Abstract
Luffa is a kind of melon crop widely cultivated in temperate regions worldwide. Browning is one of the serious factors affecting the quality of Luffa. Therefore, the molecular mechanism of Luffa browning is of great significance to study. However, the molecular diversity of Luffa cultivars with different browning-resistant abilities has not been well elucidated. In our study, we used high-throughput sequencing to determine the transcriptome of two Luffa cylindrica cultivars ‘2D-2’ and ‘35D-7’. A total of 115,099 unigenes were clustered, of which 22,607 were differentially expression genes (DEGs). Of these DEGs, 65 encoding polyphenol oxidase, peroxidase, or ascorbate peroxidase were further analyzed. The quantitative real-time PCR (RT-qPCR) data indicated that the expression levels of the LcPPO gene (Accession No.: Cluster-21832.13892) was significantly higher in ‘35D-7’ compared with that in ‘2D-2’. Several POD genes (Accession No.: Cluster-21832.19847, Cluster-21832.30619 and Cluster-48491.2) were also upregulated. Analysis of the plantTFDB database indicated that some transcription factors such as WRKY gene family may also participate in the regulation of Luffa browning. The results indicated that the divergence of genes expression related to enzymatic reaction may lead to the different browning resistances of Luffa. Our study will provide a theoretical basis for breeding of browning-resistant Luffa.
Keywords: Expression, Luffa, Polyphenol oxidase, Transcriptome, Browning
Introduction
Luffa (Luffa cylindrica), belongs to Luffa Mill. of Cucurbitaceae, is an annual herbaceous climbing plant (Oboh & Aluyor, 2009). Luffa is native to India and tropical southern Asia and is mainly distributed in Asia, Oceania, Africa, and America. Luffa has eight species worldwide. China mainly cultivates Luffa cylindrica Roem. and Luffa acutangula Roxb. (Oboh & Aluyor, 2009; Xu et al., 2018). Compared with other Cucurbitaceae plants, the most remarkable characteristic of Luffa is the abundant reticular fibers composed of vascular bundles within fruit. With its peel, seeds and pulp removed, Luffa is considered as a traditional Chinese medicine, Luffa sponge (Asma & Muhammad, 2013). Luffa contains large amounts of vitamin B, vitamin C, saponins, citrulline, and other nutrients. Therefore, Luffa has effects on scurvy prevention, antivirus, cosmetology and so on (Mazali & Alves, 2005). Since Luffa was introduced into China during the Song Dynasty, it has been widely cultivated in China due to its strong adaptability, and a variety of local cultivars have gradually been formed (Su et al., 2010). However, the skin, pulp, and juice of Luffa fruits are prone to browning during the transportation and processing; browning seriously affects the nutritional value and flavor of the fruit, leading to the decline in the commercial quality of Luffa products (Friedman, 1991; Vamosvigyazo, 1995). Therefore, cultivating browning-resistant Luffa germplasm resources is of great significance.
Browning is a common discoloration phenomenon and a beneficial step in food processing, such as the productions of soy sauce, bread, black tea and coffee, etc. (Okada, Katsura & Kobayashi, 2004; Purlis & Salvadori, 2007; Troup et al., 2015; Lee & Knag, 2017). However, browning can affect the flavor and reduce the nutritional value of most fruits and vegetables and is therefore considered the second largest cause of quality loss (Lim, Cheun & Wong, 2019). The two main types of browning are as follows: enzymatic browning, which is the result of polyphenol oxidase (PPO) that catalyzes the formation of quinone and its polymer; and non-enzymatic browning, which involves Maillard reaction and caramelization (Buera et al., 1987; Zamora & Hidalgo, 2005). The browning of Luffa is mainly due to enzymatic browning. When Luffa is stimulated by collision, friction, or heating, the content of active oxygen increases, thereby damaging the cellular structure of Luffa. The compartmental distribution of enzymes and phenolic substances will be broken and lead to the formation of quinones (Xu et al., 2018). Numerous enzymes involved in this process are polyphenol oxidase, ascorbate peroxidase (APX), and peroxidase (POD), etc. (Nicolas et al., 1994; Jiang et al., 2016). Research on the mechanism of browning has been carried out in several species (Ali et al., 2016; Liu et al., 2019). Jiang (2000) found that anthocyanins, polyphenol oxidase, and phenols participated in the lychee pericarp browning. Studies in potato showed that the antisense expression of PPO genes can inhibit the browning of potato tubers (Bachem et al., 1994). Browning in fresh eggplant is related to the phenolic content and specific activity of PPO (Mishra, Gautam & Sharma, 2013). These results provide a reference for the exploration of the mechanism of Luffa browning and make up for the lack of related studies.
The development of transcriptome sequencing techniques has led to identification of novel transcripts, splice isoforms, and expression differences in numerous species (Wu et al., 2014; Wang et al., 2015). A large amount of sequences and plant molecular information have been revealed using omics analysis (Li et al., 2018; Li et al., 2020), but the application of such research in Luffa is still lacking. In the present study, two Luffa cultivars were used. ‘2D-2’ is a kind of browning-resistant cultivar and ‘35D-7’ is a cultivar prone to browning. Both of the two cultivars are self-bred lines which were originated from ‘Early Rou Luffa’ and ‘Short Stick-like Rou Luffa’, respectively. Transcriptome analysis was conducted to investigate the divergence between the two Luffa cultivars with browning-resistant or -sensitive characteristics. Differentially expression genes (DEGs) related to Luffa browning were identified. Expression levels of related genes were also validated and analyzed. Our results can provide potential insights into the molecular regulation of Luffa browning and a basis for cultivating improved Luffa germplasm with high browning resistance.
Materials & Methods
Plant materials and sample collection
The seeds of the two Luffa cultivars, ‘2D-2’ and ‘35D-7’, were preserved in the Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences and cultivated in the farm of Luhe Base (118.62°N, 32.49°E, Nanjing, China). The field was approved for using by Jiangsu Academy of Agricultural Sciences (Contract No.: SC2018001). The fruits of each cultivar were collected after 10 days after pollination. The fruit pulps near the end of the stalk that are prone to browning were sampled for subsequent experiments. Samples were collected from three individual fruits. The samples were immersed in liquid nitrogen immediately and stored at −80 °C.
RNA extraction and quality control
The total RNA of each sample was extracted according to the instructions of the plant RNA simple total RNA (Tiangen, Beijing, China). Agarose gel electrophoresis and Agilent Bioanalyzer 2100 (Agilent, China) was used to detect the integrity of the RNA samples. RNA concentration and purity were measured by NanoPhotometer (Implen, Germany) and Qubit 2.0 Fluorometer (ThermoFisher, USA).
Construction of cDNA library and sequencing
Construction of cDNA library was conducted in accordance with the instructions of NEBNext Ultra Directional RNA Library Prep Kit for Illumina (NEB, Beijing, China). Ribosomal RNA was removed from total RNA to obtain the mRNA of the samples. Fragmentation buffer was added to break RNA into small fragments, which were used for the synthesis of the first-strand cDNA. Buffer, dNTPs (dUTP, dATP, dGTP, and dCTP), and DNA polymerase I were utilized to synthesize the second chain. The obtained double-stranded cDNA was purified by AMPure XP beads. After terminal repairing, A-tail adding, indexing adapter ligation, and fragment size selection, PCR amplification was conducted to enrich the short fragments for construction of the final cDNA library. The cDNA library was further quantified and its insert size was detected by Agilent 2100 to ensure that its quality could achieve the standard for sequencing. Sequencing was performed on the NovaSeq 6000 Sequencing System (Illumina, California, USA). Paired-end (150bp) sequencing was set as sequencing strategy.
Data filtering and de novo assembly
Adapter-related reads, low-quality reads, and reads with relatively high number of base ‘N’ were filtered to ensure the accuracy of subsequent analysis. Clean reads after raw data filtering, error rate checking, and GC content distribution checking were used for subsequent transcript assembly. Trinity 2.6.6 was used to assemble the clean reads to obtain the reference sequence (Grabherr et al., 2013). The longest cluster sequence obtained by Corset hierarchical clustering (https://code.google.com/p/corset-project/) was used as unigene for subsequent analysis.
