Significance
This study validates a path toward personalized medicine that utilizes a survey of the effective signaling state of tumors via posttranslational modification analysis as a diagnostic alternative to direct examination of genetic mutations. Here, this approach is utilized to identify biomarkers for Cdk5-driven tumors. The same approach could be adapted to identify markers of other tumor drivers. As the number of cancer types in which Cdk5 is implicated is rising and as drugs with the potential to selectively target Cdk5 are moved toward the clinic, the translational potential of this diagnostic–therapeutic coupled system will be broad.
Keywords: predictive biomarkers, neuroendocrine tumors, cyclin-dependent kinase 5
Abstract
Disparities in cancer patient responses have prompted widespread searches to identify differences in sensitive vs. nonsensitive populations and form the basis of personalized medicine. This customized approach is dependent upon the development of pathway-specific therapeutics in conjunction with biomarkers that predict patient responses. Here, we show that Cdk5 drives growth in subgroups of patients with multiple types of neuroendocrine neoplasms. Phosphoproteomics and high throughput screening identified phosphorylation sites downstream of Cdk5. These phosphorylation events serve as biomarkers and effectively pinpoint Cdk5-driven tumors. Toward achieving targeted therapy, we demonstrate that mouse models of neuroendocrine cancer are responsive to selective Cdk5 inhibitors and biomimetic nanoparticles are effective vehicles for enhanced tumor targeting and reduction of drug toxicity. Finally, we show that biomarkers of Cdk5-dependent tumors effectively predict response to anti-Cdk5 therapy in patient-derived xenografts. Thus, a phosphoprotein-based diagnostic assay combined with Cdk5-targeted therapy is a rational treatment approach for neuroendocrine malignancies.
Cyclin-dependent kinases (Cdks) are a family of proline-directed serine/threonine kinases that are required for progression of normal cell division. Typical Cdks are regulated through binding to cyclins, proteins which are expressed at varying levels at distinct stages of the cell cycle (1). As master regulators of cell division, Cdk1/2/4/6 currently serve as popular targets for cancer therapy development (2–4). Although Cdk5 shares ∼60% sequence identity with founding family members Cdk1 and Cdk2 (5), it possesses noncanonical features. Specifically, Cdk5 is not required for normal cell cycle progression and is not activated by cyclins. Cdk5 is regulated instead through interactions with cofactors p35 and p39 (6). Although Cdk5 is expressed in a broad number of tissues (5), its activators are mostly restricted to neuronal cells (7) where Cdk5 activity is important for central nervous system (CNS) development and cognitive processes such as learning and memory (8). Cleavage of p35 and p39 by calpain produces truncated activators, p25 and p29, respectively (9). These cleavage products have increased protein stability and mislocalize in cells due to removal of an N-terminal myristoylation site (10). In neurons, p25 aberrantly activates Cdk5 and has been linked to neurotoxicity, neuronal injury, and neurodegeneration (6).
Recent research suggests the aberrant activation of Cdk5 in nonneuronal cells can usurp signaling components involved in the cell cycle to drive proliferation (11). Expression of Cdk5 as well as p35 and/or p25 (p35/p25) has been shown for three types of neoplasms originating from neuroendocrine (NE) cells: medullary thyroid carcinoma (MTC) (12), small cell lung cancer (SCLC) (13), and pituitary adenomas (PAs) (14). In MTC, inhibition of Cdk5 activity decreases rates of cell growth (12, 15); in SCLC and PA, it decreases migration and invasion (16, 17). Expression of p25 in thyroid C cells produces MTC in mice (12), in part through alteration of traditional cell cycle regulatory components (18). Here we show that Cdk5 and p35/p25 expression may be an important driver of many types of NE cancer and that aberrant Cdk5 activity allows for a diagnostic-coupled treatment strategy that targets this protein kinase.
Results
Cdk5 in NE Neoplasms.
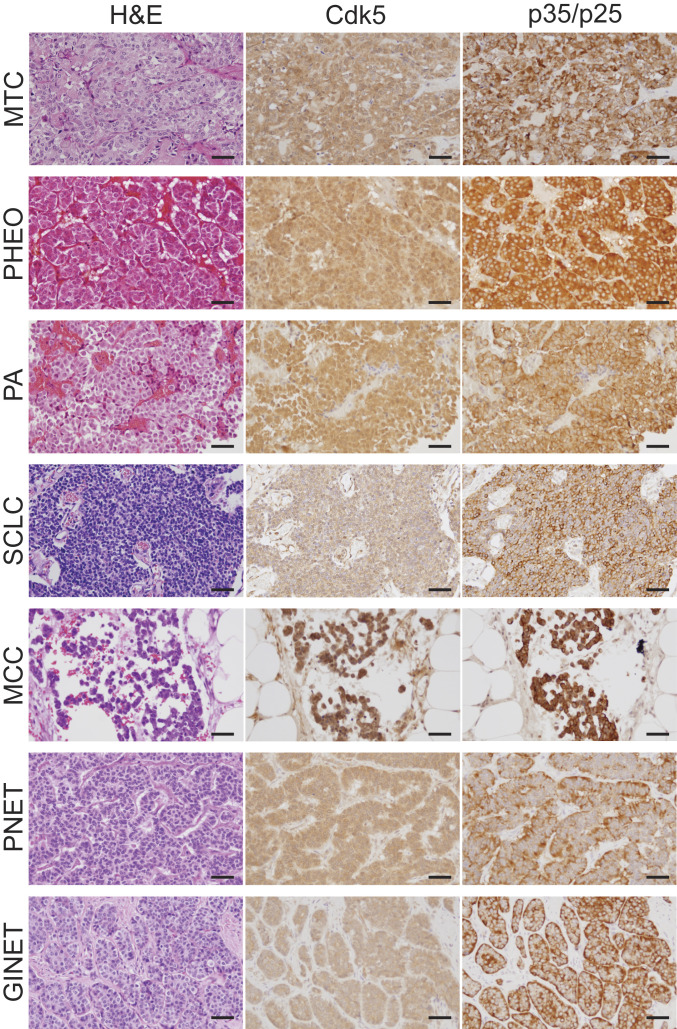
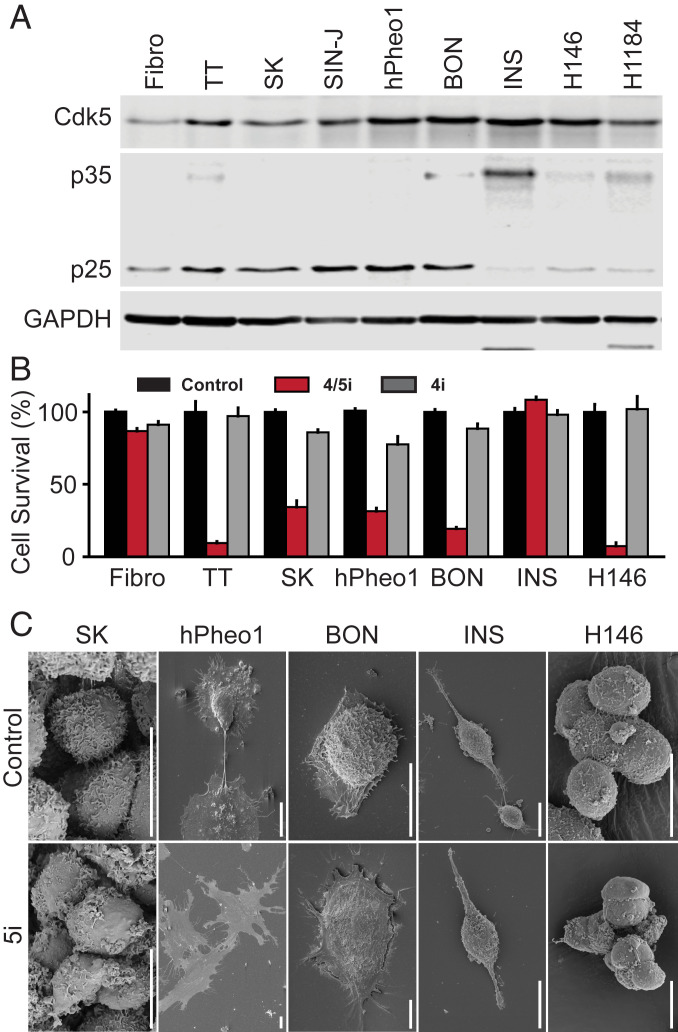
To better understand the potential role of Cdk5 across multiple forms of NE malignancies, we assessed expression of the kinase and its activating cofactors in as many different NE tumor and cancer types as we could obtain samples. Histological analysis demonstrated that Cdk5 and p35/p25 are present throughout various human NE neoplasms including MTC, pheochromocytoma (Pheo), PA, SCLC, Merkel cell carcinoma (MCC), pancreatic NE tumors (PNETs), and gastrointestinal NE tumors (GINETs) (Fig. 1). Furthermore, these proteins are present in cell lines derived from multiple types of NE neoplasms, including three human MTCs (TT, MTC-SK, and SIN-J), a human progenitor PHEO (hPheo1) (19), a human pancreatic carcinoid (BON), a rat insulinoma (INS), and two human SCLCs (H146 and H1184) (Fig. 2A and SI Appendix, Fig. S1A). Selective Cdk5 inhibition by Indolinone A (IndoA) blocked all human NE cancer cell growth more potently than it affected normal human fibroblasts or rat INS cells (Fig. 2B and SI Appendix, Fig. S2 A–H). NE cell lines treated with Cdk5 inhibitors typically showed flattening and smoothing of cell body consistent with a less malignant phenotype, except the rat INS line (Fig. 2C and SI Appendix, Fig. S1B). Interestingly, the aberrant Cdk5 activator, p25, was present in all human cell lines derived from naturally occurring tumors but not the rat INS line generated using irradiation (20) (Fig. 2A), indicating that the presence of p25 was responsible for driving aberrant proliferation downstream of Cdk5.
Fig. 1.
Cdk5 pathway components are present in human NE tumors. H&E stain, immunohistochemistry (IHC) for Cdk5, and IHC for p35/p25 in human NE tumors (Scale bars, 50 μm.)
