar]
- Simpson-Lavy K, Kupiec M. Carbon catabolite repression in yeast is not limited to glucose. Sci Rep. 2019;9:6491. doi: 10.1038/s41598-019-43032-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh G, Sinha S, Bandyopadhyay KK, Lawrence M, Paul D. Triauxic growth of an oleaginous red yeast Rhodosporidium toruloides on waste ‘extract’ for enhanced and concomitant lipid and β-carotene production. Microb Cell Factories. 2018;17:182. doi: 10.1186/s12934-018-1026-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siripong W, Wolf P, Kusumoputri TP, Downes JJ, Kocharin K, Tanapongpipat S, Runguphan W. Metabolic engineering of Pichia pastoris for production of isobutanol and isobutyl acetate. Biotechnol Biofuels. 2018;11:1. doi: 10.1186/s13068-017-1003-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sitepu I, Selby T, Lin T, Zhu S, Boundy-Mills K. Carbon source utilization and inhibitor tolerance of 45 oleaginous yeast species. J Ind Microbiol Biotechnol. 2014;41:1061–1070. doi: 10.1007/s10295-014-1447-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skrbic B, Durisic-Mladenovic N, Macvanin N. Determination of metal contents in sugar beet (Beta vulgaris) and its products: empirical and chemometrical approach. Food Sci Technol Res. 2010;16:123–134. doi: 10.3136/fstr.16.123. [DOI] [Google Scholar]
- Souffriau B, Den Abt T, Thevelein JM. Evidence for rapid uptake of d-galacturonic acid in the yeast Saccharomyces cerevisiae by a channel-type transport system. FEBS Lett. 2012;586:2494–2499. doi: 10.1016/j.febslet.2012.06.012. [DOI] [PubMed] [Google Scholar]
- Spagnuolo M, Yaguchi A, Blenner M. Oleaginous yeast for biofuel and oleochemical production. Curr Opin Biotechnol. 2019;57:73–81. doi: 10.1016/j.copbio.2019.02.011. [DOI] [PubMed] [Google Scholar]
- Subtil T, Boles E. Improving L-arabinose utilization of pentose fermenting Saccharomyces cerevisiae cells by heterologous expression of L-arabinose transporting sugar transporters. Biotechnol Biofuels. 2011;4:38. doi: 10.1186/1754-6834-4-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanimura A, Takashima M, Sugita T, Endoh R, Ohkuma M, Kishino S, Ogawa J, Shima J. Lipid production through simultaneous utilization of glucose, xylose, and l-arabinose by Pseudozyma hubeiensis: a comparative screening study. AMB Express. 2016;6:58. doi: 10.1186/s13568-016-0236-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taskin M, Ortucu S, Aydogan MN, Arslan NP. Lipid production from sugar beet molasses under non-aseptic culture conditions using the oleaginous yeast Rhodotorula glutinis TR29. Renew Energy. 2016;99:198–204. doi: 10.1016/j.renene.2016.06.060. [DOI] [Google Scholar]
- Teixeira MC, Duque P, Sá-Correia I. Environmental genomics: mechanistic insights into toxicity of and resistance to the herbicide 2,4-D. Trends Biotechnol. 2007;25:363–370. doi: 10.1016/j.tibtech.2007.06.002. [DOI] [PubMed] [Google Scholar]
- Teixeira MC, Mira NP, Sá-Correia I. A genome-wide perspective on the response and tolerance to food-relevant stresses in Saccharomyces cerevisiae. Curr Opin Biotechnol. 2011;22:150–156. doi: 10.1016/j.copbio.2010.10.011. [DOI] [PubMed] [Google Scholar]
- Tsukamoto J, Durán N, Tasic L. Nanocellulose and bioethanol production from orange waste using isolated microorganisms. J Braz Chem Soc. 2013;24:1537–1543. doi: 10.5935/0103-5053.20130195. [DOI] [Google Scholar]
- Vadkertiová R, Sláviková E. Metal tolerance of yeasts isolated from water, soil and plant environments. J Basic Microbiol. 2006;46:145–152. doi: 10.1002/jobm.200510609. [DOI] [PubMed] [Google Scholar]
- van der Klei IJ, Yurimoto H, Sakai Y, Veenhuis M. The significance of peroxisomes in methanol metabolism in methylotrophic yeast. Biochim Biophys Acta, Mol Cell Res. 2006;1763:1453–1462. doi: 10.1016/j.bbamcr.2006.07.016. [DOI] [PubMed] [Google Scholar]
- van Dijk M, Erdei B, Galbe M, Nygård Y, Olsson L. Strain-dependent variance in short-term adaptation effects of two xylose-fermenting strains of Saccharomyces cerevisiae. Bioresour Technol. 2019;292:121922. doi: 10.1016/j.biortech.2019.121922. [DOI] [PubMed] [Google Scholar]
- Vendruscolo F, Albuquerque PM, Streit F, Esposito E, Ninow JL. Apple pomace: a versatile substrate for biotechnological applications. Crit Rev Biotechnol. 2008;28:1–12. doi: 10.1080/07388550801913840. [DOI] [PubMed] [Google Scholar]
- Villas-Bôas SG, Esposito E, De Mendonça MM. Novel lignocellulolytic ability of Candida utilis during solid-substrate cultivation on apple pomace. World J Microbiol Biotechnol. 2002;18:541–545. doi: 10.1023/A:1016350612380. [DOI] [Google Scholar]