Unigene function annotation
BLAST software was used to search the unigene sequence against Kyoto Encyclopedia of Genes and Genomes (KEGG) (Minoru et al., 2007), NCBI non-redundant protein (NR) (Pruitt, Tatiana & Maglott, 2005), Swiss-Prot, Gene Ontology (GO) (Mi et al., 2019), Clusters of Orthologous Groups of proteins (COG) (Natale et al., 2000), and TrEMBL database (Boeckmann et al., 2003) (E-value ≤ 1E-5). The predicted amino acid sequences of the best aligning unigenes were transferred and searched by Pfam database using HMMER software to obtain the annotation information of the unigenes (El-Gebali et al., 2019). BLASTX was used for further functional categorization with the E-value threshold of 1E-5.
Identification of DEGs
DESeq2 was utilized to analyze the differential expression among samples. Non-standardized reads were counted by featureCounts. Multiple hypothesis testing of P-value was conducted based on Benjamini–Hochberg method. The transcription abundance of the samples was expressed by fragments per kilobase of transcript per million fragments mapped (FPKM). The rationale of the DEGs was set as follows: —log2fold change—≥1 and the false discovery rate (FDR) < 0.05. The function of the DEGs was annotated by KEGG, GO, and COG database, and DEGs related to browning were identified according to the annotation information. Transcription factor annotation was identified by iTAK version 1.7 software in reference of plantTFDB (Jin et al., 2017).
Validation of the DEGs through quantitative real-time PCR (RT-qPCR)
Primer Premier 6.0 was used to design the specific primers of the DEGs. The primer sequences are listed in Table 1. Lc18SRNA was selected as reference gene (Singh et al., 1998). RT-qPCR was performed on Bio-Rad CFX96 PCR platform (Bio-Rad, USA) using Hieff qPCR SYBR Green Master Mix (Yeason, Shanghai, China).The reaction mixture contains 10 µL of enzyme, 2 µL of diluted cDNA, 0.4 µL of forward or reverse primer, and sterile deionized water was used to replenish the volume to 20 µL. The reaction procedure was set as follows: 95 °C for 5 min; 95 °C for 10 s and 60 °C for 30 s, 40 cycles in total. A melting curve was established by increasing from 65 to 95 °C to verify the amplification specificity. The obtained cycle threshold values were calculated using Pfaffl method (2001). Three independent biological replicates were set for each sample. Significance analysis was carried out in Microsoft Excel 2016 according to the method of one-way ANOVA.
Table 1. Primers used for RT-qPCR.
| Gene symbol | Description | Forward primer sequence (5′to 3′) | Reverse primer sequence (5′to 3′) |
|---|---|---|---|
| Cluster-21832.13892 | polyphenol oxidase | CGAATGTTCACTGTGCCTAT | CTTCGGAGCGTCGTAGT |
| Cluster-21832.19847 | peroxidase | CAACATTGTCCGCCGTGAG | TCTGCTGTCTCTTCTTCCGTATA |
| Cluster-21832.30619 | ACTGCTTCGTCAATGGATGTG | CTGTTGGCTATTCTGCTGTCTT | |
| Cluster-21832.38395 | GGTGGCTCTGCTTCTTATTACATA | TTGGTCTCCCTCTCTTCTTCTTT | |
| Cluster-23349.0 | GCATCACAAGAGCCAGGAAG | GTGGTCGGAGCATATCTATCAAC | |
| Cluster-2660.0 | GAGCGAGAGTGAAGCCAATC | CACCTGAGAGCACAACAACAT | |
| Cluster-48491.2 | GGTGAAGTGTACGGTGATGATG | GAATCTATCTGCCTCTATCCTCCT | |
| Cluster-5215.0 | GCTTCCACCAACATACACTCC | GCCGCCTACCAGAATCACT | |
| Cluster-19973.6 | ascorbate peroxidase | ACGGATGCTAACAACGAGGAT | CTGAACTGAAGCGGATGGATG |
| Cluster-21832.12599 | GCTCCTGAAGACTACATTCT | GACCAGCATTAGCACCAT | |
| Cluster-21832.17816 | CTGCTGTTAGTGAGGAGTAT | CAATAGCCTAATAGCGATGTC | |
| Cluster-21832.12605 | GCCAACTGATGCTGCTCTC | GAATCTTCTGCTTCATTGAGTCTG | |
| Lc18SRNA | reference gene | GTGTTCTTCGGAATGACTGG | ATCGTTTACGGCATGGACTA |
Results
Sequencing and assembly of Luffa
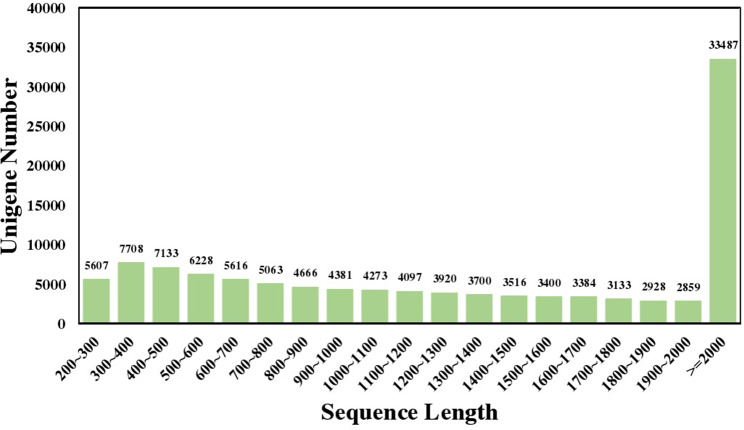
A total of 67,969,672 and 59,573,164 raw reads were detected by sequencing the two samples, ‘2D-2’ and ‘35D-7’, respectively. The sequencing raw data can be found in the Sequence Read Archive (SRA) database (Accession No.: SRP252794). Assembly sequences has been deposited at TSA database under the Accession GILZ00000000. After data filtering, 9.67 G and 8.49 G clean high-quality reads were obtained. The percentage of Q20 and Q30 of ‘2D-2’ and ‘35D-7’ were 98.53% and 95.3%, 98.52% and 95.3%, respectively. The GC percentage in high-quality reads were 44.12% and 44.74%, respectively. The number of the contigs found by assembling the sequences according to Trinity was 144,048 with a mean length of 1,311 bp and an N50 length of 2,160 bp. After clustering, the longest clusters were identified as unigenes, and the total number of unigenes was 115,099 and the mean length and N50 length was 1,573 bp and 2,235 bp, respectively. The length distribution of the unigenes is shown in Fig. 1. Most of the unigenes were over 2,000 bp in length (33,487, 29.09%), then followed by 300 to 500 bp (7,708, 6.70%).
Figure 1. Sequence length distribution of the Luffa unigene library.
Functional annotation and classification
All unigenes were annotated against seven databases (KEGG, NR, SwissProt, TrEMBL, KOG, GO and Pram), and the detailed numbers of unigenes are listed in Table 2 (Table S1). The NR database annotated most of the 115,099 unigenes (99,246, 80.14%). The GO annotation classified the unigenes according to three categories of GO (biological process, cellular component, and molecular function), the results are displayed in Fig. S1. A total of 78,549 unigenes were branched into 58 function groups.