Fig. 2.
Cdk5 promotes growth in human NE cells. (A) Immunoblot of Cdk5 pathway components in fibroblasts and NE cells. (B) Fibroblasts and NE cell lines were treated with 0.02% DMSO (control), 0.3 μM IndoA (4/5i), or 0.3 μM IndoB (4i) and monitored for effects on cell growth. Error bars represent SEM. (Full curves and IC50 values are reported in SI Appendix, Fig. S1). (C) Scanning electron microscopy of NE cells treated with control (0.02% DMSO) or 5i (hPheo1 and BON, 2 μM IndoA; SK and H146, 5 μM CP6813101) for 4 h. (Scale bars, 10 μm.)
IndoA inhibits Cdk5 activity with high affinity, but also targets Cdk4. Therefore, this compound potentially acts in cells as a Cdk4/5 inhibitor (4/5i) (21). Importantly, the structurally related Cdk4-specific inhibitor (4i), IndoB, was 6.4- to 15.4-fold less potent at preventing human NE cell proliferation (Fig. 2B and SI Appendix, Fig. S2 A–H). Growth of multiple NE cancer cell lines was likewise blocked by the broad spectrum Cdk inhibitors roscovitine and dinaciclib. CP681301, a selective inhibitor of both Cdk2 and Cdk5, also blocked growth of NE cancer cells, whereas the Cdk2-specific inhibitor CVT313 had a greatly reduced effect (SI Appendix, Fig. S2 I–L). Thus, Cdk5 inhibition was necessary to robustly inhibit proliferation of cells derived from multiple forms of NE neoplasms. Previous work demonstrated cell growth inhibition of MTC cells using kinase dead mutants of Cdk5 as well as siRNA targeting the Cdk5 activator p35 (12). Together, these data indicate that Cdk5 and p35/25 expression characterize at least a portion of all NE neoplasms and aberrant Cdk5 activity is a major contributor to the growth of NE cancer cells.
Downstream Targets of Cdk5.
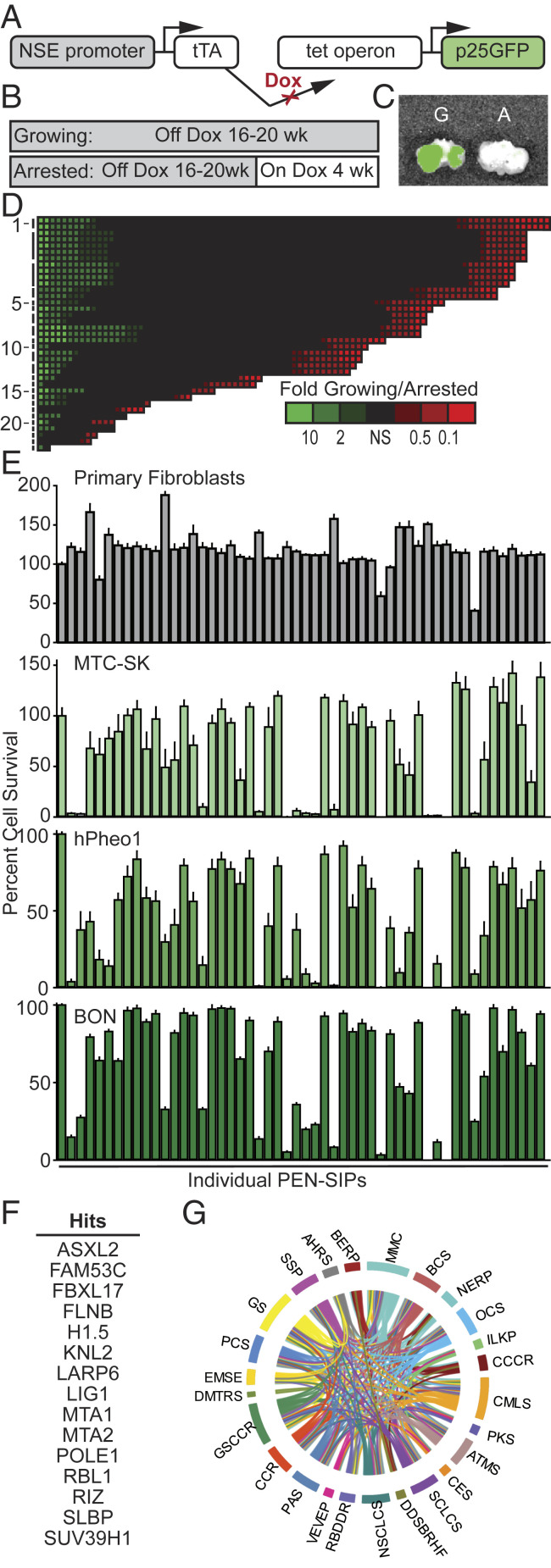
To determine which pathways Cdk5 may target to drive NE cell proliferation, we used a unique bitransgenic mouse model of MTC (NSE-p25OE mice), developed in our laboratory, in which tumors arise at the natural organ site in the presence of an intact immune system. These mice express an activator of Cdk5, p25-GFP, in C cells of the thyroid under the control of a doxycycline (dox) regulatable promotor (Fig. 3A) (12). This system can be used to generate actively growing MTC tumors through expression of p25-GFP as well as growth-arrested MTC tumors through initial expression and subsequent suppression of p25-GFP via readministration of dox (Fig. 3 A–C). As C cells only comprise 3% of a normal thyroid, this system allows generation of sufficient quantities of C cells for direct comparison of the signaling states between dividing and nondividing populations.
Fig. 3.
Identification of potential tumorigenic phosphoprotein signaling pathways. (A) Schematic of bitransgenic system for regulated tissue-specific expression of p25GFP. (B) Diagram of induction paradigm for generation of growing and arrested tumor tissue. (C) Overlay of photograph and GFP fluorescence from analysis by IVIS Spectrum (PerkinElmer) of resected trachea/esophagus with bilateral growing (G) and arrested (A) tumor tissue. (D) Phosphoproteome of mouse MTC tumors represented as a heat map of phosphopeptide levels in growing vs. arrested tumors sorted by protein group as indicated in SI Appendix, Table S1 (73) (y axis) and fold change intensity (x axis). (E) Growth/survival assay of NE cell lines treated with control (0.3% DMSO) or the PEN-SIPs (30 μM) indicated in SI Appendix, Table S2. Error bars represent SEM (n = 7 to 8). (F) Hits selected from E. (G) IPA of hits from E.
Phosphoproteomic analysis of growing vs. arrested MTC tumors was performed using PhosphoScan technology that included trypsin digest and peptide immunoprecipitation utilizing antibodies that recognize proline-directed phosphorylation of serine or threonine in the context of a MAPK or CDK consensus sequence (22). This analysis revealed global differences in the proline-directed S/T phosphosignaling network including over 250 peptides with elevated phosphorylation levels in growing tumors (Fig. 3D and SI Appendix, Table S1). From this set of phosphorylation sites, those not conserved in humans or conforming to a stringent Cdk5 phosphorylation sequence (S/T-P-x-K/H/R) were eliminated. From the remaining proteins, 50 of the most highly up-regulated phosphorylation sites, with preference for those with established or suggested roles in cancer, were selected for investigation as potential tumorigenic regulators (SI Appendix, Table S2). Short interfering peptides (SIPs) containing the phosphorylation site flanked by eight amino acids on both the N- and C-terminal sides were designed to selectively interfere with phosphorylation or function of the 50 targets. A cell-penetrating sequence (RQIKIWFQNRRMKWKK) from penetratin (PEN) was added to the N terminus of peptides to facilitate entry into cells. A high-throughput proliferation-based assay identified 15 SIPs that inhibited growth of NE cancer cells but not normal primary fibroblasts (Fig. 3 E and F and SI Appendix, Table S2), suggesting that these sites may be important for NE cancer cell growth.
We performed ingenuity pathway analysis (IPA) to ascertain major signaling cascades and pathways that are associated with these 15 proteins. Among the predicted 25 statistically enriched canonical pathways, cell cycle regulation, DNA repair, and diverse cancer signaling pathways are a predominant feature (Fig. 3F and SI Appendix, Table S3). Thus, the downstream targets identified here are associated with common cancer mechanisms.
Biomarkers of Cdk5 Activity.