- Wagner JM, Alper HS. Synthetic biology and molecular genetics in non-conventional yeasts: current tools and future advances. Fungal Genet Biol. 2016;89:126–136. doi: 10.1016/j.fgb.2015.12.001. [DOI] [PubMed] [Google Scholar]
- Wamelink MMC, Struys EA, Jakobs C. The biochemistry, metabolism and inherited defects of the pentose phosphate pathway: a review. J Inherit Metab Dis. 2008;31:703–717. doi: 10.1007/s10545-008-1015-6. [DOI] [PubMed] [Google Scholar]
- Wang G-Y, Chi Z, Song B, Wang Z-P, Chi Z-M. High level lipid production by a novel inulinase-producing yeast Pichia guilliermondii Pcla22. Bioresour Technol. 2012;124:77–82. doi: 10.1016/j.biortech.2012.08.024. [DOI] [PubMed] [Google Scholar]
- Wang Y, Gong Z, Yang X, Shen H, Wang Q, Wang J, Zhao ZK. Microbial lipid production from pectin-derived carbohydrates by oleaginous yeasts. Process Biochem. 2015;50:1097–1102. doi: 10.1016/j.procbio.2015.04.014. [DOI] [Google Scholar]
- Weninger A, Glieder A, Vogl T. A toolbox of endogenous and heterologous nuclear localization sequences for the methylotrophic yeast Pichia pastoris. FEMS Yeast Res. 2015;15:fov082. doi: 10.1093/femsyr/fov082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wikiera A, Mika M, Starzyńska-Janiszewska A, Stodolak B. Development of complete hydrolysis of pectins from apple pomace. Food Chem. 2015;172:675–680. doi: 10.1016/j.foodchem.2014.09.132. [DOI] [PubMed] [Google Scholar]
- Williams PA (2011) Natural polymers: introduction and overview. In: RSC Polymer Chemistry Series. pp 1–14
- Wirth F, Goldani LZ. Epidemiology of Rhodotorula : an emerging pathogen. Interdiscip Perspect Infect Dis. 2012;2012:1–7. doi: 10.1155/2012/465717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wisselink HW, Toirkens MJ, del Rosario Franco Berriel M, Winkler AA, van Dijken JP, Pronk JT, van Maris AJA. Engineering of Saccharomyces cerevisiae for efficient anaerobic alcoholic fermentation of L-arabinose. Appl Environ Microbiol. 2007;73:4881–4891. doi: 10.1128/AEM.00177-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y, Shen X, Yuan Q, Yan Y. Metabolic engineering strategies for co-utilization of carbon sources in microbes. Bioengineering. 2016;3:10. doi: 10.3390/bioengineering3010010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yaguchi A, Spagnuolo M, Blenner M. Engineering yeast for utilization of alternative feedstocks. Curr Opin Biotechnol. 2018;53:122–129. doi: 10.1016/j.copbio.2017.12.003. [DOI] [PubMed] [Google Scholar]
- Yapo BM, Lerouge P, Thibault J-F, Ralet M-C. Pectins from citrus peel cell walls contain homogalacturonans homogenous with respect to molar mass, rhamnogalacturonan I and rhamnogalacturonan II. Carbohydr Polym. 2007;69:426–435. doi: 10.1016/j.carbpol.2006.12.024. [DOI] [Google Scholar]
- Yapo BM, Robert C, Etienne I, Wathelet B, Paquot M. Effect of extraction conditions on the yield, purity and surface properties of sugar beet pulp pectin extracts. Food Chem. 2007;100:1356–1364. doi: 10.1016/j.foodchem.2005.12.012. [DOI] [Google Scholar]
- Yasokawa D, Murata S, Iwahashi Y, Kitagawa E, Nakagawa R, Hashido T, Iwahashi H. Toxicity of methanol and formaldehyde towards Saccharomyces cerevisiae as assessed by DNA microarray analysis. Appl Biochem Biotechnol. 2010;160:1685–1698. doi: 10.1007/s12010-009-8684-y. [DOI] [PubMed] [Google Scholar]
- Ye S, Jeong D, Shon JC, Liu KH, Kim KH, Shin M, Kim SR. Deletion of PHO13 improves aerobic l-arabinose fermentation in engineered Saccharomyces cerevisiae. J Ind Microbiol Biotechnol. 2019;46:1725–1731. doi: 10.1007/s10295-019-02233-y. [DOI] [PubMed] [Google Scholar]
- Zhang GC, Liu JJ, Kong II, Kwak S, Jin YS. Combining C6 and C5 sugar metabolism for enhancing microbial bioconversion. Curr Opin Chem Biol. 2015;29:49–57. doi: 10.1016/j.cbpa.2015.09.008. [DOI] [PubMed] [Google Scholar]
- Zhao M, Shi D, Lu X, Zong H, Zhuge B, Ji H. Ethanol fermentation from non-detoxified lignocellulose hydrolysate by a multi-stress tolerant yeast Candida glycerinogenes mutant. Bioresour Technol. 2019;273:634–640. doi: 10.1016/j.biortech.2018.11.053. [DOI] [PubMed] [Google Scholar]
- Zhou W, Widmer W, Grohmann K (2008) Developments in ethanol production from citrus peel waste. Proc Fla State Hort Soc:307–310
- Zuin VG, Segatto ML, Ramin LZ. Plants as resources for organic molecules: facing the green and sustainable future today. Curr Opin Green Sustain Chem. 2018;9:1–7. doi: 10.1016/j.cogsc.2017.10.001. [DOI] [Google Scholar]