Table 2. Summary of annotation of Luffa unigenes.
| Annotation database | Annotated number | Proportion of all unigenes (%) |
|---|---|---|
| KEGG | 73,507 | 63.86 |
| NR | 92,246 | 80.14 |
| SwissProt | 66,525 | 57.8 |
| TrEMBL | 91,475 | 79.48 |
| KOG | 57,412 | 49.88 |
| GO | 78,549 | 68.24 |
| Pfam | 70,026 | 60.84 |
| All Annotated | 92,524 | 80.39 |
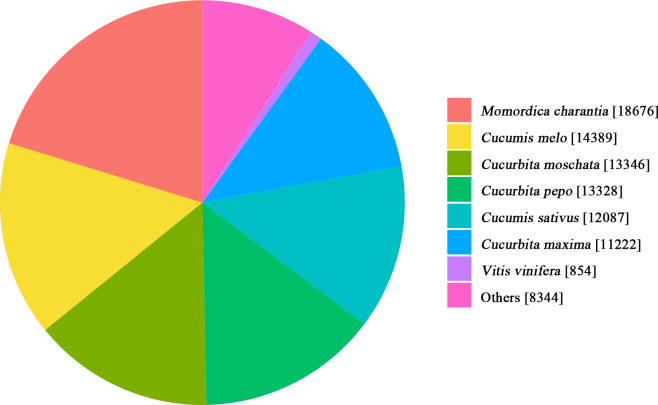
Comparison with the NR database showed similarity in the transcripts between Luffa and related species (Fig. 2). As shown in the pie chart, Luffa and Momordica charantia had the highest similarity, followed by Cucumis melo, Cucurbita moschata, Cucurbita pepo, Cucumis sativus, Cucurbita maxima and Vitis vinifera which have a great number of matches with Luffa. Except Vitis vinifera belongs to Vitaceae, the other species similar to Luffa all belong to Cucurbitaceae.
Figure 2. Species distribution of the Luffa unigene library against the NR database.
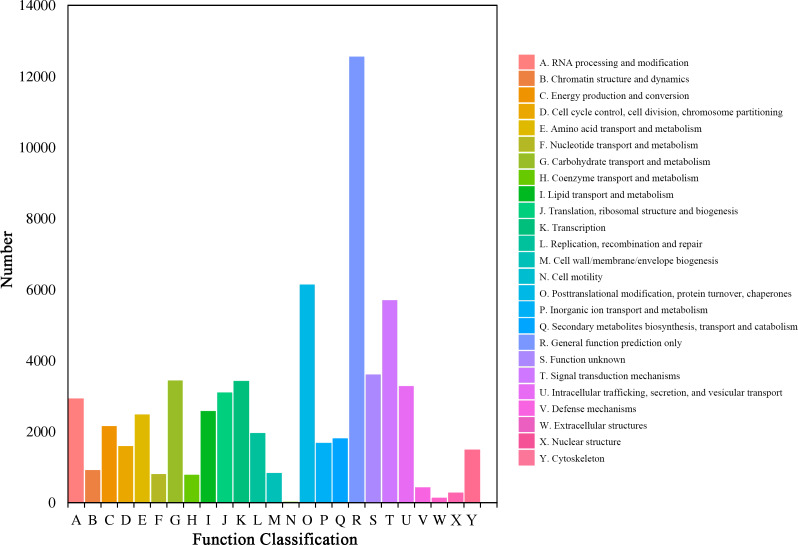
The unigenes were also clustered by the ‘eukaryotic Orthologous Groups’ (KOG) of COG database. Based on the orthologous and evolutionary relationships, homologous genes from different species were divided into different clusters and the function of unigenes was annotated (Fig. 3). The top three categories among the all 25 categories were ‘general function prediction only’ (12,558, 19.59%), ‘posttranslational modification, protein turnover, chaperones’ (6,140, 9.58%), and ‘signal transduction mechanisms’ (5,702, 8.90%), respectively. However, ‘cell motility’ (24, 0.04%), ‘extracellular structures’ (132, 0.21%), and ‘nuclear structure’ (277, 0.43%) had the lowest percentage.
Figure 3. COG function classification of the Luffa unigene library.
Identification of DEGs
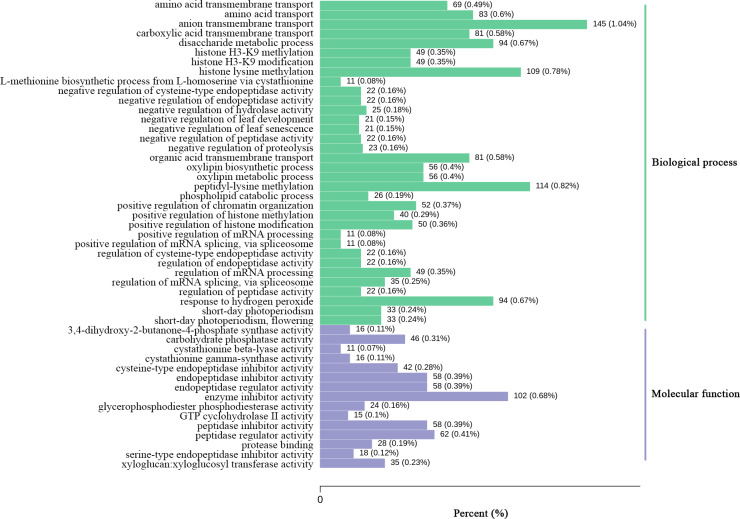
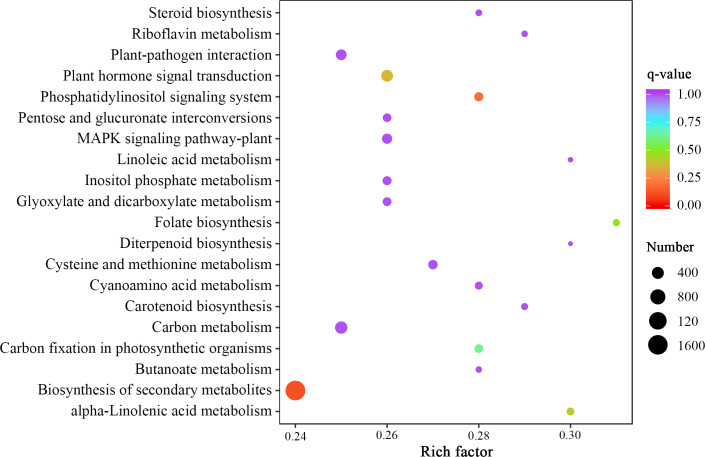
By comparing the expression levels represented by FPKM of two Luffa cultivars ‘2D-2’ and ‘35D-7’, a total of 22,607 DEGs were identified. Of the DEGs, 13,465 were down-regulated and 9,142 were up-regulated in ‘35D-7’. The overall distribution of the expression levels and annotation of DEGs was determined (Table S2). The GO database was used to annotate the potential functions of the DEGs (Fig. S2). The top 50 GO-terms that possessed the lowest q-value are shown in Fig. 4. These GO-terms were distributed into two categories, ‘biological process’ and ‘molecular function’ based on background genes. The major GO-terms related to ‘biological process’ were ‘anion transmembrane transport’ (145, 1.04%), ‘peptidyl-lysine methylation’ (114, 0.82%), and ‘histone lysine methylation’ (109, 0.78%). As for the ‘molecular function’, ‘enzyme inhibitor activity’ (102, 0.68%) and ‘peptidase regulator activity’ (62, 0.41%) were the predominate GO-terms. The result of KEGG analysis (Fig. 5, Fig. S3) indicated that most of the DEGs were enriched in the ‘biosynthesis of secondary metabolites’ (ko01110) pathway (1,649, 21.47%). Pathways ‘plant-pathogen interaction’ (ko04626) and ‘carbon metabolism’ (ko01200) contained 291 (3.79%) and 444 (5.78%) DEGs. In ‘plant hormone signal transduction’ (ko04075), 386 (5.03%) DEGs were enriched.
Figure 4. GO enrichment analysis of the DEGs identified in the unigene libraries between two Luffa cultivars.
Figure 5. Statistics of KEGG enrichment of the DEGs identified in the unigene libraries between two Luffa cultivars.