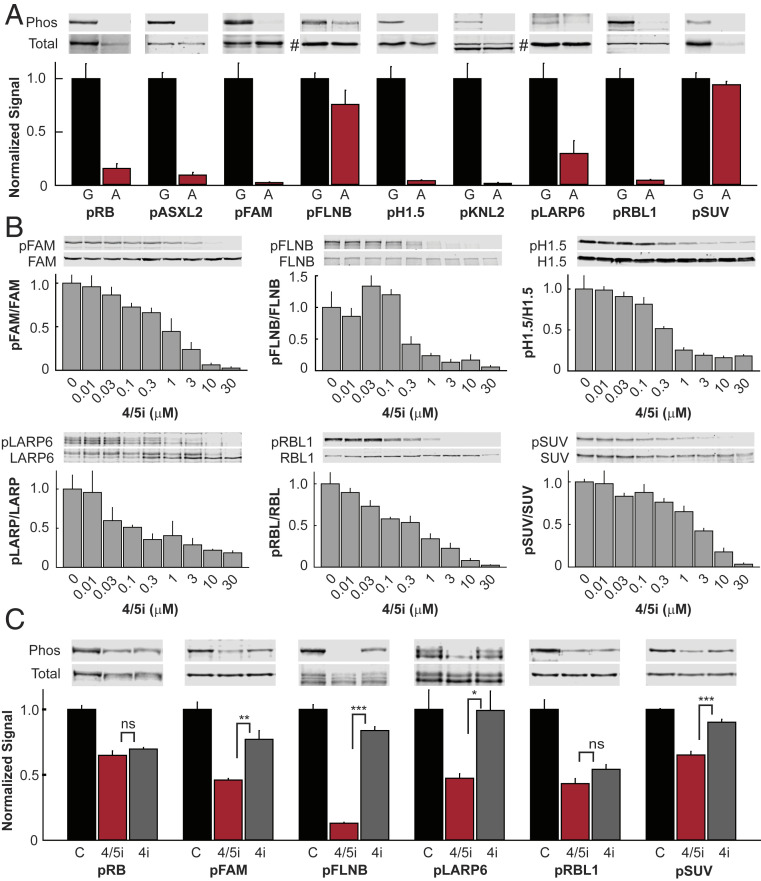
Phosphorylation state-specific antibodies were successfully generated for detection of six sites: Ser608 Additional Sex Combs Like 2 (ASXL2), Thr143 Family with Sequence Similarity 53 Member C (FAM53C), Thr709 Filamin B (FLNB), Thr202 La Ribonucleoprotein Domain Family Member 6 (LARP6), Ser110 Kinetochore-Associated Protein Homolog (KNL-2), and Ser988 Retinoblastoma-Like Protein 1 (RBL1). Phosphorylation state-specific antibodies were also generated to two sites previously identified as targets of Cdk2: Ser17 Histone 1.5 (H1.5) (23, 24) and Ser391 Suppressor of Variegation 3-9 Homolog 1 (SUV39H1) (25) (SI Appendix, Fig. S3). Phosphorylation of these eight sites, as well as the thoroughly established Ser807/Ser811 sites on retinoblastoma-associated protein (RB) (26, 27), was confirmed in mouse MTC tumors, in which growth was driven by expression of p25-GFP, and reduced in arrested tumors (Fig. 4A). In agreement, phosphorylation levels of six of these sites were dose-dependently decreased by the 4/5i, IndoA, in hPheo1 cells (Fig. 4B). Similar decreases in these phosphorylation sites were observed in human MTC-SK, TT, and BON cells (Fig. 4C and SI Appendix, Fig. S4). This effect appeared Cdk5 specific as addition of the 4i, IndoB, to multiple NE cell lines had significantly less effect on the phosphorylation states of Thr143 FAM53C, Thr709 FLNB, Ser17 H1.5, and Thr202 LARP6 compared to IndoA. In contrast, phosphorylation of Ser807/Ser811 RB, a known target of multiple Cdks, was decreased upon treatment with both inhibitors. Interestingly, the phosphorylation of the RB family member, RBL1, was more responsive to IndoA than IndoB in TT cells, whereas phosphorylation of Ser391 SUV39H1 was more responsive to IndoA than IndoB in BON cells (Fig. 4C and SI Appendix, Fig. S4). Overall, these data demonstrate that phosphorylation of these six proteins is dependent upon Cdk5 activity and suggests that these phosphorylation sites could serve as biomarkers for many types of Cdk5-driven NE tumors. Attempts to produce antibodies for the remaining seven sites identified by SIP inhibition of cell growth were unsuccessful and therefore the capacity of these sites to serve as biomarkers could not be evaluated.
Fig. 4.
Phosphoproteins are biomarkers of Cdk5 activity. (A) Immunoblot analysis of phosphoproteins in growing (G) and arrested (A) mouse MTC tumors. For pFLNB and pLARP6, total blot is actin (#); all others are normalized to each specific protein (n = 3 to 4). (B) Immunoblot analysis of phosphoproteins in human hPheo1 cells treated with increasing concentrations of IndoA (4/5i) for 4 h (n = 3 to 4 for all except 0.01, 1, and 30 μM points of pLARP6 where n = 2). (C) Immunoblot analysis of phosphoproteins in BON cells treated with 0.3% DMSO (control), 2 μM IndoA (4/5i), or 2 μM IndoB (4i) for 4 h. All error bars represent SEM. Simple comparisons between two groups of normally distributed data were performed using two-tailed Student’s t test. *P < 0.05, **P < 0.01, ***P < 0.001; ns, not significant.
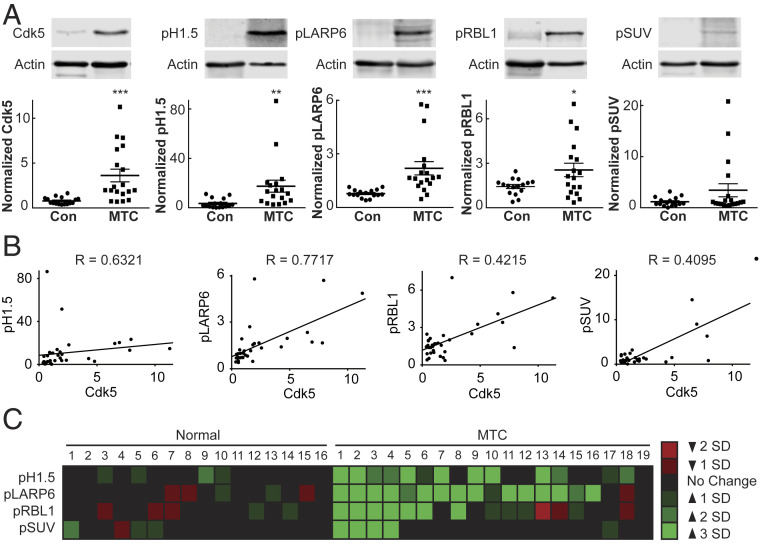
To determine if these Cdk5-dependent biomarkers occurred in human NE tumors, cohorts of MTC patient tumors and normal human thyroid tissues were compared. Immunoblot analysis revealed that three of the phosphorylation sites, Ser17 H1.5, Thr202 LARP6, and Ser988 RBL1, were significantly elevated in the total tumor population and exhibited a positive correlation with overall Cdk5 expression levels (Fig. 5 A and B). The phosphorylation state of Ser391 SUV39H1 was only increased in a small portion of tumor samples but retained a positive correlation with Cdk5 expression. Distribution of all four phosphosites varied across patients with phospho-LARP6 being the most commonly detected followed by phosphorylation of H1.5 and RBL1, respectively (Fig. 5C). Although 73% of patient tumors exhibited elevated Cdk5 levels, only 21% presented with elevation of all four biomarkers of Cdk5 pathway activity, emphasizing the fact that presence of a protein does not correlate 100% with function of that protein and highlighting the need for direct readouts of pathway activity such as phosphorylation of downstream substrates. Thus, Cdk5-dependent tumorigenic signaling may be considered patient specific. Furthermore, detection of all four biomarkers in a patient could predict positive response to a Cdk5-targeted therapeutic approach.
Fig. 5.
Phosphoprotein biomarkers are present in subgroups of human MTC patients. (A) Immunoblot analysis of Cdk5 and phosphoproteins in normal human thyroid tissue and human MTC tumors. Due to sample size, samples were processed on three separate gels, each containing a reference for normalization. Error bars represent SEM. (B) Phosphoprotein level vs. Cdk5 expression in samples represented in A, analyzed by Spearman rank-order analysis; rho (R). (C) Heat map representation of immunoblot analysis of phosphoproteins from A, relative to the average and SD of the total normal thyroid population, grouped by individual patient sample. Simple comparisons between two groups of normally distributed data were performed using two-tailed Student’s t test. *P < 0.05, **P < 0.01, ***P < 0.001.
Cdk5 Inhibitors as Effective Therapeutics in NE Models.
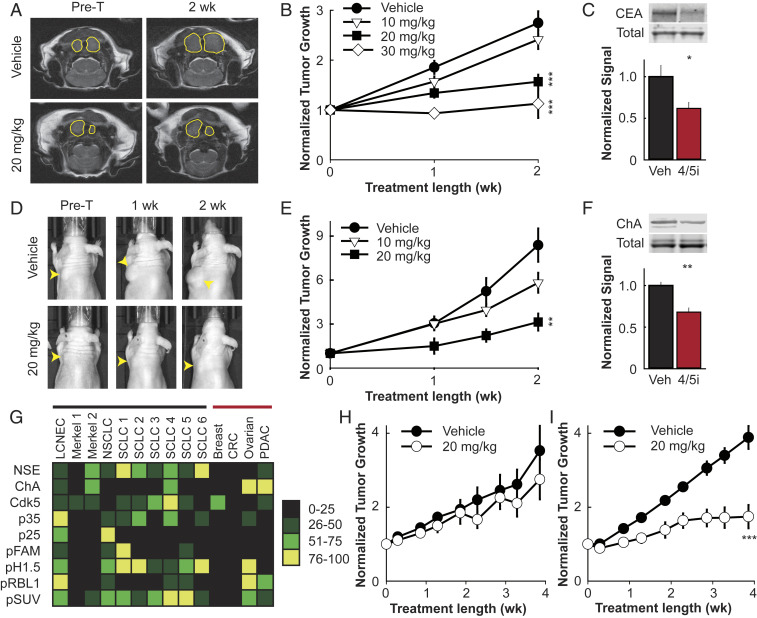
To evaluate the efficacy of anti-Cdk5 therapy when administered systemically, IndoA was utilized in multiple animal models of NE cancer. IndoA was chosen for these studies as this compound is more selective for Cdk5 than dinaciclib and more potent than roscovitine and CP681301 (SI Appendix, Fig. S2) (12, 28, 29). First, the transgenic model of MTC driven directly by activation of Cdk5, NSE-p25OE, was treated with vehicle or IndoA (10 to 30 mg/kg body weight [BW]) once every 3 d for 2 wk. Tumor growth was significantly reduced by 20 and 30 mg/kg of IndoA compared to control animals (Fig. 6 A and B). Plasma levels of Carcinoembryonic Antigen-Related Cell Adhesion Molecule 1 (CEACAM1), a marker of MTC (30), were also reduced in treated animals, indicating therapeutic efficacy (Fig. 6C). As a second model, TT cells derived from a familial case of human MTC were used to generate xenograft tumors in nude mice. As with the transgenic model, IndoA attenuated TT cell tumor progression (Fig. 6 D and E). While CEACAM1 was not detected in the plasma of these animals, levels of chromogranin A (ChA), another marker of MTC (31), were reduced (Fig. 6F). Thus, Cdk5 inhibition blocks the growth of tumors in distinct models of MTC. The efficacy of IndoA was also assessed in a xenograft tumor model generated from human pancreatic NE tumor-derived BON cells (SI Appendix, Fig. S5). IndoA was equally effective in impeding the growth of these tumors, suggesting that targeted Cdk5 therapy could be effective across multiple NE cancer types.