Identification of transcription factors among the DEGs
According to the rules of transcription factor family defined in plantTFDB database, several transcription factor families have been identified among the DEGs (Table 3). Twenty-six DEGs were classified into HB transcription factors and the number of up- and down regulated genes was similar. Meanwhile, 24 and 20 DEGs were identified as MYB transcription factors and C2H2 transcription factors, respectively, which were mostly downregulated. Other transcription factors, such as AP2, bHLH, bZIP, and NAC, were also annotated. Among the identified transcription factors, 47 DEGs only expressed in Luffa cultivar ‘35D-7’. The most significant difference in expression was found in Cluster-21832.27579 which was identified as bHLH transcription factor, followed by Cluster-10198.4 and Cluster-17143.0 belonging to Whirly and HSF family, respectively. On the contrary, a total of 107 DEGs were only detected in ‘2D-2’.
Table 3. Transcription factors identified in the DEGs.
| Transcription factor | Number of DEGs | ||
|---|---|---|---|
| Total | Up-regulated | Down-regulated | |
| AP2 | 16 | 9 | 7 |
| B3 | 17 | 6 | 11 |
| bHLH | 14 | 8 | 6 |
| bZIP | 17 | 9 | 8 |
| C2H2 | 20 | 5 | 15 |
| C3H | 11 | 3 | 8 |
| FAR1 | 11 | 3 | 8 |
| GARP | 10 | 6 | 4 |
| GRAS | 7 | 5 | 2 |
| GRF | 4 | 2 | 2 |
| HB | 26 | 14 | 12 |
| HD-ZIP | 9 | 7 | 2 |
| HSF | 11 | 7 | 4 |
| MADS | 8 | 2 | 6 |
| MYB | 24 | 10 | 14 |
| NAC | 19 | 10 | 9 |
| NF-YA | 6 | 1 | 5 |
| SBP | 14 | 4 | 10 |
| TCP | 4 | 0 | 4 |
| Trihelix | 7 | 4 | 3 |
| Whirly | 2 | 1 | 1 |
| WRKY | 17 | 9 | 8 |
Identification of browning-related DEGs in Luffa
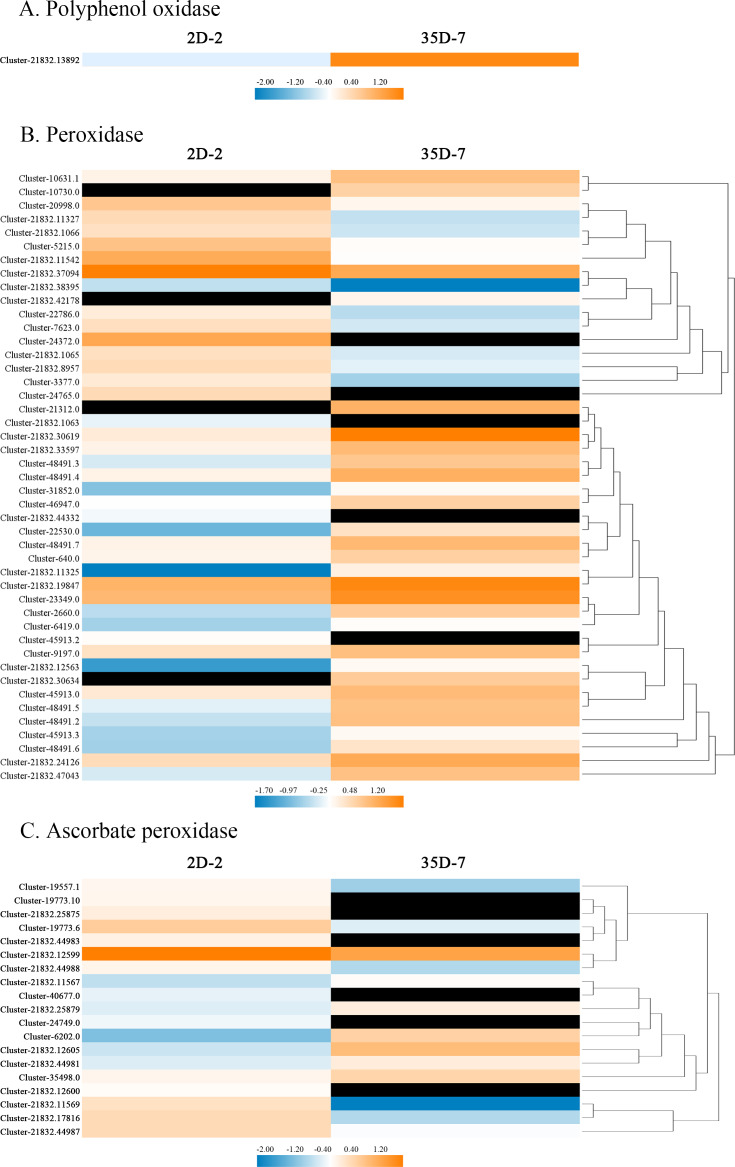
To explore the gene regulatory mechanism of Luffa browning, we compared the expression levels of browning-related genes in the two Luffa cultivars with different resistance to browning. A total of 65 DEGs encoded PPO, POD, or APX were annotated to have roles in the enzymatic reaction in Luffa browning. Figure 6 shows the heatmap indicating the expression levels of these DEGs. The expression level of LcPPO (Accession No: Custer-21832.13892) was significantly higher in the Luffa cultivar ‘35D-7’ than in ‘2D-2’. Among the 45 DEGs related to LcPOD, 17 had lower expression and 28 had higher expression in ‘35D-7’ than in ‘2D-2’. In particular, the log2fold change of two POD genes (Accession No: Cluster-21312.0 and Cluster-21832.30634, respectively) reached up to 12.16 and 11.10, respectively. Nineteen APX-related genes were also identified, but most of them were down-regulated in ‘35D-7’.
Figure 6. Heatmap of the expression levels of browning-related genes identified in the DEGs.
(A) Polyphenol oxidase, (B) peroxidase, (C) ascorbate peroxidase. Heat maps were created by the log2 relative abundance of the DEGs. Blue and orange represent low and high expression, respectively. Black represents no expression detected.
Validation of expression levels of browning-related genes in Luffa
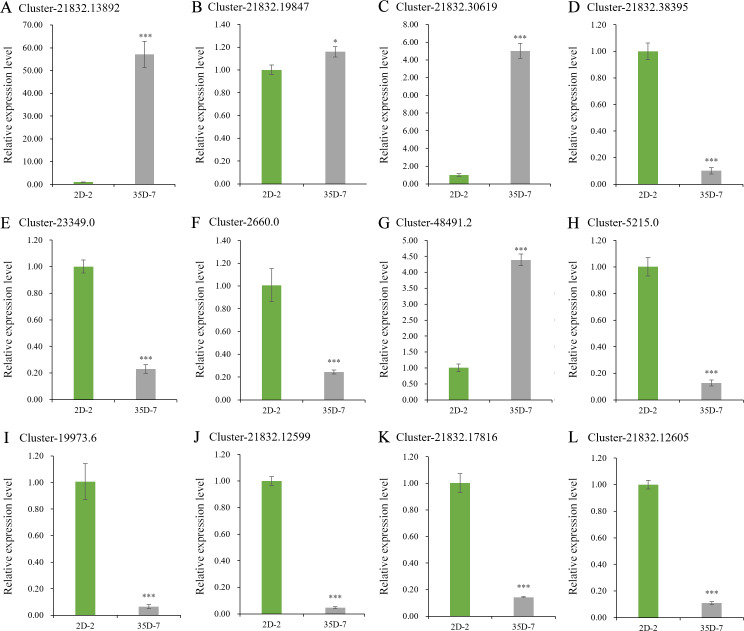
The expression levels of some DEGs, including LcPPO, seven LcPOD genes, and four LcAPX genes were detected and analyzed to validate the expression levels of DEGs related to Luffa browning (Fig. 7), their sequences are placed in the Data S1. The expression level of LcPPO (Accession No: Cluster-21832.13892) in ‘35D-7’ was 57.09 times that in ‘2D-2’. Among the selected LcPOD genes, three (Accession No: Cluster-21832.19847, Cluster-21832.30619 and Cluster-48491.2) were upregulated significantly in ‘35D-7’. The expression levels of all four LcAPX genes were lower in Luffa cultivar ‘35D-7’ than in ‘2D-2’. In general, nine of the 12 genes exhibited consistent expression trends in the results of RT-qPCR and transcriptome analysis.
Figure 7. Expression levels of browning-related genes.
The unigene ‘Cluster-21832.13892’ (A) was identified as LcPPO encoding polyphenol oxidase. The unigenes ‘Cluster-21832.19847’, ‘Cluster-21832.30619’, ‘Cluster-2183.38395’, ‘Cluster-23349.0’, ‘Cluster-2660.0’, ‘Cluster-48491.2’ and ‘Cluster-5215.0’ (B-H) were identified as LcPODs encoding peroxidase. The unigenes ‘Cluster-19973.6’, ‘Cluster-21832.12599’, ‘Cluster-21832.17816’ and ‘Cluster-21832.12605’ (I-L) were identified as LcAPXs encoding ascorbate peroxidase.
Discussion
Luffa is one of the most important melon vegetables, and its fruits are the main edible organ (Liu et al., 2017; Xu et al., 2018). Browning of fruits and vegetables is a complex process that involves several factors that work together (Ioannou & Ghoul, 2013). A number of methods, including physical, chemical, and biotechnology aspects have been studied to inhibit browning (Friedman, 1991; Chisari, Barbagallo & Spagna, 2007; Lopez-Nicolas et al., 2007; Arogundade & Mu, 2012). Studies on the browning tolerance of Luffa mainly focused on the physiological mechanism and variety selection. However, the related molecular genetic mechanism and genetic engineering have been rarely addressed. This lack of in-depth understanding affects the breeding progress of Luffa germplasm.
Transcriptome sequencing is one of the main methods used to explore gene expression, and is widely used in the fields of candidate gene discovery, functional identification and genetic improvement (Wang, Gertein & Snyder, 2009; Conesa et al., 2016). In the present study, we determined the transcriptome of two Luffa inbred cultivars ‘2D-2’ and ‘35D-7’. A total of 115,099 unigenes were clustered and annotated against seven databases. Among these unigenes, 22,607 DEGs with potential role in the regulation of Luffa browning resistance were identified. In 2017, Zhu et al. analyzed the transcriptome changes during browning and found that less unigenes (58,073) were assembled in the fresh-cut Luffa cyindrica ‘Fusi-3’ fruits (Zhu et al., 2017). These findings indicate differences in the transcriptomes of different varieties. The annotation against the KOG database was also similar to that reported in a previous study (Zhu et al., 2017). Comparison of the annotation of the unigenes shows that a large number of unigenes of Luffa were matched to Momordica charantia in addition to Cucumis melo. This result provides a new reference for the functional exploration of Luffa genes.
The identification of DEGs provides new insights about Luffa browning. Based on the results of the KEGG annotation, a large number of DEGs were enriched in the ‘biosynthesis of secondary metabolites’ and ‘carbon metabolism’ pathways. Phenols are the substrate of enzymatic browning. Phenols are important secondary metabolites formed by replacing the hydrogen atoms in the aromatic ring into hydroxyl or functional derivatives (Parr & Bolwell, 2000). The browning substrate of banana is dopamine (Griffiths, 1959). Browning in pear skin is caused by the phenols mainly composed of l-epicatechin (Siegelman, 1955). The main browning substrates of chufa are (-)-gallocatechin gallate, (-)-epicatechin gallate and (+)-catechin gallate (Sun et al., 2010). The polyphenol oxidase substrate content is likely different among different Luffa cultivars. DEGs related to plant hormone signal transduction and α-linolenic acid metabolism were also found in the two cultivars. Plant hormone is an important regulator of plant growth and development and participates in a variety of biological processes in plants (Friedman, 1991; Kulka, 2008). α-linolenic acid is a common fatty acid in plants that is mainly found in cell membrane lipids in plants (Li et al., 2016). The destruction of cellular structure occurs during browning. The different expression levels of α-linolenic acid-related genes demonstrated that cell membrane structure divergence may exist in different Luffa cultivars and may result in the easy destruction of the cell structure.
Among the identified DEGs, oxidase genes, which are thought to be involved in browning were the focus of this study. High expression levels of several genes were found in Luffa ‘35D-7’, including a LcPPO gene, a few LcPOD and LcAPX genes. RT-qPCR analysis was also performed to validate the expression of these gene in the Luffa samples. The LcPPO gene expression was extremely high in ‘35D-7’ and was more than 50 times that in ‘2D-2’. PPO is a metalloproteinase encoded by nuclear gene and can oxidize phenol substrates to quinone (Mayer, 2006). With the rapid development of cloning technology, PPO genes have been obtained from a variety of plants, including potato (Wang et al., 2003), wheat (Demeke & Morris, 2002), tea (Min, Liu & He, 2012), banana (Galeazzi, Sgarbieri & Conatantinides, 1981), etc. A close correlation exists between the expression levels of PPO genes and browning (Chi et al., 2014). In 2018, Zhu et al. cloned three PPO genes from Luffa cylindrica (L.) Roem and analyzed their expression profiles. The PPO genes expressed in different tissues of Luffa and responded to the postharvest storage and fresh cutting conditions (Zhu et al., 2018). In the present study, we found different expressions of LcPPO genes in different Luffa cultivars. This result indicated that PPO might be a key gene in regulating Luffa browning. Our results confirmed differences in the expression among different browning-resistant varieties. Several LcPOD genes were also found, but the expression differences were not as significant as LcPPO genes. In the presence of hydrogen peroxide, POD can catalyze the oxidation of phenols as a supplement to PPO (Chisari, Barbagallo & Spagna, 2007). Therefore, LcPOD genes were also involved in the browning process in Luffa. Previous study of Zhu et al. has confirmed that genes PPO, PAL (encoding phenylalanine ammonia lyase), POD, CAT (encoding catalase) and SOD (encoding superoxide dismutase) may associate with the enzymatic browning of fresh-cut Luffa (Zhu et al., 2017). We compared the expression levels of these genes in two Luffa varieties ‘2D-2’ and ‘35D-7’ and found that four genes, encoding PPO, PAL, POD and CAT enzyme, respectively, showed the same trend as the changes during browning of ‘Fusi-3’ (Table S3). The combined results demonstrated that some genes that play roles in the browning process also have effect on the browning tolerance of different Luffa cultivars. Further study on the functions of these genes will provide an important basis for the cultivation of Luffa varieties that are resistant to browning.
Transcription factors are proteins that can bind to specific sequences to regulate the expression of target genes (Katagiri & Chua, 1992; Schwechheimer, Zourelidou & Bevan, 1998). Transcription factors play an important role in plant growth and development, stress responses, and other aspects (Ramachandran, Hiratsuka & Chua, 1994; Meshi & Iwabuchi, 1995). We annotated the transcription factors according to the sequence obtained by the transcriptome analysis. A large number of transcription factors were identified among the DEGs, including some common transcription factor families AP2, MYB, WRKY etc. Our previous study found that some miRNAs related to browning can regulate downstream transcription factors (Xu et al., 2018). The result of Zhu considered that members of Group II WRKY superfamily may be related to the Luffa browning (Zhu et al., 2017). Furthermore, Liu et al. found that four WRKY genes were up-regulated in different postharvest and storage browning periods of Luffa, indicating their potential regulatory roles (Liu et al., 2017). In our study, 17 WRKY transcription factors have been identified among the DEGs. Based on the domain analysis, most of them had two WRKY domains which belong to the Group I subfamily (Table S4). The result suggested that more WRKY members may be involved in the regulation of browning. DEGs with high log2fold change including ‘Cluster-18244.0’, ‘Cluster-10926.1’ and ‘Cluster-18244.4’ need to be focused on. A large number of transcription factors were only expressed in one cultivar indicating their special roles. However, the functional identification of related transcription factor genes has not been carried out yet. Exploration of the functions of regulatory genes will be more helpful for Luffa breeding.