Fig. 6.
Cdk5-targeted therapy is effective in biomarker (+) tumors. (A) Representative MRIs of NSE-p25OE MTC model mice prior to treatment (Pre-T) or treated for 2 wk with vehicle or 20 mg/kg BW IndoA. Tumors are outlined in yellow. (B) Quantitation of tumor growth over time from MRI of NSE-p25OE MTC mice treated with vehicle (n = 6) or 10 mg/kg (n = 4), 20 mg/kg (n = 6), or 30 mg/kg (n = 3 to 4) BW IndoA. (C) Immunoblot analysis of blood plasma for CEACAM1 (CEA) in vehicle and 20 or 30 mg/kg BW IndoA (4/5i) animals from B. (D) Representative images of TT cell xenograft MTC model mice treated for 2 wk with vehicle or 20 mg/kg BW IndoA. Tumors are marked by yellow arrowheads. (E) Quantitation of tumor growth over time from caliper measurements of TT cell xenograft MTC model mice treated with vehicle or 10 mg/kg or 20 mg/kg BW IndoA (n = 5 to 7). (F) Immunoblot analysis of blood plasma in vehicle and 20 mg/kg BW IndoA (4/5i) animals from E. (G) Heat map of protein and phosphoprotein population percentile ranking from immunoblot analysis of NE tumors (black bar) and non-NE tumors (red bar) from PDX model mice. (H and I) Quantitation of tumor growth over time from biomarker-negative Merkel 1 (H; n = 8) and biomarker-positive NSCLC (I; n = 8) PDX model mice treated with vehicle or 20 mg/kg BW IndoA (20 mg/kg). All error bars represent SEM. Simple comparisons between two groups of normally distributed data were performed using two-tailed Student’s t test. *P < 0.05, **P < 0.01, ***P < 0.001.
Biomarkers Are Predictive of Response to Anti-Cdk5 Therapy.
We hypothesized that the phosphorylation states of the newly identified set of proteins could serve as biomarkers of Cdk5-driven tumors. If true, biomarker-positive tumors should be responsive to Cdk5 inhibitor therapy while biomarker-negative tumors should be nonresponsive. To test this, a cohort of tumors from patient-derived xenograft (PDX) models of mixed origins was analyzed quantitatively for the presence of the common NE markers ChA and neuronal-specific enolase (NSE) (32), Cdk5 pathway components, and putative Cdk5-dependent biomarkers (SI Appendix, Fig. S6A). Index scores were assigned based on the population quartiles (0 for quartile 1; 1 for quartile 2; 2 for quartile 3; and 3 for quartile 4), summed together, and used to generate a representative heat map (Fig. 6G and SI Appendix, Table S4). Two models were identified with high biomarker index scores; a large cell NE carcinoma (LCNEC; score 8) model and a nonsmall cell lung cancer (NSCLC; score 6) model. Two Merkel cell models (Merkel 1 and 2) were also identified that exhibited very low biomarker index scores, 0 and 1, respectively. In collaboration with Champions Oncology, cohorts of these four disparate PDX models were treated with IndoA (20 mg/kg), and tumor progression was assessed (Fig. 6 H and I and SI Appendix, Fig. S6 B and C). Initial implantation and growth characteristics of the NSCLC model and the Merkel 1 model were highly comparable and allowed for two full cohorts of eight animals per group to be monitored for 4 wk. Tumor growth of the biomarker-positive NSCLC model was notably attenuated by Cdk5 inhibitor treatment, whereas the biomarker-negative Merkel 1 model had no response (Fig. 6 H and I). Implantation and growth characteristics of the LCNEC and Merkel 2 models were highly variable and allowed only smaller cohorts of 5 to 8 tumor-bearing animals and testing for a 2-wk period. Nonetheless, growth of the biomarker-positive LCNEC model was significantly reduced, whereas growth of the biomarker-negative Merkel 2 model was not (SI Appendix, Fig. S6 B and C). Together, these data strongly support the ability of these biomarkers to predict responsiveness to anti-Cdk5 therapy.
Enhanced Drug Delivery Method.
Although transgenic MTC mouse model animals responded favorably to IndoA treatments, tumor regression was not observed. Testing higher doses of IndoA for increased potency was not feasible as cohorts receiving 30 mg/kg BW experienced 50% mortality while both 20 and 30 mg/kg-treated cohorts evidenced some level of liver or kidney toxicity as measured by aspartate aminotransferase (AST) activity (33, 34) and urea levels (35, 36), respectively. (Fig. 7 E and F). To deliver IndoA selectively to tumors and thereby circumvent toxicity issues, we tested a biomimetic nanoparticle-based drug delivery system, leukosomes (LKs), generated from a combination of synthetic phospholipids and leukocyte membrane extracts (37, 38). Traditional nanoparticle delivery systems are dependent upon passive targeting through enhanced permeability and retention (EPR) of unhealthy tumor vasculature (39). Although still benefitting from EPR, LKs actively target activated endothelium via chemotactic mechanisms analogous to those utilized by white blood cells (40). Importantly, LKs are successfully camouflaged by leukocyte membrane proteins, leading to lower total opsonization and avoidance of the rapid immune clearance observed with purely synthetic nanoparticle platforms (41–43).
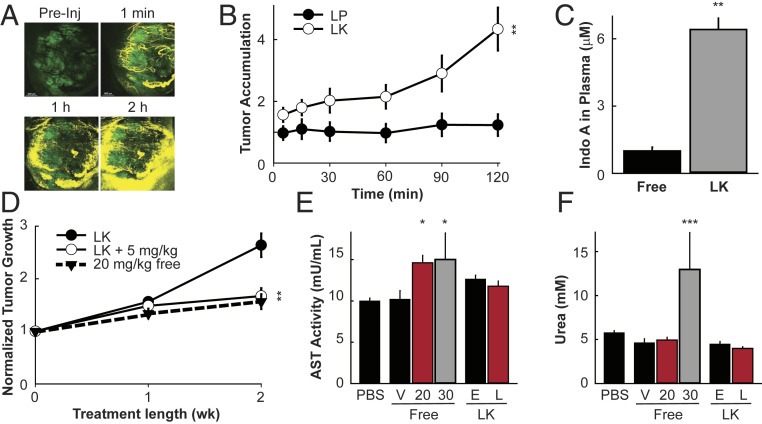
Fig. 7.
Targeted delivery of Cdk5 inhibitors via leukosomes. (A) Representative images of MTC tumors from NSE-p25OE mice injected with Cy5.5-labeled LKs and imaged in vivo over 2 h using IVM. Green, tumor cells; yellow, LKs. (B) Quantitation of accumulation of Cy5.5-labeled LPs and LKs in MTC tumors of NSE-p25OE mice over time normalized by total tumor size (n = 7 to 8). (C) HPLC-MS analysis of IndoA in blood plasma from NSE-p25OE MTC mice treated with 5 mg/kg BW IndoA (free) or 5 mg/kg BW IndoA encapsulated in LKs (LK) (n = 3). (D) Quantitation of tumor growth over time from MRI of NSE-p25 MTC mice treated with empty LKs or 5 mg/kg BW IndoA encapsulated in LKs (LK + 5 mg/kg) compared to 20 mg/kg BW free IndoA from Fig. 6B (n = 5 to 6). (E and F) Analysis of blood plasma for AST activity (E) and urea level (F) in animals treated with PBS, unencapsulated IndoA (free) (V, vehicle alone, 20 mg/kg BW, or 30 mg/kg BW), or IndoA encapsulated in LKs (E, empty LK, L, drug loaded LK). All error bars represent SEM. Simple comparisons between two groups of normally distributed data were performed using two-tailed Student’s t test. *P < 0.05, **P < 0.01, ***P < 0.001.
As tumors are generally highly inflamed and attract multiple types of immune cells (44, 45), we tested the ability of LKs to traffic to MTC tumors of NSE-p25OE mice and to stabilize lower doses of IndoA. Using intravital microscopy, LKs were verified to exhibit increased tumor localization compared to control liposomes (LPs) and to time-dependently spread from the vasculature into surrounding tumor tissue (Fig. 7 A and B). Maximum encapsulation of IndoA into LKs had no deleterious effects on particle size or homogeneity (SI Appendix, Fig. S7) and allowed a dosage of 5 mg/kg BW IndoA per retroorbital injection. High performance liquid chromatography/mass spectrometry (HPLC/MS) analysis of blood plasma demonstrated a sixfold stabilization of encapsulated drug compared to free drug at these lower dosage levels (Fig. 7C).
IndoA delivered at 10 mg/kg BW in the free form had no effect on tumor growth in transgenic MTC animals (Fig. 6B). In contrast, delivering only 5 mg/kg BW IndoA encapsulated in LKs had the same effect as delivering 20 mg/kg BW free drug (Figs. 6B and 7D). Thus, the LK delivery system generated an equivalent effect utilizing a 75% lower dosage. LKs also protected animals against hepatic and renal toxicity (Fig. 7 E and F). While complete tumor arrest or shrinkage was not observed at 5 mg/kg BW, further modifications to allow higher encapsulation of drug could provide additional benefit for this unique delivery approach.
Discussion
The development of advanced sequencing techniques has led to an explosion of information pertaining to the genomic landscape of cancer. In some cases, this information successfully progressed to the development of personalized medicine. For example, mutations at Val600 in the serine/threonine kinase B-Raf are predictive of response to B-Raf inhibitors in patients with metastatic melanoma (46, 47). However, the majority of B-Raf mutant papillary thyroid cancer and colon cancer patients do not respond to B-Raf-targeted inhibitors (48), emphasizing the difficulty designing treatment options based on single gene mutations.