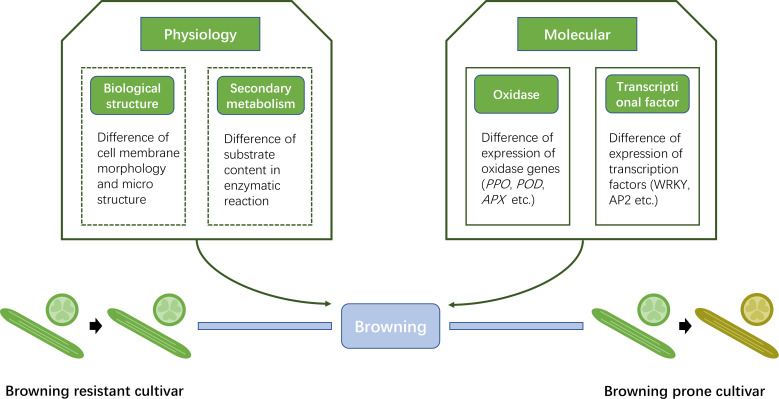
We summarized a diagram indicating the new insights into browning-resistant Luffa cultivar (Fig. 8). From the viewpoint of physiology, the cell membrane structure varies among different browning-resistant Luffa. Divergence may also exist in the content of enzymatic reaction substrate that lead to browning. These two findings need to be further confirmed. In the present study, we confirmed the molecular differences between the two different Luffa varieties. The expression of oxidase genes, including LcPPO, LcPOD and LcAPX significantly differed between the two varieties. Multiple transcription factors such as WRKY transcription factor family may also play regulatory roles. Our findings provide potential insights into the molecular regulation of Luffa browning and establish foundation to cultivate improved Luffa germplasm with high browning resistance.
Figure 8. Simplified diagram of differences between browning resistant and browning prone Luffa cultivars.
Conclusions
Our study compared the transcriptomes of two Luffa cylindrica cultivars with different resistances to browning by high-throughput sequencing. Among the differentially expression genes, 65 encoded polyphenol oxidase (PPO), peroxidase (POD) and ascorbate peroxidase (APX) were identified and further analyzed. The result of RT-qPCR confirmed the expression levels of 12 selected genes and demonstrated that these genes expressed higher in Luffa cultivar with strong browning resistance. Some transcription factors such as WRKY gene family may also participate in the regulation of Luffa browning. Our findings can establish foundation to cultivate improved Luffa germplasm with high browning resistance.
Supplemental Information
Figure S1: GO classification of the Luffa unigene library, Figure S2: GO classification of the DEGs identified in the unigene libraries between two Luffa cultivars, Figure S3: KEGG classification of the DEGs identified in the unigene libraries between two Luffa cultivars. Table S3. Comparison of unigenes identified in transcriptome during browning of ‘Fusi-3’ fruits and transcriptomes of two Luffa cultivars ‘2D-2’ and ‘35D-7’. Table S4. Information of WRKY transcription factors identified in the transcriptomes of Luffa ‘2D-2’ and ‘35D-7’.
Funding Statement
The research was supported by the grants from the Suzhou Agricultural Science and Technology Innovation Project (Research on the application of key agricultural technologies) (SNG2018061). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Additional Information and Declarations
Competing Interests
The authors declare there are no competing interests.
Author Contributions
Ya-Hui Wang conceived and designed the experiments, performed the experiments, analyzed the data, prepared figures and/or tables, and approved the final draft.
Xiao-Hong Liu and Zhi-Ming Yan performed the experiments, analyzed the data, authored or reviewed drafts of the paper, and approved the final draft.
Rong-Rong Zhang performed the experiments, prepared figures and/or tables, and approved the final draft.
Ai-Sheng Xiong analyzed the data, authored or reviewed drafts of the paper, and approved the final draft.
Xiao-Jun Su conceived and designed the experiments, analyzed the data, authored or reviewed drafts of the paper, and approved the final draft.
Data Availability
The following information was supplied regarding data availability:
The raw data for qRT-PCR are available in the Supplementary File.
The sequencing raw data are accessible in the Sequence Read Archive (SRA) database: SRP252794. Assembly sequences are available at TSA database: GILZ00000000.
References
- Ali et al. (2016).Ali HM, El-Gizawy AM, El-Bassiouny RE, Saleh MA. The role of various amino acids in enzymatic browning process in potato tubers, and identifying the browning products. Food Chemistry. 2016;192:879–885. doi: 10.1016/j.foodchem.2015.07.100. [DOI] [PubMed] [Google Scholar]
- Arogundade & Mu (2012).Arogundade LA, Mu TH. Influence of oxidative browning inhibitors and isolation techniques on sweet potato protein recovery and composition. Food Chemistry. 2012;134(3):1374–1384. doi: 10.1016/j.foodchem.2012.03.035. [DOI] [PubMed] [Google Scholar]
- Asma & Muhammad (2013).Asma S, Muhammad I. Loofa (Luffa Cylindrica) sponge: review of development of the biomatrix as a tool for biotechnological applications. Biotechnology Progress. 2013;29(3):273–600. doi: 10.1002/btpr.1702. [DOI] [PubMed] [Google Scholar]
- Bachem et al. (1994).Bachem CWB, Speckmann GJ, Vanderlinde PCG, Verheggen FTM, Hunt MD, Stefffens JC, Zabeau M. Antisense expression of polyphenol oxidase genes inhibits enzymatic browning in potato-tubers. Bio-Technology. 1994;12(11):1101–1105. doi: 10.1038/nbt1194-1101. [DOI] [Google Scholar]
- Boeckmann et al. (2003).Boeckmann B, Bairoch A, Apweiler R, Blatter MC, Estreicher A, Gaseiger E, Martin MJ, Michoud K, O’Donovan C, Phan I, Plibout S, Schneider M. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Research. 2003;31:365–370. doi: 10.1093/nar/gkg095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buera et al. (1987).Buera MD, Chirife J, Resnik SL, Lozano RD. Nonenzymatic browning in liquid model systems of high water activity - kinetics of color changes due to reaction between glucose and glycine peptides. Journal of Food Science. 1987;52(4):1068–1070. doi: 10.1111/j.1365-2621.1987.tb14277.x. [DOI] [Google Scholar]
- Chi et al. (2014).Chi M, Chagwat B, Lane WD, Tang G, Su Y, Sun R, Oomah BD, Wiersma PA, Xiang Y. Reduced polyphenol oxidase gene expression and enzymatic browning in potato (Solanum tuberosum L.) with artificial microRNAs. BMC Plant Biology. 2014;14:62. doi: 10.1186/1471-2229-14-62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chisari, Barbagallo & Spagna (2007).Chisari M, Barbagallo RN, Spagna G. Characterization of polyphenol oxidase and peroxidase and influence on browning of cold stored strawberry fruit. Journal of Agricultural & Food Chemistry. 2007;55(9):3469–3476. doi: 10.1021/jf063402. [DOI] [PubMed] [Google Scholar]
- Conesa et al. (2016).Conesa A, Madrigal P, Tarazona S, Gomez-Cabrero D, Cervera A, McPherson A, Szczesniak MW, Gaffney DJ, Elo LL, Zhang XG, Mortazavi A. A survey of best practices for RNA-seq data analysis. Genome Biology. 2016;17:1–19. doi: 10.1186/s13059-016-1047-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demeke & Morris (2002).Demeke T, Morris C. Molecular characterization of wheat polyphenol oxidase (PPO) Theoretical & Applied Genetics. 2002;104:813–818. doi: 10.1007/s00122-001-0847-3. [DOI] [PubMed] [Google Scholar]
- El-Gebali et al. (2019).El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, Qureshi M, Richardson LJ, Salazar GA, Smart A, Sonnhammer AM, Hirsh ELL, Paladin L, Piovesan L, Tosatto D, Finn SCE, Robert D. The Pfam protein families database in 2019. Nucleic Acids Research. 2019;47:D427–D432. doi: 10.1093/nar/gky995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman (1991).Friedman M. Prevention of adverse effects of food browning. Advances in Experimental Medicine and Biology. 1991;289:171–215. doi: 10.1007/978-1-4899-2626-5_15. [DOI] [PubMed] [Google Scholar]