For NE neoplasms, mutations are prominent in genes encoding the scaffolding protein menin and the receptor tyrosine kinase RET. Menin mutations were initially identified over 20 y ago in patients with multiple endocrine neoplasia (MEN) type 1 syndrome (49, 50). Menin is a broadly expressed tumor suppressor in which mutations typically cause protein truncation (51, 52). Currently there are no therapeutics with the potential to circumvent these mutations.
Mutations in the protooncogene RET were discovered over 20 y ago in patients with MEN2 syndrome (53–55). The development of vandetanib and carbozantinib, tyrosine kinase inhibitors that target RET, generated hope that patients possessing RET mutations could be successfully treated. Unfortunately, human trials revealed no correlation between the presence of a RET mutation and patient response to therapy (56–58).
As with all cancers, many mutations, in addition to RET, are present within each MTC cancer cell. These additional mutations can alter the diverse input nodes of the signaling network that drive cancer cell growth and survival. For these reasons, looking at the signaling network with a broader lens that includes posttranslational modifications could be beneficial and aid in the elimination of “false positive” nonresponders, that would be predicted responders, from genomic or proteomic information alone. The current study reveals that Cdk5 is likely a contributor to at least a portion of all NE tumor types. This study also identifies a set of phosphorylation-based biomarkers which indicate that not only are Cdk5 pathway components present, but Cdk5 is actively modulating the signaling network and regulating cancer physiology. It is important to note that additional experiments will be necessary to determine if each of these proteins are direct targets of Cdk5. Regardless, the dependence of these phosphorylation sites on Cdk5 activity validates them as biomarkers of Cdk5 pathway activation.
In addition to being biomarkers of Cdk5 pathway activity, the phosphoproteins identified here are potentially directly involved in promotion of cell growth and/or survival. For example, RBL1 is a member of the retinoblastoma (RB) family of proteins that includes the tumor suppressor RB and RBL2. The RB family plays a major role in cell cycle regulation and is also involved in modulating senescence, apoptosis, and chromosomal stability (27). Although functional compensation has been observed among the family members, some differences exist. Unlike RB, RBL1 and RBL2 do not bind to activating transcription factors E2F1-3. They interact instead with transcriptional repressors E2F4 and E2F5 (59–61). Remarkably, both RBL1 and RBL2, but not RB, are members of the DREAM complex, a regulatory unit that mediates cell entry into quiescence (62, 63).
Emerging information suggests Cdk5 may play a role in non-NE cancers as well (11). High Cdk5 expression in neuroblastoma, a rare but deadly childhood cancer (64–66), is correlated with worse overall survival, and inhibition of Cdk5 amplifies the potency of Cdk2 inhibitors in neuroblastoma cell lines (67). The Cdk5 gene is amplified in pancreatic ductal adenocarcinoma (PDAC) and blockage of Cdk5 activity inhibits migration and invasion in PDAC cell lines. Involvement of Cdk5 is also implicated in colorectal cancer (68), breast cancer (69), glioblastoma (70), and prostate cancer (71). These findings suggest that Cdk5 is a vulnerable node that can be targeted in most tissues to facilitate cancer development and/or progression and that Cdk5 likely plays a role in subgroups of many cancer types. As the heterogeneity between tumors of the same cancer type is becoming more apparent, the likelihood that treatment approaches will evolve based on individual tumor signaling states instead of general tumor type classification is increasing. The biomarkers identified in this study are not limited to use for NE cancer patients. Indeed, the ovarian and PDAC PDX models analyzed here (Fig. 6G), both non-NE by diagnosis, exhibit high levels of NE features as well as biomarkers of Cdk5 pathway activation.
To be useful clinically, antibodies to detect these biomarkers will need to be developed into an easy-to-use assay that allows reliable quantitation of biomarker levels in patient tumors, preferably in relatively small samples such as core biopsies. Current progress in multiplexed assay systems make these platforms especially attractive for clinical analysis of groups of biomarkers from relatively small biological sources. For example, bead-based multianalyte profiling has been employed for the evaluation of 22 cytokines in a single serum sample from cancer patients (72). This technology has the capacity to distinguish up to 100 different analytes per sample and is compatible with several forms of biological samples, including solid tissue lysates. Once a clinically applicable assay platform is generated, additional preclinical surveys for these biomarkers in multiple PDX models, coupled with Cdk5 inhibitor testing, could aid delineation of finite cutoffs for classification of multiple forms of cancer as predicted responders for Cdk5-targeted therapy and validate progression of such an assay, coupled with novel Cdk5 inhibitors, into clinical trials. At this stage, establishment of standard operating procedures for collection, handling, storage, and processing of human tumor samples will be critical for ensuring protein- and phosphoprotein-based biomarker integrity and will facilitate ease of implementation in the clinic. Using a multianalyte platform, analysis of biomarkers for additional oncogenic pathways could be combined with analysis for biomarkers of Cdk5-driven tumors to yield a network readout of the broader signaling state of the tissue. Assessing signaling states across a network of pathways might allow more accurate, stage-dependent, therapeutic decisions to be made for individual patients.
Materials and Methods
Animal Research.
All animal work was performed in accordance with the guidelines of the Animal Welfare Act and the Guide for the Care and Use of Laboratory Animals under approved protocols by University of Texas Southwestern (UTSW), University of Alabama at Birmingham (UAB), and Houston Methodist Research Institute (HMRI) Institutional Animal Care and Use Committees.
Antibody Production and Purification.
Phosphopeptides (SIT*SPNRTGC-ASXL2, CAPSKLW*TPIKH-FAM53C, CSY*TPVKAIK-FLNB, CAPVEK*SPAK-H1.5, CANYE*SPGKI-KNL2, CALA*TPQKNG-LARP6, CSIYI*SPHKN-RBL1, CAGLPG*SPKK-SUV39H1; *indicates a phosphorylated residue) were conjugated to Limulus polyphemus hemocyanin (Sigma H1757), emulsified with Freund’s adjuvant (Sigma F5881 or F5506), and injected s.c. into New Zealand white rabbits (Charles River Laboratories). Rabbits were boosted once and blood was collected twice over a 5-wk period for 12 mo. Blood was allowed to clot at 4 °C for 24 h, centrifuged at 1,000 × g, and plasma was isolated and stored at −20 °C. Phosphorylation state-specific antibodies were purified from plasma using phosphopeptide affinity columns by elution with 100 mM glycine pH 2.5 into 1 M Tris pH 8.6 (11:1 volume ratio, final 80 mM Tris pH ∼ 7.5). Antibodies were dialyzed into 50 mM Tris pH 7.6 plus 150 mM NaCl and stored at −20 °C.
Cell Culture and Assays.
All cells were tested and verified to be free of mycoplasma contamination. Cell lines were probed for NE markers to verify identity. Cells were cultured at 37 °C and 5% CO2 in a humidified incubator. Fibroblasts were cultured in Dulbecco's modified Eagle medium (DMEM) with 10% fetal bovine serum (FBS). TT, MTC-SK, and SIN-J cells were cultured in Ham’s F-12:medium 199 (1:1) with 10% FBS. hPheo1, H146, and H1184 cells were cultured in RPMI with 10% FBS, 1 mM Na-pyruvate, and 10 mM Hepes. BON cells were cultured in DMEM:Ham’s F-12 (1:1) with 10% FBS. INS cells were cultured in RPMI with 10% FBS, 1 mM Na-pyruvate, 10 mM Hepes, 4.5 g/L glucose, and 50 μM β-ME. TT-RLuc cells were cultured in RPMI with 20% FBS, 100 μg/mL penicillin, and 100 μg/mL streptomycin.
Growth assays.
Cells were seeded onto a black 96-well plate with a clear optical bottom and allowed to recover overnight. Growth of hPheo1 cells was measured 2 d after inhibitor treatment; and growth of Fibro, TT, MTC-SK, BON, INS, and H146 cells was measured 6 d after inhibitor treatment using Cyquant direct proliferation assay (Invitrogen) and an Optima Fluostar plate reader (BMG LabTech). For SIP experiments, cells received two SIP treatments per experiment. Growth of BON and hPheo1 cells was measured 2 d after initial SIP treatment and growth of Fibro and MTC-SK cells was measured 6 d after initial SIP treatment using procedures described above.
Immunoblot analysis.
Cells were seeded onto 6-well dishes and allowed to recover overnight. Cells were treated for 4 h with inhibitors and then lysed in 50 mM NaF and 1% sodium dodecyl sulfate (SDS) with brief sonication. Samples were diluted to equivalent total protein concentrations in 1× Laemmli buffer and separated by SDS/polyacrylamide gel electrophoresis (PAGE). Proteins were transferred onto nitrocellulose for immunoblotting utilizing in-house phosphorylation state-specific antibodies, anti-Cdk5 (sc-173), anti-p35 (sc-820), anti-GAPDH (Sigma G8795), anti-actin (Abcam ab6276), anti-ASXL2 (Abcam ab106540, Sigma sab1407639), anti-Fam53C (Abcam 105679), anti-FLNB (Abnova PAB30702), anti-H1.5 (sc-247158), anti-KNL2 (sc-162587), anti-RBL (sc-318-G), anti-LARP6 (Sigma sab1407657), and anti-SUV39H1 (Sigma S8316). Anti-goat, -rabbit, and -mouse secondary antibodies conjugated to either IRdye 680RD or IRdye 800CW (LiCor) were used for detection on a LiCor Odessey CLx. Actin and GAPDH were used as sample processing controls. Other total proteins were probed on the same membrane as phosphoproteins unless antibody conditions did not allow.
Human Tissue Analysis.
Collection.
Human tissues were collected with patient consent and in accordance with institutional review board (IRB) regulations. Samples were collected under University of Wisconsin–Madison IRB 2011-0145, MD Anderson IRB PA11-0744, University of Texas Southwestern Medical Center IRB STU102010-042 and STU102010-051, National Institute of Child Health and Human Development IRB 00-CH-0093, Louisiana State University IRB 5774, and University of Sydney LNR/13/HAWKE/424-1312-417M. All samples were deidentified prior to use.
Histology.