- Galeazzi, Sgarbieri & Conatantinides (1981).Galeazzi MAM, Sgarbieri VC, Conatantinides SM. Isolation, purification and physicochemical characterization of polyphenoloxidases (PPO) from a dwarf variety of banana (Musa cavendishii, L) Journal of Food Science. 1981;46:150–155. doi: 10.1111/j.1365-2621.1981.tb14551. [DOI] [Google Scholar]
- Grabherr et al. (2013).Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I. Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nature Biotechnology. 2013;29:644–652. doi: 10.1038/nbt.1883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffiths (1959).Griffiths LA. Detection and identification of the polyphenoloxidase substrate of the banana. Nature. 1959;184:58–59. doi: 10.1038/184058a0. [DOI] [PubMed] [Google Scholar]
- Ioannou & Ghoul (2013).Ioannou I, Ghoul M. Prevention of enzymatic browning in fruits and vegetables. European Scientific Journal. 2013;9(30):311–341. doi: 10.1021/bk-1995-0600.ch004. [DOI] [Google Scholar]
- Jiang et al. (2016).Jiang GH, Kim YM, Nam SH, Yim SH, Eun JB. Enzymatic browning inhibition and antioxidant activity of pear juice from a new cultivar of asian pear (Pyrus pyrifolia Nakai cv. Sinhwa) with different concentrations of ascorbic acid. Food Science & Biotechnology. 2016;25:153–158. doi: 10.1007/s10068-016-0023-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang (2000).Jiang YM. Role of anthocyanins, polyphenol oxidase and phenols in lychee pericarp browning. Journal of the Science of Food and Agriculture. 2000;80(3):305–310. doi: 10.1002/1097-0010(200002)80:33.0.CO;2-H. [DOI] [Google Scholar]
- Jin et al. (2017).Jin J, Tian F, Yang DC, Meng YQ, Kong L, Luo J, Gao G. PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Research. 2017;45:D1040–D1045. doi: 10.1093/nar/gkw982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katagiri & Chua (1992).Katagiri F, Chua NH. Plant transcription factors—present knowledge and future challenges. Trends in Genetics. 1992;8:22–27. doi: 10.1016/0168-9525(92)90020-5. [DOI] [PubMed] [Google Scholar]
- Kulka (2008).Kulka RG. Hormonal control of root development on epiphyllous plantlets of Bryophyllum (Kalanchoe) marnierianum: role of auxin and ethylene. Journal of Experimental Botany. 2008;59(9):2361–2370. doi: 10.1093/jxb/ern106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee & Knag (2017).Lee SG, Knag HW. Comparison of adipocyte browning effects of white tea, oolong tea, green tea, and black tea water extracts using 3T3–L1 adipocyte. FASEB. 2017;31(1_supplement):lb408. doi: 10.1096/fasebj.31.1_supplement.lb408. [DOI] [Google Scholar]
- Li et al. (2018).Li H, Liu ZW, Wu ZJ, Wang YX, Teng RM, Zhuang J. Differentially expressed protein and gene analysis revealed the effects of temperature on changes in ascorbic acid metabolism in harvested tea leaves. Horticulture Research. 2018;5:65. doi: 10.1038/s41438-018-0070-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li et al. (2020).Li MY, Feng K, Hou XL, Jiang Q, Xu ZS, Wang GL, Liu JX, Wang F, Xiong AS. The genome sequence of celery (Apium graveolens L.), an important leaf vegetable crop rich in apigenin in the Apiaceae family. Horticulture Research. 2020;7:9. doi: 10.1038/s41438-019-0235-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li et al. (2016).Li QY, Lei S, Du KB, Li LZ, Pang XF, Wang ZC, Wei M, Fu S, Hu LM, Xu L. RNA-seq based transcriptomic analysis uncovers alpha-linolenic acid and jasmonic acid biosynthesis pathways respond to cold acclimation in Camellia japonica. Scientific Reports. 2016;6:36463. doi: 10.1038/srep36463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim, Cheun & Wong (2019).Lim WY, Cheun CF, Wong CW. Inhibition of enzymatic browning in sweet potato (Ipomoea batatas (L.)) with chemical and natural anti-browning agents. Journal of Food Processing and Preservation. 2019;43:e14195. doi: 10.1111/jfpp.14195. [DOI] [Google Scholar]
- Liu et al. (2017).Liu JT, Zhu HS, Wen QF, Wang B, Zhang QR, Chen MD, Lin H, Xue ZZ. Isolation and browning analysis of WRKY transcription factor gene of Luff (Luffa cylindrical) Journal of Agricultural Biotechnology. 2017;25:1950–1960. doi: 10.3969/j.issn.1674-7968.2017.12.005. [DOI] [Google Scholar]
- Liu et al. (2019).Liu X, Wang T, Lu Y, Yang Q, Li Y, Deng X, Liu Y, Du X, Qiao L, Zheng J. Effect of high oxygen pretreatment of whole tuber on anti-browning of fresh-cut potato slices during storage. Food Chemistry. 2019;301:125287. doi: 10.1016/j.foodchem.2019.125287. [DOI] [PubMed] [Google Scholar]
- Lopez-Nicolas et al. (2007).Lopez-Nicolas JM, Perez-Lopez AJ, Carbonell-Barrachina A, Garcia-Carmona F. Use of natural and modified cyclodextrins as inhibiting agents of peach juice enzymatic browning. Journal of Agricultural and Food Chemistry. 2007;55(13):5312–5319. doi: 10.1021/jf070499h. [DOI] [PubMed] [Google Scholar]
- Mayer (2006).Mayer AM. Polyphenol oxidases in plants and fungi: going places? A review. Phytochemistry. 2006;67(21):2318–2331. doi: 10.1016/j.phytochem.2006.08.006. [DOI] [PubMed] [Google Scholar]
- Mazali & Alves (2005).Mazali IO, Alves OL. Morphosynthesis: high fidelity inorganic replica of the fibrous network of loofa sponge (Luffa cylindrica) Anais da Academia Brassileira de Ciencias. 2005;77(1):25–31. doi: 10.1590/S0001-37652005000100003. [DOI] [PubMed] [Google Scholar]
- Meshi & Iwabuchi (1995).Meshi T, Iwabuchi M. Plant transcription factors. Plant Cell Physiology. 1995;36:1405–1420. doi: 10.1093/oxfordjournals.pcp.a078903. [DOI] [PubMed] [Google Scholar]
- Mi et al. (2019).Mi H, Muruganujan A, Ebert D, Huang X, Thomas PD. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Research. 2019;47:D419–D426. doi: 10.1093/nar/gky1038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Min, Liu & He (2012).Min MH, Liu K, He Q. The characterization of tea polyphenol oxidase(PPO) Food Industry. 2012;33(11):151–154. [Google Scholar]
- Minoru et al. (2007).Minoru K, Michihiro A, Susumu G, Masahiro H, Mika H, Masumi I, Toshiaki K, Shuichi K, Shujiro O, Toshiaki T. KEGG for linking genomes to life and the environment. Nucleic Acids Research. 2007;36(suppl_1):D480–D484. doi: 10.1093/nar/gkm882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mishra, Gautam & Sharma (2013).Mishra BB, Gautam S, Sharma A. Free phenolics and polyphenol oxidase (PPO): the factors affecting post-cut browning in eggplant (Solanum melongena) Food Chemistry. 2013;139(1-4):105–114. doi: 10.1016/j.foodchem.2013.01.074. [DOI] [PubMed] [Google Scholar]