Formalin-fixed, paraffin-embedded samples were cut into 5-μm sections, deparaffinized, and subjected to microwave antigen retrieval (citrate buffer, pH 6.0). Sections were then stained using standard protocols for H&E or immunostained with antibodies recognizing p35/p25 (sc-820, Santa Cruz Biotechnology) or Cdk5 (308-Cdk5, PhosphoSolutions). For immunostaining, sections were permeabilized with 0.3% Triton X-100, quenched free of endogenous peroxidases, and blocked with 2.5% normal goat serum prior to overnight incubation with primary antibodies at 4 °C. Bound primaries were detected by sequential incubation with biotinylated-secondary antisera, streptavidin-peroxidase (Vector Laboratories), and diaminobenzidine chromagen (DAKO) following the manufacturer’s directions.
Immunoblot analysis.
Tissues were crushed while frozen then lysed in 50 mM NaF and 1% SDS with brief sonication. Samples were diluted to equivalent total protein concentrations in 1× Laemmli buffer and separated by SDS/PAGE. Proteins were transferred onto nitrocellulose for immunoblotting as described above.
Intravital Microscopy.
Intravital microscopy (IVM) was performed using an upright Nikon A1R laser scanning confocal microscope with a resonance scanner, motorized and heated stage, and Nikon long-working distance 4× and 20× dry plan-apochromat objectives housed within the IVM Core at the HMRI. For imaging, NSE-p25OE mice were anesthetized with isoflurane and the ventral surface of the neck was opened to expose the trachea, salivary glands, and MTC tumors. Tumors were positioned in direct contact with the coverslip, visualized using the GFP signal, and positions were selected for imaging. After selection of positions, Cy5.5-labeled LKs and LPs were administered via retroorbital injection and mice were imaged continuously for 2 h using the 4× objective. Images were quantified using Nikon Elements. The tumor accumulation reported was normalized by dividing the area occupied by LKs or LPs by the area occupied by the tumor within each image.
LCMS2 Analysis of IndoA.
Extraction.
Sera (25 µL) were reconstituted in 25 µL of extraction solution (25% acetonitrile [ACN]/75% H20), vortexed for 1 min, diluted with 25 µL of 100% ACN, followed by two cycles of vortexing for 1 min and incubating at room temperature (RT) for 10 min. Samples were then diluted with 200 µL of 100% ACN, vortexed for 1 min, then stored at −20 °C overnight. Samples were thawed at RT for ∼10 min, centrifuged at 14,000 × g for 10 min at 4 °C to remove cell debris, then supernatants were transferred to 2-mL tinted glass vials and dried down to ∼5 to 10 µL under argon gas at 25 °C for 25 min. Samples were immediately reconstituted (adjusted to 210 µL) in 25% ACN/1% trifluoroacetic acid (TFA)/74% H20 and centrifuged at 14,000 × g for 10 min at 4 °C to remove any residual debris. The supernatants were then transferred to tinted vials prior to analysis.
LC/MS.
Analysis was performed on a Dionex Ultimate 3000 UHPLC+ Focused Stack & Auto Sampler (Thermo Fisher Scientific/Dionex) a using a RP C18 Hypersil Gold (100 mm inner diameter [I.D.] × 4.6 mm, 5 mm 175-Ǻ pore size; Thermo Fisher Scientific) in-line with an LTQXL mass spectrometer equipped with a heated electrospray ionization source (Thermo Fisher Scientific), and all data were collected in selective reaction monitoring (SRM) mode. The HPLC was configured with binary mobile phases that include solvent A (0.1% TFA/99.9% H2O), and solvent B (0.1% TFA/15% H2O/5% ACN). The gradient program steps were run in linear mode as follows; 0 to 6 min at 75 to 50% B (200 µL/min), 6 to 7 min at 50 to 80% B (200 µL/min), 7 to 11 min at 80 to 90% B (200 µL/min), 11 to 12 min at 90 to 25% B (500 µL/min), and finally 12 to 16 min at 25% B (200 µL/min). SRM mode was optimized using a parent ion window of 453.2 ± 1.0 m/z, 20% normalized collision energy, activation energy at 0.240, activation time of 30 ms, with a daughter ion window of 306.5 ± 1.5 m/z. The resultant Xcalibur RAW files were collected in profile mode and the SRM base peak values processed and extracted using Xcalibur version 2.2 sp1.48.
Leukosome Synthesis and Characterization.
LKs were developed as previously reported (38). Briefly, 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) and 1,2-dioleoylsn-glycero-3-phosphocholine (DOPC) and cholesterol (Avanti Polar Lipids) (4:3:3 molar ratio) were dissolved in ethanol at a final lipid concentration of 9 mM and mixed with membrane proteins, previously resuspended in aqueous buffer at 1:50 protein to-lipid concentrations, using the NanoAssemblr Benchtop platform (Precision NanoSystems, Inc.). Passive loading of IndoA within LKs was obtained by dissolving the drug in the ethanol mixture containing the lipids. Size and polydispersity index were determined through dynamic light scattering analysis using a Nanosizer ZS (Malvern Instruments). Surface charge (zeta potential) was measured using a ZetaSizer Nano ZS (Malvern Instruments).
Magnetic Resonance Imaging.
MRI conducted at University of Texas Southwestern Medical Center was performed using a 7-Tesla small animal MRI system (Agilent Inc.) with a 40 mm (I.D.) radio frequency (RF) coil and a 400 mT/m gradient coil set. Animals were anesthetized by isoflurane and imaged in a supine position, head first with the thyroid centered with respect to the center of a RF coil. Two-dimensional (2D) fast spin-echo (FSE) images on three orthogonal planes (transverse, coronal, and sagittal) were first acquired to ensure the position and the orientation of the thyroid tumors. For volume measurements of the thyroid tumors, the high-resolution T2-weighted FSE axial images was acquired. Major imaging parameters were: repetition time/echo time = 2,500/40 ms, field-of-view = 25.6 × 25.6 mm, matrix size = 256 × 256, slice thickness = 1 mm, no gap, eight averages, affording 100-µm in-plane resolution.
Phosphoproteomics.
Peptide preparation.
Mouse tumors were homogenized in 8 M urea lysis buffer (20 mM Hepes pH 8.0, 9 M urea, 1 mM sodium vanadate, 2.5 mM sodium pyrophosphate, 1 mM β-glycerol-phosphate), sonicated, then centrifuged for 15 min at 4 °C at 20,000 × g. Supernatants were reduced with 4.5 mM dithiothreitol for 30 min at 55 °C followed by alkylation with 10 mM iodoacetamide. The samples were then digested with trypsin overnight at room temperature. Digests were acidified with 1% TFA and peptides desalted and purified over Sep-Pak C18 columns (Waters, WAT051910) using 40% acetonitrile in 0.1% TFA for elution. Elutes were lyophilized and stored at −80 °C.
Immunoaffinity purification of peptides.
Lyophilized peptides were dissolved in immunoaffinity purification (IAP) buffer (50 mM Mops pH 7.2, 10 mM sodium phosphate, 50 mM NaCl), sonicated, and insoluble matter was removed by centrifugation. CDK substrate motif antibody (Cell Signaling Technology, 2324) and MAPK substrate motif antibody (Cell Signaling Technology, 2325) was coupled to protein A beads (Roche). Immobilized antibody was incubated with peptide mixtures, immunoprecipitation was carried out at 4 °C overnight, then washed with IAP buffer, and eluted with 0.15% TFA. Eluates were further purified using C18 microtips (StageTips or Eppendorf C18 PerfectPure tips) with elution in 60% MeCN, 0.1% TFA, and then lyophilized.
Analysis by LC-MS/MS.
Purified peptide mixtures were loaded onto a 10 cm × 75 µm PicoFrit capillary column (New Objective) packed with Magic C18 AQ reversed-phase resin (Michrom Bioresources) using a Famos autosampler with an inert sample injection valve (Dionex). The column was developed with a 45-min gradient of acetonitrile in 0.125% formic acid (Ultimate pump, Dionex), and tandem mass spectra were collected in a data-dependent manner with a Thermo Fisher Scientific LTQ ion trap mass spectrometer equipped with electron transfer dissociation (ETD) module or with an Orbitrap mass spectrometer.
Assigning peptide sequences using Sequest.
MS/MS spectra were evaluated using TurboSequest in the Sequest Browser package supplied as part of BioWorks 3.3 (Thermo Fisher Scientific). Searches were performed against the National Center for Biotechnology Information (NCBI) human protein database. Cysteine carboxamidomethylation was specified as a static modification and phosphorylation was allowed as a variable modification on serine and threonine.
Preclinical Drug Testing.
A bitransgenic mouse model of MTC was generated as previously described by crossing NSE-rTA and TetOp-p25GFP parental lines (12). Bitransgenic litters were monitored by MRI. Beginning at 10 to 40 mm3 bilateral tumor volume, animals were treated once every 3 d by immunoprecipitation (IP) with vehicle (0.7% dimethyl sulfoxide [DMSO], 3.4% EtOH, 7.4% PEG400, 3.4% propylene glycol, 3.4% Kolliphor EL, 1.1% Tween 80 in 1× phosphate buffered saline), 10 mg/kg, 20 mg/kg, or 30 mg/kg BW IndoA. Animals were monitored by MRI for 2 wk then killed 24 h postfinal injection by CO2 euthanasia and cardiac perfusion with 0.1 mM ammonium molybdate, 5 mM ethylene glycol-bis(β-aminoethyl ether)-N,N,N′,N′-tetraacetic acid, 50 mM NaF, 2 mM Na orthovanidate, 10 mM Na pyrophosphate, and protease inhibitors (Sigma S8820) in PBS. Tissues were frozen or fixed in 4% paraformaldehyde for 24 h and submitted for paraffin embedding. Human BON and TT cell xenograft mouse models were generated by implanting 5e6 BON or TT-RLuc cells s.c. in the NU/NU nude mouse (Crl:NU-Foxn1nu) strain from Jackson Laboratories. Beginning at tumor volumes of 100 to 450 mm3 (average starting sizes between groups varied less than 40 mm3), animals were treated as described above and monitored by caliper measurement for 2 wk. Animals were killed and tissue was processed as described above. All animals were randomly assigned to treatment groups but blinding was not possible.