- Natale et al. (2000).Natale DA, Shankavaram UT, Galperin M, Wolf YI, Arivind L, Koonin EV. Towards understanding the first genome sequence of a crenarchaeon by genome annotation using clusters of orthologous groups of proteins (COGs) Genome Biology. 2000;1(5):1–19. doi: 10.1186/gb-2000-1-5-research0009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicolas et al. (1994).Nicolas JJ, Richard-Forget FC, Goupy PM, Amiot MJ, Aubert SY. Enzymatic browning reactions in apple and apple products. Critical Review in Food Science and Nutrition. 1994;34:109–157. doi: 10.1080/10408399409527653. [DOI] [PubMed] [Google Scholar]
- Oboh & Aluyor (2009).Oboh IO, Aluyor EO. Luffa cylindrica - an emerging cash crop. African Journal of Agricultural Research. 2009;4(8):684–688. doi: 10.1021/jf9008839. [DOI] [Google Scholar]
- Okada, Katsura & Kobayashi (2004).Okada T, Katsura H, Kobayashi M. Relationship between browning color and antioxidant activity of soy sauce supplemented with wheat gluten. Seibutsu-Kogaku Kais. 2004;82(11):514–518. [Google Scholar]
- Parr & Bolwell (2000).Parr AJ, Bolwell GP. Phenols in the plant and in man. The potential for possible nutritional enhancement of the diet by modifying the phenols content or profile. Journal of the Science of Food and Agriculture. 2000;80(7):985–1012. doi: 10.1002/(SICI)1097-0010(20000515)80:7<985::AID-JSFA572>3.0.CO;2-7. [DOI] [Google Scholar]
- Pfaffl (2001).Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Research. 2001;29:e45. doi: 10.1093/nar/29.9.e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pruitt, Tatiana & Maglott (2005).Pruitt KD, Tatiana T, Maglott DR. NCBI Reference Sequence (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Research. 2005;33(suppl_1):D501–D504. doi: 10.1093/nar/gki025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purlis & Salvadori (2007).Purlis E, Salvadori VO. Bread browning kinetics during baking. Journal of Food Engineering. 2007;80(4):1107–1115. doi: 10.1016/j.jfoodeng.2006.09.007. [DOI] [Google Scholar]
- Ramachandran, Hiratsuka & Chua (1994).Ramachandran S, Hiratsuka K, Chua NH. Transcription factors in plant growth and development. Current Opinion in Genetics & Development. 1994;4(5):642–646. doi: 10.1016/0959-437X(94)90129. [DOI] [PubMed] [Google Scholar]
- Schwechheimer, Zourelidou & Bevan (1998).Schwechheimer C, Zourelidou M, Bevan MW. Plant transcription factor studies. Annual Review of Plant Physiolody and Plant Molecular Biology. 1998;49(1):127–150. doi: 10.1146/annurev.arplant.49.1.127. [DOI] [PubMed] [Google Scholar]
- Siegelman (1955).Siegelman HW. Detection and identification of polyphenoloxidase substrates in apple and pear skins. Archieves of Biochemistry & Biophysics. 1955;56(1):97–102. doi: 10.1016/0003-9861(55)90337-4. [DOI] [PubMed] [Google Scholar]
- Singh et al. (1998).Singh VK, Mangalam AK, Dwivedi S, Naik S. Primer premier: program for design of degenerate primers from a protein sequence. Biotechniques. 1998;24:318–319. doi: 10.2144/98242pf02. [DOI] [PubMed] [Google Scholar]
- Su et al. (2010).Su XJ, Xu H, Chen LZ, Song B, Yuan XH, Chen JF. Analysis on the phylogenetic relationship of luffa (Luffa cylindrical (L.) Roem) germplasm based on ISSR. Journal of Nanjing Agricultural University. 2010;33(3):42–46. doi: 10.1080/00949651003724790. [DOI] [Google Scholar]
- Sun et al. (2010).Sun J, You YL, Garcia E, Long X, Wang JB. Biochemical properties and potential endogenous substrates of polyphenoloxidase from chufa (Eleocharis tuberosa) corms. Food Chemistry. 2010;118(3):799–803. doi: 10.1016/j.foodchem.2009.05.065. [DOI] [Google Scholar]
- Troup et al. (2015).Troup GJ, Navarini L, Liverani FS, Drew SC. Stable radical content and anti-radical activity of roasted Arabica coffee: from in-tact bean to coffee brew. PLOS ONE. 2015;10:e0122834. doi: 10.1371/journal.pone.0122834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vamosvigyazo (1995).Vamosvigyazo L. Prevention of enzymatic browning in fruits and vegetables - a review of principles and practice. Acs Symposium Series. 1995;600:49–62. doi: 10.1021/bk-1995-0600.ch004. [DOI] [Google Scholar]
- Wang et al. (2015).Wang GL, Jia XL, Xu ZS, Wang F, Xiong AS. Sequencing, assembly, annotation, and gene expression: novel insights into the hormonal control of carrot root development revealed by a high-throughput transcriptome. Molecular Genetics & Genomics. 2015;290(4):1379–1391. doi: 10.1007/s00438-015-0999-5. [DOI] [PubMed] [Google Scholar]
- Wang et al. (2003).Wang Q, Wang D, Huang J, Guo A. Cloning of potato polyphenol oxidase gene and constructing of antisense PPO expression vector. Acta Botanica Boreali-occidentalia Sinica. 2003;23(10):1745–1749. doi: 10.1023/A:1022289509702. [DOI] [Google Scholar]
- Wang, Gertein & Snyder (2009).Wang Z, Gertein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics. 2009;10(1):57–63. doi: 10.1038/nrg2484.RNA-Seq. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu et al. (2014).Wu ZJ, Li XH, Liu ZW, Xu ZS, Zhuang J. De novo assembly and transcriptome characterization: novel insights into catechins biosynthesis in Camellia sinensis. BMC Plant Biology. 2014;14:277. doi: 10.1186/s12870-014-0277-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu et al. (2018).Xu YY, Liu Z, Lou LN, Su XJ. Identification of browning-related microRNAs and their targets reveals complex miRNA-mediated browning regulatory networks in Luffa cylindrica. Scientific Reports. 2018;8:16242. doi: 10.1038/s41598-018-33896-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zamora & Hidalgo (2005).Zamora R, Hidalgo FJ. Coordinate contribution of lipid oxidation and Maillard reaction to the nonenzymatic food browning. Critical Reviews in Food Science. 2005;45:49–59. doi: 10.1080/10408690590900117. [DOI] [PubMed] [Google Scholar]
- Zhu et al. (2018).Zhu H, Kang J, Liu J, Chen M, Yong L, Wang B, Lin B, Wen Q. Cloning and expression analysis of polyphenol oxidase PPO gene family from Luffa cylindrical. Journal of Nuclear Agricultural Sciences. 2018;32(8):1502–1512. [Google Scholar]
- Zhu et al. (2017).Zhu HS, Liu JT, Wen QF, Chen MD, Wang B, Zhang QR, Xue ZZ. De novo sequencing and analysis of the transcriptome during the browning of fresh-cut Luffa cylindrica ‘Fusi-3′fruits. PLOS ONE. 2017;12(11):e0187117. doi: 10.1371/journal.pone.0187117. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1: GO classification of the Luffa unigene library, Figure S2: GO classification of the DEGs identified in the unigene libraries between two Luffa cultivars, Figure S3: KEGG classification of the DEGs identified in the unigene libraries between two Luffa cultivars. Table S3. Comparison of unigenes identified in transcriptome during browning of ‘Fusi-3’ fruits and transcriptomes of two Luffa cultivars ‘2D-2’ and ‘35D-7’. Table S4. Information of WRKY transcription factors identified in the transcriptomes of Luffa ‘2D-2’ and ‘35D-7’.
Data Availability Statement
The following information was supplied regarding data availability:
The raw data for qRT-PCR are available in the Supplementary File.
The sequencing raw data are accessible in the Sequence Read Archive (SRA) database: SRP252794. Assembly sequences are available at TSA database: GILZ00000000.