Scanning Electron Microscopy.
Cells were fixed on coverslips with 2.5% (vol/vol) glutaraldehyde in 0.1 M sodium cacodylate buffer overnight at 4 °C. After three rinses in 0.1 M sodium cacodylate buffer, they were postfixed with 1% osmium tetroxide in 0.1 M sodium cacodylate buffer for 45 min. Cells were rinsed with water and dehydrated with increasing concentration of ethanol, followed by increasing concentrations of hexamethyldisilazane in ethanol. Cells were air dried under the hood, mounted on SEM stubs and sputter coated with gold palladium in a Cressington 108 auto sputter coater. Images were acquired on a field-emission scanning electron microscope (Zeiss Sigma) at 3.00-kV accelerating voltage.
Statistical Analysis.
Shapiro–Wilk tests were performed to determine normality of data. Preclinical drug testing in animal models and accumulation of nanoparticles were analyzed by two-way ANOVA with repeated measures. The Holm–Sidak method was used for all ANOVA tests and all indications of significance are for the treatment group over the entire time period, not individual time points. Immunoblots containing more than two conditions, AST assays, and urea assays were analyzed by one-way ANOVA. Immunoblots of human tumors generated data that were not distributed normally. For this reason, comparisons between distributions of groups were performed using the Kolmogorov–Smirnov test and correlations between data were performed by Spearman rank-order analysis. Simple comparisons between two groups of normally distributed data were performed using two-tailed Student’s t test. Sample sizes are provided within figure legends or in results (*P < 0.05, **P < 0.01, ***P < 0.001).
Toxicity Assays.
Prior to cardiac perfusion, blood was collected from animal subjects via retroorbital bleeding. Blood was immediately mixed with (ethylenedinitrilo)tetraacetic acid (10 mM final) then centrifuged at 1,000 × g for 10 min at 4 °C to allow isolation of plasma. AST assays (Sigma MAK055) and urea assays (Sigma MAK006) were performed on plasma per the manufacturer’s instructions.
Supplementary Material
Acknowledgments
We thank Melanie Cobb, Joseph Goldstein, John Minna, Roswitha Pfragner, and Courtney Townsend for cell lines and Eric Knudsen for primary fibroblast cultures. We thank Haydn Ball and the University of Texas Southwestern Medical Center (UTSW) Protein Technology Core for peptide synthesis, the UTSW Animal Resource Center for antigen injection and blood collection, Robyn Leidel and Kate Luby-Phelps for help with SEM, Kelly Hartman for assistance with leukosome preparation, John Totenhagen and Samuria Thomas and the University of Alabama at Birmingham (UAB) Small Animal Imaging Facility for additional MRI, and HMRI Intravital Microscopy Core for IVM. We thank Champions Oncology for PDX tissue and preclinical testing in PDX models, and Pfizer for CP681301. We thank Peter Kipp for collegial advice and support. We thank Eckard Wimmer for his contribution to this work. This work was supported by an American Cancer Society Postdoctoral Fellowship (A.M.C.) and the Sackler Foundation (A.M.C. and J.A.B.); a North American Neuroendocrine Tumor Society fellowship (K. Pozo); NIH award S10OD023552-01 (M.T.); an American Thyroid Association Research Grant and the Dedman Family Scholar in Clinical Care Award (S.C.O.); an American Cancer Society Institutional Research Grant Junior Faculty Development Award (S.R.) and the SDHB Pheo/Para Coalition (S.R. and J.A.B.); a North American Neuroendocrine Tumor Society/Neuroendocrine Tumor Research Foundation Basic/Translational Science Investigator Award (R.J.-S.); the William Randolph Heart Foundation, the Robert J. Kleberg, Jr. and Helen C. Kleberg Foundation, Cancer Prevention & Research Institute of Texas Award ID RP170466, and National Cancer Institute (NCI) and the Office of Research on Women’s Health Grant 1R56CA213859 (E.T.); an American Cancer Society Research Scholars Award, and NIH awards DA033485-01, MH083711-01, NS073855-01, and R56MH116896 (J.A.B.); and NCI awards 5P30CA142543 (UTSW Simmons Comprehensive Cancer Center) and P30CA013148 (UAB O’Neal Comprehensive Cancer Center).
Footnotes
The authors declare no competing interest.
This article is a PNAS Direct Submission.
Data deposition: Phosphoproteomic data can be accessed on PhosphositePlus using the following link: https://www.phosphosite.org/Supplemental_Files.action.
This article contains supporting information online at https://www.pnas.org/lookup/suppl/doi:10.1073/pnas.2010103117/-/DCSupplemental.
Data Availability.
Phosphoproteomic data are deposited in PhosphoSitePlus (https://www.phosphosite.org/Supplemental_Files.action). Reprints and permissions information is available at www.pnas.org. Readers are welcome to comment on the online version of the paper. Correspondence and requests for materials should be addressed to J.A.B. (jbibb@uab.edu).
References
- 1.Malumbres M., Barbacid M., Mammalian cyclin-dependent kinases. Trends Biochem. Sci. 30, 630–641 (2005). [DOI] [PubMed] [Google Scholar]
- 2.Vijayaraghavan S., Moulder S., Keyomarsi K., Layman R. M., Inhibiting CDK in cancer therapy: Current evidence and future directions. Target. Oncol. 13, 21–38 (2018). [DOI] [PubMed] [Google Scholar]
- 3.Shapiro G. I., Cyclin-dependent kinase pathways as targets for cancer treatment. J. Clin. Oncol. 24, 1770–1783 (2006). [DOI] [PubMed] [Google Scholar]
- 4.Asghar U., Witkiewicz A. K., Turner N. C., Knudsen E. S., The history and future of targeting cyclin-dependent kinases in cancer therapy. Nat. Rev. Drug Discov. 14, 130–146 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dhavan R., Tsai L. H., A decade of CDK5. Nat. Rev. Mol. Cell Biol. 2, 749–759 (2001). [DOI] [PubMed] [Google Scholar]
- 6.Dhariwala F. A., Rajadhyaksha M. S., An unusual member of the Cdk family: Cdk5. Cell. Mol. Neurobiol. 28, 351–369 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tsai L. H., Delalle I., Caviness V. S. Jr., Chae T., Harlow E., p35 is a neural-specific regulatory subunit of cyclin-dependent kinase 5. Nature 371, 419–423 (1994). [DOI] [PubMed] [Google Scholar]
- 8.Angelo M., Plattner F., Giese K. P., Cyclin-dependent kinase 5 in synaptic plasticity, learning and memory. J. Neurochem. 99, 353–370 (2006). [DOI] [PubMed] [Google Scholar]
- 9.Lee M. S. et al., Neurotoxicity induces cleavage of p35 to p25 by calpain. Nature 405, 360–364 (2000). [DOI] [PubMed] [Google Scholar]
- 10.Patrick G. N. et al., Conversion of p35 to p25 deregulates Cdk5 activity and promotes neurodegeneration. Nature 402, 615–622 (1999). [DOI] [PubMed] [Google Scholar]
- 11.Pozo K., Bibb J. A., The emerging role of Cdk5 in cancer. Trends Cancer 2, 606–618 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pozo K. et al., The role of Cdk5 in neuroendocrine thyroid cancer. Cancer Cell 24, 499–511 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wei K. et al., An immunohistochemical study of cyclin-dependent kinase 5 (CDK5) expression in non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC): A possible prognostic biomarker. World J. Surg. Oncol. 14, 34 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Xie W. et al., CDK5 and its activator P35 in normal pituitary and in pituitary adenomas: Relationship to VEGF expression. Int. J. Biol. Sci. 10, 192–199 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lin H., Chen M. C., Chiu C. Y., Song Y. M., Lin S. Y., Cdk5 regulates STAT3 activation and cell proliferation in medullary thyroid carcinoma cells. J. Biol. Chem. 282, 2776–2784 (2007). [DOI] [PubMed] [Google Scholar]
- 16.Demelash A. et al., Achaete-scute homologue-1 (ASH1) stimulates migration of lung cancer cells through Cdk5/p35 pathway. Mol. Biol. Cell 23, 2856–2866 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xie W. et al., Phosphorylation of kinase insert domain receptor by cyclin-dependent kinase 5 at serine 229 is associated with invasive behavior and poor prognosis in prolactin pituitary adenomas. Oncotarget 7, 50883–50894 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Pozo K. et al., Differential expression of cell cycle regulators in CDK5-dependent medullary thyroid carcinoma tumorigenesis. Oncotarget 6, 12080–12093 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ghayee H. K. et al., Progenitor cell line (hPheo1) derived from a human pheochromocytoma tumor. PLoS One 8, e65624 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Asfari M. et al., Establishment of 2-mercaptoethanol-dependent differentiated insulin-secreting cell lines. Endocrinology 130, 167–178 (1992). [DOI] [PubMed] [Google Scholar]
- 21.Weishaupt J. H. et al., Inhibition of CDK5 is protective in necrotic and apoptotic paradigms of neuronal cell death and prevents mitochondrial dysfunction. Mol. Cell. Neurosci. 24, 489–502 (2003). [DOI] [PubMed] [Google Scholar]
- 22.Rikova K. et al., Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131, 1190–1203 (2007). [DOI] [PubMed] [Google Scholar]
- 23.Sarg B., Helliger W., Talasz H., Förg B., Lindner H. H., Histone H1 phosphorylation occurs site-specifically during interphase and mitosis: Identification of a novel phosphorylation site on histone H1. J. Biol. Chem. 281, 6573–6580 (2006). [DOI] [PubMed] [Google Scholar]
- 24.Talasz H., Sarg B., Lindner H. H., Site-specifically phosphorylated forms of H1.5 and H1.2 localized at distinct regions of the nucleus are related to different processes during the cell cycle. Chromosoma 118, 693–709 (2009). [DOI] [PubMed] [Google Scholar]
- 25.Park S. H., Yu S. E., Chai Y. G., Jang Y. K., CDK2-dependent phosphorylation of Suv39H1 is involved in control of heterochromatin replication during cell cycle progression. Nucleic Acids Res. 42, 6196–6207 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rubin S. M., Deciphering the retinoblastoma protein phosphorylation code. Trends Biochem. Sci. 38, 12–19 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Dick F. A., Rubin S. M., Molecular mechanisms underlying RB protein function. Nat. Rev. Mol. Cell Biol. 14, 297–306 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.De Azevedo W. F. et al., Inhibition of cyclin-dependent kinases by purine analogues: Crystal structure of human cdk2 complexed with roscovitine. Eur. J. Biochem. 243, 518–526 (1997). [DOI] [PubMed] [Google Scholar]
- 29.Parry D. et al., Dinaciclib (SCH 727965), a novel and potent cyclin-dependent kinase inhibitor. Mol. Cancer Ther. 9, 2344–2353 (2010). [DOI] [PubMed] [Google Scholar]
- 30.Turkdogan S. et al., Carcinoembryonic antigen levels correlated with advanced disease in medullary thyroid cancer. J. Otolaryngol. Head Neck Surg. 47, 55 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nobels F. R. et al., Chromogranin A as serum marker for neuroendocrine neoplasia: Comparison with neuron-specific enolase and the alpha-subunit of glycoprotein hormones. J. Clin. Endocrinol. Metab. 82, 2622–2628 (1997). [DOI] [PubMed] [Google Scholar]
- 32.Hirabayashi K. et al., Histopathology of gastrointestinal neuroendocrine neoplasms. Front. Oncol. 3, 2 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Giannini E. G., Testa R., Savarino V., Liver enzyme alteration: A guide for clinicians. CMAJ 172, 367–379 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Grigorian A., O’Brien C. B., Hepatotoxicity secondary to chemotherapy. J. Clin. Transl. Hepatol. 2, 95–102 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Gowda S. et al., Markers of renal function tests. N. Am. J. Med. Sci. 2, 170–173 (2010). [PMC free article] [PubMed] [Google Scholar]
- 36.Ruggiero A., Ferrara P., Attinà G., Rizzo D., Riccardi R., Renal toxicity and chemotherapy in children with cancer. Br. J. Clin. Pharmacol. 83, 2605–2614 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Molinaro R. et al., Biomimetic proteolipid vesicles for targeting inflamed tissues. Nat. Mater. 15, 1037–1046 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Molinaro R. et al., Design and development of biomimetic nanovesicles using a microfluidic approach. Adv. Mater. 30, e1702749 (2018). [DOI] [PubMed] [Google Scholar]
- 39.Kobayashi H., Watanabe R., Choyke P. L., Improving conventional enhanced permeability and retention (EPR) effects; what is the appropriate target? Theranostics 4, 81–89 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Martinez J. O. et al., Biomimetic nanoparticles with enhanced affinity towards activated endothelium as versatile tools for theranostic drug delivery. Theranostics 8, 1131–1145 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Corbo C. et al., Effects of the protein corona on liposome-liposome and liposome-cell interactions. Int. J. Nanomedicine 11, 3049–3063 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Corbo C. et al., Unveiling the in vivo protein corona of circulating leukocyte-like carriers. ACS Nano 11, 3262–3273 (2017). [DOI] [PubMed] [Google Scholar]
- 43.Corbo C. et al., The impact of nanoparticle protein corona on cytotoxicity, immunotoxicity and target drug delivery. Nanomedicine (Lond.) 11, 81–100 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Coussens L. M., Werb Z., Inflammation and cancer. Nature 420, 860–867 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Gonzalez H., Hagerling C., Werb Z., Roles of the immune system in cancer: From tumor initiation to metastatic progression. Genes Dev. 32, 1267–1284 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bollag G. et al., Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature 467, 596–599 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Flaherty K. T. et al., Inhibition of mutated, activated BRAF in metastatic melanoma. N. Engl. J. Med. 363, 809–819 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Fiskus W., Mitsiades N., B-Raf inhibition in the clinic: Present and future. Annu. Rev. Med. 67, 29–43 (2016). [DOI] [PubMed] [Google Scholar]
- 49.Chandrasekharappa S. C. et al., Positional cloning of the gene for multiple endocrine neoplasia-type 1. Science 276, 404–407 (1997). [DOI] [PubMed] [Google Scholar]
- 50.Lemmens I. et al., Identification of the multiple endocrine neoplasia type 1 (MEN1) gene. The European Consortium on MEN1. Hum. Mol. Genet. 6, 1177–1183 (1997). [DOI] [PubMed] [Google Scholar]
- 51.Lemos M. C., Thakker R. V., Multiple endocrine neoplasia type 1 (MEN1): Analysis of 1336 mutations reported in the first decade following identification of the gene. Hum. Mutat. 29, 22–32 (2008). [DOI] [PubMed] [Google Scholar]
- 52.Agarwal S. K. et al., Menin molecular interactions: Insights into normal functions and tumorigenesis. Horm. Metab. Res. 37, 369–374 (2005). [DOI] [PubMed] [Google Scholar]
- 53.Mulligan L. M. et al., Germ-line mutations of the RET proto-oncogene in multiple endocrine neoplasia type 2A. Nature 363, 458–460 (1993). [DOI] [PubMed] [Google Scholar]
- 54.Donis-Keller H. et al., Mutations in the RET proto-oncogene are associated with MEN 2A and FMTC. Hum. Mol. Genet. 2, 851–856 (1993). [DOI] [PubMed] [Google Scholar]
- 55.Hofstra R. M. et al., A mutation in the RET proto-oncogene associated with multiple endocrine neoplasia type 2B and sporadic medullary thyroid carcinoma. Nature 367, 375–376 (1994). [DOI] [PubMed] [Google Scholar]
- 56.Wells S. A. Jr. et al., Vandetanib in patients with locally advanced or metastatic medullary thyroid cancer: A randomized, double-blind phase III trial. J. Clin. Oncol. 30, 134–141 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kurzrock R. et al., Activity of XL184 (Cabozantinib), an oral tyrosine kinase inhibitor, in patients with medullary thyroid cancer. J. Clin. Oncol. 29, 2660–2666 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Sherman S. I., Lessons learned and questions unanswered from use of multitargeted kinase inhibitors in medullary thyroid cancer. Oral Oncol. 49, 707–710 (2013). [DOI] [PubMed] [Google Scholar]
- 59.Beijersbergen R. L. et al., E2F-4, a new member of the E2F gene family, has oncogenic activity and associates with p107 in vivo. Genes Dev. 8, 2680–2690 (1994). [DOI] [PubMed] [Google Scholar]
- 60.Hijmans E. M., Voorhoeve P. M., Beijersbergen R. L., van ’t Veer L. J., Bernards R., E2F-5, a new E2F family member that interacts with p130 in vivo. Mol. Cell. Biol. 15, 3082–3089 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Trimarchi J. M., Lees J. A., Sibling rivalry in the E2F family. Nat. Rev. Mol. Cell Biol. 3, 11–20 (2002). [DOI] [PubMed] [Google Scholar]
- 62.Litovchick L. et al., Evolutionarily conserved multisubunit RBL2/p130 and E2F4 protein complex represses human cell cycle-dependent genes in quiescence. Mol. Cell 26, 539–551 (2007). [DOI] [PubMed] [Google Scholar]
- 63.Sadasivam S., DeCaprio J. A., The DREAM complex: Master coordinator of cell cycle-dependent gene expression. Nat. Rev. Cancer 13, 585–595 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Weinstein J. L., Katzenstein H. M., Cohn S. L., Advances in the diagnosis and treatment of neuroblastoma. Oncologist 8, 278–292 (2003). [DOI] [PubMed] [Google Scholar]
- 65.Newman E. A., Nuchtern J. G., Recent biologic and genetic advances in neuroblastoma: Implications for diagnostic, risk stratification, and treatment strategies. Semin. Pediatr. Surg. 25, 257–264 (2016). [DOI] [PubMed] [Google Scholar]
- 66.Toyoda H., Yin J., Mueller S., Wimmer E., Cello J., Oncolytic treatment and cure of neuroblastoma by a novel attenuated poliovirus in a novel poliovirus-susceptible animal model. Cancer Res. 67, 2857–2864 (2007). [DOI] [PubMed] [Google Scholar]
- 67.Chen Z. et al., Multiple CDK inhibitor dinaciclib suppresses neuroblastoma growth via inhibiting CDK2 and CDK9 activity. Sci. Rep. 6, 29090 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Zhuang K. et al., CDK5 functions as a tumor promoter in human colorectal cancer via modulating the ERK5-AP-1 axis. Cell Death Dis. 7, e2415 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Goodyear S., Sharma M. C., Roscovitine regulates invasive breast cancer cell (MDA-MB231) proliferation and survival through cell cycle regulatory protein cdk5. Exp. Mol. Pathol. 82, 25–32 (2007). [DOI] [PubMed] [Google Scholar]
- 70.Liu R. et al., Cdk5-mediated regulation of the PIKE-A-Akt pathway and glioblastoma cell invasion. Proc. Natl. Acad. Sci. U.S.A. 105, 7570–7575 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Strock C. J. et al., Cyclin-dependent kinase 5 activity controls cell motility and metastatic potential of prostate cancer cells. Cancer Res. 66, 7509–7515 (2006). [DOI] [PubMed] [Google Scholar]
- 72.Dehqanzada Z. A. et al., Assessing serum cytokine profiles in breast cancer patients receiving a HER2/neu vaccine using Luminex technology. Oncol. Rep. 17, 687–694 (2007). [PubMed] [Google Scholar]
- 73.Carter A., et al. , Supplementary Files For Selected High Throughput Protein Modification Publications from Cell Signaling Technology. PhosphositePlus. https://www.phosphosite.org/Supplemental_Files.action. Deposited 20 March 2020.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Phosphoproteomic data are deposited in PhosphoSitePlus (https://www.phosphosite.org/Supplemental_Files.action). Reprints and permissions information is available at www.pnas.org. Readers are welcome to comment on the online version of the paper. Correspondence and requests for materials should be addressed to J.A.B. (jbibb@uab.edu).